Abstract
Iron (Fe) deficiency is a frequent nutritional problem limiting apple production in calcareous soils. The utilization of rootstock that is resistant to Fe deficiency is an effective way to solve this problem. Malus halliana is an Fe deficiency-tolerant rootstock; however, few molecular studies have been conducted on M. halliana. In the present work, a transcriptome analysis was combined with qRT-PCR and sugar measurements to investigate Fe deficiency responses in M. halliana roots at 0 h (T1), 12 h (T2) and 72 h (T3) after Fe deficiency stress. Total of 2473, 661, and 776 differentially expressed genes (DEGs) were identified in the pairs of T2 vs. T1, T3 vs. T1, and T3 vs. T2, respectively. Several DEGs were enriched in the photosynthesis, glycolysis and gluconeogenesis, tyrosine metabolism and fatty acid degradation pathways. The glycolysis and photosynthesis pathways were upregulated under Fe deficiency. In this experiment, sucrose accumulated in Fe-deficient roots and leaves. However, the glucose content significantly decreased in the roots, while the fructose content significantly decreased in the leaves. Additionally, 15 genes related to glycolysis and sugar synthesis and sugar transport were selected to validate the accuracy of the transcriptome data by qRT-PCR. Overall, these results indicated that sugar synthesis and metabolism in the roots were affected by Fe deficiency. Sugar regulation is a way by which M. halliana responds to Fe deficiency stress.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Iron (Fe) is an essential micronutrient for plants (Briat et al. 2015; Zargar et al. 2015a). Low uptake of Fe alters plant chloroplast structure, blocks the synthesis of chlorophyll and reduces photosynthesis (Niebur and Fehr 1981; Briat et al. 2015). Fe availability is very low in calcareous soils, which is caused by high pH and poor aeration (Mcgeorge et al. 1935). Moreover, calcareous soils account for approximately 30% of the world’s cultivated soils, and Fe deficiency is a widespread agricultural problem affecting crop yields (Mori 1999). An approach to preventing this problem is to use Fe deficiency-tolerant rootstocks (Rombolà and Tagliavini 2006). Malus halliana shows Fe deficiency-tolerant characteristics. The investigation of responses of Malus halliana to Fe deficiency may provide new insights into its regulation and adaptation mechanisms.
Under Fe deficiency, non-graminaceous plants experience changes in metabolic levels, including increases in several enzymes involved in the glycolytic pathway, the citrate cycle and the pentose phosphate pathway (Abadía et al. 2002; Zocchi 2006; López-Millán et al. 2009). Transcriptomic and proteomic studies have also reported an upregulation of enzymes related to the glycolytic pathway and citrate cycle in roots in Fe-deficient environments (Thimm et al. 2001; Jelali et al. 2010; Anita et al. 2012). Alteration of glycolysis seems to be an important mechanism for the adaptation of plants to Fe deficiency (Mai and Bauer 2016). Accordingly, Fe deficiency induces an accumulation of organic acids, mainly malate and citrate, that can affect Fe availability (Abadía et al. 2002). Moreover, root sugar accumulation under Fe deficiency results from starch degradation and/or the reorientation of photosynthate distribution, probably via sorbitol or sucrose (Loescher et al. 1990). This reprogramming of metabolism constitutes an anaplerotic pathway of a carbon source, which partly compensates for low photosynthetic rates (López-Millán et al. 2009).
Sugar plays a vital role in plant growth and development as well as in the response to stress. Sorbitol and sucrose are the major transport forms for photoassimilates in apple trees (Reuscher et al. 2016). Additionally, sucrose is cleaved to glucose and fructose to perform additional functions. Kircher and Schopfer (2012) reported that photosynthesis-derived sugar is necessary for the regulation of root elongation growth by light. Moreover, numerous studies have shown that Fe deficiency increases the level of sucrose in plant roots (Thoiron and Briat 1999; Rellán-Álvarez et al. 2010; Jiménez et al. 2011), which is required for regulating Fe deficiency responses in plants as a signal molecule. Exogenous applications of sucrose further stimulate Fe acquisition (Lin et al. 2016). On the other hand, the increases in root sucrose concentrations under Fe deficiency also support a metabolic shift towards fermentation (Rodríguez-Celma et al. 2013).
Changes in M. halliana roots involved in responses to Fe deficiency during the early stage of deficiency were investigated. Roots are the organs that directly contact the nutrient matrix, so it is very important to study the effects of Fe deficiency on roots. Large-scale transcriptomic analyses, such as those involving DNA microarrays and RNA sequencing (RNA-Seq), are tools to gain a global view of plant responses to abiotic stress. In this study, RNA-Seq technology was used to explore a genome-wide transcriptional characterization of Malus halliana roots regarding the response to Fe deficiency stress, and to investigate the possible molecular mechanisms of Fe deficiency tolerance in this species.
Materials and methods
Plant materials
Malus halliana apple seedlings with six true leaves were used as the test material. Uniform seedlings were selected and transferred to foam boxes that contained half-strength Han’s nutrient solution (Han et al. 1994) for a 15-d preculture. The plants were then transferred to 500-mL plastic boxes (five plants per container) filled with Han’s nutrient solution that contained either 4 µmol·L−1 (-Fe) or 40 µmol·L−1 Fe(III)–EDTA (CK). The roots were harvested after 0 h (T1), 12 h (T2) and 72 h (T3) of Fe deficiency stress for transcriptome analysis, with T1 serving as a control. Each biological replicate comprised ten plants. The timepoints were selected according to previous physiological experiments (Wang et al. 2018).
Transcriptome sequencing
The root samples were harvested and immediately immersed in liquid nitrogen for RNA extraction. The total RNA was isolated using a TRIzol kit (Invitrogen, Carlsbad, CA, USA). Two replicates of each sample were sequenced. Sequencing libraries were generated using a NEBNext® UltraTM RNA Library Prep Kit for an Illumina® device (NEB, Ipswich, MA, USA). Clustering of the index-coded samples was performed on a cBot Cluster Generation System with a TruSeq PE Cluster kit v3-cBot-HS (Illumina). After cluster generation, library preparations were sequenced on an Illumina Hiseq 4000 platform. Clean data were obtained by removing reads containing the adapter, reads containing poly-N and low-quality reads from the raw data. The clean data were aligned to the apple reference genome using Tophat2 (Kim et al. 2013). The abundances of all of the genes were normalized and calculated (via uniquely mapped reads) by the expected number of fragments per kilobase of transcript sequence per million base pairs sequenced (FPKM) method (Trapnell et al. 2010). Differential expression in the paired samples was screened by the DEG-seq method (adjusted P value (p-adj) < 0.05 and |log2(Fold Change)| > 1). The DEGs identified were then subjected to Gene Onthology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses using GO-Seq (Young et al. 2010) and KOBAS 2.0 (Xie et al. 2011), respectively. GO enrichment analyses were based on the Wallenius noncentral hypergeometric distribution (corrected P value ≤ 0.05), and KEGG pathway enrichment analyses with a false discovery rate (FDR) ≤ 0.05 were conducted.
Quantitative real-time PCR
cDNA was synthesized from total RNA using a PrimeScript™ RT reagent kit with gDNA Eraser (Perfect Real Time) (TaKaRa, Dalian, China). Quantitative real-time PCR was performed using Light Cycler® 96 Instrument (Roche, Shanghai, China) with GAPDH as a reference gene. The relative levels of genes were calculated using the 2−ΔΔCt method (Livak and Schmittgen 2001). Measurements for each plate were replicated three times. The real-time PCR primer pairs used are listed in Table S1.
Sugar content determination
Glucose content was measured with a glucose test kit (No. PT-2-Y) (Comin, Suzhou, China), sucrose content was measured with a sucrose test kit (No. ZHT-2-Y) (Comin, Suzhou, China) and fructose content was measured with a fructose test kit (No. GT-2-Y) (Comin, Suzhou, China). All operations are strictly operated according to the kit instructions.
Statistical analyses
Parameters were statistically tested by analyses of variance and comparisons of means were performed with a Duncan’s test (P < 0.05). Statistical analyses were performed with SPSS, version 22.0 (IBM, Armonk, NY, USA). Figures were made with the drawing software Origin 8.0 (OriginLab, Hampton, MA, USA).
Data availability
All Illumina sequence data have been deposited in Sequence Read Archive with the project ID PRJNA400500.
Results
RNA-Seq transcriptome of M. halliana
RNA-Seq libraries were established from the roots of Fe-deficient-treated seedlings at T1, T2 and T3. Correlations between gene expression levels in samples are important for testing the reliability of experiments. The correlation coefficients were greater than 0.93 between replicates (Fig. 1a). RNA-Seq generated more than 38.76 million raw reads for each sample, and two biological replicates were set for each time point (Table 1). Of these reads, the GC content of the libraries was approximately 46.00%. After quality control, a number of clean reads from 33.34 to 44.14 millions were produced that had an approximate Q30 of 93.00% and a number of clean reads from 24.28 to 26.60 millions were mapped to the apple genome.
Differentially expressed genes during iron deficiency
The differentially expressed genes (DEGs) at T1, T2 and T3 were examined via an adjusted P value (p-adj) < 0.05 and |log2(Fold Change)| > 1 as the threshold, and the DEGs were identified by three pairwise comparisons (Fig. 1b). Total of 2473, 661, and 776 DEGs were found in the pairs T2 vs. T1, T3 vs. T1, and T3 vs. T2, respectively. A comparison of these three datasets showed that 12 genes overlapped among T2 vs. T1, T3 vs. T1, and T3 vs. T2. The results showed that 1477 DEGs were significantly upregulated and that 996 DEGs were downregulated in the T2 library compared with the T1 library. In total, 369 DEGs were significantly upregulated, and 293 DEGs were downregulated in T3 compared with T1. In addition, 339 DEGs were upregulated while 437 genes were downregulated in T3 compared with T2 (Fig. 1c).
Functional classification of differentially expressed genes during iron deficiency
To decipher the functions of these DEGs, GO term enrichment analysis was performed for each group comparison using GO-Seq (Fig. 2). The first three categories of cellular component were photosynthetic membrane, thylakoid membrane and thylakoid, while heat-shock protein binding, oxidoreductase activity and ADP binding were the main annotations represented in the molecular function category. For the biological process category, oxidation–reduction process, single-organism metabolic process, small molecule metabolic process, carboxylic acid metabolic process and organic acid metabolic process were strongly enriched in T2 vs. T1, T3 vs. T1 and T3 vs. T2.
We also mapped the DEGs in the KEGG database to perform biochemical pathway analysis (Table 2). Four, six and five KEGG pathways were significantly enriched in T2 vs. T1, T3 vs. T1, and T3 vs. T2, respectively. The DEGs were associated with various KEGG pathways involved in photosynthesis, metabolism and protein processing. The photosynthesis pathway or photosynthesis–antenna protein pathway was induced in T2 vs. T1, T3 vs. T1 and T3 vs. T2. Interestingly, the glycolysis/gluconeogenesis pathway, tyrosine metabolism and fatty acid degradation were induced only at T3. Notably, we observed specific enrichment of genes in the glycolysis/gluconeogenesis pathway (mdm00010).
Expression profiles of glycolysis-related genes
We analyzed the glycolysis/gluconeogenesis-related genes, including phosphoglucomutase (PGM), glucose-6-phosphate isomerase (GPI), fructose-1,6-bisphosphatase (FBPase), 6-phosphofructokinase (PFK), fructose-bisphosphate aldolase (FBA), phosphoglycerate kinase (PGK), enolase (ENO) and pyruvate kinase (PK) (Fig. 3; Table S2). The expressions of most PGM genes (103439690, 103439907 and 103452245) decreased at T3, while the expression of seven genes (103401972, 103417145, 103420974, 103440430, 103440507, 103440552 and 103,440,553) encoding the GPI was upregulated at T3. The expression of FBPase (103,416,291, 103433550, 103444638, 103450517 and 103450592) genes was upregulated under Fe deficiency. The expression of two PFKase genes (103400183 and 103439066) was upregulated at T3, the expression of five genes was upregulated at T2 (103408071, 103414866, 103418180,103438916 and 103439065), that of two genes (103411637 and 103448202) was downregulated under Fe deficiency, and that of one gene (103439064) was upregulated consistently. In addition, genes coding for FBA exhibited a higher expression level than did genes coding for other enzymes. The expression of four PGK genes (103409034, 103435386, 103445876 and 103445883) showed upregulation under Fe deficiency. The expression of ENO genes (103415341, 103419156, 103431235, 103437941, 103447452, 103449384, 103453924 and 103455462) showed little response to Fe deficiency. There were 27 genes encoding PK. Among them, 15 genes (103402797, 103408335, 103410064, 103410070, 103414668, 103416875, 103420072, 103420595, 103425723, 103431059 103431990, 103438858, 103442109, 103445912 and 103449914) were upregulated at T3, 9 genes (103403377, 103412636, 103414179, 103422541, 103425559, 103429773, 103439830, 103443326 and 103451429) were downregulated at T3, and 3 genes (103424068, 103436310 and 103436870) showed no obvious changes under Fe deficiency.
Expression profiles of sugar-related genes
The genes for sugar synthesis, metabolism and transport were analyzed, including sucrose synthase (SS), sucrose phosphate synthase (SPS), sucrose transport protein (SUC), alkaline/neutral invertase (A/N Invs), plastidic glucose transporter (PGT) and sugar transporter (ST) (Fig. 4). The expression of sucrose biosynthesis genes, seven SS genes (103400901, 103409880, 103417499, 103426254, 103426255, 103449164 and 103453958), increased in the roots under the Fe-deficient environment. However, expression of the SPS genes (103400066, 103411875, 103417980 and 103456219) was downregulated at T2 and T3 compared to T1. The expression of the most of the A/N Invs genes (103402206, 103403505, 103413070, 103413827, 103427242 and 103436229) was downregulated at T3. The levels of expression of most SUC (103401916, 103410965, 103416524103422077, 103441771, 103443890 and 103444946) strongly changed at T2. The expression of most PGT genes (103402038, 103402916, 103410965, 103416524103441771 and 103443890) was upregulated at T2. Here, the ST genes comprised two types: bidirectional sugar transporter genes and sugar transporter ERD6 genes. Two bidirectional sugar transporter genes (103452184 and 103423978) showed high expression at T2. In addition, there was high expression of the sugar transporter ERD6 genes 103450477 and 103413416.
Expression profiles of photosynthesis-related genes
The expression levels of photosynthesis-related genes, such as photosystem protein, cytochrome b6-f subunit, ferredoxin, ATP synthase chain and plastocyanin, were analyzed (Fig. 5). The expression of all photosynthesis-related genes was upregulated under Fe deficiency. Two photosystem protein genes (103430669 and 103416404), one plastocyanin gene (103450341) and one ferredoxin gene (103418288) showed the highest expression at T3. In addition, other genes were expressed at the highest level at T2.
Changes in sugar content under iron deficiency
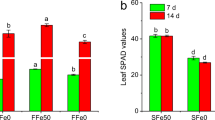
Sugar concentration was determined under Fe deficiency by examining the extracted sugars from the roots and leaves (Fig. 6). The levels of glucose were significantly decreased in the roots of both T2 and T3 compared to T1. However, there were no significant changes in glucose content in the leaves (Fig. 6a). Interestingly, the changes in fructose contents in the roots and leaves were opposite for glucose (Fig. 6b). Under Fe deficiency, the levels of fructose were significantly decreased in the leaves, but there were no clear changes in the roots. The trend of the change in sucrose content was opposite that of the other two kinds of sugar (Fig. 6c). The sucrose contents increased under Fe deficiency in both the roots and leaves; the change in the roots occurred at T2, while the change in the leaves occurred at T3.
Effect of iron deficiency on sugar levels in M. halliana roots and leaves. Glucose (a), fructose (b) and sucrose (c) contents were measured at three time points: T1, T2 and T3 under Fe deficiency. Data are presented as means ± SE (n = 3). The data with different lower-case letters in same organ show significant difference (p < 0.05)
Validation of the RNA-Seq analysis results by qRT-PCR
Moreover, to validate the accuracy of the experiments, 15 genes related to glycolysis/gluconeogenesis, sugar synthesis and sugar transport were selected for qRT-PCR analysis. The relative expression of the 15 genes was measured using independent samples with the same treatment as those subjected to RNA-Seq analysis. The measurements were replicated three times. GAPDH was used as a reference gene for data normalization. Eleven genes showed significant correlations (P = 0.05) between the qRT-PCR results and the RNA-Seq analysis results, which indicated that the RNA-Seq data were highly reliable (Fig. 7).
Discussion
Fe deficiency is one of the most frequent nutritional problems affecting apple production in calcareous soils. Plants alter their physiological processes and root architecture to cope with low availability of Fe in soils (López-Millán et al. 2000; Fu et al. 2017). A global information analysis of plant responses to Fe deficiency may provide new insights into the molecular functions and mechanisms involved. Efforts have been made to obtain a general overview of Fe deficiency-induced changes in the transcriptome of M. xiaojinensis (Wang et al. 2014), but information is scarce regarding the carbohydrate regulation especially in M. halliana. In the present study, the RNA-Seq analysis method was used to investigate the changes induced by Fe deficiency at three time points in the transcriptome profile of roots extracted from M. halliana plants grown hydroponically. The results of our transcriptomic analyses showed that more than 650 DEGs in the pairs T2 vs. T1, T3 vs. T1, and T3 vs. T2 were found (Fig. 1b). According to the GO classification, these DEGs are involved mainly in photosynthetic membrane, thylakoid, the oxidation–reduction process and acid metabolism (Fig. 2). In addition, the DEGs are enriched in pathways related to photosynthesis, photosynthesis–antenna protein, glycolysis/gluconeogenesis, tyrosine metabolism and fatty acid degradation (Table 2).
In view of gene expression profiling, the glycolysis pathway may be more important in Fe deficiency responses than other pathways. An increase in glycolysis has been widely reported in Arabidopsis thaliana (Yang et al. 2010), Medicago truncatula (López-Millán et al. 2011), sugar beet (López-Millán et al. 2000), and tomato (Anita et al. 2012). Positive modulation of glycolysis is an important mechanism for the adaptation of plants to Fe deficiency (Mai and Bauer 2016), which may be due to an increased demand for organic acids in Fe-deficient roots (López-Millán et al. 2013). In this experiment, although the citrate cycle was not significantly enriched, the glycolysis/gluconeogenesis, tyrosine metabolism, fatty acid degradation, and taurine and hypotaurine metabolism pathways all point to pyruvate, the substrate of the citrate cycle.
The enzymes PGM, GPI, FBPase, PFK, FBA, PGK, ENO and PK are involved in the glycolysis pathway. These enzymes play an important role in glycolysis; for example, PGM catalyzes the bidirectional conversion of glucose-1-phosphate (G-1-P) and G-6-P, which is the first step of glycolysis (Gururaj et al. 2004). FBA, which occupies a central position in the glycolysis pathway, is at the crossroads of several metabolic pathways that involve metabolites derived from glycolysis. The genes coding for these enzymes were altered under Fe deficiency (Table S2). In Arabidopsis, an expression analysis of genes involved in the glycolysis pathway revealed an induction of several enzymes (Thimm et al. 2001). Within 72 h of Fe-deficient growth, the expression levels of these genes increased, but no changes were found following 1 d of Fe-deficient growth (Thimm et al. 2001). In this experiment, the expression levels of PGM (103439690), GPI (10344043, 103401972, 103440430 and 103417145), and all of the FBPase, PFK, FBA (103449400 and 103453790) and PGK genes were induced regardless of whether Fe deficiency persisted for 12 or 72 h which indicates that M. halliana has a rapid response to Fe deficiency. Additionally, these transcriptional data fit well with proteomic data previously published before (Rodríguez-Celma et al. 2011) as well as the increased activity of PK and PFK recorded in cucumber roots under Fe deficiency (Espen et al. 2000). Specifically, the expression levels of ENO, FBA and PGK were found to be upregulated in Fe-deficient Prunus, Medicago, Chlorella, Solanum and Cucumis roots (Brumbarova et al. 2008; Rellán-Álvarez et al. 2010; Rodríguez-Celma et al. 2011, 2013; Kircher and Schopfer 2012).
Enhanced root glycolysis under Fe deficiency leads to sugar accumulation (Zargar et al. 2015a; Lin et al. 2016). Sucrose not only provides materials and energy for plant growth and development but also functions as a signaling molecule for the regulation of various physiological processes in plants, such as root growth and photosynthesis (Kircher and Schopfer 2012; Zargar et al. 2015a). In this experiment, sucrose accumulated in the roots and leaves under Fe deficiency (Fig. 6 C), similar to the results of previous studies in Prunus and Arabidopsis thaliana (Jiménez et al. 2011; Zargar et al. 2015b). In addition, the expression levels of SS genes increased under Fe deficiency, especially 103449164, which was upregulated almost threefold at T3 compared with T1. However, the expression levels of SPS genes were downregulated under Fe deficiency. This result may have occurred because SS plays a major role in the synthesis of sucrose under Fe deficiency. One of the SUC genes (103451844) was significantly downregulated under Fe deficiency, which also coincided with an increase in sucrose content in the roots.
Fructose and glucose are products of sucrose hydrolysis catalyzed by A/N Invs. A/N Invs enzymes play a major role in sucrose partitioning and long-distance transport (Winter and Huber 2000). In this experiment, A/N Invs (103441853 and 103452658) showed high expression at T2. However, the glucose content significantly decreased in the roots and changed little in the leaves under Fe deficiency. The alteration of fructose content is opposite to that of glucose. The response differences between glucose and fructose may due to carbohydrates partitioning under Fe deficiency. Additionally, Lin et al. (2016) find that glucose and fructose contents in Arabidopsis thaliana roots decreased under Fe deficiency. Nevertheless, a survey of the sugar concentration in Arabidopsis thaliana shoots under Fe deficiency indicated that glucose and fructose concentrations increase as sucrose concentration increased (Zargar et al. 2015b). This finding may reveal that different species and different organizations have different ways to respond to Fe deficiency stress. Moreover, in the study of three cherry rootstocks under Fe deficiency stress, Jiménez et al. (2011) found that the glucose concentration decreased in the more tolerant rootstock and increased in the more sensitive rootstock after Fe deficiency, which indicates that different resistant plants show different response modes to Fe deficiency. The glucose regulation pattern of M. halliana is similar to that of more tolerant cherry. High expression of PGT genes was induced in the roots in response to Fe deficiency, which may be part of the reason that the glucose in the roots decreased while that in leaves did not change significantly.
Overall, the results indicated that sugar metabolism is affected by Fe deficiency and might lead to the disruption of sugar synthesis and utilization. Sugar accumulation originates from starch degradation and reorientation of photosynthate partitioning, probably via sucrose (Loescher et al. 1990; Zargar et al. 2015a). Therefore, Zargar et al. (2015a) reported that reduced photosynthetic activity may be affected by high sugar concentrations. Interestingly, in this work, although the sucrose content increased, the photosynthesis and photosynthesis–antenna protein pathways were enhanced in M. halliana under Fe deficiency, and the expression of all photosynthesis-related genes was upregulated under Fe deficiency. This finding indicates that sugar regulatory effects in different plants are different under Fe deficiency. Not only sucrose metabolism but also the whole of carbohydrate metabolism may play important roles in strategies for coping with Fe deficiency stress in M. halliana and may affect photosynthesis levels.
Sugar regulation is an important part of the root response of M. halliana to Fe deficiency. Furthermore, Fe deficiency causes the upregulated expression of photosynthesis-related genes. This study provides a foundation for an improved understanding of Fe tolerance responses in apple and raises the issue of the relationship between photosynthesis and sugar under Fe deficiency. In future studies, we will focus on investigating the network of carbohydrate metabolism, transport, partitioning and photosynthesis in M. halliana under Fe deficiency and screening for key genes for cloning and functional analyses.
References
Abadía J, López-Millán AF, Rombolà A, Abadía A (2002) Organic acids and Fe deficiency: a review. Plant Soil 241(1):75–86
Anita Z, Laura Z, Nicola T, Mario P, Roberto P, Zeno V, Cesco S (2012) Genome-wide microarray analysis of tomato roots showed defined responses to iron deficiency. BMC Genom 13(1):101
Briat JF, Dubos C, Gaymard F (2015) Iron nutrition, biomass production, and plant product quality. Trends Plant Sci 20(1):33–40
Brumbarova T, Matros A, Mock HP, Bauer P (2008) A proteomic study showing differential regulation of stress, redox regulation and peroxidase proteins by iron supply and the transcription factor fer. Plant J 54(2):321–334
Espen L, Dell’Orto M, De NP, Zocchi G (2000) Metabolic responses in cucumber (Cucumis sativus L.) roots under Fe-deficiency: a 31p-nuclear magnetic resonance in-vivo study. Planta 210(6):985–992
Fu L, Zhu Q, Sun Y, Du W, Pan Z, Peng S (2017) Physiological and transcriptional changes of three citrus rootstock seedlings under iron deficiency. Front Plant Sci 8:1104
Gururaj A, Barnes CJ, Vadlamudi RK, Kumar R (2004) Regulation of phosphoglucomutase 1 phosphorylation and activity by a signaling kinase. Oncogene 23(49):8118–8127
Han ZH, Wang Q, Shen T (1994) Comparison of some physiological and biochemical characteristics between iron-efficient and inefficient species in the genus Malus. J Plant Nutr 17:230–241
Jelali N, Wissal M, Dell’Orto M, Abdelly C, Gharsalli M, Zocchi G (2010) Changes of metabolic responses to direct and induced Fe deficiency of two Pisum sativum cultivars. Environ Exp Bot 68(3):238–246
Jiménez S, Ollat N, Deborde C, Maucourt M, Rellán-Álvarez R, Moreno MA, Gogorcena Y (2011) Metabolic response in roots of Prunus rootstocks submitted to iron chlorosis. J Plant Physiol 168(5):415–423
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL (2013) Tophat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14(4):R36
Kircher S, Schopfer P (2012) Photosynthetic sucrose acts as cotyledon-derived long-distance signal to control root growth during early seedling development in Arabidopsis. P Natl Acad Sci USA 109(28):11217–11221
Lin XY, Ye YQ, Fan SK, Jin CW, Zheng SJ (2016) Increased sucrose accumulation regulates iron-deficiency responses by promoting auxin signaling in Arabidopsis plants. Plant Physiol 170(2):907
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25(4):402–408
Loescher WH, Mccamant T, Keller JD (1990) Carbohydrate reserves, translocation, and storage in woody plant roots. Hortscience 25(3):274–281
López-Millán AF, Morales F, Andaluz S, Gogorcena Y, Abadía A, Rivas JDL, Abadía J (2000) Responses of sugar beet roots to iron deficiency. changes in carbon assimilation and oxygen use. Plant Physiol 124(2):885
López-Millán AF, Morales F, Gogorcena Y, Abadía A, Abadía J (2009) Metabolic responses in iron deficient tomato plants. J Plant Physiol 166:375–384
López-Millán AF, Grusak MA, Abadía A, Abadía J (2013) Iron deficiency in plants: an insight from proteomic approaches. Front Plant Sci 4(9):254
Mai HJ, Bauer P (2016) From the proteomic point of view: integration of adaptive changes to iron deficiency in plants. Curr Plant Biol 5(C):45–56
Mcgeorge WT, Buehrer TF, Breazeale JF (1935) Phosphate availability in calcareous soils: a function of carbon dioxide and pH. J Am Soc Agron 27(5):330–335
Mori S (1999) Iron acquisition by plants. Curr Opin Plant Biol 2(3):250–253
Niebur WS, Fehr WR (1981) Agronomic evaluation of soybean genotypes resistant to iron deficiency chlorosis. Crop Sci 21(4):551–554
Rellán-Alvarez R, Andaluz S, Rodríguez-Celma J, Wohlgemuth G, Zocchi G, Alvarez-Fernández A, Fiehn O, López-Millán AF, Abadía J (2010) Changes in the proteomic and metabolic profiles of Beta vulgaris root tips in response to iron deficiency and resupply. BMC Plant Biol 10:120
Reuscher S, Fukao Y, Morimoto R, Otagaki S, Oikawa A, Isuzugawa K, Shiratake K (2016) Quantitative proteomics based reconstruction and identification of metabolic pathways and membrane transport proteins related to sugar accumulation in developing fruits of pear (Pyrus communis). Plant Cell Physiol 57(3):505–518
Rodríguez-Celma J, Lattanzio G, Grusak MA, Abadía A, Abadía J, Lópezmillán AF (2011) Root responses of Medicago truncatula plants grown in two different iron deficiency conditions: changes in root protein profile and riboflavin biosynthesis. J Proteome Res 10(5):2590–2601
Rodríguez-Celma J, Lattanzio G, Jiménez S, Briat JF, Abadía J, Abadía A, Gogorcena Y, López-Millán AF (2013) Changes induced by Fe deficiency and Fe resupply in the root protein profile of a peach-almond hybrid rootstock. J Proteome Res 12(3):1162
Rombolà AD, Tagliavini M (2006) Iron nutrition of fruit tree crops. In: Barton LL, Abadía J (eds) Iron nutrition in plants and rhizopheric micoorganisms. Springer, Dordrecht, pp 61–83
Thimm O, Essigmann B, Altmann T, Buckhout TJ (2001) Response of Arabidopsis to iron deficiency stress as revealed by microarray analysis. Plant Physiol 127(3):1030–1043
Thoiron S, Briat JF (1999) Differential expression of maize sugar responsive genes in response to iron deficiency. Plant Physiol Biochem 37:759–766
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, Pachter L (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28(5):511–515
Wang S, Lu B, Wu T, Zhang X, Xu X, Han Z, Wang Y (2014) Transcriptomic analysis demonstrates the early responses of local ethylene and redox signaling to low iron stress in Malus xiaojinensis. Tree Genet Genomes 10(3):573–584
Wang YX, Hu Y, Zhu YF, Baloch AW, Jia XM, Guo AX (2018) Transcriptional and physiological analyses of short-term iron deficiency response in apple seedlings provide insight into the regulation involved in photosynthesis. BMC Genom 19:461
Winter H, Huber SC (2000) Regulation of sucrose metabolism in higher plants: localization and regulation of activity of key enzymes. Crit Rev Plant Sci 35(4):253–289
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S, Kong L, Gao G, Li CY, Wei L (2011) KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39:W316–W322
Yang TJ, Lin WD, Schmidt W (2010) Transcriptional profiling of the Arabidopsis iron deficiency response reveals conserved transition metal homeostasis networks. Plant Physiol 152(4):2130–2141
Young MD, Wakefield MJ, Smyth GK, Oshlack (2010) Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11(2):R14
Zargar SM, Agrawal GK, Rakwal R, Fukao Y (2015a) Quantitative proteomics reveals role of sugar in decreasing photosynthetic activity due to Fe deficiency. Front Plant Sci 6(592):592
Zargar SM, Kurata R, Inaba S, Oikawa A, Fukui R, Ogata Y, Agrawal GK, Rakwal R, Fukao Y (2015b) Quantitative proteomics of Arabidopsis shoot microsomal proteins reveals a cross-talk between excess zinc and iron deficiency. Proteomics 15(7):1196
Zocchi G (2006) Metabolic changes in iron-stressed dicotyledonous plants. Iron nutrition in plants and rhizospheric microorganisms. Springer Netherlands, 359–370
Funding
This work was supported by Gansu Agricultural University Youth Postgraduate Tutor Support Fund Project (project No. GAU-2NDS-201710), Gansu Education Department University Research Project (project No. 2018A-035) and Lanzhou Science and Technology Bureau Program (project No. 2015-3-76).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that we have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hu, Y., Zhu, Yf., Guo, Ax. et al. Transcriptome analysis in Malus halliana roots in response to iron deficiency reveals insight into sugar regulation. Mol Genet Genomics 293, 1523–1534 (2018). https://doi.org/10.1007/s00438-018-1479-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-018-1479-5