Abstract
The prehistoric human settlement of the Lakshadweep islands remains a mystery for various reasons. Uncertainty about the existence of indigenous tribes in these islands and the lack of folklore records present major obstacles to the reconstruction of Lakshadweep ancestry. However, with extant population data, we seek to understand the maternal ancestry of the Kavaratti islanders. Mitochondrial control region variation analysis of 80 individuals from this island shows maternal links with the populations in the northwestern region of the South Asian mainland. The founder clade R30b2, observed in the Kavaratti islanders, is so far present only in the Scheduled Castes from the Punjab region, Jat Sikhs and Nairs. All other mainland populations carry basal R30 or R30a subclades. The presence of a specific Uralic U4 lineage in our samples, in addition to the Indo-European affinity observed in the phylogeny tree, substantiates a northwestern maternal ancestry of the Kavaratti islanders and implies an ancestral admixture with early humans in the Near East at the time of the last glacial maximum (LGM). Based on our Bayesian analysis, we furthermore propose that a group bearing mostly R30b2 during the LGM recovery, moved eastward and southward, where they received Indian-specific M haplogroups. Hence, the maternal ancestry of the Kavaratti islanders is evidently a consequence of the demographic changes in the northwestern region of the Indian subcontinent caused by the Last Glacial Maximum. The haplogroup distribution pattern and nucleotide sequence data produced in this study will enrich the forensic database of the Lakshadweep islands.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The prehistory and settlement of the Lakshadweep (Lakṣadvīpŭ) islands, long known in English and other Western languages by the tadbhava form of the native name for the archipelago as the Laccadive Islands, has interested numerous researchers and historians. However, the question as to who the earliest settlers of this archipelago were still awaits an answer. We seek to solve this puzzle with genetic data. Even in the decade of high-throughput next generation sequencing, mitochondrial DNA remains an important tool for identifying and dating deep maternal lineages of antiquity that are present in extant populations. Studies rely on haplogroup frequency distribution and the divergence times of haplogroups in order to understand the clustering patterns within and between populations. In this study, we identify haplogroups prevalent in the Kavaratti islanders (n = 80) of the Lakshadweep archipelago. We then use the haplogroup frequency distribution pattern as well as the control region sequence variation information obtained from various datasets to perform multivariate analysis. In addition, we construct maximum likelihood and Bayesian phylogenetic trees to date the coalescent age of these clusters. Interestingly, all our analyses point towards the northwestern region of the Indian subcontinent and date the maternal ancestry of the Kavaratti islanders to the Last Glacial Maximum. This study will provide an important direction for future research emphasising the repercussions of the LGM in shaping the gene pool of both northern and southern ancient populations.
The Lakshadweep Islands are located off the southwest coast of the Indian subcontinent. Kavaratti Island is one of the 36 islands of this archipelago. Kavaratti Island is surrounded by Agatti and Amini in the north and Āndrōttŭ and Kalpēni in the east (Fig. 1). The nearest distance from the city of Cochin (Kocci) on the Malabar Coast is 404 km. Kavaratti is the capital of the Lakshadweep Union Territory. In the 2011 census, the population was 11,221. Socioculturally, all the islands except Minicoy are similar, with the majority following Islam and mostly involved in coconut plantations, jute trade and fisheries. Minicoy, which is closer to the Maldives, has undergone the greater influence of Maldivian customs. The Lakshadweep population comprises four tribes, i.e. Aminidivi (Aminidvīpŭ), Kōya, Mālumi and Mēlāccēri. However, the aboriginal tribes are believed to have gone extinct with the arrival of the ancestors of the present-day inhabitants, who have characteristics of the Malabārīs of Kerala. The Kōya take pride in tracing their ancestry to Nambūdiri Brahmins and occupy the highest stratum on the islands, whereas the Mālumi and Mēlāccēri provide services to the Kōya. The Mēlāccēri dominate the northern islands. The Mālumi are believed to have descended from the seafaring Urukkār ‘boatmen’ class of the Malabar Coast. The Mēlāccēri are believed to be descendants of the Tīyyar tribe (Gabriel 1988).
The Periplus of the Erythraean Sea, dating from the first century AD, mentions the Laccadives as ‘islands off the Malabar coast’ (τὰς νήσους…αὐτῆς τῆς Λιμυρικῆς) as a source of tortoise shell, and mentions the two islands Αἰγιδίον (Aigidion) and Καινειται (Cæneitæ) as a hideout for pirates (Arrian and Strabo 1533; McCrindle 1878; Schoff 1912). Gray (1889) identified the islands Αἰγιδίον (Aigidion), Κάναθρα (Canathra), Μονάχη (Monachè) and Ἀμμίνη (Amminè), mentioned in the second century AD by Claudius Ptolemy in his Geography, with the Laccadive islands Kavaratti, Agatti, Amini and Minicoy, respectively. The Καινειται (Cæneitæ) of the Periplus and Ptolemy’s Κάναθρα (Canathra) may both represent Occidental renderings of the name of Kavaratti, and we also suggest that Ὀρνέων (Orneorum) and Καλαδαδρούα (Calandadrua) in the same archipelago may have represented Ptolemy’s Graeco-Roman renderings of the original names of the islands known today in modern Malayāḷam as Baṃkaram and Kiltān, respectively (Moletius 1562; Nobbe 1845). The Kavaḍidvīpa mentioned in the twelfth century inscription at Narendra (Barnett 1915) and in another contemporaneous inscription of the Goan Kādamba dynasty as Kāpardikadvīpa (Fleet 1870) is probably Kavaratti (Tripati 2006). Human settlement on the Laccadives no doubt dates back to antiquity. Forbes (1979) contends that a wave of Hindu immigrants from the Indian mainland settled on the island in or around the seventh century AD. The older inscriptions on the Laccadives are in the Vaṭṭeḻuttu script and are anterior in date to the inscriptions in Arabic script. The dating of the introduction of Islam to the Laccadives remains problematic, but the formerly Buddhist island realm of the Maldives, including Minicoy, were not converted to Islam until the middle of the twelfth century AD.
Not many genetic studies have been undertaken on the population of the Laccadives. Mustak et al. (2019) studied the maternal and paternal ancestry on eight islands. The study concluded that the maternal ancestry of the islanders was related to South Indian populations, whilst the paternal lineage derived overwhelmingly from North Indian and West Eurasian haplogroups. Kavaratti island was not included. In the present study, we address this gap by generating forensic data for an important island from the Lakshadweep archipelago in the form of mitochondrial DNA sequences. A recent study that focused on the Mēlāccēri tribe (Anthropological Survey of India 2021) found only two haplogroups, viz. M30 and R30, each with very low nucleotide diversity, confirming a strong bottleneck event. In order to understand the cause for such a founder effect, in the present study, we emphasise the R30 and M2 haplogroup distribution. We also assess the impact of the Last Glacial Maximum (Malyarchuk et al. 2010; Silva et al. 2017; Shamoon-Pour et al. 2019) as a possible cause for the change in population dynamics in the Near East.
Materials and methods
Population sample
Blood samples were randomly collected from 80 unrelated males residing on Kavaratti Island. Informed consent was obtained from the participants. The study was conducted after obtaining the approval of the Institutional Human Ethics Committee of Mangalore University in accordance with the guidelines in force.
DNA extraction and genotyping
DNA was extracted using the standard Phenol Chloroform method. Non-coding hypervariable segments in the control region were amplified using ABI Veriti 96-well Thermal Cycler (Applied Biosystems). For improved resolution, the coding region (11,948–12,772 and 14,856–15,978) covering 2028 bp was also amplified. Primer details are given in ESM_1 Table 1. PCR products were sequenced using Sanger sequencing technique (ABI 3130xl Genetic Analyser, Applied Biosystems). The mtDNA sequences were aligned against the reverse Cambridge Reference Sequence (rCRS) in Ugene Unipro Software (Okonechnikov et al. 2012). Variations were scored, and haplogroups were assigned according to Phylotree 17 FU1 (Dür et al. 2021), using Haplogrep 2.4.0 software (Weissensteiner et al. 2016), followed by a EMPOP database query check (Huber et al. 2018). Two samples showing ambiguous results were excluded from the analysis. Data generated during this study are available in the supplementary file ESM_2.
Statistical analysis
DnaSP6 software (Rozas et al. 2017) was used to calculate nucleotide diversity, haplotype diversity and Tajima’s D indices, based on sequence variation. For comparative analysis, pairwise Fst among different populations was calculated using Arlequin Software (Excoffier and Lischer 2010). The same software was used for AMOVA (Analysis of Molecular Variance). PAST4 software (Hammer et al. 2001) was used for multivariate analysis, such as Principal Component Analysis (PCA) and Neighbour Joining (NJ) Tree clustering. M2 and R30 haplogroup network analysis was performed using the POPART package in R (Leigh and Bryant 2015). The Phylogeny Tree was constructed using Mega 11 software (Tamura et al. 2021).
Bayesian analysis
Bayesian age estimation was performed using BEAST v2.7.4. Samples were filtered based on the haplogroups. Ancient mitogenomes (with higher coverage and C14 dates) from AmtDB were used for calibration (Ehler et al. 2019). The L2c2 mitogenome from a Moreno individual (PaMOR16007) was used as an outgroup. BEAST runs were performed using the non-coding region sequences. Trees were linked, and runs were performed with the HKY substitution model. A strict molecular clock was chosen with mutation rates of 1.292 and 0.369 mutation/site/million years for the HVS1 and HVS2 region, respectively (Connell et al. 2022). A consensus tree was generated using Treeannotator with the following settings: 50% burnin, Common Ancestor Heights. Consensus tree was prunned in iTOL. In Figtree, the tree was rerooted to L2c2 outgroup. Median Heights with 95% HPD (High Probability Densities) were considered as TMRCAs and reported. All the runs were performed with 5,000,000 burnin and 50,000,000 iterations, drawing samples at intervals of 10,000 MCMC (Markov chain Monte Carlo) steps (Olivieri et al. 2017; Brandini et al. 2018; Capodiferro et al. 2021).
Results and discussion
Genomic diversity in the Kavaratti islanders
Haplogroups R30b2 (32.1%), M2a’b (17.7%), M33a (10.3%), M3a1 + 204 (9.0%) and U4b1a1a1 (5.1%) account for about 70% of the total mtDNA variation on Kavaratti Island (Table 1). The abundance of macrohaplogroup M (51%) in this population resembles mainland populations of the Indian subcontinent (Anthropological Survey of India 2021). Macrohaplogroup R accounted for 32% of the total distribution. The U4 haplogroup occurred at 10% frequency. Viewed at the macro-level, this population carries Indian-specific M and ancient N in relatively equal proportion. These two haplogroups contrast with the mainland tribal gene pool, which shows a significant difference between the two. Interestingly, such equal distribution is also a characteristic of Northern India (Anthropological Survey of India 2021).
Great importance is ascribed to the distribution of M2 in the Indian mainland as far as autochthonous haplogroups are concerned. The frequency of haplogroup M2 in Dravidian tribes is higher than it is in Indo-European speakers (Metspalu 2004). This variation is geographically correlated as well, with a higher prevalence in central and western tribes (Kumar et al. 2008) and traces in eastern tribes like Munda and Mal Pahariya. Betta Kuruba is an exception with the highest M2 frequency (~ 40 to 60%) in the mainland. On Kavaratti island, haplogroup M2 accounts for 19.2% of the variation, suggesting its deep ancestral link with these ancient tribes (Kumar et al. 2008; Anthropological Survey of India 2021). Our data allow us to resolve the M2 haplogroup up to the M2a’b subclade with mutations at 447, 16,274 and 16,319 (van Oven and Kayser 2009). Earlier studies have shown a higher prevalence of the M2a subclade in populations in the eastern part of India (Metspalu 2004), whilst haplogroup M2b peaks at 35.65% in Betta Kuruba and is generally absent in tribes of the western region (Kumar et al. 2008; Anthropological Survey of India 2021). Therefore, the two lineages of M2 are evidently unique. Without the coding region mutations, the presence of M2a on Kavaratti Island cannot be confirmed. However, the absence of diagnostic mutations for ‘Dravidian’ M2b and M2b2 in the non-coding region, i.e. transition at 182 and poly ‘A’ at 16,180–19,182 followed by 12 ‘C’s and transition at 16,295, respectively (Kumar et al. 2008), hints at a unique ancient lineage of M2 on this island that is different from the one prevalent in southern tribes. To substantiate the uniqueness of the M2 lineage on the island, we performed network analyses of M2 individuals from various datasets. Our analyses of data show a separate cluster for the Kavaratti islanders (ESM_1 Fig. 4). Furthermore, Bayesian analysis of M2 sequences from the available databases clustered the M2 subclade found in Kavaratti island separately. This Kavaratti M2 cluster shares a common ancestor with the northwestern M2 clade, and appears to be younger (41,611 YBP, 95% HPD: 18,983–68,032) than the other clades found in the Indian subcontinent (ESM_1, Fig. 5 and Table 4) with coalescence times of 46–57 thousand years. Earlier studies have reported a broader range for the time to the most recent common ancestor for M2 subclades in India (Metspalu 2004; Kumar et al. 2008; Chandrasekar et al. 2009), with M2a dated between 21 and 50 thousand years and M2b dated between 12 and 55 thousand years. Nevertheless, with the exception of M2, which seems to be absent in our samples, the time to the most recent common ancestor for M2a fell in the pre-Last Glacial Maximum period. This time frame for the Kavaratti M2 cluster in addition to its affinity towards the northwestern M2 clade suggests that our samples share M2 ancestry with the northwestern populations.
Another haplogroup that belongs to the pre-LGM is R30. To substantiate this claim, our Bayesian analysis, involving all the available R30 sequences, showed an increase in the population size ~ 27 kya (ESM_1. Figure 11). However, the only subclade present on Kavaratti Island is R30b2 (32.1%). Our results are similar to the findings of the earlier studies, which showed a higher prevalence of haplogroup R30 on the Lakshadweep Islands (Mustak et al. 2019; Anthropological Survey of India 2021). Mustak et al. (2019) report R30 frequencies ranging between 60 and 80 percent in some of the northern islands of the archipelago. A control region sequence analysis of R30 samples in the Kavaratti islanders shows very low haplotype diversity (Hd = 0.417) and nucleotide diversity (π = 0.00152), suggesting a recent population bottleneck or a founder event (Grant and Bowen 1998). All of these pieces of evidence motivates us to conclude that R30b2 is indeed a founder haplogroup for the Lakshadweep population. The reported coalescent age of the R30b2 subclade is approximately 16 ± 8.4 KY (Chaubey 2013), corresponding to the timeline of the LGM recovery (Clark et al. 2009; Hughes and Gibbard 2015). Based on our Bayesian analysis, we establish a time to the most recent common ancestor of 17,616 (95% HPD: 6661–32,561) for R30b2 (ESM_1, Fig. 6). Discrepancies were observed in earlier studies (Chaubey 2013; Ranaweera et al. 2014) regarding the distribution of R30 in the Vedda tribe on Sri Lanka. The observed R30 haplogroup distribution is mainly due to the exclusion of the diagnostic mutation at 373, which is one of the diagnostic mutations for R30 (van Oven and Kayser 2009). When these sequences were reanalysed, we found that the Vedda tribe carry only traces of R30 lineages (~ 3%, unpublished), whereas the Urali Kuruman draw 35% of their ancestry from the R30a1c subclade, with a coalescent age of ~ 9.4 KY(Sylvester et al. 2019). We report a time to the most recent common ancestor of ~ 12,000 years for R30a1c, whilst the R30a clade dates back to ~ 39,000 YBP. While both of these tribes carrying R30a and R30b are from the southern subcontinent, clade R30 is believed to have originated in northern India (Palanichamy et al. 2004), with specific subclades in Nepal (Fornarino et al. 2009) and Iran as well (Derenko et al. 2013). Interestingly, caste groups such as Reddy, Bhargava and Thogataveera (Palanichamy et al. 2004) also carry traces of R30 subclades, strengthening the hypothesis of their ancestral links with the Near East. The R30b2 subclade found in the Kavaratti islanders also occurs in scheduled caste populations in the Punjab and in Jat Sikhs (Singh et al. 2021). All of this suggests that the founder haplogroup R30b2 must have entered the Lakshadweep islands from the mainland during the LGM recovery phase when the sea level was reduced and the island was closer to the Indian subcontinent (Clark et al. 2009; Kim et al. 2023).
Besides M2 and R30, the only other haplogroup found in our samples with a time to the most recent common ancestor anterior to the LGM is M33. Like M2 and R30, the M33 cluster is unique in Kavaratti. Moreover, this clade appears to have separated from the other mainland clades ~ 50,000 years ago (ESM_1, Fig. 7). The time to the most recent common ancestor of the M33a clade found on Kavaratti Island is 33,381 years ago (95% HPD: 11,829–56,166). Unlike R30 and M2, the subclades of M33 are spread throughout the Indian subcontinent. A commonality among these three haplogroups is their deep-rooted presence in the northwestern region of the subcontinent. A Bayesian Skyline Plot shows an increase in the population size of M33 carriers ~ 19,000 years ago. Two other subclades of M, namely M3 and M30, show a post-LGM increase in the effective population size and a time to the most recent common ancestor of ~ 20,000 YBP (ESM_1 Fig. 8 and Fig. 9), suggesting that it was indeed the LGM that had driven these haplogroups southward. Haplogroup M33 is mostly found in central and western populations with the exception of Mal Pahariya. Similarly, haplogroup M30 is predominantly present in western tribes, such as the Kathodi, Mathakur, Kathakur, Khatri, Koraga, Nihal, Bhil and Andh. Haplogroup M3 peaks at between 10 and 30% in the northwestern region of the subcontinent, encompassing Pakistan, Kashmir and the Punjab. The Jenu Kuruba and Koraga tribes from the southern region show a higher frequency of M3 along with the Kamar and Kathodi from the western region. It is plausible that whilst one group reached the Lakshadweep archipelago, ancestors of the other afore-mentioned ancient tribes moved into the Indian peninsula carrying the pre-LGM genetic footprint of the northwestern region of the subcontinent, which included these M subclades. The other possibility is that the M3 cluster, which oddly shows a time to the most recent common ancestor of 1070 YBP (95% HPD: 205–2158), entered Kavaratti island between the second and twelfth century AD, as mentioned in documented history (Forbes 1979).
The western Eurasian haplogroup U4 peaks at 24% in the Volga-Ural region north of the Caspian Sea (Bermisheva et al. 2002). Haplogroup U4 is also found at varying frequencies in Central Asia, the Near East and Anatolia (Quintana-Murci 2004). Based on the comparison of the available U4 sequences, we suggest that our U4 samples, a sample from the northwestern region of the subcontinent (Singh et al. 2021) and a few ancient European samples from AmtDB (Ehler et al. 2019) belong to a lineage with 16356C and 16362C variations that are found in the ‘Uralic’ lineage of the Eastern European U4 cluster, originating around the LGM time depth (Malyarchuk 2004; Översti and Palo 2022). The time to the most recent common ancestor of the U4 cluster found in the Kavaratti islanders is ~ 13,000 YBP. One U4a2 sample from Kavaratti shows a time to the most recent common ancestor of ~ 28,000 YBP. Although these estimates are based on the control region sequences, the time depth of increase in the U4 effective population size in the vicinity of the LGM suggests that the U4 clade found on this island certainly belongs to an older lineage and is not a consequence of recent migration. Also, on the Indian subcontinent, U4 is reported only in the Punjab region (Singh et al. 2021). The rare presence of this haplogroup in the Kavaratti islanders serves as evidence for admixture between the populations from the colder north and the ancestors of this lineage around the Near East and the northwestern region of the subcontinent during the Late Pleistocene, as suggested earlier (Richards et al. 2000; Metspalu 2004; Pala et al. 2012; Silva et al. 2017). These haplogroups with their probable temporal origin in the vicinity of the LGM are an important indicator of population dynamics within the Indian subcontinent. Kavaratti island along with other islands of the archipelago were populated by aboriginals from the mainland who had reached the archipelago during the LGM recovery phase. What motivated them to move onto an island with limited resources is yet to be established. Nonetheless, it is clear that the migration caused a bottleneck, and since then this population has remained largely isolated.
Diversity statistics
Nucleotide diversity (π) in the Kavaratti population is 0.00122, whilst the haplotype diversity (Hd) is 0.993. A lower π index and a higher Hd index suggest a population bottleneck and a rapid population expansion with the accumulation of mutations. A statistically significant Tajima’s D value of − 1.92469 indicates a recent population expansion.
Genetic affinities with island populations
The Lakshadweep archipelago population is distributed over ten islands. To get an overview of the overall Lakshadweep gene pool, we compared average pairwise differences (Fst) based on haplotype frequencies (ESM_1 Table 2). The Kavaratti population (Fst = 0.858) is a more diverse group compared to Chetlat (0.324), Bitra (0.564) and Kadmat (0.526). This enhanced diversity is primarily due to the stronger founder effect of R30 on these islands. The R30 frequency on these islands is > 60%, whilst other more diverse islands including Kavaratti show < 40%. The Lakshadweep islands are different form each other in area, vegetation and human population. To assess the relationship between founder haplogroups and island area, we conducted a Pearson correlation test. Haplogroup R30 is significantly (p < 0.05) negatively correlated with Fst (r2 = − 0.80) and island area (r2 = − 0.71). The result supports the hypothesis that the earliest settlers had a high frequency of R30 when they reached the Lakshadweep archipelago. Similarly, smaller islands have accumulated this haplogroup in high frequency. Haplogroup M2 shows insignificant positive correlation with Fst (r2 = 0.13) and island area (r2 = 0.20). There is insignificant positive correlation between the island area and Fst (r2 = 0.54, p = 0.1). In other words, the greater the area, the more diverse the population.
When Kavaratti was compared with other islands based on Nei’s genetic distance (ESM_1 Fig. 1), the genetic composition of the island turned out to be more similar to Amini (d = 0.1208), Minicoy (0.1353) and Kalpēni (0.1649). This similarity may be due to the close proximity of these islands with Kavaratti. We also constructed a Neighbour Joining Tree to understand the clustering pattern (ESM_1 Fig. 2). Besides Amini, Minicoy and Kalpēni, Āndrōttŭ too clustered closer to Kavaratti. Based on these results, it can be concluded that genetic admixture between Kavaratti and these islands occurred more frequently compared to other more distant islands. Interestingly, being a diverse group, Kavaratti and Amini retained the founder haplogroup R30 at ~ 30%, whilst Minicoy and Kalpēni lost this haplogroup gradually. Conversely, it can also be hypothesised that the northern islands including Amini, Agatti, Kiltān and Kavaratti were explored first by the R30 founders, and Minicoy was the latest. Fine structure analysis of all these islands would provide a better understanding of the time depth of migration and settlement.
We performed AMOVA grouping of the islands based on geography, founder haplogroup frequency and area (ESM_1 Table 3). We did not find any notable differences leading towards population differentiation or formation of structured populations. This finding clearly indicates that these islands, though largely isolated from the mainland, had exchanges amongst themselves, and the founder effect is relatively uniform on these islands.
Genetic affinities with mainland populations
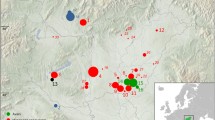
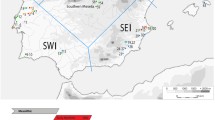
To reconstruct the maternal ancestral lineage of the Kavaratti islanders, we compared their gene pool with that of the mainland populations (Fig. 2). We used Principal Component Analysis (PCA) to visualise the clustering pattern amongst them (Fig. 3). Interestingly, Kavaratti clustered along with the Nair, neighboured by Kodava, Madia, Koraga, Kathodi and the Punjab Scheduled Castes. This cluster lies exactly in between the Western-Central-Eastern cluster and Northern-Northwestern cluster. A similar clustering pattern is seen in the neighbour joining (NJ) Tree plotted, based on Hamming’s dissimilarity and an Andamanese root (Fig. 3).
In the neighbour joining tree, Kavaratti clustered with the Punjab Scheduled Castes. This clustering is due to the R30, M2, M3, M30 and L haplogroups present in these populations. No significant affinity obtains between the Kavaratti and Śrī Laṅkan populations. However, the indigenous Śrī Laṅkan tribe called Vedda clustered more closely to Kavaratti than the general population of the island. We observed a similar pattern when larger datasets (1000 genome) were included in the Phylogeny Tree (Fig. 4). In the Phylogeny Tree, the Kavaratti islanders (KAV) clustered with Indo-European speaking language communities.
The multivariate analysis, therefore, suggests that the Kavaratti population belonged to a unique wave of migrants that neither belonged to the earliest Indian-specific M2 carriers nor to the West Eurasian descendants. A peculiar feature in the Kavaratti cluster is the presence of the M3, M30 and U2 clades. Haplogroup U2 peaks in the northwestern and northern region of the Indian subcontinent, like M3, M5 and M30. Out of these, M3 and M30 originated during or after the LGM event. This time depth is important, since the members of the Kavaratti cluster carry M3 and M30, leading us to the assumption that these populations had similar gene pools during the LGM recovery, after which bottleneck events took place. At this juncture, if an R30 bottleneck happened, then it is obvious that the group that migrated to Kavaratti belonged to a bigger group that included the ancestors of the Urali Kuruman, the Punjab Scheduled Castes, Uttar Pradesh Brahmins, Thogataveera, Reddy, Jat Sikh, Nair, etc. Therefore, we hypothesise that the Kavaratti islanders trace their maternal ancestry to the northwestern region of the Indian subcontinent. To resolve this picture further, based on the subclade information available for the R30b2 and R30 haplogroup network analysis (ESM_1 Fig. 3) and principal component analysis (Fig. 2), we suggest that the Kavaratti islanders are maternally related to Mazhabi Sikhs, Balmikis (Punjab Scheduled Castes) (Singh et al. 2016) and Nairs. All other groups carry R30a subclades. Interestingly, Nairs have two important characteristics which are similar to Lakshadweep islanders, firstly their Malabar roots, as described in oral traditions and documented history, and secondly the high status enjoyed by women (Forbes 1979). However, if the Kavaratti islanders were recent offshoots of the Nairs, then it would most likely have been mentioned in folklore. At present, no record of any direct Nair lineage to the Lakshadweep islanders, either written or oral, is known to exist (Chib 1985).
Our analyses limit our ability to trace the route of migration, as we do not find any population with R30b2 between the Punjab and Kerala. However, there are two possible routes, viz. (1) The modern route from the Malabar coast, and (2) a hypothetical “non-Malabar route”. The Malabar route is more plausible, because many indigenous tribes with founder haplogroups (Jenu Kuruba, Betta Kuruba, Urali Kuruman, Koraga, etc.) exist along the western coast of India. The lineage of the Kavaratti islanders may have migrated in isolation, following a coastal route until the bearers of this lineage explored the Lakshadweep archipelago. Fine-scale structuring using whole genome data will provide a better understanding of the deep-rooted ancestral links between the Malabar population and the Lakshadweep islanders to confirm lineage sharing between the two.
In conclusion, we identify in the Kavaratti islanders an R30b2 founder lineage arising from the northwestern region of the Indian subcontinent during the Last Glacial Maximum period. The only other populations to carry this lineage are the Scheduled Castes from the Punjab, Jat Sikhs and Nairs, who are either found in the northwestern portion of the Indian subcontinent or have recent historical links with this region. In addition, we also find unique clusters of pre-LGM M2 and M33 clades sharing common ancestors with those found in the northwestern populations. A similar trend is seen in the clustering pattern based on the M3 and M30 haplogroups that are prevalent in this region and date to the LGM period. We also report a rare occurrence of the U4 lineage dated to the LGM period that is otherwise only found in Europe. Its presence in the Kavaratti islanders only suggests that the ancestors of the founders admixed with a diverse group before drastic demographic changes that occurred during and after the LGM. Our conclusions are based on the control region of the mitochondrial DNA and, therefore, must be further substantiated with whole mitogenomes.
Data availability
The datasets generated during and/or analysed during the current study are available in the supplementary file.
References
Anthropological Survey of India (2021) Genomic diversity in people of India: focus on mtDNA and Y-Chromosome polymorphism. Springer, Singapore
Arrian H, Plutarch S (1533) Arriani & Hannonis periplus, Plutarchvs de fluminibus & montibus, Strabonius epitome. Froben, Basel
Barnett LD (1915) ‘Inscriptions at Narendra’, Epigraphia Indica and Record of the Archæological Survey of India. XIII:298–326
Bermisheva MA, Tambets K, Villems R, Khusnutdinova EK (2002) Diversity of mitochondrial DNA haplogroups in ethnic populations of the Volga-Ural region. Mol Biol 36:802–812. https://doi.org/10.1023/A:1021677708482
Brandini S, Bergamaschi P, Cerna MF et al (2018) The Paleo-Indian entry into South America according to mitogenomes. Mol Biol Evol 35:299–311. https://doi.org/10.1093/molbev/msx267
Capodiferro MR, Aram B, Raveane A et al (2021) Archaeogenomic distinctiveness of the Isthmo-Colombian area. Cell 184:1706-1723.e24. https://doi.org/10.1016/j.cell.2021.02.040
Chandrasekar A, Kumar S, Sreenath J et al (2009) Updating phylogeny of mitochondrial DNA macrohaplogroup M in India: dispersal of modern human in South Asian Corridor. PLoS ONE 4:e7447
Chaubey G (2013) Language isolates and their genetic identity: a commentary on mitochondrial DNA history of Sri Lankan ethnic people: their relations within the island and with the Indian subcontinental populations. J Hum Genet. https://doi.org/10.1038/jhg.2013.122
Chib SS (1985) Caste tribes and culture of India: Andaman Nicobar and Lakshadweep Islands, vol IX. ESS Publications, New Delhi
Clark PU, Dyke AS, Shakun JD et al (2009) The Last Glacial Maximum. Science (80-) 325:710–714. https://doi.org/10.1126/science.1172873
Connell JR, Benton MC, Lea RA et al (2022) Pedigree derived mutation rate across the entire mitochondrial genome of the Norfolk Island population. Sci Rep 12:6827. https://doi.org/10.1038/s41598-022-10530-3
Derenko M, Malyarchuk B, Bahmanimehr A et al (2013) Complete Mitochondrial DNA Diversity in Iranians. PLoS ONE 8:e80673. https://doi.org/10.1371/journal.pone.0080673
Dür A, Huber N, Parson W (2021) Fine-Tuning Phylogenetic Alignment and Haplogrouping of mtDNA Sequences. Int J Mol Sci 22:5747
Ehler E, Novotný J, Juras A et al (2019) AmtDB: a database of ancient human mitochondrial genomes. Nucleic Acids Res 47:D29–D32. https://doi.org/10.1093/nar/gky843
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567. https://doi.org/10.1111/j.1755-0998.2010.02847.x
Fleet JF (1870) Some further inscriptions relating to the Kâdamba kings of Goa. Journal of the Bombay Branch of the Royal Asiatic Society IX (XXVII):262–313
Forbes ADW (1979) Sources towards a history of the Laccadive Islands. South Asia J South Asian Stud 2(1–2):130–150. https://doi.org/10.1080/00856407908722989
Fornarino S, Pala M, Battaglia V et al (2009) Mitochondrial and Y-chromosome diversity of the Tharus (Nepal): a reservoir of genetic variation. BMC Evol Biol 9:154. https://doi.org/10.1186/1471-2148-9-154
Gabriel T (1988) Caste conflict in Kalpeni Island. Bull School Orient Afr Stud LI (3):489–499
Grant WAS, Bowen BW (1998) Shallow population histories in deep evolutionary lineages of marine fishes: insights from sardines and anchovies and lessons for conservation. J Hered 89:415–426. https://doi.org/10.1093/jhered/89.5.415
Gray A (1889) Early Notices of the Maldives, Vol. II, Part II of The Voyage of François Pyrard of Laval to the East Indies, the Maldives, the Moluccas and Brazil, translated into English from the third French edition of 1619 (3 volumes). Hakluyt Society, London, pp 423–492
Hammer Ø, Harper DAT, Ryan PDR (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4:4–9
Huber N, Parson W, Dür A (2018) Next generation database search algorithm for forensic mitogenome analyses. Forensic Sci Int Genet 37:204–214. https://doi.org/10.1016/j.fsigen.2018.09.001
Hughes PD, Gibbard PL (2015) A stratigraphical basis for the last glacial maximum (LGM). Quat Int 383:174–185. https://doi.org/10.1016/j.quaint.2014.06.006
Kim HL, Li T, Kalsi N et al (2023) Prehistoric human migration between Sundaland and South Asia was driven by sea-level rise. Commun Biol 6:150. https://doi.org/10.1038/s42003-023-04510-0
Kumar S, Padmanabham P, Ravuri RR et al (2008) The earliest settlers’ antiquity and evolutionary history of Indian populations: evidence from M2 mtDNA lineage. BMC Evol Biol 8:230. https://doi.org/10.1186/1471-2148-8-230
Leigh JW, Bryant D (2015) POPART: Full-feature software for haplotype network construction. Methods Ecol Evol 6:1110–1116. https://doi.org/10.1111/2041-210X.12410
Malyarchuk BA (2004) Differentiation of the mitochondrial subhaplogroup U4 in the populations of Eastern Europe, Ural, and Western Siberia: implication to the genetic history of the Uralic populations. Russ J Genet 40:1281–1287. https://doi.org/10.1023/B:RUGE.0000048671.32870.cb
Malyarchuk B, Derenko M, Grzybowski T et al (2010) The peopling of Europe from the mitochondrial haplogroup U5 perspective. PLoS ONE 5:16–20. https://doi.org/10.1371/journal.pone.0010285
McCrindle JW (1878) The Commerce and Navigation of the Erythræan Sea, being a Translation of Periplus Maris Erythræi by an Anonymous Writer, and of Arrian’s Account of the Voyage of Nearkhos from the Mouth of the Indus to the Head of the Persian Gulf. Thacker, Spink & Co., Calcutta
Metspalu M (2004) Most of the extant mtDNA boundaries in south and southwest Asia were likely shaped during the initial settlement of Eurasia by anatomically modern humans. BMC Genet. https://doi.org/10.1186/1471-2156-5-26
Moletius I (1562) Geographia Cl. Ptolemæi Alexandrini. Apvd Vincentivm Valgrisvm, Venetiis, p 264
Mustak MS, Rai N, Naveen MR et al (2019) The peopling of Lakshadweep Archipelago. Sci Rep 9:1–7. https://doi.org/10.1038/s41598-019-43384-3
Nobbe CFA (1845) Claudii Ptolemæi Geographia, Tom. II. Sumptibus et typis Caroli Tauchnitii, Lipsiæ
Okonechnikov K, Golosova O, Fursov M et al (2012) Unipro UGENE: a unified bioinformatics toolkit. Bioinformatics 28:1166–1167. https://doi.org/10.1093/bioinformatics/bts091
Olivieri A, Sidore C, Achilli A et al (2017) Mitogenome diversity in Sardinians: a genetic window onto an Island’s Past. Mol Biol Evol 34:1230–1239. https://doi.org/10.1093/molbev/msx082
Översti S, Palo JU (2022) Variation in the substitution rates among the human mitochondrial haplogroup U sublineages. Genome Biol Evol 14:evac097. https://doi.org/10.1093/gbe/evac097
Pala M, Olivieri A, Achilli A et al (2012) Mitochondrial DNA signals of late glacial recolonization of Europe from Near Eastern Refugia. Am J Hum Genet 90:915–924. https://doi.org/10.1016/j.ajhg.2012.04.003
Palanichamy MG, Sun C, Agrawal S et al (2004) Phylogeny of mitochondrial DNA macrohaplogroup N in India, based on complete sequencing: implications for the peopling of South Asia. Am J Hum Genet 75:966–978. https://doi.org/10.1086/425871
Quintana-Murci L (2004) Where west meets east: the complex mtDNA landscape of the southwest and Central Asian corridor. Am J Hum Genet. https://doi.org/10.1086/383236
Ranaweera L, Kaewsutthi S, Win Tun A et al (2014) Mitochondrial DNA history of Sri Lankan ethnic people: their relations within the island and with the Indian subcontinental populations. J Hum Genet 59:28–36. https://doi.org/10.1038/jhg.2013.112
Richards M, Macaulay V, Hickey E et al (2000) Tracing European founder lineages in the near Eastern mtDNA pool. Am J Hum Genet 67:1251–1276. https://doi.org/10.1016/S0002-9297(07)62954-1
Rozas J, Ferrer-Mata A, Sanchez-DelBarrio JC et al (2017) DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol Biol Evol 34:3299–3302. https://doi.org/10.1093/molbev/msx248
Schoff WH (1912) The periplus of the erythræan sea: travel and trade in the Indian Ocean by a merchant of the first century. Longmans, Green and Co, London
Shamoon-Pour M, Li M, Merriwether DA (2019) Rare human mitochondrial HV lineages spread from the Near East and Caucasus during post-LGM and Neolithic expansions. Sci Rep 9:14751. https://doi.org/10.1038/s41598-019-48596-1
Silva M, Oliveira M, Vieira D et al (2017) A genetic chronology for the Indian Subcontinent points to heavily sex-biased dispersals. BMC Evol Biol 17:88. https://doi.org/10.1186/s12862-017-0936-9
Singh G, Talwar I, Sharma R et al (2016) Genetic differentiation and population structure of five ethnic groups of Punjab (North-West India). Mol Genet Genom 291:2055–2063. https://doi.org/10.1007/s00438-016-1239-3
Singh G, Yellapu S, Sandhu HS et al (2021) Genetic characterisation of the North-West Indian populations: analysis of mitochondrial DNA control region variations. Ann Hum Biol 48:166–172. https://doi.org/10.1080/03014460.2021.1879933
Sylvester C, Krishna MS, Rao JS, Chandrasekar A (2019) Maternal genetic link of a south Dravidian tribe with native Iranians indicating bidirectional migration. Ann Hum Biol 46:175–180. https://doi.org/10.1080/03014460.2019.1599067
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Tripati S (2006) ‘Ships on hero stones from the west coast of India. Int J Naut Archaeol 35:88–96. https://doi.org/10.1111/j.1095-9270.2005.00081.x
van Oven M, Kayser M (2009) Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Hum Mutat 30:E386–E394. https://doi.org/10.1002/humu.20921
Weissensteiner H, Pacher D, Kloss-Brandstätter A et al (2016) HaploGrep 2: mitochondrial haplogroup classification in the era of high-throughput sequencing. Nucleic Acids Res 44:W58–W63. https://doi.org/10.1093/nar/gkw233
Acknowledgements
This work is supported by the Science and Engineering Research Board (SERB) of the Department of Science and Technology of the Government of India (EEQ/2021/000465) in New Delhi. AMT acknowledges the Indian Council for Cultural Relations (ICCR) for providing a fellowship. The authors thank Dr. Kumarasamy Thangaraj from the Centre for DNA Fingerprinting (CDFD) at Hyderabad for his support and guidance. The authors are grateful to the Lakshadweep Administration, the Department of Science and Technology (DST) and the Directorate of Health Services of Kavaratti Island for their support during this study. The authors also acknowledge the participants, lab technicians and research staff involved in the study.
Funding
This study was funded by the Science and Engineering Research Board (SERB), Department of Science and Technology, Government of India (EEQ/2021/000465), New Delhi, India.
Author information
Authors and Affiliations
Contributions
AMT and JJS have contributed equally in conceptualisation, mtDNA analysis, statistical analysis and lab work. MSM and IB has collected the blood samples and outlined the manuscript preparation. LK evaluated the analysis. GvD, MSM and IB have reviewed the manuscript. All the authors have contributed to the manuscript and approved the final draft.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no competing interests to declare that are relevant to the content of this article.
Ethics approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Institutional Human Ethics Committee of Mangalore University (MU/AZ/20/IHEC/2014-15).
Consent to participate
Informed consent was obtained from all individual participants included in the study.
Consent to publish
Participants signed informed consent regarding publishing their data anonymously.
Additional information
Communicated by Shuhua Xu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tayyeh, A.M., Sequeira, J.J., Kumar, L. et al. The maternal ancestry of the Kavaratti islanders and the last glacial maximum aftermath. Mol Genet Genomics 298, 1467–1477 (2023). https://doi.org/10.1007/s00438-023-02072-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-023-02072-8