Abstract
Uterine fibroids (UF) are common pelvic tumors in women, heritable, and genome-wide association studies (GWAS) have identified ~ 30 loci associated with increased risk in UF. Using summary statistics from a previously published UF GWAS performed in a non-Hispanic European Ancestry (NHW) female subset from the Electronic Medical Records and Genomics (eMERGE) Network, we constructed a polygenic risk score (PRS) for UF. UF-PRS was developed using PRSice and optimized in the separate clinical population of BioVU. PRS was validated using parallel methods of 10-fold cross-validation logistic regression and phenome-wide association study (PheWAS) in a seperate subset of eMERGE NHW females (validation set), excluding samples used in GWAS. PRSice determined pt < 0.001 and after linkage disequilibrium pruning (r2 < 0.2), 4458 variants were in the PRS which was significant (pseudo-R2 = 0.0018, p = 0.041). 10-fold cross-validation logistic regression modeling of validation set revealed the model had an area under the curve (AUC) value of 0.60 (95% confidence interval [CI] 0.58–0.62) when plotted in a receiver operator curve (ROC). PheWAS identified six phecodes associated with the PRS with the most significant phenotypes being 218 ‘benign neoplasm of uterus’ and 218.1 ‘uterine leiomyoma’ (p = 1.94 × 10–23, OR 1.31 [95% CI 1.26–1.37] and p = 3.50 × 10–23, OR 1.32 [95% CI 1.26–1.37]). We have developed and validated the first PRS for UF. We find our PRS has predictive ability for UF and captures genetic architecture of increased risk for UF that can be used in further studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Uterine fibroids (UF; also known as uterine leiomyomata) are the most common benign tumors for women affecting up to 70% of European ancestry (EA) women and 80% of African ancestry (AA) women by menopause (Baird et al. 2003). Symptoms of UF vary from asymptomatic in up to 50% of cases to severe and debilitating abdominal pain and bloating, pelvic pressure, and heavy menstrual bleeding. Additionally, UF can cause infertility, pregnancy complications, and adverse obstetrics outcomes (Coronado et al. 2000; Qidwai et al. 2006). Despite their high prevalence and health burden on women and the health care system, UF etiology remains elusiveok and many women experience barriers in obtaining a diagnosis.
There are several known risk factors for UF that include increasing proportion of AA, early onset of menarche, obesity, and environmental factors (Baird et al. 2003; Templeman et al. 2009; Terry et al. 2010). Molecular studies of UF tumors have identified somatic cytogenetic rearrangements and mutations in mediator complex subunit 12 (MED12) in 50–85% of tumors and G protein-coupled receptor 10 (GPR10) as causes of UL growth, suggesting chromosome instability similar to cancer (Markowski et al. 2012; Styer and Rueda 2016). UF initiation and growth are thought to be stimulated by fluctuations in estrogen and progesterone (Andersen 1996). Current interventions for UF include pharmaceutical and surgical therapies aimed at reducing symptoms and improving quality of life. Both approaches seek to alleviate symptoms by slowing the growth or shrinking the size of the tumor. Surgical interventions that remove UF tumors and leave the uterus in place have a tumor reoccurrence rate of 21.43%, which increases to 38.71% if a woman has more than one tumor (Radosa et al. 2014), leaving the only permanent curative surgical intervention to be hysterectomy. This has led UF to be the leading cause of hysterectomy and gynecological hospitalization in the United States, estimated to cost US $5.9 to 34.4 billion annually (Cardozo et al. 2012).
Recent advances in genomic analysis have explored germline genetic predispositions to UF. To date, nine genome-wide association studies (GWAS) have been conducted in various populations producing ~ 30 associated loci. These studies include Cha et al. (2011) that identified two loci in a Japanese cohort, later replicated in an EA population by Edwards et al. (2013). Rafnar et al. (2018) identified 16 loci in an Icelandic population, Välimäki et al. (2018) identified 22 loci in a combined UK and Finnish population, Gallagher et al. (2019) identified 8 novel loci and replicated 21 loci in EA, and a genome-wide linkage study by Eggert et al. (2012) discovered two loci. Despite identifying many loci associated with UF, how these variants collectively attribute risk for UF is not well understood.
Polygenic risk scores (PRS) are a weighted sum of the effect of each associated variant, representing the individual’s aggregated genetic predisposition for a disease. They have been recently used to investigate the shared etiology between phenotypes, to test genome-wide gene-by-environment and gene-by-gene interactions, and to assess causal inference using Mendelian Randomization (Choi et al. 2020). Here we construct a UF PRS using summary statistics from a previous EA UF GWAS performed in a subset of the Electronic Medical Record and Genomics (eMERGE) Network (GWAS set), optimized in an independent clinical population BioVU (optimization set), and validated in the eMERGE Network (previous GWAS samples excluded). We use two strategies in parallel, cross-validation and phenome-wide association study, (PheWAS) to validate the PRS and explore genetic risk for UF. PRS was then used in a hypothesis-free discovery PheWAS in non-Hispanic EA (NHW) female-only cohort of eMERGE. We performed supplemental discovery PheWAS in eMERGE populations of NHW sex combined, NHW men, non-Hispanic black (NHB) females only, and NHB sex combined. This approach allows us to probe the causes for and health consequences of UF in the context of the clinical phenome and identify disease diagnoses with shared genetic architecture with UF.
Methods
Study populations
The Electronic Medical Records and Genomics (eMERGE) Network (2007–present) is a national network of DNA repositories that are linked to electronic health records (EHR). A detailed description of the organization of the eMERGE Network has been previously published (McCarty et al. 2011). Data contained in the EHR includes diagnostic (International Classification of Disease, ICD9 and ICD10) and procedure codes, basic demographics, discharge summaries, progress notes, health history, laboratory values, imaging reports, medication orders, and pathology reports. Participants in the eMERGE network were genotyped separately, then imputed and merged. A detailed description of the genotyping, imputation, and quality control of the eMERGE phase III array dataset has been previously reported (Stanaway et al. 2019).
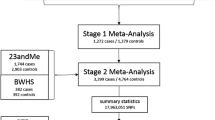
The BioVU DNA Repository is a deidentified database of EHRs that are linked to patient DNA samples at Vanderbilt University Medical Center. A detailed description about the database and how it is maintained has been published elsewhere (Roden et al. 2008). The EHR for BioVU contains the same information as stated above for eMERGE. While Vanderbilt contributes to eMERGE, samples included for this study are unique to BioVU. BioVU participants were genotyped on a custom multi-ethnic genome array (MEGA) array and genotypes aligned to the forward strand. Initial quality control of both study populations excluded samples or variant sites if they had missingness above a 2% threshold. Samples were also excluded if consent had been removed, sample was duplicated, failed sex concordance, or failed self-reported race or genetically determined race concordance. Imputation was performed on the Michigan Imputation Server using Minimac4 using the 1000 genomes phase 3 combined reference panel (Das et al. 2016). An overview of the study aims and analyses performed within each study population are in Fig. 1.
Flow chart depicting methods used in this study. Pink boxes indicate usage of eMERGE Network and the blue box indicates usage of BioVU at various stages. The summary statistics from a previous uterine fibroid (UF) GWAS performed in eMERGE were taken as input for PRSice and PRSice performed the optimization of the polygenic risk score (PRS) in an independent BioVU sample. PRSice used the optimization set to determine the p-value threshold (pT) and linkage disequilibrium (LD) pruning to produce the best PRS possible. A PRS was then calculated for all samples in the validation set. The validation set is comprised of self-reported “non-Hispanic”, “white”, females from the eMERGE network and excludes samples used in the previous GWAS. 10-fold cross validation and phenome-wide association studies (PheWAS) were used as validation methods. Additionally, we performed supplemental discovery PheWAS in various sex and self-reported racial groups
Polygenic risk score development and optimization
The UF polygenic risk score was constructed using PRSice software (version 1.23) (Euesden et al. 2015). Variant effect sizes were extracted from a previous meta-analysis GWAS of image-confirmed case/control UF GWAS by Edwards et al. (2019). Briefly, the study consisted of EA females with imaging confirmed case/control status, totaling 2651 cases and 4306 controls (GWAS set). The summary statistics contained 9,691,748 variants at p-values ranging from 1 to 4.32 × 10–8, which were pruned to remove SNPs in linkage disequilibrium (r2 > 0.2). The model was optimized in an independent set of image-confirmed UF cases (N = 429) and controls (N = 4750) from BioVU (optimization set). Case/control status was image-confirmed and determined in both the GWAS and optimization sets using a previously published algorithm (Feingold-Link et al. 2014). Variants used in the PRS and their respective weights can be found in Supplemental Table 1.
Statistical analysis
All data preparation and statistical analysis was performed in R (version 4.0.3). From the eMERGE cohort, samples used in the GWAS set and samples overlapping with BioVU were removed leaving 80,186 individuals. Ten principle components for remaining samples were calculated using SNPTEST2 (Marchini et al. 2007) and were filtered according to the following criteria to form the NHW female cohort: genetic sex is female, race is self-reported “white”, ethnicity is self-reported “non-Hispanic”, and an adult. This left 26,637 in the NHW female subset that was used as the validation set in PheWAS. Covariates of body mass index (BMI) and age were defined as adult median BMI from the entire EHR and current age were assigned to all individuals where possible. The mean and standard deviation of the covariates were calculated for the population. In the validation set, UF case/control status for females was extracted from the EHR according to PheWAS definitions using the code 218.1 ‘uterine leiomyoma’ to assign cases. UF phecode status was assigned to females and females with missing covariate data were dropped leaving 18,605 NHW females respectively for the 10-fold cross-validation method. Individuals with defined UF phecode status were used for T test, density plots, and receiver operating curve (ROC) analysis. Welch’s Two-Sample T test was used to test for a difference in mean PRS between cases and controls in validation set. Density plot of PRS case/control status in validation set was created using the R package ‘ggplot2’ version 3.3.2 (Whickam 2016).
Predictive performance of UF PRS
To assess the predictive performance of the PRS to identify cases in females, we conducted a 10-fold cross-validated logistic regression and estimated the receiver operator curve (ROC) and its area under the curve (AUC) for various models in the validation set. Modeling and ROC plot visualization was performed in R (version 4.0.3) using packages ‘pROC’ version 1.17.0.1 (Robin et al. 2011) and ‘ggplot2’ version 3.3.2 (Whickam 2016). The base model was UF ~ PRS with subsequently added covariates 10 principal components (PCs), age, and BMI to create the full model UF ~ PRS + 10PCs + Age + BMI. DeLong’s test for two correlated ROC curves was used to test if the full model had a greater AUC value than the base model.
UF PRS PheWAS
We then conducted a hypothesis-free analysis of evaluating what phenotypes in the phenome associate with genetic risk for UF using the R package ‘PheWAS’ version 2.0 (Denny et al. 2010). The method and utility of PheWAS has already been described (Denny et al. 2010). PheWAS used UF PRS as predictor for 1875 clinical disease phenotypes called phecodes from Phecode Map 1.2 (Denny et al. 2013). One unit of PRS increase represents an increase in the aggregate genetic score based on the weights of the variants derived from the previous GWAS. PRS effect sizes were scaled to represent the effect size per one standard deviation increase in PRS. Phecodes are phenotype algorithms based on ICD9 and ICD10 codes. For each phecode, the PheWAS software sorts individuals into cases, controls, or neither, based upon the presence or absence of the phecode component ICD codes in their EHR. Individuals with two or more codes are considered cases, individuals with no instances of the code are considered controls, and individuals with one code, an incompatible sex for the diagnosis, or one or more related codes (in certain scenarios), are considered neither cases nor controls and were excluded from analysis of that diagnosis.
Multiple PheWAS covariate models were assessed in the validation set and the supplemental PheWAS set using the same stepwise addition described above starting with the base model and working to the full model. In sex-combined cohorts, sex was included as a covariate. Within PheWAS regression models, each phecode (having a phecode versus not having the phecode) is the outcome and continuous PRS (raw and untransformed) is the exposure. Supplemental PheWAS sets consisted of NHW sex combined, NHW males only, and individuals who identified as “non-Hispanic” and “black” (NHB) made up the NHB sex combined, and NHB females sets. Population characteristics of each set is in Table 1. A p-value of 2.7 × 10–5 was the threshold for significance to correct for multiple testing (Bonferroni correction of 0.05/1875 phecodes tested). A suggestive significance level of 2.7 × 10–4 was used.
Results
Study populations
In total, 26,637 NHW females comprised the validation set with 18,605 of those women having phecode determined UF status. The mean BMI of the validation set was 28.50 (standard deviation [SD] = 6.94), age was 67.04 (SD = 19.14), and mean PRS was 1227.91 (SD = 14.93) (Table 1). Study populations characteristics for supplemental PheWAS sets can also be found in Table 1.
UF PRS optimization
UF PRS GWAS published summary statistics (Edwards et al. 2019) were evaluated using PRSice (Euesden et al. 2015) using the optimization set. The optimal p-value threshold was determined to be pt < 0.001. After linkage disequilibrium pruning (r2 < 0.2), this PRS consisted of 4458 SNPs. The model was determined to be significant (pseudo-R2 = 0.0018, p = 0.041).
PRS validation and predictive performance analysis
In the validation and supplemental PheWAS sets, 3616 of the 4458 SNPs were available from imputed or genotyped variants. Density plots of the PRS in the validation set stratified on case/control status reveal a right shift in the distribution of PRS for cases (Fig. 2). Welch’s two-sample T test revealed mean PRS was significantly different between cases and controls (p = 3.36 × 10–15) (Table 2).
Density plot of the uterine fibroid (UF) polygenic risk score (PRS) calculated in the validation set. The validation set consisted of 18,605 self-reported “non-Hispanic white” (NHW) females whose case/control status was determined by phecode 218.1 ‘uterine leiomyoma’ according to phenome-wide association study (PheWAS) case/control definitions. The cases had a mean PRS of 1,231.07 while controls had a mean PRS of 1,227.49. T test revealed the means are significantly different (p = 4.09 × 10–15)
Using 10-fold cross-validation logistic regression, the predictive performance of the UF PRS was evaluated in the validation set. The base model (UF ~ PRS) had an AUC value of 0.56 (95% confidence interval [CI] 0.55–0.58). Adding covariates to the model stepwise to achieve the full model (UF ~ PRS + 10PCs + Age + BMI) increased the AUC to 0.60 (95% CI 0.58–0.62) (Fig. 3). Modeling of the covariates (UF ~ 10PCs + Age + BMI) had an AUC of 0.55 (95% CI 0.53–056). DeLong’s one-sided test for correlated ROC curves revealed adding the covariates to create the full model significantly increased the AUC as compared to the base model (p = 3.87 × 10–7). In the full model, the odds ratio (OR) of a one SD increase in PRS is 1.35 (95% CI 1.34–1.37, p = 5.21 × 10–4).
Receiver operator characteristic (ROC) curves of the various 10-fold cross-validation logistic regression models produced using uterine fibroid (UF) status as the outcome and UF polygenic risk score (PRS) as the predictor in the validation set. The validation set consisted of 18,605 self-reported “non-Hispanic white” (NHW) females whose case/control status was determined by phecode 218.1 ‘uterine leiomyoma’ according to phenome-wide association study (PheWAS) case/control definitions. With only the PRS as a predictor, the area under the curve (AUC) is 0.56. Adding 10 PCs, age, and BMI as covariates iteratively increases the area to 0.63. DeLong’s test for two correlated ROC curves found the AUC values of the models to be significantly different (p = 7.74 × 10–7)
UF PRS PheWAS
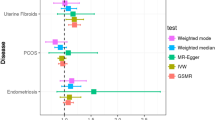
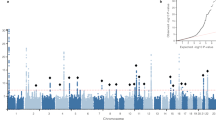
PheWAS was also used to validate the PRS in eMERGE NHW females using multiple models and here we present the results of the full model. We identified six phecodes with evidence of association with the PRS The strongest association was for phecodes 218 and 218.1, ‘benign neoplasms of the uterus’ (OR 1.31 [95% CI 1.26–1.37] p = 1.94 × 10–23) and ‘uterine leiomyoma’ (OR 1.32 [95% CI 1.26–1.37], p = 3.59 × 10–23) (Fig. 4A, Table 3) respectively. Additional associated phecodes included 654.1 ‘abnormality of organs and soft tissues of pelvis complicating pregnancy, childbirth, or the puerperium’ (OR 1.75 [95% CI 1.57–1.92], p = 3.64 × 10–10) 626 ‘disorders of menstruation and other abnormal bleeding from female genital tract’ (OR: 1.12 [95% CI 1.08–1.16] p = 7.84 × 10–9), 626.1 ‘Irregular menstrual cycle/bleeding’ (OR 1.13 [95% CI 1.09–1.17] p = 1.14 × 10–8) and 626.12 ‘Excessive or frequent menstruation’ (OR 1.17 [95% CI 1.11–1.23] p = 3.45 × 10–8) (Fig. 4A, B, Table 3). Full PheWAS summary statistics for all models can be found in Supplemental Tables 2 to 5.
A Manhattan plot of PheWAS results for eMERGE self-identified "non-Hispanic white" females (N = 26,637). Model was adjusted for age, BMI, and ten principal components. Most significantly associated phenotypes with the PRS are benign neoplasm of uterus (phecode 218) and uterine leiomyoma (phecode 218.1). Red line represents Bonferroni significance (2.7 × 10–5) and blue line represents suggested significance (2.7 × 10–4). B A zoomed in Manhattan plot of genitourinary category phenotypes from the eMERGE NHW females adjusted for age, BMI, and ten principal components PheWAS (N = 51,396). Three of the four other significant phenotypes are in this category along with phecodes sitting just below significance including ovarian cysts and endometriosis
We observed evidence of nominal association (one magnitude above the multiple testing threshold, p < 2.7 × 10–4) in the validation set full model for the following phecodes: 625 ‘Pain and other symptoms associated with female genital organs’ (OR 1.09 [95% CI 1.05–1.14], p = 1.57 × 10–4), 212 ‘Benign neoplasm of respiratory and intrathoracic organs’ (OR 1.43 [95% CI 1.23–1.63], p = 5.00 × 10–4) (Fig. 4B, Table 3). The phenotype 212 is the only phenotype to afflict both sexes and it is not significant in supplemental PheWAS of NHW sex combined (p = 0.019) (Supplemental Tables 6 to 10). The phenotype ‘ovarian cysts’ is associated with the PRS in the base model of the validation set (OR 1.13 [95% CI 1.07–1.18], p = 2.57 × 10-5) but is not significant in the full model (p = 8.91 × 10-4) (Supplement Table 9). Supplemental PheWAS performed included NHW sex combined, NHW male-only, NHB sex combined, and NHB females-only subsets. None of these analyses yielded significant or suggestive significant results except the raw PRS in NHW males with no adjustments (Supplemental Tables 10 to 21).
Discussion
Here we present the first constructed, independently optimized, and independently validated PRS for UF. We used two strategies in parallel to validate the PRS and demonstrate that the findings of our UF PRS were consistent across validation methods. In our two validation methods, 10-fold cross-validation logistic regression and PheWAS, performed in a NHW female-only subset from eMERGE, we observed consistent OR for a one standard deviation increase in PRS for the phecode 218.1 ‘uterine leiomyoma’: 1.35 (95% CI 1.34–1.37) and 1.32 (95% CI 1.26–1.37) respectively. OR estimate and its 95% confidence interval for the UF phecode was consistent across all PheWAS models tested in the NHW female-only subsets eMERGE subset (Supplement). Additionally, we found that cases in our NHW female-only subset validation sets had a statistically significant greater mean PRS than controls and observe a right shift in the distribution of PRS values in cases as compared to controls in the density plots (Fig. 2). Our results indicate the PRS we have constructed is capturing some genetic risk for UF and an increase in PRS value is associated with an increased risk for UF.
The UF PRS associated with an additional five phecodes in all PheWAS models tested in the validation set. One phecode is a parent code to 218.1 and represents UF in the EHR, one code is a pregnancy complication (“Abnormality of organs and soft tissues of pelvis complicating pregnancy, childbirth, or the puerperium”), and three codes are menstrual phenotypes. The menstrual phenotypes detected describe menstruation as “irregular” and “excessive”, which describe the known symptomology of UF (Wise and Laughlin-Tommaso 2016). It is possible we are detecting women who are experiencing symptoms of UF but do not have an official UF diagnosis. Women making these kinds of complaints to their health care provider may go on to be diagnosed with UF. Evaluating the combination of symptomology with genetic risk for UF may allow for quicker diagnosis and treatment for UF in women. The final significant phenotype is a pregnancy complication. UF is believed to be the cause of infertility and pregnancy complications in a small percentage of women, with fibroid characteristics (location, size, number) affecting the likelihood fibroids are the root cause. Women with more severe fibroids (submucosal fibroids, large, multiple fibroids) are more likely to experience infertility and pregnancy complications (Parazzini et al. 2015). We observed the largest effect size for this phenotype, indicating increasing genetic risk for fibroids places women at higher risk for this pregnancy complication.
In addition to the NHW females from eMERGE, we also performed PheWAS in the eMERGE subsets of NHW sex combined, NHW men, NHB sex combined, and NHB females. The NHW sex combined PheWAS bore similar results to the NHW females PheWAS (Supplement). There were no significant results in PheWAS for NHW men, NHB sex combined, and NHB females (Supplement). Similar findings between the NHW females and NHW sex combined while simultaneously seeing no significant results in NHW males indicate the results observed in PheWAS are being solely driven by women in the cohort. We do not observe increased risk for any phenotypes related to men in association with increased genetic UF risk. No significant results in any AA population PheWAS performed is in line with current literature. Literature has noted that PRS’s have low or no predictive power when applied to a population that is different from the ancestral population that the summary statistics are derived from (Dikilitas et al. 2020). Possible reasons for this observed phenomenon are common variants differ between populations, differences in LD structures, and differences in environment (Martin et al. 2019). Overall, this suggests the genetic architecture for UF may differ based upon race.
PRS are currently being explored for integration in different aspects of health care/prediction (Chatterjee et al. 2016). There are benefits and shortcomings to using a PRS in a clinical setting that include risk prediction, diagnostic refinement, and improved population screening, incorrect risk estimations, ancestry bias, and regulation respectively (Adeyemo et al. 2021). The PRS constructed in this study follows the benefits and risks outlined and would need more refinement before clinical implementation is possible. However, we have created a new tool for fibroids research. Future studies, such as Mendelian randomization, can use the PRS to investigate genetic architecture of UF risk. Knowledge of the genetic architecture can lead to a better understanding of the etiology of the disease and may identify novel drug targets and therapy options. Currently the prediction model proposed here of the PRS, PCs, age, and BMI performs moderately weakly for prediction. Formulating and adding a clinical or phenotype risk score for the model may increase the predictability and utility of such a model.
The greatest strength of our study is we were able to use independent datasets (no overlapping samples) for each stage (GWAS, PRS optimization, PRS validation) of this study. Additionally, the optimization of the PRS was performed in a different clinical population (BioVU) from the GWAS and validation clinical population (eMERGE). Theoretically, this should make our PRS more transferable across different clinical populations of NHW females. However, we were limited to internal validity of the eMERGE clinical population. External validity of the PRS in an independent clinical population should be performed when available. Finally, because we used eMERGE for the GWAS step and PRS validation, we had to restrict the number of usable UF cases. Many of the severe cases of UF (evidence of large tumors, multiple tumors, or tumor reoccurrence) were used in the GWAS. Setting up the study this way may overestimate the effect size of variants produced in GWAS but underestimate the effect size of the PRS in validation. Despite these shortcomings, we have produced a UF PRS that is significantly associated with UF in a clinical population.
Our results indicate that there is shared genetic architecture between UL and other disorders in the clinical phenome. Based on the other phenotypes, it is suggested that this architecture may contain genetic variants that effect endogenous sex hormones within women. Further work will need to be carried out to determine what this shared relationship is. This in turn will give rise to a better understanding of the etiology of UL and provide the bases for the development of future treatment.
Data availability
The authors declare that the data supporting the findings of this study are available within the article and its supplemental tables. Variants and their respective weights and alleles contained in the PRS are available in Supplemental Table 1. Data used to construct the PRS will be made available in PGS catalog upon publication. Full summary statistics from the UF GWAS used to construct the PRS are available in MRBase.
References
Adeyemo A, Balaconis MK, Darnes DR, Fatumo S, Granados Moreno P et al (2021) Responsible use of polygenic risk scores in the clinic: potential benefits, risks and gaps. Nat Med 27:1876–1884
Andersen J (1996) Growth factors and cytokines in uterine leiomyomas. Semin Reprod Endocrinol 14:269–282
Baird DD, Dunson DB, Hill MC, Cousins D, Schectman JM (2003) High cumulative incidence of uterine leiomyoma in black and white women: ultrasound evidence. Am J Obstet Gynecol 188:100–107
Cardozo ER, Clark AD, Banks NK, Henne MB, Stegmann BJ et al (2012) The estimated annual cost of uterine leiomyomata in the United States. Am J Obstet Gynecol 206:211.e211-219
Cha PC, Takahashi A, Hosono N, Low S-K, Kamatani N, Kubo M, Nakamura Y (2011) A genome-wide association study identifies three loci associated with susceptibility to uterine fibroids. Nat Genet 43(5):447–450
Chatterjee N, Shi J, Garcia-Closas M (2016) Developing and evaluatung polygenic risk prediction models for stratified disease prevention. Nat Rev Genet 17:392–406
Choi SW, Mak TS-H, O’Reilly PF (2020) Tutorial: a guide to performing polygenic risk score analyses. Nat Protoc 15:2759–2772
Coronado GD, Marshall LM, Schwartz SM (2000) Complications in pregnancy, labor, and delivery wither uterine leiomyomas: a population-based study. Obstet Gynecol 95:764–769
Das S, Forer L, Schonherr S, Sidore C, Locke AE, Kwong A, Vrieze SI, Chew EY, Levy S, McGue M, Schlessinger D, Stambolian D, Loh PR, Iacono WG, Swaroop A, Scott LJ, Cucca F, Kronenberg F, Boehnke M, Abecasis GR, Fuchsberger C (2016) Next-generation genotype imputation service and methods. Nat Genet 48(10):1284–1287
Denny JC, Ritchie MD, Basford MA, Pulley JM, Bastarache L et al (2010) PheWAS: demonstrating the feasibility of a phenome-wide scan to discover gene-disease associations. Bioinformatics 26:1205–1210
Denny JC, Bastarache L, Ritchie MD, Carroll RJ, Zink R et al (2013) Systematic comparison of phenome-wide association study of electronic medical record data and genome-wide association study data. Nat Biotechnol 31:1102–1111
Dikilitas O, Schaid DJ, Kosel ML, Carroll RJ, Chute CG et al (2020) Predictive utility of polygenic risk scores for coronary heart disease in three major racial and ethnic groups. Am J Hum Genet 106:707–716
Edwards TL, Hartmann KE, Velez Edwards DR (2013) Variants in BET1L and TNRC6B associate with increasing fibroid volume and fibroid type among European Americans. Hum Genet 132:1361–1369
Edwards TL, Giri A, Hellwege JN, Hartmann KE, Stewart EA et al (2019) A trans-ethnic genome-wide association study of uterine fibroids. Front Genet 10:511
Eggert SL, Huyck KL, Somasundaram P, Kavalla R, Stewart EA et al (2012) Genome-wide linkage and association analyses implicate FASN in predisposition to uterine leiomyomata. Am J Hum Genet 91:621–628
Euesden J, Lewis CM, O’Reilly PF (2015) PRSice: Polygenic Risk Score software. Bioinformatics 31:1466–1468
Feingold-Link L, Edwards TL, Jones S, Hartmann KE, Velez Edwards DR (2014) Enhancing uterine fibroid research through utilization of biorepositories linked to electronic medical record data. J Womens Health (larchmt) 23:1027–1032
Gallagher CS, Makinen N, Harris HR, Rahmioglu N, Uimari O et al (2019) Genome-wide association and epidemiological analyses reveal common genetic origins between uterine leiomyomata and endometriosis. Nat Commun 10:4857
Marchini J, Howie B, Myers S, McVean G, Donnelly P (2007) A new multipoint method for genome-wide association studies by imputation of genotypes. Nat Genet 39:906–913
Markowski DN, Bartnitzke S, Loning T, Drieschner N, Helmke BM, Bullerdiek J (2012) MED12 mutations in uterine fibroids—their relationship to cytogenetic subgroups. Int J Cancer 131(7):1528–1536 (Electronic)
Martin AR, Kanai M, Kamatani Y, Okada Y, Neale BM et al (2019) Clinical use of current polygenic risk scores may exacerbate health disparities. Nat Genet 51:584–591
McCarty CA, Chisholm RL, Chute CG, Kullo IJ, Jarvik GP et al (2011) The eMERGE Network: a consortium of biorepositories linked to electronic medical records data for conducting genomic studies. BMC Med Genom 4:13
Parazzini FMD, Tozzi LMD, Bianchi SMD (2015) Pregnancy outcome and uterine fibroids. Best practice & research. Clin Obstet Gynaecol 34:74–84
Qidwai GI, Caughey AB, Jacoby AF (2006) Obstetric outcomes in women with sonographically identified uterine leiomyomata. Obstet Gynecol 107:376–382
Radosa MP, Owsianowski Z, Mothes A, Weisheit A, Vorwergk J et al (2014) Long-term risk of fibroid recurrence after laparoscopic myomectomy. Eur J Obstet Gynecol Reprod Biol 180:35–39
Rafnar T, Gunnarsson B, Stefansson OA, Sulem P, Ingason A et al (2018) Variants associating with uterine leiomyoma highlight genetic background shared by various cancers and hormone-related traits. Nat Commun 9(1):3636
Robin X, Turck N, Hainard A, Tiberti N, Lisacek F, Sanchez J-C, Müller M (2011) pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinform
Roden DM, Pulley JM, Basford MA, Bernard GR, Clayton EW et al (2008) Development of a large-scale de-identified DNA biobank to enable personalized medicine. Clin Pharmacol Ther 84:362–369
Stanaway IB, Hall TO, Rosenthal EA, Palmer M, Naranbhai V et al (2019) The eMERGE genotype set of 83,717 subjects imputed to ~40 million variants genome wide and association with the herpes zoster medical record phenotype. Genet Epidemiol 43:63–81
Styer AK, Rueda BR (2016) The epidemiology and genetics of uterine leiomyoma. Best practice & research. Clin Obstet Gynaecol 34:3–12
Templeman C, Marshall SF, Clarke CA, Henderson KD, Largent J et al (2009) Risk factors for surgically removed fibroids in a large cohort of teachers. Fertil Steril 92:1436–1446
Terry KL, De Vivo I, Hankinson SE, Missmer SA (2010) Reproductive characteristics and risk of uterine leiomyomata. Fertil Steril 94(7):2703-2707 (Electronic)
Välimäki N, Kuisma H, Pasanen A, Heikinheimo O, Sjöberg J et al (2018) Genetic predisposition to uterine leiomyoma is determined by loci for genitourinary development and genome stability. Elife 7:e37110
Whickam H (2016) ggplot2: Elegant graphics for data analysis. Springer-Verlag, New York
Wise LA, Laughlin-Tommaso SK (2016) Epidemiology of uterine fibroids: from menarche to menopause. Clin Obstet Gynecol 59(1):2–24
Funding
This research was funded by the National Institutes of Health through the following grants: [R01HD074711, R03HD078567, R01HD093671 to D.R.V.E., T32GM080178 to J.A.P., U01HG006378 to D. R., K12HD04348 to J.N.H.]. This research has been conducted using the UK Biobank Resource (Project ID: 13869). The Mayo Clinic is funded by HG06379. This phase of the eMERGE Network was initiated and funded by the National Human Genome Research Institute through the following grants: U01HG8657 (Group Health Cooperative/University of Washington); U01HG8685 (Brigham and Women’s Hospital); U01HG8672 (Vanderbilt University Medical Center); U01HG8666 (Cincinnati Children’s Hospital Medical Center); U01HG6379 (Mayo Clinic); U01HG8679 (Geisinger Clinic); U01HG8680 (Columbia University Health Sciences); U01HG8684 (Children’s Hospital of Philadelphia); U01HG8673 (Northwestern University); U01HG8701 (Vanderbilt University Medical Center serving as the Coordinating Center); U01HG8676 (Partners Healthcare/Broad Institute); and U01HG8664 (Baylor College of Medicine).
Author information
Authors and Affiliations
Contributions
YZ, GPJ, OD, IJK, DJS, DRC, SAP, MTML, and DRVE designed the study. YZ, GPJ, OD, IJK, DJS, DRC, SAP, MTML, JCD, and DR contributed to phenotypical/clinical aspects of cohorts. YZ, GPJ, OD, IJK, DJS, DRC, SAP, MTML, and EST contributed to genotyping, quality control, imputation, and/or analysis of genotyping data. JNH constructed the UL PRS and calculated a PRS for all participants in eMERGE and BioVU. JAP performed PheWAS and Mendelian randomization. JAP and DRVE drafted the paper. JNH, YZ, OD, IJK, EST, GPJ, DJS, DRC, SAP, MTML, DR, JCD, and TLE provided critical comments on paper, draft, and analysis. All authors read and approved the final paper.
Corresponding author
Ethics declarations
Conflict of interest
All authors have no relevant financial or non-financial interests to disclose.
Ethics approval
The Institutional Review Board (IRB) of Vanderbilt University approved this study and confirmed no ethical approval is required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Piekos, J.A., Hellwege, J.N., Zhang, Y. et al. Uterine fibroid polygenic risk score (PRS) associates and predicts risk for uterine fibroid. Hum Genet 141, 1739–1748 (2022). https://doi.org/10.1007/s00439-022-02442-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-022-02442-z