Abstract
Biomphalaria spp. snails are intermediary hosts of Schistosoma mansoni, etiologic agent of intestinal schistosomiasis, one of the most important neglected tropical diseases. Biomphalaria straminea is an important intermediary host that possess a different phenotype to parasite infection but shows a large geographic distribution and high capacity of new ecologic niche invasion. Our purpose was to characterize for the first time the differentially expressed proteome in B. straminea during two times intervals after primary and secondary exposure to S. mansoni. The hemolymph was collected at 1 and 15 days after primary and secondary exposure of snails to the parasite. Total proteins were extracted and digested with trypsin. LC–MS/MS label-free quantification was performed and analyzed using Maxquant and Perseus software. Proteins were identified and annotated using Blast2GO tools. After 1 day of exposure, most of upregulated proteins are hemoglobin type 2, C and H type lectins, molecules related to cell adhesion, and response to oxidative stress. After 15 days, we found a similar pattern of upregulated proteins but some fibrinogen-related proteins (FREPs) and TEPs homologs were downregulated. Regarding the differentially expressed proteins during secondary response, the principal immune-related proteins upregulated were C and H type lectins, cellular adhesion molecules, biomphalysin, and FREP3. We noted a several upregulated biological processes during both responses that could be the one of the key points of efficacy in the immune response to parasite. Our data suggests different immune mechanisms used by B. straminea snails challenged with S. mansoni.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biomphalaria is a genus of pulmonary mollusks that inhabit freshwater environments. These planorbid snails are spread around the world, occupying mainly South America and Africa. Some species of Biomphalaria, such as Biomphalaria glabrata and Biomphalaria straminea are responsible for the transmission Schistosoma mansoni parasite in South America by acting as an intermediate host of this human pathogen. S. mansoni belongs to a group of flatworm species of the genus Schistosoma that are etiological agents of schistosomiasis that affects millions of people worldwide. Many factors contribute to the schistosomiasis establishment in a region. One of the crucial aspects is the presence of Biomphalaria snails (Colley et al. 2014).
The immune response of Biomphalaria to infection with S. mansoni is complex. The defense system of snails is translated into several molecular interactions through components such as immune receptors, immune effectors, reactive oxygen species (ROS), proteases, and antimicrobial proteins (Coustau et al. 2015). Some studies described a compatibility profile between the molecules expressed by the snail and the molecules expressed by the parasite (Mitta et al. 2012), which is responsible for different susceptibility profiles to the parasite infection depending on the snail species involved.
Expressed proteins by B. glabrata’s immune system during interaction with S. mansoni are the purpose of several studies with different approaches, mainly transcriptomics and proteomics methods (Adema et al. 2010; Bouchut et al. 2006; Vergote et al. 2005). Like in other organisms, the immune response of Biomphalaria to S. mansoni infection is divided into cellular and humoral responses (Negrão-Corrêa et al. 2012). The cellular response is based mainly on the sporocyst encapsulation by hemocytes in hemolymph and molecules that modulate that cellular response (Pila et al. 2017). The humoral response in Biomphalaria sp. is composed of a molecular variety like receptors, signalization, and effectors against pathogens.
In cell-free plasma, signal molecules, recognition receptors, and effectors against pathogens can be found as part of the snail’s humoral response. Among such components, the most studied during the immune response to schistosome infection are lectins (Wu et al. 2017).
Some of these lectins are known as fibrinogen-related proteins (FREPs) and play an essential role during snails defense, where the FREP knockdown modifies the parasite susceptibility phenotype (Hanington et al. 2012). In addition, some FREPs can form immunocomplexes with Thioester-containing proteins (TEPs) and aerolysin (biomphalysins) (Gordy et al. 2015; Li et al. 2020). C-type lectin-related proteins (CREPs) and galectin-related proteins (GREPs), similar to FREPs, also have relevant functions in the snail mechanisms through the processes of opsonization and agglutination of the parasite (Tetreau et al. 2017; Wu et al. 2017).
Besides the classic innate immune response reported in the conventional investigations, there is a change in the response profile when these organisms are challenged to a second exposure to the parasite, which fostered the appearance of hypotheses about memory or secondary response. Many studies have been carried out to clarify this more refined and specific response. The immune priming protocol is the most applied to evaluate this type of response during the parasite-host interaction in some invertebrates (Contreras-Garduño et al. 2016; Portela et al. 2013). In B. glabrata, it was observed that snails previously exposed to parasite infection can acquire resistance to a second challenge, with differences in the transcriptomic and proteomic profile between the primary and secondary responses (Pinaud et al. 2016).
Although S. mansoni can evolve in different Biomphalaria species, there is a predominance of studies with B. glabrata. Few studies limited their work to only investigate the importance of other species such as B. straminea for maintaining the parasite’s life cycle. The ultrastructural characterization of B. straminea hemocytes showed a similarity of cellular morphological pattern to B. glabrata hemocytes. Despite that, the immune response patterns against S. mansoni infection differ in these two species, suggesting a relationship between resistance profile and differentially expressed molecules during the infection in B. straminea (Cavalcanti et al. 2012).
Analyzing gene expression of FREPs candidates, we can see these differences in the humoral response between the species B. glabrata and B. straminea during exposure to S. mansoni. Despite belonging to the same genus, it was not possible to detect some of these proteins in the species B. straminea yet, since these molecules are highly variable and may have different sequences between organisms (de Melo et al. 2019).
Despite having a more resistant phenotype than B. glabrata, B. straminea has a fundamental role in disseminating and maintaining the S. mansoni cycle in different niches (Lin et al. 2020). Epidemiological researches show that these snails inhabit tropical regions, being found mainly in the Americas and China, presenting patterns of habitat changes for regions with a subtropical climate, where such climatic adaptations provide a greater diffusion of the snail and, consequently, greater risk of dispersal of the parasite (Scholte et al. 2012; Yang et al. 2018).
In this study, we aim to identify variations in the B. straminea proteome exposed to S. mansoni, with a particular focus on immune relevant proteins in the hemolymph. We performed a label-free proteomic analysis to identify the differentially expressed proteins in the primary and secondary response to the parasite, resulting in the identification respectively of 39 and 35 proteins, contributing to a more comprehensive understanding of this species’ immune mechanisms.
Materials and methods
Biomphalaria straminea snails and Schistosoma mansoni miracidia
B. straminea snails were reared on Immunopathology Laboratory Keizo Asami-LIKA/UFPE vivarium in 24 °C chlorine-free water and fed with lettuce ad libitum.
Mice were individually infected with 120 cercariae of the LE strain of S. mansoni and kept in an experimental animal facility, following the approval of the Ethics and Use of Animals Committee (CEUA) under protocol number 104/2016 of the Aggeu Magalhães Institute—FIOCRUZ/PE. Feces obtained from infected mice were macerated in distilled water, filtered, and then exposed to artificial light and heat for 2 h, allowing the miracidia to hatch. After that, the miracidia were separated, counted, and placed in 12-well plates.
Experimental exposition and “immune priming” protocol
Exposures to the parasite were performed by placing the snails in plates with individual wells where each snail was in contact with 10 S. mansoni miracidia for 1.5 h under artificial light. After exposure, confirmation of the penetration of miracidia in the B. straminea snails was carried out by certifying the absence of miracidia in the well, with the aid of a stereomicroscope.
To verify the immune priming process, a protocol described by Portela et al. (2013) was followed with some modifications. Briefly, the snails were individually exposed to 10 S. mansoni miracidia, and then 25 days later, were subjected to a new exposure. The snails were divided into four experimental groups containing 5 snails per group (Fig. 1). All experiments were carried out in triplicate.
Overview of experimental methodology. GA, individually exposed snails to 10 miracidia and 25 days after exposed in a second time to 10 miracidia; GB, individually exposed snails to 10 miracidia and 25 days after submitted only to infection stimulus without miracidia; GC, snails submitted only to infection stimulus without miracidia and 25 days after exposed to 10 miracidia; CO, snails submitted only to infection stimulus without miracidia and 25 days after exposed again to infection stimulus without miracidia; The infection stimulus is the same conditions of miracidia exposition without miracidia. dpe = days post-exposure, dpr = days post-reinfection
We used samples from the GC group (snails exposed only once to the parasite) to assess the primary response. That group corresponds to 1 day after primo exposure (1dpe) and 15 days after the primo response (15 dpe). To investigate the secondary immune response, we used GA group samples (snails exposed and reexposed) corresponding to 1 dpr (1 day post-second exposition) and 15 dpr (15 days post-second exposure). The GB and CO groups representing snails exposed once at different times and snails never exposed (naive), respectively.
Hemolymph collection
The snails had their shells cleaned with 70% alcohol and dried with absorbent paper. We collected the hemolymph by cephalopodal puncture using 27G microneedles for drilling and siliconized tips to prevent cell adhesion and packed in siliconized microtubes with a mix of protease inhibitors (Cytiva- GE Healthcare) and samples were stored at − 80 °C until use.
We collected the hemolymph at 1 day (GA1, GB1, GC1, and CO1) and 15 days (GA15, GB15, GC15, and CO15) after reexposure.
Protein extraction
B. straminea hemolymph was thawed and frozen twice in liquid nitrogen to assist the lysis of hemocytes, then protein extraction buffer (SDS 12%; 0.3 M DTT; 0.3 M Tris–HCl; pH 7.5) was added to hemolymph (1/2; v/v). The samples were heated for 5 min at 95 °C, mixed by vortex and placed in an ultrasonic bath for 1 h. The samples were centrifuged at 14000 g for 5 min, and the supernatant containing the proteins from each sample was stored separately. The protein quantification was performed in triplicate using 2D QuantKit (Cytiva-GE Healthcare) following the manufacturer’s protocol.
Protein preparing
For proteomic analysis, 40 µg of protein extract from each experimental group were used and fractionated in SDS-PAGE with 12% polyacrylamide gel and stained with Coomassie Blue R-250. The proteins were excised from the gel, bleached with 25 mM ammonium bicarbonate (NH4HCO3), 50% ethanol, followed by total dehydration with 100% ethanol, and dried in a vacuum concentrator. Proteins were reduced with 10 mM DTT, 50 mM NH4 HCO3 for 1 h at 56 °C, and then it was alkylated with 50 mM iodoacetamide, 50 mM NH4 HCO3 for 1 h at room temperature. Samples were washed in 50 mM NH4 HCO3 and dehydrated in 100% ethanol, twice. Then, incubated in trypsin 12.5 ng/ml, 50 mM NH4 HCO3 at 37 °C for 18 h. After digestion, the peptides were extracted from the gel matrix by washing twice in 30% MeCN, 3% trifluoroacetic acid, and at the end twice in 100% MeCN. The peptides were concentrated in a vacuum concentrator through evaporation of the MeCN and desalination using C18 columns made “in house.”
Label-free LC-MS/MS quantification
Peptides were analyzed in an Ultimate 3000 RSLCnano liquid chromatography, coupled to the Fusion Lumos mass spectrometer (Thermo Scientific) (mass spectrometry platform RPT02H—Carlos Chagas Institute—FIOCRUZ PARANA). Initial chromatography was carried out in a 250nL/min flow of 5 to 40% MeCN in 1% formic acid, 5% DMSO in a gradient of 120 min. The spectrometer was operated in a data-dependent (DDA) mode, changing from the acquisition of MS to MS/MS. MS1 spectra were acquired on the Orbitrap analyzer with a resolution of 120,000 at 200 m/z. The most intense ions were isolated with a target value of 50,000, fragmented by HCD, and analyzed on Orbitrap with a resolution of 15,000 at 200 m/z. Each sample was analyzed in a technical duplicate.
Data analysis
Raw data were analyzed using the MaxQuant software (version 1.5.5.1) (Cox and Mann 2008), using the Andromeda algorithm to search databases using a target-decoy approach (Cox et al. 2011). MS data were searched in a B. straminea assembled transcriptome (paper in preparation) containing 130,731 protein isoforms. The reversed sequences of the proteins were used as the decoy database. Carbamidomethylation of cysteine was defined as fixed modification and methionine oxidation as a variable modification of the peptides. Only peptides containing at least seven amino acids were accepted, and an FDR of 0.01 was applied to both peptides and proteins. The protein groups identified by MaxQuant were analyzed in the Perseus software (version 1.6.0.7) (Tyanova et al., 2016). The label-free quantification (LQF) intensities were transformed to log10 before obtaining each protein ratio in the comparison between groups. Proteins were accepted that presented valid quantification data in at least three samples from each group and at least two unique peptides.
Differential expression analysis in the primary response was performed using Student’s t-test between two samples. Analysis of variance (ANOVA) was used to evaluate innate immune memory between multiple groups to. A permutation-based FDR rate (FDR < 0.05) was used to correct p-values in all cases. Differentially expressed proteins (DEPs) with a minimum fold change (s0) equal to 0.5 and a q-value < 0.05 were considered upregulated and downregulated, respectively. The sequences of the identified proteins were annotated using INTERPROSCAN 5 and BLAST2GO® (BioBam) to analyze and predict a signal peptide, conserved domains, and transmembrane domains prediction of Gene Ontology (GO) and pathways.
Results
The primary immune response of Biomphalaria straminea exposed to Schistosoma mansoni
To investigate the differential protein expression of the primary immune response to the parasite, we used the GC and CO groups, while the GC group represents the immune response of snails exposed only once to the parasite, with the GC1 samples representing the response of 1 day post-exposure (1dpe) and GC15 the 15 day post-exposure response (15 dpe). The CO1 and CO15 samples, on the other hand, correspond to hemolymph of snails that have only undergone exposure stress, corresponding to naive snails (Fig. 1).
Respectively, groups 1 dpe and 15 dpe showed 43 and 48 valid proteins to differential statistical analysis (Supplementary 1). A total of 39 differentially expressed proteins (DEPs) were identified in the primary response; 8 exclusives to the 1 dpe samples; and 14 exclusives to the 15 dpe group (Fig. 2A) (Supplementary 1 and 2). Also, we identified 7 unique proteins in the 1 dpe snail samples and 7 in the 15 dpe samples (Fig. 2B) (Supplementary 2), suggesting that their respective abundances were higher after exposure, whereas in the control group, they are suppressed or undetectable by the methodology used.
Venn diagrams of differentially expressed proteins in primary response. Blue set represents the differentially expressed proteins exclusive 1dpe and yellow set represents differentially expressed proteins exclusive 15dpe (A). Venn diagram of proteins identified in samples of 1dpe exposed, 1dpe control, 15dpe exposed, and 15dpe control. Green set represents proteins only identified in 1dpe and pink set represents proteins only identified in 15dpe (B)
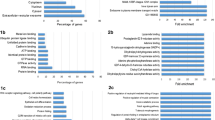
For the 1 dpe group, out of 43 valid proteins, 27 proteins were found to be differentially expressed. Among them, 18 are upregulated and 7 downregulated (Table 1). To investigate the possible functions of these proteins, they were grouped according to their main GO terms. The main molecular functions (GO) upregulated were as follows: carbohydrate binding (GO: 0,030,246), ATP binding (GO: 0,005,524), and peroxidase activity (GO: 0,004,601), while a predominance of oxygen carrier activity (GO: 0,005,344) and signaling functions of transmembrane receivers (GO: 0,004,888) were among downregulated proteins (Fig. 3A). Regarding the biological process (GO) in 1 dpe, the terms cell adhesion (GO: 0,007,155) and response to oxidative stress (GO: 0,006,979), oxygen transport (GO: 0,015,671), iron ions transport (GO: 0,006,826), and transmembrane ion transport (GO: 0,034,220) were identified as upregulated while the term oxygen transport (GO: 0,015,671) was the most frequent in downregulated proteins (Supplementary 2) (Fig. 3A).
In 15 dpe, out of 48 valid proteins, 31 proteins were differentially expressed; 26 upregulated; and 5 downregulated proteins (Table 2). The most common molecular functions (GO) upregulated were oxygen binding (GO: 0,019,825), carbohydrate binding (GO: 0,030,246), nucleotide binding (GO: 0,005,524), and peroxidase activity (GO: 0,004,601), while in the downregulated were carbohydrate binding (GO: 0,030,246) and endopeptidase inhibitor activity (GO: 0,004,866). The most frequent biological process (GO) terms in upregulated proteins at 15 dpe were oxygen transport (GO: 0,015,671), pathogenesis (GO: 0,009,405), and response to oxidative stress (GO: 0,006,979); however, only the term cell adhesion (GO: 0,007,155) was identified among the downregulated proteins (Supplementary 2) (Fig. 3B).
It is interesting to note that some proteins did not have their GO mapping properly completed, but their conserved domains were adequately identified. Such proteins belong mainly to the families of lectins, FREPs, clotting factors, and aerolysin, showing immunological relevance.
Proteomic analysis of immune primed Biomphalaria straminea challenged with Schistosoma mansoni
In 1 day post-reexposure (1dpr) group, we identified 55 proteins out of which 35 were DEPs (q-value < 0.05) (Table 3, Supplementary 2). We grouped these DEPs according to the pattern of differential expression in cluster 1: proteins upregulated exclusively following secondary exposure (GA1); cluster: 2, proteins that showed upregulation during any primary exposure (GB1 or GC1) and increased expression following secondary exposure (GA1); cluster 3: proteins downregulated in the primary exposure (GC1 or GB1) and decreased expression after secondary exposure (GA1); and cluster 4: downregulated proteins exclusively following secondary exposure (GA1) (Fig. 4A).
Heatmap of differentially expressed proteins in secondary response. A Proteins classified by family domain in Biomphalaria straminea 1 day after a secondary exposition to Schistosoma mansoni. Yellow to blue-scaled indicate the ratio of fold change value from lowest to higher. Four clusters are identified: cluster 1 (upregulated proteins related exclusively to secondary response); cluster 2 (upregulated proteins in primary exposition and increased in secondary challenge); cluster 3 (downregulated proteins in primary response and decreased in secondary challenge); cluster 4 (downregulated proteins exclusively in secondary response). B Proteins classified by family domain in Biomphalaria straminea 15 days after a secondary exposition to Schistosoma mansoni. Yellow to blue-scaled indicate the ratio of fold change value from lowest to higher. Three clusters are identified: cluster 1 (upregulated proteins related exclusively to secondary response); cluster 2 (upregulated proteins in primary exposition and increased in secondary challenge); and cluster 3 (downregulated proteins exclusively in secondary response)
In 15 days after reexposure (15 dpr), we identified 54 proteins. Among these, 10 are DEPs in 15 dpr compared to the control group (Table 4). DEPs were grouped according to the differential expression pattern in cluster 1: proteins upregulated exclusively following secondary exposure (GA15); cluster 2: upregulated proteins on primary exposure and increased expression after secondary exposure; and cluster 3: downregulated proteins exclusively following secondary exposure (Fig. 4B).
Discussion
Proteins of the primary response of B. straminea exposed to S. mansoni
As in B. glabrata, B. straminea hemoglobins are the most abundant proteins in the hemolymph. Most of these elements showed upregulation in B. straminea after 1 and 15 days post-exposure to S. mansoni. In the first group (1 dpe), we observed a greater variety of super expressed isoforms corresponding to type 2 hemoglobin. In the second time post-exposure (15 dpe), there is a change in the hemoglobin isoforms, where type 1 hemoglobin is more identified among upregulated proteins. Tetreau et al. (2017), using a proteomic approach in an investigation of the plasma of B. glabrata exposed in vitro to different pathogens, detected exclusively type 2 hemoglobin in the interaction with S. mansoni and Echinostoma caproni. The authors also referred to the presence of different molecular weights of these proteins, considered cleavage subproducts functionally actives, with unclear immune capacity this genus.
Hemocyanin is also another molecule that was found in high abundance in Biomphalaria’s hemolymph. Our results identified a significant increase in the abundance of this protein after exposure to S. mansoni. Hemocyanin functionality goes beyond carrying oxygen. This molecule is part of the type 3 copper proteins superfamily that has an important enzymatic activity in the phenoloxidase and melanization pathways related to the innate immune response (Coates and Decker 2017; Coates and Nairn 2014). Apparently, this is not its primary function in Biomphalaria snails, since in previous studies, it was not possible to detect a wide variation of phenoloxidase activity in B. straminea early response to S. mansoni infection, through the L-DOPA pathway (de Melo et al. 2019). In addition to the functions already described, hemocyanin still acts as a precursor to antimicrobial and antiviral peptides (Dolashka and Voelter 2013; Peña and Adema 2016; Qin et al. 2018). These evidences and its post-exposure expression variations suggest a strong involvement in the primary immune response.
Other molecules differentially expressed were the enzymes peroxidase and glutathione peroxidase, which showed upregulation in both times post-exposure to the parasite. The innate immune response pathway by enzymes related to oxidative stress is described in different invertebrate groups. It can be activated in following biotic and abiotic stress, being also precedents and synergists of other pathways such as cell adhesion and encapsulation (Reference). In B. glabrata, reactive oxygen species (ROS) can rapidly destroy the S. mansoni sporocyst when incubated in cell-free plasma, being produced mainly by strains with a resistant phenotype to infection by the parasite, showing that this pathway is quite efficient for trematodes (Bender et al. 2005; Fogarty et al. 2019). In addition to the importance of ROS in B. glabrata, an increase in the production of these molecules was also observed in other relationships of mollusk/trematode, such as Lymnaea stagnalis when exposed to Echinoparyphium aconiatum cercariae (Abbas et al. 2019; Buchmann 2014; Mitta et al. 2017).
One of the most relevant and studied proteins during the immune response of invertebrates is the lectin family. These molecules can bind to carbohydrates on the surface of pathogens and act in a signaling and opsonizing manner against various agents (Fujita et al. 2004). C-type lectins (CTL) are relevant in the defense pathway against trematodes, such as S. mansoni (Coustau et al. 2015). The CTL domain confers this family the ability to recognize pathogen-associated molecular patterns (PAMPs), antimicrobial activity, and induce phagocytosis or encapsulation processes. The latter is through the connection and formation of complexes with other proteins, like integrins (Wang et al. 2011, 2014). Variations in the CTL expression were observed in the B. glabrata proteome 15 days after primary exposure (Pinaud et al. 2016), and it was expressed in the plasma of both the susceptible (NMRI) and resistant (BS-90) strains of B. glabrata (Wu et al. 2017). Other studies show an increased expression of these lectins of B. glabrata in a few hours after infection by S. mansoni (Ittiprasert et al. 2010). In our results, we identified C-type lectins with increased expression in 1 dpe and 15 dpe in the primary response of B. straminea. These findings show a wide diversity of representatives of this protein family between species, in addition to suggesting that the B. straminea recognition pathways are activated continuously after exposure to the parasite.
Curiously, we identified upregulated H-type lectins in 1 and 15 dpe samples. There are few descriptions of this family among studies involving invertebrate immune response, being first described in snails Helix pomatia as H. pomatia agglutinin (HPA), with relationship to immunoprotective and reproductive aspects in the species (Sanchez et al. 2006). The H-type lectin domain can bind with high specificity to galactose (Gal) or N-acetylgalactosamine (GalNAc) carbohydrates, types expressed particularly on the surface of cancer cells, being the target of possible biomedical applications (Pietrzyk-Brzezinska and Bujacz 2020). In B. straminea-S. mansoni relationship, the expression of this specific lectin during the immune response may be evidence of a different recognition pathways used by this snail comparing with the model specie B. glabrata.
Some peculiarities were identified regarding the FREPs in B. straminea, a critical protein family in the B. glabrata-S. mansoni relationship. We detected homologs to FREPs in naive 1 dpe snail samples. In contrast, we also identified some other corresponding FREP isoforms exclusively in exposed snails and 15 dpe control samples (Supplementary 2). Portet et al. (2017), also using a label-free proteomics approach with B. glabrata, justified the absence of FREPs due to the ability of FREPs to bind to parasite antigens and precipitate being lost during the process of obtaining proteins. Our data corroborate in part with this hypothesis, considering we identified, in the 15 dpe group, the downgrade of a protein homolog to FREP12, present in the exposed snails, but with less abundance.
Immune primed Biomphalaria straminea proteomic response to Schistosoma mansoni
The secondary response, also called the innate immune response of memory, gives invertebrates success during the second contact with pathogens. The ability of some invertebrate clades to be exposed to a first challenge and then reshape their immune response in the face of a second challenge makes immune priming a relevant leap in the evolutionary process of these organisms (Sheehan et al. 2020). This issue has been studied for more than a decade in different types of pathogen-host interaction. It is evaluated from mechanisms such as microbiota interference in the Anopheles-Plasmodium relationship (Rodrigues et al. 2010), through the change in the type of immune response of the species B. glabrata with the presence of essential recognition molecules such as FREPs (Pinaud et al. 2016), until the discovery of extremely effector components presents in the secondary memory response, such as Biomphalysins (Galinier et al. 2013; Tetreau et al. 2017). To investigate the innate immune response of memory in the species B. straminea, we used an adapted method proposed by Portela et al. (2013), based on two-round of exposure with the parasite, together with several control groups (Fig. 1).
C-type and H-type lectins (Fig. 4A) are components that appear with high abundance in the primary response and increase their expression following secondary challenge in B. straminea. In 1 and 15 days after second exposure (GA1 and GA15), C-type lectins showed high expression (Fig. 4A and Fig. 4B). In proteomics and transcriptome approaches to evaluate B. glabrata snails’ secondary response, various isoforms of C-type lectins are found differentially expressed after the immune priming process (Pinaud et al. 2016; Tetreau et al. 2017). Likewise, transcripts related to this protein family are found upregulated in heterologous parasite infections, challenging B. glabrata to exposure and reexposure with different strains of S. mansoni (Pinaud et al. 2019). In B. straminea snails, CTLs seem to play an essential role during the innate and memory immune responses since they are detected in greater abundance during all times after exposure to the parasite.
Concerning H-type lectin, it has not been reported with differential expression in previous studies involving B. glabrata and S. mansoni, being identified during the primary and secondary response of B. straminea. The presence of this protein suggests that B. straminea uses different recognition molecules during the process of exposure to the parasite, which could be a key factor for the less susceptible phenotype of the species.
The change in the expression pattern of apolipoprotein B-100-like, downregulated 1 day after secondary challenge (Fig. 3A) and upregulated in 15 days after secondary challenge (Fig. 3B), suggests its relationship with the late secondary response. This protein was also found with decreased abundance in hemocytes from a susceptible strain of B. glabrata in primary interaction with S. mansoni sporocysts (Dinguirard et al. 2018). In addition to Biomphalaria snails, other studies describe the participation of apolipoprotein-like in other invertebrates such as insects and mollusks both in the primary immune response, as well as in secondary exposures to pathogens (Castillo et al. 2019; Rey-Campos et al. 2019; Stączek et al. 2018; Wu et al. 2017).
Regarding the humoral response profile, we noted in 15 dpe snails (GA15), mainly the presence of pathogen recognition receptors (lectins and FREP) and cytolytic agents (biomphalysin) (Fig. 4B). In this context, FREPs seem to play a critical role in the secondary response of B. straminea, once was upregulated only in samples of reexposed snails (Fig. 3B). FREP3 has a strong relationship with the phenotype of resistance of B. glabrata to trematodes (Hanington et al. 2012; Pila et al. 2017). This molecule may also be associated with the more resistant profile of the species B. straminea. Furthermore, the relevant increase in abundance only in B. straminea 15 dpe reinforces the idea of FREPs playing a key role during the secondary response.
We also highlight the expression pattern of biomphalysin in the secondary response in B. straminea, which was different from the profile previously described in B. glabrata. We observed an increase in the abundance of biomphalysin in the 15 dpr samples. However, Pinaud et al. (2016) reported that, in B. glabrata, this protein was not detected in the 15 days after the second challenged proteome. The authors justified that this component can be consumed quickly after exercising its role in the immune response. The different expression patterns of these immune relevant components’ give evidence of greater efficiency of innate immune response and secondary response in B. straminea.
The immunocomplexes formations between BgFREP3 and biomphalysin with other components, mainly BgTEP1, result in an efficiency of plasma-mediated response against S. mansoni by Biomphalaria (Li et al. 2020). The presence of two of the main proteins involved in these immunocomplexes in our data corroborates with the more significant responsiveness of B. straminea and can be the target of future studies for a better understanding of these specific mechanisms.
Although the knowledge about the immune response of B. straminea is limited, this work provides for the first time a proteomic overview of the host-parasite interaction during the primary and secondary immune response against S. mansoni. Several proteins identified in this study are homologous to those of the model species B. glabrata. However, other proteins, such as H-type lectins, need further elucidation of their structures and functions in B. straminea. Such data reinforce the necessity for additional studies involving this species using complementary strategies to add knowledge to the present work results.
Data availability
PRIDE dataset identifier PXD023681.
References
Abbas MN, Kausar S, Cui H (2019) The biological role of peroxiredoxins in innate immune responses of aquatic invertebrates. Fish Shellfish Immunol 89:91–97. https://doi.org/10.1016/j.fsi.2019.03.062
Adema CM, Hanington PC, Lun C-M, Rosenberg GH, Aragon AD, Stout BA, Richard MLL, Gross PS, Loker ES (2010) Differential transcriptomic responses of Biomphalaria glabrata (Gastropoda, Mollusca) to bacteria and metazoan parasites, Schistosoma mansoni and Echinostoma paraensei (Digenea, Platyhelminthes). Mol Immunol 47:849. https://doi.org/10.1016/j.molimm.2009.10.019
Bouchut A, Sautiere PE, Coustau C, Mitta G (2006) Compatibility in the Biomphalaria glabrata/Echinostoma caproni model: potential involvement of proteins from hemocytes revealed by a proteomic approach. Acta Trop 98:234–246. https://doi.org/10.1016/j.actatropica.2006.05.007
Buchmann K (2014) Evolution of innate immunity: clues from invertebrates via fish to mammals. Front Immunol 5:1–8. https://doi.org/10.3389/fimmu.2014.00459
Castillo MG, Humphries JE, Mourão MM, Marquez J, Montelongo CE, Gonzalez A, Montelongo CE (2019) Biomphalaria glabrata immunity: post-genome advances. Dev Comp Immunol 104:103557. https://doi.org/10.1016/j.dci.2019.103557
Cavalcanti MGS, Filho FC, Mendonça AMB, Duarte GR, Barbosa CCGS, De Castro CMMB, Alves LC, Brayner FA (2012) Morphological characterization of hemocytes from Biomphalaria glabrata and Biomphalaria straminea. Micron 43:285–291. https://doi.org/10.1016/j.micron.2011.09.002
Coates CJ, Decker H (2017) Immunological properties of oxygen-transport proteins: hemoglobin, hemocyanin and hemerythrin. Cell Mol Life Sci 74:293–317. https://doi.org/10.1007/s00018-016-2326-7
Coates CJ, Nairn J (2014) Diverse immune functions of hemocyanins. Dev Comp Immunol 45:43–55. https://doi.org/10.1016/j.dci.2014.01.021
Colley DG, Bustinduy AL, Secor WE, King CH (2014) Human schistosomiasis. Lancet 383:2253–2264. https://doi.org/10.1016/S0140-6736(13)61949-2
Contreras-Garduño J, Lanz-Mendoza H, Franco B, Nava A, Pedraza-Reyes M, Canales-Lazcano J (2016) Insect immune priming: ecology and experimental evidences. Ecol Entomol 41:351–366. https://doi.org/10.1111/een.12300
Coustau C, Gourbal B, Duval D, Yoshino TP, Adema CM, Mitta G (2015) Advances in gastropod immunity from the study of the interaction between the snail Biomphalaria glabrata and its parasites: a review of research progress over the last decade. Fish Shellfish Immunol 46:5–16. https://doi.org/10.1016/j.fsi.2015.01.036
Cox J, Mann M (2008) MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol 26:1367–1372. https://doi.org/10.1038/nbt.1511
Dolashka P, Voelter W (2013) Antiviral activity of hemocyanins. Invertebr Surviv J 10:120–127
Fogarty CE, Zhao M, McManus DP, Duke MG, Cummins SF, Wang T (2019) Comparative study of excretory–secretory proteins released by Schistosoma mansoni-resistant, susceptible and naïve Biomphalaria glabrata. Parasit Vectors 12:452. https://doi.org/10.1186/s13071-019-3708-0
Fujita T, Matsushita M, Endo Y (2004) The lectin-complement pathway - its role in innate immunity and evolution. Immunol Rev 198:185–202. https://doi.org/10.1111/j.0105-2896.2004.0123.x
Gordy MA, Pila EA, Hanington PC (2015) The role of fibrinogen-related proteins in the gastropod immune response. Fish Shellfish Immunol 46:39–49. https://doi.org/10.1016/j.fsi.2015.03.005
Ittiprasert W, Miller A, Myers J, Nene V, El-Sayed NM, Knight M (2010) Identification of immediate response genes dominantly expressed in juvenile resistant and susceptible Biomphalaria glabrata snails upon exposure to Schistosoma mansoni. Mol Biochem Parasitol 169:27–39. https://doi.org/10.1016/j.molbiopara.2009.09.009
Lin D, Zeng X, Sanogo B, He P, Xiang S, Du S, Zhang Y, Wang L, Wan S, Zeng X, Yang Y, Lv Z, Liang Y, Deng Z, Hui JHL, Yuan D, Ding T, Wu Z, Sun X (2020) The potential risk of schistosoma mansoni transmission by the invasive freshwater snail biomphalaria straminea in south china. PLoS Negl Trop Dis 14:1–19. https://doi.org/10.1371/journal.pntd.0008310
Mitta G, Adema CM, Gourbal B, Loker ES, Theron A (2012) Compatibility polymorphism in snail/schistosome interactions: from field to theory to molecular mechanisms. Dev Comp Immunol 37:1–8. https://doi.org/10.1016/j.dci.2011.09.002
Peña JJ, Adema CM (2016) The planorbid snail biomphalaria glabrata expresses a hemocyanin-like sequence in the albumen gland. PLoS ONE 11:1–17. https://doi.org/10.1371/journal.pone.0168665
Pietrzyk-Brzezinska AJ, Bujacz A (2020) H-type lectins – structural characteristics and their applications in diagnostics, analytics and drug delivery. Int J Biol Macromol 152:735–747. https://doi.org/10.1016/j.ijbiomac.2020.02.320
Pila EA, Li H, Hambrook JR, Wu X, Hanington PC (2017) Schistosomiasis from a snail’s perspective: advances in snail immunity. Trends Parasitol 33:845–857. https://doi.org/10.1016/j.pt.2017.07.006
Pinaud S, Portela J, Duval D, Nowacki FC, Olive MA, Allienne JF, Galinier R, Dheilly NM, Kieffer-Jaquinod S, Mitta G, Théron A, Gourbal B (2016) A shift from cellular to humoral responses contributes to innate immune memory in the vector snail Biomphalaria glabrata. PLoS Pathog 12:1–18. https://doi.org/10.1371/journal.ppat.1005361
Pinaud S, Portet A, Allienne JF, Belmudes L, Saint-Beat C, Arancibia N, Galinier R, Du Pasquier L, Duval D, Gourbal B (2019) Molecular characterisation of immunological memory following homologous or heterologous challenges in the schistosomiasis vector snail, Biomphalaria glabrata. Dev Comp Immunol 92:238–252. https://doi.org/10.1016/j.dci.2018.12.001
Portela J, Duval D, Rognon A, Galinier R, Boissier J, Coustau C, Mitta G, Théron A, Gourbal B (2013) Evidence for specific genotype-dependent immune priming in the lophotrochozoan biomphalaria glabrata snail. J Innate Immun 5:261–276. https://doi.org/10.1159/000345909
Qin Z, Babu VS, Wan Q, Muhammad A, Li J, Lan J, Lin L (2018) Antibacterial activity of hemocyanin from red swamp crayfish (Procambarus clarkii). Fish Shellfish Immunol 75:391–399. https://doi.org/10.1016/j.fsi.2018.02.010
Rey-Campos M, Moreira R, Gerdol M, Pallavicini A, Novoa B, Figueras A (2019) Immune tolerance in Mytilus galloprovincialis hemocytes after repeated contact with vibrio splendidus. Front Immunol 10:1–15. https://doi.org/10.3389/fimmu.2019.01894
Sanchez J-F, Lescar J, Chazalet V, Audfray A, Gagnon J, Alvarez R, Breton C, Imberty A, Mitchell EP (2006) Biochemical and structural analysis of Helix pomatia Agglutinin. J Biol Chem 281:20171–20180. https://doi.org/10.1074/jbc.M603452200
Scholte RGC, Carvalho OS, Malone JB, Utzinger J, Vounatsou P (2012) Spatial distribution of Biomphalaria spp., the intermediate host snails of Schistosoma mansoni, in Brazil. Geospat Health 6:S95–S101. https://doi.org/10.1016/j.jcv.2015.07.170
Sheehan G, Farrell G, Kavanagh K (2020) Immune priming: the secret weapon of the insect world. Virulence 11:238–246. https://doi.org/10.1080/21505594.2020.1731137
Stączek S, Zdybicka-Barabas A, Mak P, Sowa-Jasiłek A, Kedracka-Krok S, Jankowska U, Suder P, Wydrych J, Grygorczuk K, Jakubowicz T, Cytryńska M (2018) Studies on localization and protein ligands of Galleria mellonella apolipophorin III during immune response against different pathogens. J Insect Physiol 105:18–27. https://doi.org/10.1016/j.jinsphys.2017.12.009
Tetreau G, Pinaud S, Portet A, Galinier R, Gourbal B, Duval D (2017) Specific pathogen recognition by multiple innate immune sensors in an invertebrate. Front Immunol 8:1249. https://doi.org/10.3389/fimmu.2017.01249
Tyanova S, Temu T, Sinitcyn P, Carlson A, Hein MY, Geiger T, Mann M, Cox J (2016) The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat Methods 13:731–740. https://doi.org/10.1038/nmeth.3901
Vergote D, Bouchut A, Sautière PE, Roger E, Galinier R, Rognon A, Coustau C, Salzet M, Mitta G (2005) Characterisation of proteins differentially present in the plasma of Biomphalaria glabrata susceptible or resistant to Echinostoma caproni. Int J Parasitol 35:215–224. https://doi.org/10.1016/j.ijpara.2004.11.006
Wang L, Huang M, Zhang H, Song L (2011) The immune role of C-type lectins in molluscs. ISJ-Invertebrate Surviv J 8:241–246
Wang X-W, Zhao X-F, Wang J-X (2014) C-type lectin binds to β-integrin to promote hemocytic phagocytosis in an invertebrate. J Biol Chem 289:2405–2414. https://doi.org/10.1074/jbc.M113.528885
Wu X, Dinguirard N, Sabat G, Lui H, Gonzalez L, Gehring M, Bickham-wright U, Yoshino TP (2017) Proteomic analysis of Biomphalaria glabrata plasma proteins with binding affinity to those expressed by early developing larval Schistosoma mansoni. PLoS Pathog 13:1–30. https://doi.org/10.6019/PXD004942
Yang Ya, Cheng W, Wu X, Huang S, Deng Z, Zeng X, Yuan D, Yang Yu, Wu Z, Chen Y, Zhou Y, Jiang Q (2018) Prediction of the potential global distribution for Biomphalaria straminea, an intermediate host for Schistosoma mansoni. PLoS Negl Trop Dis 12:1–16. https://doi.org/10.1371/journal.pntd.0006548
Bender RC, Broderick EJ, Goodall CP, Christopher J, Bender RC, Broderick EJ, Goodall CP, Bayne CJ (2005) Respiratory burst of biomphalaria glabrata hemocytes : Schistosoma mansoni-resistant snails produce more extracellular H2O2 than susceptible snails published by : Allen Press on behalf of The American Society of Parasitologists Stable URL : https://www.js. J. Parasitol. 91, 275–279
Cox J, Neuhauser N, Michalski A, Scheltema RA, Olsen JV, Mann M (2011) Andromeda : A peptide search engine integrated into the MaxQuant environment. J. Proteome reserach 1794–1805. https://doi.org/10.1021/pr101065j
de Melo ES, Brayner FA, Junior NCP, França IRS, Alves LC (2019) Investigation of defense response and immune priming in Biomphalaria glabrata and Biomphalaria straminea, two species with different susceptibility to Schistosoma mansoni. Parasitol. Res. 189–201. https://doi.org/10.1007/s00436-019-06495-4
Dinguirard N, Cavalcanti MGS, Wu X-J, Bickham-Wright U, Sabat G, Yoshino TP, S Cavalcanti MG, Wu X-J, Bickham-Wright U, Sabat G, Yoshino TP (2018) Proteomic analysis of Biomphalaria glabrata hemocytes during in vitro encapsulation of Schistosoma mansoni Sporocysts. Front Immunol 9, 1–17. https://doi.org/10.3389/fimmu.2018.02773
Galinier R, Portela J, Moné Y, Allienne JF, Henri H, Delbecq S, Mitta G, Gourbal B, Duval D (2013) Biomphalysin, a new β pore-forming toxin involved in Biomphalaria glabrata immune defense against Schistosoma mansoni. PLoS Pathog. 9. https://doi.org/10.1371/journal.ppat.1003216
Hanington PC, Forys MA, Loker ES (2012) A somatically diversified defense factor, FREP3, is a determinant of snail resistance to schistosome infection. PLoS Negl Trop Dis 6. https://doi.org/10.1371/journal.pntd.0001591
Li H, Hambrook JR, Pila EA, Gharamah AA, Fang J, Wu X, Hanington P (2020) Coordination of humoral immune factors dictates compatibility between Schistosoma mansoni and Biomphalaria glabrata. Elife 9. https://doi.org/10.7554/eLife.51708
Mitta G, Gourbal B, Grunau C, Knight M, Bridger JM, Théron A (2017) The compatibility between biomphalaria glabrata snails and Schistosoma mansoni, in: Advances in Parasitology. pp. 111–145. https://doi.org/10.1016/bs.apar.2016.08.006
Negrão-Corrêa D, Mattos ACA, Pereira CAJ, Martins-Souza RL, Coelho PMZ (2012) Interaction of schistosoma mansoni sporocysts and hemocytes of biomphalaria. J Parasitol Res 6. https://doi.org/10.1155/2012/743920
Rodrigues J, Brayner FA, Alves LC (2010) Innate immune memory in. Science (80-. ). 329, 1353–1356
Acknowledgements
The authors thank FIOCRUZ for using the Technological Platforms Network and Immunopathology Keizo Asami Laboratory (LIKA/UFPE).
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD023681.
Funding
This study was supported by Oswaldo Cruz Fundation (FIOCRUZ – PROEP APQ number 1658–2.13/15); Fundação de Amparo a Ciência do Estado de Pernambuco, Brazil (FACEPE, APQ number 0279–2.13/15); and in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior, Brasil (CAPES), as a fellowship.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception. Nairomberg Cavalcanti Portela Junior, Luiz Carlos Alves, and Elverson Soares de Melo – conceived the design of the proposal; Nairomberg Cavalcanti Portela Junior, Elverson Soares de Melo, Iasmin Lopes de Lima, and Rubens Emanoel Tavares da Rocha performed the methodology, lab experiments, and data analysis, Roberto Afonso, and José Luiz de Lima Filho provided complementary resources and analysis. Nairomberg Cavalcanti Portela Junior – writing original draft; Nairomberg Cavalcanti Portela Junior, Luiz Carlos Alves, and Fábio André Brayner and Ana Paula Sampaio Feitosa – writing, reviewing, and editing. Luiz Carlos Alves and Fábio André Brayner – funding acquisition and research supervision.
Corresponding author
Ethics declarations
Ethics approval
FIOCRUZ Ethics and Use of Animals Committee (CEUA) protocol number 104/2016.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Section Editor: Ramaswamy Kalyanasundaram
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Junior, N.C.P., de Melo, E.S., de Lima, I.L. et al. A proteomics evaluation of the primary and secondary immune response of Biomphalaria straminea challenged by Schistosoma mansoni. Parasitol Res 120, 4023–4035 (2021). https://doi.org/10.1007/s00436-021-07341-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-021-07341-2