Abstract
The spotted wing drosophila, D. suzukii, is a serious agricultural pest attacking a variety of soft fruits and vegetables. Although originating from East Asia it has recently invaded America and Europe raising major concern about its expansion potential and the consequent economic losses. Since cytogenetic information on the species is scarce, we report here the mitotic karyotype and detailed photographic maps of the salivary gland polytene chromosomes of D. suzukii. The mitotic metaphase complement contains three pairs of autosomes, one of which is dot-like, and one pair of heteromorphic (XX/XY) sex chromosomes. The salivary gland polytene complement consists of five long polytene arms, representing the two metacentric autosomes and the acrocentric X chromosome, and one very short polytene element, which corresponds to the dot-like autosome. Banding pattern as well as the most characteristic features and prominent landmarks of each polytene chromosome arm are presented and discussed. Furthermore, twelve gene markers have been mapped on the polytene chromosomes of D. suzukii by in situ hybridization. Their distribution pattern was found quite similar to that of D. melanogaster revealing conservation of synteny although the relative position within each chromosome arm for most of the genes differed significantly between D. suzukii and D. melanogaster. The chromosome information presented here is suitable for comparative cytogenetic studies and phylogenetic exploration, while it could also facilitate the assembly of the genome sequence and support the development of genetic tools for species-specific and environment-friendly biological control applications such as the sterile insect technique.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Drosophila suzukii (Diptera: Drosophilidae), also known as the spotted wing drosophila, was first described by Matsumura in Japan (Matsumura 1931). It belongs to the suzukii subgroup and is closely related to D. melanogaster and other Drosophila species of the melanogaster species group (Yang et al. 2004, 2012). The species is considered native to Southeast Asia. However, it was reported in California and Spain in 2008 (Hauser 2011; Calabria et al. 2012) and has been expanding across America (north and south) and Europe ever since (Hauser 2011; Burrack et al. 2012; Cini et al. 2012; Deprá et al. 2014; Asplen et al. 2015), proving to be highly invasive. The rapid expansion of D. suzukii in a wide temperature and climate range has raised serious concern since it is one of the very few Drosophila species that infests healthy, marketable fruits. Females of D. suzukii possess a non-typical serrated ovipositor that allows them to pierce and lay eggs inside ripening fruits (Atallah et al. 2014). The developing larvae feed on the fruit pulp, while the wounds on the fruit skin serve as entrance to bacterial and fungal pathogens, both decreasing fruit quality and value (Walsh et al. 2011; Cini et al. 2012; Hamby et al. 2012; Ioriatti et al. 2015; Asplen et al. 2015). D. suzukii is a polyphagous pest known to infest a variety of berries (blueberries, blackberries, raspberries, and strawberries), and other soft skin fruits as well, such as cherries, apricots, peaches, pears, grapes, and kiwis (Grassi et al. 2011; Lee et al. 2011; Walsh et al. 2011; Bellamy et al. 2013; Haye et al. 2016). This insect is at present causing substantial crop losses and economical damage in USA and Europe (Bolda et al. 2010; Goodhue et al. 2011; Walsh et al. 2011; Cini et al. 2012), while the danger of expanding to new territories in the future renders it a major insect pest threat worldwide. Therefore, there is an intensive need for effective management and population control of the species. Innovative biological control methods such as the Sterile Insect Technique (SIT) could be a promising practice for the efficient management of D. suzukii, as it has been successfully implemented at a large scale for other insect pests, requiring, nevertheless, extensive knowledge on the biology of the insect (Dyck et al. 2005).
Cytogenetic analysis can provide significant information on the genetics and the genomic organization of insect species. Dipteran polytene chromosomes are extremely useful for studying chromosome structure and function, as well as temporal gene activities (Zhimulev et al. 2004 and references therein). Their characteristic species-specific banding pattern is used to assess phylogenetic relationships among closely related species (Krimbas and Powell 1992; Zhao et al. 1998; Gariou-Papalexiou et al. 2007, 2016; Drosopoulou et al. 2011a, b, 2017; Zacharopoulou et al. 2011a, b, 2017; Mavragani-Tsipidou et al. 2014; Augustinos et al. 2015) or even to distinguish among members of species complexes (Lemeunier and Ashburner 1984; Mavragani-Tsipidou et al. 1992; Coluzzi et al. 2002; Gariou-Papalexiou et al. 2007; Caceres et al. 2009; Augustinos et al. 2014). They also enable the construction of detailed physical genetic maps through in situ hybridization and precise localization of genomic sequences, revealing homologies of chromosomes or chromosome segments even among more distantly related species (Zacharopoulou et al. 1992, 2017; Drosopoulou et al. 1997, 2015, 2017; Zhao et al. 1998; Zambetaki et al. 1999; Gariou-Papalexiou et al. 2002; Holt et al. 2002; Mavragani-Tsipidou 2002; Campos et al. 2007; Sharakhova et al. 2007; Tsoumani et al. 2011; Stocker et al. 2012; Garcia et al. 2015). Furthermore, cytogenetic analysis has supported the development, characterization, and improvement of genetic sexing strains (GSSs) used in effective SIT control of important pest species (Zacharopoulou and Franz 2013; Zacharopoulou et al. 2017).
In the current study, we present the mitotic karyotype and detailed photographic polytene chromosome maps of the spotted wing drosophila, D. suzukii. We also compare the chromosome organization of the above species to D. melanogaster based on in situ hybridization data. Our data are expected to contribute to the assembly of the D. suzukii genome and to the better understanding of the species phylogenetic relationships within the melanogaster subgroup. Moreover, it could prove useful for the construction and characterization of GSSs in support to the efforts for the development and application of effective SIT for the population control of this major agricultural pest.
Materials and methods
Flies
Third instar larvae of D. suzukii used in the present study came from a laboratory colony maintained at the Joint FAO/IAEA Insect Pest Control Laboratory (IPCL), Seibersdorf, Austria. The IPCL colony was established in 2014 using pupae originated from the Agricultural Entomology Unit of the Edmund Mach Foundation in San Michele All’Adige, Trento Province, Italy. Larvae and adult flies of D. melanogaster, strain Canton-S, maintained at the Department of Genetics, Development and Molecular Biology of Aristotle University of Thessaloniki, were also used.
Mitotic chromosome preparations
Mitotic chromosome preparations were made from nerve ganglia of third instar larvae, following the air-drying technique without the use of colchicine described by Mavragani-Tsipidou et al. (2014). Brain tissue was dissected in Ringer’s solution and transferred to a well slide containing 1% sodium citrate hypotonic solution for at least 15 min. Subsequently, it was transferred to fresh methanol/acetic acid 3:1 fixative solution which was replaced with new solution every minute to ensure the complete removal of the water. After 3 min, the fixative solution was removed, and 60% acetic acid was added. The material was pipetted through a thin micropipette tip several times for dispersal and dried on a clean slide placed on a pre-warmed hot plate (40–45 °C). Chromosomes were stained with 5% Giemsa in 10 mM phosphate buffer, pH 6.8. More than 20 chromosome preparations, each from an individual larva, were analyzed and about 50 well-spread metaphases were photographed using a phase contrast microscope (LEIKA DMR) and a CCD camera (ProgResCFcool; Jenoptik Jena Optical Systems, Jena, Germany).
Polytene chromosome preparations
Polytene chromosome preparations were made from salivary glands of well-fed third instar larvae following the procedure described by Mavragani-Tsipidou et al. (2014), with some modifications. Larvae were dissected in Ringer’s solution and the glands were transferred to 45% acetic acid for 2-3 min where the adhering fat body was removed. Tissue was transferred to 3 M HCl for 1 min and to lacto-acetic acid (glacial acetic acid:water:lactic acid in 3:2:1 ratio) for about 5 min before staining in lacto-aceto-orcein for 5–7 min. Excess stain was removed by washing the material in a drop of lacto-acetic acid before squashing.
Construction of photographic chromosome maps
Chromosome slides were observed with 60 × and 100 × objectives on a phase contrast microscope (LEIKA DMR) and at least 100 well-spread nuclei or isolated chromosomes were photographed using a CCD camera (ProgResCFcool; Jenoptik Jena Optical Systems, Jena, Germany). Selected chromosomal regions, providing a clear banding pattern and demonstrating the continuity of each polytene element, were assembled using the Adobe Photoshop CS6 Extended Software, to construct the composite photographic map for each chromosomal element.
In situ hybridization procedures
Polytene chromosome preparations for in situ hybridization were made from salivary glands of late third instar larvae following the protocol described by Pardue (1986) with some modifications. Larvae were dissected in Ringer’s solution and the glands were transferred to 45% acetic acid for 2–3 min where the adhering fat body was removed. Then the material was transferred into a small volume (about 15 μl) of lacto-acetic acid on an 18X18 coverslip for about 5 min. The preparation was squashed after a slide had been laid on the coverslip and turned over. The slide was placed horizontally at − 20 °C for 24 h and the coverslip was removed by a razor blade after the preparation was dipped in liquid nitrogen. Slides were dehydrated in 95% ethanol and stored at room temperature (RT).
Six of the gene probes used for in situ hybridization represented genomic or cDNA fragments previously cloned from D. melanogaster or D. auraria, while the remaining six were generated in the present study (Table 1). Primers (Table 2) were designed on selected gene sequences of D. melanogaster and D. suzukii genomes available in FlyBase, release FB2018_04 (Thurmond et al. 2019) and SpottedWingFlyBase, v 1.0 (Chiu et al. 2013), respectively. PCR amplifications on D. melanogaster DNA were performed using BIOTAQ DNA Polymerase (BIOLINE, UK). Amplification products, after purification with Exonuclease I and Shrimp Alkaline Phosphatase (NEB, USA), were cloned using the QIAGEN PCR cloning kit (QIAGEN, Germany) and sequenced by Eurofins Genomics (Germany). All sequences were confirmed by BLASTN (https://blast.ncbi.nlm.nih.gov/Blast.cgi).
Labeling and detection was performed using the DIG-DNA labeling and Detection kit (ROCHE, Germany). Hybridization was performed at 62–65 °C as previously described (Drosopoulou and Scouras 1995). Parallel hybridization of D. melanogaster polytene chromosome preparations was performed for each probe as positive control. Four to five preparations were hybridized with each probe, and at least ten well-spread nuclei per preparation were observed at 63 × or 100 × magnification with a Nikon Eclipse 80i or a Leica DMR phase contrast microscope. Photographs were captured using a Nikon DS-5M-U1 (63 ×) or a JenoptikProgRes (100 ×) CCD camera.
Results and discussion
Mitotic chromosomes
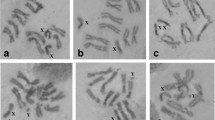
The mitotic karyotype of the D. suzukii strain analyzed consists of four pairs of chromosomes: (a) one pair of sex chromosomes, (b) two pairs of meta- or sub-metacentric autosomes and (c) one pair of dot-like autosomes (Fig. 1). The sex chromosomes are designated as the first pair of the mitotic karyotype, while the three autosomes are labeled from 2 to 4, in order of descending size. The first autosome pair (chromosome 2) has about twice the size of the second one (chromosome 3), while the third autosome (chromosome 4) is very small like a dot. The X chromosome is an acrocentric of medium size (about half the size of pair 3), and the Y is a short, rod-shaped chromosome that is heavily stained (Fig. 1).
The above-described mitotic complement is in agreement with previous descriptions of the D. suzukii mitotic karyotype (Lemeunier et al. 1986; Deng et al. 2007) and very similar to the karyotype of D. melanogaster and other species of the melanogaster species group (Lemeunier et al. 1986).
Polytene chromosomes
The analysis of the salivary gland polytene chromosomes of D. suzukii showed that the polytene complement consists of five long and one very short well-banded polytene elements (Fig. 2). The polytene chromosome arms were named X (the sex chromosome), 2L, 2R, 3L, 3R, and 4, based on the similarities of the telomeric regions and the chromosomal banding pattern to D. melanogaster and were divided into sections from 1 to 102. The detailed photographic maps of the polytene chromosomes of D. suzukii third instar larvae are shown in Fig. 3. A short description of the most prominent diagnostic landmarks for each element is given below.
X chromosome (sections 1–20, Fig. 3)
In the polytene complement of D. suzukii, the X chromosome is represented by one long polytene arm with a very characteristic torus-shaped tip and an easily recognizable proximal end. Prominent landmarks of this polytene arm are the three intense bands in section 2, the puffed structure in section 6, the dense banding pattern in section 9, the two bands in section 10 surrounded by two puffs, the three bands in section 14 followed by three puffs at sections 14–16 and the two zones at section 17.
2L chromosome arm (sections 21–40, Fig. 3)
The tip of this chromosome arm is followed by two dark bands separated by a bright interband region (section 21). Section 23 has a characteristic banding pattern. Other landmarks are the dense banding pattern in sections 29–30, the puff at section 33, the series of bands at section 36 as well as at the beginning of section 38.
2R chromosome arm (sections 41–60, Fig. 3)
The 2R chromosome arm is easily identified by its characteristic tip. Prominent landmarks of this chromosome are section 56 with poor banding pattern followed by two intense bands and the puff at section 52 also preceding two intense and clear bands in section 51. Section 47 consists of thin bands followed by a diffused area and two dark bands (the second belongs to section 46). A prominent landmark is the thin area of section 44.
3L chromosome arm (sections 61–80, Fig. 3)
Apart from its characteristic tip, the 3L polytene arm is identified by the presence of a very prominent band at the beginning of section 62, a series of three dark bands at section 63, the section 66 with two sharp bands followed by a puff and a very thick band, another thick band at section 69, the section 73 followed by the puff at section 74, the series of tree bands at 75, the puff at 77 and the fan-shaped proximal end (80).
3R chromosome arm (sections 81–100, Fig. 3)
This is the longest polytene element of the complement. It is recognized not only by its tip but also by its characteristic slightly puffed proximal end (82). The region at the borders of sections 98 and 97 is easily recognized. Other landmarks of this arm are the series of seven intense bands in section 95 and the characteristic structure of section 85. The chromosome was often found broken or stretched between sections 93 and 90.
Chromosome 4 (sections 101–102, Fig. 3)
Chromosome 4 forms a very short polytenized element tightly joined with the heterochromatic mass of the chromocenter.
Chromosome localization of molecular markers
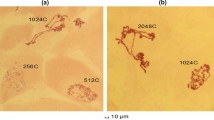
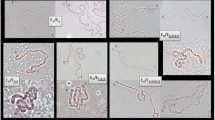
Twelve gene markers selected to represent all five long polytene elements of D. melanogaster were localized on the polytene chromosomes of D. suzukii by in situ hybridization (Table 1). Eight of the probes gave unique hybridization signals. In detail, the Sxl, w, and if genes were mapped on the X chromosome of D. suzukii at sections 7, 13 and 16, respectively (Fig. 4); the Adh and the CG17652 probes were localized at sections 22 and 29 of the 2L chromosome arm, respectively (Fig. 5a, b); Opa1 mapped at section 53 of the 2R arm (Fig. 5e) and the hsp83 and Toll-9 markers were localized on 3L chromosome arm at sections 70 and 71, respectively (Fig. 6a, b). For βtub 56D, βtub60D, aTub 84D and hsp70 probes, which represent members of the β-tubulin, α-tubulin and Hsp70 multigene families, respectively, apart from the main hybridization signals, additional secondary signals presenting lower frequency and intensity were observed both in D. suzukii and D. melanogaster. Specifically, the main hybridization signals for βtub60D and βTub56D were identified at sections 49 and 50 of the D. suzukii 2R polytene arm, respectively (Fig. 5c, d), while both probes presented weaker hybridization at each other’s main hybridization sites in several nuclei of D. suzukii (Fig. 5d) similar to D. melanogaster. The αTub84D probe gave a main hybridization signal at section 84 and a secondary signal at section 87 of the D. suzukii 3R chromosome arm (Fig. 6c). This pattern was similar to the one observed in D. melanogaster, where the main signal was identified at the locus of the αTub84D gene and a secondary one at the locus of the aTub85E gene, suggesting that the hybridization sites at 84 and 87 in D. suzukii is where the putative orthologues of the αTub84D and aTub85E genes are located, respectively (Fig. 7). Similarly, the hsp70 probe hybridized mainly at section 86 but also gave secondary signals in sections 92 and 94 of the D. suzukii 3R polytene arm (Fig. 6d). Comparing the above hybridization pattern (number, relative intensity and frequency of the hybridization signals) with the one observed in D. melanogaster, i.e. main signal at 87A and 87B at the loci of the hsp70 genes and secondary hybridization at the loci of the hsp68 and the hsc70-4 genes (sections 95D and 88E, respectively; Fig. 6e), it could be suggested that the hybridization sites in sections 86, 92 and 94 in D. suzukii indicate the location of the hsp70, hsp68 and the hsc70-4 putative orthologues in this species, respectively (Fig. 7).
In situ hybridization on polytene chromosome 2 of Drosophila suzukii.a Hybridization site of the Adh probe; b hybridization site of the CG17652 probe; c hybridization site of the βtub60D probe; d hybridization site of the βtub56D probe; e hybridization site of the Opa1 probe. Thick arrows indicate the main hybridization signals. Thin arrow on d indicates secondary hybridization
In situ hybridization on polytene chromosome 3 of Drosophila suzukii (a–d) and Drosophila melanogaster (e). a Hybridization site of the hsp83 probe; b hybridization site of the Toll-9 probe; c hybridization sites of the aTub84D probe; d, e hybridization sites of the hsp70 probe. Thick arrows indicate the main hybridization signals. Thin arrows on c–e indicate secondary hybridization
Schematic comparative representation of Drosophila suzukii (Ds) and Drosophila melanogaster (Dm) polytene chromosomes. Solid lines link the relative positions of the orthologue genes revealed by main hybridization signals. Dashed lines link the relative positions of putative orthologue genes revealed by secondary hybridization. C in grey circles indicates the centromeres
The distribution of the gene loci on the chromosomes of D. suzukii is very similar to that of D. melanogaster as diagrammatically shown in Fig. 7. The same set of genes appears to be linked on the respective chromosome arms suggesting conservation of chromosomal gene content between the two species. The above is also supported by the D. suzukii genome assembly (Chiu et al. 2013). Chromosomal arm-level synteny has been previously shown by physical mapping and genome sequencing among numerous closely or distantly related Drosophila species (Drosopoulou and Scouras 1995, 1998; Pardali et al. 1996; Drosopoulou et al. 1996, 1997, 2002; Clark et al. 2007; Bhutkar et al. 2008; Schaeffer et al. 2008; Stocker et al. 2012) proving true Muller’s hypothesis that during Drosophila evolution the six chromosomal elements A–F, maintained their structure and identity (Sturtevant and Novitski 1941). Similarly, cytogenetic and genomic studies outside the Drosophila genus were able to reveal synteny of genetic loci and correspondence of chromosome elements among species of different dipteran families (Foster et al. 1981; Zacharopoulou et al. 1992, 2017; Zhao et al. 1998; Zambetaki et al. 1999; Gariou-Papalexiou et al. 2002; Mavragani-Tsipidou 2002; Campos et al. 2007; Tsoumani et al. 2011; Drosopoulou et al. 2015, 2017; Sved et al. 2016), suggesting that the overall organization and content of chromosome elements has been conserved throughout Schizophora evolution (Sved et al. 2016).
However, within each chromosome arm the relative positions of the majority of the gene loci mapped are significantly different between D. suzukii and D. melanogaster (Fig. 7). This is not surprising since 58 out of the 160 synteny blocks identified by the D. suzukii genome assembly presented inverted direction between the two species (Chiu et al. 2013). Extensive reshuffling of genes within chromosome arms has been also revealed from comparisons among a number of Drosophila (Drosopoulou and Scouras 1995, 1998; Pardali et al. 1996; Drosopoulou et al. 1996, 1997, 2002; Clark et al. 2007; Bhutkar et al. 2008; Schaeffer et al. 2008; Stocker et al. 2012) and non drosophilid species (Foster et al. 1981; Zacharopoulou et al. 1992, 2017; Zhao et al. 1998; Zambetaki et al. 1999; Gariou-Papalexiou et al. 2002; Mavragani-Tsipidou 2002; Campos et al. 2007; Tsoumani et al. 2011; Drosopoulou et al. 2015, 2017; Sved et al. 2016). The above observations support that, unlike inter chromosomal arm rearrangements, intra-chromosomal events, such as within arm inversions, have been a common phenomena playing an important role during Diptera evolution (Ashburner et al. 1982; Ashburner 1989; Krimbas and Powell 1992; Rieseberg 2001; Schaeffer et al.2008; Stocker et al. 2012; Lee et al. 2013; Sharakhov et al. 2016; Sved et al. 2016; Zacharopoulou et al. 2017).
In summary, the first high-quality polytene chromosome maps for D. suzukii presented here could be used in comparative cytogenetic studies providing information on the phylogenetic status of the species within the melanogaster species group, while they enable the physical mapping of additional gene markers that should prove particularly useful for the assignment of scaffolds to chromosomal loci and the assembly of the genome sequence. Furthermore, linking cytogenetic with molecular knowledge could also assist the development and characterization of stable GSSs for their potential use in SIT applications against this destructive pest.
References
Ashburner M (1989) Inversions. In: Ashburner M (ed) Drosophila: a laboratory handbook. Cold Spring Harbor Laboratory Press, New York, pp 509–528
Ashburner M, Carson HL, Thompson J (1982) The genetics and biology of Drosophila. Academic Press, London
Asplen MK, Anfora G, Biondi A et al (2015) Invasion biology of spotted wing Drosophila (Drosophila suzukii): a global perspective and future priorities. J Pest Sci 88:469–494. https://doi.org/10.1007/s10340-015-0681-z
Atallah J, Teixeira L, Salazar R, Zaragoza G, Kopp A (2014) The making of a pest: the evolution of a fruit-penetrating ovipositor in Drosophila suzukii and related species. Proc Biol Sci 281:20132840. https://doi.org/10.1098/rspb.2013.2840
Augustinos AA, Drosopoulou E, Gariou-Papalexiou A, Bourtzis K, Mavragani-Tsipidou P, Zacharopoulou A (2014) The Bactrocera dorsalis species complex: comparative cytogenetic analysis in support of Sterile Insect Technique applications. BMC Genet 15(Suppl 2):S16
Augustinos AA, Drosopoulou E, Gariou-Papalexiou A, Asimakis ED, Cáceres C, Tsiamis G, Bourtzis K, Mavragani-Tsipidou P, Zacharopoulou A (2015) Cytogenetic and symbiont analysis of five members of the B. dorsalis complex (Diptera, Tephritidae): no evidence of chromosomal or symbiont-based speciation events. ZooKeys 540:273–298
Bellamy DE, Sisterson MS, Walse SS (2013) Quantifying host potentials: indexing postharvest fresh fruits for spotted wing drosophila, Drosophila suzukii. PLOS One 8(4):e61227
Bhutkar A, Schaeffer SW, Russo SM et al (2008) Chromosomal rearrangement inferred from comparisons of 12 Drosophila genomes. Genetics 179:1657–1680
Bolda M, Goodhue R, Zalom FG (2010) Spotted wing Drosophila: potential economic impact of a newly established pest. Agric Resour Econ Update 13:5–8
Burrack HJ, Smith JP, Pfeiffer DG, Koehler G, La Forest J (2012) Using volunteer-based networks to track Drosophila suzukii (Diptera: Drosophilidae) an invasive pest of fruit crops. J Integr Pest Manag 4:B1–B5
Caceres C, Segura DF, Vera MT, Wornoayporn V, Cladera JL, Teal P, Sapountzis P, Bourtzis K, Zacharopoulou A, Robinson AS (2009) Incipient speciation revealed in Anastrepha fraterculus (Diptera; Tephritidae) by studies on mating compatibility, sex pheromones, hybridization, and cytology. Biol J Linnean Soc 97:152–165. https://doi.org/10.1111/j.1095-8312.2008.01193.x
Calabria G, Maca J, Bachli G, Serra L, Pascual M (2012) First records of the potential pest species Drosophila suzukii (Diptera: Drosophilidae) in Europe. J Appl Entomol 136:139–147
Campos SRC, Rieger TT, Santos JF (2007) Homology of polytene elements between Drosophila and Zaprionus determined by in situ hybridization in Zaprionus indianus. Genet Mol Res 6:262–276
Chiu JC, Jiang X, Zhao L, Hamm CA, Cridland JM et al (2013) Genome of Drosophila suzukii, the spotted wing Drosophila. G3: Genes Genom Genet 3:2257–2271. https://doi.org/10.1534/g3.113.008185
Cini A, Ioriatti C, Anfora G (2012) A review of the invasion of Drosophila suzukii in Europe and a draft research agenda for integrated pest management. Bull Insectol 65:149–160
Clark AG, Eisen MB, Smith DR et al (2007) Evolution of genes and genomes on the Drosophila phylogeny. Nature 450:203–218
Coluzzi M, Sabatini A, della Torre A, Di Deco MA, Petrarca V (2002) A polytene chromosome analysis of the Anopheles gambiae species complex. Science 298:1415–1418
Deng Q, Zeng Q, Qian Y, Li C, Yang Y (2007) Research on the karyotype and evolution of Drosophila melanogaster species group. J Genet Genomics 34:196–213. https://doi.org/10.1016/S1673-8527(07)60021-6
Deprá M, Poppe JL, Schmitz HJ, De Toni DC, Valente VLS (2014) The first records of the invasive pest Drosophila suzukii in the South American continent. J Pest Sci 87:379–383. https://doi.org/10.1007/s10340-014-0591-5
Drosopoulou E, Scouras ZG (1995) The β-tubulin gene family evolution in the Drosophila montium subgroup of the melanogaster species group. J Mol Evol 41:293–298
Drosopoulou E, Scouras ZG (1998) The organization of the α-tubulin gene family in the Drosophila montium subgroup of the melanogaster species group. Genome 41:504–509
Drosopoulou E, Konstantopoulou I, Scouras ZG (1996) The heat shock genes in the Drosophila montium subgroup. Chromosomal localization and evolutionary implications. Chromosoma 105:104–110
Drosopoulou E, Tsiafouli M, Mavragani-Tsipidou P, Scouras ZG (1997) The glutamate dehydrogenase, E74 and putative actin gene loci in the Drosophila montium subgroup. Chromosoma 106:20–28
Drosopoulou E, Wiebauer K, Yiangou M, Mavragani-Tsipidou P, Domdey H, Scouras ZG (2002) Isolation, characterization, and localization of beta tubulin genomic clones of three Drosophila montium subgroup species. Genome 45:604–607
Drosopoulou E, Augustinos AA, Nakou I, Koeppler K, Kounatidis I, Vogt H, Papadopoulos NT, Bourtzis K, Mavragani-Tsipidou P (2011a) Genetic and cytogenetic analysis of the American cherry fruit fly, Rhagoletis cingulata (Diptera: Tephritidae). Genetica 139:1449–1464. https://doi.org/10.1007/s10709-012-9644-y
Drosopoulou E, Nestel D, Nakou I, Kounatidis I, Papadopoulos NT, Bourtzis K, Mavragani-Tsipidou P (2011b) Cytogenetic analysis of the Ethiopian fruit fly Dacus ciliatus (Diptera: Tephritidae). Genetica 139:723–732
Drosopoulou E, Nakou I, Mavragani-Tsipidou P (2015) The Bactrocera oleae genome: localization of nine genes on the polytene chromosomes of the olive fruit fly (Diptera: Tephritidae). Genome 57:573–576
Drosopoulou E, Pantelidou C, Gariou-Papalexiou A, Augustinos AA, Chartomatsidou T, Kyritsis GA, Bourtzis K, Mavragani-Tsipidou P, Zacharopoulou A (2017) The chromosomes and the mitogenome of Ceratitis fasciventris (Diptera: Tephritidae): two genetic approaches towards the Ceratitis FAR species complex resolution. Sci Rep 7:4877. https://doi.org/10.1038/s41598-017-05132-3
Dyck VA, Hendrichs J, Robinson AS (2005) Sterile insect technique: principles and practice in area-wide integrated pest management. Springer, The Netherlands
Foster G, Whitten M, Konovalov C, Arnold J, Maffi G (1981) Autosomal genetic maps of the Australian sheep blowfly, Lucilia cuprina dorsalis R.-D. (Diptera: Calliphoridae), and possible correlations with the linkage maps of Musca domestica L. and Drosophila melanogaster (Mg.). Genet Res 37:55–69
Garcia C, Delprat A, Ruiz A, Valente VLS (2015) Reassignment of Drosophila willistoni genome scaffolds to chromosome II arms. G3: Genes Genom Genet 5:2559–2566
Gariou-Papalexiou A, Gourzi P, Delprat A, Kritikou D, Rapti K, Chrysanthakopoulou B, Mintzas A, Zacharopoulou A (2002) Polytene chromosomes as tools in the genetic analysis of the Mediterranean fruit fly, Ceratitis capitata. Genetica 116:59–71
Gariou-Papalexiou A, Yannopoulos G, Robinson AS, Zacharopoulou A (2007) Polytene chromosome maps in four species of tsetse flies Glossina austeni, G. pallidipes, G. morsitans morsitans and G. m. submorsitans (Diptera: Glossinidae): a comparative analysis. Genetica 129:243–251
Gariou-Papalexiou A, Giardini MC, Augustinos AA, Drosopoulou E, Lanzavecchia SB, Cladera JL, Caceres C, Bourtzis K, Mavragani-Tsipidou P, Zacharopoulou A (2016) Cytogenetic analysis of the south American fruit fly Anastrepha fraterculus (Diptera: Tephritidae) species complex: construction of detailed photographic polytene chromosome maps of the Argentinian Af. sp.1 member. PLOS One 11(6):7192. https://doi.org/10.1371/journal.pone.0157192
Goldberg DA (1980) Isolation and partial characterization of the Drosophila alcohol dehydrogenase gene. Proc Natl Acad Sci USA 77:5794–5798. https://doi.org/10.1073/pnas.77.10.5794
Goodhue RE, Bolda M, Farnsworth D, Williams JC, Zalom FG (2011) Spotted wing Drosophila infestation of California strawberries and raspberries: economic analysis of potential revenue losses and control costs. Pest Manag Sci 67:1396–1402. https://doi.org/10.1002/ps.2259
Grassi A, Giongo L, Palmieri L (2011) Drosophila (Sophophora) suzukii (Matsumura), new pest of soft fruits in Trentino (North-Italy) and in Europe. IOBC/WPRS Bull 70:121–128
Hamby KA, Hernandez A, Boundy-Mills K, Zalom FG (2012) Associations of yeasts with spotted-wing Drosophila (Drosophila suzukii; Diptera: Drosophilidae) in cherries and raspberries. Appl Environ Microbiol 78:4869–4873
Hauser M (2011) A historic account of the invasion of Drosophila suzukii (Matsumura) (Diptera: Drosophilidae) in the continental United States, with remarks on their identification. Pest Manag Sci 67:1352–1357
Haye T, Girod P, Cuthbertson AGS, Wang XG, Daane KM, Hoelmer KA, Baroffio C, Zhang JP, Desneux N (2016) Current SWD IPM tactics and their practical implementation in fruit crops across different regions around the world. J Pest Sci 89:643–651. https://doi.org/10.1007/s10340-016-0737-8
Holt RA, Subramanian GM, Halpern A, Sutton GG, Charlab R, Nusskern DR et al (2002) The genome sequence of the malaria mosquito Anopheles gambiae. Science 298:129–149
Ioriatti C, Walton V, Dalton D, Anfora G, Grassi A, Maistri S, Mazzoni V (2015) Drosophila suzukii (Diptera: Drosophilidae) and its potential impact to wine grapes during harvest in two cool climate wine grape production regions. J Econ Entomol 108:1148–1155. https://doi.org/10.1093/jee/tov042
Kalfayan L, Wensink PC (1981) α-tubulin genes of Drosophila. Cell 24:97–106
Konstantopoulou I, Scouras ZG (1998) The heat-shock gene hsp83 of Drosophila auraria: genomic organization, nucleotide sequence, and long antiparallel coupled ORFs (LAC ORFs). J Mol Evol 46:334–343
Konstantopoulou I, Ouzounis C, Drosopoulou E, Yiangou M, Sideras P, Sander C, Scouras ZG (1995) A Drosophila hsp70 gene contains long antiparallel coupled open reading frames (LAC ORFs) conserved in homologous loci. J Mol Evol 41:414–420
Krimbas CB, Powell JR (1992) Drosophila inversion polymorphism. CRC Press, Florida
Lee JC, Bruck DJ, Dreves AJ, Ioriatti C, Vogt H, Baueld P (2011) In focus: spotted wing drosophila, Drosophila suzukii, across perspectives. Pest Manag Sci 67:1349–1351
Lee Y, Collier TC, Sanford MR, Marsden CD, Fofana A, Cornel AJ, Lanzaro GC (2013) Chromosome inversions, genomic differentiation and speciation in the African Malaria Mosquito Anopheles gambiae. PLOS One. https://doi.org/10.1371/journal.pone.0057887
Lemeunier F, Ashburner M (1984) Relationships within the melanogaster species subgroup of the genus Drosophila (Sophophora). IV. The chromosomes of two new species. Chromosoma 89(343):351
Lemeunier F, David JR, Tsacas L, Ashburner M (1986) The melanogaster species group. In: Ashburner M, Carson HL, Thompson J (eds) The genetics and biology of Drosophila, vol 3. Academic Press, London, pp 147–256
Matsumura S (1931) 6000 illustrated insects of Japan-empire (in Japanese). Tokohshoin, Tokyo
Mavragani-Tsipidou P (2002) Genetic and cytogenetic analysis of the olive fruit fly Bactrocera oleae (Diptera: Tephritidae). Genetica 116:45–57
Mavragani-Tsipidou P, Scouras ZG, Charalampidis K, Lavrentiadou S, Kastritsis CD (1992) The polytene chromosomes of Drosophila triauraria and D. quadraria, sibling species of D. auraria. Genome 35:318–326
Mavragani-Tsipidou P, Zacharopoulou A, Drosopoulou E, Augustinos AA, Bourtzis K, Marec F (2014) Tephritid Fruit Flies (Diptera). In: Sharakhov I (ed) Protocols for cytogenetic mapping of Arthropod genomes. CRC Press, Taylor and Francis Group, pp 1–62
Pardali E, Feggou E, Drosopoulou E, Konstantopoulou I, Scouras ZG, Mavragani-Tsipidou P (1996) The Afrotropical Drosophila montium subgroup: Balbiani ring 1, polytene chromosomes, and heat shock response of Drosophila vulcana. Genome 39:588–597
Pardue ML (1986) Drosophila a practical approach. In: Roberts DB (ed) In situ hybridization to DNA of chromosomes and nuclei. IRL Press, Oxford, pp 111–137
Rieseberg LH (2001) Chromosomal rearrangements and speciation. Trends Ecol Evol 16:351–358
Schaeffer SW, Bhutkar AU, McAllister BF et al (2008) Polytene chromosomal maps of 11 Drosophila species: the order of genomic scaffolds inferred from genetic and physical maps. Genetics 179:1601–1655
Sharakhov IV, Artemov GN, Sharakhova MV (2016) Chromosome evolution in malaria mosquitoes inferred from physically mapped genome assemblies. J Bioinform Comput Biol 14:1630003. https://doi.org/10.1142/S0219720016300033
Sharakhova M, Hammond MP, Lobo NF, Krzywinski J, Unger MF, Hillenmeyer ME et al (2007) Update of the Anopheles gambiae PEST genome assembly. Genome Biol 8:R5
Stocker AJ, Rusuwa BB, Blacket MJ, Frentiu FD, Sullivan M et al (2012) Physical and linkage maps for Drosophila serrata, a model species for studies of clinal adaptation and sexual selection. G3-Genes Genom Genet 2:287–297. https://doi.org/10.1534/g3.111.001354
Sturtevant AH, Novitski E (1941) The homologies of the chromosome elements in the genus Drosophila. Genetics 24:517–541
Sved JA, Chen Y, Shearman D, Frommer M, Gilchrist AS, Sherwin WB (2016) Extraordinary conservation of entire chromosomes in insects over long evolutionary periods. Evolution 70:229–234
Thurmond J, Goodman JL, Strelets VB, Attrill H, Gramates LS, Marygold SJ, Matthews BB, Millburn M, Antonazzo G, Trovisco V, Kaufman TC, Calvi BR, FlyBase Consortium (2019) FlyBase 2.0: the next generation. Nucleic Acids Res 47(D1):D759–D765
Tsoumani KT, Augustinos AA, Kakani EG, Drosopoulou E, Mavragani-Tsipidou P, Mathiopoulos KD (2011) Isolation, annotation and applications of expressed sequence tags from the olive fly, Bactrocera oleae. Mol Genet Genomics 285:33–45
Walsh DB, Bolda MP, Goodhue RE, Dreves AJ, Lee J, Bruck DJ, Walton VM, O’Neal SD, Zalom FG (2011) Drosophila suzukii (Diptera: Drosophilidae): invasive pest of ripening soft fruit expanding its geographic range and damage potential. J Integr Pest Manag 2:1–7
Yang Y, Zhang YP, Qian YH, Zeng QT (2004) Phylogenetic relationships of the Drosophila melanogaster species group deduced from spacer regions of histone gene H2A. Mol Phylogenet Evol 30:336–343
Yang Y, Hou ZH, Qian YH, Kang H, Zeng Q-T (2012) Increasing the data size to accurately reconstruct the phylogenetic relationships between nine subgroups of the Drosophila melanogaster species group (Drosophilidae, Diptera). Mol Phylogenet Evol 62:214–223
Zacharopoulou A, Franz G (2013) Genetic and Cytogenetic Characterization of Genetic Sexing Strains of Bactrocera dorsalis and Bactrocera cucurbitae (Diptera: Tephritidae). J Econ Entomol 106:995–1003
Zacharopoulou A, Frisardi M, Savakis C, Robinson AS, Tolias P, Konsolaki M, Komitopoulou K, Kafatos FC (1992) The genome of the Mediterranean fruit fly Ceratitis capitata: localization of molecular markers by in situ hybridization to salivary gland polytene chromosomes. Chromosoma 101:448–455
Zacharopoulou A, Augustinos AA, Sayed WAA, Robinson AS, Franz G (2011a) Mitotic and polytene chromosomes analysis of the oriental fruit fly, Bactrocera dorsalis (Hendel) (Diptera: Tephritidae). Genetica 139:79–90
Zacharopoulou A, Sayed WAA, Augustinos AA, Yesmin F, Robinson AS, Franz G (2011b) Analysis of mitotic and polytene chromosomes and photographic polytene chromosome maps in Bactrocera cucurbitae (Diptera: Tephritidae). Ann Entomol Soc Am 104:306–318
Zacharopoulou A, Augustinos AA, Drosopoulou E, Tsoumani K, Gariou-Papalexiou A, Franz G, Mathiopoulos KD, Bourtzis K, Mavragani-Tsipidou P (2017) A review of more than 30 years of cytogenetic studies of Tephritidae in support of Sterile Insect Technique and global trade. Entomol Exp Appl 164:204–225. https://doi.org/10.1111/eea.12616
Zambetaki A, Zacharopoulou A, Scouras ZG, Mavragani-Tsipidou P (1999) The genome of the olive fruit fly Bactrocera oleae: localization of molecular markers by in situ hybridization to the salivary gland polytene chromosomes. Genome 42:744–751. https://doi.org/10.1139/g99-017
Zhao JT, Frommer M, Sved JA, Zacharopoulou A (1998) Mitotic and polytene chromosome analyses in the Queensland fruit fly, Bactrocera tryoni (Diptera: Tephritidae). Genome 41:510–526
Zhimulev IF, Belayaeva ES, Semeshin VF et al (2004) Polytene chromosomes: 70 years of genetic research. Internat Rev Cytol 241:203–275
Acknowledgements
We would like to thank Katerina Nikolouli for providing the biological material used in the present study and Prof. Penelope Mavragani-Tsipidou for fruitful discussions on the results of the present work. The present study has been funded by the Joint FAO/IAEA Division of Nuclear Techniques in Food and Agriculture through CRP and SSA projects.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Stefan Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Drosopoulou, E., Gariou-Papalexiou, A., Karamoustou, E. et al. The chromosomes of Drosophila suzukii (Diptera: Drosophilidae): detailed photographic polytene chromosomal maps and in situ hybridization data. Mol Genet Genomics 294, 1535–1546 (2019). https://doi.org/10.1007/s00438-019-01595-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-019-01595-3