Abstract
Trans-Eurasian cultural and genetic exchanges have significantly influenced the demographic dynamics of Eurasian populations. The Hexi Corridor, located along the southeastern edge of the Eurasian steppe, served as an important passage of the ancient Silk Road in Northwest China and intensified the transcontinental exchange and interaction between populations on the Central Plain and in Western Eurasia. Historical and archeological records indicate that the Western Eurasian cultural elements were largely brought into North China via this geographical corridor, but there is debate on the extent to which the spread of barley/wheat agriculture into North China and subsequent Bronze Age cultural and technological mixture/shifts were achieved by the movement of people or dissemination of ideas. Here, we presented higher-resolution genome-wide autosomal and uniparental Y/mtDNA SNP or STR data for 599 northwestern Han Chinese individuals and conducted 2 different comprehensive genetic studies among Neolithic-to-present-day Eurasians. Genetic studies based on lower-resolution STR markers via PCA, STRUCTURE, and phylogenetic trees showed that northwestern Han Chinese individuals had increased genetic homogeneity relative to northern Mongolic/Turkic/Tungusic speakers and Tibeto-Burman groups. The genomic signature constructed based on modern/ancient DNA further illustrated that the primary ancestry of the northwestern Han was derived from northern millet farmer ancestors, which was consistent with the hypothesis of Han origin in North China and more recent northwestward population expansion. This was subsequently confirmed via excess shared derived alleles in f3/f4 statistical analyses and by more northern East Asian-related ancestry in the qpAdm/qpGraph models. Interestingly, we identified one western Eurasian admixture signature that was present in northwestern Han but absent from southern Han, with an admixture time dated to approximately 1000 CE (Tang and Song dynasties). Generally, we provided supporting evidence that historic Trans-Eurasian communication was primarily maintained through population movement, not simply cultural diffusion. The observed population dynamics in northwestern Han Chinese not only support the North China origin hypothesis but also reflect the multiple sources of the genetic diversity observed in this population.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The history of Homo sapiens occupation of East Asia can be traced back to the late Paleolithic period, and anatomically modern humans (AMHs) permanently inhabited this region 50,000 years ago (Yang et al. 2017). Evidence from aboriginal Australian genomes suggested two waves of settlements in Asia, including an initial southern Out-of-Africa migration wave approximately 60,000 years ago and a later wave approximately 45,000 years ago (Rasmussen et al. 2011). However, this model of early, independent human dispersals into Asia was questioned due to its disregard of the archaic introgression of Denisovan genetic material into modern Asian Negrito, Australian, and New Guinean populations (Mallick et al. 2016). One well-fitted phylogenetic admixture graph with two archaic admixture events further suggested that New Guineans and Australians formed the eastern Eurasian clade together with mainland East Asians, represented by Austronesian Ami and Tai–Kadai Dai (Mallick et al. 2016). In East Asia, pioneering genomic work in the HUGO Pan-Asian project that genotyped 50,000 single-nucleotide polymorphisms (SNPs) in 1900 individuals from 73 populations was finished in 2009 and provided evidence that the main Southern Migration Route played an important role in the peopling of East Asia, with little genetic contribution from the Northern Migration Route (Consortium et al. 2009). However, other uniparental evidence (based on maternally inherited mitochondrial DNA and paternally inherited Y-chromosomal genomes) illuminated the possibility of genetic contributions from northern Eurasia into East Asia via the Northern Migration Route (Su et al. 1999; Wen et al. 2004).
The ethnolinguistic and cultural diversity in eastern Eurasia and its complex history of population admixtures have promoted the exploration of genomic information from people living on the East Asian subcontinent, and the results have been widely utilized in forensics, anthropology and medical genetics, especially for the exploration of the pathogenicity of genetic variants (medically relevant novel and rare loci) in the modern precision medicine era (Stoneking and Delfin 2010; Liu et al. 2018; Cao et al. 2020). However, early comprehensive population genetic studies have mostly focused on Central/South Asian genetic diversity (Damgaard et al. 2018; Narasimhan et al. 2019), and the pattern of genetic variations in East Asia has not been fully characterized, especially for China, which has multiple language- and agriculture-oriented centers. Chinese populations have been classically categorized according to the six main language families: Austronesian, Austroasiatic, Hmong–Mien, Tai–Kadai, Sino-Tibetan and Trans-Eurasian. Fortunately, recently obtained information from ancient genomes has reshaped our understanding of the peopling of East Asia (Raghavan et al. 2014; Jeong et al. 2016; Yang et al. 2017, 2020; Lipson et al. 2018; McColl et al. 2018; Bai et al. 2020; Gakuhari et al. 2020; Ning et al. 2020; Wang et al. 2020a, b; Zhang and Fu 2020). First, genetic variations in mitochondrial and autosomal DNA from paleolithic hunter-gatherers have demonstrated differentiated genetic relationships between Upper Pleistocene East Asians and modern people (Zhang and Fu 2020); 40,000-year-old Tianyuan people formed a deeply northern East Asian lineage but made limited genetic contributions to modern East Asians (Yang et al. 2017); and the Ancient Northern Eurasian clade (24,000-year-old Mal’ta) contributed one-third of its genome variations to indigenous Americans but also made only limited genetic contributions to modern East Asians (Raghavan et al. 2014). In contrast, the 11,000-year-old Longlin and Qingshuiyuan people exhibit maternal genetic continuity with modern southern East Asians (Bai et al. 2020).
Second, the analyses of Holocene East Asian or Southeast Asian ancient genomes have documented multiple waves of migration of Chinese farmers and subsequent admixture events with resident groups in Southeast Asia (Lipson et al. 2018; McColl et al. 2018), lowland East Asia (Ning et al. 2020; Yang et al. 2020), highland Qinghai–Tibet Plateau (Jeong et al. 2016), and the Japanese Archipelago (Gakuhari et al. 2020), among others (Wang et al. 2020a), which are also referred to as Holocene expansion events. McColl et al. explored the genetic prehistory based on the genome-wide polymorphisms of 25 late Neolithic-to-historic Southeast Asians and found that the first batch of southward migration of Yangtze rice farmers disseminated the proto-Austroasiatic language and mixed with the local Southeast Asian indigenous hunter-gatherers (represented by 7,950-year-old Hoabinhian genomes) to produce the modern Austroasiatic speakers distributed fragmentarily in Southeast Asia, such as the Mlabri and Htin (McColl et al. 2018). The genetic legacy of this first layer showed a strong genetic affinity with Andamanese and Japanese Jomon, which also possessed an ancient genetic connection between coastal Southeast Asia and the Japanese archipelago (Gakuhari et al. 2020). Later, multiple southward migrations from South China to the islands of Southeast Asia during the Bronze Age were associated with Austronesian expansion via the coastal expansion route and with mainland Southeast Asia associated with Tai–Kadai/Hmong–Mien speakers via the inland expansion route (McColl et al. 2018). Lipson and his colleagues at Harvard Medical School sequenced eighteen Neolithic-to-Iron Age Southeast Asians and reconstructed the phylogenetic lineage of these ancient populations (Lipson et al. 2018). They also found that indigenous Southeast Asian hunter-gatherers, as a highly diverged Eastern Eurasian lineage, mixed with early farmers from South China to form Austroasiatic-related ancestral populations (Man_Bac). In addition, this study also recorded the ancient DNA signature of southward migration of Tibeto-Burman ancestors in the late Neolithic/Bronze Age Oakaie population (Lipson et al. 2018). In China, 10 early Neolithic genomes from the Yellow River Basin and 16 Neolithic-to-historic genomes from Fujian and the surrounding region were used to reconstruct the landscape of North-to-South population stratification from the early Neolithic period. Stronger northern East Asian affinity in late Neolithic Fujian populations demonstrated that the main migration direction of East Asians in the Holocene period was from north to south (Yang et al. 2020). The closer genetic relationship between late Neolithic Tanshishan/Xitoucun populations, 3000-year-old Vanuatu samples and modern Austronesian Ami and Atayal suggested that the initial common origin of Austronesian speakers was sited in the coastal region of South China. Coastal population intercommunications between ancient populations from Vietnam, China, Japan and far eastern Russia showed that the coastal migration route was a convenient and important gateway for ancient population movement and mixture (McColl et al. 2018; Gakuhari et al. 2020; Wang et al. 2020a; Yang et al. 2020). In addition, 55 ancient genomes from the Yellow River, western Liao River and Amur River regions also showed archeologically supported changeable subsistence strategies that allowed northern millet farmers to adapt to climate change, which was achieved via population movements in the inland migration corridor between the three river regions (Ning et al. 2020). Comprehensive analysis of ancient and modern genome data from Tibetan individuals via Wang et al. illustrated that multiple Paleolithic and Neolithic migration events have participated in the peopling of the Tibetan Plateau and revealed the different population stratifications among culturally diverse Tibetans (Wang et al. 2020b): a major influx of 2700-year-old Chokhopani ancestry into Ü-Tsang Tibetans, an additional western Eurasian influx into Ando Tibetans, and a southern East Asian influx into Kham Tibetans (Wang et al. 2020b). Furthermore, three Holocene expansion events partially or completely associated with dissemination of the Trans-Eurasian, Sino-Tibetan, and southern language macrofamilies (Austronesian, Tai–Kadai, and Austroasiatic) have also been comprehensively documented via ancient population genomics (Wang et al. 2020a).
These intensified North-to-South population interactions in China have been well documented; however, east-to-west transcontinental communications need to be comprehensively characterized from a genetic perspective. Trans-Eurasian cultural exchange has recently been extensively documented via historic records and archeological findings (Dong et al. 2020). In addition, the corresponding transcontinental population movement during the Bronze Age to Iron Age has been confirmed in the core regions of Siberia (Damgaard et al. 2018). However, the mechanisms underlying the spread of the archeologically supported western Bronze Age package in China (i.e., assimilation of ideas or movement of people), as well as the association between the westward spread of millet agriculture and eastward spread of barley/wheat farming technology and human movement, need to be genetically explored, especially in the Hexi Corridor and the surrounding regions in northwestern China. Among the modern gene pool of northwestern East Asians, population genetic studies have focused on the topics of origin, migration, admixture, and substructure based on lower-density genetic markers or limited sample sizes (He et al. 2018b; Wang et al. 2018b; Li et al. 2020). However, a comprehensive survey of the genetic diversity and finer-scale structure of northwestern East Asians is in its infancy, and much work based on different genetic markers (single-nucleotide polymorphisms (SNPs), short tandem repeats (STRs) and so on) and denser anthropological sampling should be carried out. Here, we conducted a comprehensive population genetic survey based on the genetic variations of genome-wide STRs/SNPs in northwestern East Asia and investigated their forensic features, genetic diversity, population substructure, and phylogenetic relationships based on Paleolithic-to-modern East Asian genetic variations.
Materials and methods
Samples, DNA preparation and PCR amplification and profiling
We collected samples from 599 unrelated healthy individuals (152 males and 447 males) in Gansu Province in Northwest China (Figure S1) with written informed consent. Our study and corresponding protocols were reviewed and approved via the Medical Ethics Committee of Xiamen University (XDYX201909). In addition, we also followed the recommendations of the Declaration of Helsinki (Nicogossian et al. 2014) and the regulations of the Human Genetic Resources Administration of China (HGRAC). We extracted genomic DNA using the QIAamp DNA Mini Kit (Qiagen) and quantified the extracted DNA materials via a NanoDrop-2000c instrument (Thermo Fisher Scientific). All prepared DNA templates were preserved at − 20 °C until the next step of DNA amplification. We used the Huaxia Platinum PCR amplification kit and ProFlex 96-well PCR System (Thermo Fisher Scientific) following the kit recommendations to amplify the targeted 26 genetic loci in 549 individuals. We used the Applied Biosystems 3500XL Genetic Analyzer to electrophorese and separate the amplified DNA products and GeneMapper ID-X v.1.4 (Thermo Fisher Scientific) to call and check the obtained genotype data. We used the Infinium® Global Screening Array (GSA) to genotype approximately 700 K SNPs across the whole genome in 50 male individuals. We considered the site missing rates per person or per SNPs and Hardy–Weinberg disequilibrium in the quality control with the following parameter settings (mind: 0.01, geno: 0.01, --hwe 0.001 and --maf 0.01) using PLINK 1.9 (Chang et al. 2015).
Population database
To conduct a comprehensive population genetic survey, we employed five different reference databases to make population comparisons: two STR genotype-based datasets, one STR allele frequency-based dataset, and two high-density SNP-based datasets. The first comprised genotypes from 23 autosomal STRs in 12,960 individuals (549 genotypes first reported here and 12,411 genotypes collected from the public database) from 17 Eurasian populations and was referred to as the 23STR genotype dataset. This dataset comprised seven Sinitic populations (He et al. 2018a, 2018c; Wang et al. 2018a; Liu et al. 2019; Pengyu Chen et al. 2019d, a, b, c; Li et al. 2020) (Han populations from Gansu, Chengdu, Hainan, Shanxi, Shaanxi and Zhujiang and one Wuzhong Hui), four Turkic-speaking populations (Jin et al. 2017; Chen et al. 2019b, c; Liu et al. 2019) (one Kyrgyz from Akto, three Uyghur from Artux, Urumqi and across Xinjiang), five Tibeto-Burman-speaking populations (Wang et al. 2018a; Liu et al. 2019) [one Yi group from Liangshan, two Tibetans from Tibet and two Sichuan Tibetans (Liangshan and Chengdu)] and one Central Asian population from Quetta Hazara (Chen et al. 2019a). The second dataset included 14,365 individuals from the aforementioned 17 populations and 3 Western Eurasian populations (Estonian, Polish, and Saudi Arabian) based on the STRs identified in both of 2 different forensic PCR amplification systems (Sadam et al. 2015; Alsafiah et al. 2017; Ossowski et al. 2017), which was referred to as the 20STR genotype dataset. The third included 57 worldwide populations with allele frequency distributions of 20 STRs except for D6S1043, Penta D, and Penta E, which was referred to as a frequency-based dataset (Gaviria et al. 2013; Park et al. 2013, 2016; Fujii et al. 2014; Almeida et al. 2015; Hossain et al. 2016; Wang et al. 2016; Zhang et al. 2016a, b; Choi et al. 2017; Guerreiro et al. 2017; Moyses et al. 2017; Ossowski et al. 2017; Taylor et al. 2017; Wu et al. 2017; Yang et al. 2018; Chen et al. 2019a). The other two SNP-based datasets were formed by merging our newly genotyped data with the publicly available Human-Origin and 1240 K datasets and then used to conduct the population genomics analyses for East Asian populations (Patterson et al. 2012; Jeong et al. 2019; Wang et al. 2020a). The basic dataset of Eurasian modern and ancient reference populations were collected from Reich Lab (https://reich.hms.harvard.edu/downloadable-genotypes-present-day-and-ancient-dna-data-compiled-published-papers). Additional modern and ancient reference population data of East Asians from China, Japan, Mongolia and Nepal were collected from recent publications (Jeong et al. 2016; Ning et al. 2020; Wang et al. 2020a; Yang et al. 2020).
STR-based statistical analysis
We first estimated the statistical parameters related to forensic genetics (personal identification and parentage testing) in Gansu Han. We used the online tool STRAF (Gouy and Zieger 2017) to estimate the forensic parameters [matching probability (PM), discrimination power (PD), typical paternity index (TPI), power of paternity exclusion (PE), gene diversity (GD), polymorphism information content (PIC)] and pairwise Fst genetic distances (Weir and Cockerham 1984). We subsequently used Arlequin 3.5.2 to test the Hardy–Weinberg equilibrium (HWE) with 100,000 Markov chain steps and 100,000 dememorization steps via the type of locus by locus and linkage disequilibrium among all pairs of the included 23 STR loci with 10,000 permutations and 2 initial conditions of expectation maximization (Excoffier and Lischer 2010). Arlequin 3.5.2 was also used to calculate the observed heterozygosity (Ho) and expected heterozygosity (He). We used Phylip software (Cummings 2004) to calculate Cavalli-Sforza and Nei genetic distances based on the allele frequency distributions. Multivariate Statistical Package (MVSP) software 3.22 (Kovach 2007) was used to perform principal component analysis (PCA) among 57 worldwide or 28 Eurasian populations based on allele frequency distribution, and the cmdscale function in R was used to generate multidimensional scaling plots (MDS) of the worldwide or Eurasian populations based on different genetic distance matrixes. Phylogenetic frameworks were constructed via the neighbor-joining (NJ) algorithm in MEGA 7.0 (Kumar et al. 2016). The individual ancestry composition of the two genotype-based datasets was determined using STRUCTURE (Evanno et al. 2005).
Genomic-based statistical analysis
We used two higher-density datasets to reconstruct the deep population genetic history of northwestern Han. We used the Smartpca program built into EIGENSOFT v.6.1.4 to perform PCA (Patterson et al. 2006) and ADMIXTURE v.1.3.0 (Alexander et al. 2009) to carry out the model-based ancestral composition dissection based on the Human-Origin-merged dataset following our previous default settings (He et al. 2020a). Ancient individuals were projected onto the top two principal components (numoutlieriter: 0 and lsqproject: YES). We used PLINK v.1.9 (Chang et al. 2015) to prune SNP data with strong linkage disequilibrium with the following parameters (--indep-pairwise 200 25 0.4). We used the qp3Pop program of ADMIXTOOLS to conduct admixture-f3(Source1, Source2; Han_Lanzhou) to explore the admixture source proxies and perform outgroup-f3(Source, Han_Lanzhou; Mbuti) to explore the shared genetic drift. We further used the f4-statistics in the forms f4(Eurasian1, Eurasian2; Han_Lanzhou, Outgroup) and f4(Eurasian1, Han_Lanzhou; Eurasian2, Mbuti) to study genetic affinity, continuity and admixture. We also used qpWave/qpAdm and qpGraph (Haak et al. 2015) to estimate the admixture proportion and phylogenetic splits and admixture events based on the 1240 K-based merged dataset. Eight worldwide representative populations were used as outgroups, which included five modern populations (Mbuti, Papuan, Australian, Mixe and Onge) and three Eurasian ancient people (Ust_Ishim, Kostenki14 and MA1_HG). We finally used ALDER to estimate the time of North-to-South and West-to-East admixtures with 28 years as one generation length (Loh et al. 2013).
Results
Forensic features and genetic diversity of northwestern Han Chinese individuals
Gansu Province, with a population size of over 26 million, is located between the Qinghai–Tibet Plateau and Loess Plateau in Northwest China and serves as an important corridor for the prehistoric human occupation of the Qinghai–Tibet Plateau (especially for Yellow River millet farmers). Moreover, the Hexi Corridor passed through Gansu Province, suggesting that this region also played a key role in the Trans-Eurasian exchange of genetic materials, culture, crops, livestock and technology (Dong et al. 2020). We successfully obtained genotype data for 23 autosomal STRs in 549 Gansu Han individuals and merged them with publicly available reference data. All results from STR-based population genetic analyses showed that the northwestern Han Chinese population is homogenous and has high genetic diversity. All 23 STR loci were found to be in HWE and LE after conducting Bonferroni correction (Tables S1–2). We observed 277 alleles with allele frequency distributions ranging from 0.0009 to 0.5237 (Table S3 and Figure S2). As shown in Table S3, Penta E had the maximum number of alleles (23), followed by FGA (20), and TH01 had the fewest alleles (6), followed by TPOX. GD varied from 0.6180 (TPOX) to 0.9231 (Penta E), and Ho ranged from 0.5847 to 0.9362. PD varied from 0.7887 to 0.9868, PE varied from 0.2729 to 0.8699, and PM varied from 0.0132 to 0.2113. We also found that PIC ranged from 0.5545 to 0.9169 and that TPI ranged from 1.2039 to 7.8429. For the combined powers for forensic practice, we estimated the values of two combined forensic indexes: the combined power of discrimination (CPD) and the combined probability of exclusion (CPE). The CPD and CPE in Gansu Han were 7.827E-28 and 0.9999999998, respectively. The observed highly polymorphic and informative forensic statistical indexes showed that this 23-STR-based PCR amplification system was suitable for forensic identification of individuals and parentage testing. All included statistical parameters consistently demonstrated that northwestern Han individuals possessed high genetic diversity.
Population genetic analyses among Eurasian/worldwide populations via STR genetic markers
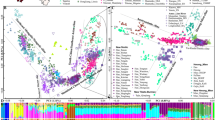
To investigate the genetic relationships between Gansu Han and other reference populations, we merged genotype data of the 23 autosomal STRs in the Han population with data from 16 other Central Asian or East Asian populations and calculated the pairwise Fst genetic distances. A total of 12,960 genotype data points were collected, and we found that Gansu Han exhibited the closest genetic relationship with Shanxi Han (0.0002), followed by Han Chinese groups from Zhujiang and Shaanxi and the Hui group from Wuzhong (Table S4). We next performed MDS among 17 Central or East Asian populations and found three genetic clusters. Turkic-speaking populations (Uyghur and Kyrgyz) were grouped with a Central Asian Hazara and localized in the right position in the MDS plot, forming one western Eurasian affinity cluster. Southern Han clustered with northern Han (including Gansu Han) and Hui in North China (Wuzhong) to form the second cluster (Sinitic-speaking cluster), localized in the top left position in the MDS. The remaining populations, localized in the bottom right position, clustered as the Tibeto-Burman-speaking cluster (Figure S3A). Patterns of genetic affinity were further confirmed by cluster analysis via heatmap (Figure S3B), NJ-based phylogenetic relationship reconstruction (Figure S3C) and STRUCTURE-based ancestral composition (Figure S3D). We further explored the genetic similarities and differences between Gansu Han and relatively closely related reference populations (Fig. 1) and merged genotype data from Central/East Asia and three populations from western Eurasia. Finally, we obtained a new dataset with genetic variation data for 20 autosomal STRs in 14,365 from 20 populations. As our outgroup, the three western Eurasian populations possessed the most distant genetic relationship with Gansu Han, which was inferred from the Fst genetic distance (Table S5). The three aforementioned clusters and one western Eurasian cluster were observed in the new MDS plot (Fig. 1a), and similar patterns of genetic relationships were also confirmed via the results from heatmap, phylogenetic relationship and STRUCTURE (Fig. 1b–d).
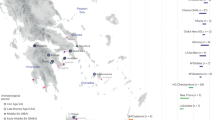
Genetic similarities and differences among 20 ethnically/geographically diverse Eurasian populations based on the genetic variations of 20 overlapping STRs. a Multidimensional scaling plots (MDS) showed genetic similarities between Lanzhou Han and other 19 Eurasian populations. b Genetic affinity among 20 populations inferred from the heatmap. c Phylogenetic relationship among the 20 populations reconstructed by the Fst genetic distance matrix. d Ancestry composition among 20 populations inferred from model-based STRUCTURE
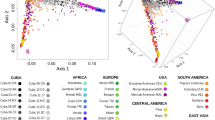
A denser sampling of reference population-based allele frequency distribution could be collected from publicly available population genetic data reports. Thus, we further performed comprehensive population genetic analyses based on the newly merged dataset with greater global population representation. We merged allele frequency data for 20 autosomal STRs in the Gansu Han population with data from 56 other reference populations. PCA based on allele frequency distribution was performed, and we found that a total of 48.484% of the variance in this population represented variation among worldwide populations and 52.590% of the variance occurred within the East Asian population, showing a strong population genetic affinity within geographically close or ethnically close populations distributed in different continental regions (Fig. 2a). Here, we could also observe the genetic affinity within the linguistically close populations within East Asians, especially for Tibeto-Burman and Sinitic speakers, and Gansu Han clustered most closely with the geographically close Wuzhong Hui. Genetic similarities between Gansu Han were also observed in the estimated pairwise Cavalli-Sforza genetic distances (Table S6 and Figure S4) and its heatmap (Figure S5), which showed the most shared ancestry (the closest genetic distance) between Gansu Han and the geographically close Shaanxi Han. We subsequently carried out MDS among 57 populations based on pairwise Nei genetic distances and observed similar patterns of genetic relationships (Fig. 2b). The Nei-based phylogenetic relationships showed a clear association between genetic affinity and geographical adjacency or linguistic affinity (Fig. 2c).
Patterns of genetic relationships between Lanzhou Han in Gansu province, Northwest China. a Result of principal component analysis (PCA) based on the allele frequency distribution. b Plots of multidimensional scaling analysis showed the genetic relationship among newly studied and the included reference populations based on the pairwise Cavalli-Sforza genetic distances. c The phylogenetic relationship between Gansu Han and worldwide reference populations was reconstructed based on the pairwise Nei’s genetic distances
Population genomic analyses revealed the genetic affinity and ancestral makeup of northwestern Han Chinese
We additionally generated genome-wide data from 50 modern Han Chinese individuals from Lanzhou in Gansu Province, northwestern China. We first conducted genome-wide data-based PCA to explore the genetic structure of Lanzhou Han under the genetic background of modern and ancient East Asians. We projected publicly available data from ancient individuals from Nepal, China, Japan, Mongolia and southern Siberia into modern PC plots. Lanzhou Han occupied the intersection of the Sino-Tibetan cline and northern Mongolic/Tungusic cline and deviated slightly toward southern Siberian populations (Fig. 3a). The previously reported middle/lower Yellow River Basin farmers during the early Neolithic-to-Iron Age were projected to be close to Lanzhou Han; however, geographically nearby upper Yellow River Basin farmers from the Ganqing region (late Neolithic Lajia and Jinchankou, and Iron Age Dacaozi) were slightly shifted to ancient Tibetan from Nepal and modern Tibetans. We further focused on the genetic backgrounds of Sino-Tibetans and other southern modern and ancient East Asians and performed one subregional East Asian PCA (Fig. 3b). We observed a clear Sino-Tibetan cline running between the highland modern Tibetan lineage represented by Lhasa Tibetan and the intersection region of the Hmong–Mien cline and Tai–Kadai cline, and Lanzhou Han partly overlapped with modern northern Han Chinese and ancient Yangshao and Longshan people from Henan Province, which indicated that northwestern Han Chinese, represented by Lanzhou Han, showed genetic affinity to ancient Yangshao/Longshan-related people and modern northern Han Chinese from the Shanxi, Henan and Shandong Provinces. Here, we could also identify the genetic separation of human populations from southern East Asia and Southeast Asia.
Relationship between Lanzhou Han and modern/ancient Eurasians based on the genome-wide genetic variations. a, b Principal component analysis (PCA) focused on populations from the eastern Eurasian (a) and southern East Asia (b). Ancient individuals were projected on the modern genetic backgrounds. c Ancestry composition among the newly genotyped and reference populations showed the genetic similarities and differences
Population clustering patterns inferred from the ADMIXTURE results among non-Africans showed four main ancestry components in Lanzhou Han. The minimum cross-validation value is 0.6701 when we predefined 11 ancestral population sources in the model-based clustering analyses (k = 11). As shown in Fig. 3c, we observed 2 dominant northern East Asian ancestries, represented by the Jomon lineage (0.106) and ancient Tibetan lineage (2125-year-old Mebrak, 0.423), and 2 southern East Asian ancestries, represented by the coastal proto-Austronesian lineage maximized in Taiwan Hanben (0.100) and the inland proto-Hmong–Mien lineage enriched in Hmong (0.218), with 11 predefined ancestral populations. In addition, we identified a small genetic contribution from the Baikal ancient lineage represented by early Bronze Age Ust-Belaya and proto-Austroasiatic Htin/Mang lineage (0.045) into northwestern Han. A model-based cluster of 12 ancestral sources also confirmed that Lanzhou Han derived their primary ancestry from Mebrak (0.417), Jomon (0.105), Hmong (0.220), and Hanben (0.099). Interestingly, we identified a small proportion (0.021) of ancestry descended from Basque or western Eurasian steppe pastoralist-related populations, as well as low amounts of gene flow from Htin (0.043) and Transbaikal Evenk (0.069) populations.
Consistent with the shared mosaic genetic components observed in the ADMIXTURE results and patterns of genetic variations in PCA, the shared genetic drift among modern Eurasian populations determined via outgroup-f3(Source1, Han_Lanzhou; Mbuti) showed that Lanzhou Han had a close genetic affinity with modern northern Han Chinese from geographically different regions (Fig. 4a). The stronger genomic affinity was further confirmed via outgroup-f3(Ancient Eurasian, Han_Lanzhou; Mbuti), which pointed out that middle Neolithic-to-Iron Age people from the Yellow River Basin in northern East Asia showed the most shared genetic drift with Lanzhou Han, especially for Luoheguxiang people in Henan (Fig. 4b). To further explore plausible ancestral sources for Lanzhou Han, we calculated admixture-f3 statistics in the form f3(Source1, Source2; Lanzhou Han) using 70,949 SNPs in 192 Eurasian modern and 177 ancient populations. After excluding 7401 ancient source pairs with fewer than 10,000 overlapping SNPs, we found that 8873 out of 66,519 pairs displayed significant admixture signals (negative-f3 values with Z-scores less than -3). In detail, the composition of northern and southern East Asians always produced the most negative admixed signatures, pointing to the main ancestral sources of Lanzhou Han from two lineages related to northern and southern East Asians, respectively, which may be associated with millet farmer predecessors from the Yellow River Basin and rice farmer predecessors from the Yangtze River Basin (Fig. 4c–e). Geographically distinct Han Chinese populations combined with one of the western Eurasian populations produced the most negative-f3 values, for example, f3(Chuvash, Han_Nanchong; Han_Lanzhou) = − 17.620*SE. We also identified negative-f3 values for combinations of one western modern Eurasian and one East Asian population, such as steppe pastoralists (Yamnaya, Afanasievo, Okunevo, and Andronovo) combined with Asians or ancient Yellow/Yangtze River Basin populations combined with western Eurasians (Fig. 4f–h).

source for Lanzhou Han when we assumed ancestral northern East Asians (c, d), ancestral southern East Asian Hanben (e), Western Eurasian steppe pastoralist as one of the source populations (f–h)
Results of three-population tests. a, b The shared genetic drift between Lanzhou Han and modern Eurasian reference populations (a) and ancient populations (b). c–h The plausible ancestral
Ancestral origins and genetic history reconstruction of northwestern Han via modern/ancient DNA perspectives
We performed affinity-four-population statistics to study the asymmetric genetic relationship between Lanzhou Han and other modern/ancient Eurasians in the form f4(Modern/ancient Eurasian reference1, Modern/ancient Eurasian reference2; Han_Lanzhou, Mbuti). The included reference populations could be categorized into six groups (Fig. 5a) based on the patterns of shared derived alleles. We observed significant negative-f4 values when we used group1 as reference population1 (populations listed on the left of the heatmap), including paleolithic East Asian lineage (40,000-year-old Tianyuan), Bronze Age western Eurasian pastoralists (Sintashta, Andronovo, Afanasievo, and Srubnaya people), Xinjiang ancient Shirenzigou people and modern Uyghur, Nepal Kusunda and the deeply diverged southern Eurasian lineage of Onge, which pointed to stronger northern East Asian affinity of Lanzhou Han. Compared with all other reference populations, we observed statistically positive f4 values in f4(Reference group6, Modern/ancient Eurasian reference2; Han_Lanzhou, Mbuti), indicating that Lanzhou Han shared the most derived alleles with group6, which comprised lowland Sino-Tibetan and ancient northern East Asians. We subsequently calculated the f4(Reference population, Han_Lanzhou; Ancestral source candidates, Mbuti) to validate the genetic continuity and admixture of northwestern Han. We assumed that if one ancestral source candidate A is the direct ancestor of Lanzhou Han, more shared alleles (negative-f4 values) between them should be observed. If ancestral source candidate A is the only direct ancestor of Lanzhou Han, there should be nonsignificant f4 values in f4(Ancestral source candidate, Han_Lanzhou; Reference population, Mbuti). As shown in Fig. 5b, we observed strong affinity signals in f4(Reference population, Han_Lanzhou; Ancestral source candidate, Mbuti); when we used ancient East Asians or modern northern East Asians as possible ancestral source candidates, we could observe the negative-f4 values, including the geographically close late Neolithic Shimao people from Shaanxi, late Neolithic Qijia people (Jinchankou and Lajia) and Iron Age Dacaozi from Gansu Province. The validation test of the unique ancestral population of Lanzhou Han was further validated using the 1240 K-based merged dataset, and no additional gene flow events were identified based on our included reference populations, which suggested a stronger genetic affinity between northern Ancient East Asians and modern Lanzhou Han. Marginal Z-scores could be produced when we used the Ami (Z-Score: − 2.692), Thai (− 2.507), Dai (− 2.383), Atayal (− 2.224), Sintashta (− 2.007) and Shirenzigou (− 1.835) as references and Lajia as an ancestral source, suggesting that compared with late Neolithic people in Gansu, modern people may have additional small contributions of genetic materials from southern East Asians and western Eurasians. These additional admixture signals were further evidenced via f4-statistics based on the Human-Origin-merged dataset (Table S7–8).
Genomic affinity between Lanzhou Han and other modern and ancient Eurasians. a Results from f4(Reference population1, reference population2; Lanzhou Han, Mbuti) showed the unequal derived alleles between Lanzhou Han and other modern/ancient populations compared with other East Asians. b Genetic admixture and continuity between Lanzhou Han and other modern and ancient source candidates inferred via f4(Reference population1, Lanzhou Han; Reference population2, Mbuti).
The results from the qpWave focused on Lanzhou Han showed that it could be fitted via the two-way admixture model and could be modeled as the admixture result of 0.862 ± 0.038 Lajia-related ancestry and 0.138 ± 0.038 Hanben-related ancestry (p_rank1: 0.1393), 0.883 ± 0.047 Shimao-related ancestry and 0.117 ± 0.047 Hanben-related ancestry (p_rank1: 0.5014), or 0.861 ± 0.049 Miaozigou-related ancestry and 0.139 ± 0.049 Hanben-related ancestry (p_rankl: 0.1068). To elucidate the genetic affinity between Lanzhou Han and western Eurasians, we also applied qpAdm modeling and a three-way admixture model to quantify the proportion of Western Eurasian ancestry in northwestern Han Chinese. When we used a French population as the western source proxy, Lanzhou Han was observed to be better fitted as an admixture of 0.815 ± 0.063 ancestry related to Iron Age Luoheguxiang people, 0.163 ± 0.056 ancestry related to Hanben, and 0.022 ± 0.014 related to French (p_rank2: 0.377). This three-way admixture model could also be well fitted if the middle and late steppe pastoralists of Andronovo were used as the western Eurasian source (0.816, 0.163, and 0.021; p_rank2: 0.343). In addition, three-way admixture models of Longtoushan–Hanben–French (0.739–0.244–0.016) and Longtoushan–Hanben–Andronovo (0.734–0.248–0.018) could also provide a good fit for Lanzhou Han’s admixture history. We also explored the phylogenetic relationships between Lanzhou Han and the surrounding modern and ancient Eurasian populations with the events of population splits and gene flow using graphics-based qpGraph modeling (Fig. 6). The two best-fitted qpGraph models showed the close genetic affinity between modern Lanzhou Han and ancient northern East Asian lineages represented by millet farmers in the Yellow River Basin. When we used the late Neolithic Shimao people as the proxy for northern sources, we were able to model 8% of northwestern Han derived from western Eurasians (Fig. 6a). When we considered the archaic genetic materials in the Non-African and Australasian groups, the northwestern Han could be modeled as a mixture of southern East Asian ancestry related to Hanben and a northern lineage close to northern East Asians and the southern Siberian lineage (Fig. 6b), which may contain western Eurasian admixture-derived alleles. Here, the pattern of genetic structure illuminated the admixture processes of primary ancestry derived from the admixture event between the southern East Asian lineage and northern East Asian or Siberian lineage and indicated minor genetic contributions from western Eurasia. To comprehensively characterize the formation of the gene pool of modern northwestern Han Chinese, we used ALDER to date the North-to-South and West-to-East admixture events based on the decay of admixture-induced linkage disequilibrium (Fig. 7). We detected an ancient admixture between southern East Asia and northern East Asia estimated at approximately the middle to historic Neolithic period with different ancestral source candidates (approximately 5000 BCE–1500 CE). We also tested the French, Basque and Greek populations as the western source and obtained a time of admixture of 30 generations ago (approximately 1000 years ago, during the Tang and Song Dynasties). We obtained contiguous intervals ranging from 24 to 30 generations ago for different western sources.
Admixture graph model of northwestern Han Chinese with the order of population splits and positions of admixture events. The largest deviation between empirical and theoretical f-statistics less than |Z|= 2.748, which indicated a good fit considering the large number of the f-statistics analyzed. Admixture events were labeled as dotted lines with corresponding admixture proportion and parameters of branch length were marked as 1000 times of f2 values
Uniparental genetic landscape of northwestern Han Chinese
We successfully obtained uniparental haplogroups for 49 male individuals (Table S9). For the mtDNA haplogroup, we assigned 49 mitochondrial genomes into 36 terminal haplogroups with frequencies ranging from 0.0204 to 0.0612 (F1a1,3), and D4 (14/49) was the dominant maternal lineage in the northwestern lineage. For the male-inherited Y chromosome, we obtained 38 terminal Y haplogroups with frequencies ranging from 0.0240 to 0.0816 (Q1a1a1a1a ~). We found that southern Siberian-dominant lineages (C2b, C2c, N1, and Q1a) appeared in our northwestern Chinese Han. The patterns of genetic diversity also showed the multiple genetic sources of modern northwestern Han Chinese.
Discussion
The Hexi Corridor and its surrounding regions are well known for the famous Majiayao culture in middle and late Neolithic times and subsequent control by the Rongdi tribe before the Han dynasty (Dong et al. 2017). In addition, this region was the main region of intersection of the eastward spread of barley/wheat agriculture and westward spread of millet technology in the Neolithic (Leipe et al. 2019). The westward migration of Han Chinese and their ancestors mainly occurred within the historic period, and the descendants of these migrants resided here permanently. Northwest China is the cradle of Trans-Eurasian cultural and genetic exchange (Dong et al. 2020). However, the genetic diversity, fine-scale genetic structure, and western Eurasian admixture signal of northwestern populations should be fully surveyed. Here, we conducted one genetic survey based on autosomal STR and genome-wide SNP analyses among 599 individuals to reconstruct the population genomic history of northwestern Chinese populations.
First, due to the controversy about the origin of Han Chinese and the model of the formation of this population, we uncovered that the main lineage of northwestern Han descended from the ancient northern East Asian lineage related closely to middle/upper Yellow River millet farmers or hunter-gatherers from the Mongolian Plateau (Ning et al. 2020; Wang et al. 2020a; Yang et al. 2020). This genomic affinity between modern northwestern Han and ancient northern East Asians supports North China as the origin of Sinitic-speaking populations. This is consistent with the hypothesis of co-origination of the Sino-Tibetan language family in North China, evidenced via the shared cognates and common homeland in the Bayesian-based phylogenetic relationship reconstruction (Sagart et al. 2019; Zhang et al. 2019). We also identified the gene flow from southern East Asian populations into the northwestern Han based on f-statistics and qpAdm/qpWave admixture models, suggesting a complex admixture pattern underlying the formation of the modern Han. Previous paternal/maternal DNA-based findings have demonstrated that the demic diffusion model promoted the formation of modern observed genetic diversity and variation in Han Chinese individuals (Wen et al. 2004). The fine-scale genetic structure presented here further suggests a revised model of North China origin and range expansion with local admixture, emphasizing the incorporation of genetic material from additional ancestral populations during the process of Han Chinese expansion.
Second, focusing on the West-To-East genetic connection, we identified a western Eurasian admixture signature in northwestern Han via f3/f4-statistics, which was confirmed in the qualitative analyses via qpAdm/qpGraph models. Here, we found that modern northwestern Han Chinese populations were derived from three ancestral populations: two major eastern Eurasian components (ancient northern East Asians related to the Yellow River millet farmers and ancient southern East Asians related to the Yangtze rice farmers) and one western Eurasian component. This complex pattern of ancestral admixture is especially interesting in that it is significantly different from the two-way admixture model of southern Han (He et al. 2020a, b). However, the most proximate ancestral sources of western Eurasian sources and corresponding admixture dates remain controversial. Previous ancient Neolithic genomes from Baikal Lake (de Barros Damgaard et al. 2018; Sikora et al. 2019) and the Yellow River Basin (Ning et al. 2020; Wang et al. 2020a; Yang et al. 2020) have demonstrated limited genetic contributions of western Eurasia to eastern Eurasia. Different from the complex mixing pattern observed in Europe (three-way admixture model of local hunter-gatherers, incoming Anatolian farmers and westward-spreading Yamnaya pastoralists) (Lazaridis et al. 2014), eastern Eurasia possessed relatively high genetic stability during the Neolithic revolution. Here, we also provide one possible process describing the introduction of western Eurasian ancestry into the northeastern Han. We suspect that this low level of Western Eurasian signal may have been introduced into North China after the Bronze Age via extensive population interactions during the globalization process. Indeed, our estimated date of western-eastern admixture mainly spanned from 1500 years ago to 500 years ago in historic time. It is known that the ALDER admixture time was estimated based on single admixture events; however, actual mixing events are continuous (Loh et al. 2013). Thus, these estimated dates are later than the initial admixture time. In addition, recent ancient genomes from Shirenzigou in Xinjiang Province identified significant western Eurasian Yamnaya ancestry 2000 years ago (Ning et al. 2019). The extent of the influence of Yamnaya-related ancestry on inland populations needs further genetic testing to determine. In total, clearer processes of the introduction of Western Eurasian ancestry into North China will be fully illuminated via population-scale analyses of additional ancient genomes to characterize temporally different populations from the Hexi Corridor in the future, although our current estimated date of admixture is consistent with the development of communication on the Silk Road.
In summary, our results falsified the hypothesis that the historic Trans-Eurasian populations around the Hexi Corridor during the Silk Road development period interacted with Han Chinese populations via cultural diffusion alone. Instead, the homogeneous genetic structure observed in modern northwestern Han Chinese harbors some extent of western Eurasian admixture (2% via qpAdm-based or 8% in qpGraph-based models) dating to approximately 1000 CE, suggesting that local populations (Ancestral Han) mixed with incoming western Eurasians, along with the adoption of technology and culture. In addition, no direct genetic continuity between geographically close-knit late Neolithic-to-historic Ganqing ancient populations (Lajia, Jinchankou and Dachaozi) was identified. The genomic affinity between northwestern Han and ancient northern East Asians demonstrated that the primary ancestry of northwestern Han Chinese populations was derived from ancestral populations related to northern millet farmers in the Yellow River Basin, suggesting their common origin in North China and recent northwestward expansion. Stronger affinity with southern modern and ancient East Asians compared with northern Neolithic populations, revealed via f3/f4, qpAdm/qpWave, and qpGraph, suggested the northward migration of ancient southern rice-farmer-related ancestral populations and their genetic contributions to modern northwestern Han Chinese populations. In conclusion, our results suggested that modern northwestern Han derived their ancestry from three different ancestral sources: two major East Asian groups, associated with millet and rice farmers, and one minor Western source.
Data availability
We submitted allele frequency data and other numerical data that underlies graphs or summary statistics in the supplementary material. Following the regulations and informed consent of this project approved via the Medical Ethics Committee of Xiamen University and the regulations of the Human Genetic Resources Administration of China (HGRAC), the obtained raw data can be shared via personal communication with corresponding authors. We make the data available upon request by asking the person requesting the data to agree in writing to the following restrictions: (I) the data can be only used for studying population history; (II) the data cannot be used for commercial purposes; (III) the data cannot be used as identify the sample donors; (IV) the data cannot be used for studying natural/cultural selections, medical or other related studies.
References
Alexander DH, Novembre J, Lange K (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19:1655–1664
Almeida C, Ribeiro T, Oliveira AR, Porto MJ, Costa Santos J, Dias D, Dario P (2015) Population data of the GlobalFiler((R)) express loci in South Portuguese population. Forensic Sci Int Genet 19:39–41
Alsafiah HM, Goodwin WH, Hadi S, Alshaikhi MA, Wepeba PP (2017) Population genetic data for 21 autosomal STR loci for the Saudi Arabian population using the GlobalFiler((R)) PCR amplification kit. Forensic Sci Int Genet 31:e59–e61
Bai F, Zhang X, Ji X, Cao P, Feng X, Yang R, Peng M, Pei S, Fu Q (2020) Paleolithic genetic link between Southern China and Mainland Southeast Asia revealed by ancient mitochondrial genomes. J Hum Genet 65(12):1125–1128
Cao Y, Li L, Xu M, Feng Z, Sun X, Lu J, Xu Y, Du P, Wang T, Hu R, Ye Z, Shi L, Tang X, Yan L, Gao Z, Chen G, Zhang Y, Chen L, Ning G, Bi Y, Wang W, China MAPC (2020) The ChinaMAP analytics of deep whole genome sequences in 10,588 individuals. Cell Res 30:717–731
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ (2015) Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4:7
Chen P, Adnan A, Rakha A, Wang M, Zou X, Mo X, He G (2019a) Population background exploration and genetic distribution analysis of Pakistan Hazara via 23 autosomal STRs. Ann Hum Biol 46:514–518
Chen P, Zou X, Wang B, Wang M, He G (2019b) Genetic admixture history and forensic characteristics of Turkic-speaking Kyrgyz population via 23 autosomal STRs. Ann Hum Biol 46:498–501
Chen P, Zou X, Wang M, Gao B, Su Y, He G (2019c) Forensic features and genetic structure of the Hotan Uyghur inferred from 27 forensic markers. Ann Hum Biol 46:589–600
Chen P, Wu J, Luo L, Gao H, Wang M, Zou X, Luo H, Yu L, Han Y, Jia F, He G (2019d) Population genetic analysis of modern and ancient DNA variations yields new insights into the formation, genetic structure and phylogenetic relationship of Northern Han Chinese. Foront Genet 10:1045
Choi EJ, Park KW, Lee YH, Nam YH, Suren G, Ganbold U, Kim JA, Kim SY, Kim HM, Kim K, Kim W (2017) Forensic and population genetic analyses of the GlobalFiler STR loci in the Mongolian population. Genes Genomics 39:423–431
Consortium HP-AS, Abdulla MA, Ahmed I, Assawamakin A, Bhak J, Brahmachari SK, Calacal GC, Chaurasia A, Chen CH, Chen J, Chen YT, Chu J, Cutiongco-de la Paz EM, De Ungria MC, Delfin FC, Edo J, Fuchareon S, Ghang H, Gojobori T, Han J, Ho SF, Hoh BP, Huang W, Inoko H, Jha P, Jinam TA, Jin L, Jung J, Kangwanpong D, Kampuansai J, Kennedy GC, Khurana P, Kim HL, Kim K, Kim S, Kim WY, Kimm K, Kimura R, Koike T, Kulawonganunchai S, Kumar V, Lai PS, Lee JY, Lee S, Liu ET, Majumder PP, Mandapati KK, Marzuki S, Mitchell W, Mukerji M, Naritomi K, Ngamphiw C, Niikawa N, Nishida N, Oh B, Oh S, Ohashi J, Oka A, Ong R, Padilla CD, Palittapongarnpim P, Perdigon HB, Phipps ME, Png E, Sakaki Y, Salvador JM, Sandraling Y, Scaria V, Seielstad M, Sidek MR, Sinha A, Srikummool M, Sudoyo H, Sugano S, Suryadi H, Suzuki Y, Tabbada KA, Tan A, Tokunaga K, Tongsima S, Villamor LP, Wang E, Wang Y, Wang H, Wu JY, Xiao H, Xu S, Yang JO, Shugart YY, Yoo HS, Yuan W, Zhao G, Zilfalil BA, Indian Genome Variation C (2009) Mapping human genetic diversity in Asia. Science 326:1541–1545
Cummings MP (2004) PHYLIP (Phylogeny Inference Package). Wiley, Hoboken
Damgaard PB, Marchi N, Rasmussen S, Peyrot M, Renaud G, Korneliussen T, Moreno-Mayar JV, Pedersen MW, Goldberg A, Usmanova E, Baimukhanov N, Loman V, Hedeager L, Pedersen AG, Nielsen K, Afanasiev G, Akmatov K, Aldashev A, Alpaslan A, Baimbetov G, Bazaliiskii VI, Beisenov A, Boldbaatar B, Boldgiv B, Dorzhu C, Ellingvag S, Erdenebaatar D, Dajani R, Dmitriev E, Evdokimov V, Frei KM, Gromov A, Goryachev A, Hakonarson H, Hegay T, Khachatryan Z, Khaskhanov R, Kitov E, Kolbina A, Kubatbek T, Kukushkin A, Kukushkin I, Lau N, Margaryan A, Merkyte I, Mertz IV, Mertz VK, Mijiddorj E, Moiyesev V, Mukhtarova G, Nurmukhanbetov B, Orozbekova Z, Panyushkina I, Pieta K, Smrcka V, Shevnina I, Logvin A, Sjogren KG, Stolcova T, Taravella AM, Tashbaeva K, Tkachev A, Tulegenov T, Voyakin D, Yepiskoposyan L, Undrakhbold S, Varfolomeev V, Weber A, Wilson Sayres MA, Kradin N, Allentoft ME, Orlando L, Nielsen R, Sikora M, Heyer E, Kristiansen K, Willerslev E (2018) 137 ancient human genomes from across the Eurasian steppes. Nature 557:369–374
de Barros DP, Martiniano R, Kamm J, Moreno-Mayar JV, Kroonen G, Peyrot M, Barjamovic G, Rasmussen S, Zacho C, Baimukhanov N, Zaibert V, Merz V, Biddanda A, Merz I, Loman V, Evdokimov V, Usmanova E, Hemphill B, Seguin-Orlando A, Yediay FE, Ullah I, Sjögren K-G, Iversen KH, Choin J, de la Fuente C, Ilardo M, Schroeder H, Moiseyev V, Gromov A, Polyakov A, Omura S, Senyurt SY, Ahmad H, McKenzie C, Margaryan A, Hameed A, Samad A, Gul N, Khokhar MH, Goriunova OI, Bazaliiskii VI, Novembre J, Weber AW, Orlando L, Allentoft ME, Nielsen R, Kristiansen K, Sikora M, Outram AK, Durbin R, Willerslev E (2018) The first horse herders and the impact of early Bronze age steppe expansions into Asia. Science 360:eaar7711
Dong G, Yang Y, Liu X, Li H, Cui Y, Wang H, Chen G, Dodson J, Chen F (2017) Prehistoric trans-continental cultural exchange in the Hexi Corridor, northwest China. Holocene 28:621–628
Dong G, Du L, Wei W (2020) The impact of early trans-Eurasian exchange on animal utilization in northern China during 5000–2500 BP. The Holocene:0959683620941169
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Fujii K, Iwashima Y, Kitayama T, Nakahara H, Mizuno N, Sekiguchi K (2014) Allele frequencies for 22 autosomal short tandem repeat loci obtained by PowerPlex Fusion in a sample of 1501 individuals from the Japanese population. Leg Med (Tokyo) 16:234–237
Gakuhari T, Nakagome S, Rasmussen S, Allentoft ME, Sato T, Korneliussen T, Chuinneagain BN, Matsumae H, Koganebuchi K, Schmidt R, Mizushima S, Kondo O, Shigehara N, Yoneda M, Kimura R, Ishida H, Masuyama T, Yamada Y, Tajima A, Shibata H, Toyoda A, Tsurumoto T, Wakebe T, Shitara H, Hanihara T, Willerslev E, Sikora M, Oota H (2020) Ancient Jomon genome sequence analysis sheds light on migration patterns of early East Asian populations. Commun Biol 3:437
Gaviria A, Zambrano AK, Morejon G, Galarza J, Aguirre V, Vela M, Builes JJ, Burgos G (2013) Twenty two autosomal microsatellite data from Ecuador (Powerplex Fusion). Forensic Sci Int Genet Suppl Ser 4:e330–e333
Gouy A, Zieger M (2017) STRAF-A convenient online tool for STR data evaluation in forensic genetics. Forensic Sci Int Genet 30:148–151
Guerreiro S, Ribeiro T, Porto MJ, Carneiro de Sousa MJ, Dario P (2017) Characterization of GlobalFiler loci in Angolan and Guinean populations inhabiting Southern Portugal. Int J Legal Med 131:365–368
Haak W, Lazaridis I, Patterson N, Rohland N, Mallick S, Llamas B, Brandt G, Nordenfelt S, Harney E, Stewardson K, Fu Q, Mittnik A, Banffy E, Economou C, Francken M, Friederich S, Pena RG, Hallgren F, Khartanovich V, Khokhlov A, Kunst M, Kuznetsov P, Meller H, Mochalov O, Moiseyev V, Nicklisch N, Pichler SL, Risch R, Rojo Guerra MA, Roth C, Szecsenyi-Nagy A, Wahl J, Meyer M, Krause J, Brown D, Anthony D, Cooper A, Alt KW, Reich D (2015) Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 522:207–211
He G, Wang M, Liu J, Hou Y, Wang Z (2018a) Forensic features and phylogenetic analyses of Sichuan Han population via 23 autosomal STR loci included in the Huaxia Platinum System. Int J Legal Med 132:1079–1082
He G, Wang Z, Wang M, Luo T, Liu J, Zhou Y, Gao B, Hou Y (2018b) Forensic ancestry analysis in two Chinese minority populations using massively parallel sequencing of 165 ancestry-informative SNPs. Electrophoresis 39:2732–2742
He G, Wang Z, Wang M, Zou X, Liu J, Wang S, Hou Y (2018c) Genetic variations and forensic characteristics of Han Chinese population residing in the Pearl River Delta revealed by 23 autosomal STRs. Mol Biol Rep 45:1125–1133
He G, Wang Z, Guo J, Wang M, Zou X, Tang R, Liu J, Zhang H, Li Y, Hu R, Wei LH, Chen G, Wang CC, Hou Y (2020a) Inferring the population history of Tai-Kadai-speaking people and southernmost Han Chinese on Hainan Island by genome-wide array genotyping. Eur J Hum Genet 28:1111–1123
He GL, Li YX, Wang MG, Zou X, Yeh HY, Yang XM, Wang Z, Tang RK, Zhu SM, Guo JX, Luo T, Zhao J, Sun J, Xia ZY, Fan HL, Hu R, Wei LH, Chen G, Hou YP, Wang CC (2020b) Fine-scale genetic structure of Tujia and central Han Chinese revealing massive genetic admixture under language borrowing. J Syst Evol 59(1):1–20
Hossain T, Hasan M, Mazumder AK, Momtaz P, Sufian A, Khandaker JA, Akhteruzzaman S (2016) Genetic polymorphism studies on 22 autosomal STR loci of the PowerPlex Fusion System in Bangladeshi population. Leg Med (Tokyo) 23:44–46
Jeong C, Ozga AT, Witonsky DB, Malmstrom H, Edlund H, Hofman CA, Hagan RW, Jakobsson M, Lewis CM, Aldenderfer MS, Di Rienzo A, Warinner C (2016) Long-term genetic stability and a high-altitude East Asian origin for the peoples of the high valleys of the Himalayan arc. Proc Natl Acad Sci U S A 113:7485–7490
Jeong C, Balanovsky O, Lukianova E, Kahbatkyzy N, Flegontov P, Zaporozhchenko V, Immel A, Wang CC, Ixan O, Khussainova E, Bekmanov B, Zaibert V, Lavryashina M, Pocheshkhova E, Yusupov Y, Agdzhoyan A, Koshel S, Bukin A, Nymadawa P, Turdikulova S, Dalimova D, Churnosov M, Skhalyakho R, Daragan D, Bogunov Y, Bogunova A, Shtrunov A, Dubova N, Zhabagin M, Yepiskoposyan L, Churakov V, Pislegin N, Damba L, Saroyants L, Dibirova K, Atramentova L, Utevska O, Idrisov E, Kamenshchikova E, Evseeva I, Metspalu M, Outram AK, Robbeets M, Djansugurova L, Balanovska E, Schiffels S, Haak W, Reich D, Krause J (2019) The genetic history of admixture across inner Eurasia. Nat Ecol Evol 3:966–976
Jin X, Wei Y, Chen J, Kong T, Mu Y, Guo Y, Dong Q, Xie T, Meng H, Zhang M, Li J, Li X, Zhu B (2017) Phylogenic analysis and forensic genetic characterization of Chinese Uyghur group via autosomal multi STR markers. Oncotarget 8:73837–73845
Kovach WL (2007) MVSP-A MultiVariate statistical package for windows, ver. 3.1. Kovach Computing Services, Pentraeth
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lazaridis I, Patterson N, Mittnik A, Renaud G, Mallick S, Kirsanow K, Sudmant PH, Schraiber JG, Castellano S, Lipson M, Berger B, Economou C, Bollongino R, Fu Q, Bos KI, Nordenfelt S, Li H, de Filippo C, Prufer K, Sawyer S, Posth C, Haak W, Hallgren F, Fornander E, Rohland N, Delsate D, Francken M, Guinet JM, Wahl J, Ayodo G, Babiker HA, Bailliet G, Balanovska E, Balanovsky O, Barrantes R, Bedoya G, Ben-Ami H, Bene J, Berrada F, Bravi CM, Brisighelli F, Busby GB, Cali F, Churnosov M, Cole DE, Corach D, Damba L, van Driem G, Dryomov S, Dugoujon JM, Fedorova SA, Gallego Romero I, Gubina M, Hammer M, Henn BM, Hervig T, Hodoglugil U, Jha AR, Karachanak-Yankova S, Khusainova R, Khusnutdinova E, Kittles R, Kivisild T, Klitz W, Kucinskas V, Kushniarevich A, Laredj L, Litvinov S, Loukidis T, Mahley RW, Melegh B, Metspalu E, Molina J, Mountain J, Nakkalajarvi K, Nesheva D, Nyambo T, Osipova L, Parik J, Platonov F, Posukh O, Romano V, Rothhammer F, Rudan I, Ruizbakiev R, Sahakyan H, Sajantila A, Salas A, Starikovskaya EB, Tarekegn A, Toncheva D, Turdikulova S, Uktveryte I, Utevska O, Vasquez R, Villena M, Voevoda M, Winkler CA, Yepiskoposyan L, Zalloua P, Zemunik T, Cooper A, Capelli C, Thomas MG, Ruiz-Linares A, Tishkoff SA, Singh L, Thangaraj K, Villems R, Comas D, Sukernik R, Metspalu M, Meyer M, Eichler EE, Burger J, Slatkin M, Paabo S, Kelso J, Reich D, Krause J (2014) Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513:409–413
Leipe C, Long T, Sergusheva EA, Wagner M, Tarasov PE (2019) Discontinuous spread of millet agriculture in eastern Asia and prehistoric population dynamics. Sci Adv 5:eaax6225
Li L, Zou X, Zhang G, Wang H, Su Y, Wang M, He G (2020) Population genetic analysis of Shaanxi male Han Chinese population reveals genetic differentiation and homogenization of East Asians. Mol Genet Genomic Med 8:e1209
Lipson M, Cheronet O, Mallick S, Rohland N, Oxenham M, Pietrusewsky M, Pryce TO, Willis A, Matsumura H, Buckley H, Domett K, Nguyen GH, Trinh HH, Kyaw AA, Win TT, Pradier B, Broomandkhoshbacht N, Candilio F, Changmai P, Fernandes D, Ferry M, Gamarra B, Harney E, Kampuansai J, Kutanan W, Michel M, Novak M, Oppenheimer J, Sirak K, Stewardson K, Zhang Z, Flegontov P, Pinhasi R, Reich D (2018) Ancient genomes document multiple waves of migration in Southeast Asian prehistory. Science 361:92–95
Liu S, Huang S, Chen F, Zhao L, Yuan Y, Francis SS, Fang L, Li Z, Lin L, Liu R, Zhang Y, Xu H, Li S, Zhou Y, Davies RW, Liu Q, Walters RG, Lin K, Ju J, Korneliussen T, Yang MA, Fu Q, Wang J, Zhou L, Krogh A, Zhang H, Wang W, Chen Z, Cai Z, Yin Y, Yang H, Mao M, Shendure J, Wang J, Albrechtsen A, Jin X, Nielsen R, Xu X (2018) Genomic analyses from non-invasive prenatal testing reveal genetic associations, patterns of viral infections, and Chinese population history. Cell 175(347–359):e314
Liu J, Wang Z, He G, Wang M, Hou Y (2019) Genetic polymorphism and phylogenetic differentiation of the Huaxia Platinum System in three Chinese minority ethnicities. Sci Rep 9:3371
Loh PR, Lipson M, Patterson N, Moorjani P, Pickrell JK, Reich D, Berger B (2013) Inferring admixture histories of human populations using linkage disequilibrium. Genetics 193:1233–1254
Mallick S, Li H, Lipson M, Mathieson I, Gymrek M, Racimo F, Zhao M, Chennagiri N, Nordenfelt S, Tandon A, Skoglund P, Lazaridis I, Sankararaman S, Fu Q, Rohland N, Renaud G, Erlich Y, Willems T, Gallo C, Spence JP, Song YS, Poletti G, Balloux F, van Driem G, de Knijff P, Romero IG, Jha AR, Behar DM, Bravi CM, Capelli C, Hervig T, Moreno-Estrada A, Posukh OL, Balanovska E, Balanovsky O, Karachanak-Yankova S, Sahakyan H, Toncheva D, Yepiskoposyan L, Tyler-Smith C, Xue Y, Abdullah MS, Ruiz-Linares A, Beall CM, Di Rienzo A, Jeong C, Starikovskaya EB, Metspalu E, Parik J, Villems R, Henn BM, Hodoglugil U, Mahley R, Sajantila A, Stamatoyannopoulos G, Wee JT, Khusainova R, Khusnutdinova E, Litvinov S, Ayodo G, Comas D, Hammer MF, Kivisild T, Klitz W, Winkler CA, Labuda D, Bamshad M, Jorde LB, Tishkoff SA, Watkins WS, Metspalu M, Dryomov S, Sukernik R, Singh L, Thangaraj K, Paabo S, Kelso J, Patterson N, Reich D (2016) The simons genome diversity project: 300 genomes from 142 diverse populations. Nature 538:201–206
McColl H, Racimo F, Vinner L, Demeter F, Gakuhari T, Moreno-Mayar JV, van Driem G, Gram Wilken U, Seguin-Orlando A, de la Fuente CC, Wasef S, Shoocongdej R, Souksavatdy V, Sayavongkhamdy T, Saidin MM, Allentoft ME, Sato T, Malaspinas AS, Aghakhanian FA, Korneliussen T, Prohaska A, Margaryan A, de Barros DP, Kaewsutthi S, Lertrit P, Nguyen TMH, Hung HC, Minh Tran T, Nghia Truong H, Nguyen GH, Shahidan S, Wiradnyana K, Matsumae H, Shigehara N, Yoneda M, Ishida H, Masuyama T, Yamada Y, Tajima A, Shibata H, Toyoda A, Hanihara T, Nakagome S, Deviese T, Bacon AM, Duringer P, Ponche JL, Shackelford L, Patole-Edoumba E, Nguyen AT, Bellina-Pryce B, Galipaud JC, Kinaston R, Buckley H, Pottier C, Rasmussen S, Higham T, Foley RA, Lahr MM, Orlando L, Sikora M, Phipps ME, Oota H, Higham C, Lambert DM, Willerslev E (2018) The prehistoric peopling of Southeast Asia. Science 361:88–92
Moyses CB, Tsutsumida WM, Raimann PE, da Motta CH, Nogueira TL, Dos Santos OC, de Figueiredo BB, Mishima TF, Candido IM, de Oliveira Godinho NM, Beltrami LS, Lopes RK, Guidolin AF, Mantovani A, Dos Santos SM, de Souza CA, Gusmao L (2017) Population data of the 21 autosomal STRs included in the GlobalFiler((R)) kits in population samples from five Brazilian regions. Forensic Sci Int Genet 26:e28–e30
Narasimhan VM, Patterson N, Moorjani P, Rohland N, Bernardos R, Mallick S, Lazaridis I, Nakatsuka N, Olalde I, Lipson M, Kim AM, Olivieri LM, Coppa A, Vidale M, Mallory J, Moiseyev V, Kitov E, Monge J, Adamski N, Alex N, Broomandkhoshbacht N, Candilio F, Callan K, Cheronet O, Culleton BJ, Ferry M, Fernandes D, Freilich S, Gamarra B, Gaudio D, Hajdinjak M, Harney E, Harper TK, Keating D, Lawson AM, Mah M, Mandl K, Michel M, Novak M, Oppenheimer J, Rai N, Sirak K, Slon V, Stewardson K, Zalzala F, Zhang Z, Akhatov G, Bagashev AN, Bagnera A, Baitanayev B, Bendezu-Sarmiento J, Bissembaev AA, Bonora GL, Chargynov TT, Chikisheva T, Dashkovskiy PK, Derevianko A, Dobes M, Douka K, Dubova N, Duisengali MN, Enshin D, Epimakhov A, Fribus AV, Fuller D, Goryachev A, Gromov A, Grushin SP, Hanks B, Judd M, Kazizov E, Khokhlov A, Krygin AP, Kupriyanova E, Kuznetsov P, Luiselli D, Maksudov F, Mamedov AM, Mamirov TB, Meiklejohn C, Merrett DC, Micheli R, Mochalov O, Mustafokulov S, Nayak A, Pettener D, Potts R, Razhev D, Rykun M, Sarno S, Savenkova TM, Sikhymbaeva K, Slepchenko SM, Soltobaev OA, Stepanova N, Svyatko S, Tabaldiev K, Teschler-Nicola M, Tishkin AA, Tkachev VV, Vasilyev S, Veleminsky P, Voyakin D, Yermolayeva A, Zahir M, Zubkov VS, Zubova A, Shinde VS, Lalueza-Fox C, Meyer M, Anthony D, Boivin N, Thangaraj K, Kennett DJ, Frachetti M, Pinhasi R, Reich D (2019) The formation of human populations in South and Central Asia. Science 365:eaat7487
Nicogossian A, Kloiber O, Stabile B (2014) The revised world medical association’s declaration of Helsinki 2013: enhancing the protection of human research subjects and empowering ethics review committees. World Med Health Policy 6:1–3
Ning C, Wang C-C, Gao S, Yang Y, Zhang X, Wu X, Zhang F, Nie Z, Tang Y, Robbeets M, Ma J, Krause J, Cui Y (2019) Ancient genomes reveal Yamnaya-related ancestry and a potential source of Indo-European speakers in iron age Tianshan. Curr Biol 29:2526-2532.e2524
Ning C, Li T, Wang K, Zhang F, Li T, Wu X, Gao S, Zhang Q, Zhang H, Hudson MJ, Dong G, Wu S, Fang Y, Liu C, Feng C, Li W, Han T, Li R, Wei J, Zhu Y, Zhou Y, Wang CC, Fan S, Xiong Z, Sun Z, Ye M, Sun L, Wu X, Liang F, Cao Y, Wei X, Zhu H, Zhou H, Krause J, Robbeets M, Jeong C, Cui Y (2020) Ancient genomes from northern China suggest links between subsistence changes and human migration. Nat Commun 11:2700
Ossowski A, Diepenbroek M, Szargut M, Zielinska G, Jedrzejczyk M, Berent J, Jacewicz R (2017) Population analysis and forensic evaluation of 21 autosomal loci included in GlobalFiler PCR Kit in Poland. Forensic Sci Int Genet 29:e38–e39
Park JH, Hong SB, Kim JY, Chong Y, Han S, Jeon CH, Ahn HJ (2013) Genetic variation of 23 autosomal STR loci in Korean population. Forensic Sci Int Genet 7:e76-77
Park HC, Kim K, Nam Y, Park J, Lee J, Lee H, Kwon H, Jin H, Kim W, Kim W, Lim S (2016) Population genetic study for 24 STR loci and Y indel (GlobalFiler PCR Amplification kit and PowerPlex(R) Fusion system) in 1000 Korean individuals. Leg Med (Tokyo) 21:53–57
Patterson N, Price AL, Reich D (2006) Population structure and eigenanalysis. PLoS Genet 2:e190
Patterson N, Moorjani P, Luo Y, Mallick S, Rohland N, Zhan Y, Genschoreck T, Webster T, Reich D (2012) Ancient admixture in human history. Genetics 192:1065–1093
Raghavan M, Skoglund P, Graf KE, Metspalu M, Albrechtsen A, Moltke I, Rasmussen S, Stafford TW Jr, Orlando L, Metspalu E, Karmin M, Tambets K, Rootsi S, Magi R, Campos PF, Balanovska E, Balanovsky O, Khusnutdinova E, Litvinov S, Osipova LP, Fedorova SA, Voevoda MI, DeGiorgio M, Sicheritz-Ponten T, Brunak S, Demeshchenko S, Kivisild T, Villems R, Nielsen R, Jakobsson M, Willerslev E (2014) Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature 505:87–91
Rasmussen M, Guo X, Wang Y, Lohmueller KE, Rasmussen S, Albrechtsen A, Skotte L, Lindgreen S, Metspalu M, Jombart T, Kivisild T, Zhai W, Eriksson A, Manica A, Orlando L, De La Vega FM, Tridico S, Metspalu E, Nielsen K, Avila-Arcos MC, Moreno-Mayar JV, Muller C, Dortch J, Gilbert MT, Lund O, Wesolowska A, Karmin M, Weinert LA, Wang B, Li J, Tai S, Xiao F, Hanihara T, van Driem G, Jha AR, Ricaut FX, de Knijff P, Migliano AB, Gallego Romero I, Kristiansen K, Lambert DM, Brunak S, Forster P, Brinkmann B, Nehlich O, Bunce M, Richards M, Gupta R, Bustamante CD, Krogh A, Foley RA, Lahr MM, Balloux F, Sicheritz-Ponten T, Villems R, Nielsen R, Wang J, Willerslev E (2011) An Aboriginal Australian genome reveals separate human dispersals into Asia. Science 334:94–98
Sadam M, Tasa G, Tiidla A, Lang A, Axelsson EP, Pajnic IZ (2015) Population data for 22 autosomal STR loci from Estonia. Int J Legal Med 129:1219–1220
Sagart L, Jacques G, Lai Y, Ryder RJ, Thouzeau V, Greenhill SJ, List JM (2019) Dated language phylogenies shed light on the ancestry of Sino-Tibetan. Proc Natl Acad Sci U S A 116:10317–10322
Sikora M, Pitulko VV, Sousa VC, Allentoft ME, Vinner L, Rasmussen S, Margaryan A, de Barros DP, de la Fuente C, Renaud G, Yang MA, Fu Q, Dupanloup I, Giampoudakis K, Nogues-Bravo D, Rahbek C, Kroonen G, Peyrot M, McColl H, Vasilyev SV, Veselovskaya E, Gerasimova M, Pavlova EY, Chasnyk VG, Nikolskiy PA, Gromov AV, Khartanovich VI, Moiseyev V, Grebenyuk PS, Fedorchenko AY, Lebedintsev AI, Slobodin SB, Malyarchuk BA, Martiniano R, Meldgaard M, Arppe L, Palo JU, Sundell T, Mannermaa K, Putkonen M, Alexandersen V, Primeau C, Baimukhanov N, Malhi RS, Sjogren KG, Kristiansen K, Wessman A, Sajantila A, Lahr MM, Durbin R, Nielsen R, Meltzer DJ, Excoffier L, Willerslev E (2019) The population history of northeastern Siberia since the Pleistocene. Nature 570:182–188
Stoneking M, Delfin F (2010) The human genetic history of East Asia: weaving a complex tapestry. Curr Biol 20:R188-193
Su B, Xiao J, Underhill P, Deka R, Zhang W, Akey J, Huang W, Shen D, Lu D, Luo J, Chu J, Tan J, Shen P, Davis R, Cavalli-Sforza L, Chakraborty R, Xiong M, Du R, Oefner P, Chen Z, Jin L (1999) Y-Chromosome evidence for a northward migration of modern humans into Eastern Asia during the last Ice Age. Am J Hum Genet 65:1718–1724
Taylor D, Bright JA, McGovern C, Neville S, Grover D (2017) Allele frequency database for GlobalFiler STR loci in Australian and New Zealand populations. Forensic Sci Int Genet 28:e38–e40
Wang Z, Zhou D, Jia Z, Li L, Wu W, Li C, Hou Y (2016) Developmental Validation of the Huaxia Platinum System and application in 3 main ethnic groups of China. Sci Rep 6:31075
Wang M, Wang Z, He G, Jia Z, Liu J, Hou Y (2018a) Genetic characteristics and phylogenetic analysis of three Chinese ethnic groups using the Huaxia Platinum System. Sci Rep 8:2429
Wang Z, He G, Luo T, Zhao X, Liu J, Wang M, Zhou D, Chen X, Li C, Hou Y (2018b) Massively parallel sequencing of 165 ancestry informative SNPs in two Chinese Tibetan-Burmese minority ethnicities. Forensic Sci Int Genet 34:141–147
Wang C-C, Yeh H-Y, Popov AN, Zhang H-Q, Matsumura H, Sirak K, Cheronet O, Kovalev A, Rohland N, Kim AM, Bernardos R, Tumen D, Zhao J, Liu Y-C, Liu J-Y, Mah M, Mallick S, Wang K, Zhang Z, Adamski N, Broomandkhoshbacht N, Callan K, Culleton BJ, Eccles L, Lawson AM, Michel M, Oppenheimer J, Stewardson K, Wen S, Yan S, Zalzala F, Chuang R, Huang C-J, Shiung C-C, Nikitin YG, Tabarev AV, Tishkin AA, Lin S, Sun Z-Y, Wu X-M, Yang T-L, Hu X, Chen L, Du H, Bayarsaikhan J, Mijiddorj E, Erdenebaatar D, Iderkhangai T-O, Myagmar E, Kanzawa-Kiriyama H, Nishino M, Shinoda K-i, Shubina OA, Guo J, Deng Q, Kang L, Li D, Li D, Lin R, Cai W, Shrestha R, Wang L-X, Wei L, Xie G, Yao H, Zhang M, He G, Yang X, Hu R, Robbeets M, Schiffels S, Kennett DJ, Jin L, Li H, Krause J, Pinhasi R, Reich D (2020a) The Genomic formation of human populations in East Asia. bioRxiv:2020.2003.2025.004606
Wang M, Zou X, Ye H-Y, Wang Z, Liu Y, Liu J, Wang F, Yao H, Chen P, Tao R, Wang S, Wei L-H, Tang R, Wang C-C, He G (2020b) Peopling of Tibet Plateau and multiple waves of admixture of Tibetans inferred from both modern and ancient genome-wide data. bioRxiv:2020.2007.2003.185884
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Wen B, Li H, Lu D, Song X, Zhang F, He Y, Li F, Gao Y, Mao X, Zhang L, Qian J, Tan J, Jin J, Huang W, Deka R, Su B, Chakraborty R, Jin L (2004) Genetic evidence supports demic diffusion of Han culture. Nature 431:302–305
Wu L, Pei B, Ran P, Song X (2017) Population genetic analysis of Xiamen Han population on 21 short tandem repeat loci. Leg Med (Tokyo) 26:41–44
Yang MA, Gao X, Theunert C, Tong H, Aximu-Petri A, Nickel B, Slatkin M, Meyer M, Paabo S, Kelso J, Fu Q (2017) 40,000-Year-old individual from Asia provides insight into early population structure in Eurasia. Curr Biol 27(3202–3208):e3209
Yang L, Zhang X, Zhao L, Sun Y, Li J, Huang R, Hu L, Nie S (2018) Population data of 23 autosomal STR loci in the Chinese Han population from Guangdong Province in southern China. Int J Legal Med 132:133–135
Yang MA, Fan X, Sun B, Chen C, Lang J, Ko YC, Tsang CH, Chiu H, Wang T, Bao Q, Wu X, Hajdinjak M, Ko AM, Ding M, Cao P, Yang R, Liu F, Nickel B, Dai Q, Feng X, Zhang L, Sun C, Ning C, Zeng W, Zhao Y, Zhang M, Gao X, Cui Y, Reich D, Stoneking M, Fu Q (2020) Ancient DNA indicates human population shifts and admixture in northern and southern China. Science 369:282–288
Zhang M, Fu Q (2020) Human evolutionary history in Eastern Eurasia using insights from ancient DNA. Curr Opin Genet Dev 62:78–84
Zhang H, Xia M, Qi L, Dong L, Song S, Ma T, Yang S, Jin L, Li L, Li S (2016a) Forensic and population genetic analysis of Xinjiang Uyghur population on 21 short tandem repeat loci of 6-dye GlobalFiler PCR Amplification kit. Forensic Sci Int Genet 22:22–24
Zhang H, Yang S, Guo W, Ren B, Pu L, Ma T, Xia M, Jin L, Li L, Li S (2016b) Population genetic analysis of the GlobalFiler STR loci in 748 individuals from the Kazakh population of Xinjiang in northwest China. Int J Legal Med 130:1187–1189
Zhang M, Yan S, Pan W, Jin L (2019) Phylogenetic evidence for Sino-Tibetan origin in northern China in the Late Neolithic. Nature 569:112–115
Funding
HBY was supported by the National Natural Science Foundation of China (31760309), Foundation for Humanities and Social Sciences Research Of the Ministry of Education (18YJAZH116), Scientific research project of Colleges and universities in Gansu Province (2017B-34), Gansu University of Political Science and Law major scientific research projects (2017XZD10), Lanzhou Talent Innovation and Entrepreneurship Project (2018-RC-113) and Gansu province guides science and technology innovation special project (2018ZX03). CCW was supported by the Nanqiang Outstanding Young Talents Program of Xiamen University (X2123302), National Natural Science Foundation of China (31801040), and Fundamental Research Funds for the Central Universities (ZK1144). YXL was supported by the National Postdoctoral Program for Innovative Talents (BX20180180). GLH was funded by China Postdoctoral Science Foundation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All the author declare no conflict of interest.
Ethical approval
This study and corresponding protocols were reviewed and approved via the Medical Ethics Committee of Xiamen University (XDYX201909).
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Communicated by Shuhua Xu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
438_2021_1767_MOESM1_ESM.xlsx
Supplementary file1 Table S1 p values of linkage-disequilibrium of 23 autosomal STRs in Gansu Han population (XLSX 16 KB)
438_2021_1767_MOESM4_ESM.xlsx
Supplementary file4 Table S4 Pairwise Fst genetic distance between Gansu Han and other 16 eastern Eurasian reference population based on genetic variations of 23 STRs (XLSX 13 KB)
438_2021_1767_MOESM5_ESM.xlsx
Supplementary file5 Table S5 Pairwise Fst genetic distance between Gansu Han and other 19 Eurasian reference population based on genetic variations of 20 STRs (XLSX 14 KB)
438_2021_1767_MOESM6_ESM.xlsx
Supplementary file6 Table S6 Pairwise Cavalli-Sforza genetic distance between Gansu Han and other 56 worldwide reference population based on allele frequency of 20 STRs (XLSX 36 KB)
438_2021_1767_MOESM7_ESM.xlsx
Supplementary file7 Table S7 Results of f4(Reference population1, Reference population2; Han_Lanzhou, Mbuti) based on Human-Origin-merge dataset (XLSX 1109 KB)
438_2021_1767_MOESM8_ESM.xlsx
Supplementary file8 Table S8 Results of f4(Reference population1, Han_Lanzhou; Reference population2, Mbuti) based on Human-Origin-merge dataset (XLSX 1075 KB)
438_2021_1767_MOESM10_ESM.pdf
Supplementary file10 Figure S1. Geographical position of Lanzhou, in northwest China. Figure S2. Allele frequencies of twenty-three short tandem repeats (STRs) in Lanzhou Han. Figure S3. Genetic relationships among geographically/ethnically diverse East Asians inferred from 23 autosomal STRs. (A) Multidimensional scaling plots (MDS) showed the genetic similarities among 17 East Asian populations; (B) Heat map based on the pairwise genetic distance (Fst) showed the genetic affinity among the studied and the reference populations. (C) The phylogenetic relationship base on the Fst genetic matrix among 17 populations showed genetic phylogeny. (D) Genetic similarities and differences inferred by model-based ancestry dissection. Figure S4. The pairwise Cavalli-Sforza genetic distances between Gansu Han and other 56 worldwide reference populations.Figure S5. Genetic similarities and differences among Lanzhou Han and other 56 worldwide reference populations (PDF 1026 KB)
Rights and permissions
About this article
Cite this article
Yao, H., Wang, M., Zou, X. et al. New insights into the fine-scale history of western–eastern admixture of the northwestern Chinese population in the Hexi Corridor via genome-wide genetic legacy. Mol Genet Genomics 296, 631–651 (2021). https://doi.org/10.1007/s00438-021-01767-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-021-01767-0