Abstract
Among the 26 human type II keratins, K78 is the only one that has not yet been explored with regard to its expression characteristics. Here, we show that, at both the transcriptional and translational levels, K78 is strongly expressed in the basal and parabasal cell layers with decreasing intensity in the lower suprabasal cells of keratinising and non-keratinising squamous epithelia and keratinocyte cultures. The same pattern has been detected at the transcriptional level in the corresponding mouse epithelia. Murine K78 protein, which contains an extraordinary large extension of its tail domain, which is unique among all known keratins, is not detectable by the antibody used. Concomitant studies in human epithelia have confirmed K78 co-expression with the classical basal keratins K5 and K14. Similarly, K78 co-expression with the differentiation-related type I keratins K10 (epidermis) and K13 (non-keratinising epithelia) occurs in the parabasal cell layer, whereas that of the corresponding type II keratins K1 (epidermis) and K4 (non-keratinising epithelia) unequivocally starts subsequent to the respective type I keratins. Our data concerning K78 expression modify the classical concept of keratin pair K5/K14 representing the basal compartment and keratin pairs K1/K10 or K4/K13 defining the differentiating compartment of stratified epithelia. Moreover, the K78 expression pattern and the decoupled K1/K10 and K4/K13 expression define the existence of a hitherto unperceived early differentiation stage in the parabasal layer characterized by K78/K10 or K78/K13 expression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fifty four keratins are encoded by the type I and type II keratin multi-gene families on human chromosomes 17q21.2 and 12q13.13, respectively (The Human Genome Sequencing Consortium 2004; Schweizer et al. 2006). Studies carried out in our and other laboratories during the last 20 years have led to the characterisation of the complex expression and interaction patterns of most human keratins. Currently, three human keratins, the type II keratin K78 (previously designated Kb40 or K5b; Hesse et al. 2001, 2004) and the type I keratins K23 and K24 (previously Ka23 and Ka24, respectively; Hesse et al. 2004) have only been characterised at the gene/cDNA level, whereas detailed expression studies of the respective mRNAs and proteins in human epithelia are still missing. The present study explores the expression profile of the last type II keratin, K78, in order to pave the way for further investigations into its function.
At present, only sporadic, mostly uncommented data from various gene array analyses exist regarding the expression of K78 in human or murine epithelia. The complete human K78 cDNA was originally isolated from a scalp skin cDNA library and strong K78 mRNA expression was found in human tongue epithelium (Rogers et al. 2005). Up-regulation of K78 has been observed during wound re-epithelialisation in human skin (Nuutila 2013), in various ichthyoses (Rice et al. 2013), in skin arrays of Helicobacter-infected mice (Toyoda et al. 2013) or eosinophilic oesophagitis (Kiran et al. 2015). Moreover, reverse transcription plus the polymerase chain reaction (RT-PCR) analyses suggest the presence of K78 transcripts in corneal epithelium, although the respective protein has not been demonstrated there (Lu et al. 2006). Expressed sequence tag (EST) databases indicate the strongest K78 expression in oral mucosa, oesophagus, larynx, pharynx and tumours thereof but also in the eye and heart (NCBI: EST-, GEO-profiles).
Down-regulation of K78 has been reported in oral squamous cell carcinomas (Chen et al. 2008; Sakamoto et al. 2011; Bundela et al. 2014) and p53-mutated carcinomas (Garritano et al. 2013) and upon alterations of the p63 pathway in the epidermis of Satb1-KO mice (Fessing et al. 2011) and during skin wounding in humans (Nuutila 2013) and mice (Ansell 2012). Moreover, K78 down-regulation has been observed in the pancreas after hypothalamic lesion (Kiba et al. 2010).
Here, we compare human K78 with homologues in other species and perform a comprehensive analysis of its expression relative to a set of other constitutively expressed keratins that are commonly used as keratinocyte differentiation markers. Our study shows the expression of K78 in non-cornifying and cornifying stratified human epithelia in which K78 transcript and protein occur both in basal and in parabasal cell compartments and are clearly maintained, albeit with gradually decreasing intensity, in the mid-suprabasal compartments. These data are in contrast to the widely accepted view of an abrupt switch from basal versus suprabasal keratin expression in stratified epithelia and suggest keratin patterns and the resulting pairing possibilities are definitely more complex than conventionally assumed.
Materials and methods
Tissues and cultured cells
Our study included human adult (i.e., scalp, forearm, footsole) and 14th week of pregnancy (WP14) skin (i.e., palm, scalp, back), various internal epithelia (i.e., dorsal tongue, oesophagus, vagina, duodenum) and one case of a well-differentiated oral squamous cell carcinoma. Tissues were obtained either during surgery for medical reasons or from cadavers during pathological investigations (National Tissue Collection, University of Heidelberg, Germany) under institutional approval and included adherence to the Declaration of Helsinki (Ethics Committee, Medical Faculty of the University of Heidelberg, vote: S-221-2012 to L.L.). Keratinising and non-keratinising mouse epithelia were from adult wildtype C57BL/6 mice. Animals were killed in accordance with the UK Animals Scientific Procedures Act, 1986. Immediately after excision, tissues were snap-frozen in isopentane pre-cooled in liquid nitrogen and stored at -70 °C (for details, see Langbein et al. 2004). Cultures of normal human epidermal keratinocytes (NHEK) and immortalised keratinocytes of the human line HaCaT (Boukamp et al. 1988) were also investigated. Cells were freshly seeded, grown to sub-confluency and confluency with focal onset of stratification.
Primary antibodies used for indirect immunofluorescence (IIF) or Western blots and their respective dilutions are given in Table 1. In cases of double-labelling, the two appropriately diluted primary antibodies were mixed before incubation. Several established antibodies against distinct keratins (see Table 1) were used to control and verify unexpected staining results (not shown).
Secondary antibodies
IgG or IgG + IgM used for IIF were: goat anti-guinea pig, goat anti-mouse, coupled to Cy3 (1:500, red fluorescence) or Alexa 488 (1:200, green fluorescence; all Molecular Probes). In cases of double-labelling, appropriately diluted secondary antibodies of the respective species were mixed before incubation. For enhanced chemoluminescence (ECL) detection, horseradish-peroxidase-coupled rabbit anti-guinea pig IgG (H + L) were used (Dianova, 1:10,000).
IIF microscopy
Cryosections were fixed in acetone (-20 °C; 10 min), permeabilised with TRIS-buffered saline plus Triton-X100 (TBST: TBS + 0.001 % Triton-X100) and blocked with 5 % normal goat serum in phosphate-buffered saline (PBS). Sections were incubated with primary antibodies for 1 h and, after being rinsed in PBS, incubated with secondary antibodies (dilution of 1:200-500) for 30 min. After being washed in PBS, sections were dried and mounted. Visualisation and documentation were performed with a photomicroscope (Axiophot equipped with AxioCam camera and AxioVision software; Carl Zeiss, Oberkochen, Germany; for details, see Langbein et al. 2004, 2005) and, in cases of cultured keratinocytes, as confocal images with a laser scanning microscope (Zeiss LSM700; ZEN software; 488 nm, 405 nm lasers; all the microscope settings, such as laser power, gain, offset, 1 AU pinhole and others, were kept unchanged for all samples with the same staining combination). Exclusion of the specific primary antibody or its replacement with normal serum of untreated animals of the same species or with PBS was used as a negative control and also for secondary antibody background evaluation. DNA counterstaining of nuclei was performed with DAPI (4,6-diamidino-2-phenylindole).
Tissue extraction, gel electrophoresis and Western blots
Tissue extracts were prepared from cryosections and resolved by sodium dodecylsulfate-polyacrylamide gel electrophoresis (SDS-PAGE; 10 % polyacrylamide; Langbein et al. 2004, 2005). For Western blots, gels were transferred to polyvinylidene difluoride membranes (Immobilon-P, Millipore, Eschborn, Germany) by wet blotting. After steps involving staining (0.1 % Coomassie blue R250), destaining and blocking with 5 % non-fat milk powder in TRIS-buffered saline, membranes were incubated with the respective primary and secondary antibodies (see above; for further details, see Langbein et al. 2004, 2005).
RNA isolation and quantitative RT-PCR analyses
Total RNA was isolated from fresh cryosections by using TRIzol Reagent (Invitrogen, Carlsbad, Calif., USA) by homogenisation (FastPrep FP120 Homogenizer, ThermoSavant). After 5 min of incubation (room temperature), chloroform was added and the preparation was further incubated (3 min, room temperature). Following centrifugation of the supernatant, the RNA was precipitated and washed with 75 % ethanol. The dried pellet was dissolved in RNase-free water. Total RNA (1 μg) was reverse-transcribed (ReverseAid H Minus First Strand cDNA Synthesis Kit: Thermo SCIENTIFIC Rockford, Ill., USA) and quantitative RT-PCR (qRT-PCR) was performed in a LightCycler 480 II (Roche) with the LightCycler 480 Probes Master mix, forward and reverse primers and the UPL-probe (Universal ProbeLibrary; Roche Applied Science, Mannheim, Germany). After preincubation at 95 °C for 10 min, 45 amplification cycles with denaturation (95 °C, 10s) and annealing (60 °C, 30s) were applied. Fluorescence (533 nm) was monitored by LightCycler 480 software 1.5 after each cycle. PCR was stopped by cooling to 40 °C.
For quantification, housekeeping genes (human: PBD-G; mouse: b2-microglobulin, respectively) were analysed in parallel. The ratio of relative mRNA expression of control versus sample was normalised to the housekeeping gene. For verification of the quality of primers/probes combination and the efficiency of the PCR, standard curves were performed for each test. As negative controls, water instead of cDNA was used. Primers (forward, f-; reverse, r-) and, where applicable, UPL-probes were used: human hKRT78 (f-agctggaggctctgagagagta, r-atggacagcaccacagacg), hPBD-G (f-agccacatcgctcagacac, r-gcccaatacgaccaaatcc), mouse mKRT78 (f-aggcaacctggagtctctga, r-ctagtccgcgtgattctcctc), and mb2-microglobulin (f-tgccagagaagagtgtggtg, r-agccgggtgttgaggttt). Because of the limited access to human tissue samples, only technical replicates (n = 3) could be made.
In situ hybridisation
A 218-bp PCR product (annealing temperature 58 °C; forward: attctgggctctgggaaggacc; reverse: gggcagcggacacggagtt) of the 3’-non-coding region of the human keratins KRT78 (for details, see Rogers et al. 2005), KRT14, KRT5, KRT10 and KRT1 (see Langbein et al. 2004) were used to prepare 35S-labelled riboprobes. In situ hybridisation (ISH) on cryostat sections was carried out as described previously in detail (Langbein et al. 2004). Briefly, following overnight hybridisation at 42 °C, sections were repeatedly washed with SSC (standard sodium citrate)/formamide at 50 °C, digested with RNaseA at 37 °C and washed with SSC/formamide/dithiothreitole at 50 °C. After being dipped in photo-emulsion (NTB-2; Kodak), sections were exposed for 2 to 3 days and stained with haematoxylin. Regular internal positive controls were KRT14 and KRT5 antisense probes from the 3′-non-coding region and negative controls were the respective sense probes, hybridisation buffer instead of the probe or RNAse digestion before hybridisation (not shown). For the recording of the ISH signals by reflection microscopy, a ZEISS CLSM 510 Meta was used, which allowed the simultaneous visualisation of ISH in epi-illumination for the detection of reflection signals (red channel) and transmitted light in bright field for haematoxylin staining (green channel) and the combination of the two signal channels by an overlay by using ZEISS LSM Imaging Browser or ZEN software (for further details, see Langbein et al. 2004).
For verification and extended analyses, non-radioactive ISH was carried out with digoxigenin (Dig)-labelled Locked Nucleic Acid (LNA)-enhanced oligonucleotides of human KRT78 (5DigN-ATACCAATAGAACAGCTCTGGA-3DigN) and mouse Krt78 (5DigN-TCTTCCGACTTGATTCCACTGTCTT-3DigN; Exiqon, Vedbaek, Denmark). Both were used at a hybridisation temperature of 54 °C, following the instructions of the manufacturer. As a positive control, we used a Dig-labelled β-actin LNA mRNA ISH probe (Exiqon) and, for negative probes, we employed a Dig-labelled scramble LNA mRNA ISH probe (Exiqon) or hybridisation buffer instead of a specific probe. Signals were detected by using an anti-Dig antibody labelled with alkaline phosphatase and BCIP (5-bromo-4-chloro-3-indolyl-phosphate) as a substrate for the visualisation of enzyme activity, largely following the instructions of the manufacturer (Exiqon).
Results
Gene locus, evolution and sequence characteristics of K78
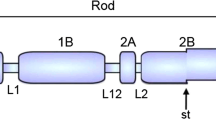
The KRT78 gene is located between the KRT8 and KRT79 genes at the telomeric end of the type II keratin gene cluster on human chromosome 12q13.13 (Fig. 1a; for the complete type II cluster, see Supplementary Fig. S1a). Although KRT8 is expressed in simple epithelia, KRT79 Footnote 1 to KRT2 are typical for suprabasal layers of various stratified epithelia (for a review, see Moll et al. 2008) and KRT73 to KRT71 are typical for defined layers of the hair follicle (Langbein et al. 2006; for a review, see Langbein and Schweizer 2013). Human KRT78 encodes a protein of 520 amino acids (Fig. 1b) with a calculated molecular weight of 56.8 kDa and an isoelectric point of 5.9 (Rogers et al. 2005).
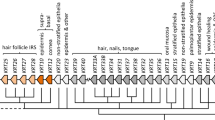
Analysis of the human K78 gene. a Location of the KRT78 gene within the type II keratin gene domain on chromosome 12q13.13. The scheme is incomplete at the centromeric end (right; see also Supplementary Fig. S1a). Note the atypical location of the reversely oriented type I keratin gene KRT18 at the telomeric end of the type II keratin gene domain. b Amino acid sequence of K78 (bold letters α-helical rod domain, blue glycine–rich [GGX] motifs, red cysteine residues, green proline residues). c Representation of the phylogeny (left) of discussed species and corresponding structures of K78 proteins (right). Major events in the evolution of K78 are indicated
To determine the degree of evolutionary conservation of KRT78 and its molecular features, we derived amino acid sequences of KRT78 gene homologues from the GenBank database and compared the sequences of the predicted proteins. K78 is present in a broad range of mammalian species including the Tasmanian devil, a representative of marsupials (Fig. 1c; Supplementary Fig. S2a). Because of the incompleteness of the genome sequence of monotreme mammals (platypus) and because of the ambiguity about the orthology relationship of mammalian and non-mammalian keratin gene clusters (Vandebergh and Bossuyt 2012), the evolutionary origin of KRT78 (presumably by duplication from a more ancestral gene and subsequent accumulation of mutations) could not be mapped precisely on the phylogenetic tree of vertebrates. As both marsupials and placental mammals have retained KRT78, the minimum evolutionary age of KRT78 equals the duration of their separate evolution of approximately 170 million years (Kumar and Hedges 1998).
Interestingly, the murine K78 comprises a large insert in the carboxy-terminal tail region, which is absent in human K78 and its homologues of evolutionary more distant species. This insert is encoded by a stretch of 51 repeats of a segment of 33 nucleotides within the last exon of Krt78. Consequently, the length of the tail domain (following the LEGE/LESE motif at the end of the rod domain) of murine K78 is increased to 654 amino acid residues relative to 100 residues in human K78 (Supplementary Fig. S2b). By comparing the K78 tail sequences of mammals with known phylogenetic relationships, we could map the peculiar extension of the tail domain to the last common ancestor of the clade Muroidea (comprising more than 1300 species, including mice, rats, hamsters and blind mole rats), which diverged from other rodents between 55 and 60 million years ago (Fig. 1c; Huchon et al. 2007). The peculiar insert shows considerable variation in the sequence of the repeat unit while having been conserved in mouse, rat, hamster and blind mole rat (Fig. 1c).
The 51-copy-repeat structure is composed of three distinct but nonetheless highly conserved, 11-residue motifs (Supplementary Fig. S2b, c). Searches of the structure and sequence databases have revealed no close matches (except for a “Sporozoite p67 surface antigen; pfam05642”, NCBI, with unknown function) to these repeats and, thus, little can presently be ascertained as to their likely structure or function. Nonetheless, each 11-residue quasi-repeat can be expected to have the same tertiary conformation based on a conserved β-bend and these structural elements can be expected to be helically related to one another (Crane 1950; Fraser and Parry 2014). Consequently, the overall conformation is predicted to be rod-like with the key cysteine, serine and tyrosine residues being periodically distributed along the length and available for interaction.
Within a rod domain-based tree of the human type II keratins, K78 appears as a single-branched keratin, adjacent to K80-K86 (Supplementary Fig. S1b) whose genes are, however, located far away from the KRT78 gene at the centromeric end of this domain (Supplementary Fig. S1a). Both K78 and K80 possess both relatively high numbers of cysteine and proline residues and few (K78) or no (K80) GGG or GGX repeats in their non-α-helical head and tail domains (Fig. 1b, in red, green and blue respectively), i.e., properties that are clearly intermediate between those of epithelial and hair keratins (Langbein et al. 2010; for cysteine residue function in K14 and K5, see Feng and Coulombe 2015). In contrast, because of the repetitive sequences in the tail domain of the murine K78, its carboxyterminal GGX motif frequency is considerably increased (Supplementary Fig. S2c). GGG/X motifs are thought to be involved in regulatory interactions (Omary et al. 2006; Thandapani et al. 2013), whereas cysteine residues facilitate intra- and intermolecular covalent crosslinking (Feng and Coulombe 2015).
Expression of human K78 as determined by ISH and IIF
mRNA and ISH
The expression pattern of K78 transcripts was determined by ISH with antisense probes annealing to the 3’ untranslated region of the human K78 mRNA. Negative controls with sense probes of the same sequence or with hybridisation buffer alone gave no signals (not shown). Probes of other keratins (e.g., K14, K5, K10, K1; see below) showed the expected signals thereby confirming the specificity of the method and the probes. Both non-radioactive (Fig. 2a-i) and radioactive (Supplementary Fig. S3) ISH clearly revealed K78 transcripts in the basal cell layer of the oesophageal epithelium. A similar labelling intensity was observed in para- and lower suprabasal layers but gradually decreased and finally disappeared in mid-suprabasal cells (Fig. 2a; Supplementary Fig. S3a, a’). In contrast, K14 transcripts were restricted to the basal cell layer (Fig. 2b, Supplementary Fig. S3b). In addition, transcripts of both K78 and K14 also appeared in the stromal invaginations from the lamina propria (stromal papillae) labelling both basal and lower suprabasal cells (Fig. 2a, b, arrows). Comparable results were obtained in interpapillary dorsal tongue epithelium (Fig. 2c for K78, Fig. 2d for K14, Supplementary Fig. S4c, c’ for K78, Supplementary Fig. S4d for K14), interfollicular scalp epidermis (Fig. 2e, Supplementary Fig. S3e for K78, Fig. 2f, Supplementary Fig. S3f for K14) and the infundibulum (Fig. 2e for K78, Fig. 2f for K14). K78 and K14 transcripts were also present in the entire outer root sheath (ORS) of the hair follicles (Fig. 2e, g, Supplementary Fig. S3g for K78, Fig. 2f, Supplementary Fig. S3h for K14). K78 transcripts were also detectable in the non-differentiated cells of sebaceous glands (Fig. 2h) and in the peripheral duct cells of the eccrine sweat glands (Fig. 2i). In the stratified epithelium of the facial (Supplementary Fig. S3j for K78, Supplementary Fig. S3k for K14) and plantar (Supplementary Fig. S3l for K78, Supplementary Fig. S3m for K14) epidermis, the K78 mRNA expression pattern was essentially identical to that described above for the oesophageal epithelium. Transcripts of K5, e.g., on scalp (Supplementary Fig. S3i), were also concentrated in the basal layer and widely comparable with that of K14.
Expression pattern of keratin K78 in human epithelia. a-i Non-radioactive in situ hybridisation (ISH): strong labelling of K78 mRNA in basal and parabasal layers and, with decreasing intensity, in early suprabasal layers of the oesophagus (a, arrows positive stromal papillae), tongue (c), scalp (e, ORS outer root sheath of hair follicle [see also g], inf infundibulum, ife interfollicular scalp epidermis), sebaceous glands (h, sg) and peripheral cells of middle/ upper eccrine sweat gland ducts (i, asterisks negative luminal duct cells). K14 mRNA is restricted to the basal layer of oesophagus (b, arrows positive mucous gland ducts), tongue (d) and interfollicular scalp epidermis (f, ife). In the lower infundibulum (f, inf) and the upper ORS (f), early suprabasal cells were also labelled. For corresponding radioactive ISH studies, see Supplementary Fig. S3. j-p’ Indirect immunofluorescence (IIF): the K78 antibody mirrors the ISH data showing strong protein labelling in basal, parabasal and lower suprabasal cell layers and considerably lower staining in the mid-/upper suprabasal layers of the squamous non-keratinising epithelia: oesophagus (j), vagina (k), tongue with filiform papillae (l, fp) and interpapillary epithelium (l, ip) and keratinising epithelia: scalp (m, ife, inf), scalp sebaceous glands (m’, sg), scalp peripheral eccrine sweat gland duct (m’’, gd gland duct with positive luminal and peripheral cells, ud upper duct, single asterisks positive peripheral but negative luminal cells, me myoepithelial cells, double asterisks negative secretory gland cells; for a summary, see Supplementary Fig. S6), hair follicle ORS (n) and plantar skin (o). Non-stratified “simple” epithelia (e.g., ileum) were K78 negative (p) and K8 positive (p’). q Western blots of scalp extracts. Both K78 and K14 antibodies labelled single bands at the respective molecular weights (Co Coomassie-Blue-G-stained Western blot membrane). j–o Nuclear staining with DAPI (blue). Bars 20 μm (m’, n, p, p’), 50 μm (g, i–k, m, o), 100 μm (h, l, m’’), 200 μm (a–e)
In the mouse, K78 transcripts were detected by ISH in the basal layer and, with decreasing intensity, in the para- and lower suprabasal layers of the oesophageal epithelium (Supplementary Fig. S4a) and of the tongue (Supplementary Fig. S4b). qRT-PCR screening of murine tissues showed predominant K78 mRNA expression in the non-keratinising stratified epithelia of the tongue, oesophagus, forestomach and vagina and distinctly lower amounts of K78 transcripts in the various skin types, whereas all simple epithelia were K78-negative (Supplementary Fig. S4c). Thus, these data indicated the conserved K78 mRNA expression in the basal and lower suprabasal layers of non-cornifying stratified epithelia of human and mouse and the K78 expression in the equivalent cell layers of the human epidermis. Furthermore, the strongest Northern signal in the human tongue (Rogers et al. 2005) was clearly confirmed by our K78 qRT-PCR data, which also showed the highest values in mouse tongue but were still high in vagina, oesophagus and forestomach. This indicated that the higher stratified squamous epithelia contained higher amounts of K78 transcripts compared with the less stratified skin samples. In principle, these values were also revealed by the ISH images (e.g., human: Supplementary Fig. S3; mouse: Supplementary Fig. S4a, b) similarly to and quantitatively confirmed by qRT-PCR (Supplementary Fig. S4c) of mouse tissues.
Protein, IIF and Western blot
Despite many years of experience and numerous attempts, we failed to raise specific antibodies against N- or C-terminal peptides of K78. Screening of a plethora of commercially available “K78 antibodies” suggested that most of them lacked specificity according to stringent criteria (see Griffiths and Lucocq 2014). Finally, we selected a rabbit antiserum raised against a peptide from the head domain of human K78 (amino acids 12-61; Supplementary Fig. S2b, underlined; Abcam, ab179546Footnote 2). On probing an extract of scalp skin, this K78 antiserum revealed a single band at the predicted molecular weight of ~57 kDa by Western blot analysis, whereas the K14-specific guinea pig antiserum (see Table 1) showed a single band at ~52 kDa (Fig. 2q), although not clearly detectable as a sharp band in the Coomassie-Blue-G-stained blotted membranes (Fig. 2q, Co). Since this K78 antibody did not react with murine K78 (data not shown; for a comparison of the respective human and mouse sequence, see Supplementary Fig. S2b, underlined), our subsequent studies were limited to human tissues.
The expression patterns revealed by the K78 antibody faithfully mirrored the ISH data and demonstrated strong K78 protein labelling in the basal and parabasal/lower suprabasal cell layers and considerably weaker staining in the mid/upper suprabasal layers of the non-keratinising oesophageal (Fig. 2j) and vaginal (Fig. 2k) epithelium. Likewise, in the partially keratinised dorsal tongue epithelium (Fig. 2l), in the scalp (Fig. 2m) and in the plantar epidermis (Fig. 2o), K78 staining was clearly seen in a gradient from the basal to lower suprabasal layers. Obviously and in agreement with the ISH data, the extension of K78 expression into the suprabasal compartments was distinctly more pronounced in non-keratinising epithelia, as compared with the keratinising epithelia and the epidermis (Fig. 2j,k).
Moreover, in scalp sections, K78 was present both in the interfollicular epidermis (Fig. 2m) and the infundibulum (Fig. 2m) including the entire ORS of the hair follicles (Fig. 2n) and in the sebocytes (Fig. 2m’). In sweat glands, both peripheral and luminal cells of the lower (glandular) duct (Fig. 2m’’, gd) were positive. Remarkably, peripheral cells of the middle/upper duct were positive (Fig. 2m’’, ud), whereas the luminal cells were negative (Fig. 2m’, asterisks). Throughout, K78 was undetectable by IIF in simple epithelia as shown representatively for the ileum (Fig. 2p), which in turn was positive for K8 (Fig. 2p’).
This finding was supported by qRT-PCR, which showed that K78 mRNA was more abundant by at least three orders of magnitude in the highly stratified epithelia of the human tongue and foot sole than in the non-stratified epithelium of the duodenum (not shown) and in mouse (Supplementary Fig. S4c).
Coexpression of K78 with classical epithelial keratins
The peculiar expression profile of K78 in the basal and lower suprabasal layers of stratified epithelia, which (to the best of our knowledge) is a pattern hitherto unprecedented in the literature on keratins, raised the question as to the respective type I partner(s) for the hetero-polymerisation of K78 into stable intermediate filaments (IFs). Independent of the type of squamous epithelia, the most obvious candidate is K14, i.e., the type I member of the classical basal keratin pair K5/K14 (for a review, see Moll et al. 2008). In accordance with the ISH data (for K14: Fig. 2b, d, f, Supplementary Fig. S3b, d, f, h, k, m), double-label IIF with K14 antibodies clearly revealed strong K14 (green)/K78 (red) co-expression (merged yellow) in the basal cell layer of oesophagus (Fig. 3a), tongue (Fig. 3b) and vaginal (Fig. 3c and insert) epithelia and in the strongly keratinising palmar (Fig. 3d) and scalp (Fig. 3e) epidermis. Notably and in agreement with the ISH results, K78 was also detected in the lower suprabasal layers in which K14 signals were absent or only focally but more weakly detected in the lowermost suprabasal layers (Fig. 3e). Again confirming the ISH data, ubiquitous K14 (green)/K78 (red) co-expression was seen over the entire ORS epithelium (Fig. 3f) and was best visible for K78 by double-staining with the companion layer-specific K75 (Fig. 3h, green; Winter et al. 1998). Co-staining of K78 and K14 was also seen in the sebaceous gland (Fig. 3g).
Expression analysis of K78 relative to the classical basal keratins K14 and K5 by double-label IIF. Double-labelling of K78 (red) and K14 (a–g, green) or K5 (a’-e’, green) in squamous epithelia of oesophagus (a), tongue (b, ip interpapillary region, fp filiform papillae) and vagina (c) and in palmar (d) and scalp (e) epidermis shows that K14 and K5 are most prominent in the basal layer and decrease often in variable degree in the lower suprabasal layers, whereas K78 clearly extends into the suprabasal layers (yellow merged label and red staining, respectively). Coexpression with K14 is also seen in the ORS (f) and sebaceous glands (g, sg). In the hair follicle (h), the K75-positive/companion layer (green) is next to the K78-positive ORS (red). Nuclei are stained (blue) with DAPI. Bars 20 μm (a, a’, c–e, c’–e’, f–h), 50 μm (b, b’
We also analysed K78 expression relative to that of K5, i.e., the classical basal type II keratin and partner of K14. In agreement with the ISH data for K5 transcripts (Supplementary Fig. S3i), IIF labelling of non-keratinising squamous epithelia showed strong K5 expression (green) in the basal cell layer and, similar to K14, focally and weakly in the lower suprabasal compartment, where it co-localised with K78 (red, merged yellow) in the oesophagus (Fig. 3a’), tongue (Fig. 3b’) and vagina (Fig. 3c’). The same held true for the keratinising palmar (Fig. 3d’) and scalp (Fig. 3e’) epidermis.
Interestingly, the comparison of K78 expression with that of the respective differentiation-specific keratins K4/K13 (non-keratinising squamous epithelia) showed that, in the oesophagus (Fig. 4a, a’) and tongue (Fig. 4b, b’, and inserts) epithelium, the suprabasal K13 expression (green) clearly started in the parabasal cells, leading to a locally limited coexpression with suprabasal K78 (red in Fig. 4a, b and, in insert, merged yellow). Surprisingly, the corresponding studies for the respective type II keratin K4 revealed a different pattern. In both oesophagus (Fig. 4a’) and tongue (Fig. 4b’ and insert) epithelium, K4 (type II) expression started later than that of K13 (type I) and virtually spared the typical para- to lower suprabasal zone of temporary K13/K78 co-expression. This finding could be confirmed by K13 (red)/K4 (green) double-labelling of tongue and tongue epithelium (Fig. 4c, d).
Expression analysis of K78 relative to the classical suprabasal keratins K13/K4 and K10/K1 by double-label IIF. Double-labelling of K78 (red)/K13 (green) and K78 (red)/K4 (green), respectively, in squamous epithelia of oesophagus (a, a’, c, i) and tongue (b, b’, d, j, ip interpapillary region, fp filiform papillae) shows that K13 expression starts in the parabasal layer (a, b), whereas that of K4 (green) begins a few layers later (a’, b’) yielding a broad red-stained K78 area (cf. inserts in b, b’). The delay of expression between K13 and K4 is readily seen by double-labelling of these keratins in the oesophagus (c) and tongue (d). Similar results are obtained by double-labelling of K78 (red)/K10 (green) and K78 (red)/K1 (green), respectively, in epidermal tissues of scalp (e, e’) and palmar (f, f’) skin. The delay of expression between K10 (red) and K1 (green) is also readily seen in scalp (g) and palmar (h) skin by using double-labelling of these keratins. Note that single wedge-shaped cells in the basal layer are K10-positive (g, arrowheads). Remarkably, even in non-keratinised squamous epithelia, keratins K10 (red) and K1 (green) are occasionally seen alone or together (merged yellow) in single cells or cell groups (i, j). In eccrine sweat glands, K78 is expressed in both myoepithelial cells (k, me; asterisks secretory gland cells are negative) and middle/upper peripheral duct cells (k, md), whereas K10 occurs in K78-negative luminal duct cells. Nuclei are stained (blue) with DAPI. Bars 20 μm ( a, a’, c, e, e’, g, f’, h, k), 50 μm (b, b’, d, f, i, j)
In accordance with ISH data from plantar and scalp epidermis by using either K10-specific (Supplementary Fig. S5a-d) or K1-specific (Supplementary Fig. S5e-f’) probes, comparable results were obtained for the combination of antibodies against K10 (green)/K78 (red) and K1 (green)/K78 (red) in keratinising epithelia such as in the scalp epidermis (Fig. 4e, e’) and the much thicker palmar epidermis (Fig. 4f, f’) in which K78/K10 co-expression in the parabasal cells was evident (Fig. 4f, merged yellow). In analogy to the delayed onset of K4 expression relative to that of K13 in the oesophagus (Fig. 4a, a’) and the tongue (Fig. 4b, b’), the expression of K1 (type II) started later than that of K10 (type I) in scalp and plantar epidermis (cf. Figs. 4f, f’). This finding was confirmed by K10 (red)/K1 (green) double-labelling (Fig. 4g, h). To strengthen these data and to avoid possible epitope artefacts, we used several K1 and K10 antibodies (Table 1, see also Byrne et al. 1994), all yielding comparable results. Regularly, single wedge-shaped cells in the basal layer were also K10-positive (Fig. 4g, arrowheads) indicating that these cells are just leaving the basal compartment, are committed to the start of differentiation and are “on transit” into the first differentiated parabasal compartment.
Surprisingly, a subset of suprabasal cells of the non-keratinising epithelia of the oesophagus (Fig. 4i) and the tongue (Fig. 4j) express either K10 (green) or K1 (red), with a few of these cells co-expressing (merged yellow) both keratins indicating a “decoupled” expression of both keratins in these epithelia.
In eccrine sweat glands, K78 (red) was detectable in the peripheral cells of the middle/upper duct (Fig. 4f, k, md) and in glandular myoepithelial cells (Fig. 4k, me), whereas K10 was present only in luminal duct cells (Fig. 4k, green). Moreover, K78 expression (see also Fig. 2m’’) further supported our concept of a “vertical” differentiation of the sweat gland duct cells along the duct axis (Langbein et al. 2005; for summary of keratins in sweat glands and ducts, see Supplementary Fig. S6).
Expression of K78 in cultured keratinocytes, during embryogenesis and in squamous cell carcinoma
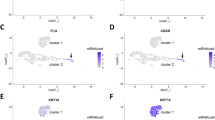
To allow for comparisons with the expression patterns of other keratins, we extended our K78 immunolocalisation studies to keratinocyte cultures in vitro, embryonic skin and tumour samples. In subconfluent (Fig. 5a) and confluent (Fig. 5b, c) cultures of normal human epidermal keratinocytes (NHEK), all cells were positive for K78 (red) and K14 (green; merged yellow in Fig. 5c, c’ and insert) and exhibited a fine fibrillar keratin network (Fig. 5a, c’). In contrast, spontaneously immortalised HaCaT keratinocytes were only focally positive for K78 (Fig. 5d). At confluence, roughly half of the cells expressed K78 (Fig. 5d), whereas all were K14 positive (Fig. 5e, K78 [red]/K14 [green] and merged yellow). Both keratins showed a fine filamentous network (Fig. 5e, insert) attached to desmoplakin-stained desmosomes (Fig. 5f and insert with K78 [red]/dp [green]). The staining pattern (green) was seen as a broad punctuated area of desmosomal contacts from overlapping cellular zones, typical for densely grown keratinocytes. The observed expression pattern of K78 in HaCaT cells was completely independent of cellular proliferation as monitored by K78/Ki-67 double-staining showing randomly scattered K78-positive/negative and Ki67-positive/negative cells (Fig. 5g, K78 [red]/Ki-67 [green]). This suggested that the regulation of K78 expression was different from that of its potential type II partner K14 and indicated the existence of various cell clones with their own differentiation potentials in HaCaT cultures, as also observed during Ca2+-induced differentiation (Micallef et al. 2008, 2010), or were found to be cell-density-dependent (Ryle et al. 1989).
Expression of K78 in cultures of human normal and immortalised keratinocytes, during embryonic development and in a squamous cell carcinoma of the mouth. In subconfluent and confluent cultures, all normal epidermal keratinocytes (NEHK) were positive for K78 (a–c’, red) and K14 (c, c’, green) and showed a fine fibrillar network staining (a, c’). Immortalised keratinocytes of line HaCaT (d-g) expressed K78 (red in d, e) only partially, whereas all cells were positive for K14 (green in e) in a typical IF network attached to desmosomes (f, insert; green desmoplakin; asterisks in insert K78-negative cell) at cellular margins. K78 (red) was independent of cell proliferation (g, green Ki-67). At 14th week of pregnancy, in embryonic scalp (h, i) and palmar skin (j, k), K78 (red) was expressed in basal and lower suprabasal layers leaving the uppermost layer unstained (h, j, arrows). Periderm cells were K78-negative but K4 positive (h, j, green). Double-labelling of K78 (i, k, red) and K10 (i, k, green) revealed parabasal coexpression (i, k, merged yellow) followed by suprabasal staining of K10 (i, k, green). Periderm cells are negative (i, k, asterisks) for both keratins. In a well-differentiated squamous cell carcinoma, peripheral tumour areas with cell nests without squamous maturation were positive for both K78 (l-n, red) and K14 (l, green, merged yellow/orange). Artefactual K14 staining in necrotic residues is indicated (asterisks in k). Central areas with higher squamous maturation were positive for K10 (m, green) and K1 (n, green). Note that, in the serial sections, larger areas were labelled for K10 than for K1 (cf. m, n). Nuclei are stained (blue) with DAPI. Bars 20 μm (a–g), 50 μm (h–k), 100 μm (l–n)
At the 14th week of pregnancy, the human epidermis consists of only three layers covered by the periderm (Holbrook 2006; Holbrook and Odland 1975; Langbein et al. 2013). In accordance with adult epidermis, K78 (red) was detectable in the basal and parabasal layer and also in cells lining incipient hair pegs (Fig. 5h), whereas it was absent from the uppermost epidermal cell layer (Fig. 5h, arrows) and from the K4-expressing periderm (Fig. 5h, green). K10 expression (Fig. 5i, green) was present in the second layer of the epidermis in which it was co-expressed with K78 (red, merged yellow) and in the third layer but was absent from the basal epidermal layer and the periderm cells (Fig. 5i, asterisks). These patterns were also detected in the palmar skin at the same developmental age (Fig. 5j, k).
To explore whether K78 (red) expression might add useful information in the differential diagnosis of epithelial pathologies, we initially analysed a well-differentiated oral squamous cell carcinoma with various degrees of maturation of the tumour cells in individual tumour cell nests. Peripheral tumour areas showed nests of cells without squamous maturation (excluding necrotic parts, see asterisks in Fig. 5l) that were positive for K14 (Fig. 5l, red) and less strongly also for K78 (Fig. 5l, green, merged yellow to orange color). K78 immunoreactivity disappeared in central areas of tumour cell nests showing higher squamous maturation (Fig. 5m, n, green). The basal layer of the tumour was K78-positive and negative for K10, whereas some lower suprabasal areas were positive for both K78 and K10 (Fig. 5m, green). In contrast, no coexpression of K78 and K1 was observed (Fig. 5l, green). These preliminary data on squamous cell carcinoma indicated that the expression pattern of these keratins was similar in stratified epithelial cells of both tumours and normal tissues. K78 labelling of non- and early differentiated cells could be helpful in the refinement of differential histopathological diagnoses in general by providing a more detailed characterisation of the maturation grade of squamous carcinoma cells.
Discussion
Within the framework of our efforts to unveil the expression characteristics of the members of the human keratin family, we recently investigated two type II keratins, K80 and K77, which are both expressed in a differentiation-related manner, either in fully differentiated cells of virtually all types of epithelia (K80; Langbein et al. 2010) or in the human (embryonic) and mouse (embryonic and adult) epidermis and eccrine sweat gland ducts (K77; Langbein et al. 2013). The present study of K78 has now led to the completion of the expression catalogue of the keratin type II subfamily.
Akin to K80 and K77, K78 has also proved to be a constitutively expressed type II keratin in non-keratinising and keratinising human squamous epithelia. However, in contrast to K80 and K77, its expression has turned out to be a very early keratin whose strongest mRNA and protein synthesis occurs in the basal and parabasal cell layers. Its weaker suprabasal expression rapidly vanishes in the epidermis, whereas it is maintained up to the mid/upper cell layers of non-keratinising epithelia. Identical mRNA expression patterns have been found in the non-keratinising squamous epithelia of the mouse.
Further studies aimed at investigating K78 expression in the context of the known keratin expression profiles of both types of epithelia are summarised in Fig. 6. Their interpretation is based on previous studies of keratin expression during early mouse embryogenesis, studies that have provided strong evidence that the prerequisite for type I + type II keratin heterotypic pairing consists in the coincident supply of newly synthesised keratin monomers of opposite types from the respective mRNAs (Lu et al 2005; Lee and Coulombe 2009; Yamada et al. 2002; Bray et al. 2015). Because of a proteolytic sensitivity of keratins in the absence of their binding partner, a mRNA might be synthesised, whereas the encoded protein is degraded (e.g., K19; Vijayaraj et al. 2009). Furthermore, this model is notably focused on keratins that are constitutively expressed at normal keratinocyte differentiation stages. Keratins found only in single cells or in cell groups in the basal cell layer (i.e., K7, K8, K19, K20) and keratins being induced under “activating conditions” in the suprabasal compartment (i.e., K6, K16, K17) have not been considered here but definitely increase the complexity of keratin expression and interactions at least in a focal pattern.
Sequential keratin expression in (a) human epidermis and (b) non-keratinising squamous epithelia. Numbers delineate zones of active mRNA synthesis of major keratins. Grey-shaded areas cover the corresponding zones of demonstrable major keratin protein. Additional focal or single-cell expression of type II keratins K7 and/or K8 and type I keratins K15, K18, K19 and/or K20 in the basal cell layer are indicated (see b #). Fetal (foetal) expression of K77 only occurs during embryonic development (Langbein et al. 2013). Regional (regional) expression of keratin K9 (Langbein et al. 1993) occurs in palmoplantar epidermis. Focally/scattered occurring keratins of proliferative “activated” epithelia, namely K6, K16 and K17, are not shown. Instead of the terms Stratum basale, Stratum spinosum and Stratum granulosum, the usage of the terms basal-, parabasal-, lower-, middle- and upper suprabasal layers allows a finer description of the various complex differentiation stages, whereas to date, the Stratum spinosum uniformly comprises all layers from the parabasal up to the Stratum granulosum. The earliest step of differentiation occurs in the parabasal layer in which the cells are K10-positive (b basal layer [undifferentiated], pb parabasal layer [early differentiated], lsb lower suprabasal layers [advanced differentiated], msb mid-suprabasal layers [late differentiated], usb upper suprabasal layers [fully differentiated])
According to the model outlined in Fig. 6a, K5 and K78 compete, in the basal layer of the epidermis, for the single type I keratin K14 for filament formation. In the parabasal cell layer, the active synthesis of keratins K5 and K14 ceases (although the proteins are still weakly detectable) in favour of the onset of K10 synthesis, which is then confronted with two type II keratins, the still actively synthesised K78 and, in fetal human (and fetal and adult mouse) epidermis, K77 (Langbein et al. 2013). In the lower suprabasal layers, this pattern is supplemented by the type II keratin K1 and, subsequently, in mid-suprabasal layers, is reduced by the cessation of active K78 synthesis. The most complex competition of three type II keratins for K10 is observed in the uppermost epidermal layers in which, in addition to K1, K77 and K2, the new K80 (Langbein et al. 2010) is ultimately expressed (Fig. 6a). In palmoplantar epidermis, this situation is even more complex, because of the additional synthesis of the type I keratin K9 (Langbein et al. 1993; Swensson et al. 1998), which now competes with K10 for the type II keratins in the upper epidermal layers (Fig. 6a). As keratin proteins are of high stability, dynamic aspects of their expression can be better evaluated by analysing the respective transcripts (Roop et al 1988). The non- and radioactive ISH data of K78:K14 or K78:K5 amounts in the basal layer (Fig. 2a:b, c:d, e:f; Supplementary Fig. S3a:b, c:d, e:f, j:k) seem to be widely visually comparable. This also holds true for the amounts of K78:K10 in the parabasal and lower suprabasal layers (cf. respective images in Fig. 2 and Supplementary Fig. S3). Taken together, these data imply that the current view of essentially two distinct keratin pairs, i.e., K5/K14 and K1/K10, which simply discriminate the proliferative (basal) against the differentiating (suprabasal) epidermal compartments is no longer tenable. Rather, at the transition of the basal to the differentiating layers, the new basal keratin pattern K5/K78//K14 underlies successive steps of loss of constituent keratins and gain of new keratins. Within this scenario, the type of keratin K10 represents a sort of backbone along the vertical axis of differentiation and successively undergoes multiple changes of pairing with type II keratins, among which, in humans, the combination K1/K10 is preponderant in adult epidermis but competes with the alternative combination K77/K10 in fetal epidermis (Fig. 6a).
By substituting the keratin pair K1/K10 through K4/K13, the above reasoning can be extrapolated step by step to squamous non-keratinising epithelia (Fig. 6b). Differences to the epidermal pattern are limited to the absence of keratins K2, K77 (in fetal skin) and K9 (in ridged skin) in the uppermost cell layers. The resulting numerically reduced keratin pattern can be attributed to the lack of exposure of these inner epithelia to various external burdens and mechanical stress. This confirms the notion that the total number of keratins expressed in an epithelium correlates with the respective requirements placed upon it (i.e., resilience, protection), an aspect that is particularly evident for hard-keratinising structures such as hair (Langbein et al. 2005, 2013).
What can be deduced from the unforeseen expression pattern of K78 in keratinising and non-keratinising epithelia? We observed unexpected features at the transition from the basal (K78/K5//K14) to the parabasal and then to the suprabasal cell layers of both types of epithelia. Contrary to the current concept of a continuously maintained co-expression of keratin pairs K1/K10 or K4/K13 during differentiation in all living suprabasal (including parabasal) layers of the epidermis or non-keratinising epithelia, respectively, our study revealed that the respective type I keratin is expressed slightly earlier than its type II partner. This results in a continuous parabasal cell layer in which the type I keratins K10 or K13 resort to the type II keratin K78 for filament formation (Fig. 6a, b).
Notwithstanding the strong co-expression of K78 with K5, K1 and/or K4 in squamous epithelia, it is obviously unable to compensate for these keratins in the corresponding gene knockout mice (Ness et al. 1998; Lu et al. 2006; Vijayaraj et al. 2007; Wallace et al. 2012). At present, we are aware of only two examples of a successful replacement and functional compensation of one keratin by another and both have occurred during evolution. Thus, the absence of the suprabasal type II keratin K3 in murine corneal epithelium has been shown to be substituted by keratin K5 whose expression is extended from the basal into the suprabasal compartment (Lu et al. 2006). Similarly, during human evolution, the type I hair cortex keratin gene KRT41 has been mutated into a transcribed pseudogene, KRT41P (formerly ψHaA), concomitant with its full compensation by hair keratin K31 (Winter et al. 2001). Otherwise, mutations leading to defective keratin(s) or to the loss of keratin(s) by knocking them out might induce new keratin(s) that never replace the defect or lost keratin(s) and their functions: this would result in pathologic phenotypes.
Growing evidence indicates that keratin synthesis in epithelia is regulated at the transcriptional level and that each keratin gene is controlled by a fine-tuned plethora of transcription factors and DNA-binding sites of its own (www.genecard.org; Rorke et al. 2015). For example, a comparison of regulatory transcription-factor-binding sites in the gene promoter revealed no consensus in theoretically relevant sites between KRT78 (e.g., CP2, FOXJ2, LUN-1/Foxf2, c-Myb, AP-4, Msx-1 STAT1 and -2 but without, for exmaple, AP1) and KRT5 (e.g., Sp1, AP1, RAR, GR, AP2, NfκB and P53; www.genecard.org; Blumenberg 2000; Rorke et al. 2015) with potential consequences in their gene regulation. Without functional studies, nothing can be known about their functional relevance in KRT78. This suggests that, during the step-wise differentiation of stratified epithelia, each phase of differentiation is characterised by the expression of an intimately orchestrated collective of keratins (for a review, see Blumenberg 2000). Hence, the forced silencing of a distinct keratin gene can apparently not be compensated by the mere presence of a potential, even tightly co-expressed substitute but might require additional factors such as the timely simultaneous expression of the involved type I and type II partners at roughly equimolar concentrations. Thus, insufficient upregulation (K1 in K2-deficient mice; Fischer et al. 2014) or insufficient levels of the constitutive expression of an alternative keratin (K78 in mice deficient in K5, K1 or K4) might be responsible for the failure to substitute for the inactivated keratins.
It stands to reason that mutations in K78 might be pathogenic. In fact, various reports of diseases resembling epidermolysis bullosa (EBS) have appeared in which no mutations in K5 or K14 have been detected (Rugg et al. 1994; Lane and McLean 2004; see also a recent review by Fine et al. 2014). Should some of these cases be indeed attributable to K78 mutations, then the determination of whether K78 is also expressed in the lower suprabasal layers will be of interest as it might lead to subtle alterations in the clinical picture when compared with that of K5- or K14-basal layer-related EBS cases.
In summary, the equally strong basal-parabasal and decreasing mid-suprabasal expression of K78 unveiled here in keratinising and non-keratinising squamous epithelia challenges the prevailing view that the basic keratin pattern in stratified squamous epithelia can principally be described by the classical basal keratin pair K5/K14 and the respective differentiation-specific keratin pairs of the squamous epithelia. The specific expression of K14 and K10 in the basal and suprabasal layers, respectively, has been confirmed by this study, suggesting that the current view of the transition from basal to suprabasal keratin gene expression is valid for type I keratins. In contrast, the plethora of type II keratins in these epithelia entails an impressive dynamic with regard to the sequential keratin expression and, thus, pairing possibilities. Most importantly, the additional formation of an “early differentiation stage” in parabasal cells is suggested such that the switch from the undifferentiated to the differentiated type II keratin gene expression level occurs later than the corresponding switch in type I keratin gene expression (Fig. 6).
As to whether the described scenario constitutes the ultimate complex keratin pattern of keratinising and non-keratinising squamous epithelia must await the elucidation of the expression of the last still uncharacterised type I keratins, namely K23 and K24.
References
Ansell D (2012) The role of hair follicles in cutaneous wound healing. PhD thesis. University of Manchester, Cell Biology, Faculty of Life Sciences
Blumenberg M (2000) Transcriptional control of keratin expression. Madame Curie Bioscience database (internet). NCBI Bookshelf, http://www.ncbi.nlm.nih.gov/books/NBK6213/
Boukamp P, Petrussevska RT, Breitkreutz D, Hornung J, Markham A, Fusenig NE (1988) Normal keratinization in a spontaneously immortalized aneuploid human keratinocyte cell line. J Cell Biol 106:761–771
Bray DJ, Walsh TR, Noro MG, Notman R (2015) Complete structure of an epithelial keratin dimer: implications for intermediate filament assembly. PLoS One 10:e0132706. doi:10.1371/journal.pone.0132706
Bundela S, Sharma A, Bisen PS (2014) Potential therapeutic targets for oral cancer: ADM, TP53, EGFR, LYN, CTLA4, SKIL, CTGF, CD70. PLoS One 9:e102610
Byrne C, Tainsky M, Fuchs E (1994) Programming gene expression in developing epidermis. Development 120:2369–2383
Chen C, Méndez E, Houk J, Fan W, Lohavanichbutr P, Doody D, Yueh B, Futran ND, Upton M, Farwell DG, Schwartz SM, Zhao LP (2008) Gene expression profiling identifies genes predictive of oral squamous cell carcinoma. Cancer Epidemiol Biomarkers Prev 17:2152–2162
Crane HR (1950) Principles and problems of growth. Sci Month 70:376–389
Feng X, Coulombe PA (2015) A role for disulfide bonding in keratin intermediate filament organization and dynamics in skin keratinocytes. J Cell Biol 209:59–72
Fessing MY, Mardaryev AN, Gdula MR, Sharov AA, Sharova TY, Rapisarda V, Gordon KB, Smorodchenko AD, Poterlowicz K, Ferone G, Kohwi Y, Missero C, Kohwi-Shigematsu T, Botchkarev VA (2011) p63 regulates Satb1 to control tissue-specific chromatin remodelling during development of the epidermis. J Cell Biol 194:825–839
Fine JD, Bruckner-Tuderman L, Eady RA, Bauer EA, Bauer JW, Has C, Heagerty A, Hintner H, Hovnanian A, Jonkman MF, Leigh I, Marinkovich MP, Martinez AE, McGrath JA, Mellerio JE, Moss C, Murrell DF, Shimizu H, Uitto J, Woodley D, Zambruno G (2014) Inherited epidermolysis bullosa: updated recommendations on diagnosis and classification. J Am Acad Dermatol 70:1103–1126
Fischer H, Langbein L, Reichelt J, Praetzel-Wunder S, Buchberger M, Ghannadan M, Tschachler E, Eckhart L (2014) Loss of keratin K2 expression causes aberrant aggregation of K10, hyperkeratosis and inflammation. J Invest Dermatol 134:2579–2588
Fraser RDB, Parry DAD (2014) Keratin intermediate filaments: differences in the sequences of the Type I and Type II chains explain the origin of the stability of an enzyme-resistant four-chain fragment. J Struct Biol 185:317–326
Garritano S, Inga A, Gemignani F, Landi S (2013) More targets, more pathways and more clues for mutant p53. Oncogenesis 2:e54. doi:10.1038/oncsis.2013.15
Griffiths G, Lucocq JM (2014) Antibodies for immunolabeling by light and electron microscopy: not for the faint hearted. Histochem Cell Biol 142:347–360
Hesse M, Magin TM, Weber K (2001) Genes for intermediate filament proteins and the draft sequence of the human genome: novel keratin genes and a surprisingly high number of pseudogenes related to keratin genes 8 and 18. J Cell Sci 114:2569–2575
Hesse M, Zimek A, Weber K, Magin TM (2004) Comprehensive analysis of keratin gene clusters in humans and rodents. Eur J Cell Biol 83:19–26
Holbrook KA (2006) Embryogenesis of the skin. In: Irvine AD, Hoeger PH, Yan AC (eds) Harper’s textbook of pediatric dermatology, 2nd edn. Wiley, Hoboken, pp 21-241
Holbrook KA, Odland GF (1975) The fine structure of developing human epidermis: light, scanning, and transmission electron microscopy of the periderm. J Invest Dermatol 65:16–38
Huchon D, Chevret P, Jordan U, Kilpatrick CW, Ranwez V, Jenkins PD, Brosius J, Schmitz J (2007) Multiple molecular evidences for a living mammalian fossil. Proc Natl Acad Sci U S A 104:7495–7499
Kiba T, Kintaka Y, Suzuki Y, Ishizuka N, Ishigaki Y, Inoue S (2010) Gene expression profiling in rat pancreas after ventromedial hypothalamic lesioning. Pancreas 39:627–632
Kiran KC, Rothenberg E, Sherrill JD (2015) In vitro model for studying esophageal epithelial differentiation and allergic inflammatory responses identifies keratin involvement in eosinophilic esophagitis. PLoS One 10:e0127755, doi:10.1371/ journal phone 012775
Kumar S, Hedges SB (1998) A molecular timescale for vertebrate evolution. Nature 392:917–920
Kunisada M, Cui CY, Piao Y, Ko MS, Schlessinger D (2009) Requirement for Shh and Fox family genes at different stages in sweat gland development. Hum Mol Genet 18:1769–1778
Lane EB, McLean WH (2004) Keratins and skin disorders. J Pathol 204:355–366
Langbein L, Schweizer J (2013) The keratins of the human hair follicle. In: Camacho FM, Tosti A, Price VH, Randall VA (eds) Montagna Tricología Enfermedades del folículo pilosebáceo. Editorial Aula Medica, Madrid, pp 73–108
Langbein L, Heid HW, Moll I, Franke WW (1993) Molecular characterization of the body site-specific human cytokeratin 9—cDNA cloning, amino acid sequence and tissue specificity of gene expression. Differentiation 55:57–73
Langbein L, Spring H, Rogers MA, Praetzel S, Schweizer J (2004) Hair keratins and hair follicle-specific epithelial keratins. Methods Cell Biol 78:413–451
Langbein L, Rogers MA, Praetzel S, Cribier B, Peltre B, Gassler N, Schweizer J (2005) Characterization of a novel human type II epithelial keratin K1b, specifically expressed in eccrine sweat glands. J Invest Dermatol 125:428–444
Langbein L, Rogers MA, Praetzel-Wunder S, Helmke B, Schirmacher P, Schweizer J (2006) K25 (K25irs1), K26 (K25irs2), K27 (K25irs3) and K28 (K25irs4) represent the type I inner root sheath (IRS) keratins of the human hair follicle. J Invest Dermatol 126:2377–2386
Langbein L, Eckhart L, Rogers MA, Praetzel-Wunder S, Schweizer J (2010) Against the rules: human keratin K80: two functional alternative splice variants, K80 and K80.1, with special cellular localization in a wide range of epithelia. J Biol Chem 285:36909–36921
Langbein L, Reichelt J, Eckhart L, Praetzel-Wunder S, Kittstein W, Gassler N, Schweizer J (2013) New facets of keratin K77: interspecies variations of expression and different intracellular location in embryonic and adult skin of man and mouse. Cell Tissue Res 354:793–812
Lee CH, Coulombe PA (2009) Self-organization of keratin intermediate filaments into cross-linked networks. J Cell Biol 186:409–421
Lu H, Hesse M, Peters B, Magin TM (2005) Type II keratins precede type I keratins during early embryonic development. Eur J Cell Biol 84:709–718
Lu H, Zimek A, Chen J, Hesse M, Büssow H, Weber K, Magin TM (2006) Keratin 5 knockout mice reveal plasticity of keratin expression in the corneal epithelium. Eur J Cell Biol 85:803–811
Micallef L, Belaubre F, Pinon A, Jayat-Vignoles C, Delage C, Charveron M, Simon A (2008) Effects of extracellular calcium on the growth-differentiation switch in immortalized keratinocyte HaCaT cells compared with normal human keratinocytes. Exp Dermatol 18:143–151
Micallef L, Battu S, Pinon A, Cook-Moreau J, Cardot PJ, Delage C, Simon A (2010) Sedimentation field-flow fractionation separation of proliferative and differentiated subpopulations during Ca2+-induced differentiation in HaCaT cells. J Chromatogr B Anal Technol Biomed Life Sci 878:1051–1058
Moll R, Divo M, Langbein L (2008) The human keratins: biology and pathology. Histochem Cell Biol 129:705–733
Ness SL, Edelmann W, Jenkins TD, Liedtke W, Rustgi AK, Kucherlapati R (1998) Mouse keratin 4 is necessary for internal epithelial integrity. J Biol Chem 273:23904–23911
Nuutila K (2013) Gene expression profiling of human skin donor site wound healing to guide novel regenerative therapies. Academic Dissertation. Faculty of Medicine, Institute of Biomedicine, Pharmacology, Department of Plastic Surgery, Helsinki Burn Centre, University of Helsinki
Omary MB, Ku NO, Tao GZ, Toivola DM, Liao J (2006) “Heads and tails” of intermediate filament phosphorylation: multiple sites and functional insights. Trends Biochem Sci 31:383–394
Rice RH, Bradshaw KM, Durbin-Johnson BP, Rocke DM, Eigenheer RA, Phinney BS, Schmuth M, Gruber R (2013) Distinguishing ichthyoses by protein profiling. PLoS One 8:e75355. doi:10.1371/journal.pone.0075355
Rogers MA, Edler L, Winter H, Langbein L, Beckmann I, Schweizer J (2005) Characterization of new members of the human type II keratin gene family and a general evaluation of the keratin gene domain on chromosome 12q1313. J Invest Dermatol 124:536–544
Roop DR, Krieg TM, Mehrel T, Cheng CK, Yuspa SH (1988) Transcriptional control of high molecular weight keratin gene expression in multistage mouse skin carcinogenesis. Cancer Res 48:3245–3452
Rorke EA, Adhikary G, Young CA, Rice RH, Elias PM, Crumrine D, Meyer J, Blumenberg M, Eckert RL (2015) Structural and biochemical changes underlying a keratoderma-like phenotype in mice lacking suprabasal AP1 transcription factor function. Cell Death Dis 6:e1647. doi:10.1038/cddis.2015.21
Rugg EL, McLean WH, Lane EB, Pitera R, McMillan JR, Dopping-Hepenstal PJ, Navsaria HA, Leigh IM, Eady RA (1994) A functional “knockout” of human keratin 14. Genes Dev 8:2563–2573
Ryle CM, Breitkreutz D, Stark HJ, Leigh IM, Steinert PM, Roop D, Fusenig NE (1989) Density-dependent modulation of synthesis of keratins 1 and 10 in the human keratinocyte line HaCaT and in ras-transfected tumorigenic clones. Differentiation 40:42–54
Sakamoto K, Aragaki T, Morita K, Kawachi H, Kayamori K, Nakanishi S, Omura K, Miki Y, Okada K, Katsube K, Takizawa T, Yamaguchil A (2011) Down-regulation of keratin 4 and keratin 13 expression in oral squamous cell carcinoma and epithelial dysplasia: a clue for histopathogenesis. Histopathology 58:531–542
Schweizer J, Bowden PE, Coulombe PA, Langbein L, Lane EB, Magin TM, Maltheis L, Omary MB, Parry DAD, Rogers MA, Wright M (2006) A new consensus nomenclature for mammalian keratins. J Cell Biol 174:169–174
Swensson O, Langbein L, McMillan JR, Churchill LJ, Leigh IM, McLean WHI, Lane EB, Eady RAJ (1998) Specialized keratin expression pattern in human ridged skin as an adaptation to high physical stress. Br J Dermatol 139:767–775
Thandapani P, O'Connor TR, Bailey TL, Richard S (2013) Defining the RGG/RG motif. Mol Cell 50:613–623
The International Human Genome Sequencing Consortium (2004) Finishing the euchromatic sequence of the human genome. Nature 431:931–945
Toyoda T, Tsukamoto T, Yamamoto M, Ban M, Saito N, Takasu S, Shi L, Saito A, Ito S, Yamamura Y, Nishikawa A, Ogawa K, Tanaka T, Tatematsu M (2013) Gene expression analysis of a Helicobacter pylori-infected and high-salt diet-treated mouse gastric tumour model: identification of CD177 as a novel prognostic factor in patients with gastric cancer. BMC Gastroenterol 35:177–190
Vandebergh W, Bossuyt F (2012) Recurrent functional divergence of early tetrapod keratins in amphibian toe pads and mammalian hair. Mol Biol Evol 29:995–1004
Veniaminova NA, Vagnozzi AN, Kopinke D, Do TT, Murtaugh LC, Maillard I, Dlugosz AA, Reiter JF, Wong SY (2013) Keratin 79 identifies a novel population of migratory epithelial cells that initiates hair canal morphogenesis and regeneration. Development 140:4870–4880
Vijayaraj P, Söhl G, Magin TM (2007) Keratin transgenic and knockout mice: functional analysis and validation of disease-causing mutations. Methods Mol Biol 360:203–251
Vijayaraj P, Kröger C, Reuter U, Windoffer R, Leube RE, Magin TM (2009) Keratins regulate protein biosynthesis through localization of GLUT1 and -3 upstream of AMP kinase and Raptor. J Cell Biol 187:175–184
Wallace L, Roberts-Thompson L, Reichelt J (2012) Deletion of K1/K10 does not impair epidermal stratification but affects desmosomal structure and nuclear integrity. J Cell Sci 125:1750–1758
Winter H, Langbein L, Praetzel S, Jacobs M, Rogers MA, Leigh IM, Tidman N, Schweizer J (1998) A novel human type II epithelial keratin, K6hf, specifically expressed in the companion layer of the hair follicle. J Invest Dermatol 111:955–962
Winter H, Langbein L, Krawczak M, Cooper DN, Jave-Suarez LF, Rogers MA, Praetzel S, Heidt PJ, Schweizer J (2001) Human type I hair keratin pseudogene ΨhHaA has functional orthologs in the chimpanzee and gorilla: evidence for recent inactivation of the human gene after the Pan-Homo divergence. Hum Genet 108:37–42
Yamada S, Wirtz D, Coulombe PA (2002) Pairwise assembly determines the intrinsic potential for self-organization and mechanical properties of keratin filaments. Mol Biol Cell 13:382–391
Acknowledgments
We thank Hermann Stammer for his support with human qPCR data and Iris Martin for HaCaT and NEHK keratinocyte cultures (both German Cancer Research Center, DKFZ, Heidelberg, Germany). Damir Krunic (DKFZ, Light Microscopy Core Facility) is gratefully acknowledged for support with the cLS microscopy. We also thank Ingrid Hausser-Siller (EM lab Dermatology, University of Heidelberg, Germany) and Rudolf Leube (Molecular Cell Anatomy, RWTH Aachen University, Germany) for fruitful discussions.
Funding
We are grateful for generous funding from the Wilhelm-Sander Stiftung, Munich (2007.133.2 to L.L.); L.E. was supported by the Austrian Science Fund (FWF): P23801.
Conflicts of interest
No conflicts of interest are declared.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Langbein, L., Eckhart, L., Fischer, H. et al. Localisation of keratin K78 in the basal layer and first suprabasal layers of stratified epithelia completes expression catalogue of type II keratins and provides new insights into sequential keratin expression. Cell Tissue Res 363, 735–750 (2016). https://doi.org/10.1007/s00441-015-2278-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-015-2278-5