Abstract
Histone modification regulates plant development events by epigenetically silencing or activating gene expression, and histone methylation is regulated by histone lysine methyltransferases (KMTs) and histone lysine demethylases (KDMs). The JmjC domain proteins, an important KDM family, erase methyl marks (CH3-) from histones and play key roles in maintaining homeostasis of histone methylation in vivo. Here, we analyzed 169 JmjC domain proteins from whole genomes of plants ranging from green alga to higher plants together with 36 from two animals (fruit fly and human). The plant JmjC domain proteins were divided into seven groups. Group-I KDM4/JHDM3 and Group-V JMJD6 were found in all the plant species and the other groups were detected mainly in vascular or seed plants. Group-I KDM4/JHDM3 was potentially associated with demethylation of H3K9me2/3, H3K27me2/3, and H3K36me1/2/3, Group-II KDM5A with H3K4me1/2/3, Group-III KDM5B with H3K4me1/2/3 and H3K9me1/2/3, Group-V JMJD6 with H3R2, H4R3, and hydroxylation of H4, and Group-VII KDM3/JHDM2 with H3K9me1/2/3. Group-IV/Group-VI JmjC domain-only A/B proteins were involved in hydroxylation and demethylation of unknown substrate sites. The binding sites for the cofactors Fe(II) and α-ketoglutarate in the JmjC domains also were analyzed. In the α-ketoglutarate binding sites, Thr/Phe/Ser and Lys were conserved and in the Fe(II) binding sites, two His and Glu/Asp were conserved. The results show that JmjC domain proteins are a conserved family in which domain organization and cofactor binding sites have been modified in some species. Our results provide insights into KDM evolution and lay a foundation for functional characterization of KDMs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Epigenetics is the term used to describe heritable changes in gene function that are not caused by changes in the DNA sequence (Wu and Morris 2001; Dupont et al. 2009). Histones (H3, H4, H2A, H2B) form the core of the nucleosome that is the fundamental unit of chromatin. Over 60 different histone residues that can be modified by at least eight distinct types have been reported (Kouzarides 2007; Shahbazian and Grunstein 2007; Lennartsson and Ekwall 2009; Banerjee and Chakravarti 2011; Bannister and Kouzarides 2011). Histone modification, one of the main epigenetic regulatory mechanisms, plays critical roles in various biological processes, including regulation of chromatin structure dynamics and gene expression at the whole-genome level (van Leeuwen and Gottschling 2002; Kouzarides 2007; Gelato and Fischle 2008).
Histone lysine methylation status is controlled dynamically by histone lysine methyltransferases (KMTs) and histone lysine demethylases (KDMs). H3K4 or H3K36 methylation is generally associated with active transcription, whereas H3K9 or H3K27 methylation is associated with gene silencing (Pontvianne et al. 2010; Binda 2013). KMTs catalyze mono-, di-, or trimethylation of lysine residues, and these modifications are involved in most plant development processes (Pontvianne et al. 2010; Thorstensen et al. 2011; Ay et al. 2014).
Histone lysine methylation is a reversible chromatin mark (Mosammaparast and Shi 2010) and to balance the methylation level of histone lysines, distinct histone demethylases are required (Chen et al. 2011). Bacteria and human lysine-specific demethylase 1 (LSD1) was the first KDM reported to specifically target histone H3 lysine 4 (H3K4) and histone H3 lysine 9 (H3K9) (Shi et al. 2004; Metzger et al. 2005). Arabidopsis LDL1 and LDL2 are homologs of LSD1 and they have been found to be involved in flowering time control (Jiang et al. 2007). The JmjC (Jumonji C) family of KDMs is involved in histone lysine or arginine demethylation through an oxidative reaction (Tsukada et al. 2006; Rotili and Mai 2011). A JmjC domain protein was first reported by Takeuchi and colleagues (1995); now there are more than 25,000 JmjC domain protein sequences in UniProt (http://www.ebi.ac.uk/uniprot) and SMART (http://smart.embl-heidelberg.de/). For their activity, JmjC domain proteins require iron Fe(II) and α-ketoglutarate (αKG) as cofactors. The JmjC domain has eight β-sheets, three conserved amino-acid residues in the binding site of Fe(II), and two residues in the binding site of αKG (Klose et al. 2006; Lan et al. 2008; Lu et al. 2008) also are important for Fe(II) binding. Thr185/Phe185 and Lys206 are required for αKG binding (Chen et al. 2006; Klose et al. 2006; Lu et al. 2008). JmjC domain proteins often harbor a JmjN (Jumonji N) domain, which as an integral core of all KDM catalytic domains (Pilka et al. 2015).The JmjC domain histone demethylases (JHDMs) can remove mono-, di-, and tri-methylation of histone lysine (Klose et al. 2006) and different KDMs exhibit different substrate specificities: KDM2A/JHDM1A are specific for H3K36me1/2, KDM5/JARID1 for H3K4me1/2/3, KDM4/JHDM3 for H3K9me2/3 and H3K36me2/3, KDM6 for H3K27me2/3, and KDM3/JHDM2 for H3K9me1/2 (Klose et al. 2006; Allis et al. 2007; Lu et al. 2008; Hou and Yu 2010). JmjC domain proteins play important roles in epigenetic processes, gene expression, and plant development by their involvement in the interplaying between histone modifications and DNA methylation (Lu et al. 2008; Saze et al. 2008; Chen et al. 2011; Luo et al. 2014).
Previously, JmjC domain proteins have been classified into eight groups in animals and five groups in plants (Lu et al. 2008). Recently, Luo et al. (2014) showed that Arabidopsis JmjC domain proteins could be divided into five groups: KDM4/JHDM3 group (AtJMJ11-13), KDM5/JARID1 group (AtJMJ14-19), JMJD6 group (AtJMJ21/22), KDM3/JHDM2 group (AtJMJ24-29), and JmjC domain-only group (AtJMJ20 and AtJMJ30-32). The functions and substrate specificities of each group also have been suggested (Luo et al. 2014). AtJmjC11/AtJMJ11/ELF6 (EARLY FLOWERING 6) was reported to act as a repressor in the photoperiod pathway by erasing H3K4 methylation deposited in FT (Flowering Locus T) chromatin thereby preventing precocious flowering (Noh et al. 2004; Jeong et al. 2009). AtJmjC12/AtJMJ12/REF6 (RELATIVE OF EARLY FLOWERING 6) is a FLC (Flowering Locus C) repressor by demethylating H3K27me2/3 (Noh et al. 2004; Lu et al. 2011a). Additionally, mutations in AtJMJ11 or AtJMJ12 were found to lead to brassinosteroid-related phenotypes and increased H3K9me3 levels (Yu et al. 2008). AtJMJ13 could act redundantly with REF6 (Holec and Berger 2012). AtJMJ14 was shown to act downstream of the Argonaute effector complex to demethylate histone H3K4 at the target of RNA silencing and repress flowering (Searle et al. 2010; Yang et al. 2010). In contrast, AtJMJ15 and AtJMJ18 were reported to accelerate flowering by demethylating H3K4me3/H3K4me2 at FLC (Yang et al. 2012a, b). AtJMJ30 may be a novel clock component involved in controlling the circadian period (Lu et al. 2011b). In rice, 20 JmjC domain proteins have been identified (Lu et al. 2011a). OsJMJ703 was shown to regulate stem elongation and plant growth by demethylating H3K4me1/me2/me3 (Chen et al. 2013), OsJMJ705 was found to function as an H3K27me2/3 demethylase involved in response to pathogen infection (Li et al. 2013), and OsJMJ706 was reported to encode a heterochromatin-associated H3K9 demethylase involved in the regulation of flower development (Sun and Zhou 2008). In addition, BcJMJ30 (Brassica campestris Jumonji 30) was implicated in pollen development and fertilization (Li et al. 2012). Therefore, it is well established that JmjC proteins play crucial roles in various plant species and developmental processes.
Only a few plant JmjC domain proteins have been analyzed at the whole-genome level (Lu et al. 2008). Zhou and Ma (2008) analyzed the distinct evolutionary patterns of histone demethylases in some lineages, but several representative species, such as the Selaginella, Ostreococcus, and gymnosperms, were not included in their study; moreover, the enzymatic activity of each subfamily remained unclear. Here we identified and analyzed JmjC domain proteins from representative green lineages. The results provide insights into the evolutionary conservation and diversification of JmjC domain proteins and putative catalytic targets, which may contribute to the future functional characterization of these important epigenetic regulators in plants.
Materials and methods
Identification of JmjC domain proteins
JmjC domain protein sequences from Arabidopsis thaliana, Populus trichocarpa, Brachypodium distachyon, Oryza sativa, Selaginella moellendorffii, Physcomitrella patens, Homo sapiens, and Drosophila melanogaster were retrieved from Chromdb database (http://www.chromdb.org/). Arabidopsis AtJMJ11-30 were used as queries to blastp against official websites of Brassica rapa (http://brassicadb.org/brad/, The syntenic genes between Arabidopsis and cabbage are also detected), Picea abies (http://congenie.org/) and Ostreococcus tauri or Volvox carteri (http://genome.jgi-psf.org).
Domain organization analysis
The protein sequences were analyzed for domain organization using NCBI-CD searches (http://ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi). The low-complexity filter was turned off, and the Expect Value was set at 10 to detect short domains or regions of less conservation in this analysis. Domains were also verified and named according to the SMART database (http://smart.embl-heidelberg.de/). Proteins with JmjC domain were used to construct phylogenetic trees. And sequences of JmjC domain were used to align and logos analysis.
Phylogenetic and sequence alignment analysis
Multiple sequence alignments were performed using the ClustalW program (Thompson et al. 1994). The resulting file was subjected to phylogenic analysis using the MEGA 6.0 program (Tamura et al. 2013). The trees constructed setting was depended on proteins sequences or JmjC domain sequences. JmjC domains of each group were aligned on line (https://npsa-prabi.ibcp.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_clustalw.html), and logos analysis was carried out (http://weblogo.berkeley.edu/logo.cgi).
Results
Identification and phylogenetic analysis of JmjC domain proteins
We screened 29 JmjC domain proteins in B. rapa, 25 in P. trichocarpa, 21 in A. thaliana, 20 in O. sativa, 19 in B. distachyon, 16 in P. abies, 14 in S. moellendorffii/P. patens, six in O. tauri, and five in V. carteri, together with 25 in H. sapiens and 11 in D. melanogaster. We have used the nomenclature of lysine demethylases proposed by the Chromdb database (JMJ, http://www.chromdb.org/index.html) (Online Resource 1).
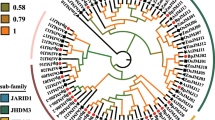
Our phylogenetic analysis based on 169 JmjC domain proteins in 10 plants and 36 in fruit fly and human showed that the plant JmjC domain proteins could be divided into seven groups rather than the five groups reported in a recent study (Luo et al. 2014). Group-III KDM5B proteins were separated from Group-KDM5/JARID1 proteins based on their domain organization. Group-IV JmjC domain-only A was separated from Group-JmjC domain-only and was closely related to Group-JMJD6 whose homologs shared a similar domain organization with JmjC domain-only (Fig. 1). The origin and substrate targets for each group were inferred based on the phylogenetic tree and the functions were inferred from the annotated Arabidopsis and human KDMs (Table 1).
Phylogenetic tree of JmjC domain proteins based on sequences of JmjC domain. This tree concludes 169 JmjC domain proteins from P. trichocarpa, A. thaliana, B. rapa, O. sativa, B. distachyon, S. moellendorffii, P. patens, V. carteri, and O. tauri genomes, and 36 from animals. The JmjC domain proteins can be grouped into 7 groups based on the phylogenetic tree and domain organization. Square shows JmjC domain proteins from model plant Arabidopsis, triangle show plant JmjC domain proteins homologs from human and fruit fly and diamond show animal JmjC domain proteins from human and fruit fly. Different colors show different groups. The proteins named according to web (http://www.chromdb.org/index.html). JmjC domain protein sequences were aligned using ClustalW, and the phylogenetic tree analysis was performed using MEGA6. The trees were constructed with the following settings: tree inference as neighbor-joining; include sites as pairwise deletion option for total sequences analysis; Substitution model: p-distance; and Bootstrap test of 1000 replicates for internal branch reliability
Group-I KDM4/JHDM3
Group-I included three Arabidopsis (AtJMJ11-13), six P. trichocarpa, five O. sativa, four B. distachyon/P. abies/S. moellendorffii/human, three B. rapa, three V. carteri, two P. patens/fruit fly, and one O. tauri JmjC domain proteins (Fig. 2a; Online Resource 1). It is remarkable that some genes were doubled in poplar and rice but not in cabbage even though whole genome duplication has occurred after cabbage and Arabidopsis divergence (Wang et al. 2011). The Group-I KDM4/JHDM3 members have been predicted to act as H3K9me2/3, H3K27me2/3, and H3K36me2/3 demethylases and some of them play roles in flowering time control (Noh et al. 2004; Jeong et al. 2009; Yang et al. 2010; Searle et al. 2010; Lu et al. 2011a; Luo et al. 2014). KDM4/JMJD2 was shown to epigenetically control herpes virus infection and reactivation from latency, but whether it is involved in the plant defense system remains unknown (Liang et al. 2013). JmjC, JmjN, and C2H2/C5H2 are the domains specific to this group. The JmjN domain and its interaction with the JmjC catalytic domain were found to be important for Jhd2 (also known as KDM5), a H3K4-specific demethylase in budding yeast (Huang et al. 2010; Quan et al. 2011). C2H2/C5H2 are small DNA-binding peptide motifs, which suggests that Group-I KDM4/JHDM3 might bind directly to a specific DNA sequence (Klug 1999; Lu et al. 2008).
Phylogenetic and domain analysis of Group-I KDM4/JHDM3 based on full-length sequences. Group-I KDM4/JHDM3 includes AtJMJ11, AtJMJ12, AtJMJ13 and their homologues in all organisms and can be divided into 3 sub-groups. Subgroup-I and II character JmjC, JmjN and C2H2 domains while subgroup-III characters JmjC, JmjN and C5H2 domains. Domains came from SMART V 6.0 database except C5H2 zinc finger from CDD V3.11 database. a Phylogeny and domain organization. b Logos analysis of JmjC domain. Fe(II) binding site are showed in red triangle and those in the αKG binding site are showed in black triangle. The trees were constructed with the following settings: Tree inference as neighbor-joining; include sites as complete deletion option for total sequences analysis; substitution model: Poisson model; and Bootstrap test of 1000 replicates for internal branch reliability
The Group-I JmjC proteins were divided into three subgroups based on their phylogenetic relationship and domain organization. Subgroups-I and II contained AtJMJ11 and AtJMJ12 plant homologs, respectively. Subgroup-I proteins were found only in angiosperms, while Subgroup-II proteins were found in higher plants. The domain organization of the Subgroup-I/II proteins was highly conserved, except that the B. distachyon protein BdJMJ11301 had an additional Cdt1_m domain. Subgroup-III contained AtJMJ13 homologs that were found in all the plants and animals studied. The C5H2 zinc finger was specific to this group (Fig. 2a). The C2H2/C5H2 domains were absent in SmJMJ1605, VcJMJ6402/640, and OtJMJ3606, indicating that these proteins may bind indirectly to a specific chromatin region by interacting with other factors.
Sequence alignment and logos analysis of the JmjC domain showed that the αKG and Fe(II) binding sites were highly conserved in the group; however, some proteins from pine, Lycophyta, and V. carteri were the exceptions. For example, in the αKG binding site of SmJMJ1601, Phe was replaced by Trp, and in the Fe(II) binding site of SmJMJ1608, His was replaced by Arg, whereas in VcJMJ6401, Phe and Lys in the αKG binding site were replaced by Thr and Arg, respectively, and His was replaced by Leu in the Fe(II) binding site (Fig. 2b; Online Resource 2).The conserved binding sites suggest that most members of Group-I may be active histone demethylases that can target tri- and di-methylated H3K9 and H3K36 (Lu et al. 2011b). AtJMJ11, which has conserved binding sites for αKG and Fe(II), was shown to have H3K27 demethylase activity (Lu et al. 2011a).
Group-II KDM5A
The phylogenetic analysis showed that KDM5/JARID1 could be divided into two groups, Group-II KDM5A and Group-III KDM5B (Fig. 1), even though they have similar substrates. Group-II KDM5A included five A. thaliana (AtJMJ14/16/18/19), nine B. rapa, five P. trichocarpa, two O. sativa/B. distachyon/P. abies/S. moellendorffii, and one P. patens JmjC domain proteins. No Group-II KDM5A proteins were found in green alga and animals (Fig. 3a; Online Resource 1). The KDM5/JARID1 proteins in animals belonged to Group-III KDM5B. KDM5/JARID1 proteins are histone demethylases of H3K4me1/2/3. Animal JMRID1 was reported to be involved in cell fate determinacy (Benevolenskaya 2007; Secombe and Eisenman 2007). In plants, the KDM5/JARID1 proteins were found to regulate flowering time, AtJMJ14 was shown to promote flowering by activating FT, the solute carrier SOC1, the floral homeotic protein AP1, and LEAFY (LFY), while AtJMJ15 promoted flowering by reducing the H3K4me3 of FLC (Lu et al. 2010; Searle et al. 2010; Yang et al. 2010, 2012a). Moreover, overexpression of the H3K4 demethylase gene AtJMJ15 was found to enhance salt tolerance in Arabidopsis (Shen et al. 2014). AtJMJ18 displayed demethylase activity toward H3K4me3 and H3K4me2 of FLC and accelerated flowering (Yang et al. 2012b).
Phylogenetic and domain analysis of Group-II KDM5A based on full-length sequences. Group-II includes AtJMJ14-16, AtJMJ18/19 and their homologues in vascular plants and can be divided into 3 sub-groups. Subgroup-I and II character JmjC, JmjN and FYRN-FYRC domains while subgroup-III characters only JmjC, JmjN domains. a Phylogeny and domain organization. b Logos analysis of JmjC domain. Fe(II) binding site are showed in red triangle and those in the αKG binding site are showed in black triangle. The trees were constructed with the following settings: tree inference as neighbor-joining; include sites as complete deletion option for total sequences analysis; substitution model: p-distance; and Bootstrap test of 1000 replicates for internal branch reliability
The FYRN (FY-rich domain N-terminal) and FYRC (FY-rich domain C-terminal) domains, which are founded in a variety of chromatin-associated proteins and are particularly common in histone H3K4 methyltransferases (García-Alai et al. 2010), were specific to proteins in Group-II KDM5A; AtJMJ19 homologs were an exception (Fig. 3a). Group-II KDM5A could be divided into three subgroups (Fig. 3). Subgroup-I contained the AtJMJ14/15/18 homologs in higher plants and had highly conserved domain organization. The exceptions were AtJMJ15, which had additional FDH-GDH (formate dehydrogenase-glycerate dehydrogenase) and PMEI (plant invertase/pectin methylesterase inhibitor) domains, AtJMJ18, which had a Knot1 domain (knottins), BrJMJ10829 (Bra039869), which had a PMEI domain, and BrJMJ10815 (Bra021937), which had a PU domain (furin-like repeats). PMEI inhibits pectin methylesterase and invertase activity by forming a non-covalent complex and functions in the regulation of wounding, osmotic stress, senescence, seed development, and plant defense mechanisms (Hong et al. 2010). In this subgroup, the KDM5A proteins of cabbage had the most copies. We found that BrJMJ10829 (Bra039869) and BrJMJ10816 (Bra023136) were syntenic genes of AtJMJ18, indicating a onetime duplication event may have occurred. Additionally, BrJMJ10815 (Bra021937), but not BrJMJ10811 (Bra017281), was found to be a syntenic gene of AtJMJ15, implying that BrJMJ10811 may be a new original gene. Subgroup-II contained AtJMJ16 homologs in flowering plants, and all the genes were duplicated except in the monocots. Two copies of AtJMJ16 were found in cabbage, and BrJMJ10820 (Bra030735) with conserved length and domain organization was syntenic to AtJMJ14, whereas BrJMJ10814 (Bra018612) was not. The BrJMJ10814 sequence was much longer than the other sequences in this subgroup and had additional HELICc (helicase superfamily C-terminal) and DEXDc (DEAD-like helicases superfamily) domains, suggesting that it may exhibit additional helicase activity. Subgroup-III was only found in dicots. The protein sequences in this subgroup were much shorter than the other sequences in Group-II and contained only JmjC and JmjN domains. Cabbage had two syntenic genes of AtJMJ19, BrJMJ10801 (Bra000108) and BrJMJ10804 (Bra005068).
Sequence alignment and logos analysis showed that the JmjC domains were highly conserved in Subgroups-I and II, while in Subgroup-III the binding sites for αKG and Fe(II) were variable. Phe and Lys were replaced by Gln and Arg in the αKG binding site, and two His residues were replaced by Lys and Tyr in the for Fe(II) binding site in the Subgroup-III proteins. PtJMJ910 in Subgroup-III was an exception and was similar to proteins in Subgroups-I and II, that is, it had Phe and Lys in the αKG binding site and two His residues in the Fe(II) binding site (Fig. 3b; Online Resource 3).
Group-III KDM5B
The phylogenetic tree and domain organization showed that KDM5A/5B belonged to a branch, but had different domain organization (Figs. 1, 3, 4). Group-III KDM5B contained one member from all the studied plants except for pine and V. carteri (Fig. 4a; Online Resource 1). It was surprising that Group-II KDM5A included the pine KDMs, while the alga and animal KDMs were absent. Group-III KDM5B contained all the plant and animal KDMs except those of pine and V. carteri. AtJMJ17 was shown to have H3K4me1/2/3 and H3K9me1/2/3 demethylase activity and was reported to be involved in the plant’s defense system against virulent pathogens (Lu et al. 2008; Erickson 2012).
Phylogenetic and domain analysis of Group-III KDM5B based on full-length sequences. Group-III includes AtJMJ17 and their homologues in all organisms except pine and V.carteri. JmjC, JmjN, PHD, and BRIGHT are the special domains. a Phylogeny and domain organization. b Logos analysis of JmjC domain. Fe(II) binding site are showed in red triangle and those in the αKG binding site are showed in black triangle. The trees were constructed with the following settings: tree inference as neighbor-joining; include sites as complete deletion option for total sequences analysis; substitution model: no. of differences; and Bootstrap test of 1000 replicates for internal branch reliability
Unlike the Group-II KDM5A proteins, the Group-III KDM5B proteins contain PHD (plant homeodomain) and BRIGHT domains that are specific to them (Fig. 4a). BRIGHT, also called ARID (AT-rich interaction domain), is a helix-turn-helix motif-based DNA-binding domain that is conserved in a wide variety of species and plays important roles in development, tissue-specific gene expression, and proliferation control (Kim et al. 2004; Patsialou et al. 2005). The PHD finger domain is a C4HC3 zinc-finger-like motif that is found in nuclear proteins where it forms an integral part of the enzymatic core of the HAT (histone acetyltransferases) domain of CBP (CREB binding protein) and is involved in epigenetics and chromatin-mediated transcriptional regulation (Kalkhoven et al. 2002). SmJMJ1603 has an additional RRM (RNA recognition motif) domain that was found to be important for binding a large variety of RNA sequences and proteins (Maris et al. 2005). PpJMJ1504 and DmJMJ401 have an additional HMG17 (high mobility group HMG14 and HMG17) domain that was reported to facilitate transcription by repressing histone activity (Bustin et al. 1995). It should be noted that all these extra domains in these proteins were involved in binding to RNAs or proteins. HsJMJ503 and HsJMJ504 have a CHAD domain, but VcJMJ6402, which is much short, contained only the JmjC domain (Fig. 4a).
The sequences of the JmjC domain in Group-III proteins were highly conserved (Fig. 4b; Online Resource 4). Sequence alignment and logos analysis of the JmjC domain showed that the Group-III proteins had two conserved His residues and Glu in the Fe(II) binding site, and Phe and Lys in the αKG binding site, suggesting that all the proteins in this group may have similar function.
Group-IV JmjC domain-only A
In phylogenetic tree, Groups-IV, V and VI belonged to different branches, but cluster together in large clade (Fig. 1). The JmjC domain-only group included AtJMJ20 and AtJMJ30-32 and their homologs. Although most of them contained only the JmjC domain (Klose et al. 2006; Lu et al. 2008), the AtJMJ20 group belonged to a distant branch that was close to Group-V JMJD6 (Fig. 1), the same results were obtained even when different methods were used to construct the trees, suggesting that the origin of Group-IV JmjC domain-only A was different from that of Group-VI JmjC domain-only B. Group-IV JmjC domain-only A contained one member from all the plants except pine, moss, and green alga. All the proteins in this group only have a JmjC domain, except HsJMJ22, which had an additional Cupin_2 domain with unknown function (Fig. 5a). The HsJMJ522/JMJD4 proteins catalyze carbon 4 (C4) lysyl hydroxylation of the eukaryotic release factor eRF1 (Feng et al., 2014). AtJMJ20 may affect seed germination by removal of repressive histone arginine methylations at GA3ox1/GA3ox2 (Cho et al. 2012).
Phylogenetic and domain analysis of Group-IV JmjC domain-only A based on full-length sequences. Group-IV includes AtJMJ20 and their homologues in vascular plants except pine. a Phylogeny and domain organization. b Logos analysis of JmjC domain. Fe(II) binding site are showed in red triangle and those in the αKG binding site are showed in black triangle. The trees were constructed with the following settings: tree inference as neighbor-joining; include sites as pairwise deletion option for total sequences analysis; substitution model: Poisson model; and Bootstrap test of 1000 replicates for internal branch reliability
In the JmjC domain, the two His residues in the Fe(II) binding site and Lys in the αKG binding site were conserved, but Glu was replaced by Asp in the Fe(II) binding site, and Phe was replaced by Thr in the αKG binding site (Fig. 5b, Online Resource 5). In addition, it was suggested that the JmjC domain-only proteins may be active demethylases, but their substrates were not determined (Lu et al. 2008; Chen et al. 2011).
Group-V JMJD6
Based on the phylogenetic analysis, Group-V JMJD6 was close to Group-IV and contained AtJMJ21/22 homologs, two members in all the plants, except for three in P. patens and one in P. trichocarpa/O. tauri/fruit fly/human, indicating that no obvious gene duplication event had occurred (Figs. 1, 6a). Mouse JMJD6 was reported to show histone arginine demethylase activity of histone H3R2 and H4R3 (Chang et al. 2007). In addition, JMJD6 was found to be unable to remove the methyl group from histone arginine residues but it could hydroxylate the histone H4 tail at lysine residues in a 2-oxoglutarate (2-OG)- and Fe(II)-dependent manner (Han et al. 2012; Wang et al. 2014). It has been suggested that JMJD6 might associate preferentially with RNA/RNA complexes and to a lesser extent with chromatin (Hahn et al. 2010). AtJMJ22 was found to be involved in seed germination dependent on HYB activation (Cho et al. 2012).
Phylogenetic and domain analysis of Group-V JMJD6 based on full-length sequences. Group-V includes AtJMJ21/22 and their homologues in all organisms and can be divided into 2 sub-groups which character JmjC, FBOX and Cupin_2 domains. a Phylogeny and domain organization. b Logos analysis of JmjC domain. Fe(II) binding site are shown in red triangle and those in the αKG binding site are shown in black triangle. The trees were constructed with the following settings: tree inference as neighbor-joining; include sites as pairwise deletion option for total sequences analysis; substitution model: p-distance; and Bootstrap test of 1000 replicates for internal branch reliability
Most Group-V JMJD6 members have additional F-box and cupin_2 domains (Fig. 6a). The F-box domain, named based on its first identification at the N-terminal region of cyclin F, contains about 40–50 amino acids and has been shown to be required for protein–protein interactions involved in many processes (Craig and Tyers 1999; Wang et al. 2004; Jain et al. 2007). Group-V JMJD6 proteins were divided into two subgroups. Among Subgroup-I, OsJMJ711 had an additional Sec7 domain, which is a conserved elongated, all-helical sequence, and is required for proper protein transport through the Golgi (Mossessova et al. 1998; Cox et al. 2004). PpJMJ1506 had a hydrolase domain that is found in bacteria and eukaryotes and is approximately 110 amino acids long. In subgroup-II, monocots and green alga lacked the F-box and cupin_2 domains. SmJMJ1607 had an additional G2F domain (G2 fragment) that contained binding sites for collagen IV and perlecan, and PpJMJ1507 had an AXH (ataxin-1 and HBP1 module) domain, which is a protein–protein and RNA binding motif in ATX1 (ataxin-1) (de Chiara et al. 2003; Chen et al. 2004).
Similar to Group-IV, the JmjC domains of the Group-V proteins had the two conserved His residues and Aspin the Fe(II) binding sites, and Lys in the αKG binding sites, but the Phe/Thr sites varied (Fig. 6b; Online Resource 6).
Group-VI JmjC domain-only B
Group-VI JmjC domain-only B contained AtJMJ30-32 and their homologs, four in O. sativa/B. distachyon/S. moellendorffii, three in B. rapa/P. patens/O. tauri/human, two in P. trichocarpa/fruit fly, and one in P. abies, but none in V. carteri (Figs. 1 and 7a). The functions of this group of proteins are largely unclear. HsJMJ525/JMJD5 was proposed to be involved in embryonic cell proliferation acting as H3K36me2 histone demethylase (Ishimura et al. 2012; Zhu et al. 2014), AtJMJ30/JMJD5 was found to show hydroxylase activity, but no histone demethylation ability (Del Rizzo et al. 2012), and AtJMJ30 was ubiquitously expressed in different tissues and involved in Circadian clocks (Lu et al. 2008; Jones et al. 2010; Jones and Harmer. 2011; Lu et al. 2011a).
Phylogenetic and domain analysis of Group-VI JmjC domain-only B based on full-length sequences. Group-VI includes AtJMJ30-32 and their homologues in all plants and can be divided into 3 sub-groups. Subgroup-I characters JmjC and _2 domains, and Subgroup-II/III character JmjC domains. a Phylogeny and domain organization. b Logos analysis of JmjC domain. JmjC domains of AtJMJ31, OsJMJ713, BdJMJ11314, SmJMJ1602/10/14 are detected by SMART (http://smart.embl-heidelberg.de/) because them cannot be detected by NCBI. Fe(II) binding site are showed in red triangle and those in the αKG binding site are showed in black triangle. The trees were constructed with the following settings: tree inference as neighbor-joining; include sites as pairwise deletion option for total sequences analysis; substitution model: no. of differences; and bootstrap test of 1000 replicates for internal branch reliability
Group-VI was divided into three subgroups (Fig. 7a). Subgroup-I contained AtJMJ30 homologs, one homolog in dicot, Lycophyta, and moss, and two homologs in the monocots and O. tauri. A few of JMJC proteins in Subgroup-I had JmjC and cupin_2 domains. AtJMJ30 had an additional BRO1_Alix_like (protein-interacting Bro1-like domain of mammalian Alix and related domains) domain and BrJMJ10824 (Bra035742) had a TAP_C (TAP C-terminal) domain that was found to be important for binding to FG repeat-containing nuclear pore proteins (Bachi et al. 2000). Subgroup-II contained AtJMJ31 homologs, one homolog in each species, except for three homologs in Lycophyta and none in pine and green alga, and had a highly conserved domain organization. Subgroup-III proteins were found in the higher plants except Lycophyta, and BrJMJ10827 (Bra038271) had an additional PTX (pentraxins) domain that is found in CRP (C-reactive protein, also called PTX1) and SAP (serum amyloid P component), which has been regarded as innate antibodies (Lu et al. 2013a).
The amino acids in the Fe(II) binding sites and Lys in the αKG binding sites in the JmjC domain of this group were similar to those in the JmjC domain of Group-V. Glu was replaced by Asp, and Phe/Thr was replaced by Ser in the αKG binding sites in Subgroup-II HsJMJ525 and DmJMJ408 (Fig. 7b; Online Resource 7).
Group-VII KDM3/JHDM2
Group-VII KDM3/JHDM2 was the largest group and contained JmjC domain proteins that demethylate H3K9me2 and H3K9me1, but not H3K9me3, comprising six members in Arabidopsis (AtJMJ24-29) and their homologs, 10 in B. rapa, nine in P. trichocarpa, seven in P. abies, five in O. sativa/B. distachyon, four in P. patens and human, and one in fruit fly, but none in Lycophyta and green alga (Klose et al. 2006; Lu et al. 2008; Luo et al. 2014). Most of the known functions of KDM3 are attributed to the human homologs. For example, HsJMJ516/JMJD1A may regulate cell proliferation and renewal (Loh et al. 2007; Park et al. 2013), cell migration and invasion (Tee et al. 2014), and sex determination (Kuroki et al. 2013). HsJMJ518/JMJD1C was shown to regulate DNA repair through demethylating MDC1 (mediator of DNA damage checkpoint 1) (Lu and Matunis 2013b; Watanabe et al. 2013). AtJMJ24 was reported to interact with other JmjC and DCL proteins and increase root length, and cotyledon and floral organ size (Audonnet 2014). AtJMJ25/IBM1 was shown to be involved in leaf breadth, floral organ and embryo morphogenesis, pollen grain fertility, and seed reproduction, and was regulated by DDM1 (decrease in DNA methylation1) and KYP (kryptonite); it also activated RDR2 (RNA-dependent RNA polymerase2) and DCL3 (Dicer-like3) through H3K9me2 and DNA methylation (Saze et al. 2008; Fan et al. 2012).
Besides the JmjC domain, the RING (really interesting new gene) domain was the second main domain in this group, and the ZZ domain also was found in some members of Subgroups I–III. The RING domain consists of two zinc fingers with the sequence C-X2-C-X[9–39]-C-X[1–3]-H–X[2–3]-C-X2-C-X[4–48]-C-X2-C and it often functions in ubiquitin ligase activity (Lorick et al. 1999; Joazeiro and Weissman 2000; Smit et al. 2012). The ZZ domain binds two zinc ions and has the Cys-X2-Cys motif found in other zinc finger domains (Ponting et al. 1996; Legge et al. 2004). Group-VII was divided into four subgroups. Subgroup-I contains AtJMJ25/26/29 and their homologs in higher plants. Monocots had four copies of AtJMJ25, but no copies of AtJMJ26/29. The AT-hook has been identified in HMG (high mobility group) DNA-binding proteins from plants (Klosterman and Hadwiger 2002). PtJMJ916/918 had a B-box. Surprisingly, Subgroup-II contained AtJMJ27 and its three homologs in cabbage and seven in pine, but no copies in monocots and Lycophyta. Subgroup-III contained AtJMJ24 and its homologs in flowering plant, and Subgroup-IV contained only AtJMJ28 and its homologs in dicots. In many members of Subgroup-III/IV the JmjC domain was not detected by SMART (Fig. 8a). The JmjC domains proteins of animals with only JmjC domain belonged to the same branch on the phylogenetic tree (Fig. 8a). In cabbage, the AtJMJ28/29 homologs were diploid and the AtJMJ27 homologs were triploid (Fig. 8). BrJMJ10808 (Bra013461)/BrJMJ10828 (Bra038775) were syntenic genes of AtJMJ28/29, andBrJMJ10802 (Bra000935) and BrJMJ10826 (Bra037394) were syntenic genes of AtJMJ27, but BrJMJ10803 (Bra000936) was not.
Phylogenetic and domain analysis of Group-VII KDM3/JHDM2 based on full-length sequences. Group-VII includes AtJMJ24-29 and their homologues in seed plants and moss and can be divided into 4 sub-groups. JmjC and RING are the special domains. a Phylogeny and domain organization. b Logos analysis of JmjC domain. JmjC domains of PtJMJ913/914, AtJMJ28, BrJMJ10809/21/28, OsJMJ716, and BdJMJ11320 are detected by SMART (http://smart.embl-heidelberg.de/) because they cannot be detected by NCBI. Fe(II) binding site are showed in red triangle and those in the αKG binding site are showed in black triangle. The trees were constructed with the following settings: tree inference as neighbor-joining; include sites as partial deletion option for total sequences analysis; substitution model: poisson model; and bootstrap test of 1000 replicates for internal branch reliability
Sequence alignments and logos analysis of the JmjC domain showed that Thr and Lys in the αKG binding sites, and the two His residues and Asp in the Fe(II) binding sites were highly conserved in Subgroups-I/II. Lys in the αKG binding sites was conserved in Subgroups-III/IV, but the Phe and Thr varied and one of the His in the Fe(II) binding site was replaced by either Tyr of Phe (Fig. 8b; Online Resource 8).
Discussion
JmjC domain proteins constitute the largest histone demethylase family that contributes to histone methylation levels in vivo. Here, we screened 169 JmjC domain proteins from the whole genomes of green plants and 36 from animals (Fig. 1; Online Resource 1). The phylogenetic analysis showed that JmjC domain proteins were an ancient and conserved family that could be divided into seven groups, which displayed different histone demethylases activity in the green lineage. In all seven groups, the Fe(II) and αKG binding sites within the JmjC domain were highly conserved and when replacements occurred, most were by amino acid residues with similar properties.
Histone methylation mainly occurs in the Lys residues of histone H3 (K4, K9, K27, K36, K79) and H4 (K20), and are carried out by KMTs, except H3K79,which is methylated by the DOT1L (DOT1-Like) methyltransferase (Tschiersch et al. 1994; Jones et al. 2008; Nguyen and Zhang 2011). Previous study reported 37 KMTs in Arabidopsis,49 in cabbage, and 36 in rice and these proteins were involved in the methylation of H3K4, H3K9, H3K27, and H3K36 (Huang et al. 2011). Here, we found only 21 JmjC histone demethylases in Arabidopsis, 20 in rice, and 29 in cabbage, and these proteins targeted H3K4, H3K9, H3K27, and H3K36 (Fig. 1; Online Resource 1). The JmjC domain proteins have been divided previously into eight groups: five found in both plants and animals (KDM5/JARID1, KDM4/JHDM3, KDM3/JHDM2, JMJD6, and JmjC domain-only); and three found only in animals (KDM6/JMJD3, KDM2/JHDM1, and PHF) (Lu et al. 2008). We also did not find homologous proteins of KDM6/JMJD3 (HsJMJ14/15, DmJMJ405), KDM2/JHDM1(HsJMJ12/13, DmJMJ407) and PHF (HsJMJ10/11/24, DmJMJ408/409) in plants. Moreover, the animal JARID2 proteins (HsJMJ505, DmJMJ402) were distant from the homologous plant proteins (Fig. 1). KDM6/JMJD3 was reported to be involved in H3K27me3 (Pasini et al. 2010; Chen et al. 2012), and PHF8 was found to demethylate H4K20me1 in zebrafish brain and craniofacial development (Qi et al. 2010). KDM2/JHDM1 was found to be involved in H3K36me2 and JARID2 was reported to affect the H3K27me3 levels of target genes acting as an essential component of PcG (Polycomb group) complex (Tsukada et al. 2006; Pasini et al. 2010). We also noted that animal KDM2/JHDM1 and PHF were much closer to Group-VI JmjC domain-only B (Fig. 1). Generally, the total number of JmjC domain proteins increased with plant evolution; lower plants such as green alga had fewer JmjC domain proteins than the higher plants in our study (Fig. 1; Online Resource 1). Only six KDMs (possibly involved in K3K4, H3K9, and H3K36) were found in O. tauri. Overall the numbers of JmjC domain proteins were much less than the numbers of KMTs found in plants, suggesting that JmjC domain proteins might share a variety of demethylation roles.
Although the numbers of JmjC domain proteins were relatively small, some genes seemed to have been duplicated. Arabidopsis and cabbage both belong to the family Brassicaceae, and whole genome duplication is known to have occurred after cabbage and Arabidopsis divergence, making these two plants ideal material for gene duplication studies (Wang et al. 2011). Many KMT genes were found to have duplicated, especially Group-KMT1 genes with H3K9me activity and Group-KMT2/KMT7 genes with H3K4me activity (Huang et al. 2011). Here, the cabbage KDMs were found to have duplicated in Group-II, which was involved in the demethylation of H3K4me1/2/3, and in Group-VII, which was involved in the demethylation of H3K9me1/2/3, suggesting a distinct functional division. Most of the duplicated genes, except BrJMJ10803 and BrJMJ10811, shared synteny with their Arabidopsis homologs, suggesting that they were derived from chromosome/genome segment duplications. Some duplication proteins display similar domain organization, while others increase or lose domain(s).
Based on the proteins phylogenetic tree, domain organization, and their relationship to known functions proteins from model plant Arabidopsis and animal human/fruit fly, the substrate specifies of the plant JmjC domain proteins was inferred (Table 1): Group-I KDM4/JHDM3, demethylation of H3K9me2/3, H3K27me3 and H3K36me1/2/3; Group-II KDM5A, demethylation of H3K4me1/2/3; Group-III KDM5B, demethylation of H3K4me1/2/3 and H3K9me1/2/3; Group-IV JmjC domain-only A and Group-V JmjC domain-only B, hydroxylation and demethylation; Group-VI JMJD6, demethylation of H3R2 and H4R3 and hydroxylation of H4; and Group-VII KDM3/JHDM2, demethylation of H3K9me1/2/3. Based on the Arabidopsis and human annotations, we know that the Group-I KDM4/JHDM3 and Group-II KDM5A proteins might function in flowering time control, Group-V JMJD6 might play a role in seed germination, and Group-VII KDM3/JHDM2 might be involved in leaf, flower, and embryo development, and pollen grain fertility and seed reproduction (Noh et al. 2004; Saze et al. 2008; Jeong et al. 2009; Searle et al. 2010; Yang et al. 2010, Yang et al. 2012a, b; Lu et al. 2011a; Fan et al. 2012; Luo et al. 2014). It is surprising that no JmjC domain protein associated with H3K4me1/2/3 demethylation had been found in V. carteri.
The JmjC domain is the core of JmjC domain proteins, and iron Fe(II) and αKG act as cofactors (Tsukada et al. 2006; Rotili and Mai. 2011). His188, Glu190 and His276 are important for Fe(II) binding and Thr185/Phe185 and Lys206 are required for αKG binding (Chen et al. 2006; Klose et al. 2006; Lu et al. 2008). The sequence alignments of the JmjC domains showed that two His residues in the Fe(II) binding sites and Lys in the αKG binding site were highly conserved in all the JmjC domain proteins, whereas Glu was often replaced by Asp in the Fe(II) binding sites (Online Resource 5–8), and Thr/Phe was often replaced by Ser/Ala/Lys (Online Resource 6–8). Thus, the replacements often occurred among the same types of amino acid residues; Glu and Asp have similar acidic groups, and Thr and Ser each have a hydroxyl group. However, different replacements did occur; for example, one of the His in the Fe(II) binding site of AtJMJ19 and its homologs in Group-II was replaced by Tyr, and Thr/Phe in the αKG binding site was replaced by Gln (Online Resource 3). The AtJMJ24/28 homologs in Group-VII also displayed non-conservative replacements; in particular, Glu and His were replaced by Lys and Phe in the Fe(II) binding sites. These non-conservative replacements occurred mainly in flowering plant JmjC domain proteins with as yet unidentified functions (Online Resource 8).
JmjC domain proteins involved in most of plant development events function by regulating the activity of target genes by balancing the methylation status of the associated histones. Our results show that JmjC domain proteins are an ancient and conserved family in which the domain organization and Fe(II) and αKG binding sites in the JmjC domains have been modified in some species. We have shown that the JmjC domain proteins can be divided into distinct groups that target specific substrates. The results will be useful to further examine the functional conservation and divergence of JmjC domain proteins.
References
Allis CD, Berger SL, Cote J, Dent S, Jenuwien T, Kouzarides T, Pillus L, Reinberg D, Shi Y, Shiekhattar R, Shilatifard A, Workman J, Zhang Y (2007) New nomenclature for chromatin-modifying enzymes. Cell 131(4):633–636
Audonnet L (2014) Caractérisation fonctionnelle de JMJ24, une déméthylase d’histone de la famille JUMONJI, chez Arabidopsis thaliana (D). Université Paris Sud
Ay N, Janack B, Humbeck K (2014) Epigenetic control of plant senescence and linked processes. J Exp Bot 65(14):3875–3887
Bachi A, Braun IC, Rodrigues JP, Panté N, Ribbeck K, von Kobbe C, Kutay U, Wilm M, Görlich D, Carmo-Fonseca M, Izaurralde E (2000) The C-terminal domain of TAP interacts with the nuclear pore complex and promotes export of specific CTE-bearing RNA substrates. RNA 6(1):136–158
Banerjee T, Chakravarti D (2011) A peek into the complex realm of histone phosphorylation. Mol Cell Biol 31(24):4858–4873
Bannister AJ, Kouzarides T (2011) Regulation of chromatin by histone modifications. Cell Res 21(3):381–395
Benevolenskaya EV (2007) Histone H3K4 demethylases are essential in development and differentiation. Biochem Cell Biol 85(4):435–443
Binda O (2013) On your histone mark, SET, methylate. Epigenetics 8(5):457–463
Bustin M, Trieschmann L, Postnikov YV (1995) The HMG-14/-17 chromosomal protein family: architectural elements that enhance transcription from chromatin templates. Semin Cell Biol 6(4):247–255
Chang B, Chen Y, Zhao Y, Bruick RK (2007) JMJD6 is a histone arginine demethylase. Science 318(5849):444–447
Chen YW, Allen MD, Veprintsev DB, Löwe J, Bycroft M (2004) The structure of the AXH domain of spinocerebellar ataxin-1. J Biol Chem 279(5):3758–3765
Chen Z, Zang J, Whetstine J, Hong X, Davrazou F, Kutateladze TG, Simpson M, Mao Q, Pan CH, Dai S, Hagman J, Hansen K, Shi Y, Zhang G (2006) Structural insights into histone demethylation by JMJD2 family members. Cell 125(4):691–702
Chen X, Hu Y, Zhou DX (2011) Epigenetic gene regulation by plant Jumonji group of histone demethylase. Biochim Biophys Acta 1809(8):421–426
Chen S, Ma J, Wu F, Xiong LJ, Ma H, Xu W, Lv R, Li X, Villen J, Gygi SP, Liu XS, Shi Y (2012) The histone H3 Lys 27 demethylase JMJD3 regulates gene expression by impacting transcriptional elongation. Genes Dev 26(12):1364–1375
Chen Q, Chen X, Wang Q, Zhang F, Lou Z, Zhang Q, Zhou DX (2013) Structural basis of a histone H3 lysine 4 demethylase required for stem elongation in rice. PLoS Genet 9(1):e1003239
Cho JN, Ryu JY, Jeong YM, Park J, Song JJ, Amasino RM, Noh B, Noh YS (2012) Control of seed germination by light-induced histone arginine demethylation activity. Dev Cell 22(4):736–748
Cox R, Mason-Gamer RJ, Jackson CL, Segev N (2004) Phylogenetic analysis of Sec7-domain-containing Arf nucleotide exchangers. Mol Biol Cell 15(4):1487–1505
Craig KL, Tyers M (1999) The F-box: a new motif for ubiquitin dependent proteolysis in cell cycle regulation and signal transduction. Prog Biophys Mol Biol 72(3):299–328
de Chiara C, Giannini C, Adinolfi S, de Boer J, Guida S, Ramos A, Jodice C, Kioussis D, Pastore A (2003) The AXH module: an independently folded domain common to ataxin-1 and HBP1. FEBS Lett 551(1–3):107–112
Del Rizzo PA, Krishnan S, Trievel RC (2012) Crystal structure and functional analysis of JMJD5 indicate an alternate specificity and function. Mol Cell Biol 32(19):4044–4052
Dupont C, Armant DR, Brenner CA (2009) Epigenetics: definition, mechanisms and clinical perspective. Semin Reprod Med 27(5):351–357
Erickson D (2012) Role of histone demethylases in pathogen defense. Syracuse University Honors Program Capstone Projects. p140
Fan D, Dai Y, Wang X, Wang Z, He H, Yang H, Cao Y, Deng XW, Ma L (2012) IBM1, a JmjC domain-containing histone demethylase, is involved in the regulation of RNA-directed DNA methylation through the epigenetic control of RDR2 and DCL3 expression in Arabidopsis. Nucleic Acids Res 40(18):8905–8916
Feng T, Yamamoto A, Wilkins SE, Sokolova E, Yates LA, Münzel M, Singh P, Hopkinson RJ, Fischer R, Cockman ME, Shelley J, Trudgian DC, Schödel J, McCullagh JS, Ge W, Kessler BM, Gilbert RJ, Frolova LY, Alkalaeva E, Ratcliffe PJ, Schofield CJ, Coleman ML (2014) Optimal translational termination requires C4 lysyl hydroxylation of eRF1. Mol Cell 53(4):645–654
García-Alai MM, Allen MD, Joerger AC, Bycroft M (2010) The structure of the FYR domain of transforming growth factor beta regulator 1. Protein Sci 19(7):1432–1438
Gelato KA, Fischle W (2008) Role of histone modifications in defining chromatin structure and function. Biol Chem 389(4):353–363
Hahn P, Wegener I, Burrells A, Böse J, Wolf A, Erck C, Butler D, Schofield CJ, Böttger A, Lengeling A (2010) Analysis of Jmjd6 cellular localization and testing for its involvement in histone demethylation. PLoS One 5(10):e13769
Han G, Li J, Wang Y, Li X, Mao H, Liu Y, Chen CD (2012) The hydroxylation activity of Jmjd6 is required for its homo-oligomerization. J Cell Biochem 113(5):1663–1670
Holec S, Berger F (2012) Polycomb group complexes mediate developmental transitions in plants. Plant Physiol 158(1):35–43
Hong MJ, Kim DY, Lee TG, Jeon WB, Seo YW (2010) Functional characterization of pectin methylesterase inhibitor (PMEI) in wheat. Genes Genet Syst 85(2):97–106
Hou H, Yu H (2010) Structural insights into histone lysine demethylation. Curr Opin Struct Biol 20(6):739–748
Huang F, Chandrasekharan MB, Chen YC, Bhaskara S, Hiebert SW, Sun ZW (2010) The JmjN domain of Jhd2 is important for its protein stability, and the plant homeodomain (PHD) finger mediates its chromatin association independent of H3K4 methylation. J Biol Chem 285(32):24548–24561
Huang Y, Liu C, Shen WH, Ruan Y (2011) Phylogenetic analysis and classification of the Brassica rapa SET-domain protein family. BMC Plant Biol 11:175
Ishimura A, Minehata K, Terashima M, Kondoh G, Hara T, Suzuki T (2012) Jmjd5, an H3K36me2 histone demethylase, modulates embryonic cell proliferation through the regulation of Cdkn1a expression. Development 139(4):749–759
Jain M, Nijhawan A, Arora R, Agarwal P, Ray S, Sharma P, Kapoor S, Tyagi AK, Khurana JP (2007) F-box proteins in rice. Genome-wide analysis, classification, temporal and spatial gene expression during panicle and seed development, and regulation by light and abiotic stress. Plant Physiol 143(4):1467–1483
Jeong JH, Song HR, Ko JH, Jeong YM, Kwon YE, Seol JH, Amasino RM, Noh B, Noh YS (2009) Repression of Flowering Locus T chromatin by functionally redundant histone H3 lysine 4 demethylases in Arabidopsis. PLoS One 4(11):e8033
Jiang D, Yang W, He Y, Amasino RM (2007) Arabidopsis relatives of the human lysine-specific Demethylase1 repress the expression of FWA and Flowering Locus C and thus promote the floral transition. Plant Cell 19(10):2975–2987
Joazeiro CA, Weissman AM (2000) RING finger proteins: mediators of ubiquitin ligase activity. Cell 102(5):549–552
Jones MA, Harmer S (2011) JMJD5 Functions in concert with TOC1 in the Arabidopsis circadian system. Plant Signal Behav 6(3):445–448
Jones B, Su H, Bhat A, Lei H, Bajko J, Hevi S, Baltus GA, Kadam S, Zhai H, Valdez R, Gonzalo S, Zhang Y, Li E, Chen T (2008) The histone H3K79 methyltransferase Dot1L is essential for mammalian development and heterochromatin structure. PLoS Genet 4(9):e1000190
Jones MA, Covington MF, DiTacchio L, Vollmers C, Panda S, Harmer SL (2010) Jumonji domain protein JMJD5 functions in both the plant and human circadian systems. PNAS 107(50):21623–21628
Kalkhoven E, Teunissen H, Houweling A, Verrijzer CP, Zantema A (2002) The PHD type zinc finger is an integral part of the CBP acetyltransferase domain. Mol Cell Biol 22(7):1961–1970
Kim S, Zhang Z, Upchurch S, Isern N, Chen Y (2004) Structure and DNA-binding sites of the SWI1 AT-rich interaction domain (ARID) suggest determinants for sequence-specific DNA recognition. J Biol Chem 279(16):16670–16676
Klose RJ, Kallin EM, Zhang Y (2006) JmjC-domain-containing proteins and histone demethylation. Nat Rev Genet 7(9):715–727
Klosterman SJ, Hadwiger LA (2002) Plant HMG proteins bearing the AT-hook motif. Plant Sci 162(6):855–866
Klug A (1999) Zinc finger peptides for the regulation of gene expression. J Mol Biol 293(2):215–218
Kouzarides T (2007) Chromatin modifications and their function. Cell 128(4):693–705
Kuroki S, Matoba S, Akiyoshi M, Matsumura Y, Miyachi H, Mise N, Abe K, Ogura A, Wilhelm D, Koopman P, Nozaki M, Kanai Y, Shinkai Y, Tachibana M (2013) Epigenetic regulation of mouse sex determination by the histone demethylase Jmjd1a. Science 341(6150):1106–1109
Lan F, Nottke AC, Shi Y (2008) Mechanisms involved in the regulation of histone lysine demethylases. Curr Opin Cell Biol 20:316–325
Legge GB, Martinez-Yamout MA, Hambly DM, Trinh T, Lee BM, Dyson HJ, Wright PE (2004) ZZ domain of CBP: an unusual zinc finger fold in a protein interaction module. J Mol Biol 343(4):1081–1093
Lennartsson A, Ekwall K (2009) Histone modification patterns and epigenetic codes. Biochim Biophys Acta 1790(9):863–868
Li Y, Qiu L, Huang L, Cao J (2012) BcJMJ30, the gene encoding jmjC domain-containing histone demethylase is associated with pollen development and fertilization in Brassica campestris ssp. chinensis. Plant Mol Biol Rep 30(3):529–538
Li T, Chen X, Zhong X, Zhao Y, Liu X, Zhou S, Cheng S, Zhou DX (2013) Jumonji C domain protein JMJ705-mediated removal of histone H3 lysine 27 trimethylation is involved in defense-related gene activation in rice. Plant Cell 25(11):4725–4736
Liang Y, Vogel JL, Arbuckle JH, Rai G, Jadhav A, Simeonov A, Maloney DJ, Kristie TM (2013) Targeting the JMJD2 histone demethylases to epigenetically control herpesvirus infection and reactivation from latency. Sci Transl Med 5(167):167ra5
Loh YH, Zhang W, Chen X, George J, Ng HH (2007) Jmjd1a and Jmjd2c histone H3 Lys 9 demethylases regulate self-renewal in embryonic stem cells. Genes Dev 21(20):2545–2557
Lorick KL, Jensen JP, Fang S, Ong AM, Hatakeyama S, Weissman AM (1999) RING fingers mediate ubiquitin-conjugating enzyme (E2)-dependent ubiquitination. PNAS 96(20):11364–11369
Lu J, Matunis MJ (2013) A mediator methylation mystery: JMJD1C demethylates MDC1 to regulate DNA repair. Nat Struct Mol Biol 20(12):1346–1348
Lu F, Li G, Cui X, Liu C, Wang XJ, Cao X (2008) Comparative analysis of JmjC domain-containing proteins reveals the potential histone demethylases in Arabidopsis and rice. J Integr Plant Biol 50(7):886–896
Lu F, Cui X, Zhang S, Liu C, Cao X (2010) JMJ14 is an H3K4 demethylase regulating flowering time in Arabidopsis. Cell Res 20:387–390
Lu F, Cui X, Zhang S, Jenuwein T, Cao X (2011a) Arabidopsis REF6 is a histone H3 lysine 27 demethylase. Nat Genet 43(7):715–719
Lu SX, Knowles SM, Webb CJ, Celaya RB, Cha C, Siu JP, Tobin EM (2011b) The Jumonji C domain-containing protein JMJ30 regulates period length in the Arabidopsis circadian clock. Plant Physiol 155(2):906–915
Lu J, Marjon KD, Mold C, Du Clos TW, Sun PD (2013) Pentraxins and Fc receptors. Immunol Rev 250(1):230–238
Luo M, Hung FY, Yang S, Liu X, Wu K (2014) Histone lysine demethylases and their functions in plants. Plant Mol Biol Rep 32(2):558–565
Maris C, Dominguez C, Allain FH (2005) The RNA recognition motif, a plastic RNA-binding platform to regulate post transcriptional gene expression. FEBS J 272(9):2118–2131
Metzger E, Wissmann M, Yin N, Müller JM, Schneider R, Peters AH, Günther T, Buettner R, Schüle R (2005) LSD1 demethylates repressive histone marks to promote androgen-receptor-dependent transcription. Nature 437(7057):436–439
Mosammaparast N, Shi Y (2010) Reversal of histone methylation: biochemical and molecular mechanisms of histone demethylases. Annu Rev Biochem 79:155–179
Mossessova E, Gulbis JM, Goldberg J (1998) Structure of the guanine nucleotide exchange factor Sec7 domain of human arno and analysis of the interaction with ARF GTPase. Cell 92(3):415–423
Nguyen AT, Zhang Y (2011) The diverse functions of Dot1 and H3K79 methylation. GenesDev 25(13):1345–1358
Noh B, Lee SH, Kim HJ, Yi G, Shin EA, Lee M, Jung KJ, Doyle MR, Amasino RM, Noh YS (2004) Divergent roles of a pair of homologous jumonji/zinc-finger-class transcription factor proteins in the regulation of Arabidopsis flowering time. Plant Cell 16(10):2601–2613
Park SJ, Kim JG, Son TG, Yi JM, Kim ND, Yang K, Heo K (2013) The histone demethylase JMJD1A regulates adrenomedullin-mediated cell proliferation in hepatocellular carcinoma under hypoxia. Biochem Biophys Res Commun 434(4):722–727
Pasini D, Cloos PA, Walfridsson J, Olsson L, Bukowski JP, Johansen JV, Bak M, Tommerup N, Rappsilber J, Helin K (2010) JARID2 regulates binding of the Polycomb repressive complex 2 to target genes in ES cells. Nature 464(7286):306–310
Patsialou A, Wilsker D, Moran E (2005) DNA-binding properties of ARID family proteins. Nucleic Acids Res 33(1):66–80
Pilka ES, James T, Lisztwan JH (2015) Structural definitions of Jumonji family demethylase selectivity. Drug Discov Today. doi:10.1016/j.drudis.2014.12.013
Ponting CP, Blake DJ, Davies KE, Kendrick-Jones J, Winder SJ (1996) ZZ and TAZ: new putative zinc fingers in dystrophin and other proteins. Trends Biochem Sci 21(1):11–13
Pontvianne F, Blevins T, Pikaard CS (2010) Arabidopsis histone lysine methyltransferases. Adv Bot Res 53:1–22
Qi HH, Sarkissian M, Hu GQ, Wang Z, Bhattacharjee A, Gordon DB, Gonzales M, Lan F, Ongusaha PP, Huarte M, Yaghi NK, Lim H, Garcia BA, Brizuela L, Zhao K, Roberts TM, Shi Y (2010) Histone H4K20/H3K9 demethylase PHF8 regulates zebrafish brain and craniofacial development. Nature 466(7305):503–507
Quan Z, Oliver SG, Zhang N (2011) JmjN interacts with JmjC to ensure selective proteolysis of Gis1 by the proteasome. Microbiology 157(Pt 9):2694–2701
Rotili D, Mai A (2011) Targeting histone demethylases: a new avenue for the fight against cancer. Genes Cancer 2(6):663–679
Saze H, Shiraishi A, Miura A, Kakutani T (2008) Control of genic DNA methylation by a jmjC domain-containing protein in Arabidopsis thaliana. Science 319(5862):462–465
Searle IR, Pontes O, Melnyk CW, Smith LM, Baulcombe DC (2010) JMJ14, a JmjC domain protein, is required for RNA silencing and cell-to-cell movement of an RNA silencing signal in Arabidopsis. Genes Dev 24(10):986–991
Secombe J, Eisenman RN (2007) The function and regulation of the JARID1 family of histone H3 lysine 4 demethylases: the Myc connection. Cell Cycle 6(11):1324–1328
Shahbazian MD, Grunstein M (2007) Functions of site-specific histone acetylation and deacetylation. Annu Rev Biochem 76:75–100
Shen Y, Conde E, Silva N, Audonnet L, Servet C, Wei W, Zhou DX (2014) Over-expression of histone H3K4 demethylase gene JMJ15 enhances salt tolerance in Arabidopsis. Front Plant Sci 5:290
Shi Y, Lan F, Matson C, Mulligan P, Whetstine JR, Cole PA, Casero RA, Shi Y (2004) Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119(7):941–953
Smit JJ, Monteferrario D, Noordermeer SM, van Dijk WJ, van der Reijden BA, Sixma TK (2012) The E3 ligase HOIP specifies linear ubiquitin chain assembly through its RING-IBR-RING domain and the unique LDD extension. EMBO J 31(19):3833–3844
Sun Q, Zhou DX (2008) Rice jmjC domain-containing gene JMJ706 encodes H3K9 demethylase required for floral organ development. PNAS 105(36):13679–13684
Takeuchi T, Yamazaki Y, Katoh-Fukui Y, Tsuchiya R, Kondo S, Motoyama J, Higashinakagawa T (1995) Gene trap capture of a novel mouse gene, jumonji, required for neural tube formation. Genes Dev 9(10):1211–1222
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30(12):2725–2729
Tee AE, Ling D, Nelson C, Atmadibrata B, Dinger ME, Xu N, Mizukami T, Liu PY, Liu B, Cheung B, Pasquier E, Haber M, Norris MD, Suzuki T, Marshall GM, Liu T (2014) The histone demethylase JMJD1A induces cell migration and invasion by up-regulating the expression of the long noncoding RNA MALAT1. Oncotarget 5(7):1793–1804
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22(22):4673–4680
Thorstensen T, Grini PE, Aalen RB (2011) SET domain proteins in plant development. Biochim Biophys Acta 1809(8):407–420
Tschiersch B, Hofmann A, Krauss V, Dorn R, Korge G, Reuter G (1994) The protein encoded by the Drosophila position-effect variegation suppressor gene Su(var)3-9 combines domains of antagonistic regulators of homeotic gene complexes. EMBO J 13:3822–3831
Tsukada Y, Fang J, Erdjument-Bromage H, Warren ME, Borchers CH, Tempst P, Zhang Y (2006) Histone demethylation by a family of JmjC domain-containing proteins. Nature 439(7078):811–816
van Leeuwen F, Gottschling DE (2002) Genome-wide histone modifications: gaining specificity by preventing promiscuity. Curr Opin Cell Biol 14(6):756–762
Wang L, Dong L, Zhang Y, Zhang Y, Wu W, Deng X, Xue Y (2004) Genome-wide analysis of S-Locus F-box-like genes in Arabidopsis thaliana. Plant Mol Biol 56(6):929–945
Wang X, Wang H, Wang J, Sun R, Wu J, Liu S, Bai Y, Mun JH, Bancroft I, Cheng F, Huang S, Li X, Hua W, Wang J, Wang X, Freeling M, Pires JC, Paterson AH, Chalhoub B, Wang B, Hayward A, Sharpe AG, Park BS, Weisshaar B, Liu B, Li B, Liu B, Tong C, Song C, Duran C, Peng C, Geng C, Koh C, Lin C, Edwards D, Mu D, Shen D, Soumpourou E, Li F, Fraser F, Conant G, Lassalle G, King GJ, Bonnema G, Tang H, Wang H, Belcram H, Zhou H, Hirakawa H, Abe H, Guo H, Wang H, Jin H, Parkin IA, Batley J, Kim JS, Just J, Li J, Xu J, Deng J, Kim JA, Li J, Yu J, Meng J, Wang J, Min J, Poulain J, Wang J, Hatakeyama K, Wu K, Wang L, Fang L, Trick M, Links MG, Zhao M, Jin M, Ramchiary N, Drou N, Berkman PJ, Cai Q, Huang Q, Li R, Tabata S, Cheng S, Zhang S, Zhang S, Huang S, Sato S, Sun S, Kwon SJ, Choi SR, Lee TH, Fan W, Zhao X, Tan X, Xu X, Wang Y, Qiu Y, Yin Y, Li Y, Du Y, Liao Y, Lim Y, Narusaka Y, Wang Y, Wang Z, Li Z, Wang Z, Xiong Z, Zhang Z (2011) The genome of the mesopolyploid crop species Brassica rapa. Nat Genet 43(10):1035–1039
Wang F, He L, Huangyang P, Liang J, Si W, Yan R, Han X, Liu S, Gui B, Li W, Miao D, Jing C, Liu Z, Pei F, Sun L, Shang Y (2014) JMJD6 promotes colon carcinogenesis through negative regulation of p53 by hydroxylation. PLoS Biol 12(3):e1001819
Watanabe S, Watanabe K, Akimov V, Bartkova J, Blagoev B, Lukas J, Bartek J (2013) JMJD1C demethylates MDC1 to regulate the RNF8 and BRCA1-mediated chromatin response to DNA breaks. Nat Struct Mol Biol 20(12):1425–1433
Wu CT, Morris JR (2001) Genes, genetics, and epigenetics: a correspondence. Science 293(5532):1103–1105
Yang W, Jiang D, Jiang J, He Y (2010) A plant-specific histone H3 lysine 4 demethylase represses the floral transition in Arabidopsis. Plant J 62(4):663–673
Yang H, Mo H, Fan D, Cao Y, Cui S, Ma L (2012a) Overexpression of a histone H3K4 demethylase, JMJ15, accelerates flowering time in Arabidopsis. Plant Cell Rep 31(7):1297–1308
Yang H, Han Z, Cao Y, Fan D, Li H, Mo H, Feng Y, Liu L, Wang Z, Yue Y, Cui S, Chen S, Chai J, Ma L (2012b) A companion cell-dominant and developmentally regulated H3K4 demethylase controls flowering time in Arabidopsis via the repression of FLC expression. PLoS Genet 8(4):e1002664
Yu X, Li L, Li L, Guo M, Chory J, Yin Y (2008) Modulation of brassinosteroid-regulated gene expression by Jumonji domain-containing proteins ELF6 and REF6 in Arabidopsis. PNAS 105(21):7618–7623
Zhou X, Ma H (2008) Evolutionary history of histone demethylase families: distinct evolutionary patterns suggest functional divergence. BMC Evol Biol 8:294
Zhu H, Hu S, Baker J (2014) JMJD5 regulates cell cycle and pluripotency in human embryonic stem cells. Stem Cells 32(8):2098–2110
Acknowledgments
This work was funded in part by Open Fund of Innovation Platform in University of Hunan (14K046), Hunan Provincial Education Department of China (13B045) and Natural Science Foundation of Hunan Province of China (2015JJ3075).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical standards
This article does not contain any studies with animals performed by any of the authors.
Additional information
Communicated by S. Hohmann.
Authors Y. Huang and D. Chen contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Huang, Y., Chen, D., Liu, C. et al. Evolution and conservation of JmjC domain proteins in the green lineage. Mol Genet Genomics 291, 33–49 (2016). https://doi.org/10.1007/s00438-015-1089-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-015-1089-4