Abstract
During the sexual life cycle of filamentous fungi, multicellular fruiting bodies are generated for the dispersal of spores. The filamentous ascomycete Sordaria macrospora has a long history as a model system for studying fruiting body formation, and two collections of sterile mutants have been generated. However, for most of these mutants, the underlying genetic defect remains unknown. Here, we investigated the mutant spadix (spd) that was generated by X-ray mutagenesis in the 1950s and terminates sexual development after the formation of pre-fruiting bodies (protoperithecia). We sequenced the spd genome and found a 22 kb deletion affecting four genes, which we termed spd1-4. Generation of deletion strains revealed that only spd4 is required for fruiting body formation. Although sterility in S. macrospora is often coupled with a vegetative hyphal fusion defect, Δspd4 was still capable of fusion. This feature distinguishes SPD4 from many other regulators of sexual development. Remarkably, GFP-tagged SPD4 accumulated in the nuclei of vegetative hyphae and fruiting body initials, the ascogonial coils, but not in sterile tissue from the developing protoperithecium. Our results point to SPD4 as a specific determinant of fruiting body formation. Research on SPD4 will, therefore, contribute to understanding cellular reprogramming during initiation of sexual development in fungi.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Multicellularity permits the formation of specialized cells and tissues, and multicellular organisms have evolved independently several times in different eukaryotic clades. Filamentous ascomycetes have been shown to be excellent model organisms for studying multicellular development. These fungi are characterized by the formation of long tubular cells, the hyphae, which are compartmentalized by septa that permit cytoplasmic and organellar movement through septal pores (Gull 1978). In contrast to yeasts, filamentous fungi are truly multicellular, in that they form large colonies of interconnected hyphae, the mycelium. Mycelia contain different cell types, such as leading hyphae at the colony periphery and highly vacuolated trunk hyphae in the colony interior (Bistis et al. 2003).
The fungal mycelium is able to generate higher order structures, such as conidiophores for asexual propagation, fruiting bodies for sexual propagation, and sclerotia for long-term survival. The generation of these structures has been studied in a number of model ascomycetes. For example, conidiophore development has been analyzed in Aspergillus nidulans, Aspergillus fumigatus, and Penicillium chrysogenum (e.g., Hoff et al. 2010; Harting et al. 2013; Cai et al. 2015; Chi and Craven 2016), fruiting body formation in A. nidulans, Neurospora crassa, Podospora anserina, and Sordaria macrospora (e.g., Fleissner et al. 2009; Kim et al. 2009, 2012; Lord and Read 2011; Coppin et al. 2012; Voigt and Pöggeler 2013; Lehr et al. 2014; Sarikaya-Bayram et al. 2014; Teichert et al. 2014b), and sclerotia formation in Botrytis cinerea and Sclerotinia sclerotiorum (e.g., Duan et al. 2013; Siegmund et al. 2015). Among these higher order structures, fruiting bodies are the most complex and harbor many cell types that are not present in the vegetative mycelium (Bistis et al. 2003; Lord and Read 2011). Therefore, vast changes in the hyphal architecture and function are a prerequisite for fruiting body formation (Pöggeler et al. 2006).
In this study, we investigated sexual development in the filamentous ascomycete S. macrospora, which has a long history as a model for fruiting body formation and meiotic recombination (Esser and Straub 1956; Esser and Straub 1958; Arnaise et al. 1984). S. macrospora is a saprophytic fungus that forms closed pear-shaped fruiting bodies (perithecia) after 7 days of growth under laboratory conditions. Sexual development starts with the formation of curled hyphae, termed ascogonia, which represent the female gametangia. Subsequently, ascogonia are wrapped by enveloping hyphae, leading to the formation of the spherical protoperithecium (Esser 1982; Lord and Read 2011). Though the young protoperithecium is loosely tangled, maturing protoperithecia develop a dense outer layer of adhered hyphae termed peridium.
Many factors required for perithecia formation were described recently for S. macrospora. These include transcription factors, mating type proteins, signaling components, subunits of the striatin-interacting phosphatase and kinase (STRIPAK) complex, and proteins involved in autophagy (reviewed in Kück et al. 2009, 2016; Teichert et al. 2014a). Interestingly, most of these factors are also required for vegetative hyphal fusion (VHF), a process assumed to enable a fungal colony to rapidly establish a mycelial network due to fast distribution of nutrients, signaling molecules, and organelles (Aanen et al. 2008; Read et al. 2009; Simonin et al. 2012). A general phenomenon in ascomycetes seems to be that proteins required for VHF are also required for other developmental processes. For example, NADPH oxidase 1, MAP kinase MAK1 of the cell wall integrity pathway, and the scaffold protein PRO40/SOFT are not only involved in VHF, but also in fruiting body formation, symbiotic or pathogenic life style, and/or the development of epigenetically controlled growth phenotypes in diverse ascomycetes (Malagnac et al. 2004; Fleißner et al. 2005; Kicka et al. 2006; Engh et al. 2007; Rech et al. 2007; Prados Rosales and Di Pietro 2008; Lichius 2010; Charlton et al. 2012; Lichius et al. 2012; Kayano et al. 2013; Dirschnabel et al. 2014; Teichert et al. 2014b; Tong et al. 2014; Becker et al. 2015; Turra et al. 2015).
The strong correlation of a sexual developmental defect and a VHF defect has led to the hypothesis that VHF is a prerequisite for fruiting body formation. However, recent results challenge this hypothesis; mutants have been identified that are either sterile and fusion-competent or fertile and fusion-deficient (for an overview, see Lichius and Lord 2014). For example, the S. macrospora autophagy mutants ΔSmatg4, ΔSmatg8, and ΔSmjlb1, as well as N. crassa prm-1, belong to the sterile, fusion-competent category (Fleissner et al. 2009; Voigt et al. 2013; Voigt and Pöggeler 2013), and N. crassa mutants ham-4 and ham-11 belong to the fertile, fusion-deficient category (Simonin et al. 2010; Fu et al. 2011; Leeder et al. 2013). The question of how the specific factors control fruiting body formation and regulate others and themselves is still unanswered.
In this study, we analyzed the sexual mutant spadix (spd) from S. macrospora that was generated by X-ray mutagenesis and previously described to form aberrant ascogonia and few very small protoperithecia (Esser and Straub 1958; Lord and Read 2011). We found that spd has a pleiotropic phenotype exhibiting sterility, pigment leakage into the medium, and cell lysis, but the mutant is capable of VHF. We identified the mutations underlying the spd phenotype and describe spd4, a new gene essential for sexual fruiting body formation.
Materials and methods
Strains and growth conditions
All S. macrospora strains used in this study are listed in Table S1. Unless stated otherwise, growth conditions were as described previously (Kamerewerd et al. 2008; Dirschnabel et al. 2014). Transformation was carried out by protoplast formation as described (Dirschnabel et al. 2014), and transformants were selected on medium containing either nourseothricin (50 µg/ml) or hygromycin B (80 U/ml). For measuring vegetative growth, strains were pre-cultured on corn meal-malt fructification medium (BMM) for 2 days, and standard inocula were transferred to synthetic Westergaard’s (SWG) medium (Nowrousian et al. 2007). The growth front was marked after one and two days, and the distance between these two marks was measured. Data are from three independent experiments with three technical replicates each.
DNA preparation, Illumina sequencing, and mapping
Sterile mutant lu/spd from our laboratory collection was back-crossed several times to wild-type, developmental mutants, or brown-spored fus to reduce unrelated background mutations (Nowrousian et al. 2012). For whole-genome sequencing, the spd isolate S102018 was crossed to fus (Fig. S1). DNA was extracted from 40 fertile and 40 sterile progeny of this cross as described previously (Nowrousian et al. 2012). Five micrograms of pooled genomic DNA from each spd and wt was subjected to 50 bp single read Illumina/Solexa sequencing with a HiSeq2000 at GATC Biotech (Konstanz, Germany). Cleaning of raw data, mapping to the S. macrospora reference genome (Nowrousian et al. 2010; Teichert et al. 2012), analysis of sequence variants, and detection of uncovered regions were performed as described (Nowrousian et al. 2012) using the Burrows Wheeler Alignment tool (Li and Durbin 2009), SAMtools (Li et al. 2009), and custom-made Perl scripts. Raw sequence data from sequencing mutant spd and wildtype were submitted to the NCBI sequence read archive (accession no. SRX1868445 and SRX1867979).
Generation of deletion strains
Plasmids and oligonucleotides used in this study are listed in Tables S2 and S3, respectively. To generate deletion strains for SMAC_01961 (spd1), SMAC_01962 (spd2), and SMAC_01963 (spd3), plasmids pKO-spd1, pKO-spd2, and pKO-spd3, respectively, were cloned by yeast recombination. For this purpose, 5′ and 3′ flanking regions were amplified from S. macrospora genomic DNA for spd1 (5′, 1961-5fw/1961-5rv, 1065 bp; 3′, 1961-3fw/1961-3rv, 956 bp), spd2 (5′, 1962-5fw_IT/1962-5rv_IT, 987 bp; 3′, 1962-3fw/1963-3rv, 1000 bp), and spd3 (5′, 1963-5fw/1963-5rv, 1000 bp; 3′, 1963-3fw/1963-3rv, 1000 bp), and transformed into yeast strain PJ69-4a (James et al. 1996) together with EcoRI/XhoI-digested pRS426 (Christianson et al. 1992) and a 1.4 kb hph cassette, derived from pDrivehph after EcoRI hydrolysis (Nowrousian and Cebula 2005).
For generation of a spd4 deletion strain, plasmid pFlip-spd4 was constructed as follows: 5′ and 3′ flanking regions of spd4 were amplified from S. macrospora genomic DNA, using primer pairs 1964-KO1-EcoRV/1964-KO2-PstI (1099 bp) and 1964-KO3-HindIII/1964-KO4-BglII (1037 bp), respectively, and sub-cloned into pDrive (Qiagen, Germany), generating pD5-spd4 and pD3-spd4. The 3′ flanking region was cut from pD3-spd4 with HindIII and BglII and ligated into pFlip (Bloemendal et al. 2014), generating pFlip3-spd4. The 5′ flanking region was cut from pD5-spd4 with EcoRV and PstI and ligated into the corresponding sites of pFlip3-spd4, generating pFlip-spd4.
Plasmids pKO-spd1, pKO-spd2, and pKO-spd3 were digested with EcoRI, and each construct was transformed into S. macrospora Δku70 (Pöggeler and Kück 2006). Preparation of DNA and Southern hybridization were performed as described (Kamerewerd et al. 2008). PCR-verified primary transformants (see Figs. S2–S4 for oligonucleotides) were crossed to spore color mutant fus (Nowrousian et al. 2012), and ascospore isolates showing hygromycin B resistance and nourseothricin sensitivity were analyzed by PCR and Southern hybridization (Figs. S2–S4). To construct a Δspd4 strain, pFLIP-spd4 was digested with EcoRV and BglII, the deletion cassette was transformed into Δku70, and primary transformants were analyzed by PCR. The spd4 deletion cassette contains an flp recombinase gene controlled by the inducible Smxyl promoter for marker recycling (Bloemendal et al. 2014), and thus, tetrad analysis was employed to identify hygromycin B- and nourseothricin-sensitive homokaryotic deletion strains. For verification, these strains were analyzed by PCR and Southern hybridization (Fig. S5).
Restoration of the spd1 ORF in the spd mutant background
To complete the spd1 ORF in the spd background, we first generated spd/Δku70 by crossing the single mutants to generate an spd strain favoring homologous recombination. Plasmid pKI1961 was generated by yeast recombination, containing the complete spd1 ORF, an hph resistance cassette, and the region located downstream of the spd deletion. PCR fragments were amplified from genomic DNA using primer pairs 1961-KI-01/1961-KI-02 (2546 bp) and 1961-KI-03/1961-KI-04 (1231 bp) and transformed into PJ69-4A together with EcoRI/XhoI-digested pRS426 (Christianson et al. 1992) and a 1.4 kb hph cassette, derived from pDrivehph after EcoRI hydrolysis (Nowrousian and Cebula 2005). pKI1961 was digested with MunI and XhoI and transformed into spd/Δku70. PCR-verified primary transformants (see Fig. S6 for oligonucleotides) were crossed to fus, and ascospore isolates showing hygromycin B resistance and nourseothricin sensitivity were tested by PCR and Southern analyses (Fig. S6) to confirm the restoration of spd1. Strain S131717 was used for further analysis and designated spd::spd1.
Generation of plasmids
Cloning and propagation of plasmids were performed using the standard laboratory protocols (Sambrook and Russel 2001) and Escherichia coli XL1 Blue MRF’ (Jerpseth et al. 1992) as a host strain. Alternatively to restriction-ligation-mediated cloning, yeast recombination was applied as described previously (Colot et al. 2006; Bloemendal et al. 2012) using Saccharomyces cerevisiae PJ69-4A (James et al. 1996) as a host.
All plasmids used in this study are listed in Table S2. Vectors with PCR fragments missing in the 22 kb deletion in spd were generated by amplifying fragments from S. macrospora wild-type DNA and sub-cloning into pDrive (Qiagen) or pTOPO (LifeTechnologies). Specifically, pRR343-3 and pRR345-1 are based on pDrive and were generated with primer pairs spd_07/spd_10 and spd_13/spd_04, respectively. pRR350-10, pRR351-6, and pRR352-4 are based on pTOPO and were generated with primer pairs spd_09/spd_12, spd_11/spd_18, and spd_14/spd_17, respectively.
Vectors p1935_OE, p-SPD1_OE, p-SPD2_OE, and pSPD3_OE were generated by homologous recombination in yeast. For SMAC_01935, a 1956 bp PCR fragment was generated with primers 1935-01/1935-02. For spd1, a 1530 bp PCR fragment was generated with primers 1961-01/1961-02. For spd2, a 2218 bp and a 2756 bp PCR fragment were generated with primers 1962-01/1962-02 and 1962-03/1962-04, respectively. For spd3, a 1152 bp PCR fragment was generated with primers 1963-01/1963-02. The PCR fragments for each gene were transformed into yeast together with HindIII-digested pDS23 (Schindler and Nowrousian 2014) as recipient vector.
Vector pSPD4_OE was generated by amplification of the complete spd4 ORF using primer pair 1964-09/1964-10, sub-cloning into pDrive, and ligation of the NotI and SpeI cut 2032 bp fragment into NotI and SpeI sites of pEHN1nat, carrying the A. nidulans gpd promoter and trpC terminator and the nat1 resistance gene (Dreyer et al. 2007). Vector pSPD4_NA, carrying spd4 together with native upstream and downstream regions, was generated by yeast recombination of PvuII-digested pDS23 (Schindler and Nowrousian 2014) and three PCR fragments amplified from S. macrospora genomic DNA (1964-11/spd_13, 1970 bp; 1964-12/1964-13, 1380 bp) and pSPD4_OE (1964-09/1964-10, 2029 bp).
For localization of SPD4, GFP fusions were generated as follows: Vector pDS23 was linearized with NotI and BglII and transformed into yeast together with an NotI/BglII digested egfp fragment from pDS23, and two PCR fragments amplified from S. macrospora genomic DNA (1964-11/1964P-GFP, 1857 bp; GFP-1964T/1964-13, 1360 bp) to generate pSPD4PT-GFP. N- and C-terminally egfp-fused spd4 constructs pGFP-SPD4 and pSPD4-GFP were generated by yeast recombination using BglII and NotI-linearized pSPD4PT-GFP, respectively, and PCR fragments amplified from S. macrospora genomic DNA (GFP-1964/spd_33 and spd_14/1964-GFP, respectively).
Microscopy
Microscopic investigations were carried out with an AxioImager M.1 microscope (Zeiss) equipped with a CoolSnap HQ camera (Roper Scientific) and a SpectraX LED lamp (Lumencor). GFP fluorescence was analyzed using filter set 41017 (Chroma Technology; HQ470/40, HQ525/50, Q495lp). To analyze sexual development over time, strains were grown on BMM-covered slides (Engh et al. 2007) for two to seven days. For detection of VHF, strains were grown on cellophane-covered MMS plates (Rech et al. 2007) for two days, and pieces of cellophane were cut and used for microscopy. Images were taken and edited with MetaMorph (version 7.7.0.0; Universal Imaging).
Perithecia formation was assayed on BMM plates after 7 days of growth using a Stemi 2000-C stereomicroscope (Zeiss) equipped with a AxioCamERc5 s digital camera (Zeiss) and AxioVision software (Zeiss). Images were processed with Adobe Creative Suite 4 (Adobe Corp.).
Results
Sequencing of the spd genome reveals a 22 kb deletion
The S. macrospora spd mutant was previously shown to lack fruiting bodies and thus being sterile (Esser and Straub 1958; Lord and Read 2011), but the underlying genetic defect had not been identified yet. Here, we sequenced the genome of spd using a pipeline previously established for mutant genome sequencing in S. macrospora (Nowrousian et al. 2012) (Table S4). The mutant was crossed with spore color mutant fus. DNA from 40 sterile and 40 fertile progeny was pooled and subjected to whole-genome re-sequencing. Comparing the spd genome to the wild-type reference genome (Nowrousian et al. 2010; Teichert et al. 2012), we identified a missense mutation (G502A) in an open reading frame (ORF), SMAC_01935, resulting in amino-acid substitution D133N at the protein level. SMAC_01935 encodes a putative mitochondrial external NAD(P)H dehydrogenase. Further searches for regions not covered in the spd genome identified a 22 kb deletion (contig 2.5:1,274,642–1,296,071) comprising part of the SMAC_01961 ORF and the complete SMAC_01962, SMAC_01963, and SMAC_01964 ORFs (Fig. 1). All mutations are located on scaffold 2.5 of the S. macrospora reference genome. PCR and Southern analysis confirmed the point mutation and deletion in spd (Fig. 1), and the sterile phenotype co-segregated with the point mutation and the deletion in 40 progeny from a cross of spd and wildtype (Figs. S7 and S8). Primer walking analysis verified the genomic sequence of the deleted region.
Mutant spd shows a 22 kb deletion and a point mutation in SMAC_01935. a Schematic representation of the region affected by the deletion and point mutation in wildtype (wt) and spd. ORFs are indicated by arrows; Locus tag numbers are given below the arrows. B and X denote restriction sites for BamHI and XbaI, respectively. Probes and predicted signal sizes for Southern analysis are indicated. G502A gives the location of the point mutation in the SMAC_01935 ORF. b G502A substitution in SMAC_01935 in the spd mutant genome leads to a D133 N amino-acid substitution. c Southern analysis of spd and wildtype (wt) verifies the 22 kb deletion in the mutant. Probes were as indicated in (a)
We renamed SMAC_01961, SMAC_01962, SMAC_01963, and SMAC_01964 as spd1, spd2, spd3, and spd4, respectively. SPD1 is a homolog of A. nidulans E1 SumO activating enzyme AosA (Harting et al. 2013), SPD2 contains a deleted in azoospermia-associated protein 2 (DAZAP2) (Tsui et al. 2000) domain, SPD3 is a putative S-adenosylmethionine-dependent methyltransferase, and SPD4 contains a domain of unknown function.
Using data from a recent RNA-seq approach (Teichert et al. 2012), we investigated the transcriptional expression of spd1-4. Samples were obtained from vegetative and sexual mycelia, as well as from protoperithecia. For technical reasons, reads from protoperithecia samples tended to map preferentially to the 3′ end of the mRNA, and often could not be mapped correctly, if the 3′ untranslated regions (UTRs) were not annotated. Therefore, we manually reannotated the UTRs of the spd genes. Compared with sexual mycelium, spd1 and spd4 were up-regulated in protoperithecia, whereas spd4 was down-regulated in vegetative mycelium compared to sexual mycelium (Fig. S9).
Several genes are responsible for the pleiotropic spd phenotype
We pursued several strategies to complement the sterile phenotype of spd. Transformation of spd with N. crassa and S. macrospora cosmids harboring the region deleted in spd whole or in part, S. macrospora PCR fragments covering different parts of the region deleted in spd, and five plasmids carrying the candidate genes expressed from the constitutive A. nidulans gpd promoter (Table S2) did not yield any fertile strains (Fig. S10). We thus individually deleted each of the genes affected by the 22 kb deletion in spd (Fig. 2a). We identified five to 12 strains for each deleted gene (Table S1), and strains having the same deletion exhibited the same phenotype. In contrast to the other deletion strains, Δspd4 is marker-free, because we employed a recently developed one-step FLP/FRT recombination system for its generation (Bloemendal et al. 2014).
Characterization of spd genes. a Schematic representation of the region affected by the spd deletion in wildtype (wt) and spd as well as four deletion strains each lacking one of the four spd genes. A boxed H indicates a hygromycin resistance cassette used to replace the particular spd gene. Δspd4 does not contain a resistance cassette due to marker recycling. b Phenotypic analysis of deletion strains and transformants with plasmids carrying spd4. Strains were grown on BMM fructification medium for 7 days. Mature perithecia are marked by arrowheads. Of the four deletion strains, only Δspd4 has a defect in perithecia formation. Introduction of the spd4 gene expressed from its native promoter (NA) or the A. nidulans gpd promoter (OE) restores perithecia formation in Δspd4, but not spd. The scale bar is 1 mm
The deletion strains were analyzed for sexual development after 7 days of growth. The wildtype formed black, pear-shaped fruiting bodies, and similar fruiting bodies were observed for Δspd1, Δspd2, and Δspd3 (Fig. 2b). However, like spd, Δspd4 exhibited no fruiting bodies (Fig. 2b). Thus, the spd4 deletion likely causes the protoperithecial arrest. For restoration experiments, we used plasmids pSPD4_OE and pSPD4_NA in which spd4 was under the control of A. nidulans gpd and the native promoter, respectively. Transformation of these plasmids into Δspd4 resulted in the restoration of fruiting body formation (Fig. 2b). However, the plasmids were unable to restore fertility in the spd mutant.
Our results indicate that additional genes are responsible for the pleiotropic spd phenotype. We hypothesized that the N-terminal part of the spd1 gene still present in the spd mutant (Fig. 2a) interfered with the complementation approach. We thus restored the complete spd1 ORF in the spd background (see Materials and Methods and Fig. S6 for details). These spd::spd1 strains still exhibited an spd-like sterile phenotype, but full fertility was regained after transformation of spd::spd1 with cosmid A12, carrying the entire region deleted in the spd mutant (Fig. S10). However, transformation of spd::spd1 with plasmids pSPD4_OE und pSPD4_NA did not result in perithecia formation, indicating that additional genomic regions deleted in spd are required for fruiting body formation. Interestingly, RNA-seq analysis revealed a strongly transcribed region in the intergenic region of spd2 and spd3 (Teichert et al. 2012). This region does not overlap any annotated features, and tblastx searches did not reveal homology to annotated fungal genes. Therefore, it might encode a non-protein coding RNA or a small peptide essential for sexual development. The SMAC_01935 mutation seems to be unrelated to the spd phenotype and was most probably carried along because of its close proximity to the 22 kb deletion.
Mutant Δspd4 is sterile and exhibits lysis of ascogonia and protoperithecia
Our deletion approach revealed that spd4 is required for fruiting body formation. Therefore, we performed a time-course analysis of Δspd4 sexual development compared to wildtype and spd. The wild-type formed ascogonia, protoperithecia, and perithecia within the predicted period of 7 days (Fig. 3a). As described previously (Lord and Read 2011), spd generated only aberrant ascogonia and small aberrant protoperithecia (Fig. 3a). We further noticed profound lysis of protoperithecia. The Δspd4 strain exhibited the same phenotype as the spd mutant. Wildtype-like fruiting body development was restored by transformation with pSPD4_OE and pSPD4_NA.
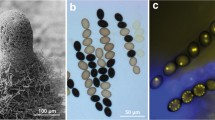
Phenotypic characterization of Δspd4. a Sexual development was assessed microscopically after 2 to 7 days of growth on BMM-covered microscope slides. The wildtype (wt) forms ascogonia (day 2), non-melanized (day 3), and melanized protoperithecia (day 4), and mature perithecia (day 7). Mutants spd and Δspd4 form only ascogonia and non-melanized protoperithecia-like structures that show profound lysis. Perithecia are never observed. Sterility can be complemented in Δspd4 by transformation with plasmids pSPD4_NA (strain M17) and pSPD4_OE (strain M847), expressing spd4 from native and the A. nidulans gpd upstream regions, respectively. The black scale bar is 20 µm, the white scale bar is 100 µm. b VHF in wildtype (wt) and Δspd4. Strains were analyzed after 2 days of growth on cellophane-covered MMS. Arrowheads indicate fusion bridges. The scale bar is 20 µm
We further analyzed vegetative growth of Δspd4 and wildtype on SWG plates. Growth of both strains was comparable, with 36.7 ± 1.1 and 37.1 ± 0.5 mm/day for wildtype and Δspd4, respectively. As mentioned above, most S. macrospora mutants with developmental arrest at the protoperithecia stage have a defect in VHF. However, Δspd4 is still capable of VHF, as is mutant spd (Figs. 3b and S11).
SPD4 localizes to the nucleus
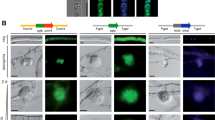
Spd4 is conserved in ascomycetes, but its function has not yet been elucidated. To gain insight into SPD4 function, we performed in silico analysis of the predicted polypeptide using the ELM resource (Dinkel et al. 2016), revealing a nuclear localization signal (NLS), a nuclear export signal (NES), and a domain of unknown function (DUF2841). This domain structure is similar to the uncharacterized yeast homolog YDR124W (Fig. 4a). We performed localization studies with SPD4 containing an N-terminal (pGFP-SPD4) or C-terminal (pSPD4-GFP) tag. Both constructs were able to restore fertility in Δspd4 (Fig. 4b). GFP-tagged SPD4 localized to spherical structures that stained with the nuclear dye NucBlue (Fig. 4c). This nuclear localization was consistent in hyphal tips and hyphae from the colony periphery (Fig. 4d). When analyzing older regions of mycelial colonies, we found that nuclear fluorescence accumulated in ascogonial coils (Fig. 4e) and was restricted to ascogonial tissues in the developing protoperithecium (Fig. 4f). Fluorescence was not observed in melanized protoperithecia, which may be due to either pigment accumulation or signal dissipation (Fig. 4g).
Localization of SPD4. a Domain structure of SPD4 and its yeast homolog. Protein domains predicted in silico are shown in different colors. NLS, nuclear localization signal; NES, nuclear export sequence; DUF, domain of unknown function. b N-terminally (M1444) and C-terminally (M1238) GFP-tagged versions of SPD4 are able to restore fertility in Δspd4. Scale bar is 1 mm. c GFP-tagged SPD4 (Δspd4 + pGFP-SPD4) localizes to spherical structures that are identified as nuclei by staining with the nuclear dye NucBlue. GFP-SPD4 localizes to nuclei in hyphal tips (apical region) and vegetative hyphae from the peripheral region (d) as well as ascogonial coils (e), but not in peridial tissue of non-melanized (f) and melanized protoperithecia (g). Strains carrying EGFP alone (wt + pDS23) or EYFP-tagged histone H2A (wt + pYH2A) are shown for comparison in (d-g). In these strains, cytoplasmic and nuclear fluorescence, respectively, appears evenly distributed through all cells. The scale bar is 10 µm in (c-g)
As a control for cellular localization, we analyzed two strains with cytoplasmic EGFP and nuclear EYFP fluorescence. Strain S106352 (Teichert et al. 2012) expresses egfp from the A. nidulans gpd promoter, which led to cytoplasmic fluorescence in vegetative hyphae and evenly distributed fluorescence in different stages of fruiting body development (Fig. 4d–g). The nuclear fluorescence by strain S107299 carrying the H2A-EYFP fusion protein (Rech et al. 2007) was evenly distributed throughout hyphae, ascogonia, and protoperithecia (Fig. 4d–g). Therefore, during sexual development, SPD4 specifically localizes to nuclei in ascogonial cells.
Discussion
In this study, we identified the developmental gene spd4 required for fruiting body development by genome sequencing of the sterile spd mutant. While restoration of sexual development in spd requires additional genomic regions besides spd4, the spd4 deletion mimics the fruiting body defect of spd and thus likely causes the developmental arrest in the mutant.
Expression studies have shown that spd4 transcript levels are up-regulated in protoperithecia compared with sexual mycelium (Teichert et al. 2012). In S. cerevisiae, expression of the uncharacterized gene YDR124W, which is homologous to spd4, has been shown to be induced by α factor mating pheromone (Harris et al. 2001). We found no regulation of spd4 in Δppg1 lacking the α-factor-like pheromone gene ppg1 (M. Lutomski and I. Teichert, unpublished), but the Δppg1 mutant from S. macrospora is fertile. Only strains lacking components of both pheromone/pheromone receptor pairs are sterile in S. macrospora (Mayrhofer et al. 2006).
We observed nuclear localization of SPD4 in vegetative hyphae and ascogonial coils, which is consistent with the presence of an NLS in SPD4. Interestingly, we did not detect SPD4 in peridial tissues. To the best of our knowledge, this is the first description of a protein that exhibits tissue-specific nuclear localization during fruiting body formation. It was suggested previously that distinct nuclear populations and developmental proteins are present in the fertile tissue inside the fruiting body (hymenium) and others in the peridium (Johnson 1976; Debuchy et al. 2010). For example, analysis of genetic mosaics of P. anserina indicated that PaNox-1 and IDC1, which is homologous to N. crassa ham-5 recently described to be a scaffold for the MAK-2 MAPK pathway during VHF (Dettmann et al. 2014; Jonkers et al. 2014), are required in the peridium, whereas thioredoxin genes are required in the hymenium (Jamet-Vierny et al. 2007; Malagnac et al. 2007). Localization of SPD4 to ascogonia may indicate that this protein is required in the hymenium. However, the mutant is blocked much earlier than the beginning of hymenium development, forming abnormal ascogonial coils and rudimentary protoperithecia. SPD4 may be required for proper ascogonium formation (Lord and Read 2011) or the ascogonium to protoperithecium transition, which requires the formation of enveloping hyphae that adhere to form the peridium (Kück et al. 2009; Debuchy et al. 2010; Lord and Read 2011). Ascogonium-derived signals have been suggested to be required for the formation of these hyphae (Bloemendal et al. 2010; Debuchy et al. 2010). Further studies have to clarify the nature of this signal and whether it is SPD4-dependent.
The question remains as to whether VHF is a prerequisite for fruiting body development or whether the finite number of signaling proteins in the fungal cell is just re-used for different signaling processes with additional factors conferring specificity to signaling outputs. A prominent example of a reusable signaling protein is the yeast MAPKKK STE11, which participates in three MAPK pathways: pheromone, pseudohyphal growth, and osmostress (reviewed by Saito 2010). In this case, signaling specificity is provided by diverse mechanisms, including scaffold proteins and cross-pathway inhibition, leading to interactions with different downstream effectors. A growing number of fungal proteins have been identified that regulate just one of the two processes, VHF or fruiting body formation. Similar to SPD4, S. macrospora autophagy proteins SmATG4, SmATG8, and SmJLB1, as well as N. crassa PRM-1, are essential for completion of the sexual cycle, but not VHF (Fleissner et al. 2009; Voigt et al. 2013; Voigt and Pöggeler 2013), though the underlying mechanisms may be different. Autophagy probably provides the fruiting body with energy and nutrients from the underlying mycelium (Khan et al. 2012). Similarly, peroxisomes mobilize reserve compounds and are necessary during fruiting body formation (Peraza-Reyes and Berteaux-Lecellier 2013). Interestingly, S. macrospora mutants lacking the transcription factor gene Smjlb1 form either protoperithecia or perithecia devoid of ascospores, depending on the complexity of the medium (Voigt et al. 2013). In contrast, the N. crassa prm-1 mutant is sterile in sexual crosses, because it lacks croziers, hook-shaped dikaryotic hyphal tips that give rise to ascus initials. PRM-1 has been shown to be involved in membrane merger after fusion, and the mutant exhibits a 50 % reduction in germling fusion and fusion of the conidium and trichogyne (a protoperithecial protrusion) (Fleissner et al. 2009). Therefore, PRM-1 and autophagy proteins may be required at a much later stage during sexual development than SPD4.
In summary, we identified SPD4 as a nuclear protein essential for sexual development. Though numerous developmental factors have already been identified, SPD4 is involved specifically in the early sexual fruiting body formation, but not VHF. Furthermore, SPD4 is restricted to ascogonial cells in the developing protoperithecium. Future studies should analyze its role as a determinant of cellular signaling specificity during the early sexual development, thereby enhancing our understanding of this reprogramming step during the fungal life cycle.
References
Aanen DK, Debets AJM, de Visser JAGM, Hoekstra RF (2008) The social evolution of somatic fusion. Bioessays 30:1193–1203
Arnaise S, Leblon G, Lares L (1984) A system for the detection of chromosomal rearrangements using Sordaria macrospora. Mutat Res 125:33–42
Becker Y, Eaton CJ, Brasell E, May KJ, Becker M, Hassing B, Cartwright GM, Reinhold L, Scott B (2015) The fungal cell-wall integrity MAPK cascade is crucial for hyphal network formation and maintenance of restrictive growth of Epichloe festucae in symbiosis with Lolium perenne. Mol Plant Microbe Interact 28:69–85
Bistis GN, Perkins DD, Read ND (2003) Different cell types in Neurospora crassa. Fungal Genet Newsl 50:17–19
Bloemendal S, Lord KM, Rech C, Hoff B, Engh I, Read ND, Kück U (2010) A mutant defective in sexual development produces aseptate ascogonia. Eukaryot Cell 9:1856–1866
Bloemendal S, Bernhards Y, Bartho K, Dettmann A, Voigt O, Teichert I, Seiler S, Wolters DA, Pöggeler S, Kück U (2012) A homolog of the human STRIPAK complex controls sexual development in fungi. Mol Microbiol 84:310–323
Bloemendal S, Löper D, Terfehr D, Kopke K, Kluge J, Teichert I, Kück U (2014) Tools for advanced and targeted genetic manipulation of the beta-lactam antibiotic producer Acremonium chrysogenum. J Biotechnol 169:51–62
Cai ZD, Chai YF, Zhang CY, Qiao WR, Sang H, Lu L (2015) The Gβ-like protein CpcB is required for hyphal growth, conidiophore morphology and pathogenicity in Aspergillus fumigatus. Fungal Genet Biol 81:120–131
Charlton ND, Shoji JY, Ghimire SR, Nakashima J, Craven KD (2012) Deletion of the fungal gene soft disrupts mutualistic symbiosis between the grass endophyte Epichloe festucae and the host plant. Eukaryot Cell 11:1463–1471
Chi MH, Craven KD (2016) RacA-mediated ROS signaling is required for polarized cell differentiation in conidiogenesis of Aspergillus fumigatus. PLoS One 11:e0149548
Christianson TW, Sikorski RS, Dante M, Shero JH, Hieter P (1992) Multifunctional yeast high-copy-number shuttle vectors. Gene 110:119–122
Colot HV, Park G, Turner GE, Ringelberg C, Crew CM, Litvinkova L, Weiss RL, Borkovich KA, Dunlap JC (2006) A high-throughput gene knockout procedure for Neurospora reveals functions for multiple transcription factors. Proc Natl Acad Sci USA 103:10352–10357
Coppin E, Berteaux-Lecellier V, Bidard F, Brun S, Ruprich-Robert G, Espagne E, Ait-Benkhali J, Goarin A, Nesseir A, Planamente S, Debuchy R, Silar P (2012) Systematic deletion of homeobox genes in Podospora anserina uncovers their roles in shaping the fruiting body. PLoS One 7:e37488
Debuchy R, Berteaux-Lecellier V, Silar P (2010) Mating systems and sexual morphogenesis in ascomycetes. In: Borkovich KA, Ebbole DJ (eds) Cellular and molecular biology of filamentous fungi. ASM Press, Washington, D.C., pp 501–535
Dettmann A, Heilig Y, Valerius O, Ludwig S, Seiler S (2014) Fungal communication requires the MAK-2 pathway elements STE-20 and RAS-2, the NRC-1 adapter STE-50 and the MAP kinase scaffold HAM-5. PLoS Genet 10:e1004762
Dinkel H, Van Roey K, Michael S, Kumar M, Uyar B, Altenberg B, Milchevskaya V, Schneider M, Kühn H, Behrendt A, Dahl SL, Damerell V, Diebel S, Kalman S, Klein S, Knudsen AC, Mäder C, Merrill S, Staudt A, Thiel V, Welti L, Davey NE, Diella F, Gibson TJ (2016) ELM 2016-data update and new functionality of the eukaryotic linear motif resource. Nucleic Acids Res 44:D294–D300
Dirschnabel DE, Nowrousian M, Cano-Dominguez N, Aguirre J, Teichert I, Kück U (2014) New insights into the roles of NADPH oxidases in sexual development and ascospore germination in Sordaria macrospora. Genetics 196:729–744
Dreyer J, Eichhorn H, Friedlin E, Kürnsteiner H, Kück U (2007) A homologue of the Aspergillus velvet gene regulates both cephalosporin C biosynthesis and hyphal fragmentation in Acremonium chrysogenum. Appl Environ Microbiol 73:3412–3422
Duan Y, Ge C, Liu S, Wang J, Zhou M (2013) A two-component histidine kinase Shk1 controls stress response, sclerotial formation and fungicide resistance in Sclerotinia sclerotiorum. Mol Plant Pathol 14:708–718
Engh I, Würtz C, Witzel-Schlömp K, Zhang HY, Hoff B, Nowrousian M, Rottensteiner H, Kück U (2007) The WW domain protein PRO40 is required for fungal fertility and associates with Woronin bodies. Eukaryot Cell 6:831–843
Esser K (1982) Cryptogams—cyanobacteria, algae, fungi, lichens. Cambridge University Press, London
Esser K, Straub J (1956) Fertility in the heterokaryon from two sterile mutants of Sordaria macrospora Auersw. Z Indukt Abstamm Vererb 87:625–626
Esser K, Straub J (1958) Genetische Untersuchungen an Sordaria macrospora Auersw., Kompensation und Induktion bei genbedingten Entwicklungsdefekten. Z Vererb 89:729–746
Fleißner A, Sarkar S, Jacobson DJ, Roca MG, Read ND, Glass NL (2005) The so locus is required for vegetative cell fusion and postfertilization events in Neurospora crassa. Eukaryot Cell 4:920–930
Fleissner A, Diamond S, Glass NL (2009) The Saccharomyces cerevisiae PRM1 homolog in Neurospora crassa is involved in vegetative and sexual cell fusion events but also has postfertilization functions. Genetics 181:497–510
Fu C, Iyer P, Herkal A, Abdullah J, Stout A, Free SJ (2011) Identification and characterization of genes required for cell-to-cell fusion in Neurospora crassa. Eukaryot Cell 10:1100–1109
Gull K (1978) Form and function of septa in filamentous fungi. In: Smith JE, Berry DR (eds) The filamentous fungi III. Developmental mycology. Wiley, New York, pp 78–93
Harris K, Lamson RE, Nelson B, Hughes TR, Marton MJ, Roberts CJ, Boone C, Pryciak PM (2001) Role of scaffolds in MAP kinase pathway specificity revealed by custom design of pathway-dedicated signaling proteins. Curr Biol 11:1815–1824
Harting R, Bayram O, Laubinger K, Valerius O, Braus GH (2013) Interplay of the fungal sumoylation network for control of multicellular development. Mol Microbiol 90:1125–1145
Hoff B, Kamerewerd J, Sigl C, Mitterbauer R, Zadra I, Kürnsteiner H, Kück U (2010) Two components of a velvet-like complex control hyphal morphogenesis, conidiophore development, and penicillin biosynthesis in Penicillium chrysogenum. Eukaryot Cell 9:1236–1250
James P, Halladay J, Craig EA (1996) Genomic libraries and a host strain designed for highly efficient two-hybrid selection in yeast. Genetics 144:1425–1436
Jamet-Vierny C, Debuchy R, Prigent M, Silar P (2007) IDC1, a pezizomycotina-specific gene that belongs to the PaMpk1 MAP kinase transduction cascade of the filamentous fungus Podospora anserina. Fungal Genet Biol 44:1219–1230
Jerpseth B, Greener A, Short JM, Viola J, Kretz PL (1992) XL1-Blue MRF` E. coli cells: mcrA-, mcrCB-, mcrF-, mmr-, hsdR- derivative of XL1-Blue cells. Strateg Mol Biol 5:81–83
Johnson TE (1976) Analysis of pattern formation in Neurospora perithecial development using genetic mosaics. Dev Biol 54:23–36
Jonkers W, Leeder AC, Ansong C, Wang Y, Yang F, Starr TL, Camp DG 2nd, Smith RD, Glass NL (2014) HAM-5 functions as a MAP kinase scaffold during cell fusion in Neurospora crassa. PLoS Genet 10:e1004783
Kamerewerd J, Jansson M, Nowrousian M, Pöggeler S, Kück U (2008) Three alpha-subunits of heterotrimeric G proteins and an adenylyl cyclase have distinct roles in fruiting body development in the homothallic fungus Sordaria macrospora. Genetics 180:191–206
Kayano Y, Tanaka A, Akano F, Scott B, Takemoto D (2013) Differential roles of NADPH oxidases and associated regulators in polarized growth, conidiation and hyphal fusion in the symbiotic fungus Epichloe festucae. Fungal Genet Biol 56:87–97
Khan IA, Lu JP, Liu XH, Rehman A, Lin FC (2012) Multifunction of autophagy-related genes in filamentous fungi. Microbiol Res 167:339–345
Kicka S, Bonnet C, Sobering AK, Ganesan LP, Silar P (2006) A mitotically inheritable unit containing a MAP kinase module. Proc Natl Acad Sci USA 103:13445–13450
Kim HR, Chae KS, Han KH, Han DM (2009) The nsdC gene encoding a putative C2H2-type transcription factor is a key activator of sexual development in Aspergillus nidulans. Genetics 182:771–783
Kim H, Wright SJ, Park G, Ouyang S, Krystofova S, Borkovich KA (2012) Roles for receptors, pheromones, G proteins, and mating type genes during sexual reproduction in Neurospora crassa. Genetics 190:1389–1404
Kück U, Pöggeler S, Nowrousian M, Nolting N, Engh I (2009) Sordaria macrospora, a model system for fungal development. In: Anke T, Weber D (eds) The Mycota XV. Springer, Heidelberg, pp 17–39
Kück U, Beier AM, Teichert I (2016) The composition and function of the striatin-interacting phosphatases and kinases (STRIPAK) complex in fungi. Fungal Genet Biol 90:31–38
Leeder AC, Jonkers W, Li J, Glass NL (2013) Early colony establishment in Neurospora crassa requires a MAP kinase regulatory network. Genetics 195:883–898
Lehr NA, Wang Z, Li N, Hewitt DA, Lopez-Giraldez F, Trail F, Townsend JP (2014) Gene expression differences among three Neurospora species reveal genes required for sexual reproduction in Neurospora crassa. PLoS One 9:e110398
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079
Lichius A (2010) Cell fusion in Neurospora crassa. PhD thesis, The University of Edinburgh, UK
Lichius A, Lord KM (2014) Chemoattractive Mechanisms in Filamentous Fungi. Open Mycol J 8:28–57
Lichius A, Lord KM, Jeffree CE, Oborny R, Boonyarungsrit P, Read ND (2012) Importance of MAP kinases during protoperithecial morphogenesis in Neurospora crassa. PLoS One 7:e42565
Lord KM, Read ND (2011) Perithecium morphogenesis in Sordaria macrospora. Fungal Genet Biol 48:388–399
Malagnac F, Lalucque H, Lepere G, Silar P (2004) Two NADPH oxidase isoforms are required for sexual reproduction and ascospore germination in the filamentous fungus Podospora anserina. Fungal Genet Biol 41:982–997
Malagnac F, Klapholz B, Silar P (2007) PaTrx1 and PaTrx3, two cytosolic thioredoxins of the filamentous ascomycete Podospora anserina involved in sexual development and cell degeneration. Eukaryot Cell 6:2323–2331
Mayrhofer S, Weber JM, Pöggeler S (2006) Pheromones and pheromone receptors are required for proper sexual development in the homothallic ascomycete Sordaria macrospora. Genetics 172:1521–1533
Nowrousian M, Cebula P (2005) The gene for a lectin-like protein is transcriptionally activated during sexual development, but is not essential for fruiting body formation in the filamentous fungus Sordaria macrospora. BMC Microbiol 5:64
Nowrousian M, Frank S, Koers S, Strauch P, Weitner T, Ringelberg C, Dunlap JC, Loros JJ, Kück U (2007) The novel ER membrane protein PRO41 is essential for sexual development in the filamentous fungus Sordaria macrospora. Mol Microbiol 64:923–937
Nowrousian M, Stajich JE, Chu M, Engh I, Espagne E, Halliday K, Kamerewerd J, Kempken F, Knab B, Kuo HC, Osiewacz HD, Pöggeler S, Read ND, Seiler S, Smith KM, Zickler D, Kück U, Freitag M (2010) De novo assembly of a 40 Mb eukaryotic genome from short sequence reads: Sordaria macrospora, a model organism for fungal morphogenesis. PLoS Genet 6:e1000891
Nowrousian M, Teichert I, Masloff S, Kück U (2012) Whole-genome sequencing of Sordaria macrospora mutants identifies developmental genes. G3 (Bethesda) 2:261–270
Peraza-Reyes L, Berteaux-Lecellier V (2013) Peroxisomes and sexual development in fungi. Front Physiol 4:244
Pöggeler S, Kück U (2006) Highly efficient generation of signal transduction knockout mutants using a fungal strain deficient in the mammalian ku70 ortholog. Gene 378:1–10
Pöggeler S, Nowrousian M, Kück U (2006) Fruiting-body development in ascomycetes. In: Kües U, Fischer R (eds) The Mycota I. Springer, Berlin, pp 325–355
Prados Rosales RC, Di Pietro A (2008) Vegetative hyphal fusion is not essential for plant infection by Fusarium oxysporum. Eukaryot Cell 7:162–171
Read ND, Lichius A, Shoji JY, Goryachev AB (2009) Self-signalling and self-fusion in filamentous fungi. Curr Opin Microbiol 12:608–615
Rech C, Engh I, Kück U (2007) Detection of hyphal fusion in filamentous fungi using differently fluorescence-labeled histones. Curr Genet 52:259–266
Saito H (2010) Regulation of cross-talk in yeast MAPK signaling pathways. Curr Opin Microbiol 13:677–683
Sambrook J, Russel DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Sarikaya-Bayram O, Bayram O, Feussner K, Kim JH, Kim HS, Kaever A, Feussner I, Chae KS, Han DM, Han KH, Braus GH (2014) Membrane-bound methyltransferase complex VapA-VipC-VapB guides epigenetic control of fungal development. Dev Cell 29:406–420
Schindler D, Nowrousian M (2014) The polyketide synthase gene pks4 is essential for sexual development and regulates fruiting body morphology in Sordaria macrospora. Fungal Genet Biol 68:48–59
Siegmund U, Marschall R, Tudzynski P (2015) BcNoxD, a putative ER protein, is a new component of the NADPH oxidase complex in Botrytis cinerea. Mol Microbiol 95:988–1005
Simonin AR, Rasmussen CG, Yang M, Glass NL (2010) Genes encoding a striatin-like protein (ham-3) and a forkhead associated protein (ham-4) are required for hyphal fusion in Neurospora crassa. Fungal Genet Biol 47:855–868
Simonin A, Palma-Guerrero J, Fricker M, Glass NL (2012) Physiological significance of network organization in fungi. Eukaryot Cell 11:1345–1352
Teichert I, Wolff G, Kück U, Nowrousian M (2012) Combining laser microdissection and RNA-seq to chart the transcriptional landscape of fungal development. BMC Genom 13:511
Teichert I, Nowrousian M, Pöggeler S, Kück U (2014a) The filamentous fungus Sordaria macrospora as a genetic model to study fruiting body development. Adv Genet 87:199–244
Teichert I, Steffens EK, Schnass N, Fränzel B, Krisp C, Wolters DA, Kück U (2014b) PRO40 is a scaffold protein of the cell wall integrity pathway, linking the MAP kinase module to the upstream activator protein kinase C. PLoS Genet 10:e1004582
Tong LC, Silar P, Lalucque H (2014) Genetic control of anastomosis in Podospora anserina. Fungal Genet Biol 70:94–103
Tsui S, Dai T, Roettger S, Schempp W, Salido EC, Yen PH (2000) Identification of two novel proteins that interact with germ-cell-specific RNA-binding proteins DAZ and DAZL1. Genomics 65:266–273
Turra D, El Ghalid M, Rossi F, Di Pietro A (2015) Fungal pathogen uses sex pheromone receptor for chemotropic sensing of host plant signals. Nature 527:521–524
Voigt O, Pöggeler S (2013) Autophagy genes Smatg8 and Smatg4 are required for fruiting-body development, vegetative growth and ascospore germination in the filamentous ascomycete Sordaria macrospora. Autophagy 9:33–49
Voigt O, Herzog B, Jakobshagen A, Pöggeler S (2013) bZIP transcription factor SmJLB1 regulates autophagy-related genes Smatg8 and Smatg4 and is required for fruiting-body development and vegetative growth in Sordaria macrospora. Fungal Genet Biol 61:50–60
Acknowledgments
This paper is dedicated to Karl Esser, who generated the spd mutant, on the occasion of his 92nd birthday. We thank Ingeborg Godehardt, Kerstin Kalkreuter, Regina Krampe, and Susanne Schlewinski for excellent technical assistance. We thank Gabriele Frenßen-Schenkel for graphical work. This work was funded by the Deutsche Forschungsgemeinschaft (KU517/12-2, KU 517/11-2, NO407/5-1).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study was funded by the Deutsche Forschungsgemeinschaft (KU517/12-2, KU 517/11-2, NO407/5-1).
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Teichert, I., Lutomski, M., Märker, R. et al. New insights from an old mutant: SPADIX4 governs fruiting body development but not hyphal fusion in Sordaria macrospora . Mol Genet Genomics 292, 93–104 (2017). https://doi.org/10.1007/s00438-016-1258-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-016-1258-0