Abstract
The genus Actinidia, also called kiwifruit, is characterized with abundant balanced nutritional metabolites, including exceptionally high vitamin C content. However, the traditional classification could not fully reflect the actual Actinidia species’ relationships, which need further revision through more accurate approaches. Compared to the nuclear genome, the chloroplast genome has simple heredity characteristics, conserved genome structure and small size, suitable for deciphering complicated species’ phylogenetic relationships. Here, the genome-wide comprehensive comparative analyses were performed over 29 independent chloroplast genomes’ sequences derived from 25 Actinidia taxa. The average genome size is 156,673.38 bp, with an average 37.20% GC content. The long repeat sequences rather than SSRs (simple sequence repeats) in Actinidia were revealed to be the causal agent leading to the chloroplast genome size expansion. The clpP gene sequences with exon merge and intron deletion were annotated in all the 29 chloroplast genomes tested, which has been previously reported to be lost in Actinidia species. Comprehensive sequence analyses indicated the distinct variation at the clpP gene locus was Actinidiaceae-specific, emerging after the Actinidiaceae-other Ericales species divergence. Four highly divergent sequences (i.e., rps16 ~ trnQ-UUG, rps4 ~ trnT-UGU, petA ~ psbJ, and rps12 ~ psbB) evolved in the LSC (large single-copy) and SSC (small single-copy) regions embodying rps12 ~ psbB (including clpP gene and its up/downstream noncoding sequence) were identified as variation hot spots in Actinidia species. Based on either LSC region alone, combined sequences of LSC and SSC or the whole chloroplast genome sequences, three identical phylogenetic trees of the 25 Actinidia taxa with relatively improved resolution were reconstructed, consistently supporting the reticulate evolutionary lineage in Actinidia. Our findings could help to better understand the evolution characteristics of chloroplast genomes and phylogenetic relationships among Actinidia species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Actinidia, commonly known as kiwifruit, ‘the king of fruits’, includes economically important horticultural species, such as A. chinensis and A. chinensis var. deliciosa that have been extensively cultivated worldwide (Testolin et al. 2016). The genus Actinidia, together with other two sister genera, Clematoclethra and Saurauia, belongs to Actinidiaceae that is located on the basal asterids, Ericales. Based on the morphological characteristics of fruit, pith and hair, Actinidia has been classified into four intrageneric sections, Leiocarpae (Lei), Maculatae (Mac), Stellatae (Ste), and Strigosae (Str) (Chat et al. 2004; Testolin et al. 2016). Given the traditional classification system could poorly reflect the actual relationships among Actinidia species, more accurate approaches need to be recruited to define their evolutionary lineage (Liu et al. 2017; Tang et al. 2019b).
Meanwhile, molecular approaches have been employed to establish distinct phylogenetic relationships for Actinidia taxa by using makers of RAPD (randomly amplified polymorphic DNA) (Huang et al. 2002), ITS (internal transcribed spacers) (Li et al. 2002), and/or sequence fragments from chloroplast and mitochondrion genomes (Chat et al. 2004). Due to lack of genome-wide sequences, with few markers including limited nucleotide information, the reconstructed phylogenetic relationships remain either incompletely resolved or weakly supported. Nevertheless, along with public releases of nuclear genome of A. chinensis “Hongyang” (Huang et al. 2013), using genome-wide SNPs (single nucleotide polymorphisms), an improved phylogenetic tree of 26 Actinidia species was reconstructed, clustering into five main groups (Liu et al. 2017), including A. chinensis complex, A. arguta complex, the A. polygama, A. rufa clade and other hairy and/or spotted fruit taxa. More recently, a comprehensive phylogenetic relationship was reconstructed on the basis of four noncoding intergenic sequences alone from chloroplast genomes of 59 Actinidia taxa (Tang et al. 2019b).
However, the subdivisions in Actinidia based on molecular phylogenetic relationships are apparently in conflict with morphological classification. Molecular phylogenetic reconstructions demonstrated that the four morphologically defined intrageneric sections were not monophyletic, probably because of natural interspecific hybridization/introgression facilitated by the sympatric distributions of Actinidia species (Chat et al. 2004; Liu et al. 2017).
Compared to nuclear genomes, the plants’ chloroplast genomes are more suitable for deciphering phylogenetic relationships in the complicated plant families, due to the hereditary characteristics, conserved genome structure and small size (Martin et al. 2005; Daniell et al. 2016). The land plants’ chloroplast genomes are mainly inherited from maternal parents and possess a highly conserved genome structure with four independent parts, including an LSC (large single-copy) region, an SSC (small single-copy) region, and two separated inverted repeat regions (IRa and IRb) between LSC and SSC (Daniell et al. 2016).
In this study, using the chloroplast genome sequences of 137 Ericales species downloaded from the NCBI genome database (http://www.ncbi.nlm.nih.gov/genome), including 25 Actinidia species available for A. zhejiangensis (Ai and Liu 2019), A. callosa var. henryi (Wu et al. 2019), A. callosa var. strigillosa (Liu et al. 2020), A. chinensis (Yao et al. 2015), A. chinensis var. deliciosa (Yao et al. 2015), A. chinensis var. setosa (Lin et al. 2019), A. lanceolata (Zhang and Liu 2019), A. arguta (Li et al. 2018; Lin et al. 2018), A. arguta var. giraldii (Ding et al. 2021), A. eriantha (Tang et al. 2019a), A. kolomikta (Lan et al. 2017), A. polygama (Wang et al. 2016), A. tetramera (Wang et al. 2016), A. rufa (Kim et al. 2018), A. valvata (Chen et al. 2020; Lin et al. 2020), A. cylindrica var. cylindrica, A. cylindrica var. reticulata, A. styracifolia (Yang et al. 2020), A. macrosperma (Chen et al. 2019), A. fulvicoma (Zhang et al. 2019), A. hubeiensis, A. hemsleyana (Xiaoqiong et al. 2021), A. indochinensis, A. latifolia, and A. rubus (Xu et al. 2020), the chloroplast genomes’ characteristics and divergent regions, as well as the evolutionary lineage were explored by comprehensive genome-wide comparative analyses in terms of genome structure, gene organization, boundaries between IR, SSC and LSC regions, SSRs (simple sequence repeats), long repeat sequences and sequence synteny and diversity. Interestingly, a seemingly widespread clpP gene loss event reported previously (Yao et al. 2015; Wang et al. 2016) was carefully inspected, and its bona fide existence and expression were redefined. Based on LSC, LSC plus SSC regions’ sequences or complete chloroplast genome sequences, distinct phylogenetic relationships among 25 Actinidia taxa were reconstructed, respectively. Our findings would provide insights for refining evolutionary relationships among Actinidia taxa, and potential molecular markers to further resolve the complicated phylogenetic lineage in genus Actinidia.
Materials and methods
The chloroplast genome data sets
The complete chloroplast genome sequences of 137 species from Ericales, including 25 Actinidia species, were downloaded from NCBI genome database. The detailed information of chloroplast genomes was listed in Table S1.
Genome structure, gene organization and repeat sequences
The genes in each chloroplast genome sequences were re-annotated using PGA (Plastid Genome Annotator) (Qu et al. 2019), GeSeq (Tillich et al. 2017) and CPGAVAS2 (Shi et al. 2019), respectively. Subsequently, the annotation results from three programs were merged. The gene organization, including total gene number, gene copy and intron number in each gene, was analyzed using our Python scripts.
SSRs were detected using MISA Perl script with thresholds of 10, 6, 5, 5, 5 and 5 repeats as a unit, respectively, for mono-, di-, tri-, tetra-, penta-, and hexanucleotide SSRs. Long repeats, including forward, reverse, palindromic and complement repeats, in Actinidia chloroplast genomes were identified using REPuter (Kurtz et al. 2001). For all repeat types, the Hamming distance was 3, which meant that two repeat copies had at least 90% similarity. The minimum repeat length was 30 bp, and the maximum number of repeated sequences displayed was 1,000.
Comparative analyses of boundaries between LSC, SSC and IR regions
Mummer 3.0 (Delcher et al. 2003) was used to align each chloroplast genome sequence to itself, to confirm the boundaries between the LSC, SSC and IR regions. If the inverted repeat region is not 100% similar, we manually adjust the position of the inverted repeat region based on Mummer’s alignment results. The boundaries’ visualization between LSC, SSC and IR regions was implemented using the SVG module in Perl.
Identification of hypervariable regions
The aligned chloroplast genome sequences of Actinidia species were imported into program DnaSP (Rozas et al. 2017) to calculate the nucleotide polymorphism. In sliding window analysis, the window length and step size were set to 600 bp and 200 bp, respectively. Meanwhile, the multiple sequence alignment of the 25 Actinidia species’ complete chloroplast genomes was also visualized using mVISTA (Frazer et al. 2004).
Analyses of clpP gene sequence and its surrounding syntenic region
Using clpP encoding protein sequence in other Ericales species as query, the tBlastn analyses were performed against the chloroplast genome sequences of 25 Actinidia species. The ORFs (open reading frames) were predicted in the similar nucleotide sequence in each tested Actinidia species, respectively. The multiple sequence alignment of predicted clpP encoding protein sequence in 25 Actinidia species, and another two Actinidiaceae species, Saurauia tristyla in genus Saurauia, and Clematoclethra scandens in genus Clematoclethra were performed using MAFFT (Katoh et al. 2019).
The syntenic regions surrounding clpP gene were retrieved from 25 Actinidia species’ chloroplast genome sequences and subsequently compared. The genes distributed in the syntenic regions were visualized using SVG module in Perl.
Phylogenetic relationship reconstruction
The phylogenetic tree among 25 Actinidia species including 27 independent chloroplast genome sequences was reconstructed, using two Actinidiaceae species, S. tristyla in genus Saurauia and C. scandens in genus Clematoclethra, as an outgroup (Table S1). The ML (maximum likelihood) phylogenetic tree was constructed, using whole chloroplast genome sequences, LSC and LSC plus SSC regions’ sequences, respectively.
Additionally, three other phylogenetic trees were reconstructed with 29 independent chloroplast genome sequences derived from 25 Actinidia species, including an additional two sequences from A. chinensis “AC017” (tetraploid) (Genbenk accession number: KP297243) and A. chinensis var. deliciosa “AD019” (hexaploid) (Genbenk accession number: KP297245), respectively. The ML (Maximum likelihood) phylogenetic tree was constructed, using whole chloroplast genome sequences, and sequences of LSC alone and LSC plus SSC regions, respectively.
In each phylogenetic relationship analysis, the nucleotide sequences were aligned by MAFFT and subsequently adjusted by trimAl (Capella-Gutierrez et al. 2009). The tree construction was performed by IQ-TREE 1.6.12 (Nguyen et al. 2015) with 1000 bootstrap replicates. The suitable model for each tree construction was determined by ModelFinder (Kalyaanamoorthy et al. 2017) integrated in IQ-TREE 1.6.12 (Nguyen et al. 2015).
Results
The summary of chloroplast genomes in Actinidia species
The 29 independent chloroplast genome sequences of 25 Actinidia species were downloaded from NCBI genome database, including A. zhejiangensis, A. callosa var. henryi, A. callosa var. strigillosa, A. chinensis, A. chinensis var. deliciosa, A. chinensis var. setosa, A. lanceolata, A. arguta, A. arguta var. giraldii, A. eriantha, A. kolomikta, A. polygama, A. tetramera, A. rufa, A. valvata, A. cylindrica var. cylindrica, A. cylindrica var. reticulata, A. styracifolia, A. macrosperma, A. fulvicoma, A. hubeiensis, A. hemsleyana, A. indochinensis, A. latifolia, and A. rubus (Table 1, Table S1). In our study, there are four independent chloroplast genomes of A. chinensis, from three diploids, “AC011”, “Hongyang”, and “Jinguo”, and one tetraploid “AC017”, respectively. Two independent chloroplast genomes were also collected in A. chinensis var. deliciosa, from “AD006” (tetraploid) and “AD019” (hexaploid). The detailed information of chloroplast genomes for 25 Actinidia species is presented in Table 1, including species name, Genbank accession number, size of LSC, SSC, IR region or whole chloroplast genome, number of genes coding for proteins, tRNAs or rRNAs, as well as GC content.
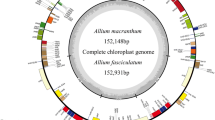
The Actinidia species’ chloroplast genomes comprised four independent parts, i.e., LSC, SSC, IRa and IRb (Fig. 1). Among the 25 Actinidia species, the average genome size is 156,673.38 bp, with an average 37.20% GC content. A. indochinensis has the smallest genome size (155,931 bp), while the largest genome size in A. tetramera is up to 157,659 bp with the lowest GC content (37.03%) (Table 1, Fig. 1).
Genome map representing the chloroplast genome structure in Actinidia species. The PCG genes in different categories, tRNA and rRNA are labeled in different color box, respectively. The genes located in inner and outer circle represent the location in plus and minus DNA strand, respectively. Red characters represent gene copy number variation. The bold in italics represents variation of introns’ number
Additionally, the genome size of A. zhejiangensis (156,717 bp) and A. callosa var. henryi (156,826 bp) is close to those in most other Actinidia species, but these two genomes encode the smallest number of genes (128 genes). Seemingly there is no association between genome size and gene number in chloroplast genome sequences of the 25 Actinidia species (Table 1).
Gene content and exon–intron structure in Actinidia species’ chloroplast genomes
The genes encoded by chloroplast genome include three types, PCG (protein-coding gene), tRNA and rRNA (Fig. 1). Except A. rubus (82 genes), A. styracifolia (82) and A. zhejiangensis (82), the other 22 Actinidia species have 83 or 84 or 85 PCGs. Additionally, the tRNA genes number varies from 37 to 41 (Table 1). As illustrated in Fig. 1 and Table 2, gene doubling took place at loci of all the four rRNAs, tRNAs and PCGs, including rps12, ndhB, psbA, ycf2, ycf15, trnA-UGC, trnI-CAU, trnI-GAU, trnL-CAA, trnN-GUU, trnR-ACG, trnV-GAC, trnH-GUG, trnM-CAU, and trnfM-CAU (Table 2). Furthermore, rps12, psbA, ycf15, trnH-GUG, trnM-CAU, and trnfM-CAU were doubled in several Actinidia species. These analyses suggested that a considerable portion of total gene number variation might be evolved from gene doubling in the chloroplast genomes of the 25 Actinidia species (Table 2).
The analysis on exon–intron structures showed that most of PCGs, tRNAs and rRNAs simply contained a single exon without intron, while a few genes had one or two introns (Table 2). Seven tRNAs (trnA-UGC, trnI-GAU, trnL-UAA, trnV-UAC, trnG-UCC, trnG-GCC and trnK-UUU) and 11 PCGs (rps12, rps16, rpl2, rpl16, rpoC1, ndhA, ndhB, petB, petD, atpF and ycf2) have one intron. By contrast, ycf3, a PCG gene, contains two introns with a more complicated exon–intron structure (Table 2). Nevertheless, some orthologs are divergent in exon–intron structure among different Actinidia species, including two PCGs, rps16 (Fig. S1) and petB (Fig. S2). For example, petB in A. callosa var. henryi has no intron, whereas the orthologous gene in other 24 Actinidia taxa contains one intron (Fig. S2).
Furthermore, the gene member expansion seems to be associated with varied exon–intron structure. The ycf2 has been doubled in 25 Actinidia species, both copies containing an intron in A. kolomikta (Fig. S3) in contrast to no intron existing in the both copies of other 24 Actinidia species. Twenty one out of 25 Actinidia species have two copies of rps12, each containing an intron (Fig. S4). Interestingly, cultivars ‘Hongyang’ and ‘Jinguo’ have two copies of rps12 instead of a single copy found in ‘AC011’ and ‘AC017’, although they all belong to A. chinensis.
The exon merge and loss of intron of clpP gene in Actinidia species’ chloroplast genomes
The clpP gene coding for the proteolytic subunit of Clp protease has been reported to be completely lost in the chloroplast genomes of Actinidia and other Actinidiaceae species and implicated to be transferred into nucleus in A. chinensis during chloroplast evolution (Yao et al. 2015).
To test whether clpP gene loss is synapomorphy in Actinidia genus or even other Actinidiaceae species, the clpP gene sequence was searched in the chloroplast genomes of 25 Actinidia species and another two Actinidiaceae species, S. tristyla in genus Saurauia and C. scandens in genus Clematoclethra. Consequently, NCBI genome annotation files of the tested 27 Actinidiaceae species indicated that a clpP gene was present only in S. tristyla, containing two exons and an intron. Subsequently, using the protein sequence of clpP gene in other Ericales species as query, the tBlastn analyses indicated that DNA sequence fragments showing high similarity were identified in 25 Actinidia species and C. scandens. The ORF (open reading frame) analyses indicated just two exons existed in 25 Actinidia species and C. scandens (Fig. 2, Table S2), encoding a 196–208 aa length clpP protein (Fig. 3). Additionally, using the predicted exon sequence of clpP gene in A. chinensis “AC011” as query, the matched Illumina raw reads from fruits and leaves transcriptome in SRA database could be identified through Blast analyses (Table S3), suggesting clpP gene may be constitutively expressed in Actinidia taxa.
To track the evolutionary variations of clpP gene, the clpP gene structures were compared among the 27 Actinidiaceae taxa and other 106 Ericales species with sequenced chloroplast genomes downloaded from NCBI Genbank database. However, multiple sequence alignment of the 137 clpP encoding protein sequences in Ericales species demonstrated that amino acids’ variation in clpP encoding protein upstream sequences just occurred in Actinidiaceae species, including 25 Actinidia species, and C. scandens, with the 19–31 upstream amino acids residues varied (Fig. S5).
Subsequently, compared to those in 27 Actinidiaceae species, the other 104 Ericales species’ clpP genes have three exons and two introns, except those in Huodendron biaristatum (1 exon) and Alniphyllum pterospermum (two exons), respectively (Table S2). For the 104 clpP members with three exons and two introns, additional Blast analyses showed the second and third exon merged with intron loss in 27 tested Actinidiaceae species.
Interestingly, the first intron’s sequences of the 104 clpP members could also be traced around the intron sequences of clpP genes in 25 Actinidia species and C. scandens (Fig. 2, Fig. S6, Table S4), but absent in S. tristyla. Comprehensive Blast analyses in NCBI Nt and Nr database showed the varied clpP sequence including two exons is Actinidiaceae-specific, implicating the exon merge and losses of intron in clpP gene might occur after the Actinidiaceae-other Ericales species divergence.
Boundaries between IR, SSC and LSC region in Actinidia species’ chloroplast genomes
Generally, there were mainly three different types of boundaries between IR, LSC or SSC regions in Actinidia species with little difference (Fig. 4, Fig. S7). Type I was found in A. arguta, A. arguta var. giraldii, A. chinensis “AC011”, A. chinensis “AC017”, A. chinensis “Jinguo”, A. chinensis “Hongyang”, A. chinensis var. deliciosa “AD006”, A. chinensis var. deliciosa “AD019”, A. cylindrica var. cylindrica, A. indochinensis, A. eriantha, A. hemsleyana, A. kolomikta, A. polygama, A. rubus, A. rufa, A. styracifolia, A. valvata, A. macrosperma, A. tetramera, A. zhejiangensis and A. fulvicoma (red labeled in Fig. 4, Fig. S7). Among Type I members, each trnN is located in IRa and IRb region, respectively, close to SSC region. Additionally, ycf1 resides at the overlapping region of SSC and IRa (Fig. 4, Fig. S7). Type II was detected in A. chinensis var. setosa, A. latifolia, and A. valvata (green labeled in Fig. 4, Fig. S7). In Type II members, ycf1 locating in SSC region is a representative characteristic.
In type III members (blue labeled in Fig. 4, Fig. S7), such as A. callosa var. strigillosa, A. hubeiensis, A. cylindrica var. reticulata and A. lanceolata, the boundary compositions between IR, LSC and SSC regions are similar to those in type II members, while a large difference is that the trnI and trnH occur at IRa/b and LSC regions besides the boundaries, respectively.
There are four species complexes in our study, including A. callosa complex (A. callosa var. henryi, A. callosa var. strigillosa), A. arguta complex (A. arguta, A. arguta var. giraldii), A. cylindrica complex (A. cylindrica var. cylindrica, A. cylindrica var. reticulata) and A. chinensis complex (A. chinensis, A. chinensis var. deliciosa, A. chinensis var. setosa). Except A. arguta complex, obvious boundary divergence could be found within the other three species complexes. A. callosa var. henryi, A. cylindrica var. cylindrica, A. chinensis and A. chinensis var. deliciosa were located in Type I (Fig. 4, Fig. S7). Whereas, A. chinensis var. setosa in Type II, and A. callosa var. strigillosa and A. cylindrica var. reticulata in type III were also observed.
SSRs in Actinidia species’ chloroplast genomes
The SSRs, including mono-, di-, tri-, tetra-, penta-, and hexanucleotide types, were analyzed in 25 Actinidia species’ chloroplast genomes and consequently, 24 (325 bp)–46 SSRs (536 bp) were identified (Fig. 5a, Table S5). A. callosa var. henryi has the largest number (46 SSRs), while A. arguta var. giraldii has the smallest (24 SSRs), respectively.
The SSR and long repeat sequences in Actinidia species’ chloroplast genomes. a The statistics of total number and size of SSR in Actinidia species. b The statistics of total number and size of long repeat sequences. c The long repeat sequences’ distribution in 70–80 kb range of the chloroplast genomes in A. tetramera, A. kolomikta, and A. callosa var. henryi, respectively
In Actinidia species, four types’ SSRs were detected, including mono-, di-, tri-, and hexa-nucleotide type. Detailed analyses indicated that the detected SSRs in Actinidia species were mainly mono-nucleotide type, accounting for 87.50–95.56% of total SSRs (Fig. S8, Table S5). Furthermore, the A/T type is the most abundant mono-nucleotide SSRs in Actinidia species, with C/G type accounting for a very small proportion (Table S5). Interestingly, the hexanucleotide SSR only exists in A. tetramera (1 SSR) and A. callosa var. henryi (1 SSR), respectively (Table S5). Our result is in accordance with previous reports that most SSRs in land plants’ chloroplasts genomes were mono- and/or di-nucleotide type, with few tri-, tetra-, penta-, and hexanucleotide type SSRs (Cui et al. 2019; Nie et al. 2019; Park et al. 2019; Huang et al. 2020; Tyagi et al. 2020). Nevertheless, compared to those in many sequenced plants’ chloroplast genomes (Cui et al. 2019), the totally detected SSRs accounted for obviously lower percentage of whole chloroplast genomes, ranging from 0.21 to 0.34% in Actinidia species.
Long repeat sequences in Actinidia species’ chloroplast genomes
A large number of long repeats, including forward, reverse, palindromic, and complementary repeats, were identified in chloroplast genomes of Actinidia species, ranging from 115 (5148 bp) to 482 (29,010 bp) (Fig. 5b, Table S6), with forward and palindromic repeats accounting for the largest portion in Actinidia species. The complementary repeats were detected only in A. callosa var. henryi, A. lanceolata and A. chinensis var. setosa, with a single copy in each species (Fig. S9, Table S6). Compared to SSRs, the total number and size of long repeat sequences in each Actinidia species largely exceeded those of SSRs, respectively (Fig. 5a, b). Similar observations were reported in Pterocarpus (Hong et al. 2020) and Aristolochia (Li et al. 2019).
The number of long repeats in A. tetramera is largely greater than that in other 24 Actinidia species (Table S6). A. tetramera has up to 482 long repeats, including 427 forward, 49 palindromic and 6 reverse repeats (Table S6). By contrast, A. lanceolata had the fewest long repeats, 115 in total, including 84 forward, 28 palindromic, 2 reverse and 1 complementary repeats (Table S6). Interestingly, 261 long repeats identified in A. chinensis var. deliciosa “AD019”, largely exceeded that of the other species in A. chinensis complex (A. chinensis, A. chinensis var. deliciosa, and A. chinensis var. setosa).
Among 25 Actinidia species, the length of the long repeat majorities is shorter than 100 bp (Table S7), predominantly ranging between 30 and 40 bp (Table S8). For long repeats’ length exceeding 100 bp, 15 out of 25 Actinidia species has less than 15 long repeats (Table S7), whereas A. kolomikta, A. fulvicoma, A. hubeiensis, A. arguta var. giraldii, A. cylindrica var. reticulata, A. latifolia, A. chinensis “AC011”, A. chinensis “AC017”, A. chinensis var. deliciosa “AD006”, A. chinensis var. deliciosa “AD019”, A. chinensis “Jinguo”, A. chinensis “Hongyang”, A. chinensis var. setosa or A. rubus own 40, 30, 19, 18, 16, 15, 28, 34, 34, 64, 32, 24, 33 or 39 long repeats with length exceeding 100 bp, respectively, ranging in size predominantly between 100 and 300 bp (Table S8).
Furthermore, the distribution of long repeat sequences displayed a species-specific enrichment in Actinidia taxa (Fig. S10). Using 10 kb sequences as a statistics unit, there were three peaks of long repeat sequences’ distribution in ranges of 50–60 kb, 70–80 kb and 130–140 kb, respectively. Specifically in A. tetramera, the majority of long repeats are located in the ranges of 50–60 kb and 70–80 kb (Fig. S10). In 70–80 kb alone, the majority of long repeats are derived from three species, A. tetramera, A. kolomikta, and A. callosa var. henryi. Further syntenic sequence analyses indicated these long repeat sequences of 70–80 kb are mainly located at the intergenic region between rps12 and psbB (Fig. 5c).
Divergent sequence regions in Actinidia species’ chloroplast genomes
To characterize the divergence, the chloroplast genome sequence alignments of Actinidia species are present by mVISTA, using A. chinensis “AC011” as reference. High sequence similarities among 25 Actinidia species were revealed by sequence identity plots of the chloroplast genome sequences (Fig. S11). The majority of sequence variations are distributed in intergenic regions, whereas the PCGs, rRNAs and tRNAs contain comparatively less sequence fluctuations. The most divergent coding regions are located in genes accD and ycf1 (Fig. S11).
To further investigate the variable nucleotides, especially the hot spots possibly involved in evolution, the sequence diversity was calculated for 25 Actinidia species tested. As a result, the average value of nucleotide diversity (Pi) is 0.00559, and the average Pi value of LSC (0.00664) and SSC (0.00814) is much higher than that in the IR (0.00249).
Detailed Pi value demonstrated the many variable regions are located in LSC and SSC regions, with the IR regions remaining relatively conserved across Actinidia genus (Fig. 6a). In LSC, SSC and IR regions, there are nine, two and zero DNA fragments showing relatively high nucleotide diversity (Pi value > 0.016) (Table S9). In IR regions, Pi value of the most divergent sequences is 0.0105. In LSC and SSC regions, there are four highly divergent regions, rps16 ~ trnQ-UUG, rps4 ~ trnT-UGU, petA ~ psbJ and rps12 ~ psbB, which exhibit remarkably higher Pi values (> 0.02) (Fig. 6a). Furthermore, rps12 ~ psbB, exclusively embodying clpP gene and its up/down-stream noncoding sequence, is the most divergent region, with Pi value > 0.03 (Fig. 6a). We checked the genome location of three divergent regions, including rps4 ~ trnT-UGU, petA ~ psbJ and rps12 ~ psbB, which are distributed in a sytenic region between 46,390 and 75,519 bp of the chloroplast genomes in 25 Actinidia species. This 29 kb region included 31 genes, including 24 PCGs and 7 tRNAs (Fig. 6b). The abundant variable nucleotide sites in the 29 kb region could provide suitable molecular markers for further phylogenetic studies of Actinidia species.
Divergent hot spots in Actinidia species chloroplast genomes. a The nucleotide diversity (Pi value) in chloroplast genomes of 25 Actinidia species. Four most divergent hot spots (Pi values > 0.02) are labeled, respectively. b The detailed gene distribution in the genomic regions spanning three highly divergent regions among 25 Actinidia species
Phylogenetic reconstruction in Actinidia
Using two Actinidiaceae species, S. tristyla and C. scandens as outgroup, phylogenetic relationships among 25 Actinidia species were reconstructed. Based on the chloroplast genome sequences, the ML phylogenetic tree was constructed among the 27 Actinidiaceae species (Fig. 7a).
The reconstructed ML phylogenetic tree of 25 Actinidia species. The ML (Maximum likelihood) phylogenetic tree of 25 Actinidia species based on the whole chloroplast genome sequences (a) and LSC plus SSC regions (b), respectively. Four infrageneric sections in Actinidia, including Leiocarpae (Lei), Maculatae (Mac), Stellatae (Ste), and Strigosae (Str), are labeled besides each species
In the phylogenetic tree, 25 Actinidia species could be classified into three main groups, Group I (7 species), Group II (11) and Group III (7). Group II and Group III represented closer phylogenetic relationships in comparison with Group I located in the outer (Fig. 7a). In Group III, A. chinensis, A. chinensis var. deliciosa, A. chinensis var. setosa, A. indochinensis and A. callosa var. strigillosa clustered together, whereas A. zhejiangensis and A. rufa formed another independent cluster.
Group II included two independent clades. A. cylindrica var. cylindrica, A. rubus, A. hubeiensis, and A. callosa var. henryi were clustered in one clade. A. styracifolia, A. eriantha, A. fulvicoma, A. cylindrica var. reticulata, A. hemsleyana, and A. latifolia, showed closely phylogenetic relationships in another clade (Fig. 7a).
Considering the abundant variable nucleotides in LSC and SSC regions (11 regions with Pi value > 0.016) (Table S9), two other ML phylogenetic trees of 25 Actinidia species was constructed based on the sequences of LSC plus SSC regions (Fig. 7b) and LSC alone (Fig. S12), respectively, showing consistent phylogenetic relationships with that reconstructed through the chloroplast genome sequences. Furthermore, based on chloroplast genome sequences (Fig. 7a), LSC plus SSC regions (Fig. 7b) or LSC alone (Fig. S12), our reconstructed relationships among the 25 Actinidia species are mainly in accordance with previous reports in Actinidia species (Liu et al. 2017; Tang et al. 2019b).
In addition, another three phylogenetic trees were also reconstructed based on whole chloroplast genome sequences (Fig. S13a), LSC plus SSC regions (Fig. S13b) or LSC alone (Fig. S14) of 31 independent chloroplast genomes from 25 Actinidia species, by adding another two chloroplast genomes from polyploid species, A. chinensis “AC017” (tetraploid) and A. chinensis var. deliciosa “AD019” (hexaploid). All the three phylogenetic trees also displayed consistent topology with our other trees based on either the whole chloroplast genome sequences or SSC and/or LSC regions alone, respectively.
Discussions
In this study, the genome-wide comparative genomic analyses were performed among chloroplast genomes of 25 Actinidia species. The clpP gene sequence with exon merge and intron deletion was identified in all the 29 tested chloroplast genomes tested from 25 Actinidia species. Four highly divergent sequence regions, including rps16 ~ trnQ-UUG, rps4 ~ trnT-UGU, petA ~ psbJ and rps12 ~ psbB were identified. Based on either sequences of LSC, combined SSC and LSC or the whole chloroplast genome sequences, the consensus phylogenetic tree with improved distinct resolution for 25 Actinidia taxa was reconstructed.
The chloroplast genomes of Actinidia species could represent genus specific evolution characteristics. In the chloroplast genomes of Actinidia species, three out of a total four highly divergent sequence regions, including rps4 ~ trnT-UGU, petA ~ psbJ and rps12 ~ psbB, were defined in a syntenic region, ranging from 46,390 to 75,519 bp. To compare the high variation sequence regions with other Ericales species, the nucleotide polymorphisms were also calculated in the chloroplast genomes of species in family Balsaminaceae (4 members), Ebenaceae (11), Pentaphylacaceae (3), Primulaceae (31), Sapotaceae (5), Styracaceae (22) and Theaceae (29), respectively (Fig. S15). Consequently, the highly divergent sequence regions in the aforementioned Ericales species’ chloroplast genomes are distinct from those in Actinidia species, representing a different evolutionary process in genus Actinidia.
Furthermore, rps12 ~ psbB region could be implicated as the most important evolutionary hot spot in genus Actinidia. The rps12 ~ psbB region exclusively embody varied clpP gene and its up/down-stream noncoding sequence (Fig. 2). Our comprehensive analyses indicate the varied clpP sequence including two exons is just Actinidiaceae-specific (Fig. 3, Fig. S5). The nucleotide variation (Pi value ˃ 0.3) demonstrates rps12 ~ psbB is the most divergent region in chloroplast genomes of Actinidia species (Fig. 6). This region is also one enriched with long repeats, mainly derived from A. tetramera, A. kolomikta, and A. callosa var. Henryi (Fig. 5c).
In the previous studies, due to lack of sufficient chloroplast genome sequences, the phylogenetic analyses of Actinidia species were mainly based on variant sites of limited nucleotide sequence fragments derived from nuclear, chloroplast and mitochondrion genomes (Huang et al. 2002; Li et al. 2002; Chat et al. 2004). Our phylogenetic studies have been performed at chloroplast genome-level among 25 Actinidia species, including sufficient nucleotides polymorphism for phylogenetic relationship reconstruction. Most of the bootstrap values besides the tree branches are 100 (Fig. 7). Significantly, our phylogenetic tree of 25 Actinidia species based on whole chloroplast genome, LSC plus SSC, or LSC alone, showed consistent phylogenetic relationships, further demonstrating the accuracy and reliability of our method and results (Fig. 7, Figs. S12, S13, S14).
Morphologically classified Actinidia taxa includes four infrageneric sections, Leiocarpae (Lei), Maculatae (Mac), Stellatae (Ste), and Strigosae (Str) (Chat et al. 2004; Testolin et al. 2016). Apparently, our phylogenetic tree is largely in accordance with the four sections, including nine species from section Ste, eight from section Lei, six from section Mac and two from section Str (Fig. 7).
All the members from section Ste and section Mac are clustered to form neighboring Group I and Group II that represent relatively closer phylogenetic relationships. Specifically, five Ste members, four Mac members and two Str members are clustered together in Group II. Adjacent to four Ste members, two from section Mac and one from Section Lei are clustered within Group III. Intriguingly, seven out of eight members from section Lei, including A. kolomikta, A. valvata, A. polygama, A. macrosperma, A. arguta, A. arguta var. giraldii, and A. tetramera, are consecutively located in Group I, consistently supporting the basal positions of most Lei species in Actinidia genus (Fig. 7). A major discrepancy is that the A. rufa, a member of Lei, is clustered with A. zhejiangensis to form an independent cluster located in Group III. But this exception seems not in conflict with another two investigations using either SNPs of nuclear genomes (Liu et al. 2017) or four polymorphic intergenic spacers sequences derived from the chloroplast genomes (Tang et al. 2019b).
Recently, two phylogenetic studies based on genome-wide SNPs (Liu et al. 2017) or four intergenic spacers sequences of the chloroplast genomes were reported (Tang et al. 2019b), respectively. Our phylogenetic tree is largely consistent with that based on genome-wide SNPs (Liu et al. 2017), supporting an improved resolution in determining the interspecific relationships of Actinidia species using whole chloroplast genome sequences in our study. An exception is that in the genome-wide SNPs phylogenetic tree (Liu et al. 2017), A. zhejiangensis is closely clustered with A. latifolia, A. eriantha, A. fulvicoma, A. cylindrica, A. callosa var. henryi and A. lanceolata, in contrast to our tree wherein A. zhejiangensis shows close relationship with A. chinensis complex (A. chinensis, A. chinensis var. deliciosa, A. chinensis var. setosa), A. callosa var. strigillosa, A. indochinensis, and A. rufa to form a monophyletic clade (Fig. 7).
Distinct from the trees of ours (Fig. 7) and on the basis of SNPs of nuclear genomes (Liu et al. 2017) that A. valvata is closely clustered with A. polygama, A. valvata shows the closest lineage with A. tetramera using four intergenic spacers sequences of the chloroplast genomes (Tang et al. 2019b). In addition, our data and previous studies (Liu et al. 2017; Tang et al. 2019b) indicated A. macrosperma together with other section Lei species are grouped into the basal clade of Actinidia species (Fig. 7). Interestingly, A. macrosperma shows different interspecific sister relationships in the three studies, including A. macrosperma/A. kolomikta (Liu et al. 2017), A. macrosperma/A. polygama (Tang et al. 2019b) and A. macrosperma/A. arguta of ours.
Additionally, for A. chinensis complex (A. chinensis, A. chinensis var. deliciosa, and A. chinensis var. setosa), A. arguta complex (A. arguta, and A. arguta var. giraldii), and A. cylindrica complex (A. cylindrica var. cylindrica, and A. cylindrica var. reticulata), both the tree of ours and on the basis of SNPs of nuclear genomes (Liu et al. 2017) support closer relationships of the species in each species complex (Fig. 7), with members from each species complex clustered in a main clade in both trees, respectively. Interestingly, A. indochinensis other than members in A. chinensis complex shows interspecific sister relationships with A. chinensis in both trees (Fig. 7) (Liu et al. 2017). In both studies’ results, similar findings also exist in A. arguta complex and A. cylindrica complex. It was demonstrated that the largely divergent evolution process might occur in the members of each Actinidia species complex.
We believe all the discrepancies could happen due to the occurrences of naturally interspecific hybridization and/or introgression events originating many times resulting in distant cytoplasm–nuclear hybridizations and reticulate evolution events in Actinidia (Chat et al. 2004; Testolin et al. 2016), as well as the independent evolution directions of the chloroplast and nuclear genomes.
Conclusion
In this study, chloroplast genome-wide comparative analyses were performed in 25 Actinidia species. The average chloroplast genome size is 156,673.38 bp, with average 37.20% GC content. The total gene number variation mainly resulted from gene copy number variations and gene losses. The long repeat sequences other than SSRs are the main repeats resulting genome size expansion. The most hypervariable regions involving evolutionary hot spots in Actinidia species is rps12 ~ psbB wherein the clpP gene sequence with exon merge and intron loss was discovered and implicated in differentiation of Actinidiaceae. The phylogenetic relationships of 25 Actinidia taxa are refined as well.
Data availability
The data presented in this study are available in the article and Supplementary Materials.
References
Ai F, Liu H (2019) The complete chloroplast genome sequence of Actinidia zhejiangensis. Mitochondrial DNA Part B 4(1):690–691. https://doi.org/10.1080/23802359.2019.1573117
Capella-Gutierrez S, Silla-Martinez JM et al (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25(15):1972–1973. https://doi.org/10.1093/bioinformatics/btp348
Chat J, Jauregui B et al (2004) Reticulate evolution in kiwifruit (Actinidia, Actinidiaceae) identified by comparing their maternal and paternal phylogenies. Am J Bot 91(5):736–747. https://doi.org/10.3732/ajb.91.5.736
Chen Y, Xu Y et al (2019) The complete chloroplast genome of Actinidia macrosperma. Mitochondrial DNA Part B 4(2):4188–4189. https://doi.org/10.1080/23802359.2019.1692733
Chen Y-T, Lai R-l et al (2020) The complete chloroplast genome sequence of actinidia valvata. Mitochondrial DNA Part B 5(3):2072–2073. https://doi.org/10.1080/23802359.2020.1764402
Cui Y, Chen X et al (2019) Comparison and phylogenetic analysis of chloroplast genomes of three medicinal and edible Amomum species. Int J Mol Sci 20(16):4040. https://doi.org/10.3390/ijms20164040
Daniell H, Lin CS et al (2016) Chloroplast genomes: diversity, evolution, and applications in genetic engineering. Genome Biol 17(1):134. https://doi.org/10.1186/s13059-016-1004-2
Delcher AL, Salzberg SL et al (2003) Using MUMmer to identify similar regions in large sequence sets. Curr Protoc Bioinform 1: 10.3.1–10.3.18. https://doi.org/10.1002/0471250953.bi1003s00
Ding F, Zhang L et al (2021) The complete chloroplast genome sequence of Actinidia arguta var. giraldii. Mitochondrial DNA B Resour 6(2):413–414. https://doi.org/10.1080/23802359.2020.1870884
Frazer KA, Pachter L et al (2004) VISTA: computational tools for comparative genomics. Nucleic Acids Res 32(Web Server issue):W273–W279. https://doi.org/10.1093/nar/gkh458
Hong Z, Wu Z et al (2020) Comparative analyses of five complete chloroplast genomes from the genus Pterocarpus (Fabacaeae). Int J Mol Sci 21(11):3758. https://doi.org/10.3390/ijms21113758
Huang H, Li Z et al (2002) Phylogenetic relationships in Actinidia as revealed by RAPD analysis. J Am Soc Hortic Sci 127(5):759–766
Huang S, Ding J et al (2013) Draft genome of the kiwifruit Actinidia chinensis. Nat Commun 4:2640. https://doi.org/10.1038/ncomms3640
Huang J, Yu Y et al (2020) Comparative chloroplast genomics of Fritillaria (Liliaceae), inferences for phylogenetic relationships between Fritillaria and Lilium and plastome evolution. Plants (basel) 9(2):133. https://doi.org/10.3390/plants9020133
Kalyaanamoorthy S, Minh BQ et al (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14(6):587–589. https://doi.org/10.1038/nmeth.4285
Katoh K, Rozewicki J et al (2019) MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform 20(4):1160–1166. https://doi.org/10.1093/bib/bbx108
Kim S-C, Lee J-W et al (2018) The complete chloroplast genome sequence of Actinidia Rufa (Actinidiaceae). Mitochondrial DNA Part B 3(2):564–565. https://doi.org/10.1080/23802359.2018.1450676
Kurtz S, Choudhuri JV et al (2001) REPuter: the manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res 29(22):4633–4642. https://doi.org/10.1093/nar/29.22.4633
Lan Y, Cheng L et al (2017) The complete chloroplast genome sequence of Actinidia kolomikta from north China. Conserv Genet Resour 10(3):475–477. https://doi.org/10.1007/s12686-017-0852-8
Li J, Huang H et al (2002) Molecular phylogeny and infrageneric classification of Actinidia (Actinidiaceae). Syst Bot 27(2):408–415
Li W, Lu Y et al (2018) The complete chloroplast genome sequence of Actinidia arguta: gene structure and genomic resources. Conserv Genet Resour 10(3):423–425. https://doi.org/10.1007/s12686-017-0840-z
Li X, Zuo Y et al (2019) Complete chloroplast genomes and comparative analysis of sequences evolution among seven Aristolochia (Aristolochiaceae) medicinal species. Int J Mol Sci 20(5):1045. https://doi.org/10.3390/ijms20051045
Lin M, Qi X et al (2018) The complete chloroplast genome sequence of Actinidia arguta using the PacBio RS II platform. PLoS One 13(5):e0197393. https://doi.org/10.1371/journal.pone.0197393
Lin H, Jiang L et al (2019) Assembly and phylogenetic analysis of the complete chloroplast genome sequence of Actinidia setosa. Mitochondrial DNA Part B 4(2):3679–3680. https://doi.org/10.1080/23802359.2019.1678423
Lin H, Xu Y et al (2020) The complete chloroplast genome of Actinidia valvata (Actinidiaceae). Mitochondrial DNA Part B 5(2):1607–1608. https://doi.org/10.1080/23802359.2020.1745105
Liu YF, Li DW et al (2017) Rapid radiations of both kiwifruit hybrid lineages and their parents shed light on a two-layer mode of species diversification. New Phytol 215(2):877–890. https://doi.org/10.1111/nph.14607
Liu Y, Xie X et al (2020) Phylogenetic relationship and characterization of the complete chloroplast genome of Actinidia callosa var. strigillosa. Mitochondrial DNA Part B 5(3):3420–3421. https://doi.org/10.1080/23802359.2020.1823258
Martin W, Deusch O et al (2005) Chloroplast genome phylogenetics: why we need independent approaches to plant molecular evolution. Trends Plant Sci 10(5):203–209. https://doi.org/10.1016/j.tplants.2005.03.007
Nguyen LT, Schmidt HA et al (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32(1):268–274. https://doi.org/10.1093/molbev/msu300
Nie L, Cui Y et al (2019) Gene losses and variations in chloroplast genome of parasitic plant Macrosolen and phylogenetic relationships within Santalales. Int J Mol Sci 20(22):5812. https://doi.org/10.3390/ijms20225812
Park I, Song JH et al (2019) Cuscuta species identification based on the morphology of reproductive organs and complete chloroplast genome sequences. Int J Mol Sci 20(11):2726. https://doi.org/10.3390/ijms20112726
Qu XJ, Moore MJ et al (2019) PGA: a software package for rapid, accurate, and flexible batch annotation of plastomes. Plant Methods 15:50. https://doi.org/10.1186/s13007-019-0435-7
Rozas J, Ferrer-Mata A et al (2017) DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol Biol Evol 34(12):3299–3302. https://doi.org/10.1093/molbev/msx248
Shi L, Chen H et al (2019) CPGAVAS2, an integrated plastome sequence annotator and analyzer. Nucleic Acids Res 47(W1):W65–W73. https://doi.org/10.1093/nar/gkz345
Tang P, Shen R et al (2019a) The complete chloroplast genome sequence of Actinidia eriantha. Mitochondrial DNA Part B 4(2):2114–2115. https://doi.org/10.1080/23802359.2019.1623111
Tang P, Xu Q et al (2019b) Phylogenetic relationship in Actinidia (Actinidiaceae) based on four noncoding chloroplast DNA sequences. Plant Syst Evol 305(9):787–796. https://doi.org/10.1007/s00606-019-01607-0
Testolin R, Huang H et al (2016) The kiwifruit genome. Springer International Publishing, Berlin
Tillich M, Lehwark P et al (2017) GeSeq—versatile and accurate annotation of organelle genomes. Nucleic Acids Res 45(W1):W6–W11. https://doi.org/10.1093/nar/gkx391
Tyagi S, Jung JA et al (2020) Comparative analysis of the complete chloroplast genome of mainland Aster spathulifolius and other Aster species. Plants (basel) 9(5):568. https://doi.org/10.3390/plants9050568
Wang WC, Chen SY et al (2016) Chloroplast genome evolution in Actinidiaceae: clpP loss, heterogenous divergence and phylogenomic practice. PLoS One 11(9):e0162324. https://doi.org/10.1371/journal.pone.0162324
Wu H, Li M et al (2019) The complete chloroplast genome sequence of Actinidia callosa var. henryi. Mitochondrial DNA Part B 4(1):652–653. https://doi.org/10.1080/23802359.2018.1561223
Xiaoqiong Q, Xiaodong X et al (2021) Characterization of the complete chloroplast genome of Actinidia hemsleyana. Mitochondrial DNA B Resour 6(11):3259–3260. https://doi.org/10.1080/23802359.2021.1993100
Xu Y-S, Zhang C-G et al (2020) The complete chloroplast genome of Actinidia rubus (Actinidiaceae). Mitochondrial DNA Part B 5(1):366–367. https://doi.org/10.1080/23802359.2019.1703571
Yang A, Liu S et al (2020) The complete chloroplast genome sequence of Actinidia styracifolia C. F. Liang. Mitochondrial DNA Part B 5(1):90–91. https://doi.org/10.1080/23802359.2019.1698337
Yao X, Tang P et al (2015) The first complete chloroplast genome sequences in Actinidiaceae: genome structure and comparative analysis. PLoS One 10(6):e0129347. https://doi.org/10.1371/journal.pone.0129347
Zhang J, Liu H (2019) The complete choloroplast genome sequence of Actinidia lanceolate. Mitochondrial DNA Part B 4(1):1187–1188. https://doi.org/10.1080/23802359.2019.1591200
Zhang F, Yan Z et al (2019) The complete chloroplast genome of Actinidia fulvicoma. Mitochondrial DNA Part B 4(2):4089–4090. https://doi.org/10.1080/23802359.2019.1691949
Funding
This work was supported by grants from the National Natural Science Foundation of China (Grant nos. 31972474, 31671259 and 31440028).
Author information
Authors and Affiliations
Contributions
LW, BL and YY: methodology, data analysis, visualization, validation, and writing—original draft preparation and writing. QZ and SC: data analysis and visualization. YL and SH: conceptualization, supervision, project administration, funding acquisition, writing—original draft preparation and writing—review and editing. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All the authors declare no conflicts of interest.
Additional information
Communicated by Bing Yang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, L., Liu, B., Yang, Y. et al. The comparative studies of complete chloroplast genomes in Actinidia (Actinidiaceae): novel insights into heterogenous variation, clpP gene annotation and phylogenetic relationships. Mol Genet Genomics 297, 535–551 (2022). https://doi.org/10.1007/s00438-022-01868-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-022-01868-4