Abstract
Non-ectomycorrhizal fungi that associate with typical ectomycorrhizae often remain hidden, and their localization inside ectomycorrhizal (ECM) roots has remained uncharacterized. In this study, the fungal community associated with the ectomycorrhizae of Castanopsis cuspidata was investigated using a culture-dependent isolation technique. Additionally, the species composition and localization were determined using molecular techniques. The results of the isolation and identification of fungal species revealed the predominance of a few species belonging to the order Helotiales. Furthermore, the fungal community structures were significantly different depending on the taxa of the ectomycorrhiza-forming fungi. A taxon-specific probe was developed to analyze the localization of one dominant Hyaloscyphaceae (Helotiales) species in ECM tissues by in situ hybridization. Hybridization signals were detected on the surface of the fungal mantle and around the ECM fungal cells within the mantle. Hyphal penetration into ECM hyphal cells of fungal mantles was also observed. Signals were not detected in the Hartig net or plant tissues inside the mantle in healthy ectomycorrhizae. These findings suggest that the analyzed species interact not only with host plant as root endophyte but also directly with the ECM fungi.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Within an ectomycorrhizal (ECM) community, non-ECM fungi have often been detected during field surveys (Kernaghan et al. 2003; Rosling et al. 2003; Urban et al. 2008; Leski et al. 2010). Such non-ECM fungi seem to ubiquitously colonize ECM roots as they have been frequently detected with molecular-based or culture-dependent approaches (Kernaghan and Patriquin 2011; Vohník et al. 2013). The non-ECM fungi commonly belong to the order Helotiales (phylum: Ascomycota), which represents the largest group in the class Leotiomycetes with 13 families and 395 genera (Wang et al. 2006). These species cover a broad range of niches and have been described as saprobes, plant pathogens, endophytes, and mycorrhizal fungi. Dark septate root endophytes, such as the Phialocephala fortinii s.l.–Acephala applanata species complex (Grünig et al. 2008) or the ericoid mycorrhizal Hymenoscyphus ericae aggregate (Vrålstad et al. 2000), have been extensively investigated as the major colonizers of the ectomycorrhiza-forming roots of diverse host plants, including conifers and angiosperms. However, in some cases, different types of ectomycorrhiza-associated fungi with non-melanized and unidentified hyphae are the predominant species (Kernaghan and Patriquin 2011, 2015). Members of the helotialean group have recently been identified as the dominant species in the roots of Fagaceae trees in the temperate and subtropical forests of Japan (Toju et al. 2013a, b, 2014). These results may illustrate the complex colonization patterns and variability of fungal root endophyte species in different hosts and environmental conditions.
Despite the increasing number of studies on ectomycorrhiza-associated helotialean species, their ecological role remains unclear. Considering ECM fungi are important for the growth and regeneration of host trees, ectomycorrhiza-associated fungi likely also affect forest ecosystems. Some ectomycorrhiza-associated helotialean species have been described as ECM fungi (Vrålstad et al. 2002a; Villarreal-Ruiz et al. 2004; Tedersoo et al. 2008; Wang et al. 2011; Huang et al. 2014), while others are regarded as putative endophytes, mycoparasites, plant pathogens, or entomopathogens (Tedersoo et al. 2009). The behaviors of ectomycorrhiza-associated fungi have been analyzed on ectomycorrhizae-forming trees under experimental conditions. The fungal isolates were often endophytic (Hashimoto and Hyakumachi 2001; Vrålstad et al. 2002a; Bergero et al. 2003; Grelet et al. 2009; Vohník et al. 2013). The endophytic fungi on ECM roots appear to coexist with ECM fungi. However, it is unclear how root-associated helotialean fungi interact with ECM fungi in the field. There is currently limited information regarding the localization of ectomycorrhiza-associated fungi in ECM roots because of the technical difficulties associated with targeting the fungi.
The objective of this study was to clarify the colonization patterns of helotialean fungi on ECM tissues in Castanopsis cuspidata-dominated forests. Castanopsis cuspidata (family: Fagaceae) is an evergreen tree and it is one of the major canopy trees in southwestern Japanese secondary forest (Tagawa 1995). To reveal the potential interactions between ectomycorrhiza-associated fungi and ECM fungi and/or plant hosts, we characterized a community of ectomycorrhiza-associated fungi by isolating and identifying species. We also visualized their localization patterns in ECM roots using a species-specific in situ hybridization technique with a newly developed probe.
Materials and methods
Sampling method
Rhizosphere soil was sampled from the 100-m2 study site in the Kodaiji-san Mountain National Forest (N 34° 59′ 49″, E 135° 47′ 8″; 150 m above sea level) in Kyoto, Japan. The experimental stands supported Castanopsis cuspidata as sole ectomycorrhizae-forming trees with ground vegetation consisting of Camellia japonica, Damnacanthus indicus, Pieris japonica, and C. cuspidata seedlings. The study site was divided into 25 4-m2 subplots. Approximately 100-ml soil cores (5 cm diameter × 5.1 cm depth), including the organic layer, were collected from the center of 12 subplots, which were alternately positioned at the sampling site. If a sample lacked or contained only a few ectomycorrhizae, we arbitrarily selected another point within the same subplot that contained a sufficient number of ectomycorrhizae. Samples for the fungal isolation and localization tests were collected in June 2013 and June to July 2017, respectively.
Fungal isolation from ectomycorrhizae samples
All soil samples were transported to the laboratory and stored at 4 °C until used. Fungi were isolated within 1 week of sampling. All ectomycorrhizae in soil cores were dissected and classified based on color, surface texture, and density of extraradical mycelium for each of individual soil cores. The ectomycorrhizae were transferred to Petri dishes for random selections. They were cleaned by removing debris and then stirring for 30 min in flasks with 100 ml deionized water containing three drops of Tween 80. Representative samples of each ECM morphotype were stored in 2× cetyltrimethylammonium bromide (CTAB) buffer [2% (w/v) CTAB, 1.4 M NaCl, 20 mM EDTA (pH 8.0), 100 mM Tris HCl (pH 8.0), and 1% (w/v) polyvinylpyrrolidone] at − 20 °C for the molecular identification of ectomycorrhiza-forming fungi. All cleaned ectomycorrhizae were surface-sterilized in 5% (w/v) Ca(ClO)2 for 2 min and then rinsed three times in sterilized deionized water. The ECM root tips were cut into two pieces, and one piece was aseptically placed onto Modified Norkran’s C medium (Yamada and Katsuya 1995) supplemented with 100 mg/l chloramphenicol and incubated at 25 °C for 1 month. The obtained isolates were used for molecular identifications. Fast-growing isolates were not identified as they were considered contaminants. However, morphotypes repeatedly isolated across several subplots were used for molecular identifications even if the isolates were fast-growing and potentially contaminants.
Fungal DNA extraction for molecular identifications
We extracted DNA from fungal cultures isolated from ectomycorrhizae as described by Izumitsu et al. (2012). A small amount of mycelia was collected from each isolated colony using a sterilized toothpick and suspended in 100 μl 10-fold diluted Tris-EDTA (TE) buffer in a microtube. Samples were microwaved (100 V, 600 W) twice for 1 min each and then centrifuged at 10,000 rpm for 5 min. The supernatants were used as templates for polymerase chain reaction (PCR) analyses.
A CTAB method was used to extract DNA from ECM root tips. Briefly, ECM root tips in 2× CTAB buffer were frozen in liquid nitrogen and heated three times at 65 °C in a block incubator. The root tips were crushed with a homogenizer pestle and then incubated at 65 °C for 30 min. Samples were purified with phenol/chloroform/isoamyl alcohol and chloroform/isoamyl alcohol solutions. The DNA was precipitated with isopropyl alcohol and rinsed with 70% ethanol. Pellets were dried with a CC-105 centrifugal concentrator (TOMY Digital Biology, Tokyo, Japan) and dissolved in 50 μl 10-fold diluted TE buffer. The extracts were stored at − 20 °C until used for molecular identifications.
PCR amplification and species identification by restriction fragment length polymorphisms and nucleotide sequencing
Representative morphotype species of ectomycorrhiza-forming fungi were identified based on the nucleotide sequences of the internal transcribed spacer (ITS) region. The PCR amplification was completed with the ITS1F (5′-CTTGGTCATTTAGAGGAAGTAA-3′) (Gardes and Bruns 1993) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) (White et al. 1990) primer pair and a T100 Thermal Cycler (Bio-Rad, Hercules, California, USA). The 10-μl PCR mix contained 1 μl 10× Blend Taq buffer, 0.4 μM each primer, 0.2 mM dNTP mixture, 0.25 U Blend Taq polymerase (TOYOBO, Osaka, Japan), and 1 μl template DNA. The PCR cycling conditions were as follows: 95 °C for 5 min; 35 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 1 min; 72 °C for 5 min.
Isolates obtained from ECM root tips were classified based on colony morphology and restriction fragment length polymorphism (RFLP) analysis of the ribosomal ITS sequence. The NSA3 (5′-AAACTCTGTCGTGCTGGGGATA-3′) and NLC2 (5′-GAGCTGCATTCCCAAACAACTC-3′) primer pair (Martin and Rygiewicz 2005) was used to PCR amplify DNA extracts from fungal cultures prior to the RFLP analysis. The PCR cycling conditions were as follows: 95 °C for 7 min; 35 cycles of 95 °C for 30 s, 60 °C for 40 s, and 72 °C for 40 s; 72 °C for 5 min. The PCR products were subsequently used for RFLP analyses involving the restriction endonucleases AluI and HinfI. Diluted PCR products were digested with each enzyme at 37 °C for 6 h. The PCR products and their digested fragments were separated by 1.0% agarose gel electrophoresis, stained with ethidium bromide, and visualized under UV light to analyze their banding patterns.
Representative ectomycorrhiza-forming fungi and one to nine samples from each RFLP type were analyzed by DNA sequencing. For ECM fungi identification, each of 1 to 12 ECM root tips was separately used for DNA extraction for each ECM morphotype. The PCR products were purified by gel electrophoresis and sequenced using the BigDye Terminator v3.1 Cycle Sequencing Kit (Thermo Fisher Scientific, Waltham, MA, USA) with the ITS1F and ITS4 primer pair and the 3130xl Genetic Analyzer (Thermo Fisher Scientific). To identify isolates, we searched for the most similar sequences using the BLAST online tool (http://www.ncbi.nlm.nih.gov/) (GenBank, NCBI). For the sequences from the ECM root tips, we searched the UNITE database (http://unite.ut.ee/). PCR products with different lengths, i.e., samples representing two or more bands were analyzed separately. Putative non-ECM fungi, e.g., Hyaloscyphaceae sp. 1 or Helotiales sp. 1 were excluded from the ECM species candidates. Also, samples that generated multiple PCR products which could not be separated were excluded from further analysis.
Statistical analysis
Data were analyzed using the R software. The fungal community structures (especially the proportions of Hyaloscyphaceae sp. 1 and Helotiales sp. 1) of the ectomycorrhizae of Cortinarius obtusus, Russula spp., and Lactarius sp. were obviously different. Thus, the proportions of Hyaloscyphaceae sp. 1 (i.e., the target species for in situ visualization) for all root tips used for fungal isolations were compared using a chi-squared test. A pairwise comparison with Bonferroni correction was then completed for each combination of the abovementioned three ECM groups.
Design of oligonucleotide probes
Because the isolation test results revealed the predominance of Hyaloscyphaceae sp. 1 at the study site, this species was targeted during the in situ visualization experiment. Nucleotide sequences of the D1/D2 region of the 26S rRNA region were determined using the NL1 (5′-GCATATCAATAAGCGGAGGAAAAG-3′) and NL4 (5′-GGTCCGTGTTTCAAGACGG-3′) primer pair (Vilela et al. 2005) as described above. The D1/D2 sequences from major putative-helotialean species (i.e., LC189052: Hyaloscyphaceae sp. 1, LC189055: Helotiales sp. 1, LC189053: Hyaloscyphaceae sp. 2, and LC189054: Helotiales sp. 2) were aligned using the ClustalW program. About 20 bases of a species-specific region were used to design the specific probe (i.e., G2-2: 5′-GTGCACCAGTGAGAACACCG-3′). The complementary sequence was used to design the negative control probe (i.e., non-G2-2: 5′-CGGTGTTCTCACTGGTGCAC-3′). The probes were synthesized by Fasmac (Kanagawa, Japan). The G2-2 probe was unable to discriminate between Hyaloscyphaceae sp. 1 and Hyaloscyphaceae sp. 2 because they were too closely related (i.e., 97.4% similarity in the ITS2 sequence), with no detectable sequence difference in the selected region. A fungal universal probe (i.e., R898: 5′-ATCCAAGAATTTCACCTCT-3′) (Tanaka 2009) targeting 18S rRNA was used as a positive control in the probe specificity test. Oligonucleotides were labeled with digoxigenin (DIG) using the DIG Oligonucleotide 3′-end Labeling Kit (Roche Diagnostics, Mannheim, Germany) according to the manufacturer’s protocol.
Probe specificity test and in situ hybridization
Fungal isolates were cultured in liquid Modified Norkran’s C medium for 2 weeks prior to the probe specificity test. In addition to Hyaloscyphaceae sp. 1 (LC189052), isolates at the experimental site (i.e., Helotiales sp. 1, LC189053 and Hyaloscyphaceae sp. 3 LC190974) and another five helotialean isolates which were isolated from surface-sterilized roots of Castanopsis cuspidata and Quercus spp. collected in Tottori and Kyoto prefecture in 2015 by the authors (i.e., Hyaloscyphaceae sp. 5: LC314065, Hyaloscyphaceae sp. 6: LC314066, Hyaloscyphaceae sp. 7: LC314067, Helotiales sp. 4: LC314068 and Lachnum sp.: LC314069) were used as reference samples. The in situ hybridization procedure was conducted as described by Tanaka et al. (2016) with a slight modification. The hybridization was conducted in a 1.5-ml microtube. The pretreatment reagents were treated with diethylpyrocarbonate and autoclaved. Approximately 0.5–1 mg (wet weight) of cultivated hyphae fixed in 4% PFA in PBS was deaerated for 10 min. The hyphae were then treated with 0.2 N HCl at room temperature for 20 min, and proteinase K (10 μg/ml) at 37 °C for 45 min. Samples were dehydrated with an ethanol series and dried using the CC-105 centrifugal concentrator. The tubes containing hyphae were placed in an HL-2000 HybriLinker hybridization chamber (UVP, Upland, CA, USA), and samples were saturated with hybridization reagent, which included hybridization buffer, DIG-labeled probe, and DNA, MB grade (Roche Diagnostics) as the carrier DNA. Samples were incubated at 45 °C for 16 h. The hybridization buffer contained the following components: 25% formamide, 4× SSC, 50 mM NaH2PO4/Na2HPO4 buffer (pH 7.0), 1 mM EDTA, and 5× Denhardt’s solution. After the hybridization, samples were washed twice for 5 min with 2× SSC containing 0.1% sodium dodecyl sulfate at room temperature. They were then washed twice for 45 min with 0.5× SSC containing 0.1% sodium dodecyl sulfate at 45 °C while being rotated. The samples were analyzed using the DIG Nucleic Acid Detection kit (Roche Diagnostics) according to the manufacturer’s protocol.
The permeability of cells for the probes may differ among fungal strains, which can affect the hybridization and staining results. Therefore, a dot blot hybridization and in silico similarity test was used to confirm the specificity of the probe for the Hyaloscyphaceae sp. 1 sequence. Fungal genomic DNA extracted using the CTAB method was dissolved in TE buffer. The DNA solution was heated at 100 °C for 5 min and cooled on ice. A 1-μl aliquot was blotted on the Biodyne PLUS 0.45 μm nylon membrane (PALL Corporation, Port Washington, NY, USA), which was then exposed to UV irradiation for cross-linking. The hybridization steps were completed as described above. The probe specificity was also tested using the BLAST online tool (http://www.ncbi.nlm.nih.gov/).
For in situ hybridizations, ectomycorrhizae sections were prepared again for five of the 12 subplots used for the isolation test. The ECM root tips were embedded in Tissue-Tek (Sakura Finetek Japan), frozen with liquefied carbon dioxide, and sliced into segments (25–30 μm thick) using the HM 400 R sliding microtome (Microm Laborgeräte, Walldorf, Germany) with a C35-type microtome blade (FEATHER Safety Razor, Osaka, Japan). Sliced sections were immediately fixed in 4% PFA in PBS. The pretreatment of sections, hybridization, and detection were conducted in a 1.5-ml tube as described above.
The frequency of hybridization signals was determined by dividing the number of ECM sections that produced signals by the total number of analyzed ECM sections. Ectomycorrhizae formed by the most dominant ECM species at the study site (i.e., C. obtusus) were identified according to morphological characteristics. We collected 5-mm tips from five ECM roots at each subplot. The hybridization procedure was conducted as described above. The number of the sections that produced hybridization signals was counted with a tally counter.
Results
Species composition of the ectomycorrhiza-associated fungal community
To determine the major fungal species associated with ECM roots, fungi were isolated from ECM root tips. We identified 359 of the 421 cultures isolated from 834 ECM root tips (Table 1). With 155 isolates, Hyaloscyphaceae sp. 1 was the most common fungus (i.e., > 36% of all isolates). Thus, this species was selected as the target for the in situ hybridization localization test. Additionally, 105 and 26 isolates corresponded to Helotiales sp. 1 and Hyaloscyphaceae sp. 2, respectively. Thus, about 68% of all isolates were one of these three Helotiales species. Furthermore, the sequences of 15, 10, and 8 isolates were similar to those of Nemania, Oidiodendron, and Cladophialophora species, respectively.
Varying ectomycorrhiza-associated fungal community with ectomycorrhiza-forming species
The ECM root tips of each subplot were classified into three to seven morphotypes based on color, surface texture, and density of extraradical mycelium. The 31 root tips that generated ITS1–5.8S–ITS2 fragments were used for sequencing analyses. We observed that the ECM root tips successfully analyzed were formed by the following 14 species (in descending order of root tip number): Cortinarius obtusus, Cortinarius sp., Russula sp. 1, Russula sp. 2, Russula sp. 3, Russula sp. 4, Lactarius sp., Thelephoraceae sp., Gymnascella sp., Boletellus sp., Boletaceae sp., Xerocomus sp., Coltriciella sp., and Tylopilus felleus. Morphological characters by which ectomycorrhizae was categorized and the number of ECM root tips successfully used for DNA identification (in parentheses) are shown for each ECM morphotype as follows. C. obtusus: white to brown, extraradical mycelium developed densely around ectomycorrhiza, irregularly branching, partly hydrophobic, (5). Cortinarius sp.: white, rhizomorphs abundant, irregularly branching, hydrophobic, (1). Russula spp.: white to brown, extraradical mycelium abundant or sparse, monopodial pinnate or no branching, (12). Lactarius sp.: light brown, no extraradical mycelium, no branching, smooth surface, (4). Thelephoraceae sp.: white, no extraradical mycelium, no branching, cystidia present, (2). Gymnascella sp.: dark brown, extraradical mycelium sparsely present, no branching, smooth surface, (1). Boletellus sp.: white, no extraradical mycelium, no branching, hydrophobic, (1). Boletaceae sp.: white, no extraradical mycelium, no branching, hydrophobic, (1). Xerocomus sp.: white, rhizomorphs present, no branching, hydrophobic, (1). Coltriciella sp.: dark brown, no extraradical mycelium, no branching, cystidia present, (1). Tylopilus felleus: brown, no extraradical mycelium, no branching, hydrophobic, (2). Although molecular analyses revealed clear differences among the four Russula species, it was difficult to differentiate between these species based on morphological features. Therefore, we treated the Russula species as one group. The ectomycorrhizae of Gymnascella sp., Boletellus sp., Boletaceae sp., Xerocomus sp., and Coltriciella sp. were not used for fungal isolations because of an insufficient number of samples. The ectomycorrhizae formed by C. obtusus yielded 124 fungal isolates, including 75 isolates of Hyaloscyphaceae sp. 1, six isolates of Helotiales sp. 1, 28 isolates of Hyaloscyphaceae sp. 2, and 28 isolates of other species. The ectomycorrhizae of Russula spp. yielded 158 isolates, including 36, 56, and 7 isolates of Hyaloscyphaceae sp. 1, Helotiales sp. 1, and Hyaloscyphaceae sp. 2, respectively. The ectomycorrhizae of Lactarius sp. yielded 105 isolates, including 40 Hyaloscyphaceae sp. 1 isolates, 36 Helotiales sp. 1 isolates, and a single Hyaloscyphaceae sp. 2 isolate. The ectomycorrhizae of the other species, including Thelephoraceae sp., Tylopilus felleus, and Cortinarius sp., yielded four isolates of Hyaloscyphaceae sp. 1, six isolates of Helotiales sp. 1, and two isolates of Hyaloscyphaceae sp. 2. Because most of the ectomycorrhizae were formed by three fungal groups (i.e., C. obtusus, Russula spp., and Lactarius sp.), we compared the corresponding ECM fungal communities (Fig. 1). The proportion of Hyaloscyphaceae sp. 1, which was the dominant species, was significantly different among the three ectomycorrhiza-forming species according to the chi-squared test (p < 0.01). Additionally, a pairwise comparison with Bonferroni correction revealed that the proportion of Hyaloscyphaceae sp. 1 in the C. obtusus ectomycorrhizae was significantly higher than that of the other two groups (p < 0.01). There was no significant difference in the proportion of Hyaloscyphaceae sp. 1 for the Russula spp., and Lactarius sp. ectomycorrhizae.
Species composition of ectomycorrhiza-associated fungal isolates obtained from each ECM taxon. The number of isolates of three major ectomycorrhiza-associated fungi (i.e., Hyaloscyphaceae sp. 1, Helotiales sp. 1, and Hyaloscyphaceae sp. 2) and of other species are presented for each of the following three dominant ECM taxa at the study site: Cortinarius obtusus, Russula spp., and Lactarius sp. Regarding Hyaloscyphaceae sp. 1, C. obtusus ectomycorrhizae yielded 75 isolates from 255 root tips, Russula spp. yielded 36 isolates from 278 root tips, and Lactarius sp. yielded 40 isolates from 229 root tips. The proportion of Hyaloscyphaceae sp. 1 was significantly different among the three ectomycorrhiza-forming species (chi-squared test, p < 0.01), and that of C. obtusus was significantly different from the other two groups (pairwise comparison with Bonferroni correction, p < 0.01)
Specificity test
To visualize fungal localizations in ectomycorrhizae by in situ hybridization, we designed a probe targeting the most frequent isolate, Hyaloscyphaceae sp. 1, and evaluated the viability of the probe. The specificity and suitability of the probe were confirmed using fungal hyphae and dot blots on a nylon membrane. However, Hyaloscyphaceae sp. 2 produced the same signaling pattern as Hyaloscyphaceae sp. 1 (data not shown) because they share the same nucleotide sequences in the putative targeted rDNA region (GenBank accession: LC189052 and LC189053). In the specificity test using cultured fungal hyphae, the Hyaloscyphaceae sp. 1-specific probe (i.e., G2-2 probe) generated hybridization signals for Hyaloscyphaceae sp. 1 (Fig. 2a) and Hyaloscyphaceae sp. 2 hyphae. In contrast, the negative control probe (i.e., non-G2-2 probe) did not generate clear signals (Fig. 2b). Neither probe generated a signal for the reference species (i.e., Helotiales sp. 1, Hyaloscyphaceae sp. 3, Hyaloscyphaceae sp. 5, Hyaloscyphaceae sp. 6, Hyaloscyphaceae sp. 7, Helotiales sp. 4 and Lachnum sp.), all of which were isolated from healthy fagaceous roots (Fig. 2a, b). The dot blots involving DNA extracts from all eight fungal species revealed that the G2-2 probe hybridized only to the Hyaloscyphaceae sp. 1 target DNA, and no signal was detected in tests with non-target DNA samples (Fig. 3). The probe sequence was searched in the GenBank using the BLAST. The probe showed 100% similarity with targeted species but also with some species of lichen, leaf endophytes, and marine and arctic soil-inhabiting fungi.
Probe specificity test with cultured fungal bodies. Eight fungal isolates (i.e., 1: Hyaloscyphaceae sp. 1, 2: Helotiales sp. 1, 3: Hyaloscyphaceae sp. 3, 4: Hyaloscyphaceae sp. 5, 5: Hyaloscyphaceae sp. 6, 6: Hyaloscyphaceae sp. 7, 7: Helotiales sp. 4, and 8: Lachnum sp.) were used to optimize pretreatment conditions and evaluate the signal specificity of the newly-designed G2-2 probe (a) and the negative control non-G2-2 probe (b). Cytoplasmic hybridization signals were detected only when Hyaloscyphaceae sp. 1 was tested with the G2-2 probe. Bars correspond to 10 μm
Probe specificity test by dot blot hybridization with fungal genomic DNA. Genomic DNA extracts of eight fungal species (i.e., 1: Hyaloscyphaceae sp. 1, 2: Helotiales sp. 1, 3: Hyaloscyphaceae sp. 3, 4: Hyaloscyphaceae sp. 5, 5: Hyaloscyphaceae sp. 6, 6: Hyaloscyphaceae sp. 7, 7: Helotiales sp. 4, and 8: Lachnum sp.) were tested with a positive control probe R898 (left) and the newly-designed G2-2 probe (right). The G2-2 probe generated a clear hybridization signal only with the target DNA derived from Hyaloscyphaceae sp. 1. The brightness and contrast of the image was adjusted to improve visibility
In situ hybridization
The DIG-labeled G2-2 probe hybridized to the target DNA and generated clear signals in field-collected ECM samples (Fig. 4). In contrast, no signals were detected for the non-G2-2 complementary negative control probe (data not shown). In ECM sections, hybridization signals were observed on the surface of fungal mantle (Fig. 4a). Also, hyphal penetration into the intercellular space (Fig. 4b) as well as penetration into ECM hyphal cells (Fig. 4c–e) of the mantle were observed. Signals were rarely observed in the inner layer of the mantle (Fig. 4g, h) and the Hartig net under the mantle (Fig. 4h). The frequency of hybridization signals was calculated by dividing the number of tissues with hybridization signals by the total number of ECM sections. Approximately 2% of the analyzed ECM sections (i.e., 19 of 945 sections) exhibited clear hybridization signals. Additionally, the proportion of sections with signals varied from 0 to 7.7% depending on the subplot. Of these sections, which originated from two of five subplots, approximately 15.8% (i.e., 3 of 19 sections) produced signals inside the ECM mantle. No signals were observed in the Hartig net or plant tissues.
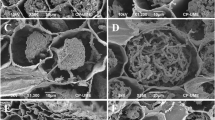
Optical micrographs of in situ hybridization of ECM sections with the taxon-specific G2-2 probe. All presented ECM sections are 25–30 μm thick. Signals indicating the presence of Hyaloscyphaceae sp. 1 or Hyaloscyphaceae sp. 2 appear as a blue–purple stain on the surface of ECM fungal mantle (a) and in the intercellular space of the mantle (b). Signals were also detected within mantle cells (c–e) but sometimes unclear whether signals were inter- or intracellular space of the mantle tissue (f). In rare cases, signals were observed around plant epidermal cells, inner mantle (g and h, arrow), and in the Hartig net (h, broken line arrow). Arrows and arrowheads indicate hybridization signals of hyphal cells and ectomycorrhizal fungal cell walls, respectively. a–c, f ECM mantle tissues. d, e, g, h Mantle tissues and the Hartig net around plant epidermal cells. The brownish parts in d, e, g, and h correspond to plant tissues. M, mantle; E, plant epidermal cell. Bars correspond to 10 μm
Discussion
At the Castanopsis-dominated study site, the ectomycorrhiza-associated fungal community was predominantly composed of a few helotialean species, including Hyaloscyphaceae sp. 1 and Helotiales sp. 1. These two species accounted for over 60% of the community. This suggests that we succeeded in selectively isolating the ectomycorrhiza-associated fungi from a complex soil fungal community that included free-living saprobes. However, the detected population may have been biased because of differences in individual culturabilities. The major isolated helotialean species could not be identified at the species or genus level using a DNA database because the closest matches were ericoid root colonizers or broad-leaf tree colonizers. Some studies on fungal root endophytes, most of which focused on boreal forests, revealed the dominance of the P. fortinii s.l.–A. applanata species complex or the Rhyzoscyphus ericae aggregate (Vrålstad et al. 2002b; Menkis et al. 2005; Kernaghan and Patriquin 2011). However, at our study site, there were relatively few darkly pigmented fungi, and the lineages mentioned above were not isolated. These findings suggest the fungal community status differs between our Castanopsis forest and a boreal, mainly coniferous forest, supposedly because some putative root endophytic species exhibit host plant specificity (Kernaghan and Patriquin 2011; Toju et al. 2013a, 2014; Yamamoto et al. 2014).
Bergero et al. (2000) isolated an Oidiodendron species from the ECM root tips of Quercus ilex and determined that it formed ericoid mycorrhizae under laboratory conditions. The genus Oidiodendron includes a group of ericoid mycorrhizal fungi. In our study, nine Oidiodendron spp. isolates were observed to inhabit the roots of C. cuspidata. The fact that we detected an ericoid mycorrhizal host (i.e., Pieris japonica) at our study site implies that the ericoid mycorrhizal fungi identified in ectomycorrhiza-forming host plants also potentially colonized ericaceous host plants. However, the possible saprotrophic activities of Oidiodendron spp. (Rice and Currah 2002) may have resulted in the ubiquitous presence of these species (Bergero et al. 2003).
The community structure of ectomycorrhiza-associated fungi depended on the fungal taxa that formed the ectomycorrhizae used for isolations (Fig. 1). Regarding the major species, Hyaloscyphaceae sp. 1 tended to occur at relatively high proportions in the ECM community of C. obtusus, but at relatively low proportions in the ECM communities of Russula spp. and Lactarius sp. This result is consistent with those of some studies that concluded that root-associated species were not randomly distributed, but tended to co-occur with certain ectomycorrhiza-forming species or an ECM morphotype (Urban et al. 2008; Tedersoo et al. 2009; Yamamoto et al. 2014). Our results imply that these two ecological types of fungi may interact with each other. Meanwhile, the reason for this biased occurrence should be carefully considered because the dense extraradical C. obtusus mycelium that formed the hyphal mats observed at our study site may have affected the microbial community in the hyphal mats via biochemical factors as described by Kluber et al. (2010, 2011). Environmental conditions surrounding ectomycorrhizae formed by C. obtusus may be considerably different from those surrounding the ectomycorrhizae of the other two groups because of enzymatic activities (Bödeker et al. 2009; Hobbie et al. 2013; Bödeker et al. 2014). This difference might be responsible for the diversity in the associated fungal community.
Cultured hyphae were used for the specificity test to examine the probe efficiency under conditions similar to those of the ensuing analysis. Moreover, considering the possibility that a pretreatment of cultured hyphae for in situ hybridizations may affect the apparent signal selectivity, we conducted dot blot hybridizations to confirm the specificity of the probe for the genomic DNA of the target isolates. Hyaloscyphaceae sp. 1 and Hyaloscyphaceae sp. 2 were specifically visualized during the specificity test with cultured hyphae. Additionally, the dot blot hybridization results confirmed the probe specificity. These observations suggest that the newly-designed G2-2 probe is suitable for practical use in the selective detection of Hyaloscyphaceae sp. 1 and/or Hyaloscyphaceae sp. 2 by hybridizing 26S rRNA, and not 26S rDNA. Unfortunately, Hyaloscyphaceae sp. 1 and Hyaloscyphaceae sp. 2 could not be distinguished by in situ hybridization because of their molecular similarity. However, these two species are close relatives that share habitats. Thus, they can be assumed to have the same or similar ecological status. Regarding this point, our results presented herein may be relevant to discussions of the ecology of a very limited spectrum of root-associated fungal species. Unfortunately, we could not eliminate the probability of cross-reactivity because the probe showed 100% similarity with some non-targeted fungal species in the database. However, none of these non-targeted species were reported as root colonizer and none was detected in the study site. Also, the majority of the isolates in the study site was Hyaloscyphaceae species. Therefore, we considered that the hybridization signals could be regarded as the targeted species, Hyaloscyphaceae sp. 1 and/or sp. 2.
Detectable in situ hybridization signals were concentrated mainly around the fungal mantle and partly observed in the mantle tissues (Fig. 4). This indicates that Hyaloscyphaceae sp. 1 and/or Hyaloscyphaceae sp. 2 colonized the mantle tissues and likely survived the surface sterilization process completed before the isolation test. There were a limited number of sections whose signals apparently penetrated into the mantle tissues (i.e., 15.8%; 3 of 19 sections with signals, Fig. 4b–h). The other signals appeared to be on the surface of the mantles (Fig. 4a) or it was difficult to distinguish whether they penetrated into the mantle tissues. Hyphal penetration into mantle tissues was relatively common among subplots (i.e., two of three subplots signals were detected) despite the rarity of signals in the sections (i.e., 2.0%; 19 of 945 sections). The proportion of hyphal penetration was not determined because the sections which clearly showed signal localization were very limited, and in many cases, it was difficult to recognize whether the signals were inter- or intrahyphal (Fig. 4f). Additionally, signals penetrated into the fungal mantle, but rarely reached the Hartig net (Fig. 4h) and absent in the plant tissue surrounded by the mantle. Vohník et al. (2013) reported that non-mycorrhizal mycelium occurred inside and around plant cells in senescent ectomycorrhizae, whereas it was usually absent in the healthy and young ectomycorrhizae of Norway spruce. This observation is consistent with our in situ hybridization results, which revealed that signals specific to Hyaloscyphaceae sp. 1 and Hyaloscyphaceae sp. 2 were often observed in the mantle tissues and that fungal colonizations of plant tissues did not occur in healthy-looking ectomycorrhizae.
The observed hyphal penetration into ECM hyphae may indicate possible direct interaction between these fungi within the ECM tissues. Also, their biased occurrence depending on the ECM taxa may suggest there is an ecological association between these Hyaloscyphaceae species and ECM fungi. So far, the physiological evidence for mycoparasitism in this group is lacking. However, the information of chitinolytic ability in helotialean root endophytes (Heinonsalo et al. 2016) and fungicolous tendency of some hyaloscyphaceous teleomorphs (Hosoya 2002; Hosoya 2013; Huhtinen et al. 2008) implies a potentially complex lifestyle including mycoparasitism and endophytism in these species. A certain dark septate endophyte parasitizes arbuscular mycorrhizal fungi in plant roots and penetrates into the arbuscular mycorrhizal hyphae (Mandyam and Jumpponen 2008). The phenomenon that we observed resembles their observation, during which fungal cells of other species are penetrated. Although mycoparasitism has been reported in ECM roots (Summerbell 1987; Pöder and Scheuer 1994; Olsson et al. 2000), it is difficult to distinguish mycoparasites from other modes of life. Additionally, there are some complex modes related to nutrient acquisition of saprophytic, mycoparasitic, and even nematophagous status (e.g., Rubner 1996; Komon-Zelazowska et al. 2007). In the present study, we can only speculate about the biological mechanisms regulating ectomycorrhiza-associated fungi because of their wide host range from plants to ECM fungi. Additionally, although we observed that the hyphae of ectomycorrhiza-associated fungi penetrate ECM fungal cells, we must emphasize that these species are usually isolated from surface-sterilized non-mycorrhizal roots as well (data not shown). In the present study, over 18% of ectomycorrhizae harbored Hyaloscyphaceae sp. 1. To clarify the reasons for the prevalence of this species at the research site, the ecological significance of this species on the ECM community will need to be characterized.
In conclusion, our findings revealed the prevalence of helotialean species in ectomycorrhizae formed on C. cuspidata roots as well as the potential of these species to interact with ECM fungi. In contrast to our understanding of their behavior as endophytes, the dominant Hyaloscyphaceae species at the study site colonized the inter- and intracellular spaces of the ECM fungal mantle tissues and hyphal penetration into ECM hyphal cells was observed. Although we focused only on a limited number of culturable species from the ectomycorrhiza-associated fungal community, our findings should help to characterize the ecological status of hyaloscyphaceous root-associated fungi and the complex network of root-associated fungal communities in forest soil ecosystems.
References
Bergero R, Perotto S, Girlanda M, Vidano G, Luppi AM (2000) Ericoid mycorrhizal fungi are common root associates of a Mediterranean ectomycorrhizal plant (Quercus ilex). Mol Ecol 9:1639–1649. https://doi.org/10.1046/j.1365-294x.2000.01059.x
Bergero R, Girlanda M, Bello F, Luppi A, Perotto S (2003) Soil persistence and biodiversity of ericoid mycorrhizal fungi in the absence of the host plant in a Mediterranean ecosystem. Mycorrhiza 13:69–75. https://doi.org/10.1007/s00572-002-0202-9
Bödeker IT, Nygren CM, Taylor AF, Olson Å, Lindahl BD (2009) Class II peroxidase-encoding genes are present in a phylogenetically wide range of ectomycorrhizal fungi. ISME J 3:1387–1395. https://doi.org/10.1038/ismej.2009.77
Bödeker I, Clemmensen KE, Boer W, Martin F, Olson Å, Lindahl BD (2014) Ectomycorrhizal Cortinarius species participate in enzymatic oxidation of humus in northern forest ecosystems. New Phytol 203:245–256. https://doi.org/10.1111/nph.12791
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes-application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
Grelet GA, Johnson D, Paterson E, Anderson IC, Alexander IJ (2009) Reciprocal carbon and nitrogen transfer between an ericaceous dwarf shrub and fungi isolated from Piceirhiza bicolorata ectomycorrhizas. New Phytol 182:359–366. https://doi.org/10.1111/j.1469-8137.2009.02813.x
Grünig CR, Duò A, Sieber TN, Holdenrieder O (2008) Assignment of species rank to six reproductively isolated cryptic species of the Phialocephala fortinii s.l.-Acephala applanata species complex. Mycologia 100:47–67. https://doi.org/10.3852/mycologia.100.1.47
Hashimoto Y, Hyakumachi M (2001) Effects of isolates of ectomycorrhizal fungi and endophytic Mycelium radicis atrovirens that were dominant in soil from disturbed sites on growth of Betula platyphylla var. japonica seedlings. Ecol Res 16:117–125. https://doi.org/10.1046/j.1440-1703.2001.00377.x
Heinonsalo J, Buée M, Vaario LM (2016) Root-endophytic fungi cause morphological and functional differences in Scots pine roots in contrast to ectomycorrhizal fungi. Botany 95:203–210. https://doi.org/10.1139/cjb-2016-0161
Hobbie EA, Ouimette AP, Schuur EA, Kierstead D, Trappe JM, Bendiksen K, Ohenoja E (2013) Radiocarbon evidence for the mining of organic nitrogen from soil by mycorrhizal fungi. Biogeochemistry 114:381–389. https://doi.org/10.1007/s10533-012-9779-z
Hosoya T (2002) Hyaloscyphaceae in Japan (6)**: the genus Hyphodiscus in Japan and its anamorph Catenulifera gen. nov. Mycoscience 43:47–57. https://doi.org/10.1007/s102670200008
Hosoya T (2013) Enumeration of remarkable Japanese discomycetes (3): first records of three inoperculate helotialean discomycetes in Japan. Bull Natl Mus Nat Sci 35:113–121
Huang J, Nara K, Zong K, Wang J, Xue S, Peng K, Shen Z, Lian C (2014) Ectomycorrhizal fungal communities associated with Masson pine (Pinus massoniana) and white oak (Quercus fabri) in a manganese mining region in Hunan Province, China. Fungal Ecol 9:1–10. https://doi.org/10.1016/j.funeco.2014.01.001
Huhtinen S, Hawksworth DL, Ihlen PG (2008) Observations on two glassy-haired lichenicolous discomycetes. Lichenologist 40:549–557. https://doi.org/10.1017/S002428290800827X
Izumitsu K, Hatoh K, Sumita T, Kitade Y, Morita A, Gafur A, Ohta A, Kawai M, Yamanaka T, Neda H, Ota Y, Tanaka C (2012) Rapid and simple preparation of mushroom DNA directly from colonies and fruiting bodies for PCR. Mycoscience 53:396–401. https://doi.org/10.1007/S10267-012-0182-3
Kernaghan G, Patriquin G (2011) Host associations between fungal root endophytes and boreal trees. Microb Ecol 62:460–473. https://doi.org/10.1007/s00248-011-9851-6
Kernaghan G, Patriquin G (2015) Diversity and host preference of fungi co-inhabiting Cenococcum mycorrhizae. Fungal Ecol 17:84–95. https://doi.org/10.1016/j.funeco.2015.05.001
Kernaghan G, Sigler L, Khasa D (2003) Mycorrhizal and root endophytic fungi of containerized Picea glauca seedlings assessed by rDNA sequence analysis. Microbial Ecol 45:128–136. https://doi.org/10.1007/s00248-002-1024-1
Kluber LA, Tinnesand KM, Caldwell BA, Dunham SM, Yarwood RR, Bottomley PJ, Myrold DD (2010) Ectomycorrhizal mats alter forest soil biogeochemistry. Soil Biol Biochem 42:1607–1613. https://doi.org/10.1016/j.soilbio.2010.06.001
Kluber LA, Smith JE, Myrold DD (2011) Distinctive fungal and bacterial communities are associated with mats formed by ectomycorrhizal fungi. Soil Biol Biochem 43:1042–1050. https://doi.org/10.1016/j.soilbio.2011.01.022
Komon-Zelazowska M, Bissett J, Zafari D, Hatvani L, Manczinger L, Woo S, Lorito M, Kredics L, Kubicek CP, Druzhinina IS (2007) Genetically closely related but phenotypically divergent Trichoderma species cause green mold disease in oyster mushroom farms worldwide. Appl Environ Microb 73:7415–7426. https://doi.org/10.1128/AEM.01059-07
Leski T, Pietras M, Rudawska M (2010) Ectomycorrhizal fungal communities of pedunculate and sessile oak seedlings from bare-root forest nurseries. Mycorrhiza 20:179–190. https://doi.org/10.1007/s00572-009-0278-6
Mandyam K, Jumpponen A (2008) Seasonal and temporal dynamics of arbuscular mycorrhizal and dark septate endophytic fungi in a tallgrass prairie ecosystem are minimally affected by nitrogen enrichment. Mycorrhiza 18:145–155. https://doi.org/10.1007/s00572-008-0165-6
Martin KJ, Rygiewicz PT (2005) Fungal-specific PCR primers developed for analysis of the ITS region of environmental DNA extracts. BMC Microbiol 5:28. https://doi.org/10.1186/1471-2180-5-28
Menkis A, Vasiliauskas R, Taylor AF, Stenlid J, Finlay R (2005) Fungal communities in mycorrhizal roots of conifer seedlings in forest nurseries under different cultivation systems, assessed by morphotyping, direct sequencing and mycelial isolation. Mycorrhiza 16:33–41. https://doi.org/10.1007/s00572-005-0011-z
Olsson PA, Münzenberger B, Mahmood S, Erland S (2000) Molecular and anatomical evidence for a three-way association between Pinus sylvestris and the ectomycorrhizal fungi Suillus bovinus and Gomphidius roseus. Mycol Res 104:1372–1378. https://doi.org/10.1017/S0953756200002823
Pöder R, Scheuer C (1994) Moserella radicicola gen. et sp. nov., a new hypogeous species of Leotiales on ectomycorrhizas of Picea abies. Mycol Res 98:1334–1338. https://doi.org/10.1016/S0953-7562(09)80307-9
Rice AV, Currah RS (2002) New perspectives on the niche and holomorph of the myxotrichoid hyphomycete, Oidiodendron maius. Mycol Res 106:1463–1467. https://doi.org/10.1017/S0953756202006767
Rosling A, Landeweert R, Lindahl BD, Larsson KH, Kuyper TW, Taylor AFS, Finlay RD (2003) Vertical distribution of ectomycorrhizal fungal taxa in a podzol soil profile. New Phytol 159:775–783. https://doi.org/10.1046/j.1469-8137.2003.00829.x
Rubner A (1996) Revision of predacious hyphomycetes in the Dactylella-Monacrosporium complex. Stud Mycol 39:1–134
Summerbell RC (1987) The inhibitory effect of Trichoderma species and other soil microfungi on formation of mycorrhiza by Laccaria bicolor in vitro. New Phytol 105:437–448. https://doi.org/10.1111/j.1469-8137.1987.tb00881.x
Tagawa H (1995) Distribution of lucidophyll oak-laurel forest formation in Asia and other areas. Tropics 5(1/2):1–40. https://doi.org/10.3759/tropics.5.1
Tanaka E (2009) Specific in situ visualization of the pathogenic endophytic fungus Aciculosporium take, the cause of witches’ broom in bamboo. Appl Environ Microb 75:4829–4834. https://doi.org/10.1128/AEM.00635-09
Tanaka E, Kumagawa T, Ito N, Nakanishi A, Ohta Y, Suzuki E, Adachi N, Hamada A, Ashizawa T, Ohara T, Tsuda M (2016) Colonization of the vegetative stage of rice plants by the false smut fungus Villosiclava virens, as revealed by a combination of species-specific detection methods. Plant Pathol. https://doi.org/10.1111/ppa.12540
Tedersoo L, Jairus T, Horton BM, Abarenkov K, Suvi T, Saar I, Kõljalg U (2008) Strong host preference of ectomycorrhizal fungi in a Tasmanian wet sclerophyll forest as revealed by DNA barcoding and taxon-specific primers. New Phytol 180:479–490. https://doi.org/10.1111/j.1469-8137.2008.02561.x
Tedersoo L, Pärtel K, Jairus T, Gates G, Põldmaa K, Tamm H (2009) Ascomycetes associated with ectomycorrhizas: molecular diversity and ecology with particular reference to the Helotiales. Environ Microbiol 11:3166–3178. https://doi.org/10.1111/j.1462-2920.2009.02020.x
Toju H, Yamamoto S, Sato H, Tanabe AS (2013a) Sharing of diverse mycorrhizal and root-endophytic fungi among plant species in an oak-dominated cool-temperate forest. PLoS One 8:e78248. https://doi.org/10.1371/journal.pone.0078248
Toju H, Yamamoto S, Sato H, Tanabe AS, Gilbert GS, Kadowaki K (2013b) Community composition of root-associated fungi in a Quercus-dominated temperate forest: “codominance” of mycorrhizal and root-endophytic fungi. Ecol Evol 3:1281–1293. https://doi.org/10.1002/ece3.546
Toju H, Sato H, Tanabe AS (2014) Diversity and spatial structure of belowground plant-fungal symbiosis in a mixed subtropical forest of ectomycorrhizal and arbuscular mycorrhizal plants. PLoS One 9:e86566. https://doi.org/10.1371/journal.pone.0086566
Urban A, Puschenreiter M, Strauss J, Gorfer M (2008) Diversity and structure of ectomycorrhizal and co-associated fungal communities in a serpentine soil. Mycorrhiza 18:339–354. https://doi.org/10.1007/s00572-008-0189-y
Vilela R, Mendoza L, Rosa PS, Belone AFF, Madeira S, Opromolla DVA, de Resende MA (2005) Molecular model for studying the uncultivated fungal pathogen Lacazia loboi. J Clin Microbiol 43(8):3657–3661. https://doi.org/10.1128/JCM.43.8.3657-3661.2005
Villarreal-Ruiz L, Anderson IC, Alexander IJ (2004) Interaction between an isolate from the Hymenoscyphus ericae aggregate and roots of Pinus and Vaccinium. New Phytol 164:183–192. https://doi.org/10.1111/j.1469-8137.2004.01167.x
Vohník M, Mrnka L, Lukešová T, Bruzone MC, Kohout P, Fehrer J (2013) The cultivable endophytic community of Norway spruce ectomycorrhizas from microhabitats lacking ericaceous hosts is dominated by ericoid mycorrhizal Meliniomyces variabilis. Fungal Ecol 6:281–292. https://doi.org/10.1016/j.funeco.2013.03.006
Vrålstad T, Fossheim T, Schumacher T (2000) Piceirhiza bicolorata—the ectomycorrhizal expression of the Hymenoscyphus ericae aggregate? New Phytol 145:549–563. https://doi.org/10.1046/j.1469-8137.2000.00605.x
Vrålstad T, Schumacher T, Taylor AF (2002a) Mycorrhizal synthesis between fungal strains of the Hymenoscyphus ericae aggregate and potential ectomycorrhizal and ericoid hosts. New Phytol 153:143–152. https://doi.org/10.1046/j.0028-646X.2001.00290.x
Vrålstad T, Myhre E, Schumacher T (2002b) Molecular diversity and phylogenetic affinities of symbiotic root-associated ascomycetes of the Helotiales in burnt and metal polluted habitats. New Phytol 155:131–148. https://doi.org/10.1046/j.1469-8137.2002.00444.x
Wang Z, Binder M, Schoch CL, Johnston PR, Spatafora JW, Hibbett DS (2006) Evolution of helotialean fungi (Leotiomycetes, Pezizomycotina): a nuclear rDNA phylogeny. Mol Phylogenet Evol 41:295–312. https://doi.org/10.1016/j.ympev.2006.05.031
Wang Q, Gao C, Guo LD (2011) Ectomycorrhizae associated with Castanopsis fargesii (Fagaceae) in a subtropical forest, China. Mycol Prog 10:323–332. https://doi.org/10.1007/s11557-010-0705-2
White TJ, Bruns T, Lee SJWT, Taylor JL (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR protocol 18(1):315–322
Yamada A, Katsuya K (1995) Mycorrhizal association of isolates from sporocarps and ectomycorrhizas with Pinus densiflora seedlings. Mycoscience 36:315–323. https://doi.org/10.1007/BF02268607
Yamamoto S, Sato H, Tanabe AS, Hidaka A, Kadowaki K, Toju H (2014) Spatial segregation and aggregation of ectomycorrhizal and root-endophytic fungi in the seedlings of two Quercus species. PLoS One 9:e96363. https://doi.org/10.1371/journal.pone.0096363
Acknowledgments
This work was financially supported by Japan Society for the Promotion of Science KAKENHI Grant Number 24380081.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Nakamura, N., Tanaka, E., Tanaka, C. et al. Localization of helotialean fungi on ectomycorrhizae of Castanopsis cuspidata visualized by in situ hybridization. Mycorrhiza 28, 17–28 (2018). https://doi.org/10.1007/s00572-017-0803-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00572-017-0803-y