Abstract
Velvet antler displays the fastest and most robust tissue proliferation in the animal world, it is a model for a complete organ development/regeneration, and alternative medicine, tonic made from velvet antler, was beneficial for human. The weight of velvet antler had high biomedical and economic value, but the related regulation mechanisms controlling velvet antler weight remain unclear. In this study, extremely heavy and light velvet antler groups were selected from a sika deer population of 100 individuals with extreme velvet antler weight. A combination of full-length transcriptome sequencing and microRNA sequencing to the proliferation zone in the tip of velvet antler was applied. A total of 55306 transcripts and 1082 microRNAs were identified. Some highly expressed genes (COL1A1, COL1A2, COL3A1, FN1, and ATP6) and microRNAs (miR-21, let-7i, and miR-27b) were highly correlated with the physiological and growth characteristics of velvet antlers. Among the 334 differentially expressed genes, we found that most of the genes were located in the developmental process, especially animal organ development process. It is exciting to see that more blood vessels were found in the growing tip of heavy velvet antler through histological observation, and GO term of blood vessel development was also significant different between two groups. The combination analysis with mRNA and microRNA data in velvet antler showed a specific regulation network involved in the development of bone, mesenchyme, cartilage, and blood vessel, and helped us clearly find out the candidate 14 genes and 6 microRNAs, which could be used for selecting significant DNA markers of velvet antler weight.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Velvet antler, a special animal organ endemic to deer, displays the fastest and most robust tissue proliferation in the animal world, and it is because deer need bigger antlers to compete for spouses in shorter breeding season. After that, the deer shed their useless antlers, the annual regeneration of the antler is a model for a complete organ development/regeneration with pointing toward medical research/applications. Moreover, velvet antlers had been found to have a strong health care function, including inflammation resistance, oxidation resistance, immune enhancement, osteoporosis prevention, and fecundity enhancement (Shin et al. 1999; Huang et al. 2010; Zha et al. 2012, 2013; Wu et al. 2013; Sui et al. 2014; Tseng et al. 2014), and alternative medicine, tonic made from velvet antler, was developed since ancient time in the Far East. Due to its high biological and medical value, sika deer are commonly raised in captivity in eastern Asia. In the domestic population, velvet antler presents an increasingly heavy phenotype as a result of artificial selection. However, the molecular mechanism regulating the weight of velvet antler has not been well explained. The growth of velvet antler was closely related to genetic factors. It was reported that, despite substantial age- and environment-related variation, deer antler size was heritable (Kruuk et al. 2002). In addition, under the same management conditions, the weight of velvet antler by artificial breeding was increased by 30–60% than uncultivated deer (Li 2003). There was a strong correlation between multiple measurements of velvet antler weight, the repeatability was 0.53, 0.49, and 0.36 in Xifeng, Shuangyang, and Changbai Mountain sika deer populations, and genetic factors played an important role in velvet antler weight variation, the ration could reach as high as 0.67, 0.63, and 0.64 among the above populations, respectively (Li 1997). These results provide a reliable theoretical and practical basis for the genetic explanation of the weight trait of velvet antler.
Velvet antler development is a highly complicated and genetically programmed process, including the growth of bone, skin, blood vessels, and nerves. The proliferation and differentiation of cells in antler growth tip are regulated by some intracellular and extracellular factors and signal pathways. Which gene plays an important role in the growth of velvet antler? It is not clear whether a gene plays a leading role or a combination of multiple genes. The initial research found that multiple genes are expressed in the epidermis, mesenchyme, pre-cartilage, and cartilage regions of velvet antler (Francis and Suttie 1998). And then, many antler growth-related factors were obtained by different methods, and most of these factors are associated with animal organ development, especially bone development (Gao et al. 2010; Steger et al. 2010; Hao et al. 2011; Yao et al. 2012b). However, the weight-associated genes are still not determined, and the underlying regulation mechanism remains unknown.
In this study, based on high-throughput sequencing of the velvet antler, we investigated the differentially expressed genes (DEGs) and microRNAs between heavy and light velvet antler. The analyses revealed underlying factors associated with velvet antler weight traits by integrating enrichment analysis, cluster analysis, and regulation network analysis. These results will provide new insights into the gene-regulation network of velvet antler development.
Materials and methods
Ethics statement
All procedures concerning animals were organized to accord with the guidelines of care and use of experimental animals established by the Ministry of Agriculture of China, and all protocols were approved by the Institutional Animal Care and Use Committee of Institute of Special Economic Animal and Plant Sciences, Chinese Academy of Agricultural Sciences, Changchun, China.
Animals
Animals used in this study were from a sika deer population of 100 individuals with extreme variation of velvet antler weight (Hu et al. 2017). The deer were raised at the commercial farm (Changchun City, China) under same conditions and were fed the same diet. 3 individuals with extremely heavy velvet antler and 3 individuals with extremely light velvet antler were selected from the opposite tails of the 100 deer population for comparative transcriptome analysis. Another 6 individuals with extreme velvet antler weight were chosen for verifying the accuracy of gene expression using quantitative real-time PCR.
Measurement of velvet antler weight
Combining the data of growth rate, shedding time, and age, we found that the fastest growing period of velvet antler ended at 75 days after shedding, indicating that this stage was the key stage in velvet antler development. Thus, for each individual, anesthesia was used, and velvet antlers were harvested for weighing and sample collection at 75 days of growth.
Histological observation of growing tip of velvet antler
To fully investigate gene expression of heavy and light velvet antler, we have a morphological and histological observation of the growing tip of the velvet antler. One part was selected and cut away from the tip by micro-dissection: the proliferation zone (PZ). Histological section of this part was examined to evaluate the difference between the two groups.
Sample collection, RNA extraction, and sequencing
After anesthesia, a total of 6 samples of 2 groups (Table 1) were collected, each group with three independent repetitions. Three experiments were conducted (Table 1). First, total RNA of six samples were extracted using TRIzol (Life Technologies Corp.), respectively, sequencing libraries of each RNA were constructed by NEBNext® Ultra™ RNA Library Prep Kit for Illumina® (NEB, USA), and then, they were sequenced on Hiseq X Ten instrument. Second, six RNAs were pooled to provide total velvet antler RNA, sequencing library was constructed by the Clontech SMARTer PCR cDNA Synthesis Kit and the BluePippin Size Selection System, and then, they were sequenced on Pacific Biosciences RS sequencing instrument. Third, 6 RNAs of 2 groups were equivalently mixed to form 2 RNA pools, sequencing libraries of 2 RNA pools were constructed by NEB Next® Multiplex Small RNA Library Prep Set for Illumina® (NEB, USA.), and then, they were sequenced on Hiseq 2500 instrument. All the sequencing works were carried out at Novogene Co., LTD (Tianjin, China). The raw data were processed using the standard methods. Full-length transcriptome and microRNA sequences were obtained and annotated.
Differential expression analysis
Full-length transcripts obtained from Pacific Biosciences RS sequencing instrument were used as reference sequence, Hiseq sequencing reads of each sample were mapped to reference by Bowtie, and statistical analysis of readcount was conducted on the mapping results by RSEM. Gene differential expression between groups was analyzed by the DESeq software using readcount data. To decrease the false discovery rate, the P values were adjusted using the Benjamini and Hochberg’s approach, and the screening threshold for differential expression genes (DEGs) was 0.05. Clustering software package pheatmap in R was used for DEGs clustering.
The microRNA expression was analyzed by TPM (transcript per million) using the normalized formula: Normalized expression = mapped readcount/Total reads × 1,000,000. Differential expression analysis between groups was carried out by the DEGseq software. Adjusted P value < 0.05 and |log2(foldchange)| > 1 were considered as the screening threshold for differential expression microRNAs.
Target genes prediction
The potential target genes of differentially expressed microRNAs were predicted by miRanda v3.3a (Score ≥ 200, Energy E ≤ − 20) and RNAhybird 2.1.2 (Energy E ≤ − 20, P ≤ 0.05), respectively. The intersections of target genes predicted by two methods were considered as candidate target genes of microRNAs.
Quantitative real-time PCR verification
A series of DEGs and microRNAs were selected for quantitative real-time PCR on a Roche 480 Real-Time PCR System using LightCycler 480 SYBR Green I Master (Roche Pharmaceutical Ltd), and there were three replicates for each group. Primers for DEGs were designed with NCBI primer-BLAST, and primers for microRNAs were designed by stem loop methods (Table S10). Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) and U6 were used as internal reference, respectively. The results were analyzed by the 2−∆∆CT method, and data were showed as means ± standard error.
GO and KEGG enrichment analysis
Gene Ontology (GO) enrichment analysis and hierarchical network diagram of GO terms were conducted by BiNGO plug-in of the Cytoscape software v3.3.0 to understand DEGs biological functions. Hypergeometric test was used for statistics, and it was corrected by Benjamini and Hochberg False Discovery Rate (FDR), the significance level of GO terms was defined as corrected P values ≤ 0.01. The DAVID 6.8 functional annotation program was used to execute Kyoto Encyclopaedia of Genes and Genomes (KEGG) pathway analysis with the following settings: gene count 5, ease 0.05.
Interaction network and expression pattern analysis
Interaction network of microRNAs and target genes was constructed by Cytoscape v3.3.0. Prime component analysis (PCA) and canonical variates analysis (CVA) were performed using R package, first, gene expression was normalized, PCA was carried out using prcomp (R-3.5.1), and PCA diagram was drew using ggbiplot package. CVA was performed using CVA function of Morpho package (R-3.5.1), and no special functions were used.
Results
Variation between two extreme groups
Velvet antler weight was significantly different in 100 sika deer (Fig. 1a, b, Table S1), and 6 sika deer were selected and divided into 2 groups based on velvet antler weight (H, 5.92 ± 0.15; L, 2.56 ± 0.04, P < 0.01). Non-metric multidimensional scaling (NMDS) plot of individual transcriptomes was performed to identify similarities among samples (Fig. 1c). Samples from the same group had similar NMDS plots, and samples between groups were different in the third dimension.
Phenotypic characterization of two extreme groups. a Light and heavy velvet antler after 75 day of growth; b two extreme groups of velvet antler weight, H group, heavy velvet antler group, L group, light velvet antler group; c non-metric multidimensional scaling (NMDS) plot based on individual transcriptomes, the relationship of samples in heavy (H) and light (L) velvet antler group is similar (orange circles). (Color figure online)
Histological observation of growing tip of velvet antler
In the central portion of a sagittally cut antler tip, proliferation zone (pre-cartilage layer, PZ) was readily identified on the cut surface (Fig. 2a, b), velvet antler growth is achieved by rapid proliferation and differentiation of cells in this region, so PZ is the growth center of velvet antler. From the histological section, we observed that, at proliferation zone (PZ), the cell morphology is uniform, the volume is small, and the cell density is high. In heavy velvet antler group, there were many small and irregular blood vessels of in this region, the wall of the vessel is thicker and consists of multi-layer cells (Fig. 2c), while at light velvet antler group, the number of blood vessels decreased, and the blood vessels are small and even difficult to be found (Fig. 2d). This finding suggested that there are differences in histological structure between heavy and light velvet antler groups in the growing tip of velvet antler.
Anatomical and ultrastructure of the growing tip of velvet antler. a Anatomical structure of heavy velvet antler; b anatomical structure of light velvet antler, arrows refer to proliferation zone (pre-cartilage layer, PZ); c histological structure of heavy velvet antler; and d histological structure of light velvet antler, arrows refer to small and irregular blood vessels
Full-length transcriptome and microRNA sequencing
Sequencing of six mRNA samples from two groups (heavy velvet antler and light velvet antler) produced 344,100,992 raw reads (Table S2). Full-length transcriptome sequencing of six pooled RNA samples produced 1,202,336 raw reads (15.2 billion bases), after filtering, 789 012 subreads representing 14.2 billion bases were obtained (Table S2), and 55 306 transcripts were produced, with an N50 value of 2397 bases (Table S2). The obtained transcripts are used as unigenes and annotated for subsequent analysis.
Sequencing of two small RNA libraries of heavy velvet antler (H library, n = 3) and light velvet antler (L library, n = 3) produced 15,895,383 and 21,487,445 raw reads, representing 380,719 and 599,238 unique reads, respectively. After aligning with reference mRNA, 8,905,044 and 11,289,383 reads were mapped on mRNA (Table S2). The sequence length distributions of H and L library was mostly the same, and 22 nt was the main small RNA category (Figure S1A). All raw sequence reads were deposited in NCBI Sequence Read Archive (SRA, http://www.ncbi.nlm.nih.gov/Traces/sra) with SRA accession: SRP145255.
Gene annotation and expression analysis
A total of 55,306 transcripts were annotated using 7 databases (Nr, Nt, Pfam, KOG/COG, Swiss-prot, KEGG and GO). Using the results of Nr database, the species distribution map can be calculated, 17% and 11% genes had high similarity with sheep and cattle (Figure S1 B), respectively. The expression level of each gene was analyzed, and the highest expression genes in the proliferation zone of velvet antler were COL1A1, COL1A2, COL3A1, FN1, and ATP6 (Table S3). It confirmed the high activity of mineralization genes in the velvet antler (Steger et al. 2010).
Known and novel microRNAs’ identification
To identify the microRNAs in sika deer velvet antler, all mammalian microRNAs in miRBase20.0 were named background database. After aligning with miRBase, 1,440,368 and 1,762,386 reads were identified in the H and L library (Table S2); among them, 855 and 868 were detected as mature microRNAs, respectively, and 743 microRNAs were co-expressed. The species distribution map showed that 10% microRNAs had high similarity with cattle (Figure S1 C). Among these known microRNAs, 7 microRNAs (hsa-miR-21-5p, sha-miR-21, mmu-let-7i-5p, gga-let-7i, ssc-let-7i, efu-miR-27b, and mmu-miR-27b-3p) were highly expressed with more than 100,000 reads in both libraries (Table S4). A total of 88 and 90 novel microRNAs were identified in the H and L library, respectively (Table S5). In summary, a total of 1082 microRNAs were identified.
Transcriptome difference between heavy and light velvet antler
The read count for each sample was standardized by DESeq then used for differential analysis, using a recognized threshold of adjusted P value < 0.05, and a total of 334 DEGs were identified between heavy and light velvet antler (Table S6). As shown in Fig. 3a, DEGs in the groups had similar expression patterns, while they were different between groups. Of these DEGs, 233 genes were up-regulated in heavy velvet antler group, 101 genes were down-regulated (Fig. 3b), and the number of up-regulated DEGs was more than the number of down-regulated DEGs, indicating that these genes played a crucial role in promoting the development of velvet antler.
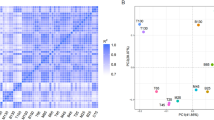
Differential expression analyses of genes and microRNAs in two extreme groups. a Cluster analyses of differential expression genes, columns and rows indicate individuals and genes, respectively. Color scale represents FPKM after standard normalization. b Number of up- and down-regulated DEGs in heavy velvet antler group. DEGs differentially expressed genes. c Read counts of the 18 differentially expressed microRNAs between heavy velvet antler (H) and light velvet antler (L) libraries. Black columns represent read count of each differentially expressed microRNA in the L library, and white columns represent read count of each differentially expressed microRNA in the H library. The Y-axis represents abundance (read count) of each microRNA in the H and L libraries
All identified microRNAs were normalized using TPM (transcript per million) based on read count. A total of 18 differentially expressed microRNAs were identified using a recognized threshold of q value < 0.05 and |log2 (fold change)| ≥ 1 (Table S7). Compared to light velvet antler, 16 microRNAs were down-regulated in heavy velvet antler (Fig. 3c), and they belong to 7 microRNA families, namely, miR-27, miR-29, miR-203, miR-205, miR-365, miR-378, and let-7.
The potential target genes of differentially expressed microRNAs were predicted based on mature sequence. In total, 3634 target genes were obtained by merging the data obtained from miRanda and RNAhybrid (Table S8). Among these differentially expressed microRNAs, efu-miR-27a has the most target genes (1697 genes). A total of 87 up-regulated target genes were obtained by merging the data from target genes of down-regulated microRNAs and up-regulated DEGs (Fig. 4).
Consistency analyses of microRNA target genes and differentially expressed genes. Venn diagram of microRNA target genes and differentially expressed genes. Target genes, down-regulated microRNA target genes; Up-regulated DEGs, up-regulated differentially expressed genes; AOD genes, differentially expressed genes involved in animal organ development
Quantitative real-time PCR verification
In a biologically independent experiment, full-length transcriptome and microRNA sequencing data were confirmed using quantitative real-time PCR analysis. A total of 12 up-regulated differentially expressed genes and 4 down-regulated microRNAs revealed by high-throughput sequencing were selected to verify the sequencing results. The quantitative real-time PCR results indicated that all the 12 genes were up-regulated in heavy velvet antler group, and 4 microRNAs were down-regulated in heavy velvet antler group (Figure S3), and there was a good correlation between quantitative real-time PCR results and high-throughput sequencing data.
Functional analysis
Gene Ontology (GO) enrichment analysis showed that the 334 differentially expressed genes (with 219 non-redundant gene symbols) were distributed into 68 Go terms based on biological process (Table S9). Most of the genes were located in the developmental process, cellular component organization, or biogenesis, and there were 72 and 86 genes involved, respectively. These relationships could also be reflected from directed acyclic graph of GO terms (Fig. 5), and it could be found that there were significant differences between heavy and light velvet antler groups in every GO term levels from developmental process to endochondral ossification, indicating that these processes were crucial in regulating the velvet antler weight. Among them, animal organ development involving 48 genes was the most relevant GO term, and 34 genes were up-regulated in this term (Fig. 4). At the next level of GO term, it contained cartilage development, bone development, and mesenchyme development, which were closely related to the development of velvet antler. Besides, GO term of blood vessel development was also significant different between two groups, and it was consistent with the histological observation results (Fig. 2c, d).
Directed acyclic graph of GO terms of differentially expressed genes. Most of the genes were located in the developmental process, cellular component organization, or biogenesis, and there were significant differences between heavy and light velvet antler groups in every GO term levels from developmental process to endochondral ossification. Among them, animal organ development involving 48 genes was the most relevant GO term (red bold italics in the figure); at the next level of GO term, it contained cartilage development, bone development, and mesenchyme development (bold in the figure), which were closely related to the development of velvet antler. (Color figure online)
The KEGG enrichment analysis for the differentially expressed genes is shown in Figure S2, the DEGs were grouped into 12 signal pathways, among them, PI3K-Akt-signaling pathway was most relevant to velvet antler development (Li et al. 2012), of the genes in this pathway, 12 genes were up-regulated in heavy velvet antler, and 3 genes were down-regulated (Table S6).
Interaction network of differentially expressed genes and microRNAs
An interaction network with 37 nodes and 65 edges was constructed (Fig. 6), using 26 up-regulated target genes (Fig. 4), ACTN1, BAG6, COL1A2, COL11A1, COL11A2, CTNNB1, DOCK1, EXOC4, GLG1, LOX, LOXL2, LOXL3, MMP14, MYH9, PHOSPHO1, PLCD1, PTN, PYGO2, RHOA, RTCB, SMARCA4, SNAI2, SPARC, TBC1D20, TNC, and WLS, directly annotated in animal organ development, and 11 down-regulated microRNAs, bta-miR-378b, cgr-miR-205, efu-miR-27a, ggo-miR-29a, hsa-miR-27a-5p, hsa-miR-29a-3p, mmu-miR-205-5p, oan-miR-205-5p, oar-miR-29a, rno-miR-203b-5p, and tgu-miR-365-1-5p. As shown in the network, there was a coordinated regulation between these microRNAs. The core microRNAs and genes were identified, with 6 microRNAs (hsa-miR-27a-5p, efu-miR-27a, tgu-miR-365-1-5p, mmu-miR-205-5p, oan-miR-205-5p, and cgr-miR-205) exhibiting important regulatory functions in the network and 14 genes (PHOSPHO1, MMP14, SPARC, GLG1, LOXL3, EXOC4, CTNNB1, LOXL2, COL1A2, LOX, MYH9, PLCD1, COL11A1, and COL11A2) regulated by at least 2 microRNAs. Among them, PHOSPHO1, MMP14, SPARC, GLG1, and RHOA were involved in bone development, LOXL3, EXOC4, SNAI2, CTNNB1, and LOXL2 were involved in mesenchyme development, COL1A2, LOX, MYH9, CTNNB1, PLCD1, andLOXL2 were involved in blood vessel development, and COL11A1, COL11A2, and SNAI2 were involved in cartilage development, respectively (Table S9). It was suspected that in heavy velvet antler group, the down-regulated microRNAs increased the expression of their target genes, and these genes were involved in the development of bone, mesenchyme, cartilage, and blood vessel, and then promoted the rapid growth of velvet antler, so the velvet antler weight was heavier than the light group.
Gene-regulation network of velvet antler in sika deer with extremely different velvet antler weight. A network with 37 nodes and 65 edges was constructed, consisting by 26 up-regulated target genes directly annotated in animal organ development, and 11 down-regulated microRNAs. There was a coordinated regulation between these microRNAs. The core microRNAs and genes were identified, with 6 microRNAs exhibiting important regulatory functions in the network and 14 genes regulated by at least 2 microRNAs
Expression pattern of differentially expressed genes in heavy and light velvet antler groups
We evaluated the expression patterns of 26 up-regulated target genes (Fig. 4) by PCA and CVA statistical analyses using R package. As shown in Fig. 7c, CVA analysis discriminated unambiguously between heavy and light velvet antler groups. As indicated by PCA ordination, most of the gene expression was strongly correlated, such as the correlation between expressions of PHOSPHO1 and SPARC, CTNNB1, and GLG1, COL11A1, and SNAI2 (Fig. 7a). It was also seen that no L was found within the area of the H triangle (Fig. 7b), indicating that the expression pattern for heavy and light velvet antler groups was obviously different. It could add significant value and credit for the sequencing result.
Expression patterns of 26 up-regulated target genes by PCA and CVA statistical analyses. a PCA biplot shows simultaneous ordination of individuals and genes, the individuals appear as points, and the genes are shown as vectors. The strength of influence of one gene on the others and the correlations are expressed by the vector length and the angle between them; b PCA ordination for individuals only, much smaller area of the triangle around the L group indicate that the gene expression pattern is much less varying for L group than for H group, none of one group’s expression pattern overlaps the other’s, indicating that the two group have separated expression pattern; c CVA statistical analyses, gene expressions are down-regulated in L individuals (white bars) as compared to H individuals (black bars). The scores for the gene expression patterns of L and H group separate widely. L, light velvet antler group; H, heavy velvet antler group
Discussion
Velvet antler weight is crucial to production performance in domestic sika deer (Dale and McCullough 2009), and much of the interest has focused on the association analysis of single-nucleotide polymorphisms, RAPD markers, and microsatellite polymorphisms with velvet antler weight, which provide the characteristic molecular marker to the heavy velvet antler (Hu et al. 2017). However, due to difficulty in measuring, the lack of standards and high economic costs, the common factors related to the weight of velvet antler were less. More importantly, complex traits cannot be completely controlled by a single gene, and it requires a series of genes (Shen et al. 2012). If we want to pinpoint the related genes of complex traits, we must use high-throughput omics data. Transcriptome sequencing technology provides a powerful tool for finding genes related to economic traits, and it has located many genes related to weight trait (Zhang et al. 2013, 2014; Vlad et al. 2014; Yuan et al. 2017). For this purpose, a number of transcriptome studies based on growing tip velvet antler had been reported (Yao et al. 2012a, b; Zhao et al. 2013; Jia et al. 2016); however, the previous studies were limited by the length of the sequencing data, and lack of comparison of different weight traits. In this study, by leveraging a full-length transcript profile and microRNA sequencing, a much more complete transcriptome of the velvet antler was generated. We investigated candidate genes and interaction network that are associated with velvet antler weight in sika deer. The combination analysis with mRNA and microRNA data showed a full regulation network in velvet antler, and increased the credibility of the results.
Some highly expressed genes found in the proliferation zone of velvet antlers were highly correlated with the physiological and growth characteristics of velvet antlers. Collagen is abundant in velvet antler. Collagens I, II A, II B, and X are expressed in all layers of velvet antlers (Price et al. 1996), and expression levels of col1A1, col1A2, and col3A1 in the ossified part of the velvet antler exceeded the skeletal levels 10-30-fold or more (Steger et al. 2010). FN1 participates in the regulation of type I collagen deposition by osteoblasts. ATP6 was related to ATP synthesis and metabolism, it ensured the requirement for high metabolic level of velvet antler during rapid growth period, and it was consisted with the report that the elevated expression of ATP6 in the antler cartilage tissues over the fetal growth plate (Steger et al. 2010). Some highly expressed microRNAs were also detected, such as miR-21 (Zheng et al. 2011), let-7i (Esquela-Kerscher et al. 2008), and miR-27b (Wang and Xu 2010), and it was these microRNAs with high expression that promote the rapid division and proliferation of mesenchyme cells without carcinogenesis.
Velvet antler, as a special animal organ endemic to deer, its growth process is similar to organ development, including the growth of bone, skin, blood vessels, and nerves. Among the DEGs, we found that most of the genes were located in the developmental process, especially animal organ development process, that were directly conformed to the growth characteristics of velvet antler. In the process of deer evolution, to use antlers as a weapon for fighting spouses in the breeding time, deer must finish rapidly velvet antler growth in a limited time (Goss 1983). The unique structure of cartilage in velvet antler meets this demand, and there are a large number of vascular network in the cartilage (Berg et al. 2007; Li et al. 2009) that ensured the requirement for high metabolic level of velvet antler during rapid growth period. In velvet antler tip, mesenchymal cells can differentiate into vascular endothelial cells and form vascular system in hypoxic environment (Clark et al. 2006). The faster the velvet antler growth, the more the blood vessels are formed. It is exciting to see that more blood vessels were found in the growing tip of heavy velvet antler through our study, and GO term of blood vessel development was also significant different between two groups, the results were consistent with histological level and gene expression level. It could be suspected that genes involved in animal organ development play a crucial role in velvet antler weight regulation.
In the previous transcriptome studies, it was found that canonical Wnt pathway played an important regulatory role in the development of velvet antler (Liu 2013). In this study, we found that an important gene in Wnt pathway, CTNNB1, which encodes beta-catenin, key downstream component of the canonical Wnt-signaling pathway, was highly expressed in the heavy velvet antler group compared to the light velvet antler group. It was reported that the concentration of the beta-catenin signal in the cytoplasm determines the activation and closure of the classical Wnt-signaling pathway. The stable expression of beta-catenin in differentiated osteoblasts can increase bone mass and regulate bone formation (Mount et al. 2006). In addition, CTNNB1 correlating with LRP4 played an important role in bone development, especially in age-related osteoporosis (Borsy et al. 2009). Therefore, we speculate that CTNNB1 is one of the key genes regulating the weight of velvet antler. Besides, we also found that 15 DEGs were identified to be associated with the PI3K-Akt-signaling pathway, PI3K-AKT-signaling pathway can promote bone growth and endochondral ossification (Rokutanda et al. 2009; Wu et al. 2017, 2018), and PI3K-AKT pathway is commonly existed in the pedicle periosteum of the antler (Li et al. 2012). Therefore, DEGs associated with the PI3K-Akt-signaling pathway are important candidates for determining the differences in velvet antler weight in sika deer. Genes RunX2, Osterix, and Osteocalcin play pivotal roles in the developing velvet antlers; however, they were not differentially expressed genes in heavy and light velvet antler groups, and their role in controlling the weight of velvet antler needs further verification.
The growth of velvet antler is a complex process of biological metabolism, which is affected by a series of genes and microRNAs. The interaction network shows a detailed picture of the gene-regulation network involved in velvet antler development, and helps us clearly find out the candidate 14 genes and 6 microRNAs related to velvet antler weight. Thus, the integrated analysis of full-length transcriptome and microRNA sequencing helped us to generate precise and informative data for a comprehensive understanding of biological mechanism on the regulation of velvet antler development. Furthermore, the genes and microRNAs identified in this study could be used for screening significant DNA markers associated with velvet antler weight in sika deer. The result would facilitate further studies, such as complex, polygenic, quantitative phenotypes, and the genetic background of QTL-s in deer.
Data availability
All raw sequence reads were deposited in NCBI Sequence Read Archive (SRA, http://www.ncbi.nlm.nih.gov/Traces/sra) with SRA accession: SRP145255.
References
Berg DK, Li C, Asher G, Wells DN, Oback B (2007) Red deer cloned from antler stem cells and their differentiated progeny. Biol Reprod 77:384–394
Borsy A, Podani J, Steger V, Balla B, Horvath A, Kosa JP, Gyurjan I, Molnar A, Szabolcsi Z, Szabo L, Jako E, Zomborszky Z, Nagy J, Semsey S, Vellai T, Lakatos P, Orosz L (2009) Identifying novel genes involved in both deer physiological and human pathological osteoporosis. Mol Genet Genomics 281:301–313
Clark DE, Li CY, Wang WY, Martin SK, Suttie JM (2006) Vascular localization and proliferation in the growing tip of the deer antler. Anat Rec Part A 288A:973–981
Dale R. McCullough ST, Koichi K (2009) Sika Deer: biology and management of native and introduced populations [M]. Springer, Japan
Esquela-Kerscher A, Trang P, Wiggins JF, Patrawala L, Cheng A, Ford L, Weidhaas JB, Brown D, Bader AG, Slack FJ (2008) The let-7 microRNA reduces tumor growth in mouse models of lung cancer. Cell Cycle 7:759–764
Francis SM, Suttie JM (1998) Detection of growth factors and proto-oncogene mRNA in the growing tip of red deer (Cervus elaphus) antler using reverse-transcriptase polymerase chain reaction (RT-PCR). J Exp Zool 281:36–42
Gao LA, Tao DY, Shan YC, Liang Z, Zhang LH, Huo YS, Zhang YK (2010) HPLC-MS/MS shotgun proteomic research of deer antlers with multiparallel protein extraction methods. J Chromatogr B: Anal Technol Biomed Life Sci 878:3370–3374
Goss RJ (1983) Deer antlers: regeneration, function and evolution [M]. Academic Press, New York
Hao L, Li HP, Yan L (2011) [Analysis of expressed sequence tags (ESTs) in sika deer (Cervus nippon hortulorum) velvet tip tissue]. Yi Chuan 33:371–377
Hu PF, Xu JP, Ai C, Shao XJ, Wang HL, Dong YM, Cui XZ, Fuhe Y, Xiumei X (2017) Screening weight related genes of velvet antlers by whole genome re-sequencing. Yi Chuan 39:1090–1101
Huang FJ, Ji JX, Qian J, Yang KY, Zhang Y, Xin YQ, Jing ZG, Wang MH, Chen TY, Wu WT (2010) Isolation, characterization and biological activity of a novel peptide from sika (Cervus nippon Temminck) antler. Pharm Biotechnol 17:151–156
Jia BY, Ba HX, Wang GW, Yang Y, Cui XZ, Peng YH, Zheng JJ, Xing XM, Yang FH (2016) Transcriptome analysis of sika deer in China. Mol Genet Genomics 291:1941–1953
Kruuk LEB, Slate J, Pemberton JM, Brotherstone S, Guinness F, Clutton-Brock T (2002) Antler size in red deer: Heritability and selection but no evolution. Evolution 56:1683–1695
Li H (2003) Study on the performance of the velvet antler of Sika Deer. Zhongguo Xumu Zazhi 39:31–32
Li HPZX, Bing GL, Li S, Zhao M, Chen XJ (1997) Specific character analysis on artificial breed and strain of deer in China. Yichuan 19:76–78
Li CY, Yang FH, Sheppard A (2009) Adult stem cells and mammalian epimorphic regeneration-insights from studying annual renewal of deer antlers. Curr Stem Cell Res Ther 4:237–251
Li C, Harper A, Puddick J, Wang W, McMahon C (2012) Proteomes and signalling pathways of antler stem cells. PLoS One 7:e30026. https://doi.org/10.1371/journal.pone.0030026
Liu M (2013) Expression changes of classical signal transduction system during velvet antler growth and development[D]. Changchun Univ Chin Med
Mount JG, Muzylak M, Allen S, Althnaian T, McGonnell IM, Price JS (2006) Evidence that the canonical Wnt signalling pathway regulates deer antler regeneration. Dev Dyn 235:1390–1399
Price JS, Oyajobi BO, Nalin AM, Frazer A, Russell RGG, Sandell LJ (1996) Chondrogenesis in the regenerating antler tip in red deer: Expression of collagen types I, IIA, IIB, and X demonstrated by in situ nucleic acid hybridization and immunocytochemistry. Dev Dyn 205:332–347
Rokutanda S, Fujita T, Kanatani N, Yoshida CA, Komori H, Liu W, Mizuno A, Komori T (2009) Akt regulates skeletal development through GSK3, mTOR, and FoxOs. Dev Biol 328:78–93
Shen YY, Lim BK, Liu HQ, Liu J, Irwin DM, Zhang YP (2012) Multiple episodes of convergence in genes of the dim light vision pathway in bats. PLoS One 7(4):e34564. https://doi.org/10.1371/journal.pone.0034564
Shin KH, Yun-Choi HS, Lim SS, Won DH, Kim JK (1999) Immuno-stimulating, anti-stress and anti-thrombotic effects of unossified velvet antlers. Nat Prod Sci 5:54–59
Steger V, Molnar A, Borsy A, Gyurjan I, Szabolcsi Z, Dancs G, Molnar J, Papp P, Nagy J, Puskas L, Barta E, Zomborszky Z, Horn P, Podani J, Semsey S, Lakatos P, Orosz L (2010) Antler development and coupled osteoporosis in the skeleton of red deer Cervus elaphus: expression dynamics for regulatory and effector genes. Mol Genet Genomics 284:273–287
Sui ZG, Zhang LH, Huo YS, Zhang YK (2014) Bioactive components of velvet antlers and their pharmacological properties. J Pharm Biomed Anal 87:229–240
Tseng SH, Sung CH, Chen LG, Lai YJ, Chang WS, Sung HC, Wang CC (2014) Comparison of chemical compositions and osteoprotective effects of different sections of velvet antler. J Ethnopharmacol 151:352–360
Vlad D, Kierzkowski D, Rast MI, Vuolo F, Dello Ioio R, Galinha C, Gan XC, Hajheidari M, Hay A, Smith RS, Huijser P, Bailey CD, Tsiantis M (2014) Leaf shape evolution through duplication, regulatory diversification, and loss of a homeobox gene. Science 343:780–783
Wang T, Xu Z (2010) miR-27 promotes osteoblast differentiation by modulating Wnt signaling. Biochem Biophys Res Commun 402:186–189
Wu FF, Li HQ, Jin LJ, Li XY, Ma YS, You JS, Li SY, Xu YP (2013) Deer antler base as a traditional Chinese medicine: a review of its traditional uses, chemistry and pharmacology. J Ethnopharmacol 145:403–415
Wu X, Li S, Xue P, Li Y (2017) Liraglutide, a glucagon-like peptide-1 receptor agonist, facilitates osteogenic proliferation and differentiation in MC3T3-E1 cells through phosphoinositide 3-kinase (PI3K)/protein kinase B (AKT), extracellular signal-related kinase (ERK)1/2, and cAMP/protein kinase A (PKA) signaling pathways involving β-catenin. Exp Cell Res 360:281–291
Wu X, Zheng S, Ye Y, Wu Y, Lin K, Su J (2018) Enhanced osteogenic differentiation and bone regeneration of poly(lactic-co-glycolic acid) by graphene via activation of PI3K/Akt/GSK-3beta/beta-catenin signal circuit. Biomater Sci 6(5):1147–1158
Yao B, Zhao Y, Wang Q, Zhang M, Liu M, Liu H, Li J (2012a) De novo characterization of the antler tip of Chinese Sika deer transcriptome and analysis of gene expression related to rapid growth. Mol Cell Biochem 364:93–100
Yao BJ, Zhao Y, Zhang HS, Zhang M, Liu MC, Liu HL, Li J (2012b) Sequencing and de novo analysis of the Chinese Sika deer antler-tip transcriptome during the ossification stage using Illumina RNA-Seq technology. Biotechnol Lett 34:813–822
Yuan Z, Liu E, Liu Z, Kijas JW, Zhu C, Hu S, Ma X, Zhang L, Du L, Wang H, Wei C (2017) Selection signature analysis reveals genes associated with tail type in Chinese indigenous sheep. Anim Genet 48:55–66
Zha EH, Gao SY, Pi YZ, Li XX, Wang YT, Yue XQ (2012) Wound healing by a 3.2 kDa recombinant polypeptide from velvet antler of Cervus nippon Temminck. Biotechnol Lett 34:789–793
Zha EH, Li XX, Li DD, Guo XS, Gao SY, Yue XQ (2013) Immunomodulatory effects of a 3.2 kDa polypeptide from velvet antler of Cervus nippon Temminck. Int Immunopharmacol 16:210–213
Zhang YC, Yu Y, Wang CY, Li ZY, Liu Q, Xu J, Liao JY, Wang XJ, Qu LH, Chen F, Xin PY, Yan CY, Chu JF, Li HQ, Chen YQ (2013) Overexpression of microRNA OsmiR397 improves rice yield by increasing grain size and promoting panicle branching. Nat Biotechnol 31(9):848–852
Zhang LC, Liang J, Luo WZ, Liu X, Yan H, Zhao KB, Shi HB, Zhang YB, Wang LG, Wang LX (2014) Genome-wide scan reveals LEMD3 and WIF1 on SSC5 as the candidates for porcine ear size. PLoS One 9(7):e102085. https://doi.org/10.1371/journal.pone.0102085
Zhao Y, Yao BJ, Zhang M, Wang SM, Zhang H, Xiao W (2013) Comparative analysis of differentially expressed genes in Sika deer antler at different stages. Mol Biol Rep 40:1665–1676
Zheng J, Xue H, Wang T, Jiang Y, Liu B, Li J, Liu Y, Wang W, Zhang B, Sun M (2011) miR-21 downregulates the tumor suppressor P12 CDK2AP1 and stimulates cell proliferation and invasion. J Cell Biochem 112:872–880
Acknowledgements
Thanks are due to Changchun Dongda deer Industry Co., Ltd. for its support in sample collection. This study was funded by the Agricultural Science and Technology Innovation Program of China [grant number CAAS-ASTIP-201X-ISAPS]; and the Special Economic Animals Sharing Platform in China.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Pengfei Hu declares that he/she has no conflict of interest. Tianjiao Wang declares that he/she has no conflict of interest. Huamiao Liu declares that he/she has no conflict of interest. Jiaping Xu declares that he/she has no conflict of interest. Lei Wang declares that he/she has no conflict of interest. Pei Zhao declares that he/she has no conflict of interest. Xiumei Xing declares that he/she has no conflict of interest.
Ethical approval
All procedures concerning animals were organized to accord with the guidelines of care and use of experimental animals established by the Ministry of Agriculture of China, and all protocols were approved by the Institutional Animal Care and Use Committee of Institute of Special Economic Animal and Plant Sciences, Chinese Academy of Agricultural Sciences, Changchun, China.
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Figure S1
Category of annotated genes and microRNAs. A, The overall length distributions of small RNA in heavy and light velvet antler group; B, The species distribution map of annotated genes; C, The species distribution map of annotated known microRNAs (TIF 739 KB)
Figure
S2 Statistics of the enriched KEGG categories for the differentially expressed genes. The differentially expressed genes were grouped into 12 signal pathways, among them, PI3K-Akt-signaling pathway was most relevant to velvet antler development (TIF 654 KB)
Figure S3
Validation of sequencing results by quantitative real-time RT-PCR. Transcript levels of 12 genes and 4 microRNAs, of which involve in animal organ development, in qRT-PCR analysis. The y-axis shows the relative gene expression levels analyzed by qRT-PCR. The bars represent SE (n=3). L group, light velvet antler group; H group, heavy velvet antler group (TIF 898 KB)
Rights and permissions
About this article
Cite this article
Hu, P., Wang, T., Liu, H. et al. Full-length transcriptome and microRNA sequencing reveal the specific gene-regulation network of velvet antler in sika deer with extremely different velvet antler weight. Mol Genet Genomics 294, 431–443 (2019). https://doi.org/10.1007/s00438-018-1520-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-018-1520-8