Abstract
Cryptosporidium spp. are key gastrointestinal protists in humans and animals worldwide. Infected cattle are considered the main source of cryptosporidiosis outbreaks in humans. However, little is known about the genetic makeup of Cryptosporidium populations in Shanxi province, China. We analyzed 858 fecal samples collected from farms in Shanxi. The presence of Cryptosporidium spp. was determined via polymerase chain reaction and subsequent sequence analysis of the small subunit rRNA gene as well as restriction fragment length polymorphism analysis. Cryptosporidium parvum was subtyped following sequence analysis of the 60 kDa glycoprotein gene (gp60). The overall prevalence of Cryptosporidium in cattle was 11.19%, with a prevalence of 13.30% and 8.67% in Lingqiu and Yingxian, respectively. The overall prevalence of Cryptosporidium in dairy and beef cattle was 10.78% and 11.50%, respectively. Cryptosporidium infection was detected across all analyzed age groups. The overall prevalence of Cryptosporidium in diarrhea and nondiarrhea samples was 18.24% and 9.72%, respectively, whereas that in intensively farmed and free-range cattle was 17.40% and 3.41%, respectively. We identified five Cryptosporidium species, with C. andersoni being the dominant species. Further, two cases of mixed infections of Cryptosporidium species were detected. All identified C. parvum isolates belonged to the subtype IIdA17G1.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cryptosporidium infects various vertebrates, including humans (Fayer 2010). It is one of the four major pathogens responsible for severe diarrhea in infants and young children (Kotloff et al. 2013), the second leading cause of diarrhea and death in children after rotavirus (Zhao et al. 2019), and the fifth most important foodborne parasite worldwide (Ryan et al. 2021). Individual animals with weakened immune function are more susceptible to Cryptosporidium infection than healthy adult animals (Feng et al. 2018), and Cryptosporidium infection in cattle may manifest as persistent watery diarrhea (Wang et al. 2018), significant weight loss (Sarkar et al. 2014), and decreased growth rate after recovery (Thomson et al. 2019). During the environmental stage, the oocyst is highly resistant and difficult to inactivate (Blanchard 2012), and no vaccine or effective therapeutic drug is currently available (Ashigbie et al. 2021).
Since the first report of Cryptosporidium in 1907 (Tyzzer 1907), at least 44 valid Cryptosporidium species and approximately 120 genotypes have been described globally (Uran-Velasquez et al. 2022). Cattle are an important host (Ryan et al. 2014), and at least 12 Cryptosporidium species (Wang et al. 2011b), predominantly Cryptosporidium parvum, C. andersoni, C. ryanae, and C. bovis, have been reported in cattle worldwide (Fayer 2010; Wang et al. 2018; Yang et al. 2020). C. parvum is the most important cause of zoonotic cryptosporidiosis (Feng et al. 2018). Over 20 subtype families of C. parvum gp60 have been identified, with IIa, IIc, and IId being the most predominant subtype families (Wang et al. 2022). Only C. parvum infections caused by the IId subtype family have been identified in cattle in China, with IIdA15G1 and IIdA19G1 being the most common subtypes (Cai et al. 2017; Wang et al. 2017).
The global prevalence of cryptosporidiosis in cattle ranges from 6.25 to 39.65% (Tarekegn et al. 2021), with a global pooled prevalence of 29.1% (Hatam-Nahavandi et al. 2019). In China, the pooled prevalence has been estimated as 14.50% (Wang et al. 2017) and 11.9% (Gong et al. 2017). To date, only the infection rate of Cryptosporidium in rodents (3.8%, 2/53) has been reported in Shanxi province, China (Ni et al. 2021). We performed the molecular epidemiological survey of cattle in Shanxi to determine the prevalence and molecular characterization of Cryptosporidium.
Materials and methods
Study areas and sample collection
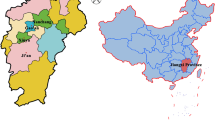
From March 2021 to September 2022, 858 fresh stool samples (30 g each) were collected from Lingqiu and Yingxian in Shanxi province, China (37°27′–38°25′N, 111°30′–113°09′E). The samples were collected from 48 preweaned calves (0–60 days old), 128 postweaned calves (61–180 days old), 186 young cattle (181–450 days old), and 496 adult cattle (>450 days old). Overall, 148 cattle had diarrhea. Each stool sample was placed inside a clean, labeled, and sterile plastic zipper bag, and the information was recorded (sampling date, age, and fecal consistency). Then, all samples were transported back to the laboratory for storage at 4°C and were processed as soon as possible.
DNA extraction, polymerase chain reaction, and sequence analysis
Total genomic DNA was extracted from each sample using E.Z.N.A Stool DNA Kit (Omega Bio-Tek Inc., Norcross, GA, USA) according to the manufacturer’s instructions. The final volume of each DNA sample was 200 μL. The samples were stored at −20°C until polymerase chain reaction (PCR) amplification.
To determine the prevalence of Cryptosporidium spp., a fragment of the small subunit (SSU) rRNA gene (~840 bp) was amplified via nested PCR (Feng et al. 2007). The 25-μL PCR mixture comprised 12.5 μL of Premix Taq (TaKaRa Taq Version 2.0 plus dye; TaKaRa Bio Inc., Kusatsu, Shiga, Japan), 0.5 μL each of forward and reverse primers, 1 μL of template DNA or first-round PCR product, and 10.5 μL of sterilized double distilled water. The first-round PCR parameters were as follows: predenaturation at 94°C for 5 min; followed by 35 cycles of denaturation at 94°C for 45 s, annealing at 55°C for 45 s, and extension at 72°C for 1 min; extension at 72°C for 7 min; and insulation at 4°C. The second-round PCR conditions were identical to those of the first-round PCR, except that the annealing temperature was 58°C. All rounds of PCR amplification included positive and negative controls. The products from the second-round PCR were analyzed via 1.5% agarose gel electrophoresis, and all positive PCR products were sent to a commercial company for bidirectional sequencing (Sangon Biotech, Shanghai, China). The sequencing results were assembled and corrected using SeqMan software (DNASTAR Inc., Madison, USA) for sequence analysis.
Subsequently, restriction fragment length polymorphism (RFLP) analysis of PCR-positive products was performed using SspI and MboII restriction endonucleases to further determine the species and status of mixed infections of Cryptosporidium. The presence of C. parvum DNA was confirmed via the abovementioned enzyme digestion and sequence analysis, and gp60 nested PCR was performed (Alves et al. 2003) at annealing temperatures of 52°C and 50°C for the first- and second-round PCR, respectively. A PCR amplification product of approximately 850 bp was identified as C. parvum, and the subtype was identified following sequencing (Sulaiman et al. 2005).
All sequences were aligned with reference sequences downloaded from GenBank (https://www.ncbi.nlm.nih.gov/genbank/) using MEGA v11.0 software (http://www.megasoftware.net/). The sequencing results were analyzed using the BLAST online platform (https://blast.ncbi.nlm.nih.gov/Blast.cgi?PROGRAM=blastn&PAGE_TYPE=BlastSearch&LINK_LOC=blasthome). To comprehensively investigate the relationship among different isolates, we constructed phylogenetic trees using the neighbor-joining algorithm based on a matrix of evolutionary distances, which were calculated using the Kimura two-parameter model via MEGA. We performed bootstrap analysis (1000 replicates) to assess the robustness of the clusters.
Statistical analysis and GenBank accession numbers
Chi-square test was performed, and 95% confidence interval (CI) values were determined using SPSS Statistics v21.0 (IBM Corp., New York, NY, USA) to compare Cryptosporidium infection rates among different sampling sites, age groups, breeds, and feeding methods as well as between diarrheal and nondiarrheal groups. A two-tailed P-value of <0.05 was considered to indicate statistical significance.
Results
Prevalence of Cryptosporidium spp.
Based on SSU rRNA PCR results, we revealed that 11.19% (96/858) of the samples were positive for Cryptosporidium. The overall prevalence of Cryptosporidium in cattle was 13.30% (62/466) and 8.67% (34/392) in Lingqiu and Yingxian, respectively, with a statistically significant difference between the two regions (odds ratio [OR]: 1.616, 95% CI: 1.039–2.514, P = 0.032). The overall prevalence of Cryptosporidium in dairy and beef cattle was 10.78% (40/371) and 11.50% (56/487), respectively, with no significant difference between them (OR: 0.930, 95% CI: 0.605–1.430, P = 0.741). Among the four age groups, postweaned calves (25.78%, 33/128) had the highest infection rate, followed by preweaned calves (18.75%, 9/48). Young (14.52%, 27/186) and adult (5.44%, 27/496) cattle showed the lowest infection rates. The overall prevalence of Cryptosporidium in preweaned calves, postweaned calves, and young cattle was significantly higher than that in adult cattle (OR: 4.009, 95% CI: 1.762–9.120, P < 0.001; OR: 6.034, 95% CI: 3.466–10.504, P < 0.001; and OR: 2.950, 95% CI: 1.680–5.179, P < 0.001, respectively). The overall prevalence of Cryptosporidium in diarrhea and nondiarrhea samples was 18.24% (27/148) and 9.72% (69/710), respectively, with the infection rate being significantly higher in diarrhea samples (OR: 2.073, 95% CI: 1.276–3.368, P = 0.003). The overall prevalence of Cryptosporidium in intensively farmed and free-range cattle was 17.40% (83/477) and 3.41% (13/381), respectively, with the infection rate being significantly higher in intensively farmed cattle (OR: 5.963, 95% CI: 3.267–10.884, P < 0.001) (Table 1).
Distribution of Cryptosporidium species and its subtypes
Sequencing of 96 Cryptosporidium PCR-positive products yielded 95 sequences from 5 Cryptosporidium species (C. andersoni, 49; C. bovis, 30; C. parvum, 8; C. ryanae, 7; and C. ubiquitum, 1) (Table 1). Phylogenetic analysis of the PCR products revealed five branches (Fig. 1). C. ubiquitum was only identified in Yingxian, whereas the other four Cryptosporidium species were detected in both Lingqiu and Yingxian. C. andersoni was the dominant species in both regions. Further, C. andersoni and C. parvum were identified across all four age groups and were the dominant species in preweaned calves. C. bovis was the dominant species in postweaned calves, whereas C. andersoni was the dominant species in young and adult cattle. Furthermore, C. andersoni was the dominant species in dairy and beef cattle as well as diarrhea and nondiarrhea samples. Additionally, RFLP analysis of the PCR products revealed two cases of mixed infections in Lingqiu: C. andersoni + C. bovis in one sample from dairy cattle and C. andersoni + C. parvum in one sample from beef cattle.
The six gp60 sequences obtained in our study were analyzed and successfully identified as C. parvum subtype IIdA17G1.
Discussion
In recent decades, an increasing number of epidemiological investigations have been conducted both domestically and internationally to assess the prevalence of cryptosporidiosis(Cai et al. 2019; Gong et al. 2017; Hatam-Nahavandi et al. 2019; Wang et al. 2017). The prevalence of bovine cryptosporidiosis varies considerably across different countries, which is associated with factors such as animal age, sampling season, sanitation conditions, total sample number, study design, diagnostic method, geographical conditions, and climate (Gong et al. 2017; Hatam-Nahavandi et al. 2019). The present study revealed that the overall prevalence of Cryptosporidium in cattle was 11.19%, which was higher than that reported in Bangladesh (5%, 31/623) (Ehsan et al. 2015) and Egypt (10.2%, 49/480) (Ibrahim et al. 2016) but lower than that reported in Italy (38.8%, 57/147) (Díaz et al. 2018), USA (24.2%, 60/248) (Peng et al. 2003), and Estonia (23.0% (112/486) (Santoro et al. 2019); further, it was lower than the global pooled prevalence in cattle (29.1%) (Hatam-Nahavandi et al. 2019). Regarding the domestic market, the prevalence of Cryptosporidium in cattle in the present study was lower than that in Taiwan (37.6%, 173/460) (Watanabe et al. 2005), Shaanxi (20.2%, 52/258) (Qi et al. 2015a), Hubei (15.6%, 53/339) (Fan et al. 2017), and Jiangxi (12.8%, 71/556) (Li et al. 2021); moreover, it was lower than that observed in two reports of pooled prevalence in Chinese cattle: 14.5% (5265/36316) (Wang et al. 2017) and 11.9% (2623/22,051) (Gong et al. 2017).
In the present study, the Cryptosporidium infection rate in dairy cattle was 10.78% (40/371), which was lower than that reported in Heilongjiang (47.68%, 72/151) (Zhang et al. 2013); Xinjiang (38.4%, 39/232 (Wu et al. 2020); 16.0%, 86/514 (Qi et al. 2015b)); Shanghai (37%, 303/818) (Cai et al. 2017); Henan (21.5%, 172/801) (Wang et al. 2011b); Anhui, Jiangsu, and Shanghai (18.82%, 387/2056) (Chen and Huang 2012); and Sichuan (14.4%, 40/278) (Zhong et al. 2018). Further, it was lower than the previously reported pooled prevalence of Cryptosporidium in Chinese dairy cattle of 11.7% (3901/33,313) (Cai et al. 2019) and 13.98% (4405/31,504) (Wang et al. 2017). The prevalence of Cryptosporidium in dairy cattle in the present study was higher than that in Ningxia (1.61%, 22/1366) (Huang et al. 2014), Beijing (2.55%, 21/822) (Li et al. 2016), Gansu (4.2%, 60/1414) (Wang et al. 2020b), Guangdong (4.38%, 63/1440) (Liang et al. 2019), Northwest China (Gansu and Ningxia; 5.09%, 150/2945) (Zhang et al. 2015), Henan (7.9%, 104/1315) (Wang et al. 2011a), and Shaanxi (2.61%, 32/1224) (Zhao et al. 2013); moreover, it was higher than the pooled prevalence of Cryptosporidium in Chinese dairy cattle (10.44%, 1330/12743) (Cai et al. 2019). The Cryptosporidium infection rate in beef cattle in our study was 11.50% (56/487), which was lower than the reported prevalence in beef cattle in Henan (26.5%, 44/166) (Ma et al. 2015) and Heilongjiang (17.53%, 71/405) (Zhao et al. 2014), and higher than the pooled prevalence in Chinese beef cattle (8.09%, 82/1013 (Gong et al. 2017); 10.47%, 122/1165 (Wang et al. 2017)). Thus, geographical differences exist in Cryptosporidium distribution.
Host age is an important factor affecting the pathogenicity of Cryptosporidium (Helmy et al. 2013), and the prevalence of Cryptosporidium in preweaned calves is generally higher than that in other age groups (Liang et al. 2019). Compared with calves, the incidence of cryptosporidiosis is several times lower in older cattle (Santín and Trout 2007). However, our study revealed that the Cryptosporidium infection rate of postweaned calves was higher than that of preweaned calves, possibly due to higher levels of maternal antibodies in preweaned calves (Cai et al. 2019). Additionally, the age groups varied among studies. In our study, the preweaned calves were <60 days old, whereas the preweaned calves in some studies were 0–90 days old (Watanabe et al. 2005; Liu et al. 2009; Zhao et al. 2013; Zhang et al. 2015; Wang et al. 2020b); thus, some of them would have been considered postweaned calves in our study.
In the present study, the overall prevalence of Cryptosporidium in bovine diarrhea samples (18.24%) was significantly higher than that in nondiarrhea samples (9.72%). This is consistent with a comprehensive report in China revealing that the prevalence of Cryptosporidium was higher in cattle with diarrhea than in those without diarrhea (Cai et al. 2019; Wang et al. 2017). In our study, the overall prevalence of Cryptosporidium (17.40%) in intensively farmed cattle was significantly higher than that in free-range cattle (3.41%), which might be attributed to a high feeding density. Several studies have reported that pathogen transmission is considerably more likely within a concentrated animal feeding operation than in extensive farming systems (Hatam-Nahavandi et al. 2019). As reported in previous reviews in China, owing to the emergence of mega farms in some regions, the prevalence of C. parvum increased from 26.0% in 2011–2016 to 46.8% in 2017–2021 (Guo et al. 2022).
The Cryptosporidium species detected in our study were C. andersoni, C. bovis, C. parvum, C. ryanae, and C. ubiquitum, which included the top four species infecting cattle worldwide (Fayer 2010; Yang et al. 2020). C. andersoni was the dominant species in dairy and beef cattle in our study, consistent with reports from many regions of China and India (Liu et al. 2009; Paul et al. 2009; Wang et al. 2011b; Zhao et al. 2013). In contrast, C. bovis has been reported as the dominant species in beef cattle in Australia, Japan, and France (Abeywardena et al. 2013; Murakoshi et al. 2012; Rieux et al. 2013). C. andersoni usually infects adult cattle (Thomson et al. 2019), consistent with our study. This species is associated with gastritis, decreased milk production, and poor weight gain (Wang et al. 2020a). Additionally, although the C. ubiquitum infection rate in our study was low, its importance must not be neglected. In Ethiopia, C. ubiquitum was detected in a 12-year-old girl who had daily contact with adult cattle and sheep (Kifleyohannes et al. 2022), and investigations in the same study area revealed the presence of C. ubiquitum in calves, lambs, and goat kids (Kifleyohannes et al. 2021), indicating that C. ubiquitum poses a threat to human health. Additionally, C. ubiquitum has been detected in urban sewage in Harbin, China (Liu et al. 2011). Therefore, our study supports the fact that the impact of C. ubiquitum on humans cannot be neglected in China.
Several reports of age-related infections in different Cryptosporidium species have been published (Huang et al. 2014; Langkjær et al. 2007; Liang et al. 2019). In China, preweaned calves are mainly infected with C. bovis (Wang et al. 2017), as reported in Henan (Wang et al. 2011b), Heilongjiang (Zhang et al. 2013), and Shaanxi (Qi et al. 2015a). Some studies have suggested that C. parvum is the most common cause of Cryptosporidium infection in preweaned calves in various countries (Geurden et al. 2007; Plutzer and Karanis 2007). A similar phenomenon was observed in Ningxia (Cui et al. 2014; Huang et al. 2014) and Xinjiang (Qi et al. 2020) in China. In our study, the prevalence of C. parvum was similar to that of C. andersoni in preweaned calves. In the present study, C. bovis was the dominant species in postweaned calves, which differed from the findings of Wang et al. (2011a) and Qi et al. (2015b), who reported that C. andersoni was the dominant species in preweaned calves in China. Our study revealed that C. andersoni was the dominant species causing Cryptosporidium infections in both young and adult cattle, consistent with the finding of a previous study in Heilongjiang, China (Liu et al. 2009). Therefore, differences in the age distribution of samples from different studies might have led to differences in the identified dominant species.
In the present study, mixed infections were detected in Lingqiu. Mixed infections of different Cryptosporidium species are common, as observed in our previous study in Inner Mongolia (Zhao et al. 2023). The high prevalence of Cryptosporidium in Lingqiu also contributes to the emergence of mixed infections. A previous study stated that the first parasite to arrive can gain a marked reproductive advantage or induce cross-reaction immunity; however, it can also pave the way for subsequent infections by disrupting the associated defenses and through immunosuppression (Herczeg et al. 2021).
There are 14 subtype families of C. parvum, of which IIa and IId are the two main zoonotic subtype families in humans and animals (Wang et al. 2014), with other subtype families occasionally appearing in humans and animals (Wang et al. 2011b). In most industrialized countries, the IIa subtype family is relatively common (Holzhausen et al. 2019; Smith et al. 2014), particularly IIaA15G2R1 (Gharieb et al. 2019; Kváč et al. 2011), which is a common zoonotic subtype (Feng et al. 2013; Heckler et al. 2015; Imre et al. 2011). IIdA15G1 and IIdA19G1 are the dominant subtypes detected in dairy cattle infected with C. parvum in China (Feng and Xiao 2017; Wang et al. 2017). The C. parvum subtype in our study was identified as IIdA17G1, which was first detected in dairy cattle in Beijing, China, in 2016 (Li et al. 2016). Thus, our study verified the existence of this subtype in China. Additionally, IIdA17G1 was detected in British European hedgehogs (Sangster et al. 2016), preweaned lambs, and goat kids in northwestern Spain (Díaz et al. 2015) as well as raw and treated water in Portugal (Lobo et al. 2009). It has also been detected in humans in the UK (Chalmers et al. 2011), Qatar (Boughattas et al. 2017), Portugal (Alves et al. 2006), and Northern Spain (Azcona-Gutiérrez et al. 2017). Thus, this subtype is associated with a risk of zoonotic disease.
In summary, Cryptosporidium and C. parvum are widely distributed in cattle in China. Effective measures must be taken to prevent the spread of Cryptosporidium and C. parvum IId subtype in China as well as restrict the introduction of the C. parvum IIa subtype. Further biological and molecular epidemiological research should be conducted to improve our understanding of the host specificity and transmission of Cryptosporidium.
Data availability
All the sequences obtained in our laboratory have been uploaded to the GenBank database under the accession numbers OR460649 to OR460678, OR460682 to OR460689, OR460692 to OR460699, OR460744 to OR460792, and OR474477 to OR474482.
Abbreviations
- gp60 :
-
60 kDa glycoprotein
- SSU rRNA :
-
small subunit ribosomal RNA
- RFLP :
-
restriction fragment length polymorphism
- PCR :
-
polymerase chain reaction
- CI :
-
confidence interval
- OR :
-
odds ratio
References
Abeywardena H, Jex AR, Firestone SM, Mcphee S, Driessen N, Koehler AV, Haydon SR, von Samson-Himmelstjerna G, Stevens M, Gasser RB (2013) Assessing calves as carriers of Cryptosporidium and Giardia with zoonotic potential on dairy and beef farms within a water catchment area by mutation scanning. Electrophoresis 34(15):2259–2267
Alves M, Xiao L, Antunes F, Matos O (2006) Distribution of Cryptosporidium subtypes in humans and domestic and wild ruminants in Portugal. Parasitol Res 99:287–292
Alves M, Xiao L, Sulaiman I, Lal AA, Matos O, Antunes F (2003) Subgenotype analysis of Cryptosporidium isolates from humans, cattle, and zoo ruminants in Portugal. J Clin Microbiol 41(6):2744–2747
Ashigbie PG, Shepherd S, Steiner KL, Amadi B, Aziz N, Manjunatha UH, Spector JM, Diagana TT, Kelly P (2021) Use-case scenarios for an anti- Cryptosporidium therapeutic. PLoS Negl Trop Dis 15(3):e0009057
Azcona-Gutiérrez JM, de Lucio A, Hernández-de-Mingo M, García-García C, Soria-Blanco LM, Morales L, Aguilera M, Fuentes I, Carmena D (2017) Molecular diversity and frequency of the diarrheagenic enteric protozoan Giardia duodenalis and Cryptosporidium spp. in a hospital setting in Northern Spain. PloS one 12(6):e0178575
Blanchard PC (2012) Diagnostics of dairy and beef cattle diarrhea. Vet Clin North Am Food Anim Pract 28(3):443–464
Boughattas S, Behnke JM, Al-Ansari K, Sharma A, Abu-Alainin W, Al-Thani A, Abu-Madi MA (2017) Molecular analysis of the enteric protozoa associated with acute diarrhea in hospitalized children. Front Cell Infect Microbiol 7:343
Cai M, Guo Y, Pan B, Li N, Wang X, Tang C, Feng Y, Xiao L (2017) Longitudinal monitoring of Cryptosporidium species in pre-weaned dairy calves on five farms in Shanghai, China. Vet Parasitol 241:14–19
Cai Y, Zhang NZ, Gong QL, Zhao Q, Zhang XX (2019) Prevalence of Cryptosporidium in dairy cattle in China during 2008–2018: a systematic review and meta-analysis. Microb Pathog 132:193–200
Chalmers RM, Smith RP, Hadfield SJ, Elwin K, Giles M (2011) Zoonotic linkage and variation in Cryptosporidium parvum from patients in the United Kingdom. Parasitol Res 108:1321–1325
Chen F, Huang K (2012) Prevalence and molecular characterization of Cryptosporidium spp. in dairy cattle from farms in China. J Vet Sci 13(1):15–22
Cui Z, Wang R, Huang J, Wang H, Zhao J, Luo N, Li J, Zhang Z, Zhang L (2014) Cryptosporidiosis caused by Cryptosporidium parvum subtype IIdA15G1 at a dairy farm in Northwestern China. Parasit Vector 7(1):1–4
Díaz P, Quílez J, Prieto A, Navarro E, Pérez-Creo A, Fernández G, Panadero R, López C, Díez-Baños P, Morrondo P (2015) Cryptosporidium species and subtype analysis in diarrhoeic pre-weaned lambs and goat kids from north-western Spain. Parasitol Res 114:4099–4105
Díaz P, Varcasia A, Pipia A, Tamponi C, Sanna G, Prieto A, Ruiu A, Spissu P, Díez-Baños P, Morrondo P (2018) Molecular characterisation and risk factor analysis of Cryptosporidium spp. in calves from Italy. Parasitol Res 117(10):3081–3090
Ehsan AM, Geurden T, Casaert S, Parvin SM, Islam TM, Ahmed UM, Levecke B, Vercruysse J, Claerebout E (2015) Assessment of zoonotic transmission of Giardia and Cryptosporidium between cattle and humans in rural villages in Bangladesh. PloS one 10(2):e0118239
Fan Y, Wang T, Koehler AV, Hu M, Gasser RB (2017) Molecular investigation of Cryptosporidium and Giardia in pre- and post-weaned calves in Hubei Province, China. Parasit Vectors 10(1):519
Fayer R (2010) Taxonomy and species delimitation in Cryptosporidium. Exp Parasitol 124(1):90–97
Feng Y, Ortega Y, He G, Das P, Xu M, Zhang X, Fayer R, Gatei W, Cama V, Xiao L (2007) Wide geographic distribution of Cryptosporidium bovis and the deer-like genotype in bovines. Vet Parasitol 144(1-2):1–9
Feng Y, Ryan UM, Xiao L (2018) Genetic diversity and population structure of Cryptosporidium. Trends Parasitol 34(11):997–1011
Feng Y, Torres E, Li N, Wang L, Bowman D, Xiao L (2013) Population genetic characterisation of dominant Cryptosporidium parvum subtype IIaA15G2R1. Int J Parasitol 43(14):1141–1147
Feng Y, Xiao L (2017) Molecular epidemiology of cryptosporidiosis in China. Front Microbiol 8:1701
Geurden T, Berkvens D, Martens C, Casaert S, Vercruysse J, Claerebout E (2007) Molecular epidemiology with subtype analysis of Cryptosporidium in calves in Belgium. Parasitology 134(14):1981–1987
Gharieb RM, Bowman DD, Liotta JL, Xiao L (2019) Isolation, genotyping and subtyping of single Cryptosporidium oocysts from calves with special reference to zoonotic significance. Vet Parasitol 271:80–86
Gong C, Cao XF, Deng L, Li W, Huang XM, Lan JC, Xiao QC, Zhong ZJ, Feng F, Zhang Y (2017) Epidemiology of Cryptosporidium infection in cattle in China: a review. Parasite 24:1
Guo Y, Ryan U, Feng Y, Xiao L (2022) Emergence of zoonotic Cryptosporidium parvum in China. Trends Parasitol 38:335–343
Hatam-Nahavandi K, Ahmadpour E, Carmena D, Spotin A, Bangoura B, Xiao L (2019) Cryptosporidium infections in terrestrial ungulates with focus on livestock: a systematic review and meta-analysis. Parasit Vector 12:1–23
Heckler RP, Borges DG, Bacha FB, Onizuka MK, Teruya L, Neves JP, Leal CR, de Lemos RA, Meireles MV, Borges Fde A (2015) First genetic identification of Cryptosporidium parvum subtype IIaA14G2R1in beef cattle in Brazil. Prev Vet Med 121(3-4):391–394
Helmy YA, Krücken J, Nöckler K, von Samson-Himmelstjerna G, Zessin K-H (2013) Molecular epidemiology of Cryptosporidium in livestock animals and humans in the Ismailia province of Egypt. Vet Parasitol 193(1-3):15–24
Herczeg D, Ujszegi J, Kásler A, Holly D, Hettyey A (2021) Host–multiparasite interactions in amphibians: a review. Parasite Vector 14:296
Holzhausen I, Lendner M, Göhring F, Steinhöfel I, Daugschies A (2019) Distribution of Cryptosporidium parvum gp60 subtypes in calf herds of Saxony, Germany. Parasitol Res 118:1549–1558
Huang J, Yue D, Qi M, Wang R, Zhao J, Li J, Shi K, Wang M, Zhang L (2014) Prevalence and molecular characterization of Cryptosporidium spp. and Giardia duodenalisin dairy cattle in Ningxia, northwestern China. BMC Vet Res 10(1):1–5
Ibrahim M, Abdel-Ghany A, Abdel-Latef G, Abdel-Aziz S, Aboelhadid S (2016) Epidemiology and public health significance of Cryptosporidium isolated from cattle, buffaloes, and humans in Egypt. Parasitol Res 115(6):2439–2448
Imre K, Lobo LM, Matos O, Popescu C, Genchi C, Dărăbuş G (2011) Molecular characterisation of Cryptosporidium isolates from pre-weaned calves in Romania: is there an actual risk of zoonotic infections? Vet Parasitol 181(2-4):321–324
Kifleyohannes T, Nødtvedt A, Debenham JJ, Terefe G, Robertson LJ (2021) Cryptosporidium and Giardia in livestock in Tigray, Northern Ethiopia and associated risk factors for infection: a cross-sectional study. Front Vet Sci 8:825940
Kifleyohannes T, Nødtvedt A, Debenham JJ, Tysnes KR, Terefe G, Robertson LJ (2022) Cryptosporidium and Giardia infections in humans in Tigray, Northern Ethiopia: an unexpectedly low occurrence of anthropozoonotic transmission. Acta Trop 231:106450
Kotloff KL, Nataro JP, Blackwelder WC, Nasrin D, Farag TH, Panchalingam S, Wu Y, Sow SO, Sur D, Breiman RF (2013) Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): a prospective, case-control study. The Lancet 382(9888):209–222
Kváč M, Hromadová N, Květoňová D, Rost M, Sak B (2011) Molecular characterization of Cryptosporidium spp. in pre-weaned dairy calves in the Czech Republic: absence of C. ryanae and management-associated distribution of C. andersoni, C. bovis and C. parvum subtypes. Vet Parasitol 177(3-4):378–382
Langkjær RB, Vigre H, Enemark HL, Maddox-Hyttel C (2007) Molecular and phylogenetic characterization of Cryptosporidium and Giardia from pigs and cattle in Denmark. Parasitology 134(3):339–350
Li F, Wang H, Zhang Z, Li J, Wang C, Zhao J, Hu S, Wang R, Zhang L, Wang M (2016) Prevalence and molecular characterization of Cryptosporidium spp. and Giardia duodenalis in dairy cattle in Beijing, China. Vet Parasitol 219:61–65
Li S, Zou Y, Wang P, Qu MR, Zhu XQ (2021) Prevalence and multilocus genotyping of Cryptosporidium spp. in cattle in Jiangxi Province, southeastern China. Parasitol Res 120(4):1281–1289
Liang N, Wu Y, Sun M, Chang Y, Lin X, Yu L, Hu S, Zhang X, Zheng S, Cui Z (2019) Molecular epidemiology of Cryptosporidium spp. in dairy cattle in Guangdong Province, South China. Parasitology 146(1):28–32
Liu A, Hong J, Wang E, Liu J, Xiao L, Shen Y, Li Y, Zhang W, Hong L (2011) Molecular identification and distribution of Cryptosporidium and Giardia duodenalis in raw urban wastewater in Harbin, China. Parasitol Res 109(3):913–918
Liu A, Wang R, Li Y, Zhang L, Shu J, Zhang W, Feng Y, Xiao L, Ling H (2009) Prevalence and distribution of Cryptosporidium spp. in dairy cattle in Heilongjiang Province, China. Parasitol Res 105(3):797–802
Lobo ML, Xiao L, Antunes F, Matos O (2009) Occurrence of Cryptosporidium and Giardia genotypes and subtypes in raw and treated water in Portugal. Lett Appl Microbiol 48(6):732–737
Ma J, Li P, Zhao X, Xu H, Wu W, Wang Y, Guo Y, Wang L, Feng Y, Xiao L (2015) Occurrence and molecular characterization of Cryptosporidium spp. and Enterocytozoon bieneusi in dairy cattle, beef cattle and water buffaloes in China. Vet Parasitol 207(3-4):220–227
Murakoshi F, Xiao L, Matsubara R, Sato R, Kato Y, Sasaki T, Fukuda Y, Tada C, Nakai Y (2012) Molecular characterization of Cryptosporidium spp. in grazing beef cattle in Japan. Vet Parasitol 187(1-2):123–128
Ni HB, Sun YZ, Qin SY, Wang YC, Zhao Q, Sun ZY, Zhang M, Yang D, Feng ZH, Guan ZH (2021) Molecular detection of Cryptosporidium spp. and Enterocytozoon bieneusi infection in wild rodents from six provinces in China. Front Cell Infect Microbiol 11:783508
Paul S, Chandra D, Tewari A, Banerjee P, Ray D, Raina O, Rao J (2009) Prevalence of Cryptosporidium andersoni: a molecular epidemiological survey among cattle in India. Vet Parasitol 161(1-2):31–35
Peng MM, Wilson ML, Holland RE, Meshnick SR, Lal AA, Xiao L (2003) Genetic diversity of Cryptosporidium spp. in cattle in Michigan: implications for understanding the transmission dynamics. Parasitol Res 90(3):175–180
Plutzer J, Karanis P (2007) Genotype and subtype analyses of Cryptosporidium isolates from cattle in Hungary. Vet Parasitol 146(3-4):357–362
Qi M, Fang Y, Wang X, Zhang L, Wang R, Du S, Guo Y, Jia Y, Yao L, Liu Q (2015a) Molecular characterization of Cryptosporidium spp. in pre-weaned calves in Shaanxi Province, north-western China. J Med Microbiol 64(1):111–116
Qi M, Wang H, Jing B, Wang D, Wang R, Zhang L (2015b) Occurrence and molecular identification of Cryptosporidium spp. in dairy calves in Xinjiang, Northwestern China. Vet Parasitol 212(3-4):404–407
Qi M, Zhang K, Huang M, Wang S, Xu C, Wang T, Jing B, Li J (2020) Longitudinal detection of Cryptosporidium spp. in 1–10-week-old dairy calves on a farm in Xinjiang, China. Parasitol Res 119(11):3839–3844
Rieux A, Chartier C, Pors I, Paraud C (2013) Dynamics of excretion and molecular characterization of Cryptosporidium isolates in pre-weaned French beef calves. Vet Parasitol 195(1-2):169–172
Ryan U, Fayer R, Xiao L (2014) Cryptosporidium species in humans and animals: current understanding and research needs. Parasitology 141(13):1667–1685
Ryan UM, Feng Y, Fayer R, Xiao L (2021) Taxonomy and molecular epidemiology of Cryptosporidium and Giardia–a 50 year perspective (1971–2021). Int J Parasitol 51:1099–1119
Sangster L, Blake DP, Robinson G, Hopkins TC, Sa RC, Cunningham AA, Chalmers RM, Lawson B (2016) Detection and molecular characterisation of Cryptosporidium parvum in British European hedgehogs (Erinaceus europaeus). Vet Parasitol 217:39–44
Santín M, Trout JM (2007) Livestock Cryptosporidium and cryptosporidiosis. CRC Press, pp 451–484
Santoro A, Dorbek-Kolin E, Jeremejeva J, Tummeleht L, Orro T, Jokelainen P, Lassen B (2019) Molecular epidemiology of Cryptosporidium spp. in calves in Estonia: high prevalence of Cryptosporidium parvum shedding and 10 subtypes identified. Parasitology 146(2):261–267
Sarkar R, Kattula D, Francis MR, Ajjampur SS, Prabakaran AD, Jayavelu N, Muliyil J, Balraj V, Naumova EN, Ward HD (2014) Risk factors for cryptosporidiosis among children in a semi urban slum in southern India: a nested case-control study. Am J Trop Med Hyg 91(6):1128
Smith R, Clifton-Hadley F, Cheney T, Giles M (2014) Prevalence and molecular typing of Cryptosporidium in dairy cattle in England and Wales and examination of potential on-farm transmission routes. Vet Parasitol 204(3-4):111–119
Sulaiman IM, Hira PR, Zhou L, Al-Ali FM, Al-Shelahi FA, Shweiki HM, Iqbal J, Khalid N, Xiao L (2005) Unique endemicity of cryptosporidiosis in children in Kuwait. J Clin Microbiol 43(6):2805–2809
Tarekegn ZS, Tigabu Y, Dejene H (2021) Cryptosporidium infection in cattle and humans in Ethiopia: a systematic review and meta-analysis. Parasite Epidemiology and Control 14:e00219
Thomson S, Innes EA, Jonsson NN, Katzer F (2019) Shedding of Cryptosporidium in calves and dams: evidence of re-infection and shedding of different gp60 subtypes. Parasitology 146(11):1404–1413
Tyzzer E (1907) A sporozoan found in the peptic glands of the common mouse. Proc Soc Exp Biol Med 5(1):12–13
Uran-Velasquez J, Alzate JF, Farfan-Garcia AE, Gomez-Duarte OG, Martinez-Rosado LL, Dominguez-Hernandez DD, Rojas W, Galvan-Diaz AL, Garcia-Montoya GM (2022) Multilocus sequence typing helps understand the genetic diversity of Cryptosporidium hominis and Cryptosporidium parvum isolated from Colombian patients. Plos one 17(7):e0270995
Wang G, Wang G, Li X, Zhang X, Karanis G, Jian Y, Ma L, Karanis P (2018) Prevalence and molecular characterization of Cryptosporidium spp. and Giardia duodenalis in 1–2-month-old highland yaks in Qinghai Province, China. Parasitol Res 117(6):1793–1800
Wang R, Ma G, Zhao J, Lu Q, Wang H, Zhang L, Jian F, Ning C, Xiao L (2011a) Cryptosporidium andersoni is the predominant species in post-weaned and adult dairy cattle in China. Parasitol Int 60(1):1–4
Wang R, Wang H, Sun Y, Zhang L, Jian F, Qi M, Ning C, Xiao L (2011b) Characteristics of Cryptosporidium transmission in preweaned dairy cattle in Henan. China. J Clin Microbiol 49(3):1077–1082
Wang R, Zhang L, Axén C, Bjorkman C, Jian F, Amer S, Liu A, Feng Y, Li G, Lv C (2014) Cryptosporidium parvum IId family: clonal population and dispersal from Western Asia to other geographical regions. Sci Rep 4(1):4208
Wang R, Zhao G, Gong Y, Zhang L (2017) Advances and perspectives on the epidemiology of bovine Cryptosporidium in China in the past 30 years. Front Microbiol 8:1823
Wang SN, Sun Y, Zhou HH, Lu G, Qi M, Liu WS, Zhao W (2020a) Prevalence and genotypic identification of Cryptosporidium spp. and Enterocytozoon bieneusi in captive Asiatic black bears (Ursus thibetanus) in Heilongjiang and Fujian provinces of China. BMC Vet Res 16(1):1–7
Wang T, Wei Z, Zhang Y, Zhang Q, Zhang L, Yu F, Qi M, Zhao W (2022) Molecular detection and genetic characterization of Cryptosporidium spp. and Enterocytozoon bieneusi in captive Asiatic black bears (Ursus thibetanus) in Heilongjiang and Fujian provinces of China. BMC Vet Res 16(1):1–7
Wang Y, Cao J, Chang Y, Yu F, Zhang S, Wang R, Zhang L (2020b) Prevalence and molecular characterization of Cryptosporidium spp. and Giardia duodenalis in dairy cattle in Gansu, northwest China. Parasite 27:62
Watanabe Y, Yang CH, Ooi HK (2005) Cryptosporidium infection in livestock and first identification of Cryptosporidium parvum genotype in cattle feces in Taiwan. Parasitol Res 97(3):238–241
Wu Y, Zhang K, Zhang Y, Jing B, Chen Y, Xu C, Wang T, Qi M, Zhang L (2020) Genetic diversity of Cryptosporidium parvum in neonatal dairy calves in Xinjiang, China. Pathogens 9(9):692
Yang X, Huang N, Jiang W, Wang X, Li N, Guo Y, Kváč M, Feng Y, Xiao L (2020) Subtyping Cryptosporidium ryanae: a common pathogen in bovine animals. Microorganisms 8(8):1107
Zhang W, Wang R, Yang F, Zhang L, Cao J, Zhang X, Ling H, Liu A, Shen Y (2013) Distribution and genetic characterizations of Cryptosporidium spp. in pre-weaned dairy calves in Northeastern China’s Heilongjiang Province. PLoS One 8(1):e54857
Zhang XX, Tan QD, Zhou DH, Ni XT, Liu GX, Yang YC, Zhu XQ (2015) Prevalence and molecular characterization of Cryptosporidium spp. in dairy cattle, northwest China. Parasitol Res 114(7):2781–2787
Zhao GH, Ren WX, Gao M, Bian QQ, Hu B, Cong MM, Lin Q, Wang RJ, Qi M, Qi MZ (2013) Genotyping Cryptosporidium andersoni in cattle in Shaanxi province, Northwestern China. PloS one 8(4):e60112
Zhao L, Chai HL, Wang MY, Zhang ZS, Han WX, Yang B, Wang Y, Zhang S, Zhao WH, Ma YM, Zhan YJ, Wang LF, Ding YL, Wang JL, Liu YH (2023) Prevalence and molecular characterization of Cryptosporidium spp. in dairy cattle in Central Inner Mongolia, Northern China. BMC Vet Res 19:134
Zhao W, Wang R, Zhang W, Liu A, Cao J, Shen Y, Yang F, Zhang L (2014) MLST subtypes and population genetic structure of Cryptosporidium andersoni from dairy cattle and beef cattle in northeastern China’s Heilongjiang Province. PLoS One 9(7):e102006
Zhao W, Zhou H, Huang Y, Xu L, Rao L, Wang S, Wang W, Yi Y, Zhou X, Wu Y (2019) Cryptosporidium spp. in wild rats (Rattus spp.) from the Hainan Province, China: molecular detection, species/genotype identification and implications for public health. Int J Parasitol: Parasites and Wildlife 9:317–321
Zhong Z, Dan J, Yan G, Tu R, Tian Y, Cao S, Shen L, Deng J, Yu S, Geng Y (2018) Occurrence and genotyping of Giardia duodenalis and Cryptosporidium in pre-weaned dairy calves in central Sichuan province, China. Parasite 25:45
CRediT authorship contribution
LZ, MYW, and YHL conceived and designed the study and critically revised the manuscript. LFW, LZ, MYW, YW, SZ, and YHL performed the samples collection. MYW and YHL prepared Fig. 1. MYW, LFW, YW, SZ, ZSZ, HLC, WJF, CY, YLD, JLW, and JS conducted the laboratory experiments. All the authors read and approved the final manuscript.
Funding
This study was funded by the National Natural Science Foundation of China (32260887) and supported by the National Natural Science Foundation of Inner Mongolia (2022MS03023).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
Our study was performed in strict accordance with the international standards published in the Guide to the Feeding, Management and Use of Experimental Animals (8th Edition) and followed the Regulations on the Management of Experimental Animals and other relevant laws and regulations. The study was approved by the Biomedical Research Ethics Committee of Inner Mongolia Agricultural University (approval no. 2020 [081]). Additionally, permission was obtained from the farm owners prior to specimen collection, and all efforts were made to minimize animal suffering.
Competing interests
The authors declare no competing interests.
Additional headings
All authors consent to participate and consent to publish.
Additional information
Section Editor: Lihua Xiao
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, L., Wang, M., Wang, L. et al. Prevalence and molecular characterization of Cryptosporidium spp. in dairy and beef cattle in Shanxi, China. Parasitol Res 123, 8 (2024). https://doi.org/10.1007/s00436-023-08058-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00436-023-08058-0