Abstract
Objectives
Increasing evidence suggests that long non-coding RNAs (lncRNAs) may play a crucial role in many biological processes in a variety of cancers and serve as the basis for many clinical applications including prognostic biomarkers and potential therapeutic targets. The aim of this study is to develop a prognostic lncRNA signature with RNA-seq data in lung adenocarcinomas.
Methods
LncRNA expression profiles and clinical data of lung adenocarcinoma patients from The Cancer Genome Atlas (TCGA) were analyzed. Univariate Cox proportional regression model was used to identify prognostic lncRNAs, and then multivariate Cox proportional regression model was used to develop a prognostic signature. Survivals were compared using log-rank test, and the biological implications of prognostic lncRNAs were analyzed using the KEGG pathway functional enrichment analysis.
Results
We identified eight lncRNAs which had prognostic association with p value <0.01 in a TCGA lung adenocarcinoma cohort of 478 patients. Then a novel prognostic signature with the eight lncRNAs was developed using Cox regression model. Signature high-risk cases had worse overall survival (OS, median 85.97 vs. 38.34 months, p < 0.001) and disease-free survival (DFS, median 44.02 vs. 26.58 months, p = 0.007) than low-risk cases. Multivariate Cox regression analysis suggested that the eight-lncRNA signature was independent of clinical and pathological factors. KEGG pathway functional enrichment analysis indicated potential functional roles of the eight prognostic lncRNAs in tumorigenesis.
Conclusions
Our findings suggest that the eight-lncRNA signature might provide an effective independent prognostic model for the prediction of lung adenocarcinoma patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer is the most common cause of cancer-related mortality worldwide, leading to over one million deaths each year (Torre et al. 2015). Adenocarcinoma is currently the predominant histological subtype of lung cancer, and the rate is still increasing (Travis et al. 2015). Although multidisciplinary cancer therapies have achieved remarkable progress in the past decade, the overall prognosis for lung cancer patients remains poor (Miller et al. 2016). It is important to discover specific prognostic factors for lung adenocarcinomas to foresee clinical outcomes and improve treatment efficacies.
In the age of genomics, gene expression profiling studies have been shown to provide prognostic information beyond conventional parameters in a variety of cancers (Niedzwiecki et al. 2016), and some have further influenced treatment guidelines (Cardoso et al. 2016). However, previous technologies used in these studies were not reflective of the current view of the genome, as most of them only analyzed protein-coding genes (Chen et al. 2007a; Endoh 2004). Genome-wide researches were able to identify tens of thousands of long non-coding RNAs (lncRNAs), which were a novel class of RNAs defined by length >200 nucleotides with no protein-coding potential (Kung et al. 2013; Schmitt and Chang 2016). LncRNAs were reported to play crucial roles in various aspects of cellular processes, including proliferation, survival, migration or genomic stability (Iyer et al. 2015). Based on the large number and expression specificity across different cancer types, lncRNAs are likely to serve as the basis for many clinical applications in oncology (Wahlestedt 2013; Yan et al. 2015). However, the understanding of lncRNAs in lung adenocarcinoma development and clinical outcome in previous studies was relatively limited.
The Cancer Genome Atlas (TCGA) project provides a collection of RNA sequence, DNA copy number variation, DNA methylation and corresponding clinical data for most cancer types. Here, we utilized RNA-seq dataset of TCGA project to identify potential prognostic lncRNAs of lung adenocarcinomas by analyzing the lncRNA expressions of lung adenocarcinomas and adjacent non-tumor tissues. Then, we developed a prognostic lncRNA signature and analyzed the biological pathway of the lncRNA signature.
Materials and methods
Clinical cohorts and lncRNA expression data procession
LncRNA expression datasets and corresponding clinical information of lung adenocarcinoma patients were obtained from publicly available TCGA database. In details, lncRNA expression information of TCGA RNA sequencing database for lung adenocarcinomas was downloaded from the Atlas of non-coding RNAs in Cancer (TANRIC, http://bioinformatics.mdanderson.org/main/TANRIC:Overview, up to Aug 16, 2016) (Li et al. 2015), following a quality filtering by excluding lncRNAs with missing data exceeded 20% of all subject. The full clinical dataset of corresponding lung adenocarcinoma patients was obtained from cBioPortal (http://www.cbioportal.org/, up to Aug 11, 2016) (Cerami et al. 2012; Gao et al. 2013). Overall, 478 patients with primary tumor and 58 of them with normal tissue were included in the current study with lncRNA expression data and corresponding clinical information.
Statistical and data mining analyses of TCGA LUAD lncRNA profiles
Differentially expressed lncRNAs between lung adenocarcinomas and normal tissues were identified with “limma” package, and significance level was set as 0.01 as default to control the false discovery rate (FDR) (Kaczkowski et al. 2016). Heatmap of differentially expressed lncRNAs was plotted with “pheatmap” package, respectively.
The association between differentially expressed lncRNAs and patient overall survival was analyzed by univariate Cox proportional regression model. Difference was considered statistically significant if p value was <0.01. Then the selected lncRNAs were fitted in a multivariate Cox proportional regression model in the dataset. A risk score formula was constructed by these prognostic lncRNAs, weighted by their estimated Cox regression coefficients (Kawaguchi et al. 2012). Patients with assigned risk score were classified into high-risk or low-risk group by using the median as the cutoff point. Overall and disease-free survival of patients in high-risk or low-risk groups were estimated using the Kaplan–Meier method. The log-rank test was used to determine survival differences between groups. Independent prognostic factors were identified through the Cox proportional hazards regression model. All tests were two-tailed. Statistical significance was set as p < 0.05. All data were analyzed using the SPSS Version 19.0 Software (SPSS Inc., Chicago, IL, USA) and R version 3.2.5.
The receiver operating characteristic (ROC) curve was constructed using R package pROC to evaluate the sensitivity and specificity of the survival prediction for the lncRNA signature risk score, age and TNM stage. Area under the curve (AUC) values were calculated from the ROC curves.
Gene set enrichment analysis was performed by the GSEA (http://www.broadinstitute.org/gsea) using MSigDB C2 CP: KEGG gene sets collection (186 gene sets available). Gene sets with a false discovery rate (FDR) value <0.01 after performing 1000 permutations were considered to be significantly enriched (Subramanian et al. 2005).
Results
Identification of prognostic lncRNAs in lung adenocarcinomas from TCGA dataset
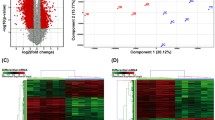
The expression of lncRNAs in the 478 lung adenocarcinomas and 58 adjacent non-tumor tissues were investigated, and a total of 132 lncRNAs were found to be expressed differentially with p value <0.01 (Fig. 1). Univariate Cox proportional regression model was performed on the lncRNA expression data, and eight lncRNAs were identified to be significantly associated with overall survival (p < 0.01, Supplementary Table 1). Among the eight lncRNAs, six (LINC00857, RP11-284F21.7, TMPO-AS1, RP11-284F21.9, LINC01137, and RP11-253E3.3) with larger hazard ratios (hazard ratio larger than 1) were associated with shorter survival; and two (RP11-344B5.2 and CTC-429P9.1) with lower hazard ratios (hazard ratio smaller than 1) were associated with longer survival.
Development of a lncRNA signature associated with survival of lung adenocarcinoma patients
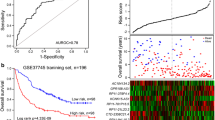
A risk score formula was established to compute a prognostic index for each patient by multivariate Cox proportional regression model according to the expressions of these significant lncRNAs and respective coefficients: Risk Score = (0.095 × LINC00857 + 0.031 × RP11-284F21.7 + 0.066 × TMPO-AS1 + 0.042 × RP11-284F21.9 + 0.061 × LINC01137 + 0.408 × RP11-253E3.3) + (−0.071 × RP11-344B5.2) + (−0.241 × CTC-429P9.1). We calculated the eight-lncRNA expression signature risk score for each patient. Patients were assigned into two groups(high-risk and low-risk groups) by their risk scores, and the median risk score served as cutoff point. Patients in the high-risk group had significantly shorter overall survival (OS, median 85.97 vs. 38.34 months, p < 0.001) and disease-free survival (DFS, median 44.02 vs. 26.58 months, p = 0.007) than patients in the low-risk group (Fig. 2). The distribution of risk score, survival status and lncRNA expression level of each individual were also analyzed (Fig. 3).
Expression of eight lncRNAs, risk score distribution and survival in TCGA cohort. The risk scores (a) for all patients in TCGA cohort are plotted in ascending order and marked as low risk (blue) or high risk (red), as divided by median score (vertical black line). Following up and survival of each patient are shown in b, and alive or dead patient is marked as blue or red, respectively. Expression distribution of eight lncRNAs in TCGA cohort by z score are shown in c, with red indicating higher expression and green indicating lower expression
Associations of the eight-lncRNA signature model with clinical parameters and patient outcome
An analysis was made of the associations between the eight-lncRNA signature model and clinical parameters in lung adenocarcinoma patients. The results showed that the lncRNA signature was not associated with age, smoking history, tumor location, surgical margin, KRAS/EGFR mutation, adjuvant or neoadjuvant therapy (Table 1). High-risk score was significantly associated with male sex p < 0.001) and late AJCC TNM stage (p < 0.001).
We performed Cox univariate and multivariate analysis to ascertain whether the eight-lncRNA expression signature could be an independent predictor for lung adenocarcinoma patients. The univariate analysis results showed that lncRNA risk score and AJCC TNM stage were associated with recurrence and mortality (results not shown). The effect of risk score, age, sex, smoking history, KRAS/EGFR mutation status and TNM stage on lung cancer patient survival time was further analyzed by multivariate Cox proportional hazard model in the cohort. The statistical results indicated that risk score might be an independent predictor of overall survival when adjusted by these factors (Table 2).
We also analyzed the relationship between the eight-lncRNA signature and the AJCC TNM Stage. As shown in Fig. 4, mean risk score increased with the tumor stage in the cohort, suggesting that the eight-lncRNA risk score model might provide a liable aid to predict the clinical outcome of lung adenocarcinoma patients. To test the performance of the signature, we conducted ROC analysis and calculated the AUC on the signature and traditional prognostic factors identified by multivariate Cox proportional hazard model (AJCC TNM stage and age). Our analysis showed that the lncRNA signature might have better prognostic value than traditional prognostic factors in predicting 5-year OS (Fig. 5, AUC, 0.689 vs. 0.661 and 0.532).
KEGG pathway analysis for eight-lncRNA signature target genes
The lncRNAs selected in the present study were correlated with several cancers and diseases by previous studies. Furthermore, we checked whether any KEGG pathways were enriched with target genes of the eight lncRNAs to reveal the biological relevance, and the target genes resulted to participate in both cancer-related and non-cancer-related pathways. The target gene enrichment of KEGG pathways in the present study is listed in Fig. 6. Pathways for ascorbate and aldarate metabolism, cell cycle, DNA replication, porphyrin and chlorophyll metabolism, pentose and glucuronate interconversions, proteasome, homologous recombination and spliceosome were enriched with the eight-lncRNA target genes. Expression level of important genes involved in cell cycle and DNA replication pathways in the cohort are shown in Supplementary Figure 1. Among these genes, Pearson correlation analysis revealed that expression of some genes, for example, PLK1, CDC20, CCNB1, BUB1B, CCNA2, CCNB2, CDC45 and CHEK1, was related with the lncRNA risk score, as listed in Supplementary Table 2. Correlation between expression level of representative genes and lncRNA risk score is shown in Supplementary Figure 2. Thus, KEGG pathway analysis has identified potential targets and biological processes known to be involved in cancer, which provided the biological relevance of the eight-lncRNA signature.
Discussion
The conventional view of cancer has been refreshed in the past decade with the rapid progression of genomic and transcriptional research. TCGA dataset has been demonstrated to be a powerful approach to link clinical features to genetic and transcriptional changes, leading to the identification of new prognostic markers and potentially novel therapeutic targets. LncRNAs are suggested to be promising biomarkers in various cancer types. In the present study, we mined and analyzed lncRNA expression profiles from the TCGA database. By analyzing the association between gene expression profiling and clinical outcome of lung adenocarcinoma patients, we identified a tumor-specific eight-lncRNA signature significantly related to the overall survival of lung adenocarcinoma patients. We further demonstrated the eight-lncRNA signature is independent predictor for lung adenocarcinoma after adjusting to the various variables including age, sex, KRAS/EGFR mutation status and AJCC pathological stage. The use of this eight-lncRNA signature should be investigated in prospective patient cohorts in the future.
Lung adenocarcinoma is characterized by higher incidence rate and worse survival outcomes than many other cancers, with a 5-year overall survival rate at approximately 20% (Miller et al. 2016). Therapy efficacy intensification for high-risk patients is therefore needed. In fact, numerous microarray methods were developed to use the identification of prognostic signatures in lung cancer since 2002 (Beer et al. 2002). Beer and colleagues firstly developed a gene expression profile based on microarray analysis which can be used to predict survival in early-stage lung adenocarcinoma patients (Beer et al. 2002). Traditionally patients were selected for adjuvant chemotherapy based on clinical criteria. Therefore, this result allowed delineation of a high-risk group that may benefit from adjuvant therapy by molecular diagnostic method. Endoh (2004) analyzed the relationship between qRT-PCR results of a total of 45 genes and clinical outcome of cancer patients and found that 8 genes of them were associated with the outcomes of lung adenocarcinoma. Chen et al. (2007b) also estimated a five-gene signature which was closely associated with survival of NSCLC patients. These substantial studies have led to ongoing clinical trials using microarray-derived qRT-PCR prognostic signature to select patients for adjuvant chemotherapy (Kratz et al. 2013). However, these previous studies only included protein-coding genes into analysis and tens of thousands of non-coding RNAs were excluded.
As an important class of non-coding RNAs, increasing evidences have indicated that lncRNAs affect various aspects of cellular homeostasis, and play key roles in cell proliferation, migration, survival, or genomic stability (Huarte 2015). A number of lncRNAs have been identified to correlate with the survival of cancer patients. However, knowledge of lncRNAs associated with lung adenocarcinoma patient survival is still limited. Zhou et al. (2015) performed an array-based transcriptional analysis of lncRNAs from the gene expression omnibus (GEO) database. An expression pattern of eight lncRNAs was found to be significantly associated with overall survival in NSCLC patients; however, the differences of genomic landscape between lung adenocarcinoma and squamous cell carcinoma were not emphasized. Shukla et al. undertook a prognostic analysis of lung adenocarcinoma RNA-seq data and generated a prognostic signature consisted of four genes, including one lncRNA. But more lncRNAs associated with outcomes of lung adenocarcinoma still need to be identified.
As for the characteristics of the eight lncRNAs according to our predictive model, the expression of two lncRNAs (RP11.344B5.2 and CTC.429P9.1) in low-risk group were found to correlate with longer survival, while other six lncRNAs (LINC00857, RP11-284F21.7, TMPO-AS1, RP11-284F21.9, LINC01137, and RP11-253E3.3) were upregulated in the high-risk group compared to low-risk group. Among the six risky lncRNAs, LINC00857 was found to be overexpressed by next generation RNA sequencing analysis of 461 lung adenocarcinomas and 156 normal lung tissues from three separate institutions (Wang et al. 2016). As the top deregulated lncRNAs, overexpression of LINC00857 was significantly associated with poor survival of lung adenocarcinoma. In vitro studies demonstrated that the overexpression of LINC00857 increased cancer cell proliferation, colony formation and invasion. Mechanistic analyses indicated that LINC00857 mediated tumor progression via cell cycle regulation. TMPO-AS1 was also found to be a prognostic biomarker in lung adenocarcinoma in another study (Li et al. 2016). Meanwhile, as a novel subset of RNA, the functions of other six lncRNAs were poorly understood. The biological and clinical studies of these lncRNAs might partially provide clues for the prognostic value, but we still need well-designed studies to validate the functions of these lncRNAs in lung adenocarcinoma. The identification of eight lncRNAs can predict the clinical outcome in patients with lung adenocarcinoma and reveal potential targets for the development of therapeutic options. LncRNAs are suggested to be promising therapeutic targets mostly by preclinical studies and by some clinical studies, and some lncRNAs have been used as therapeutic targets for the treatment in cancer patients (Chandra and Nandan 2016).
Although the eight-lncRNA signature could provide an effective independent prognostic model for the prediction of lung adenocarcinoma, the limitations of the current study should be acknowledged as well. Firstly, the patient cohort of TCGA was from multi-institutional sites, and the study results still need to be validated in other cohorts in future studies. Secondly, since non-coding RNAs were several folds of protein-coding RNAs, only lncRNAs were included in the present study. Thus, this study could not represent the whole transcription alteration associated with lung adenocarcinoma.
In conclusion, we have identified eight lncRNAs associated with survival of lung adenocarcinoma patients in a large cohort and developed a lncRNA prognostic signature. Further analysis revealed that the eight-lncRNA signature could be a prognostic factor independent of conventional clinical parameters. This can serve as a novel biomarker for lung adenocarcinoma prognostic prediction and improve treatment outcome. Further research is necessary to improve the eight-lncRNA signature in large cohorts.
References
Beer DG, Kardia SLR, Huang C, Giordano TJ, Levin AM, Misek DE, Lin L, Chen G, Gharib TG, Thomas DG, Lizyness ML, Kuick R, Hayasaka S, Taylor JMG, Iannettoni MD, Orringer MB, Hanash S (2002) Gene-expression profiles predict survival of patients with lung adenocarcinoma. Nat Med 8(8):816–824. doi:10.1038/nm733
Cardoso F, van’t Veer LJ, Bogaerts J, Slaets L, Viale G, Delaloge S, Pierga J, Brain E, Causeret S, DeLorenzi M, Glas AM, Golfinopoulos V, Goulioti T, Knox S, Matos E, Meulemans B, Neijenhuis PA, Nitz U, Passalacqua R, Ravdin P, Rubio IT, Saghatchian M, Smilde TJ, Sotiriou C, Stork L, Straehle C, Thomas G, Thompson AM, van der Hoeven JM, Vuylsteke P, Bernards R, Tryfonidis K, Rutgers E, Piccart M (2016) 70-Gene signature as an aid to treatment decisions in early-stage breast cancer. N Engl J Med 375(8):717–729. doi:10.1056/NEJMoa1602253
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, Antipin Y, Reva B, Goldberg AP, Sander C, Schultz N (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2(5):401–404. doi:10.1158/2159-8290.CD-12-0095
Chandra GS, Nandan TY (2016) Potential of long non-coding RNAs in cancer patients: from biomarkers to therapeutic targets. Int J Cancer. doi:10.1002/ijc.30546
Chen H, Yu S, Chen C, Chang G, Chen C, Yuan A, Cheng C, Wang C, Terng H, Kao S, Chan W, Li H, Liu C, Singh S, Chen WJ, Chen JJW, Yang P (2007a) A five-gene signature and clinical outcome in non-small-cell lung cancer. N Engl J Med 356(1):11–20. doi:10.1056/NEJMoa060096
Chen H, Yu S, Chen C, Chang G, Chen C, Yuan A, Cheng C, Wang C, Terng H, Kao S, Chan W, Li H, Liu C, Singh S, Chen WJ, Chen JJW, Yang P (2007b) A five-gene signature and clinical outcome in non-small-cell lung cancer. N Engl J Med 356(1):11–20. doi:10.1056/NEJMoa060096
Endoh H (2004) Prognostic model of pulmonary adenocarcinoma by expression profiling of eight genes as determined by quantitative real-time reverse transcriptase polymerase chain reaction. J Clin Oncol 22(5):811–819. doi:10.1200/JCO.2004.04.109
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, Cerami E, Sander C, Schultz N (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6(269):l1. doi:10.1126/scisignal.2004088
Huarte M (2015) The emerging role of lncRNAs in cancer. Nat Med 21(11):1253–1261. doi:10.1038/nm.3981
Iyer MK, Niknafs YS, Malik R, Singhal U, Sahu A, Hosono Y, Barrette TR, Prensner JR, Evans JR, Zhao S, Poliakov A, Cao X, Dhanasekaran SM, Wu Y, Robinson DR, Beer DG, Feng FY, Iyer HK, Chinnaiyan AM (2015) The landscape of long noncoding RNAs in the human transcriptome. Nat Genet 47(3):199–208. doi:10.1038/ng.3192
Kaczkowski B, Tanaka Y, Kawaji H, Sandelin A, Andersson R, Itoh M, Lassmann T, Hayashizaki Y, Carninci P, Forrest ARR (2016) Transcriptome analysis of recurrently deregulated genes across multiple cancers identifies new pan-cancer biomarkers. Can Res 76(2):216–226. doi:10.1158/0008-5472.CAN-15-0484
Kawaguchi A, Iwadate Y, Komohara Y, Sano M, Kajiwara K, Yajima N, Tsuchiya N, Homma J, Aoki H, Kobayashi T, Sakai Y, Hondoh H, Fujii Y, Kakuma T, Yamanaka R (2012) Gene expression signature-based prognostic risk score in patients with primary central nervous system lymphoma. Clin Cancer Res 18(20):5672–5681. doi:10.1158/1078-0432.CCR-12-0596
Kratz JR, Mann MJ, Jablons DM (2013) International trial of adjuvant therapy in high risk stage I non-squamous cell carcinoma identified by a 14-gene prognostic signature. Transl Lung Cancer Res 2(3):222–225. doi:10.3978/j.issn.2218-6751.2013.05.01
Kung JT, Colognori D, Lee JT (2013) Long noncoding RNAs: past, present, and future. Genetics 193(3):651–669. doi:10.1534/genetics.112.146704
Li J, Han L, Roebuck P, Diao L, Liu L, Yuan Y, Weinstein JN, Liang H (2015) TANRIC: an interactive open platform to explore the function of lncRNAs in cancer. Can Res 75(18):3728–3737. doi:10.1158/0008-5472.CAN-15-0273
Li DS, Ainiwaer JL, Sheyhiding I, Zhang Z, Zhang LW (2016) Identification of key long non-coding RNAs as competing endogenous RNAs for miRNA-mRNA in lung adenocarcinoma. Eur Rev Med Pharmacol Sci 20(11):2285–2295
Miller KD, Siegel RL, Lin CC, Mariotto AB, Kramer JL, Rowland JH, Stein KD, Alteri R, Jemal A (2016) Cancer treatment and survivorship statistics, 2016. CA Cancer J Clin 66(4):271–289. doi:10.3322/caac.21349
Niedzwiecki D, Frankel WL, Venook AP, Ye X, Friedman PN, Goldberg RM, Mayer RJ, Colacchio TA, Mulligan JM, Davison TS, O’Brien E, Kerr P, Johnston PG, Kennedy RD, Harkin DP, Schilsky RL, Bertagnolli MM, Warren RS, Innocenti F (2016) Association between results of a gene expression signature assay and recurrence-free interval in patients with stage II colon cancer in cancer and Leukemia Group B 9581 (alliance). J Clin Oncol 34(25):3047–3053. doi:10.1200/JCO.2015.65.4699
Schmitt AM, Chang HY (2016) Long noncoding RNAs in cancer pathways. Cancer Cell 29(4):452–463. doi:10.1016/j.ccell.2016.03.010
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci 102(43):15545–15550. doi:10.1073/pnas.0506580102
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A (2015) Global cancer statistics, 2012. CA Cancer J Clin 65(2):87–108. doi:10.3322/caac.21262
Travis WD, Brambilla E, Nicholson AG, Yatabe Y, Austin JHM, Beasley MB, Chirieac LR, Dacic S, Duhig E, Flieder DB, Geisinger K, Hirsch FR, Ishikawa Y, Kerr KM, Noguchi M, Pelosi G, Powell CA, Tsao MS, Wistuba I (2015) The 2015 world health organization classification of lung tumors. J Thorac Oncol 10(9):1243–1260. doi:10.1097/JTO.0000000000000630
Wahlestedt C (2013) Targeting long non-coding RNA to therapeutically upregulate gene expression. Nat Rev Drug Discovery 12(6):433–446. doi:10.1038/nrd4018
Wang L, He Y, Liu W, Bai S, Xiao L, Zhang J, Dhanasekaran SM, Wang Z, Kalyana-Sundaram S, Balbin OA, Shukla S, Lu Y, Lin J, Reddy RM, Carrott PJ, Lynch WR, Chang AC, Chinnaiyan AM, Beer DG, Zhang J, Chen G (2016) Non-coding RNA LINC00857 is predictive of poor patient survival and promotes tumor progression via cell cycle regulation in lung cancer. Oncotarget 7(10):11487–11499. doi:10.18632/oncotarget.7203
Yan X, Hu Z, Feng Y, Hu X, Yuan J, Zhao SD, Zhang Y, Yang L, Shan W, He Q, Fan L, Kandalaft LE, Tanyi JL, Li C, Yuan C, Zhang D, Yuan H, Hua K, Lu Y, Katsaros D, Huang Q, Montone K, Fan Y, Coukos G, Boyd J, Sood AK, Rebbeck T, Mills GB, Dang CV, Zhang L (2015) Comprehensive genomic characterization of long non-coding RNAs across human cancers. Cancer Cell 28(4):529–540. doi:10.1016/j.ccell.2015.09.006
Zhou M, Guo M, He D, Wang X, Cui Y, Yang H, Hao D, Sun J (2015) A potential signature of eight long non-coding RNAs predicts survival in patients with non-small cell lung cancer. J Transl Med 13(1):231. doi:10.1186/s12967-015-0556-3
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
No potential conflicts of interest were disclosed.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zheng, S., Zheng, D., Dong, C. et al. Development of a novel prognostic signature of long non-coding RNAs in lung adenocarcinoma. J Cancer Res Clin Oncol 143, 1649–1657 (2017). https://doi.org/10.1007/s00432-017-2411-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-017-2411-9