Abstract
An endo-1,4-β-mannanase gene (manB) from a Bacillus pumilus Nsic-2 grown in a stinky tofu emulsion was cloned and expressed in Pichia pastoris GS115. After characterized, the endo-1,4-β-mannanase (manB) show maximum activity at pH 6.0 and 50 °C with LBG as substrate and perform high stability at a range of pH 6–8. After applying for a shake flask fermentation, the specific activity of manB reached 3462 U/mg. To produce mannose, the soybean meal (SBM) was pretreated by biological fermentation for 11 days with Penicillium brevicompactum, and then hydrolyzed by manB. As a result, mannose yield reached 3.58 g per 1 kg SBM which indicated that 0.358% SBM was converted into mannose after hydrolyzation, and mean a total 20% mannan of SBM converting into mannose, while the control group demonstrated only 1.78% conversion. An effective β-mannanase for the bioconversion of mannan-rich biomasses and an efficient method to produce mannose with soybean meal were introduced.
Graphic abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hemicelluloses comprise a family of linear and branched heteropolysaccharides, including hexoses (glucose, mannose, and galactose), pentoses (xylose and arabinose) and sugar acids [1]. β-Mannan-based polysaccharides present a backbone chain of β-1,4-linked d-mannopyranose residuals or a combination of glucose and mannose residuals which can be substituted with side chains of α-1,6-linked galactose residues [2]. Complete hydrolysis of mannan requires the cooperative action of several enzymes, in which endo-1,4-β-mannanase (EC 3.2.1.78) cleaves the β-1,4-glycosidic linkage of the mannan backbone [3]. β-Mannanase is the key enzyme that catalyzes the random hydrolysis of the β-1,4-d-mannopyranosyl linkages in mannans and heteromannans to release mannooligosaccharides [4]. Based upon the amino acid sequence alignment and hydrophobic cluster analysis, most β-mannanases could be classified to glycoside hydrolase (GH) families 5, 26, 113 and 134. Endo-1,4-β-mannanases play important roles in basic research, the bioconversion of biomass materials, and diverse potential industrial applications, such as bleaching of softwood pulps, scouring and de-sizing, food, feed additives, and oil drilling [5]. Endo-1,4-β-mannanases are used in combination with xylanases in the paper and pulp industries for increasing the brightness of pulps [6]. They are also widely applied in poultry feed to reduce the anti-nutritional factor of mannan polymers found in corn-soy based feeds [7, 8]. Furthermore, β-mannanases show potential applications in the recycling of copra and coffee wastes and the processing of instant coffee [9, 10]. D-mannose, as a kind of aldose, exists abundantly in nature in the form of various mannans but the monosaccharide was relatively rare. However, the D-mannose plays a significant role in human daily life, for example, carrier for delivering the anti-cancer drugs [11], anti-inflammatory effects [12] and anti-diabetic function [13]. D-mannose has displayed stupendous application prospects in food, cosmetics, and pharmaceutical industries. For food, as an important source of glyconutrient which recommended as a dietary supplement for human health [14]. Mannose significantly affected tumor cell growth via influence the cell metabolism which less lactate was produced from glucose [15]. And for pharmaceutical industries, D-mannose demonstrates a more significant role which acts as prophylaxis of urinary tract infections and treatment of phosphor-mannose isomerase deficiency disease [16]. Therefore, d-mannose shows significant economic and industrial applications.
Mining new strains and better enzymes for excellent industrial applications will always be important to researchers [17]. However, many of the recombinant mannanase characterization from various strains have been reported including Bacillus subtilis [18], Bacteroides ovatus [19], Lactobacillus casei [20], Pediococcus acidilactici [21] and no study about Bacillus pumilus Nsic-2 grown in a stinky tofu emulsion with enzymatic property and its application was reported. In recent years, the production of mannanases in recombinant Escherichia coli has been well studied [22]. For the commercial perspective protein production, researchers had explored so many expression systems such as mammalian cells [23], E. coli [24], and P. pastoris [25] expression system. Compared with mammalian cells, Pichia does not require a complex growth medium or culture conditions, and which is genetically relatively easy to manipulate, and Pichia has a eukaryotic protein synthesis pathway [26]. At the same time, the major disadvantage of E. coli is the formation of insoluble aggregates [27]. On the contrary, the potential for secretory and soluble expression of heterologous proteins in P. pastoris provided the system a candidate for the expression of various proteins [28].
Soybean meal (SBM), as important agro-industrial biomass, is the major by-product of the soya bean oil industry. SBM, containing a lot of non-starch polysaccharides (NSP), like β-mannan, would damage to the poultry industry concerning for nutritionists due to presence of anti-nutritive properties [29]. Β-Mannan content of dehulled SBM (90% dry matter) cultivation in different region ranged from 1.02 to 1.50% and ranged from 1.33 to 2.12% for non-dehulled samples (90% dry matter) [29]. A recent study demonstrated that about 220 million tons of SBM were produced in 2017 [30]. The purposes of pretreatment were reducing the particle size of the bioresources, the protection of the hemicellulose fractions, like pentose, and the degradation of inhibitors affected the enzymatic reaction or microbe growth [31]. Biological abatement of cellulase inhibitors and an alternative in second-generation ethanol from wheat straw had achieved great success via fungal pretreatment [32]. Effects of fungal pretreatment influencing subsequent degradation of biological resources showed that pretreatment for 7 days with P. tigrinus demonstrated 50% digestibility toward hemicellulose and cellulose, while the control group displayed barely 30% digestibility [33]. In contrast to physical and chemical pretreatment, the biological method presented additional advantages as being lower cost, security, energy- saving and more environmentally friendly [32]. However, little work had been reported about the biological pretreatment of soybean meal with fungus to produce mannose via mannanase from pretreatment SBM.
Given the many disadvantages of free enzymes (instability, high cost, and recoverability), the utilization of immobilized enzymes is getting more widespread [34]. Immobilization increases the optimum temperature of the enzyme and improves thermal stability [35].
In this study, a novel alkaline GH26 family β-mannanase, named manB, was cloned and expressed in both E. coli and P. pastoris. Immobilization raise the thermostability and optimum pH of the manB, and make manB more suitable for commercial application. The pretreatment method toward SBM was selected from a variety of programs and the P. brevicompactum was employed to carry out the bio-pretreatment mission for its slower growth rate and more efficient disruption the structure of lignocellulose. The operating time of enzymatic hydrolysis and fungus pretreatment were optimized to achieve the high yield of mannose with the smashed SBM. The first work on soybean meal degraded by β-mannanase to produce mannose.
Materials and methods
Bacterial strains, plasmids, gene sequence and chemicals
The heterologous protein expression system of P. pastoris GS115 with pPIC9K was purchased from Invitrogen. The fungus strain P. brevicompactum was preserved in our lab. The gene sequence of the manB was shown in the supplement file. Isopropyl-β-d-thiogalactopyranoside (IPTG), ampicillin and geneticin G418 were purchased from Amresco (Shanghai Genebase Co., Ltd, China). DNA Mini kit and Plasmid Mini Prepare kit were purchased from Axygen Biosciences (Union City, CA, USA). The epoxy resin for immobilization was got from Xi-an Lanxiao Technology New Material Co., Ltd (Xi-an, China). The substrate Locust bean gum (LBG) was purchased from Shanghai Fu-sheng Industrial Co., Ltd. Soybean meal (SBM) (particle size 0.1–0.25 mm, non-dehulled) was got from Bang-JI Zheng-da Co., Ltd (Tianjin). And the BSA standard solution was bought from TIANGEN (TIANGEN Biotech, China). All other chemicals and reagents were of analytical grade.
Gene analysis and molecular structure modeling of manB
The nucleotide sequence and predicted amino acid sequence were analyzed using the BLAST program (NCBI). β-mannanase catalysis domain analysis was performed by the SMART domain annotation server (https://smart.embl-heidelberg.de/). The 3D model of manB was predicted via the SWISS-MODEL server according to the crystal structure of a β-mannanase (PDB ID: 3CBW) from B. subtilis (strain 168), and the models were visualized and analyzed using PDB Viewer, and the figures were constructed using Pymol. The distance tree of BLAST results was obtained with the server (https://blast.ncbi.nlm.nih.gov/Blast.cgi).
Expression of manB in P. pastoris
With the primers in the Table 1, the gene fragment was obtained by PCR. Subsequently, the obtained gene fragment was linked to the digested vector, pPIC9K, to construct the recombinant plasmid, pPIC9K-manB. The recombinant plasmid was then linearized by Sal I and transformed into P. pastoris GS115 competent cells according to the manufacturer’s instructions (Invitrogen). Recombinant GS115 clones were selected on MD plates to confirm that they were positive concerning methanol-utilizing phenotype (Mut+). After three days of incubation, transformants grown on MD plates were sacked with sterile water and the sacked transformants were then plated onto YPD solid plates containing different concentrations of G418 (1 mg/mL, 2 mg/mL, 3 mg/mL, and 4 mg/mL) [36].
After 72 h of incubation at 28 °C, selection of transformants grown on YPD solid plate with the highest concentration of G418. Eight clones were identified by 2 × Phanta Max Master Mix (Vazyme China) using the AOX1 primers, which could detect if the integration of the target gene into the genome of P. pastoris. The promising clones were then cultivated in BMGY (1% w/v yeast extract, 2% w/v peptone, 1.34% w/v yeast nitrogen base, 4 × 10–5 w/v biotin, 1% v/v glycerol, 100 mM potassium phosphate, pH 6.0) to make the OD600 reach 2–8 and induction in BMMY (1% w/v yeast extract, 2% w/v peptone, 1.34% w/v yeast nitrogen base, 4 × 10–5 w/v biotin, 1% v/v methanol, 100 mM potassium phosphate, pH 6.0) for 5 days. SDS–PAGE was then applied to detect which clone had the highest manB expression level. The sample stored was dispersed in sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) buffer and heated to 100 °C for minutes. Finally, recombinant proteins were separated by SDS–PAGE (12.5% acrylamide/bis-acrylamide) and stained with Coomassie bright blue [37] (see Table 2).
Enzyme assay and protein content
Enzyme activities of manB were measured by the 3,5-dinitrosalicylic acid (DNS) method using mannose as the standard. The reaction mixture containing 50 μL diluted enzyme, 450 μL of 5 g/L locust bean gum (LBG) in 0.05 M sodium phosphate buffer (pH 6.0) was incubated at 40 °C for 10 min, and terminated by the addition of 1 mL DNS reagent [38].
The reaction was then boiled in the water bath for 10 min, cooled to room temperature, and the absorbance was measured at 540 nm. 1 U was defined as the amount of enzyme liberating 1 μM of reducing sugar per minute under the assay conditions. Lowry’s method was employed for protein content measurement with bovine serum albumin as the standard [39].
Biochemical characterization
The optimal pH of the enzyme was determined in 50 mM different buffers with pH ranging from 3.0 to 11.0 at 40 °C, including glycine–HCl (pH 2.0–3.0), citric acid-sodium citrate (pH 4.0–5.0), sodium phosphate (pH 6.0–7.0), Tris–HCl (pH 8.0), glycine–NaOH (pH 9.0–10.6). The pH stability of β-mannanase was evaluated by assaying its residual activity in different pH buffers for 30 min at 50 °C. The optimal temperature was assessed at the temperature range of 30–70 °C in 50 mmol/L sodium phosphate (pH 6.0). The thermostability of manB was studied by determining the residual activity after incubation at temperatures for 30 min.
Effect of metal ions and other reagents on enzyme activity were determined at 50 °C in a standard assay medium containing each reagent at various concentrations. The system without any metal ions or other reagents was used as a control. The surfactant and organic solvents concentration of isopropanol, Tween-20, Tween-80, TritonX-100, methanol and DMSO were at 1% (v/v) [40, 41].
Immobilization of the recombinant β-mannanase on LX-107S epoxy resin and enzymatic properties assay
The method of immobilization was based on the study of Li et al. [42]. Enzyme immobilization was carried out under the condition which the ion concentration was from 100 mM to 2 M (100 mM, 1 M, and 2 M) and the immobilization duration was from 8 to 32 h (8 h, 20 h, and 32 h). After the epoxy resin was activated under the corresponding ion concentration, it was collected and dried. And then, an appropriate amount of epoxy resin was placed in a 50 mL Erlenmeyer flask, and a certain amount of recombinant β-mannanase solution was packed at 200 rpm and 20 °C for 2–32 h [43]. After fixation, the suspension was filtered and the resin was washed until no enzyme activity was detected in the eluate. Finally, the immobilized recombinant β-mannanase was vacuum-dried for hours and stored at 4 °C until enzymatic properties analysis [44]. Then the enzymatic properties were analyzed with the method mentioned above.
Soybean meal pretreated by Penicillium brevicompactum and Penicillium oxalicum fermentation
The enzyme load and SBM concentration were modified slightly according to the method mentioned by Hafiza Shukor [45]. All pretreatments of SBM were reacted in the 0.5 L glass reactor. The SBM was firstly smashed by grinder into small pieces in which the diameter of the treated SBM was about 0.5 μm. Accurately weighted (16 g SBM) smashed SBM and measured 200 mL distilled water was introduced to the reactor. About 107 spores per 100 mL reaction solution were inoculated into the reactor and then the reactor was maintained in the growth conditions that the temperature was 30 °C and the rotating speed was 220 rpm.
For the P. brevicompactum, sampling on the 3rd, 5th, 7th, 9th, 11th, and 13th day of the reaction and put the samples in the − 20 °C until enzymatic hydrolysis. 13 days later, melting the samples at room temperature and then putting the melted samples in the boiling water for 10 min to inactivate the fungus.
For the P. oxalicum, sampling on the 1st, 2nd, and 3rd day of the fermentation and put the samples in the − 20 °C until enzymatic hydrolysis.
The samples were analyzed by DNS method [46] to detect the reducing sugar in the fermentation supernatant.
Enzymatic hydrolysis
Efficiency of P. brevicompactum fermentation pretreatment was evaluated by subsequent enzymatic hydrolysis of which the condition was chosen by independent optimization. Adding the free enzyme which was fermented by the P. pastoris GS115 to the untreated and pretreated soybean meal solution at temperature of 50 °C without shaking for 12 h, 24 h, and 36 h. Each milliliter pretreated SBM solution was hydrolyzed with the dosage of 50 U fermented enzymes. Finally, the mixture was boiled for 10 min to inactivate the reaction and then stored at − 20 °C for analysis.
High-performance liquid chromatography (HPLC) analysis
To detect the content of the mannose in hydrolysis, enzymatic hydrolysates of SBM were analyzed by HPLC with the refractive index detector (RID; Agilent 1260). Mannose was separated on a sugar-PakTM I column (6.5 × 300 mm, Waters) with pure water as a mobile phase at a flow rate of 0.6 mL/min. The temperature of the column and detector was 75 °C and 50 °C, respectively. Products were calibrated with the mannose as standard [47].
The yield of mannose was calculated using the below equation:
The HPLC results were compared with those of mannose standard. The peak area ratios of product peaks were used as the basis for calculation. Each sample were triple.
Results
Gene analysis and molecular structure modeling of manB
Strain Nsic-2 was identified as Bacillus pumilus by 16S rDNA phylogenetic analysis and the morphology. The GeneBank accession numbers for the Bacillus pumilus Nsic-2 16S rDNA gene is KC568200. Also, strain Nsic-2 was deposited at the China Center for Type Culture collection (https://www.cctcc.org/, Wuhan, China. Accession number: CCTCC AB 2013050).
Through the alignment analysis of proteins, we obtained information about the protein sequence alignment of manB and other mannanase from NCBI. (Fig. 1a). The protein structure was viewed in Pymol Viewer (Fig. 1b). The catalytic triad (Glu297 and Glu198) was observed in the regions and shown as a stick in grey and marked in white in Fig. 1c.
Gene analysis and molecular structure modeling of manB. a Multiple sequence alignment between manB and other closely related enzymes, the analysis was accomplished with the method of NCBI. b 3D model of manB was predicted by the SWISS-MODEL. The structure of this manB could be divide to α-helix, β-sheet, random coil, and the α-helix was wrapped around the periphery with the same time the β-sheet constructed a barrel in the middle of the enzyme structure. c The catalytic triads (Glu297 and Glu198) are shown as stick
From the distance tree, evolutionarily, the closest relationship to manB was the family GH26 mannanase from B. mojavensis (Fig. 2a) and manB which was expressed in E. coli was shown as SDS-PAGE in the Fig. 2b.
The map of distance of the tree and the figure of the manB expressed in the E. coli. a The distance tree between the manB and the other mannanase from different strain. manB (this study, accession number: AGI16498.1), B. mojavensis GH26 mannanase (accession number: WP_010333200.1), B. subtilis endo-1,4-beta-mannosidase (accession number: WP_003237220.1), B. swezeyi GH26 mannanase (accession number: WP_076797482.1), B. altitudinis GH26 mannanase (accession number: WP_073414040.1), B. circulans beta-mannanase (accession number: AEO19987.1), B. velezensis mannan endo-1,4-beta-mannosidase (accession number: WP_015240845.1), Paenibacillus polymyxa Mannan endo-1,4-beta-mannosidase (accession number: VUG07555.1), Clostridium mannanase (accession number: WP_010963860.1). b The SDS-PAGE of the manB which was expressed in the E. coli. The line 1 was the buffer to dilute the cell, the line 2 was the supernatant of the induced recombinant cell, and the line 3 was the induced cell with the empty vector
High level expression and purification of the manB in P. pastoris
The original β-mannanase (manB) was successfully expressed in P. pastoris GS115. Transformant which had the highest activity growth on the G-418 YPD plate was selected and the enzyme activity reached to 502 U/mL. At the same time, the protein content of the fermentation supernatant reached 0.145 mg/mL, and the specific activity of manB was 3462 U/mg.
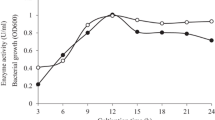
The relationship of induction days and enzymatic activity of manB which express at P. pastoris GS115 (Fig. 3a). In the first five fermentation days, the enzyme activity increased follow the fermentation time. The component of the fermentation supernatant during the induced process was analyzed by SDS-PAGE (Fig. 3b). A protein band around 41 kDa was demonstrated which was matched to the calculated molecular mass of manB (41 kDa). Clearly, the concentration of the manB in the fermentation supernatant was increased with fermentation time significantly. Addition of the protein content in the fermentation supernatant indicated an increase of the unit enzyme activity. For commercial applications, the addition of unit enzyme activity significantly reducing the cost of production.
Induced expression of manB in P. pastoris GS115. a The analysis of induction days and enzymatic activity of manB which express at P. pastoris GS115. b The SDS-PAGE analysis of induction days and expression of manB which express in P. pastoris GS115. During the first 5 days of fermentation, the expression of manB in the fermentation broth increased gradually with time. On the sixth day of fermentation, both expression of manB and unit enzyme activity began to decrease. The main reason was the degradation of enzyme which caused the decrease of the quantity and unit activity of the manB
Enzymatic properties of the purified manB
The optimal temperature of purified manB was 50 °C (Fig. 4a). The purified manB exhibited high enzyme activities over a pH range of 6.0–8.0, among which the highest optimum enzyme activity was at pH 6.0 (Fig. 4b). The enzyme retained more than 80% of the maximal activity in the pH range of 6.0 to 8.0 (Fig. 4b). It retained more than 90% of the initial activity after incubation at 50 °C for 1 h without substrate and was nearly completely inactivated at 70 °C for 150 min (Fig. 4c). And the pH stability of the manB was shown in the Fig. 4d.
Characterization of manB. a Optimum temperature of manB activity measured at pH 6.0. b Optimum pH of manB activity assayed at 50 °C in buffers ranging from pH 2.0 to 11.0. c Thermostability of manB. d pH stability of manB. After incubation at 50 °C for 30 min in buffers ranging from pH 4.0 to 10.0. The enzyme was preincubated at 30 °C, 40 °C, 50 °C, 60 °C, or 70 °C at pH 6.0, and aliquots were removed at specific time points to measure the residual activity
The effects of different metal ions and chemical reagents on the enzyme were examined by incubating 200 μL of enzyme solutions with each of the metal ions and chemicals reagents at various concentrations in pH 7.0 phosphate buffer at 50 °C for 10 min. At low concentration (1 mM), Mg2+, K+, Ni2+, and Co2+ partly inhibited or had no effects on the enzyme activity, while Zn2+ could slightly enhance the enzyme activity. However, at high concentration (5 mM and 10 mM), Mg2+ showed slightly inhibited the enzyme activity. Ca2+ and Mn2+ could be significantly or slightly enhance the 4enzyme activity at the concentration of 1 mM, 5 mM and, 10 mM. Cu2+ and Ni2+ at a concentration of 10 mM inhibited the enzyme activity severely. The manB activities were not significantly affected by isopropanol, Tween-80, TritonX-100 and completely inactivated by SDS. The reducing agent DTT enhanced the enzyme activity to 155%. Methanol and dimethyl sulfoxide (DMSO) could strongly inhibit the enzyme activity, and purified recombinant manB was significantly resistant to all tested neutral proteolytic enzymes.
Properties of immobilized enzyme
The purified manB was directly immobilized on epoxy resin lx-107 s. The effects of mannanase fixation time and buffer ionic strength on relative enzyme activity were investigated. The optimum temperature had increased from 40 to 50 °C (Fig. 5a). As the same time, the optimum pH had surged to pH 8.0 which increased by one third (Fig. 5b). And the reusability of the fixed manB was shown in the Fig. 5c. The optimal time for manB fixation is 24 h (Fig. 5d). A buffer with a higher ionic strength helps to enhance the covalent attachment of the enzyme. When the ionic strength of the buffer is higher, the mannanase conformation is easy to move and is more easily combined with the epoxy resin.
Optimization of manB immobilization and performance of lx-manB. a Effect of temperature on manB and lx-manB activity measured at pH 7.0. b Effect of pH on manB and lx-manB activity assayed at 50 °C in buffers ranging from pH 2.0 to 11.0. c Reusability of immobilized manB. d Effect of immobilization buffer ionic concentration and immobilization time on immobilization. The top X axis is the immobilization buffer ionic strength (mol/L) and the bottom axis is the immobilization time (h). In figure (a), it seems that higher temperature is needed to activate the activity of immobilized enzyme, because the activity of immobilized enzyme at 40 °C is only half of that at 50 °C
Optimization of mannose production from SBM
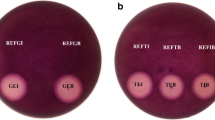
The same enzymatic hydrolysis time, pretreatment broth at different times as a substrate for enzymatic hydrolysis, more products were obtained with increasing fermentation time (Fig. 6a). However, with the fermentation broth on 13th day as the substrate, the product obtained was significantly less than the enzymatic hydrolysis result of the last sample. The yield with same manB hydrolysis on the 11th day fermentation was almost twice that on the 5th. Following our work, 24 h hydrolysis of the pre-fermentation SBM at 50 °C would get a max yield which was no significant change when the hydrolysis time increased to 36 h (Fig. 6b). That indicated the mannan and the manno-oligosaccharide in the pretreatment SBM were completely hydrolyzed by manB at the 24th hour. At the same time, product content of the un-treatment one showed no max yield with the hydrolysis of 36 h indicating that manB would have a better result towards the treated one. From the above figures, it can be found that when the fermentation time is 11 days and the hydrolysis time is 24 h, the max products are obtained. As the enzymatic hydrolysis time increased, the amount of the product increased significantly. Under the condition of enzymatic hydrolysis for 24 h, more mannose could be obtained with the increase of pre-fermentation time (Fig. 6c).
Optimization of mannose production from SBM. a The effect of fermentation time. b The effect of enzymatic hydrolysis time. c Mannose yield tendency of soybean meal pre-treated by P. brevicompactum after enzymatic hydrolysis. d Mannose yield tendency of soybean meal pre-treated by P. oxalicum after enzymatic hydrolysis. For the rapid growth of P. oxalicum and the expression of mannanase, little mannose could be obtained from the pre-fermentation soybean meal by manB. On the contrary, due to the slow growth of the P. brevicompactum, the soybean meal pre-fermentation by P. brevicompactum had higher mannose yield after being hydrolyzed by manB
In addition, pre-fermentation with the P. oxalicum, the maximum yield was reached at 24th hours, but it was still only a third of the maximum yield of P. brevicompactum. With the fermentation progresses, the growth of the fungi has begun to significantly deplete the mannan in the medium, so that the enzymatic substrate reduced and the product also reduced (Fig. 6d).
With comparison of treated and untreated SBM that hydrolyzed under the same conditions, the method introduction above was suitable for pretreatment soybean meal (SBM).
High-performance liquid chromatography (HPLC) analysis
To evaluate the applicability and accuracy of the proposed method, 60 samples were considered. On the condition mentioned above, samples were analyzed to check if there were any product peaks. 20% mannan of SBM were converted into mannose, while the control group demonstrated only 1.78% conversion. The results were demonstrated in the Fig. 7.
As a result, mannose yield reached to 0.0572 g per 16 g SBM which indicated that 0.358% SBM was converted into mannose after hydrolyzation which means total 20% mannan of SBM converting into mannose. According to Hsiao et al., the mannan content of non-dehulled SBM from North America is 1.79%, and we obtained 0.358% of mannose monosaccharide in the above experiment, so the yield ratio is 20%. The Fig. 7a was the mannose standard with HPLC analysis. Compared with the Fig. 7b and the Fig. 7c, for the control one, the mannose yield was only 0.0051 g per 16 g SBM, which mean that the conversion ratio for control group was 1.78%.
Discussion
Stinky tofu is a well-known and popular traditional fermented Chinese snack. Previous studies indicated that stinky brine fermentation is a type of alkaline fermentation because of the production of NH3 and referred to stinky tofu as an alkaline-fermented food [22]. The pH of the food rises to between about 8.0 and 9.0 because of ammonia production during the fermentation process and Bacillus sp. commonly participates in the fermentation process. Mannanase was generally used in the feed and food industries, and its product safety was widely concerned. Obviously, it was known in the literature that E. coli was commonly used host strain for recombinant protein expression and protein purification. However, the major disadvantage of E. coli was the formation of insoluble aggregates, at the same time, the extraction of recombinant proteins has also become a difficulty in the subsequent purification of proteins [27]. P. pastoris utilized methanol as a carbon source in the industry, providing the possibility of cheap and large-scale production of recombinant proteins. At the same time, its secretory expression characteristic endued it an unparalleled advantage in subsequent protein purification compared to E. coli expression system.
Demonstrated from the experimental statistic, the enzyme cloned from Bacillus pumilus was an alkaline prone beta-mannanase in which the highest enzymatic activity was at pH 6.0 and retained over 80% retained at the pH 6.0–8.0, meaning a wider range while maintaining higher relative activity than that of beta-mannanase from Bacillus sp. N16-5 ManAR2 (9–10) [22], Bacillus clausii BcManA (9–10) [48], Thermobifida fusca ManB (8–10) [49], Bacillus sp. N16-5 MAN493 (9–10) [50] and also wider than Streptomyces sp Mn428 (9–10) [51]. Among the enzymatic properties shown above, the manB had a better relative enzyme activity in a wider range of pH units, playing an important role in its application. With the alkaline prone character, the enzyme was applied for detergent production in commercial application [52] (see Table 3).
Pretreatment methods of biomass resources had been greatly enriched in recent years. Pretreatment methods for biomass resources had been greatly enriched in recent years, such as microwave-diluted acid treatment, acid treatment, steam explosion, and biological pretreatment. In a study on the effects of fungal pretreatment of wheat straw on the degradation of cellulose and hemicellulose, the researchers compared the effects of fungal pretreatments on subsequent cellulose and hemicellulose degradation rates. Among them, P. tigrinus pretreatment has the best effect on subsequent degradation [33] (see Table 4).
In our work, we also compared the enzymatic hydrolysis efficiency under two different fungi pretreatment of soybean meal, P. brevicompactum and P. oxalicum, the genome contained both cellulase and mannanase, as the best choice for pretreatment of soybean meal firstly, due to it is a rapider growth rate, two endogenous enzymes expressed synergistically to ensure the rapid growth. This causes the cellulose at the periphery of the soybean meal to be broken and relaxed, and the internal mannan is also substantially consumed so that the enzyme activity of the manB is detected slightly in the fermentation broth after the pretreatment. The strain P. brevicompactum industrially applied to produce mycophenolic acid, an immunosuppressant, rather than a strain that was a source of enzyme preparation resulting in softening the cellulose on the surface of the soybean meal without affecting the mannan. After 11 days of fungal pretreatment, the yeast-expressed mannanase was applied to hydrolyze pretreated SBM at 50 °C for 24 h to obtain 0.00358 g mannose per g dry SBM, leading a yield ratio to 0.358% which means total 20% mannan of SBM converted into mannose.
However, the conversion ratio was still low, which made it difficult for industrial applications. In the optimization of the reaction system, we also have many shortcomings, such as enzyme load and reaction time. Further investigations will focus on the mutation of the manB. Also, further optimization of soybean meal pre-treatment will be a consideration. In addition, optimization of enzymatic hydrolysis, such as enzyme dosage, should be considered.
References
Saha BC (2003) J Ind Microbiol Biotechnol 30(5):279–291
Kuhad RC, Singh A, Eriksson KEL (1997) In: Eriksson KEL et al (eds) Microorganisms and enzymes involved in the degradation of plant fiber cell walls. Springer, Berlin, Heidelberg
Moreira LRS, Filho EXF (2008) Appl Microbiol Biotechnol 79:165–178
Sittikijyothin W, Torres D, Gonçalves MP (2005) Carbohyd Polym 59:339–350
Mendoza NS, Arai M, Kawaguchi T, Cubol FS, Panerio EG, Yoshida T, Joson LM (1994) World J Microbiol Biotechnol 10(1):51–54
Benech RO, Li X, Patton D, Powlowski J, Storms R, Bourbonnais R, Tsang A (2007) Enzyme Microb Technol 41(6–7):740–747
Sallam AE, Almisherfi HM, El‐Feky MMM et al (2020) Aquaculture Nutrition
Mohapatra BR (2020) Biomass Convers Biorefinery
Jiang Z, Wei Y, Li D, Li L, Chai P, Kusakabe I (2006) Carbohyd Polym 66(1):88–96
Favaro CP, Baraldi IJ, Casciatori FP et al (2020) Biomolecules 10(2):227
Ikehara Y, Kojima N (2007) Curr Opin Mol Ther 9(1):53–61
Assreuy AMS, Shibuya MDD, Martins GJ, De Souza MLP, Cavada BS, Moreira RA, Flores CA (1997) Mediat Inflamm 6(3):201–210
Vuksan V, Jenkins DJ, Spadafora P, Sievenpiper JL, Owen R, Vidgen E, Bruce-Thompson C (1999) Diabetes Care 22(6):913–919
Hu X, Shi Y, Zhang P, Miao M, Zhang T, Jiang B (2016) Compr Rev Food Sci Food Saf 15(4):773–785
El-Nakkady SS, Hanna MM, Roaiah HM, Ghannam IA (2012) Eur J Med Chem 47:387–398
De Lonlay P, Seta N (2009) Biochim Biophys Acta (BBA) Mol Basis Dis 1792(9):841–843
Vorapreeda T, Thammarongtham C, Cheevadhanarak S, Laoteng K (2015) Microbiology 161(8):1613–1626
Su J, Li DS, Yan T, Wang JH (2007) J Microbiol 5:7
Bågenholm V, Wiemann M, Reddy SK, Bhattacharya A, Rosengren A, Logan DT, Stålbrand H (2019) J Biol Chem 294(23):9100–9117
Zhao D, Wang Y, Na J, Ping W, Ge J (2019) Prep Biochem Biotechnol 49(2):202–207
Nadaroglu H, Adiguzel G, Adiguzel A, Sonmez Z (2017) Eur Food Res Technol 243(2):193–201
Zheng H, Yu Z, Fu X, Li S, Xu J, Song H, Ma Y (2016) J Ind Microbiol Biotechnol 43(7):977–987
Chen S, Gray D, Ma J, Subramanian S (1998) Curr Protoc Protein Sci 12(1):5–10
Chittibabu G, Nath K, Das D (2006) Process Biochem 41(3):682–688
Chen L, Mohsin A, Chu J, Zhuang Y, Liu Y, Guo M (2017) Biotechnol Bioprocess Eng 22(6):767–773
Baghban R, Farajnia S, Ghasemi Y, Mortazavi M, Zarghami N, Samadi N (2018) Curr Pharm Biotechnol 19(6):451–467
Bracke A, Hoogewijs D, Dewilde S (2018) Anal Biochem 543:62–70
Ahmad M, Hirz M, Pichler H, Schwab H (2014) Appl Microbiol Biotechnol 98(12):5301–5317
Arsenault RJ, Lee JT, Latham R, Carter B, Kogut MH (2017) Poult Sci 96(12):4307–4316
LaFrance JT (2008) J Economet 147(2):336–349
National Research Council (US) (2000) Biobased Industrial Products: Priorities for Research and Commercialization. National Academies Press (US), Washington (DC)
Cao G, Ximenes E, Nichols NN, Zhang L, Ladisch M (2013) Biores Technol 146:604–610
Salvachúa D, Prieto A, López-Abelairas M, Lu-Chau T, Martínez ÁT, Martínez MJ (2011) Biores Technol 102(16):7500–7506
Thangaraj B, Solomon PR (2019) ChemBioEng Reviews 6(5):157–166
Shakeel T, Gupta M, Fatma Z, Kumar R, Kumar R, Singh R, Yazdani SS (2018) J Biol Chem 293(24):9148–9161
Chen S-Q, Cai X-H, Xie J-L, et al (2017) Starch‐Strke 69(1–2)
Norizan NABM, Halim M, Tan JS et al (2020) Molecules 25(15):3516
Park, Jungwoo, Knape et al (2019) J Appl Poultry Res 28(2):447–453
Lowry OH, Rosebrough NJ, Farr AL et al (1951) J Biol Chem 193(1):265–275
Xie J, Pan L, He Z et al (2020) Process Biochem 88(Jan.):51–59
Li Z, Yu Y, Sun J, Li D, Huang Y, Feng Y (2016) BioRes 11(1):54–70
Palomo JM, Muñoz G, Fernández-Lorente G, Mateo C, Fuentes M, Guisan JM, Fernández-Lafuente R (2003) J Mol Catal B Enzym 21(4–6):201–210
Mateo C, Palomo JM, Fernandez-Lorente G, Guisan JM, Fernandez-Lafuente R (2007) Enzyme Microb Technol 40(6):1451–1463
Shukor H, Abdeshahian P, Al-Shorgani NKN, Hamid AA, Rahman NA, Kalil MS (2016) Bioresour Technol 218:257–264
Hu R, Lin L, Liu T, Ouyang P, He B, Liu S (2008) J Biobased Mater Bioenergy 2(2):156–161
Miller GL (1959) Anal Chem 31(3):426–428
Lei X, Sun G, Xie J, Wei D (2013) Int J Syst Evol Microbiol 63(7):2501–2505
Zhou C, Xue Y, Ma Y (2018) Microb Cell Fact 17(1):124
Wang Y, Shu T, Fan P, Zhang H, Turunen O, Xiong H, Yu L (2017) Process Biochem 61:73–79
Pan X, Zhou J, Tian A, Le K, Yuan H, Xue Y, Lu H (2011) Biotechnol Lett 33(3):565–570
Pradeep GC, Cho SS, Choi YH, Choi YS, Jee JP, Seong CN, Yoo JC (2016) World J Microbiol Biotechnol 32(5):84
Srivastava PK, Kapoor M (2017) Biotechnol Adv 35(1):1–19
Germec M, Demirel F, Tas N, Ozcan A, Yilmazer C, Onuk Z, Turhan I (2017) Cellulose 24(10):4337–4353
Mathew AK, Parameshwaran B, Sukumaran RK, Pandey A (2016) Biores Technol 199:13–20
Li YX, Yi P, Liu J, Yan QJ, Jiang ZQ (2018) Bioresour Technol 256:30–37
Salvachúa D, Prieto A, López-Abelairas M et al (2011) Bioresour Technol 102(16):7500–7506
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declared that they have no conflicts of interest to this work. We declare that we do not have any commercial or associative interest that represents a conflict of interest in connection with the work submitted.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, M., Wang, J., Lin, L. et al. High-level expression of a β-mannanase (manB) in Pichia pastoris GS115 for mannose production with Penicillium brevicompactum fermentation pretreatment of soybean meal. Bioprocess Biosyst Eng 44, 549–561 (2021). https://doi.org/10.1007/s00449-020-02467-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-020-02467-6