Abstract
Spinal muscular atrophy (SMA) is caused by a deletion or mutation of the survival motor neuron 1 (SMN1) gene. Reduced SMN levels lead to motor neuron degeneration and muscular atrophy. SMN protein localizes to the cytoplasm and Cajal bodies. Moreover, in myofibrils from Drosophila and mice, SMN is a sarcomeric protein localized to the Z-disc. Although SMN participates in multiple functions, including the biogenesis of spliceosomal small nuclear ribonucleoproteins, its role in the sarcomere is unclear. Here, we analyzed the sarcomeric organization of SMN in human control and type I SMA skeletal myofibers. In control sarcomeres, we demonstrate that human SMN is localized to the titin-positive M-band and actin-positive I-band, and to SMN-positive granules that flanked the Z-discs. Co-immunoprecipitation assays revealed that SMN interacts with the sarcomeric protein actin, α-actinin, titin, and profilin2. In the type I SMA muscle, SMN levels were reduced, and atrophic (denervated) and hypertrophic (nondenervated) myofibers coexisted. The hypertrophied myofibers, which are potential primary targets of SMN deficiency, exhibited sites of focal or segmental alterations of the actin cytoskeleton, where the SMN immunostaining pattern was altered. Moreover, SMN was relocalized to the Z-disc in overcontracted minisarcomeres from hypertrophic myofibers. We propose that SMN could have an integrating role in the molecular components of the sarcomere. Consequently, low SMN levels might impact the normal sarcomeric architecture, resulting in the disruption of myofibrils found in SMA muscle. This primary effect might be independent of the neurogenic myopathy produced by denervation and contribute to pathophysiology of the SMA myopathy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Proximal spinal muscular atrophy (SMA) is an autosomal recessive neurodegenerative disorder that represents one of the most common genetic causes of infant mortality (Burghes and Beattie 2009; Coady and Lorson 2011). This devastating neuromuscular disorder is caused by deletion or mutation in the survival motor neuron (SMN1) gene, which encodes the survival motor neuron (SMN) protein (Lefebvre et al. 1995). SMN deficiency produces the selective degeneration of α-motor neurons (α-MNs) with the denervation of the affected motor units. Muscle denervation leads to a neurogenic secondary myopathy, characterized by progressive atrophy and weakness of the proximal muscles of the limbs and trunk, which usually culminate in lethal respiratory failure in the most severe form of the disease (Ogino and Wilson 2004).
The full-length human SMN (hereafter SMN) is a multifunctional protein of 294 amino acids that is expressed in both the cytoplasm and nucleus. Within the nucleus, it is concentrated in Cajal bodies (Carvalho et al. 1999; Förthmann et al. 2013; Hebert et al. 2001; Lafarga et al. 2009) and nuclear “gems” (gemini of Cajal bodies) (Liu and Dreyfuss 1996; Navascues et al. 2004). The SMN protein acquires a large functional range, forming homopolymers that associate with Gemin2-8 and Unrip proteins to form the SMN complex (Otter et al. 2007; Pellizzoni et al. 2002; Zhang et al. 2011).
SMN protein is mainly transcribed from the telomeric SMN1 gene, located on chromosome 5q13 (Brzustowicz et al. 1990; Melki et al. 1990). This gene contains nine exons (1, 2a, 2b, 3–8), but only exons 1–7 are translated into the SMN protein. An inverted duplication in the region of SMN1 resulted in a centromeric copy, the SMN2 gene, an evolutionarily recent event unique to Homo sapiens (Rochette et al. 2001). SMN2 differs from SMN1 in a translationally silent transition at + 6 in exon 7 (c.840 C to T) that weakens the exon splice site, favoring the skipping of exon 7 during mRNA splicing. Thus, the majority of SMN2 transcripts lack exon 7 (SMNΔ7) and encode a truncated, inactive and highly unstable protein that is quickly degraded by the ubiquitin-proteasome pathway (Lorson et al. 1998; Monani et al. 1999; Vitte et al. 2007). Only a small number of SMN2 transcripts are correctly spliced and produce functional SMN protein. Therefore, the SMN1 gene is almost the only source of the SMN protein under physiological conditions. Given the reduced production of SMN from the SMN2 gene, the SMN2 gene copy number in SMA patients is a key determinant of disease severity (Feldkötter et al. 2002).
The best-known function of SMN is the biogenesis of spliceosomal small nuclear ribonucleoproteins (snRNPs), essential splicing factors that are assembled into the spliceosome (for a review, see Matera and Wang (2014). Beyond its role in nuclear RNA metabolism, SMN is a multifunctional protein involved in multiple pathways, including mRNA translation (Bernabò et al. 2017; Tapia et al. 2017), the axonal transport of certain mRNAs such as β-actin mRNA (Rossoll et al. 2003), the organization of actin cytoskeleton dynamics through its interaction with profilin2 of the RhoA-ROCK pathway (Giesemann et al. 1999; Hensel and Claus 2018; Hensel et al. 2014; Nölle et al. 2011), endocytosis, autophagy, mitochondrial bioenergetics, and the ubiquitin pathways (Chaytow et al. 2018; Singh et al. 2017).
Although SMA has been traditionally considered an archetypal and selective disorder of α-MNs, there is abundant evidence that other extraneural tissues and organs are either overtly or subclinically affected in SMA patients and animal models (Coque et al. 2014; Hamilton and Gillingwater 2013; Simone et al. 2016). They include the skeletal muscle (Boyer et al. 2013; Castillo-Iglesias et al. 2019; Martínez-Hernández et al. 2009; Mutsaers et al. 2011; Walker et al. 2008), liver and pancreas (Bowerman et al. 2012; Szunyogova et al. 2016), spleen (Deguise et al. 2017), blood vessels (Somers et al. 2016), and heart (Bevan et al. 2010; Heier et al. 2010; Shababi et al. 2010).
Skeletal muscle provides an excellent model for research on the impact of SMN depletion on an organ outside the CNS. First, myofibers develop fully coupled with α-MNs through neuromuscular junctions (Greensmith and Vrbová 1997). Accordingly, SMN deficiency in SMA can intrinsically affect α-MNs, myofibers, and neuromuscular junctions (Arnold et al. 2004; Guettier-Sigrist et al. 2002; Kariya et al. 2008; Mutsaers et al. 2011; Tisdale and Pellizzoni 2015). Second, each muscle is a multitissue organ in which myofibers are organized with blood vessels, peripheral nerve fibers, and adipose and connective tissues of the endomysium, which also contribute to the structural and functional organization of the muscle (Görgens et al. 2015; Pedersen and Febbraio 2008). Consequently, SMN deficiency offers a wide range of possibilities to investigate whether skeletal myofibers and other muscle components can primarily contribute to SMA pathophysiology.
The early onset of symptomatic muscle weakness in SMA patients, before α-MN degeneration, advocates that, in addition to neurogenic myopathy caused by denervation, there is an intrinsic muscle pathology (Braun et al. 1995; Mutsaers et al. 2011; Sumner 2007). In fact, skeletal myofibers are smaller and severely affected in SMA fetuses, indicating that there is a delay in muscle growth and maturation (Martínez-Hernández et al. 2009). Similarly, SMN knockdown C2C12 myoblasts show defective development, fusion defects, cytoskeletal disorganization, and reduced proliferation capacity (Shafey et al. 2005). Furthermore, mouse models of SMA show a defective development of skeletal muscles, linking muscle weakness with fewer and smaller skeletal myofibers, even in the absence of α-MNs loss (Boyer et al. 2013; Bricceno et al. 2014; Cifuentes-Diaz et al. 2001; Guettier-Sigrist et al. 2002).
A previous study in Drosophila myofibrils reported that dSMN protein is located at both the actin-rich I-band and the α-actinin-rich Z-disc of the sarcomere and that the latter localization was conserved in mouse skeletal myofibrils (Rajendra et al. 2007). Later, it was shown that the entire mouse SMN complex localizes to the sarcomeric Z-disc (Walker et al. 2008). Moreover, in Drosophila, the SMN mutant myofibers fail to form thin filaments of actin, resulting in a disturbance of myofibrillogenesis (Rajendra et al. 2007). In this line, previous in vitro studies in neuronal models of SMN deficiency have shown the aberrant polymerization of actin through the activation of the ROCK-profilin or ROCK-ERK pathways, supporting a role of SMN in actin cytoskeleton dynamics (Bowerman et al. 2007; Hensel et al. 2014; Nölle et al. 2011). Interestingly, in vivo direct interaction between SMN and profilin-2 was demonstrated in the motoneuron cell line NSC34 (Sharma et al. 2005).
To gain a deeper understanding of the role of SMN in the organization of human skeletal myofibers, our first objective was to determine the sarcomeric distribution of human SMN and its putative molecular interactions with protein partners of the sarcomere. Likewise, we wanted to advance the knowledge of the potential primary effect of SMN deficiency on the architecture of the sarcomere in type I SMA hypertrophic myofibers, which are not directly affected by α-MN degeneration and subsequent denervation. Our results demonstrate that human SMN is a sarcomeric protein located in the I-band and M-band and provide substantial support to the view that SMN is involved in the architectural organization of the sarcomere (Rajendra et al. 2007). In fact, reduced SMN levels in type I SMA hypertrophic myofibers are associated with focal or segmental alterations of actin and other cytoskeletal proteins, as well as with the formation of minisarcomeres where SMN is relocated to Z-discs. These findings suggest that SMN deficiency in type I SMA primarily contributes to the severity of neurogenic SMA myopathy.
Material and methods
Human tissue samples
Human tissue samples were obtained following the Ethics Committee rules of the “Marqués de Valdecilla” University Hospital and Health Research Institute (IDIVAL). For this study, we used, as young adult controls, biopsy samples from vastus medialis (quadriceps) from two normal patients aged 18 and 21 years. As infant control, we used muscle samples from the autopsy of a 6-day-old postnatal subject that died from membrane hyaline disease without any neurological sign. The SMA muscle material was obtained from a male infant born at term that died of pneumonia at the age of 3 months (Tapia et al. 2012). The molecular genetic analysis for SMA had revealed the homozygous depletion of exons 7 and 8 of the SMN1 gen, and two copies of SMN2. In both cases, autopsy was performed 6 h after death.
Personal and family verbal consent was obtained for the biopsy and autopsy samples from control patients, respectively. Family written consent was obtained for the necropsy samples of the type I SMA patient.
Immunofluorescence and confocal microscopy
Longitudinal vastus medialis samples were pinned to cork, immersed in ice-cold relaxing buffer (100 mM NaCl, 2 mM KCl, 2 mM MgCl2, 6 mM K3PO4, 1 mM EGTA, and 0.1% glucose; pH 7.0) and relaxed at 4 °C until fixed in 3.7% paraformaldehyde (Gokhin et al. 2010). Muscle fragments were cryoprotected in 15% and 30% sucrose in PBS (phosphate saline buffer; 137 mM NaCl, 2.7 mM KCl, 8 mM Na2HPO4, and 2 mM KH2PO4; pH 7.4) until they sank, embedded in Tissue-Tek OCT Compound (Sakura Finetek USA), and frozen on a chilled metal block in dry ice. Cryosections, 8-μm thick, were mounted on SuperFrost slides and stored at − 80 °C until use. For immunofluorescence staining, sections were washed in ice-cold PBS, permeabilized for 45 min in PBS plus 0.5% Triton X-100, and blocked for 1 h in 4% BSA plus 1% goat serum (ab138478, Abcam) in PBS. Tissue-sections were immunolabeled with primary antibodies diluted in blocking buffer overnight at 4 °C, washed three times in PBS plus 0.05% Tween-20 (PBST), and then labeled with specific FITC-, Texas Red-, or Cy3-conjugated secondary antibodies in blocking buffer for 1 h at room temperature (RT). Some samples were counterstained with phalloidin-FITC (1:100; Sigma, UK) for the demonstration of actin filaments. Sections were then washed three times in PBS and mounted with the antifading medium Vectashield (Vector, USA).
The following primary antibodies were used in this study: mouse monoclonal anti-SMN (clone 8, BD Transduction Laboratories), anti-α-actinin (Abcam), anti-pan actin (Boehringer Mannheim), anti-gemin2 (Sigma, St. Louis, MO, USA), and anti-myosin (Developmental Studies Hybridoma Bank); rabbit polyclonal anti-SMN (Santa Cruz Biotechnologies, USA) and anti-profilin2 (Abcam). Rabbit affinity-purified polyclonal anti-titin M8/M9 antibody, raised against the C-terminal region of titin located at the M-band, was kindly donated by Prof. Siegfried Labeit, EMBL, Germany (Centner et al. 2000).
The study of the spinal cord α-MNs was performed on formalin-fixed tissue from control and SMA samples. Tissue fragments from the anterior horn were isolated and washed in PBS and cryoprotected with 30% sucrose. Cryosections, 10-μm thick, were counterstained with propidium iodide for the morphological and quantitative analysis of α-MNs. The number of α-MNs was counted in four sections of the control (postnatal subject) and SMA patient at segment L5.
Samples were examined with a LSM510 laser-scanning microscope (Zeiss, Germany) equipped with an Ar (488 nm), HeNe (543 nm), and HeNe (633 nm) ion lasers and using a × 63 immersion oil objective (1.4 NA). For double labeling experiments, images of the same confocal plane were sequentially recorded, and pseudo-colored images were generated and superimposed. TIFF images were transferred to Adobe Photoshop CC 2014 software (Adobe Systems Inc.) for presentation. For the fluorescence profile analysis, we used the image-profiling tool from the Zeiss software (LSM 5.1) where a cross-section line is drawn to obtain X profiles of fluorescence intensity patterns and to compare the intensity of FITC and Cy3 fluorescence signals within the same image line. Moreover, a quantitative analysis was performed for the colocalizations by calculating Mander’s overlap coefficients (MOC) using the JACoP ImageJ plugin (Bolte and Cordelières 2006). MOC determines the ratio of pixels in the green channel, corresponding to the actin, titin (M8/M9 epitope), α-actinin, or profilin2 signal, to pixels in red channel, corresponding to SMN, that are overlapped in double immunolabeled confocal images (n = 4) of control vastus medialis muscle samples.
Transmission electron microscopy
Muscle fascicles were fixed in 3% glutaraldehyde in 0.12 M phosphate buffer (0.12 M Na2HPO4 and 0.12 M NaH2PO4; pH 7.2) followed by tissue dissection of small fragments that were incubated overnight in same fixative at 4 °C. Tissue samples were washed in 0.12 M phosphate buffer, further fixed in 1% OsO4 for 2 h, washed again, contrasted with uranyl acetate, and dehydrated in increasing concentrations of acetones followed by propylene oxide and embedded Araldite (Electron Microscopy Sciences). Semithin sections (1 μm) of control and SMA patient were stained with toluidine blue to assess preservation of tissue structure and diagnosis of pathological changes of SMA myopathy compared with control on a light microscope. Subsequently, ultrathin sections were mounted on copper grids and stained with uranyl acetate and lead citrate for examination with a JEOL 1011 electron microscope operating at 80 kV.
Immunogold electron microscopy
Muscle fascicles were fixed in 3.7% paraformaldehyde in 0.1 M cacodylate buffer pH 7.4 for 3 h at room temperature. Samples were then washed in 0.1 M cacodylate buffer, dehydrated in increasing concentrations of methanol (at − 20 °C), embedded in Lowicryl K4M, and polymerized with ultraviolet irradiation at − 20 °C for 7 days. Ultrathin sections were mounted on nickel grids and sequentially incubated with 0.1 M glycine in PBS (15 min), 5% BSA in PBS (30 min), and the primary rabbit anti-SMN antibody (1:25) diluted in 50 mM Tris-HCl, pH 7.6, containing 5% BSA for 2 h at 37 °C. After washing, the sections were incubated with the goat anti-rabbit IgG antibody coupled to 15-nm gold particles (BioCell, UK; diluted 1:50 in PBS containing 1% BSA) for 1 h at RT. Following immunogold labeling, the grids were stained with uranyl acetate and examined with a JEOL 1011 electron microscope. As controls, ultrathin sections were treated as described above but with the primary antibodies omitted.
Western blotting
Tissue samples were homogenized with a Polytron PT-2000 (Kinematica, Luzern-Switzerland) in 10 volumes of ice-cold extraction buffer (50 mM Tris-HCl, 500 mM NaCl, 1 mM EDTA, 1 mM MgCl2, 1 mM dithiothreitol, and 2% NP-40; pH 8), and supplemented with EDTA-free complete protease inhibitor cocktail and PhosSTOP (Roche). Samples were left on ice for 20 min, and clarified by 10 min of centrifugation at 13,000 rpm. Protein lysates were boiled in 2X SDS sample buffer for 10 min, separated on 4–20% NuPage TG SDS-PAGE gels (Invitrogen), and transferred to nitrocellulose membranes using standard procedures. Membranes were blocked in PBST plus 5% non-fat dried milk (BioRad) and incubated with the following primary antibodies: mouse monoclonal anti-SMN (clone 8, BD Transduction Laboratories) and anti-α-actinin (ab9465, Abcam), goat polyclonal anti-pan actin (sc1616, Santa Cruz), and rabbit polyclonal anti-SMN (sc15320, Santa Cruz), anti-profilin2 (ab96676, Abcam) and anti-RhoGDI (ab133248, Abcam). After several washes in PBST, blots were developed with anti-mouse IRDye800DX or anti-goat or anti-rabbit IRDye680DX (Rockland Immunochemicals, USA) secondary antibodies. Protein bands were detected with an Odyssey™ Infrared-Imaging System (Li-Cor Biosciences) according to the Odyssey™ Western blotting protocol.
Co-immunoprecipitation
For the co-immunoprecipitation (co-IP) assays, control muscle samples were homogenized with a Polytron PT-2000 (Kinematica, Luzern-Switzerland) in 10 volumes of ice-cold IP buffer (50 mM Tris-HCl, 150 mM NaCl, 5 mM EDTA, and 1% NP-40; pH 8) supplemented with EDTA-free complete protease inhibitor cocktail and PhosSTOP (Roche), left on ice for 20 min, and clarified by 10 min of centrifugation at 13,000 rpm. Protein concentration of the supernatant was estimated using the Qubit assay (Invitrogen). A total of 500 μg of total muscle lysates were incubated with 30 μg of Gamma-bind-Sepharose beads (GE Healthcare) for 1 h in a rotatory wheel at 4 °C. After centrifugation at 6000 rpm for 3 min, the supernatants were incubated with 1 μg of primary antibody, either mouse anti-SMN for the co-IP of SMN complexes and anti-GFP as IgG control or rabbit anti-titin (Sigma) for the co-IP of titin complexes and 10 μl of rabbit serum (ab7487, Abcam) as IgG control. After 4 h, 100 μg of Gamma-bind-Sepharose beads were added and incubated for an additional 2 h. Immunocomplexes bound to the sheparose beads were pelleted and washed twice with complete IP buffer and 5 times in IP buffer without NP-40. Immunocomplexes were eluted by boiling the beads in 50 μl of 1X SDS sample buffer for 10 min. The input extract (diluted in 2X SDS sample buffer) and the eluted protein complexes were then subjected to SDS-PAGE and western blotting analysis, as described in the previous section. After incubation with horseradish peroxidase-coupled light chain specific secondary antibodies diluted at 1:5000 (#115-035-174 and #211-032-171, Jackson Immunoresearch, UK), immunocomplexes were detected by enhanced chemiluminescence reaction (Amersham Biosciences).
Statistical analysis
The morphometric analysis of individual fiber diameter size was performed in transversally sectioned muscle cryosections from the infant control and the SMA patient muscle samples stained with phalloidin-FITC. For maximum accuracy of the myofiber measurement, we adapted the method of the lesser diameter, defined as “the maximum diameter across the lesser aspect of the muscle fiber” described by Dubowitz and Sewry (2007). Thus, measurement of the lesser diameter of each myofiber in the field was performed on randomly collected confocal images using the ImageJ software (National Institutes of Health). Values, standard error (SD), and statistical analyses were calculated and plotted using the Prism 7 (GraphPad software). Two-tailed unpaired t test was conducted to determine the significance caused by SMN depletion in SMA samples compared with the infant control. Statistical significance was set at p < 0.05.
Results
Sarcomeric distribution of the SMN in human skeletal myofibers
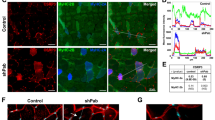
To determine the subcellular localization of SMN in control human myofibers, longitudinal cryosections of the vastus medialis muscle were immunostained with monoclonal or polyclonal anti-SMN antibodies. Irrespective of the antibody used, we observed that human SMN exhibited a striated staining pattern in bands along the myofibers (Figs. 1a, S1a). Furthermore, SMN also accumulated in granules within the SMN-positive bands (Figs. 1c–o, S1c, g). As negative controls, tissue sections were incubated only with the corresponding Texas Red- or Cy3-conjugated secondary antibody (Fig. S1c).
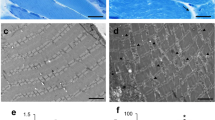
Human SMN localizes in the I- and M-bands and interacts with actin, α-actinin, and profilin2. a Longitudinal skeletal muscle cryosections of a young adult control subjected to immunofluorescence imaging using mouse monoclonal anti-SMN antibody. b Electron micrograph of the control sarcomere. c–q Confocal images of longitudinal cryosections of human vastus medialis control muscle double immunostained for SMN and either actin (c–e), the M-line epitope of titin (g–i), α-actinin (k–m), or profilin2 (o–q). Note the presence of numerous SMN-positive granules at the flanks of α-actinin-positive Z-discs (m). f, j, n, r Plot profiles along of a line of fluorescence intensity signals of SMN (red signal) and either actin (f), titinM8/M9 (j), α-actinin (n), or profilin2 (r) (green signals). s Three-dimensional reconstruction of confocal images illustrating a Z-disc immunolabeled for α-actinin (green signal) embraced by several SMN-positive granules (red signal). In panels o–q, the arrows illustrate the spatial association between the SMN-positive granules and profilin2 granules nearby the Z-discs. t, u Immunogold electron microscopy of SMN distribution in the human sarcomere. t Gold particles specifically decorate thin filaments of the I-band and are clustered in two SMN granules flanking a Z-disc. u Gold particles are localized along the M-band and its flanking domains (delimited by dotted lines), the called “M region,” described with high-resolution electron microscopy analysis of negatively stained ultrathin cryosections (Sjöström and Squire 1977). v–x Human muscle extracts (1 mg of total protein) were incubated with mouse anti-SMN (v and x) or rabbit anti-titinM8/M9 (w) antibodies, and the isolated immunocomplexes were analyzed by SDS-PAGE and Western blotting using anti-SMN (v–x), anti-α-actinin (v, w), anti-pan-actin (v), or anti-profilin2 (x) antibodies. Input lanes represent 5% of total lysate used for the co-IP analysis. As control for non-specific proteins bound to the antibodies, protein extracts were incubated with mouse monoclonal anti-GFP antibodies (IgG lane) (v and x) or with rabbit serum (w). Arrowhead in w indicates the SMN-specific band and asterisk in x indicates IgG light chain band (~ 25 kDa). Scale bars: a 5 μm; b 1 μm; c–q 2.5 μm; s 500 nm; t 120 nm; u 150 nm
To define the precise correlation of the SMN-positive striated bands with the structural components of the sarcomere (Figs. 1b, S1b), we performed double immunolabeling for SMN in combination with antibodies directed to actin (I-band), α-actinin (Z-disc), myosin (A-band), and titin (M-band). As negative controls, tissue sections were incubated only with the corresponding FITC-conjugated secondary antibody (Fig. S1c). Our results demonstrated that SMN-positive bands correspond to the actin-positive I-band (Fig. 1c–f) and titin-positive M-band (Fig. 1g–j), respectively, whereas the α-actinin-positive Z-disc (Fig. 1k–n) and myosin-positive A-band were negative (Fig. S1e-g). It was noteworthy, however, that SMN granules were tightly associated with both flanks of the Z-disc (Fig. 1k–m), as confirmed by confocal microscopy serial section 3-D reconstruction that showed solid SMN microloops of 1.36 ± 0.12 μm in diameter (mean ± SD, n = 100) embracing the Z-disc (Fig. 1s, S1d). Likewise, the localization of SMN in the I-band, Z-disc-attached granules, and M-band, including its specific flanking domains called the “M region” (Sjöström and Squire 1977), was confirmed by immunogold electron microscopy (Fig. 1t, u). The plots of the fluorescence intensity profiles of SMN (red) and the molecular markers of sarcomeric structures (green) across a line along the longitudinal axis of the myofiber confirmed the precise location of human SMN in the I-band and M-band (Fig. 1f, j) and its exclusion from the Z-disc and A-band (Fig. 1n). Moreover, the calculation of MOC validated colocalizations of SMN in the I-band (actin-SMN = 0.79 ± 0.05) and in the M-band (titin-SMN = 0.70 ± 0.06), as well as the very weak SMN signal in the Z-disc (α-actinin-SMN = 0.15 ± 0.04) and in the A-band (myosin-SMN = 0.26 ± 0.03). Regarding the nuclear distribution of SMN, we observed a diffuse nucleoplasmic signal of SMN with isolated spots of higher fluorescent intensity presumably corresponding to gems (Fig. S1n). Typical Cajal bodies immunoreactive for coilin were also observed in myonuclei (Fig. S1o).
Next, by co-immunoprecipitation (co-IP) assays in control muscle lysates, we demonstrated that SMN directly or indirectly interacts with α-actinin and actin (Fig. 1v). Furthermore, we also analyzed the putative interaction of SMN with titin. The titin co-IP assay was technically challenging due to the very large molecular size of titin (3000 kDa). In fact, only the immunoprecipitation of titin revealed that this protein establishes molecular interactions with SMN (Fig. 1w). As a control of the titin co-IP assay, we used the detection of the well-known interacting partner of titin, α-actinin (Atkinson et al. 2001) (Fig. 1w).
The SMN complex and the actin-binding protein profilin2 are localized in the sarcomere
To determine whether the human sarcomeric SMN forms a part of the SMN complex, as previously demonstrated in mouse myofibrils (Walker et al. 2008), we performed double immunolabeling for SMN and gemin2. We used gemin2 as an SMN complex marker since its direct interaction with SMN is crucial for SMN complex assembly (Lorson et al. 1998; Ogawa et al. 2007). Our results showed a strict colocalization of both molecules in the I-band and the M-band that was validated by the linear intensity profiles of the SMN and gemin2 fluorescence signals (Fig. S1i-l) as well as with the MOC (Gemin2-SMN = 0.68 ± 0.07). Furthermore, co-IP assay experiments in control muscle lysates demonstrated that SMN and gemin2 interact, indicating that the sarcomeric SMN forms part of the SMN complex (Fig. S1m).
Previous studies have established that SMN, through the polyproline stretches present at the C-terminal domain, directly interacts with the actin-binding proteins profilin1 and profilin2 (Giesemann et al. 1999; Nölle et al. 2011). This prompted us to investigate the sarcomeric localization of profilin2, which is highly expressed in skeletal muscle (Bowerman et al. 2007), and its possible correlation with the SMN distribution pattern. Double immunolabeling experiments revealed that profilin2 was located in the A-band and formed granules associated with the Z-disc, whereas SMN was detected at the I-band and M-band, as well as in Z-disc-associated granules (Fig. 1o–r). Curiously, some of these profilin2-positive granules were spatially associated with the SMN granules (Fig. 1o–r). In fact, the MOC of profilin2-SMN was 0.55 ± 0.07 in SMN-positive granules. Furthermore, the co-IP assay demonstrated the existence of a direct or indirect molecular interaction between SMN and profilin2 in the human control muscle (Fig. 1x).
The degeneration of α-MNs in type I SMA induces the formation of both “mixed-type grouping” and “group atrophy” muscle fascicles
The strict organization of SMN in the human sarcomere, as well as its interactions with some sarcomeric protein components, prompted us to investigate the potential impact of SMN depletion on the architecture of the sarcomere. We wondered whether in type I SMA muscle exists, in addition to the well-established neurogenic myopathy (for a review, see Dubowitz and Sewry (2007), an intrinsic SMN-dependent primary myopathy, which can be analyzed in hypertrophic (innervated) myofibers.
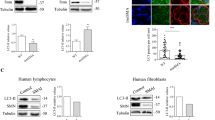
First, we confirmed the existence of α-MN loss by performing a quantitative analysis of neuronal body number in transverse cryosections of the spinal cord (L5 level) stained with propidium iodide. The study showed a loss of approximately 70% of α-MNs in the SMA patient compared with the control (Fig. 2a, b). Moreover, some of the remaining α-MNs exhibited signs of chromatolysis, a disruption of the protein synthesis machinery (Nissl bodies), compared with the prominent Nissl bodies found in control α-MNs (Fig. 2a, b, insets).
α-MN loss induces the formation of “group atrophy” and “mixed-type grouping” muscle fascicles in human type I SMA. a–b Cryosections of the spinal cord anterior horn from the infant control (a) and SMA patient (b) stained with propidium iodide (PI) illustrating the typical loss of α-MN in the SMA sample. Insets are representative examples of control (a, inset) and chromatolytic (b, inset) α-MNs. c–d Transversal cryosections of the vastus medialis muscle from the infant control (c) and the SMA patient (d) stained with Phalloidin-FITC. c Representative image of control muscle fascicles with myofibers of regular diameter. d In contrast, SMA muscle exhibits two categories of fascicles: “atrophic group,” composed exclusively of atrophic fibers, and “mixed-type grouping” where atrophic and hypertrophic fibers coexist. e Quantitative analysis of myofiber diameters measured in transversal cryosections from control and SMA muscle samples stained with phalloidin-FITC. The distribution of muscle fiber diameters was determined by measuring the smallest diameter of 100 transversal fibers selected randomly on three different images of each muscle sample. The horizontal black line within the scatter dot plot represents the mean for each group being the control 8.8 ± 1.38 μm. Note that in SMA, due the bimodal distribution of the measurements, two means were calculated 4.7 ± 1.74 μm and 21.9 ± 3.3 μm, corresponding to atrophic and hypertrophic myofibers, respectively. Data are mean ± SD. f–g Western blotting analysis of SMN protein expression level in spinal cord (f) and skeletal muscle (g) samples from the infant control and the SMA patient. Protein levels of actin (f) or RhoGDI (g) were used as loading controls, respectively. Scale bar: a–d 35 μm
The cardinal feature of the neurogenic myopathies caused by α-MN degeneration is the presence of myofibers forming both “group atrophy” and “mixed-type grouping” fascicles (Ince and Wharton 2007; Jokela et al. 2016). In the “group atrophy” fascicles, all myofibers are atrophic, reflecting a massive loss of the α-MNs that innervated these fascicles, a condition that is typically observed in type I SMA myopathy (Dubowitz and Sewry 2007; Ince and Wharton 2007). In the “mixed-type grouping” fascicles, the atrophic (denervated) myofibers are intermingled with scattered hypertrophic (innervated) ones belonging to functional motor units. The “mixed-type grouping” fascicles have particular relevance in the present study since they allow us to investigate whether the SMN deficiency intrinsically impacts on the organization of the contractile cytoskeleton using as a biological model the hypertrophic fibers.
In control vastus medialis muscle, cross-cryosections stained with phalloidin-FITC, a marker of actin filaments, showed typical fascicles with myofibers of regular size (Fig. 2c). In contrast, the type I SMA muscle cryosections exhibited atrophic myofibers of reduced diameter, forming both “group atrophy” and “mixed-type grouping” fascicles, in addition to scattered hypertrophic myofibers (Fig. 2d). The quantitative analysis of myofiber diameter revealed a unimodal distribution in control muscle, while the SMA muscle showed a bimodal distribution corresponding to the coexistence of atrophic (2 to 8 μm in diameter) and hypertrophic (> 20 μm in diameter) myofibers (Fig. 2e).
Western blot analysis revealed reduced SMN protein levels in both SMA vastus medialis muscle and spinal cord lysates compared with control samples (Fig. 2f, g). Interestingly, the enormous polyploidy of the SMA myofibers, as well as the existence of two SMN2 gene copies in the studied SMA patient, was not sufficient to ensure physiological SMN protein levels in SMA muscle. Therefore, SMN deficiency might primarily contribute to SMA myopathy.
Alterations of hypertrophic myofibers associated with reduced SMN levels in human type I SMA
To determine whether SMN deficiency is primarily associated with alterations in hypertrophic myofibers, we analyzed semithin sections stained with toluidine blue from control and SMA vastus medialis muscle (Figs. 3a, S2a-e). In toluidine blue-stained longitudinal sections from control muscle, myofibers exhibited a regular diameter and the characteristic cross-striated banding pattern of light (I-band) and dark (A-band) bands (Fig. S2a, b). The electron microscopic analysis confirmed the perfect parallelism of myofibrils and the lateral alignment of sarcomeres in which the I-band, A-band, M-band, and Z-disc were clearly defined (Fig. S2c, d).
Focal damage in hypertrophic myofibers of the human type I SMA. a–b Toluidine blue (a) and phalloidin-FITC (b) stained hypertrophic myofibers with focal damage, affecting a column with the bandwidth of a I-band (arrows). c Electron micrograph illustrating the disarray of the actin filaments from the I-band in focal damaged regions. Note the disruption of the Z-disc. d, e High-magnification electron microscopy of focal damage. Note the enlargement and disarray of the I-band. f, g Immunogold electron microscopy for the detection of actin (f) and myosin (g). Note the abundance of disordered actin-immunolabeled thin filaments (f) and the absence of myosin thick filaments (g) in focal damage sites. h–j Electron micrographs of representative electron-dense bodies nucleated at the Z-disc. k Immunogold electron microscopy for actin showing an electron-dense body decorated with numerous gold particles of actin immunoreactivity. Scale bars: a, b 7.5 μm; c, d, e 1.25 μm; f, g 300 nm; h, i 1 μm; j, k 600 nm
In longitudinal semithin sections and cryosections of the type I SMA vastus medialis muscle, both toluidine blue and phalloidin-FITC staining revealed that a high proportion of non-atrophic and hypertrophied myofibers (approximately 60%) exhibited focal or segmental structural damage, according to the extension of the affected sarcomeres in the longitudinal axis (Figs. 3a, b, S2e-i). Since these lesions must affect the contractile properties of the SMA myofiber, they might represent an important component in the pathogenesis of SMA myopathy.
The focal damage appeared as a narrow transverse column of disarrayed I-bands flanked by normal segments (Figs. 3a, b, S2e-f). A detailed examination by conventional electron microscopy and immunogold electron microscopy for the detection of actin and skeletal muscle myosin revealed the disruption of the I-band with disorganization of actin filaments and absence of myosin filaments (Fig. 3c–g). Furthermore, Z-discs were also disrupted or unrecognizable in sites of focal damage (Fig. 3c–f). Another ultrastructural feature was the presence of electron-dense bodies that were nucleated at the Z-disc and partially extended into the A-band, distorting the neighboring sarcomeres (Fig. 3h–j). These bodies consisted of an electron-dense core of Z-disc-like material and a peripheral halo of thin and thick myofilaments (Fig. 3j). By immunogold electron microscopy, it was shown that the electron-dense core was decorated by numerous gold particles of actin immunoreactivity (Fig. 3k).
Segmental abnormalities affected multiple consecutive sarcomeres with different degrees of myofibrillar alterations along the longitudinal axis of the myofiber (Figs. 4a, d, S2e, g-i). Longitudinal semithin sections stained with toluidine blue showed myofiber segments with misalignment and disruption of myofibrils, as well as the presence of multiple overcontracted minisarcomres (Fig. 4a, d). These alterations were confirmed by electron microscopy analysis (Fig. 4b, c). In particular, minisarcomeres exhibited striking and strictly regular shortening (< 1.5 μm vs ≈ 2.5 μm, in control sarcomeres) (Fig. 4e). Unlike the normal sarcomeres (Fig. 4f), minisarcomeres appeared demarked by widening Z-discs, and the I-band and A-band were indistinguishable (Fig. 4g). In fact, both actin and myosin filaments appeared to be directly attached to Z-discs, preserving their parallel arrangement to each other (Fig. 4 g). Minisarcomeres were commonly flanked by relaxed “macrosarcomeres” (> 4 μm in length), where Z-discs frequently adopted a jagged configuration, although the parallel arrangement of myofilaments was preserved (Fig. 4h). Notably, long segments with a complete misalignment of the Z-discs were also found in hypertrophic myofibers (Fig. 4i, j). In these segments, subsarcomeric rods of closely packed myofilaments without distinguishable sarcomeric bands were occasionally found (Fig. 4k, l).
Segmental damage in hypertrophic myofibers of the human type I SMA. a Panoramic image of a toluidine blue-stained semithin section of a human SMA vastus medialis muscle. Asterisks point to myofiber segments with structural alterations. b, c Electron micrographs of segmental damage with disrupted myofibrils. d Toluidine blue-stained semithin section of a myofiber showing a long segment of “minisarcomeres.” e Ultrastructural detailed of “minisarcomeres” in a hypertrophic myofiber. Note that I- and A-bands are indistinguishable and the widening of the Z-discs. f–h Detailed electron micrograph of a normal sarcomere (f), minisarcomere with no visible I-band and thick filaments directly anchored to the Z-disc (g), and a relaxed macrosarcomere exhibiting jagged Z-discs. i, j Electron micrographs of myofiber segments with a severe disruption of myofibrill organization. The Z-discs have lost their regular alignment and appeared scattered throughout the myofiber. k, l Panoramic (k) and detailed (l) electron microscopy images of a large subsarcomeric rod (asterisk) composed of closely packed myofilaments. Scale bars: a 6 μm; b, c 500 nm; d 10 μm; e 1.5 μm; f–h 300 nm; i 1.5 μm; j 300 nm; k 600 nm; l 150 nm
It should be noted that atrophic myofibers in the “mixed-type grouping” fascicles, in addition to their very reduced diameter, showed segmental alterations some of them similar to those found in hypertrophic myofibers in response to the SMN deficit. These alterations include the presence of (1) minisarcomeres (Fig. S3a, b), (2) twisted helically arranged myofibrils (Fig. S3c, d), (3) severe disruption of the sarcomere architecture (Fig. S3f-h), and (4) electron-dense bodies (Fig. S3i). We suggest that, in atrophic myofibers, the main neurogenic effect by denervation should be reinforced by an intrinsic, SMN-dependent effect, which clinically contribute to maximize the SMA myopathy.
SMN expression in hypertrophic and atrophic myofibers of the human type I SMA
Next, we analyzed the myofibrillar distribution pattern of SMN in type I SMA atrophic and hypertrophic myofibers. For this purpose, we used double immunolabeling for SMN and α-actinin in cryosections of longitudinally sectioned SMA myofibers. In both atrophic and hypertrophic myofibers, SMN was normally located in the I-band and M-band of myofiber segments with a preserved structure of sarcomeres (Fig. 5a–c, e–g). This distribution pattern was validated by plotting the fluorescence intensity profiles of SMN and α-actinin (Fig. 5d, h). However, SMN changed its normal myofiber distribution in pathological focal or segmental regions (Fig. 5i–o). Importantly, SMN was relocated to the Z-disc in (1) atrophic myofibers with helically arranged myofibrils and widened Z-discs (Fig. 5i–k), (2) minisarcomeres of hypertrophic myofibers with segmental damage (Fig. 5m–o), and (3) jagged Z-discs of relaxed macrosarcomeres in hypertrophic myofibers (Fig. 5m–o). The Z-disc redistribution of SMN was validated by the linear profiles of the fluorescence intensity of SMN and α-actinin (Fig. 5l, p).
SMN expression in human type I SMA myofibers. a–r Double immunofluorescence for SMN (red) and α-actinin (green) in longitudinally sectioned SMA myofibers. a–d In myofiber segments from atrophic myofibers with a preserved structure of the sarcomere, SMN was detected in the I-band and M-band. Note that a thinner segment with unstructured sarcomeres (abnormal segment) shows a weak and diffuse staining of both α-actinin and SMN. e–h Hypertrophic myofiber with focal damage (arrow) flanked by normally structured segments. At the focal damage region, SMN appeared irregularly distributed, whereas it was found at the I-band and M-band in the adjacent normal myofiber segments. i–k A twisted atrophic myofiber showing the colocalization of SMN and α-actinin in the widened Z-discs (i). m–o Hypertrophic myofiber with two clearly distinguishable segmental damage regions, overcontracted “minisarcomeres” (left) and relaxed “macrosarcomeres” (right). p Plot profile of the fluorescent SMN and α-actinin signals across a line passing through the overcontracted and relaxed sarcomeres illustrates the relocalization of SMN to Z-disc in both types of sarcomeres. q–r Representative example of an immature myoblast/myotube double immunolabeled for SMN (q) and α-actinin (r) is also found in human SMA type I skeletal muscle samples. Scale bars: a–c, i–k 6 μm; e–g, m–o 10 μm; q, r 4 μm
Finally, we want to point out the occasional presence of small-sized and unstructured myofibers characterized by the presence of multiple nuclear foci of SMN immunoreactivity, but with weak or undetectable expression in the sarcoplasm (Fig. 5q–r). These myofibers likely represent a stock of cells stopped in myogenesis, well in phase with the fusion-competent myoblast or in nascent myotube maturation, when its innervation has not yet occurred (Hindi et al. 2013; Martínez-Hernández et al. 2009).
Discussion
Low levels of SMN have been reported in skeletal muscle from both type I SMA patients and mouse models of the disease (Ahmad et al. 2016; Burlet et al. 1998; Coovert et al. 1997; Groen et al. 2018) and present results, suggesting that SMN deficiency can affect the function of this protein in skeletal myofibers and contribute to primary SMA myopathy. In this line, our immunocytochemical and co-IP results indicate that SMN is a sarcomeric protein in control human skeletal myofibers. The localization of SMN in the I-band and M-band, as well as its interactions with actin, profilin2, and titin, suggests a role of SMN in the maintenance of the sarcomere architecture, particularly in cytoskeletal actin dynamics. This view is consistent with a previous study in Drosophila showing that reduced SMN expression causes defects in myofibrillogenesis, resulting in severe neuromuscular disturbances (Rajendra et al. 2007). Moreover, our results also show that the focal or segmental alterations of myofibers found in the human type I SMA muscle are associated with changes in the sarcomeric distribution of the SMN. This study provides the first evidence of alterations in SMN localization in myopathies with muscle atrophy and denervation.
Previous investigations in purified myofibrils from Drosophila have shown that SMN is a sarcomeric protein that interacts with α-actinin and colocalizes with the I-band and Z-disc (Rajendra et al. 2007). Later, it was shown that in mouse myofibrils, sarcomeric SMN is recruited only in the Z-disc as part of the entire SMN complex (Rajendra et al. 2007; Walker et al. 2008). Like the Drososphila myofibers, we demonstrate that in human myofibers from the vastus medialis muscle, SMN is located in the I-band as part of the SMN complex. However, the human SMN offers three important differences in its sarcomeric compartmentalization relative to the fly sarcomeres (Rajendra et al. 2007): (1) the localization in the M-band, (2) the formation of SMN-positive granules flanking Z-disc, and (3) the exclusion of SMN from the Z-disc. This apparent discrepancy in sarcomeric SMN location may be partially due to species-specific differences and the methodology used. In mice and flies, the location of the SMN was determined in purified myofibrils (Rajendra et al. 2007; Walker et al. 2008), whereas in the present study, we analyzed SMN expression in complete myofibers treated with a relaxing buffer that prevents postmortem contraction (Gokhin et al. 2010). Likely, these physiological conditions have allowed us to localize human SMN in both I-bands and M-bands and Z-disc-associated granules as well as to determine its absence from the Z-disc.
The novel location of the SMN complex in the M-band has not been previously described. The M-band, which is situated in the middle of the A-band (Figs. 1b, S1b), is the place where the terminal tails of myosin light chains are superposed in a bipolar manner (for a review, see Lange et al. 2019). In addition, the M-band contains other proteins that allow the structural and functional coupling of the sarcomere with the sarcolemma, sarcoplasmic reticulum, and nuclear envelope (Agarkova and Perriard 2005). Among the proteins located in the M-band, particularly interesting is titin, a relevant giant protein (~ 3–4 MDa) associated with myosin. One single titin molecule spans a half-sarcomere from the Z-disc to the M-line (Fig. S1b) (Gautel et al. 1999). Due to its spatial organization, titin is considered a key scaffolding protein essential for the maintenance of the sarcomere integrity during the relaxation-contraction cycle (Agarkova and Perriard 2005; Kontrogianni-Konstantopoulos et al. 2009; Lange et al. 2019). Although an SMN domain of interaction with titin has not yet been identified, our results clearly demonstrate the localization of SMN in the M-band, and the co-IP assay indicates a direct or indirect interaction between SMN and titin. This raises the possibility that SMN may play a role in M-band maintenance. In fact, in SMA hypertrophic myofibers, SMN relocalizes to the Z-disc whenever the M-band is lost or indistinguishable in overcontracted minisarcomeres of damaged segmental regions. We suggest that the spatial rearrangement of myofilaments in minisarcomeres could potentiate the molecular interaction of SMN and α-actinin as revealed by the co-IP assays in Drosophila, mouse (Rajendra et al. 2007), and human skeletal muscle (present results).
It is noteworthy that human SMN protein localizes in the I-band of normal isometric sarcomeres, but it is absent in actin-free sarcomeric regions, such as the sub-band H, within of the A-band, and Z-disc. Nevertheless, SMN may aggregate in granules flanking the Z-disc. Consistent with the colocalization of actin and SMN in the I-band, our co-IP assays in control muscle lysates showed an interaction between these two proteins. These findings suggest that the sarcomeric SMN may be involved in the architecture of the sarcomere via its direct or indirect interaction with the actin cytoskeleton. Indeed, in hypertrophic SMA myofibers with focal or segmental damage, SMN immunostaining appears blurred in sites with a disarray of actin filaments, whereas it preserves its typical staining pattern in the normal sarcomeres. These findings suggest an SMN-dependent primary dysfunction of actin proteostasis in myofiber damage sites. Interestingly, the focal disruption of the I-band in SMA hypertrophic myofibers can cause eccentric contraction and, consequently, an impairment of normal myofiber function, contributing to SMA myopathy. However, this focal damage of the I-band cannot be considered a specific indicator of SMA myopathy since similar muscle lesions have been reported in biopsies of healthy athletes subjected to sustained eccentric contraction (Newham et al. 1983).
The view that SMN deficiency may intrinsically affect actin cytoskeleton dynamics (Hensel and Claus 2018) is reinforced by our finding of aberrant accumulations of actin filaments in electron-dense bodies. The presence of actin-dense bodies has been previously reported in SMA (Dubowitz and Sewry 2007) and is also common in other myopathies characterized by a dysfunction of actin expression and dynamics resulting in a pathological accumulation of thin filaments. They include progressive actin-accumulation myopathy, as well as other muscle disorders caused by mutations in the ACTA1 (Actin Alpha 1, Skeletal Muscle) gene that give rise to different forms of congenital myopathies (Bornemann et al. 1996; Goebel et al. 1997; North and Laing 2008; Sharma and Goebel 2005).
Regarding profilin2, our results demonstrate its localization in the A-band and in granules associated with the Z-disc. In particular, the lateral ends of the profilin2-positive A-band correspond to the zone where the bipolar extremes of the actin filaments extend into the A-band (Fig. S1b). Since profilin isoforms are major regulator of actin polymerization and dynamics (Hensel and Claus 2018; Ono 2010; Suarez et al. 2015), the sarcomeric distribution of profilin2 suggests a putative regulatory role of the thin filament length. Curiously, the rapid growth of the sarcomeric actin filament mainly occurs at the minus end (Nicholson-Dykstra et al. 2005; Ono 2007; Paavilainen et al. 2004) that extends into the profilin2-positive A-band. In this localization, profilin2 might promote the polymerization of G-actin into F-actin filaments. Moreover, we found by co-IP analysis that profilin2 and SMN interact in human muscle lysates, as has also been reported by co-IP assays and in vitro binding studies in cultured HeLa cells and spinal cord α-MNs (Giesemann et al. 1999). Interestingly, SMN has also been found to colocalize with profilin2 in neurite-like extensions (Sharma et al. 2005).
Previous studies, using an in vivo protein interaction system, have shown that SMN directly interacts with profilin2, regulating its phosphorylation by the RhoA/ROCK pathway, a key modulator of the actin dynamics (Luo et al. 1997), and its catalytic activity in actin polymerization (Nölle et al. 2011). Moreover, SMN knockdown or loss induces a decrease in SMN-profin2a complexes and enhanced activation of the RhoA/ROCK pathway, resulting in the hyperphosphorylation of profilin2 and increased F-actin (Hensel and Claus 2018; Nölle et al. 2011). Similarly, the RhoA-ROCK pathway is abnormally upregulated in the spinal cord and skeletal muscle of SMA mouse models with reduced SMN levels (Bowerman et al. 2007, 2012; Simone et al. 2016). The existence of a functional link between SMN and profilin2 is also supported by the observation that SMN depletion with SMN knockdown in PC12 cells upregulates the expression and phosphorylation of profilin2, resulting in a perturbation of the actin cytoskeleton and defective neuritogenesis (Bowerman et al. 2007; Hall 1998; Nölle et al. 2011). Taken together, low SMN levels seem to deregulate the formation of profilin2-SMN complexes, resulting in a perturbation of actin polymerization. We suggest that the dysfunction of actin dynamics could contribute to the focal or segmental alterations of the sarcomeric architecture found in the type I SMA myofibers. Further studies are required to explore the complexity and importance of the SMN- and ROCK-dependent misregulation of profilin2 in the actin cytoskeleton disarray of SMA muscle.
It is noteworthy that the reduction in SMN levels in SMA muscle, in addition to actin dynamics, might also perturb other essential SMN functions for myofiber homeostasis, such as the biogenesis of spliceosomal snRNPs, mRNA processing, transport and translation, endocytosis, mitochondrial homeostasis, and the maintenance of the myoneural junctions (for a review, see Chaytow et al. 2018; Li et al. 2014; Singh et al. 2017). In fact, we have recently shown that the reduction in SMN in type I SMA myofibers is associated with a severe perturbation of global pre-mRNA transcription and splicing. In particular, we have observed a dramatic heterochromatinization (gene silencing) of extensive genome domains and the accumulation of splicing factors in large nuclear speckles (splicing inhibition) (Castillo-Iglesias et al. 2019). In this context, our observation of focal or segmental cytoskeletal alterations raises the possibility of a differential affectation of “myonuclear domains,” the amount of cytoplasm supported by a single myonucleus (Allen et al. 1999; Hall and Ralston 1989), in type I SMA hypertrophic myofibers. Further studies in mouse models of SMA will be necessary to address whether the focal or segmental myofiber alterations correspond to one or several fragile myonuclear domains with higher perturbation of mRNA transcription, processing, and translation required to maintain the contractile apparatus.
On the other hand, recent evidence indicates that SMN physically interacts with both TRAF6 (TNF receptor associated factor 6) and the IKK (IkB kinase) complex and functions as an inhibitor for TRF6-mediated NF-kB signaling (Kim and Choi 2017). Moreover, TRAF6-mediated activation of NF-kB induces the activation of the ubiquitin-proteasome and autophagy systems in skeletal myofibers under denervation and atrophy (Paul et al. 2010). In this context, in SMA patients with severely reduced expression of SMN, the potential activation of the TRAF6-NF-kB signaling axis might contribute to the pathogenesis of SMA myopathy through the dysregulation of major proteolytic pathways.
Concluding remarks
In conclusion, our results in human skeletal myofibers and others in Drosophila and mouse myofibrils (Rajendra et al. 2007; Walker et al. 2008) are consistent with an important role for SMN in the maintenance of sarcomeric architecture and function. Therefore, SMN deficiency may primarily contribute to the focal or segmental disarray of actin filaments and sarcomeric architecture found in SMA hypertrophic (innervated) myofibers. Since these lesions must affect the contractile properties of the SMA myofibers, they might contribute to the pathogenesis of SMA myopathy.
Low SMN levels in SMA myofibers could also impact transcription and pre-RNA splicing machinery (Castillo-Iglesias et al. 2019), leading to a dysfunction of cytoskeletal protein synthesis required for the myofiber homeostasis (Görgens et al. 2015; Lee and Jun 2019; Pedersen and Febbraio 2008).
Abbreviations
- SMN1 :
-
Survival Motor Neuron 1 gene
- SMN2 :
-
Survival Motor Neuron 2 gene
- SMN:
-
Survival motor neuron protein
- SMA:
-
Spinal muscular atrophy
- α-MNs:
-
α-Motor neurons
- snRNP:
-
Spliceosomal small nuclear ribonucleoproteins
- MOC:
-
Mander´s overlap coefficient
References
Agarkova I, Perriard J-C (2005) The M-band: an elastic web that crosslinks thick filaments in the center of the sarcomere. Trends Cell Biol 15:477–485
Ahmad S, Bhatia K, Kannan A, Gangwani L (2016) Molecular mechanisms of neurodegeneration in spinal muscular atrophy. J Exp Neurosci 10:39–49
Allen DL, Roy RR, Edgerton VR (1999) Myonuclear domains in muscle adaptation and disease. Muscle Nerve 22:1350–1360
Arnold A-S, Gueye M, Guettier-Sigrist S, Courdier-Fruh I, Coupin G, Poindron P, Gies J-P (2004) Reduced expression of nicotinic AChRs in myotubes from spinal muscular atrophy I patients. Lab Investig 84:1271–1278
Atkinson RA, Joseph C, Kelly G, Muskett FW, Frenkiel TA, Nietlispach D, Pastore A (2001) Ca 2+ -independent binding of an EF-hand domain to a novel motif in the α-actinin–titin complex. Nat Struct Biol 8:853–857
Bernabò P, Tebaldi T, Groen EJN, Lane FM, Perenthaler E, Mattedi F, Newbery HJ, Zhou H, Zuccotti P, Potrich V et al (2017) In vivo translatome profiling in spinal muscular atrophy reveals a role for SMN protein in ribosome biology. Cell Rep 21:953–965
Bevan AK, Hutchinson KR, Foust KD, Braun L, McGovern VL, Schmelzer L, Ward JG, Petruska JC, Lucchesi PA, Burghes AHM et al (2010) Early heart failure in the SMNDelta7 model of spinal muscular atrophy and correction by postnatal scAAV9-SMN delivery. Hum Mol Genet 19:3895–3905
Bolte S, Cordelières FP (2006) A guided tour into subcellular colocalization analysis in light microscopy. J Microsc 224:213–232
Bornemann A, Petersen MB, Schmalbruch H (1996) Fatal congenital myopathy with actin filament deposits. Acta Neuropathol 92:104–108
Bowerman M, Shafey D, Kothary R (2007) Smn depletion alters profilin II expression and leads to upregulation of the RhoA/ROCK pathway and defects in neuronal integrity. J Mol Neurosci 32:120–131
Bowerman M, Swoboda KJ, Michalski JP, Wang GS, Reeks C, Beauvais A, Murphy K, Woulfe J, Screaton RA, Scott FW et al (2012) Glucose metabolism and pancreatic defects in spinal muscular atrophy. Ann Neurol 72:256–268
Boyer JG, Ferrier A, Kothary R (2013) More than a bystander: the contributions of intrinsic skeletal muscle defects in motor neuron diseases. Front Physiol 4:356
Braun S, Warter JM, Poindron P, Croizat B, Lagrange MC (1995) Constitutive muscular abnormalities in culture in spinal muscular atrophy. Lancet 345:694–695
Bricceno KV, Martinez T, Leikina E, Duguez S, Partridge TA, Chernomordik LV, Fischbeck KH, Sumner CJ, Burnett BG (2014) Survival motor neuron protein deficiency impairs myotube formation by altering myogenic gene expression and focal adhesion dynamics. Hum Mol Genet
Brzustowicz LM, Lehner T, Castilla LH, Penchaszadeh GK, Wilhelmsen KC, Daniels R, Davies KE, Leppert M, Ziter F, Wood D et al (1990) Genetic mapping of chronic childhood-onset spinal muscular atrophy to chromosome 5q1 1.2?13.3. Nature 344:540–541
Burghes AHM, Beattie CE (2009) Spinal muscular atrophy: why do low levels of survival motor neuron protein make motor neurons sick? Nat Rev Neurosci 10:597–609
Burlet P, Huber C, Bertrandy S, Ludosky MA, Zwaenepoel I, Clermont O, Roume J, Delezoide AL, Cartaud J, Munnich A et al (1998) The distribution of SMN protein complex in human fetal tissues and its alteration in spinal muscular atrophy. Hum Mol Genet 7:1927–1933
Carvalho T, Almeida F, Calapez A, Lafarga M, Berciano MT, Carmo-Fonseca M (1999) The spinal muscular atrophy disease gene product, SMN: a link between snRNP biogenesis and the Cajal (coiled) body. J Cell Biol 147:715–728
Castillo-Iglesias MS, Berciano MT, Narcis JO, Val-Bernal JF, Rodriguez-Rey JC, Tapia O, Lafarga M (2019) Reorganization of the nuclear compartments involved in transcription and RNA processing in myonuclei of type I spinal muscular atrophy. Histochem Cell Biol 152:227–237
Centner, T., Fougerousse, F., Freiburg, A., Witt, C., Beckmann, J.S., Granzier, H., Trombitás, K., Gregorio, C.C., and Labeit, S. (2000). Molecular tools for the study of titin’s differential expression. In Advances in Experimental Medicine and Biology, pp. 35–52
Chaytow H, Huan H, Gillingwater TH, Faller KME (2018) The role of survival motor neuron protein (SMN) in protein homeostasis. Cell Mol Life Sci 75:3877–3894
Cifuentes-Diaz C, Frugier T, Tiziano FD, Lacène E, Roblot N, Joshi V, Moreau MH, Melki J (2001) Deletion of murine SMN exon 7 directed to skeletal muscle leads to severe muscular dystrophy. J Cell Biol 152:1107–1114
Coady TH, Lorson CL (2011) SMN in spinal muscular atrophy and snRNP biogenesis. Wiley Interdiscip Rev RNA 2:546–564
Coovert DD, Le TT, McAndrew PE, Strasswimmer J, Crawford TO, Mendell JR, Coulson SE, Androphy EJ, Prior TW, Burghes AHM (1997) The survival motor neuron protein in spinal muscular atrophy. Hum Mol Genet 6:1205–1214
Coque E, Raoul C, Bowerman M (2014) ROCK inhibition as a therapy for spinal muscular atrophy: understanding the repercussions on multiple cellular targets. Front Neurosci 8:271
Deguise MO, De Repentigny Y, McFall E, Auclair N, Sad S, Kothary R (2017) Immune dysregulation may contribute to disease pathogenesis in spinal muscular atrophy mice. Hum Mol Genet 26:801–819
Dubowitz V, and Sewry C (2007) Muscle biopsy: a practical approach. Saunders, Elsevier.
Feldkötter M, Schwarzer V, Wirth R, Wienker TF, Wirth B (2002) Quantitative analyses of SMN1 and SMN2 based on real-time lightCycler PCR: fast and highly reliable carrier testing and prediction of severity of spinal muscular atrophy. Am J Hum Genet 70:358–368
Förthmann B, Brinkmann H, Ratzka A, Stachowiak MK, Grothe C, Claus P (2013) Immobile survival of motoneuron (SMN) protein stored in Cajal bodies can be mobilized by protein interactions. Cell Mol Life Sci 70:2555–2568
Gautel M, Mues A, Young P (1999) Control of sarcomeric assembly: the flow of information on titin. Rev Physiol Biochem Pharmacol 138:97–137
Giesemann T, Rathke-Hartlieb S, Rothkegel M, Bartsch JW, Buchmeier S, Jockusch BM, Jockusch H (1999) A role for polyproline motifs in the spinal muscular atrophy protein SMN. Profilins bind to and colocalize with smn in nuclear gems. J Biol Chem 274:37908–37914
Goebel HH, Anderson JR, Hübner C, Oexle K, Warlo I (1997) Congenital myopathy with excess of thin myofilaments. Neuromuscul Disord 7:160–168
Gokhin DS, Lewis RA, McKeown CR, Nowak RB, Kim NE, Littlefield RS, Lieber RL, Fowler VM (2010) Tropomodulin isoforms regulate thin filament pointed-end capping and skeletal muscle physiology. J Cell Biol 189:95–109
Görgens SW, Eckardt K, Jensen J, Drevon CA, Eckel J (2015) Exercise and regulation of adipokine and myokine production. Prog Mol Biol Transl Sci 135:313–336
Greensmith L, Vrbová G (1997) Disturbances of neuromuscular interaction may contribute to muscle weakness in spinal muscular atrophy. Neuromuscul Disord 7:369–372
Groen EJN, Perenthaler E, Courtney NL, Jordan CY, Shorrock HK, van der Hoorn D, Huang Y-T, Murray LM, Viero G, Gillingwater TH (2018) Temporal and tissue-specific variability of SMN protein levels in mouse models of spinal muscular atrophy. Hum Mol Genet 27:2851–2862
Guettier-Sigrist S, Hugel B, Coupin G, Freyssinet J-M, Poindron P, Warter J-M (2002) Possible pathogenic role of muscle cell dysfunction in motor neuron death in spinal muscular atrophy. Muscle Nerve 25:700–708
Hall A (1998) Rho GTPases and the actin cytoskeleton. Science 279(80):509–514
Hall ZW, Ralston E (1989) Nuclear domains in muscle cells. Cell 59:771–772
Hamilton G, Gillingwater TH (2013) Spinal muscular atrophy: going beyond the motor neuron. Trends Mol Med 19:40–50
Hebert MD, Szymczyk PW, Shpargel KB, Matera AG (2001) Coilin forms the bridge between Cajal bodies and SMN, the spinal muscular atrophy protein. Genes Dev 15:2720–2729
Heier CR, Satta R, Lutz C, Didonato CJ (2010) Arrhythmia and cardiac defects are a feature of spinal muscular atrophy model mice. Hum Mol Genet 19:3906–3918
Hensel N, Claus P (2018) The actin cytoskeleton in SMA and ALS: how does it contribute to motoneuron degeneration? Neuroscientist 24:54–72
Hensel N, Stockbrügger I, Rademacher S, Broughton N, Brinkmann H, Grothe C, Claus P (2014) Bilateral crosstalk of rho- and extracellular-signal-regulated-kinase (ERK) pathways is confined to an unidirectional mode in spinal muscular atrophy (SMA). Cell Signal 26:540–548
Hindi SM, Tajrishi MM, Kumar A (2013) Signaling mechanisms in mammalian myoblast fusion. Sci. Signal 6:re2
Ince, P.G., and Wharton, S.B. (2007). Chapter 5 cytopathology of the motor neuron. In Handbook of Clinical Neurology, pp 89–119
Jokela M, Huovinen S, Raheem O, Lindfors M, Palmio J, Penttilä S, Udd B (2016) Distinct muscle biopsy findings in genetically defined adult-onset motor neuron disorders. PLoS One 11:e0151376
Kariya S, Park G-H, Maeno-Hikichi Y, Leykekhman O, Lutz C, Arkovitz MS, Landmesser LT, Monani UR (2008) Reduced SMN protein impairs maturation of the neuromuscular junctions in mouse models of spinal muscular atrophy. Hum Mol Genet 17:2552–2569
Kim EK, Choi E-J (2017) SMN1 functions as a novel inhibitor for TRAF6-mediated NF-κB signaling. Biochim Biophys Acta - Mol Cell Res 1864:760–770
Kontrogianni-Konstantopoulos A, Ackermann MA, Bowman AL, Yap SV, Bloch RJ (2009) Muscle giants: molecular scaffolds in sarcomerogenesis. Physiol Rev 89:1217–1267
Lafarga M, Casafont I, Bengoechea R, Tapia O, Berciano MT (2009) Cajal’s contribution to the knowledge of the neuronal cell nucleus. Chromosoma 118:437–443
Lange, S., Pinotsis, N., Agarkova, I., and Ehler, E. (2019). The M-band: the underestimated part of the sarcomere. Biochim. Biophys. Acta - Mol. Cell Res
Lee JH, Jun H-S (2019) Role of Myokines in regulating skeletal muscle mass and function. Front Physiol 10:42
Lefebvre S, Bürglen L, Reboullet S, Clermont O, Burlet P, Viollet L, Benichou B, Cruaud C, Millasseau P, Zeviani M (1995) Identification and characterization of a spinal muscular atrophy-determining gene. Cell 80:155–165
Li DK, Tisdale S, Lotti F, Pellizzoni L (2014) SMN control of RNP assembly: from post-transcriptional gene regulation to motor neuron disease. Semin Cell Dev Biol 32:22–29
Liu Q, Dreyfuss G (1996) A novel nuclear structure containing the survival of motor neurons protein. EMBO J 15:3555–3565
Lorson CL, Strasswimmer J, Yao JM, Baleja JD, Hahnen E, Wirth B, Le T, Burghes AH, Androphy EJ (1998) SMN oligomerization defect correlates with spinal muscular atrophy severity. Nat Genet 19:63–66
Luo L, Jan LY, Jan YN (1997) Rho family small GTP-binding proteins in growth cone signalling. Curr Opin Neurobiol 7:81–86
Martínez-Hernández R, Soler-Botija C, Also E, Alias L, Caselles L, Gich I, Bernal S, Tizzano EF (2009) The developmental pattern of myotubes in spinal muscular atrophy indicates prenatal delay of muscle maturation. J Neuropathol Exp Neurol 68:474–481
Matera AG, Wang Z (2014) A day in the life of the spliceosome. Nat Rev Mol Cell Biol 15:108–121
Melki J, Sheth P, Abdelhak S, Burlet P, Bachelot MF, Lathrop MG, Frezal J, Munnich A (1990) Mapping of acute (type I) spinal muscular atrophy to chromosome 5q12-q14. The French Spinal Muscular Atrophy Investigators. Lancet (London, England) 336:271–273
Monani UR, Lorson CL, Parsons DW, Prior TW, Androphy EJ, Burghes AH, McPherson JD (1999) A single nucleotide difference that alters splicing patterns distinguishes the SMA gene SMN1 from the copy gene SMN2. Hum Mol Genet 8:1177–1183
Mutsaers CA, Wishart TM, Lamont DJ, Riessland M, Schreml J, Comley LH, Murray LM, Parson SH, Lochmüller H, Wirth B et al (2011) Reversible molecular pathology of skeletal muscle in spinal muscular atrophy. Hum Mol Genet 20:4334–4344
Navascues J, Berciano MT, Tucker KE, Lafarga M, Matera AG (2004) Targeting SMN to Cajal bodies and nuclear gems during neuritogenesis. Chromosoma 112:398–409
Newham DJ, McPhail G, Mills KR, Edwards RH (1983) Ultrastructural changes after concentric and eccentric contractions of human muscle. J Neurol Sci 61:109–122
Nicholson-Dykstra S, Higgs HN, Harris ES (2005) Actin dynamics: growth from dendritic branches. Curr Biol 15:R346–R357
Nölle A, Zeug A, van Bergeijk J, Tönges L, Gerhard R, Brinkmann H, Al Rayes S, Hensel N, Schill Y, Apkhazava D et al (2011) The spinal muscular atrophy disease protein SMN is linked to the rho-kinase pathway via profilin. Hum Mol Genet 20:4865–4878
North, K.N., and Laing, N.G. (2008). Skeletal muscle alpha-actin diseases. In Advances in Experimental Medicine and Biology, pp 15–27
Ogawa C, Usui K, Aoki M, Ito F, Itoh M, Kai C, Kanamori-Katayama M, Hayashizaki Y, Suzuki H (2007) Gemin2 plays an important role in stabilizing the survival of motor neuron complex. J Biol Chem 282:11122–11134
Ogino S, Wilson RB (2004) Spinal muscular atrophy: molecular genetics and diagnostics. Expert Rev Mol Diagn 4:15–29
Ono, S. (2007). Mechanism of depolymerization and severing of actin filaments and its significance in cytoskeletal dynamics. In International Review of Cytology, pp. 1–82
Ono S (2010) Dynamic regulation of sarcomeric actin filaments in striated muscle. Cytoskeleton 67:677–692
Otter S, Grimmler M, Neuenkirchen N, Chari A, Sickmann A, Fischer U (2007) A comprehensive interaction map of the human survival of motor neuron (SMN) complex. J Biol Chem 282:5825–5833
Paavilainen VO, Bertling E, Falck S, Lappalainen P (2004) Regulation of cytoskeletal dynamics by actin-monomer-binding proteins. Trends Cell Biol 14:386–394
Paul PK, Gupta SK, Bhatnagar S, Panguluri SK, Darnay BG, Choi Y, Kumar A (2010) Targeted ablation of TRAF6 inhibits skeletal muscle wasting in mice. J Cell Biol 191:1395–1411
Pedersen BK, Febbraio MA (2008) Muscle as an endocrine organ: focus on muscle-derived Interleukin-6. Physiol Rev 88:1379–1406
Pellizzoni L, Yong J, Dreyfuss G (2002) Essential role for the SMN complex in the specificity of snRNP assembly. Science 298:1775–1779
Rajendra TK, Gonsalvez GB, Walker MP, Shpargel KB, Salz HK, Matera AG (2007) A Drosophila melanogaster model of spinal muscular atrophy reveals a function for SMN in striated muscle. J Cell Biol 176:831–841
Rochette C, Gilbert N, Simard L (2001) SMN gene duplication and the emergence of the SMN2 gene occurred in distinct hominids: SMN2 is unique to Homo sapiens. Hum Genet 108:255–266
Rossoll W, Jablonka S, Andreassi C, Kröning A-K, Karle K, Monani UR, Sendtner M (2003) Smn, the spinal muscular atrophy-determining gene product, modulates axon growth and localization of beta-actin mRNA in growth cones of motoneurons. J Cell Biol 163:801–812
Shababi M, Habibi J, Yang HT, Vale SM, Sewell WA, Lorson CL (2010) Cardiac defects contribute to the pathology of spinal muscular atrophy models. Hum Mol Genet 19:4059–4071
Shafey D, Côté PD, Kothary R (2005) Hypomorphic Smn knockdown C2C12 myoblasts reveal intrinsic defects in myoblast fusion and myotube morphology. Exp Cell Res 311:49–61
Sharma MC, Goebel HH (2005) Protein aggregate myopathies. Neurol India 53:273–279
Sharma A, Lambrechts A, Hao LT, Le TT, Sewry CA, Ampe C, Burghes AHM, Morris GE (2005) A role for complexes of survival of motor neurons (SMN) protein with gemins and profilin in neurite-like cytoplasmic extensions of cultured nerve cells. Exp Cell Res 309:185–197
Simone C, Ramirez A, Bucchia M, Rinchetti P, Rideout H, Papadimitriou D, Re DB, Corti S (2016) Is spinal muscular atrophy a disease of the motor neurons only: pathogenesis and therapeutic implications? Cell Mol Life Sci 73:1003–1020
Singh RN, Howell MD, Ottesen EW, Singh NN (2017) Diverse role of survival motor neuron protein. Biochim Biophys Acta - Gene Regul Mech 1860:299–315
Sjöström M, Squire JM (1977) Fine structure of the A-band in cryo-sections. The structure of the A-band of human skeletal muscle fibres from ultra-thin cryo-sections negatively stained. J Mol Biol 109:49–68
Somers E, Lees RD, Hoban K, Sleigh JN, Zhou H, Muntoni F, Talbot K, Gillingwater TH, Parson SH (2016) Vascular defects and spinal cord hypoxia in spinal muscular atrophy. Ann Neurol 79:217–230
Suarez C, Carroll RT, Burke TA, Christensen JR, Bestul AJ, Sees JA, James ML, Sirotkin V, Kovar DR (2015) Profilin regulates F-actin network homeostasis by favoring formin over Arp2/3 complex. Dev Cell 32:43–53
Sumner CJ (2007) Molecular mechanisms of spinal muscular atrophy. J Child Neurol 22:979–989
Szunyogova E, Zhou H, Maxwell GK, Powis RA, Francesco M, Gillingwater TH, Parson SH (2016) Survival motor neuron (SMN) protein is required for normal mouse liver development. Sci Rep 6:34635
Tapia O, Bengoechea R, Palanca A, Arteaga R, Val-Bernal JF, Tizzano EF, Berciano MT, Lafarga M (2012) Reorganization of Cajal bodies and nucleolar targeting of coilin in motor neurons of type I spinal muscular atrophy. Histochem Cell Biol 137:657–667
Tapia O, Narcís JO, Riancho J, Tarabal O, Piedrafita L, Calderó J, Berciano MT, Lafarga M (2017) Cellular bases of the RNA metabolism dysfunction in motor neurons of a murine model of spinal muscular atrophy: role of Cajal bodies and the nucleolus. Neurobiol Dis 108:83–99
Tisdale S, Pellizzoni L (2015) Disease mechanisms and therapeutic approaches in spinal muscular atrophy. J Neurosci 35:8691–8700
Vitte J, Fassier C, Tiziano FD, Dalard C, Soave S, Roblot N, Brahe C, Saugier-Veber P, Bonnefont JP, Melki J (2007) Refined characterization of the expression and stability of the SMN gene products. Am J Pathol 171:1269–1280
Walker MP, Rajendra TK, Saieva L, Fuentes JL, Pellizzoni L, Matera AG (2008) SMN complex localizes to the sarcomeric Z-disc and is a proteolytic target of calpain. Hum Mol Genet 17:3399–3410
Zhang R, So BR, Li P, Yong J, Glisovic T, Wan L, Dreyfuss G (2011) Structure of a key intermediate of the SMN complex reveals Gemin2’s crucial function in snRNP assembly. Cell 146:384–395
Acknowledgments
The authors wish to thank Raquel García-Ceballos for technical assistance.
Funding
This work was supported by the following grants: “Centro de Investigación Biomédica en Red sobre Enfermedades Neurodegenerativas” (CIBERNED; CB06/05/0037), Spain, and “Instituto de Investigación Valdecilla” (IDIVAL, Next-Val NVAL17/22), Spain.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The authors declare that they have no conflict of interest. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Berciano, M.T., Castillo-Iglesias, M.S., Val-Bernal, J.F. et al. Mislocalization of SMN from the I-band and M-band in human skeletal myofibers in spinal muscular atrophy associates with primary structural alterations of the sarcomere. Cell Tissue Res 381, 461–478 (2020). https://doi.org/10.1007/s00441-020-03236-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-020-03236-3