Abstract
Background
FGFR1 is a receptor tyrosine kinases involved in tumor growth signaling, survival, and differentiation in many solid cancer types. There is growing evidence that FGFR1 amplification might predict therapy response to FGFR1 inhibitors in squamous cell lung cancers. To estimate the potential applicability of anti FGFR1 therapies in squamous cell carcinomas of the head and neck, we studied patterns of FGFR1 amplification using fluorescence in situ hybridization (FISH).
Materials and methods
A tissue microarray was constructed from 453 primary treatment-naive squamous cell carcinomas of the head and neck regions with histopathological and clinical follow-up data [including oral cavity (n = 222), oropharynx (n = 126), and larynx (n = 105)]. FGFR1 and centromere 8 copy numbers were assessed by dual-color FISH. FGFR1 amplification was defined as a copy number ratio FGFR1: centromere 8 ≥ 2.0. HPV sequencing and p16 immunohistochemistry (IHC) were applied to FGFR1-amplified cancers.
Results
FISH analysis was successful in 297 (66%) of the 453 cancers. FGFR1 amplification was found in 6% of analyzable tumors, and was more frequent in tumors of the oral cavity (13/133 amplified, 10%), than cancers of other localizations (1/79 oropharynx, 4/85 larynx; p = 0.007 and 0.159, respectively). One out of 18 FGFR1 amplified cancers was HPV positive. No associations were found between FGFR1 amplification and tumor phenotype or p16 IHC.
Conclusions
Head and neck cancers are recurrently affected by FGFR1 amplification, with a predominance in cancers of the oral cavity. Finding only one HPV positive and FGFR1 amplified cancer argues against a causal relationship between HPV and FGFR1 amplifications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Head and neck squamous cell carcinoma (HNSCC) is worldwide the sixth leading cancer by incidence and more than 650,000 cases are diagnosed annually (Jemal et al. 2009; Kamangar et al. 2006; Lim et al. 2016). Beside alcohol and tobacco, human papilloma virus (HPV) infection is one of the main risk factors in carcinogenesis of HNSCC and seems to have better therapy response in case of positivity (Castellsague et al. 2004; Fakhry et al. 2008; Gillison and Lowy 2004; Herrero et al. 2003; Macfarlane et al. 2010). To date, the anti-EGF receptor (EGFR) monoclonal antibody, cetuximab, is the first and only molecularly targeted therapy to demonstrate a survival benefit for patients with recurrent or metastatic HNSCC (Bonner et al. 2006; Burtness et al. 2005; Markovic and Chung 2012).
A potential new molecular target in squamous cell carcinoma is fibroblast growth factor receptor 1 (FGFR1). FGFR1 is a member of the FGFR family of receptor tyrosine kinases which are involved in biological functions such as cellular proliferation, survival and differentiation (Turner and Grose 2010). Amplification of a genomic region on 8p12 which includes the FGFR1 gene locus is frequently observed in several tumor entities including squamous cell cancer of the lung and oesophagus as well as ovarian, bladder, and prostate cancer (Edwards et al. 2003; Gorringe et al. 2007; Ishizuka et al. 2002; Simon et al. 2001). However, only recently, with the experimental introduction of FGFR1-inhibitors, it was shown that amplification of FGFR1 within the 8p12 genomic region is indeed a driver mutation. FGFR1 amplification induces FGFR1 dependency in lung cancer cell lines via the mitogen-activated protein kinase (MAPK) pathway. Thus, it is suggested that targeting FGFR1 by small molecule inhibitors might become a viable therapeutic option in squamous cell carcinoma of the lung (Dutt et al. 2011; Weiss et al. 2010).
Only limited data are available on amplification of the FGFR1 gene locus in squamous cell carcinoma of head and neck. Freier et al. (2007) reported amplification of 8p12 genomic region by fluorescence in situ hybridization (FISH) in 17.4% (16/92) oral squamous cell carcinomas. Marshall et al. (2011) recently showed that a subset of HNSCC cell lines is dependent on autocrine signaling by fibroblastic growth factors (FGF) and that these cell lines are sensitive to FGFR inhibition. Recently, two other groups reported of amplification of the FGFR1 gene locus in 9.3% (10/107) analyzed squamous cell carcinomas of the tongue and 20% (9/45) analyzed sinonasal squamous cell carcinoma (Schrock et al. 2013; Young et al. 2013).
As FGFR1 amplification might represent an opportunity for targeted therapy in HNSCC, we performed an extensive analysis for FGFR1 gene copy gain in squamous cell carcinomas of the head and neck.
Materials and methods
Specimen collection and TMA construction
A tissue microarray was constructed from a total of 453 primary surgical HNSCC specimens from formalin-fixed, paraffin-embedded archived tissue samples of the Institute of Pathology at the University Medical Center Hamburg-Eppendorf as described (Kononen et al. 1998). The usage of tissue microarrays for research purposes has been approved by the local ethics committee. Only surgical specimens of tumorectomy without the previous therapy were used for tissue microarray construction. All cases included were reviewed by two pathologists (TSC and WW). The pathologic stage was obtained from the primary report of the Institute of Pathology. UICC stage was determined regarding the 7th edition of the UICC TNM classification of malignant tumors (Sobin et al. 2009). Raw survival data were available from 441 patients. The median follow-up period was 24.1 months for oral cavity, 36 months for hypo-/oropharynx, and 54 months for laryngeal carcinomas. Recurrence was defined as tumor relapse after operation with or without adjuvant therapy. Data on therapy, adjuvant therapy setting as well as smoking and drinking were not available for the cohort. An overview of clinical and pathological data of the whole cohort is shown in Table 1. Consecutive, freshly cut sections of the tissue micro arrays were used for FISH, immunohistochemical analysis, and H&E stained reference.
FGFR1 fluorescence in situ hybridization
For analysis of FGFR1 gene copy gain, a dual-color FISH probe set was used. The set consisted of a self-constructed spectrum green-labeled bacterial artificial chromosome clone (RP11-350N15; Source Bioscience, Nottingham, UK; Abbott KIT) and a spectrum red labeled commercial centromere 8 probe (Zytovision, Bremerhaven, Germany) as a reference. Freshly cut sections (4 µm) were deparaffinized and proteolytically pretreated using a commercial kit (paraffin pretreatment reagent kit; Abbott Molecular), followed by dehydration, air drying, and denaturation for 10 min at 72 °C. Hybridization was performed overnight at 37 °C in a humidified chamber. Slides were then washed and counterstained with 0.2 µmol/L of DAPI.
Screening of TMAs was performed by evaluating 20 unequivocal tumor cells. Cases with < 20 analyzable tumor cells were considered not evaluable. Gene amplification was defined as ratio FGFR1/CEP8 ≥ 2 and average FGFR1 signals > 4. All cases showing gene amplification or any ambiguity (blurred signals, high polysomy) were subjected to validation by FGFR1 FISH on sections of the original paraffin block. In addition to the above-mentioned dual-color FISH probe set, the ZytoLight SPEC FGFR1/CEN 8 dual-color FISH probe (Zytovision, Bremerhaven, Germany) was performed on all sections of original paraffin blocks as recommended by the manufacturer.
HPV detection by PCR
Detection of HPV-DNA was performed on formalin-fixed, paraffin-embedded tumor specimens. 4 μm sections were used for DNA extraction with the QIAmp DNA Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. DNA integrity was evaluated through amplification of a beta-globin sequence with primers generating an amplicon of 127 or 111 bp, respectively (forward 5′-GCCATCACTAAAGGCACCGAG-3′ and reverse 5′-TTCCCACCCTTAGGCTGCTG-3′). Detection of HPV was performed using specific primers in the L7 region of HPV16 (forward 5′-ACAAGCAGAACCGGACAGAG -3′ and reverse 5′-GCCCATTAACAGGTCTTCCA-3′; amplicon size = 127 bp) and HPV18 (forward 5′-AAGCTCAGCAGACGACCTTC-3′ and reverse 5′-CCTTCTGGATCAGCCATTGT-3′; amplicon size = 111 bp). The three reactions were performed under identical conditions: 100 ng of DNA were subjected to PCR using the AmpliTaq Gold PCR mastermix (Applied Biosystems, Darmstadt, Germany) as recommended by the manufacturer (43 cycles, annealing temperature 55 °C). In all cases with a positive result for HPV, PCR products were sequenced for confirmation of HPV type.
Immunohistochemistry
Immunohistochemical analyses were performed on 4 µm-thick TMA sections. Staining for p16[ink4a] (BD Pharmingen™; BD bioscience USA, dilution scale 1:25) was carried out after heat pretreatment in Bond Epitope Retrieval Solution 2 (pH 9; Leica Microsystems) in a bond-automated system (Leica Microsystems). Nuclear as well as cytoplasmic p16 expression was evaluated semiquantitatively and classified as previously described (Klaes et al. 2001): negative (< 1% of cells positive), sporadic (isolated cells positive, but < 5%), focal (small cell clusters, but < 25% of the cells positive), and diffuse (> 25% of the cells stained).
Statistical analysis
To study an association between FGFR1 gene copy gain to clinical–pathological parameters, contingency table analysis and Chi-square test (likelihood) were used. Analysis on recurrence free and overall survival was done using the Kaplan–Meier method and has been compared via Logrank test. To compare the follow-up time between tumors with and without successful FISH analysis, analysis of variance (ANOVA) was performed. All p values were two-sided and p values < 0.05 were considered as significant. For statistical analysis the JMP 11.0 software (SAS institute Inc., Cary, NC, USA) was used.
Results
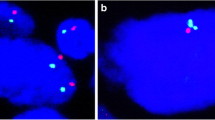
FGFR1 gene copy numbers were interpretable in 297 arrayed tumor samples. Analysis failed in 156 samples, because hybridization quality was too low, not enough tumor cells were analyzable or the entire tissue spot was missing on the tissue microarray slide. 31 cases were reevaluated on large sections of the original paraffin block by two different FGFR1 FISH probe sets. FGFR1 gene amplification defined by an FGFR1/CEP8 ratio ≥ 2 and average FGFR1 signals ≥ 4 was confirmed in 16 cases. The positive cases showed FGFR1 gene amplification with an FGFR1/CEN8 ratio > 2 and an average FGFR1 signal count ranging from 4.35 to 20.50 per tumor cell. The two probe sets used for FGFR1 FISH showed high concordance. The interpretation of the FGFR1 gene copy gain was identical with both sets in all cases. The individual results of the positive cases are shown in Table 2. Analysis on non-malignant squamous tissues (18 probes) did not show any genomic aberration in FGFR1 FISH by both probe sets. FGFR1 amplification was found in 13/133 (9.8%) tumors of the oral cavity, 1/79 (1.3%) tumors of the oropharynx, and 4/85 (4.7%) tumors of the hypopharynx/larynx. Examples of a positive and a negative case are shown in Fig. 1. FGFR1 gene amplification was found to be homogeneous in all tumor cells on the sections of the original tumor block. For this purpose, an H&E stained reference was carefully compared with the FISH slides.
FGFR1 FISH results were analyzed according to tumor localization, pathologic classification, and clinical stage. FGFR1 amplification was found in all pT stages, all pN stages, all UICC stages, and all grades of tumor differentiation. Analysis between the tumor subgroups (oral cavity, oropharynx, and hypopharynx/larynx) showed a predominance of amplified cases in the oral cavity. Detailed information can be found in Table 3. No significant correlation between FGFR1 amplification and any of the clinicopathological parameters was found in the whole HNSCC cohort or in the tumor localization subsets (p > 0.1). Survival analysis of the whole cohort and the tumor subsets of HNSCCs did not show a significant association of FGFR1 gene amplification regarding recurrent free and overall survival (p ≥ 0.38 and p ≥ 0.12, respectively). The suitability of the data for survival analysis was demonstrated by the finding of the known prognostic value of pT (p < 0.0001), pN (p = 0.0011), and UICC stage (p = 0.0001). Detailed results regarding FGFR1 gene amplification and clinico-pathologic data are summarized in Table 3.
The evaluation of p16 expression of the whole cohort did not show a significant association with FGFR1 gene amplification (p = 0.58; Table 4). All cases showing FGFR1 gene amplification were tested for HPV-DNA by polymerase chain reaction and subsequent sequencing. In one of the 18 FGFR1 amplified tumors (oral carcinoma; case #3), HPV-DNA type 16 was detected. This tumor showed diffuse expression of p16. In all other carcinomas with FGFR1 amplification, no HPV-DNA was detected.
Discussion
Our analysis of head and neck squamous cell carcinomas shows FGFR1 amplification in 18/297 tumors with a clear predominance of carcinomas of the oral cavity (13/133 tumors; p = 0.026). In total 10% of the OSCCs, 5% of the larynx and 1% of the oropharynx carcinomas showed an FGFR1 gene amplification. Regarding the available clinicopathologic data (gender, age, pT, pN, cM, Grade, UICC, radiation therapy, tumor recurrence, and overall survival), no significances were found, which is in concordance with the literature (Freier et al. 2007; Kohler et al. 2012; Reis-Filho et al. 2006; Schrock et al. 2013; Weiss et al. 2010; Young et al. 2013).
Especially, our results for the oral cavity are in line with the previous studies, where the authors reported amplification of the 8p12 locus in 16 of 92 oral squamous cell carcinomas or 10/107 squamous cell carcinomas of the tongue by FISH (Freier et al. 2007; Young et al. 2013).
An amplification of the FGFR1 gene locus (8p12) was reported for several other malignancies (squamous cell carcinoma of the lung, breast carcinoma, prostate, sinonasal undifferentiated, and ovary carcinoma) (Edwards et al. 2003; Gorringe et al. 2007; Schrock et al. 2013; Weiss et al. 2010). For squamous cell carcinoma of the lung, Weiss et al. (2010) recently proposed an FGFR1 driver mutation being regulated by transformation in the MAP kinase pathways. In addition, they reported that treatment with FGFR1- inhibitors leads to downstream inhibition and induction of apoptosis in FGFR1 amplified tumor cells using a xenograft mouse model (Weiss et al. 2010).
To date, different FGFR1 inhibitors are being tested to expand the possibilities of tumor treatment. Several studies reported of different anti-FGFR therapeutics showing an effect on different malignancies (myeloproliferative disorders, NSCLC, HNSCC, breast, prostate, and ovarian cancer) (Andre et al. 2013; Bousquet et al. 2011; Cheng et al. 2012; Dutt et al. 2011; Gozgit et al. 2012; Ledermann et al. 2011; Marshall et al. 2011; Sweeny et al. 2012; Weiss et al. 2010).
For HNSCC, Marshall et al. (2011) reported of high FGFR1 RNA levels in a cell line experiment and found out that several FGFR-specific tyrosine kinase inhibitors (TKI) led to a reduction of cell growth, suggesting that FGFR1 might be a serious therapy target for HNSCC. Experiments on HNSCC xenografts showed that treatment with inhibitors affecting FGFR1 (dovitinib, BIBF1120) led to reduced regional lymph node metastasis or inhibition of tumor growth (Hilberg et al. 2008; Sweeny et al. 2012).
Recently, the inhibitor BIBF1120 was tested in a phase I trial on prostate cancer as well as a phase II trials on ovarian cancer and squamous cell cancer of the lung, which showed therapy response (Bousquet et al. 2011; Ledermann et al. 2011).
Regarding our results of FGFR1 gene amplification in HNSCC, especially of oral squamous cell carcinoma (10% of cases), it might be very useful evaluating FGFR inhibitors on HNSCC patients to possibly improve the survival of these patients.
Contrary to squamous cell carcinomas of the lung, human papilloma virus (HPV) has been reported to be involved in tumor development of carcinomas of the head and neck regions (Castellsague et al. 2004; Gillison and Lowy 2004). Therefore, we evaluated the HPV infection and p16 expression status of all FGFR1 amplified cases, to find out if there might be crosslink of infection and gene amplification. In our analysis, only one of the 17 cases (5.6%) showed detectable HPV-DNA type 16 and was diffusely p16 positive. In addition, the comparison of p16 expression according to the FGFR1 gene status (non-/amplified) and available clinicopathologic data showed no significances for the whole cohort and tumor subsets. Due to the low amount of cases in this analysis, a precise statement on influence of HPV on FGFR1 gene amplification cannot be made.
The literature of HPV infection and its effects on FGFR1 is sparse. One study of cervical squamous cell cancer reported of increased FGFR1 gene expression levels in HPV (type 16)-transfected mice (Cheng et al. 2012). Another study analyzed a possible association of HPV infection and amplification of the FGFR1 gene locus using p16 as a surrogate marker for HPV infection and did not find any significance between p16 and FGFR1 gene status (Schrock et al. 2013).
In summary, FGFR1 gene amplification can be found in up to 10% of HNSCCs depending on the tumor site (oral cavity, oropharynx, and hypopharynx/larynx). To find out if HPV infections have any influence on the expression of FGFR1 in HNSCC, further studies have to be carried out.
The most promising fact of this study is, regarding the findings of several groups that anti-FGFR1 therapeutics had influence on tumor growth and even led to tumor reduction and lymph node metastasis, that FGFR1 might become a serious therapy target for HNSCC therapy in the future.
References
Andre F, Bachelot T, Campone M, Dalenc F, Perez-Garcia JM, Hurvitz SA, Turner N, Rugo H, Smith JW, Deudon S, Shi M, Zhang Y, Kay A, Porta DG, Yovine A, Baselga J (2013) Targeting FGFR with dovitinib (TKI258): preclinical and clinical data in breast cancer. Clin Cancer Res 19:3693–3702. doi:10.1158/1078-0432.CCR-13-0190
Bonner JA, Harari PM, Giralt J, Azarnia N, Shin DM, Cohen RB, Jones CU, Sur R, Raben D, Jassem J, Ove R, Kies MS, Baselga J, Youssoufian H, Amellal N, Rowinsky EK, Ang KK (2006) Radiotherapy plus cetuximab for squamous-cell carcinoma of the head and neck. N Engl J Med 354:567–578. doi:10.1056/NEJMoa053422
Bousquet G, Alexandre J, Le Tourneau C, Goldwasser F, Faivre S, de Mont-Serrat H, Kaiser R, Misset JL, Raymond E (2011) Phase I study of BIBF 1120 with docetaxel and prednisone in metastatic chemo-naive hormone-refractory prostate cancer patients. Br J Cancer 105:1640–1645. doi:10.1038/bjc.2011.440
Burtness B, Goldwasser MA, Flood W, Mattar B, Forastiere AA (2005) Phase III randomized trial of cisplatin plus placebo compared with cisplatin plus cetuximab in metastatic/recurrent head and neck cancer: an Eastern Cooperative Oncology Group study. J Clin Oncol 23:8646–8654. doi:10.1200/JCO.2005.02.4646
Castellsague X, Quintana MJ, Martinez MC, Nieto A, Sanchez MJ, Juan A, Monner A, Carrera M, Agudo A, Quer M, Munoz N, Herrero R, Franceschi S, Bosch FX (2004) The role of type of tobacco and type of alcoholic beverage in oral carcinogenesis. Int J Cancer 108:741–749. doi:10.1002/ijc.11627
Cheng YM, Chou CY, Hsu YC, Chen MJ (2012) Influence of HPV16 E6/7 on the expression of FGF2 and FGFR type B in cervical carcinogenesis. Reprod Sci 19:580–586. doi:10.1177/1933719111432874
Dutt A, Ramos AH, Hammerman PS, Mermel C, Cho J, Sharifnia T, Chande A, Tanaka KE, Stransky N, Greulich H, Gray NS, Meyerson M (2011) Inhibitor-sensitive FGFR1 amplification in human non-small cell lung cancer. PLoS One 6:e20351. doi:10.1371/journal.pone.0020351
Edwards J, Krishna NS, Witton CJ, Bartlett JM (2003) Gene amplifications associated with the development of hormone-resistant prostate cancer. Clin Cancer Res 9:5271–5281
Fakhry C, Westra WH, Li S, Cmelak A, Ridge JA, Pinto H, Forastiere A, Gillison ML (2008) Improved survival of patients with human papillomavirus-positive head and neck squamous cell carcinoma in a prospective clinical trial. J Natl Cancer Inst 100:261–269. doi:10.1093/jnci/djn011
Freier K, Schwaenen C, Sticht C, Flechtenmacher C, Muhling J, Hofele C, Radlwimmer B, Lichter P, Joos S (2007) Recurrent FGFR1 amplification and high FGFR1 protein expression in oral squamous cell carcinoma (OSCC). Oral Oncol 43:60–66. doi:10.1016/j.oraloncology.2006.01.005
Gillison ML, Lowy DR (2004) A causal role for human papillomavirus in head and neck cancer. Lancet 363:1488–1489. doi:10.1016/S0140-6736(04)16194-1
Gorringe KL, Jacobs S, Thompson ER, Sridhar A, Qiu W, Choong DY, Campbell IG (2007) High-resolution single nucleotide polymorphism array analysis of epithelial ovarian cancer reveals numerous microdeletions and amplifications. Clin Cancer Res 13:4731–4739. doi:10.1158/1078-0432.CCR-07-0502
Gozgit JM, Wong MJ, Moran L, Wardwell S, Mohemmad QK, Narasimhan NI, Shakespeare WC, Wang F, Clackson T, Rivera VM (2012) Ponatinib (AP24534), a multitargeted pan-FGFR inhibitor with activity in multiple FGFR-amplified or mutated cancer models. Mol Cancer Ther 11:690–699. doi:10.1158/1535-7163.MCT-11-0450
Herrero R, Castellsague X, Pawlita M, Lissowska J, Kee F, Balaram P, Rajkumar T, Sridhar H, Rose B, Pintos J, Fernandez L, Idris A, Sanchez MJ, Nieto A, Talamini R, Tavani A, Bosch FX, Reidel U, Snijders PJ, Meijer CJ, Viscidi R, Munoz N, Franceschi S (2003) Human papillomavirus and oral cancer: the International Agency for Research on Cancer multicenter study. J Natl Cancer Inst 95:1772–1783
Hilberg F, Roth GJ, Krssak M, Kautschitsch S, Sommergruber W, Tontsch-Grunt U, Garin-Chesa P, Bader G, Zoephel A, Quant J, Heckel A, Rettig WJ (2008) BIBF 1120: triple angiokinase inhibitor with sustained receptor blockade and good antitumor efficacy. Cancer Res 68:4774–4782. doi:10.1158/0008-5472.CAN-07-6307
Ishizuka T, Tanabe C, Sakamoto H, Aoyagi K, Maekawa M, Matsukura N, Tokunaga A, Tajiri T, Yoshida T, Terada M, Sasaki H (2002) Gene amplification profiling of esophageal squamous cell carcinomas by DNA array CGH. Biochem Biophys Res Commun 296:152–155
Jemal A, Siegel R, Ward E, Hao Y, Xu J, Thun MJ (2009) Cancer statistics. CA Cancer J Clin 59:225–249. doi:10.3322/caac.20006
Kamangar F, Dores GM, Anderson WF (2006) Patterns of cancer incidence, mortality, and prevalence across five continents: defining priorities to reduce cancer disparities in different geographic regions of the world. J Clin Oncol 24:2137–2150. doi:10.1200/JCO.2005.05.2308
Klaes R, Friedrich T, Spitkovsky D, Ridder R, Rudy W, Petry U, Dallenbach-Hellweg G, Schmidt D, von Knebel Doeberitz M (2001) Overexpression of p16(INK4A) as a specific marker for dysplastic and neoplastic epithelial cells of the cervix uteri. Int J Cancer 92:276–284
Kohler LH, Mireskandari M, Knosel T, Altendorf-Hofmann A, Kunze A, Schmidt A, Presselt N, Chen Y, Petersen I (2012) FGFR1 expression and gene copy numbers in human lung cancer. Virchows Arch 461:49–57. doi:10.1007/s00428-012-1250-y
Kononen J, Bubendorf L, Kallioniemi A, Barlund M, Schraml P, Leighton S, Torhorst J, Mihatsch MJ, Sauter G, Kallioniemi OP (1998) Tissue microarrays for high-throughput molecular profiling of tumor specimens. Nat Med 4:844–847
Ledermann JA, Hackshaw A, Kaye S, Jayson G, Gabra H, McNeish I, Earl H, Perren T, Gore M, Persic M, Adams M, James L, Temple G, Merger M, Rustin G (2011) Randomized phase II placebo-controlled trial of maintenance therapy using the oral triple angiokinase inhibitor BIBF 1120 after chemotherapy for relapsed ovarian cancer. J Clin Oncol 29:3798–3804. doi:10.1200/JCO.2010.33.5208
Lim SH, Sun JM, Choi YL, Kim HR, Ahn S, Lee JY, Lee SH, Ahn JS, Park K, Kim JH, Cho BC, Ahn MJ (2016) Efficacy and safety of dovitinib in pretreated patients with advanced squamous non-small cell lung cancer with FGFR1 amplification: a single-arm, phase 2 study. Cancer 122:3024–3031. doi:10.1002/cncr.30135
Macfarlane TV, Macfarlane GJ, Oliver RJ, Benhamou S, Bouchardy C, Ahrens W, Pohlabeln H, Lagiou P, Lagiou A, Castellsague X, Agudo A, Merletti F, Richiardi L, Kjaerheim K, Slamova A, Schejbalova M, Canova C, Simonato L, Talamini R, Barzan L, Conway DI, McKinney PA, Znaor A, Lowry RJ, Thomson P, Healy CM, McCartan BE, Marron M, Hashibe M, Brennan P (2010) The aetiology of upper aerodigestive tract cancers among young adults in Europe: the ARCAGE study. Cancer Causes Control 21:2213–2221. doi:10.1007/s10552-010-9641-3
Markovic A, Chung CH (2012) Current role of EGF receptor monoclonal antibodies and tyrosine kinase inhibitors in the management of head and neck squamous cell carcinoma. Expert Rev Anticancer Ther 12:1149–1159. doi:10.1586/era.12.91
Marshall ME, Hinz TK, Kono SA, Singleton KR, Bichon B, Ware KE, Marek L, Frederick BA, Raben D, Heasley LE (2011) Fibroblast growth factor receptors are components of autocrine signaling networks in head and neck squamous cell carcinoma cells. Clin Cancer Res 17:5016–5025. doi:10.1158/1078-0432.CCR-11-0050
Reis-Filho JS, Simpson PT, Turner NC, Lambros MB, Jones C, Mackay A, Grigoriadis A, Sarrio D, Savage K, Dexter T, Iravani M, Fenwick K, Weber B, Hardisson D, Schmitt FC, Palacios J, Lakhani SR, Ashworth A (2006) FGFR1 emerges as a potential therapeutic target for lobular breast carcinomas. Clin Cancer Res 12:6652–6662. doi:10.1158/1078-0432.CCR-06-1164
Schrock A, Goke F, Wagner P, Bode M, Franzen A, Huss S, Agaimy A, Ihrler S, Kirsten R, Kristiansen G, Bootz F, Lengerke C, Perner S (2013) Fibroblast-growth-factor-receptor-1 as a potential therapeutic target in sinonasal cancer? Head Neck. doi:10.1002/hed.23443
Simon R, Richter J, Wagner U, Fijan A, Bruderer J, Schmid U, Ackermann D, Maurer R, Alund G, Knonagel H, Rist M, Wilber K, Anabitarte M, Hering F, Hardmeier T, Schonenberger A, Flury R, Jager P, Fehr JL, Schraml P, Moch H, Mihatsch MJ, Gasser T, Sauter G (2001) High-throughput tissue microarray analysis of 3p25 (RAF1) and 8p12 (FGFR1) copy number alterations in urinary bladder cancer. Cancer Res 61:4514–4519
Sobin LH, Gospodarowicz MK, Wittekind C (2009) TNM classification of malignant tumors. Wiley-Blackwell, New York
Sweeny L, Zimmermann TM, Liu Z, Rosenthal EL (2012) Evaluation of tyrosine receptor kinases in the interactions of head and neck squamous cell carcinoma cells and fibroblasts. Oral Oncol. doi:10.1016/j.oraloncology.2012.06.011
Turner N, Grose R (2010) Fibroblast growth factor signalling: from development to cancer. Nat Rev Cancer 10:116–129. doi:10.1038/nrc2780
Weiss J, Sos ML, Seidel D, Peifer M, Zander T, Heuckmann JM, Ullrich RT, Menon R, Maier S, Soltermann A, Moch H, Wagener P, Fischer F, Heynck S, Koker M, Schottle J, Leenders F, Gabler F, Dabow I, Querings S, Heukamp LC, Balke-Want H, Ansen S, Rauh D, Baessmann I, Altmuller J, Wainer Z, Conron M, Wright G, Russell P, Solomon B, Brambilla E, Brambilla C, Lorimier P, Sollberg S, Brustugun OT, Engel-Riedel W, Ludwig C, Petersen I, Sanger J, Clement J, Groen H, Timens W, Sietsma H, Thunnissen E, Smit E, Heideman D, Cappuzzo F, Ligorio C, Damiani S, Hallek M, Beroukhim R, Pao W, Klebl B, Baumann M, Buettner R, Ernestus K, Stoelben E, Wolf J, Nurnberg P, Perner S, Thomas RK (2010) Frequent and focal FGFR1 amplification associates with therapeutically tractable FGFR1 dependency in squamous cell lung cancer. Sci Transl Med 2:62ra93. doi:10.1126/scitranslmed.3001451
Young RJ, Lim AM, Angel C, Collins M, Deb S, Corry J, Wiesenfeld D, Kleid S, Sigston E, Lyons B, Russell PA, Wright G, McArthur GA, Fox SB, Rischin D, Solomon B (2013) Frequency of Fibroblast Growth Factor Receptor 1 gene amplification in oral tongue squamous cell carcinomas and associations with clinical features and patient outcome. Oral Oncol. doi:10.1016/j.oraloncology.2013.01.006
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors of this manuscript declare that they have no conflict of interest.
Human and animal rights statement
The usage of archived diagnostic left-over tissues for manufacturing of tissue microarrays and their analysis for research purposes as well as patient data analysis has been approved by local laws (HmbKHG, §12,1) and by the local ethics committee (Ethics commission Hamburg, WF-049/09 and PV3652). All works have been carried out in compliance with the Helsinki Declaration. This article does not contain any studies with animals performed by any of the authors.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Clauditz, T.S., Böttcher, A., Hanken, H. et al. Prevalence of fibroblast growth factor receptor 1 (FGFR1) amplification in squamous cell carcinomas of the head and neck. J Cancer Res Clin Oncol 144, 53–61 (2018). https://doi.org/10.1007/s00432-017-2528-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-017-2528-x