Abstract
We assessed the prevalence of HPV DNA in a large series of Chinese penile cancer and examine its association with the histological subtype, p16INK4a expression, and prognosis. We pathologically categorized 226 invasive penile squamous cell carcinomas and assessed HPV genotyping by real-time PCR and p16INK4a immunohistochemistry. The results were correlated with histopathological and clinical parameters and disease-specific survival (DSS). HPV DNA was detected in 32.7% (74/226) of penile cancer cases. The most frequent genotype was HPV 16 (64/74, 86.5%), followed by HPV 18 (6/74, 8.1%). Fifty-nine (26.1%) cases were positive for the p16INK4a expression, and p16INK4a expression had a sensitivity of 56.8% (95% CI, 45.2–68.3%) and a specificity of 88.8% (95% CI, 83.8–93.9%) for defining HPV status. HPV DNA (P = 0.019), p16INK4a (P = 0.038), age (P = 0.018), grade of differentiation (P = 0.001), lymph nodes (P < 0.001), T stage (P < 0.001), M stage (P < 0.001), and lymphovascular invasion (LVI, P = 0.001) were prognostic factors for DSS. HPV-positivity (HR 0.334; 95% CI, 0.158–0.705, P = 0.004) was still a significant prognostic factor for DSS in the multivariate Cox regression model. HPV DNA was observed in one third of Chinese penile carcinoma cases. The p16INK4a expression can indicate high-risk human papillomavirus (HR-HPV). HPV-positive penile tumors confer a survival benefit over HPV-negative tumors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Penile cancer (PC) is a rare and highly disabling disease with an annual increase of 26,300 new cases worldwide [15]. The overall incidence of PC was 0.6/105 and mortality was 0.18/105 in Chinese male in 2011 reported by the National Central Cancer Registry (NCCR) of China [26]. According to previous research [11, 17], the prevalence of penile cancer has been found to be related to a variety of factors, such as HPV, phimosis, poor sanitation, smoking, multiple sexual partners, genital warts, or other sexually transmitted diseases. Human papillomavirus (HPV) infection has been identified of playing an important role in the development of penile cancer [22]. Two major causative pathways that occur in the carcinogenesis of penile cancer have been described. One has been its association with HPV infection and the other with inflammation, phimosis or sclerosing moss, and lichen planus [5]. In 2016, a new WHO classification for penile cancer was released in which penile squamous cell carcinomas are now classified as HPV- or non-HPV-related [23, 24].
The overall prevalence of HPV DNA in penile carcinoma ranges between 11.6 and 100% but often has a large between-study heterogeneity [24]. Most of the previous researches have indicated that HPV16 was the most common HPV type among HPV DNA-positive penile cancers, followed by HPV6 and HPV18 [1, 24]. Moreover, in penile invasive cancers, the HPV infection rate varied with geographical position, race, and ethnicity [1]. The assessment of HPV status was already a strong recommendation in the EAU guidelines for the pathological evaluation of penile carcinoma specimens [18].
The p16INK4a expression is often used as a surrogate marker for the presence of high-risk human papillomavirus (HR-HPV) in cervical cancer and other organs, such as the head and neck carcinoma [6, 28]. Also, it has been shown that the p16INK4a expression has a strong connection to the presence of HR-HPV in penile cancer [13, 32]. In addition to HPV DNA, p16INK4a has shown prognostication potential in penile cancer. However, the prognostic value of HPV status and P16 status in penile cancer is still inconclusive. Some studies have shown that the positivity of HPV and the P16 protein was a good predictor of prognosis [1, 24], while others had contradicting results [4, 12, 20, 32]. In a recent review, HPV- or p16-positive penile squamous cell carcinomas (SCC) demonstrated significantly better clinical outcomes (HRHPV 0.61; 95% CI, 0.38–0.98; HRp16 0.45; 95% CI, 0.30–0.69) compared with HPV-negative ones [30]. In China, Jianpo Zhai et al. (2013) has shown that the HPV DNA prevalence in Chinese patients was relatively high, at 25.9% (7/28), but its presence did not provide survival advantage [33]. However, due to the limited number of specimens used in such study, more researches are needed to elucidate their correlation in Chinese patients.
Hence, the aim of the current study is to analyze the prevalence of HPV-DNA and expression of p16INK4a in a large series of Chinese PC tissue samples and correlate these results with clinicopathological features and the patients’ survival. In addition, we also aimed to evaluate the association between the histological subtypes of the 2016 WHO penile cancer classification to that of the prevalence of HPV DNA and expression of p16INK4a.
Materials and methods
Patients
The cohort comprised of 226 patients who were treated for invasive PC from 1999 to 2013 at the Sun Yat-sen University Cancer Center (SYSUCC) (Guangzhou, China) and had tissue available for this study. The formalin-fixed paraffin-embedded (FFPE) tissue blocks of the investigated patients were preserved in the pathological archives of SYSUCC and consisted of 186 primary tumors and 40 lymph node metastatic specimens. Their respective clinical and pathological data were retrieved from our electronic medical records and tumor registry.
Pathologic evaluation
The following pathological variables were investigated by two pathologists (Chu CB, Lu JL): tumor histology, grade, pathological T and N stage, presence of lymphovascular and perineural invasion, and necrosis. The tumor histology subtypes were classified as HPV- or non-HPV-related carcinoma in whole tissue sections using the morphological criteria presented in the pathology of penile cancer [23]. The grading of the examined tumors followed the three-tiered International Society of Urological Pathology/World Health Organization system [8]. Pathological parameters were evaluated according to the Eighth Edition of the Tumor-Node-Metastasis (TNM) Staging Classification for Penile Cancer [25].
HPV DNA detection and typing
The detection of HPV on the DNA extracts from FFPE tissues was performed using a Hybribio Assay (HybriMax, Chaozhou Hybribio Limited Corp., Chaozhou, China) that detects the 23 HPV types using real-time polymerase chain reaction (PCR). The kit could identify 13 HR-HPVs (subtype, 16, 18, 31, 33, 35, 39, 45, 51, 52, 56, 58, 59, and 68), 5 low-risk HPVs (LR-HPVs) (subtype, 6, 11, 42, 43, and 44), and other HPV types commonly found in Chinese populations (subtype, 53, 66, 73, 82 and 81/CP8304). DNA was extracted from FFPE tissues using QIAamp® DNA FFPE Tissue (Q) (Cat. no. 56404, Qiagen, Hilden, Germany), and real-time PCR was performed with 2 ul of DNA buffer, 17.5 ul of Master mix (containing probes and primers), and 0.5 ul of Taq DNA polymerase. The PCR conditions were 95 °C for 10 min, 45 cycles of 95 °C for 15 s, and 60 °C for 1 min, with data collection at each cycle during the 60 °C phase on a LightCycler 480 (Roche Diagnostics and Rotor-Gene Q, Qiagen).
Immunohistochemistry for p16INK4aexpression
Immunohistochemistry was performed to determine the expression of p16INK4a according to the manufacturer’s protocol using a mouse monoclonal primary antibody p16INK4a (Clone 6H12, IgG2b/Newcastle) at a dilution of 1:100. The expression of P16INK4a was classified as four patterns: 0, no stain; 1, weak and individual; 2, moderate with small clusters; and 3, strong and diffuse (Fig. 1). Only pattern 3 was represented positive for p16INK4a expression [20].
Follow-up and statistical methods
All patients were followed for every 3 months until the second year after surgery, every 6 months in the 3rd and 4th years, and then on a yearly thereafter. Follow-up data were recorded until March 2019 and comprised of information concerning the patients’ disease status and disease-specific mortality.
Data were tabulated using Microsoft Office Excel 2016 (Microsoft Corporation, Redmond, WA, USA) and analyzed with SPSS V22 (IBM Corp., Armonk, NY, USA). Categorical variables were estimated with the chi-square test or Fisher-exact chi-square test and continuous variables with the Mann-Whitney U test. Survival analysis was estimated by the Kaplan-Meier method and compared by the log-rank test. The Cox regression proportional hazard model was used for multivariate analysis, predicting disease-specific survival (DSS). The statistical significance threshold used was P ≤ 0.05.
Results
Clinicopathological data
The patients’ median age at the time of surgery was 52 (ranging from 24 to 86) years, and the median follow-up time was 57 (2–209) months. There was no difference in the ages between HPV-positive and HPV-negative patients (Mann-Whitney U test, P = 0.833). Of the patients’ specimens, 82.3% (186/226) were primary lesion, and the rest (17.7%, 40/226) were focal lymph nodes. The most common histological subtype was usual SCC (80.4%, 144/179), and 89.4% (160/179) were found to be non-HPV-related histological subtypes. In some cases, the clinicopathological data could not be evaluated since they only received primary cancer focus resection or lymph node resection in our hospital.
HPV genotyping
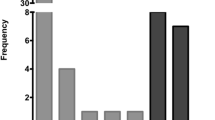
The presence of HPV DNA was detected in 74 (32.7%) of 226 samples, 80.1% (60/74) of which corresponded to only 1 HPV genotype infections. Of these, the most frequent were HPV 16 (64/74, 86.5%), followed by HPV 18 (6/74, 8.1%). Seventy-one cases (95.9%) were infected with HR-HPV, and 3 cases (4.1%) were infected with low-risk HPV (LR-HPV) (HPV6, HPV43, and HPV73 (1 type for each case)) (Fig. 2). The HR-HPV subtypes were observed either alone or together with other subtypes. Overall, 18.9% (14/74) of the investigated penile cancer cases contained HPV DNA from multiple types of HPV.
P16INK4a protein and HPV
Distribution of the p16INK4a expression patterns and the entire HPV status are shown in Fig. 3. Fifty-nine (26.1%) were positive for the p16INK4a expression, and 167 (73.9%) were negative. P16INK4a immunopositivity, considering the observed staining pattern 3, was significantly associated with HR-HPV infection (P < 0.001). p16INK4a had a sensitivity of 57.7% (95% CI, 45.5–69.2%) and a specificity of 88.4% (95% CI, 82.0–92.8%) for defining HR-HPV status. LR-HPV infection was found in 3 cases. Among them, only 1 case, who was usual SCC with HPV-43, had p16INK4a over-expression (pattern 3).p16INK4a was negative in the remaining 2 LR-HPV-positive tumors.
Association of HPV and p16INK4a protein with histopathological and clinical parameters
The clinicopathological features of the patients stratified by tumor HPV DNA status and P16INK4a protein are presented in Table 1. Tumor grade was associated with HPV presence (P = 0.029) and p16INK4a immunopositivity (P = 0.021). HPV-positive or p16INK4a immunopositivity patients tended to have higher tumor grade than HPV-negative or P16INK4a-negative patients. Pure and mixed basaloid carcinomas were observed in 18.75% (3/16 cases) of all grade 3 tumors. When basaloid carcinomas were excluded from the analyses, no significant differences were observed between tumor grade and HPV (P = 0.056) or p16INK4a (P = 0.154). The histological subtypes were correlated with P16INK4a (P = 0.007), where 62.5% of the tumor in HPV-related histologic subtypes were p16INK4a over-expression compared with 26.9% in non-HPV-related histologic subtypes. However, the difference for histologic subtypes in association with HPV positivity did not reach statistical significance (P = 0.082). In addition, age and other histological characteristics were not significantly associated with HPV or p16INK4a.
Table 2 depicts the histological subtypes, HPV status, and P16INK4a expression within these subtypes. The most common subtype was the usual carcinomas (80.4%), followed by warty carcinomas (7.8%), papillary not otherwise specified (5.0%), verrucous (3.4%), basaloid carcinoma (1.7%), and sarcomatous carcinomas (1.1%). Considering the histologic diagnosis in penile cancer, HPV and P16INK4a prevalence varied by the histologic subtypes with the highest prevalence in basaloid and warty carcinomas; 100% of pure or mixed basaloid carcinomas were p16INK4a over-expression. LR-HPV infection was found in 3 cases, of which 2 corresponded to usual SCC and the remaining 1 to warty carcinomas (all was of grade 1). From the morphologic point of view, these LR-HPV tumors were undistinguishable from HR-HPV-positive and HPV-negative tumors with similar histopathologic classification. We were unable to distinguish these LR-HPV carcinomas from HR-HPV-positive and HPV-negative carcinomas by histomorphology.
Survival analysis
The hazard ratios for disease-specific survival (DSS) to HPV DNA positivity, p16INK4a over-expression, and clinicopathological features, using Cox proportional hazard regression model, are shown in Table 3. Age (P = 0.018), grade of differentiation (P = 0.001), lymph nodes (P < 0.001), T stage (P < 0.001), M stage (P < 0.001), and lymphovascular invasion (LVI, P = 0.001) were prognostic for DSS. Regarding DSS for HPV and p16INK4a status, it was observed that HPV-positive and p16INK4a over-expression patients had a higher DSS (Fig. 4) (log-rank P = 0.019 and 0.038, respectively). In a Cox proportional hazard model, HPV-positive (HR 0.334; 95% CI, 0.158–0.705, P = 0.004) was still a significant prognostic factor for DSS, after adjustment for age, grade of differentiation, lymph node status, T stage group, M stage, and LVI (Table 3).
Discussion
This study is the largest, to our knowledge, to assess the HPV prevalence, type distribution, P16INK4a expression, and their association with clinical and pathological parameters of PC in China. The HPV DNA prevalence reported in this study for penile carcinoma (32.7%) was consistent with previous observations in a multicenter study conducted in 25 countries [1]. However, it was lower than that found in systematic reviews by Backes et al. (50%) [3], Tina Bech Olesen et al. (50.8%) [24], and one from the USA in 2014 (63%) [31]. The PC cases of this study had a higher rate of HPV positivity, in sharp contrast to what was found in Asia (13.4%) by Alemany L et al. [1]. In a recent Japanese study, 41% of penile cancer were HPV-positive while only included 34 PC samples [29]. These results indicate a significant geographical difference in HPV infection prevalence.
HPV-related penile tumors had a higher HPV DNA prevalence than non-HPV-related penile tumors. In the present study, we found that cases with warty-basaloid morphologic features were strongly related to HPV DNA positivity and p16INK4a over-expression, which is largely consistent to that previously established in other studies [7, 9, 14, 24]. Furthermore, the HPV DNA prevalence in verrucous carcinoma was similar to the previous analysis [3, 22]. It is worth noting that the usual type of SCC, which is classified as non-HPV-related according to the latest WHO classification, had a high HPV DNA prevalence of 37.5%. In consequence, it was unreasonable to assess HPV status simply by classifying penile tumors as HPV- or non-HPV-related.
Furthermore, significantly fewer well-differentiated tumors were found among the HPV-positive tumors, which was in accordance with some previous studies [13, 16]. Nevertheless, no significant differences between tumor grade and HPV (P = 0.056) or p16INK4a (P = 0.154) were found when we excluded tumors with basaloid and warty features from the analyses.
The findings of this study validated HPV16 as the most common oncogenic HPV genotype, which accounted for more than 70.4% of the single HR-HPV infections in penile cancer. HPV subtypes 6, 11, 16, 18, 31, 33, 45, 52, and 58 together accounted for approximately 90.5% of HPV DNA-positive penile cancers. According to our data, the predominance of HPV16 in penile cancer is underlined, which highlights the potential preventive effect of the available new 9-valent HPV vaccines.
The p16INK4a expression was related to the presence of HR-HPV in penile carcinoma samples, and LR-HPV cases had a lower p16INK4a upregulation than HR-HPV cases. Besides, the p16INK4a protein was related to tumor subtypes other than other histological prognostic parameters. Our data evinces that p16INK4a expression can be used as a significant marker of HR-HPV infection [10].
We examined the prognostic significance of HPV and p16INK4a status on survival in men diagnosed with penile cancer. We found that men with HPV DNA or P16-positive penile cancer had a significantly better DSS compared with those without HPV- or P16-negative penile cancer.
Our results support the hypothesis that HPV-positive and HPV-negative penile cancers differ in relation to survival outcome, which has also been found in other HPV-associated cancers including vulvar and oropharyngeal SCC [2, 27]. However, the explanation for the prognostic significance of HPV status is still unclear. It has been suggested that the presence of viral infection in HPV-associated cancers might strengthen immune surveillance, which consequently makes the HPV-positive cancers less aggressive compared with HPV-negative cancers [2, 19]. In vulvar precancerous lesions, HPV-negative lesions progress to invasive carcinoma more quickly compared with HPV-positive lesions [21], which indicated that HPV-related precancerous lesions generally develop through a slower invasive pathway and have a better prognosis in all stages of the cancerization. Furthermore, for head and neck cancers, it has been revealed that HPV-positive cancers could have a lower degree of gross genetic alterations or that the HPV status of the tumor may determine the molecular structure of the tumor, which could potentially affect the response to therapy [2].
The limitations of this study should be pointed out. First, there are inherent biases and potential errors associated with retrospective and single-center study, and a 5-year follow-up period was not achieved in all cases. Second, we used a mixed set of specimens, containing primary tumors as well as lymph node metastases. However, the histologic subtypes of lymph node metastases could not be assessed. Perhaps, this could in part explain the disappointing fact that the presence of HPV DNA did not correlate well with histologic subtypes of squamous cancer. Third, by using FFPE material, it may be that the proportion of HPV positivity was underestimated.
Conclusion
We observed that the HPV DNA prevalence was 36.4% in penile cancer and P16-positive in 26.1% of those cases. p16INK4a expression was related to HR-HPV DNA and is an important marker of HR-HPV infection. Besides, this study suggests that men with HPV DNA-positive or p16 penile cancer have a significantly better DSS compared with those with HPV/p16-negative.
References
Alemany L, Cubilla A, Halec G, Kasamatsu E, Quirós B, Masferrer E, Tous S, Lloveras B, Hernández-Suarez G, Lonsdale R, Tinoco L, Alejo M, Alvarado-Cabrero I, Laco J, Guimerà N, Poblet E, Lombardi LE, Bergeron C, Clavero O, Shin H-R, Ferrera A, Felix A, Germar J, Mandys V, Clavel C, Tzardi M, Pons LE, Wain V, Cruz E, Molina C, Mota JD, Jach R, Velasco J, Carrilho C, López-Revilla R, Goodman MT, Quint WG, Castellsagué X, Bravo I, Pawlita M, Muñoz N, Bosch FX, de Sanjosé S, group HVs (2016) Role of human papillomavirus in penile carcinomas worldwide. Eur Urol 69:953–961. https://doi.org/10.1016/j.eururo.2015.12.007
Ang KK, Harris J, Wheeler R, Weber R, Rosenthal DI, Nguyen-Tân PF, Westra WH, Chung CH, Jordan RC, Lu C, Kim H, Axelrod R, Silverman CC, Redmond KP, Gillison ML (2010) Human papillomavirus and survival of patients with oropharyngeal cancer. N Engl J Med 363:24–35. https://doi.org/10.1056/NEJMoa0912217
Backes DM, Kurman RJ, Pimenta JM, Smith JS (2009) Systematic review of human papillomavirus prevalence in invasive penile cancer. Cancer Causes Control 20:449–457. https://doi.org/10.1007/s10552-008-9276-9
Bezerra SM, Chaux A, Ball MW, Faraj SF, Munari E, Gonzalez-Roibon N, Sharma R, Bivalacqua TJ, Burnett AL, Netto GJ (2015) Human papillomavirus infection and immunohistochemical p16(INK4a) expression as predictors of outcome in penile squamous cell carcinomas. Hum Pathol 46:532–540. https://doi.org/10.1016/j.humpath.2014.12.004
Bleeker MC, Heideman DA, Snijders PJ, Horenblas S, Dillner J, Meijer CJ (2009) Penile cancer: epidemiology, pathogenesis and prevention. World J Urol 27:141–150. https://doi.org/10.1007/s00345-008-0302-z
Branca M, Ciotti M, Santini D, Di Bonito L, Giorgi C, Benedetto A, Paba P, Favalli C, Costa S, Agarossi A, Alderisio M, Syrjanen K (2004) p16(INK4a) expression is related to grade of CIN and high-risk human papillomavirus but does not predict virus clearance after conization or disease outcome Int. J Gynecol Pathol 23:354–365. https://doi.org/10.1097/01.pgp.0000139639.79105.40
Chaux A, Pfannl R, Lloveras B, Alejo M, Clavero O, Lezcano C, Muñoz N, de Sanjosé S, Bosch X, Hernández-Pérez M, Velazquez EF, Cubilla AL (2010) Distinctive association of p16INK4a overexpression with penile intraepithelial neoplasia depicting warty and/or basaloid features: a study of 141 cases evaluating a new nomenclature. Am J Surg Pathol 34:385–392. https://doi.org/10.1097/PAS.0b013e3181cdad23
Compérat E (2018) Pathology of penile cancer. Eur Urol Suppl 17:132–137. https://doi.org/10.1016/j.eursup.2017.08.005
Cubilla AL, Reuter VE, Gregoire L, Ayala G, Ocampos S, Lancaster WD, Fair W (1998) Basaloid squamous cell carcinoma: a distinctive human papilloma virus-related penile neoplasm: a report of 20 cases. Am J Surg Pathol 22:755–761. https://doi.org/10.1097/00000478-199806000-00014
Cubilla AL, Lloveras B, Alejo M, Clavero O, Chaux A, Kasamatsu E, Monfulleda N, Tous S, Alemany L, Klaustermeier J, Munoz N, Quint W, de Sanjose S, Bosch FX (2011) Value of p16(INK)(4)(a) in the pathology of invasive penile squamous cell carcinomas: a report of 202 cases. Am J Surg Pathol 35:253–261. https://doi.org/10.1097/PAS.0b013e318203cdba
Daling JR, Madeleine MM, Johnson LG, Schwartz SM, Shera KA, Wurscher MA, Carter JJ, Porter PL, Galloway DA, McDougall JK, Krieger JN (2005) Penile cancer: importance of circumcision, human papillomavirus and smoking in in situ and invasive disease. Int J Cancer 116:606–616. https://doi.org/10.1002/ijc.21009
de Araujo LA, De Paula AAP, de Paula H, Ramos JEP, de Oliveira BR, De Carvalho KPA, Guimaraes RA, de Alencar RCG, Duarte ECB, Rabelo Santos SH, Saddi VA, Carneiro M (2018) Human papillomavirus (HPV) genotype distribution in penile carcinoma: association with clinic pathological factors. PloS one 13:e0199557. https://doi.org/10.1371/journal.pone.0199557
Djajadiningrat RS, Jordanova ES, Kroon BK, van Werkhoven E, de Jong J, Pronk DT, Snijders PJ, Horenblas S, Heideman DA (2015) Human papillomavirus prevalence in invasive penile cancer and association with clinical outcome. J Urol 193:526–531. https://doi.org/10.1016/j.juro.2014.08.087
Ferrándiz-Pulido C, Masferrer E, de Torres I, Lloveras B, Hernandez-Losa J, Mojal S, Salvador C, Morote J, Ramon y Cajal S, Pujol RM, Garcia-Patos V, Toll A (2013) Identification and genotyping of human papillomavirus in a Spanish cohort of penile squamous cell carcinomas: correlation with pathologic subtypes, p16(INK4a) expression, and prognosis. J Am Acad Dermatol 68:73–82. https://doi.org/10.1016/j.jaad.2012.05.029
Forman D, de Martel C, Lacey CJ, Soerjomataram I, Lortet-Tieulent J, Bruni L, Vignat J, Ferlay J, Bray F, Plummer M, Franceschi S (2012) Global burden of human papillomavirus and related diseases. Vaccine 30(Suppl 5):F12–F23. https://doi.org/10.1016/j.vaccine.2012.07.055
Gregoire L, Cubilla AL, Reuter VE, Haas GP, Lancaster WD (1995) Preferential association of human papillomavirus with high-grade histologic variants of penile-invasive squamous cell carcinoma. J Natl Cancer Inst 87:1705–1709. https://doi.org/10.1093/jnci/87.22.1705
Hakenberg OW, Comperat EM, Minhas S, Necchi A, Protzel C, Watkin N (2015) EAU guidelines on penile cancer: 2014 update. Eur Urol 67:142–150. https://doi.org/10.1016/j.eururo.2014.10.017
Hakenberg OW, Minhas ES, Necchi A, Protzel C, Watkin N, Compérat E (2018) EAU Guidelines on Penile Cancer 2018European Association of Urology Guidelines. 2018 Edition. European Association of Urology Guidelines Office, Arnhem, The Netherlands, pp
Lont AP, Kroon BK, Horenblas S, Gallee MP, Berkhof J, Meijer CJ, Snijders PJ (2006) Presence of high-risk human papillomavirus DNA in penile carcinoma predicts favorable outcome in survival. Int J Cancer 119:1078–1081. https://doi.org/10.1002/ijc.21961
Martins VA, Pinho JD, Teixeira Junior AAL, Nogueira LR, Silva FF, Maulen VE, Khayat AS, Calixto JRR, Costa HA, Ramalho LNZ, Silva GEB (2018) P16INK4a expression in patients with penile cancer. PLoS One 13:e0205350. https://doi.org/10.1371/journal.pone.0205350
McAlpine JN, Kim SY, Akbari A, Eshragh S, Reuschenbach M, von Knebel DM, Prigge ES, Jordan S, Singh N, Miller DM, Gilks CB (2017) HPV-independent differentiated vulvar intraepithelial neoplasia (dVIN) is associated with an aggressive clinical course. Int J Gynecol Pathol 36:507–516. https://doi.org/10.1097/pgp.0000000000000375
Miralles-Guri C, Bruni L, Cubilla AL, Castellsagué X, Bosch FX, de Sanjosé S (2009) Human papillomavirus prevalence and type distribution in penile carcinoma. J Clin Pathol 62:870–878. https://doi.org/10.1136/jcp.2008.063149
Moch H, Cubilla AL, Humphrey PA, Reuter VE, Ulbright TM (2016) The 2016 WHO classification of tumours of the urinary system and male genital organs-part a: renal, penile, and testicular tumours. Eur Urol 70:93–105. https://doi.org/10.1016/j.eururo.2016.02.029
Olesen TB, Sand FL, Rasmussen CL, Albieri V, Toft BG, Norrild B, Munk C, Kjær SK (2019) Prevalence of human papillomavirus DNA and p16(INK4a) in penile cancer and penile intraepithelial neoplasia: a systematic review and meta-analysis. Lancet Oncol 20:145–158. https://doi.org/10.1016/s1470-2045(18)30682-x
Paner GP, Stadler WM, Hansel DE, Montironi R, Lin DW, Amin MB (2018) Updates in the Eighth Edition of the Tumor-Node-Metastasis Staging Classification for Urologic Cancers. Eur Urol 73:560–569. https://doi.org/10.1016/j.eururo.2017.12.018
Pang C, Guan Y, Li H, Chen W, Zhu G (2016) Urologic cancer in China. Jpn J Clin Oncol 46:497–501. https://doi.org/10.1093/jjco/hyw034
Rasmussen CL, Sand FL, Hoffmann Frederiksen M, Kaae Andersen K, Kjaer SK (2018) Does HPV status influence survival after vulvar cancer? Int J Cancer 142:1158–1165. https://doi.org/10.1002/ijc.31139
Rietbergen MM, Snijders PJ, Beekzada D, Braakhuis BJ, Brink A, Heideman DA, Hesselink AT, Witte BI, Bloemena E, Baatenburg-De Jong RJ, Leemans CR, Brakenhoff RH (2014) Molecular characterization of p16-immunopositive but HPV DNA-negative oropharyngeal carcinomas. Int J Cancer 134:2366–2372. https://doi.org/10.1002/ijc.28580
Sakamoto J, Shigehara K, Nakashima K, Kawaguchi S, Nakashima T, Shimamura M, Yasuda M, Kato T, Hasegawa T, Kobori Y, Okada H, Deguchi T, Izumi K, Kadono Y, Mizokami A (2019) Etiological role of human papillomavirus infection in the development of penile cancer. Int J Infect Dis 78:148–154. https://doi.org/10.1016/j.ijid.2018.11.003
Sand FL, Rasmussen CL, Frederiksen MH, Andersen KK, Kjaer SK (2018) Prognostic significance of HPV and p16 status in men diagnosed with penile cancer: a systematic review and meta-analysis cancer epidemiology, biomarkers & prevention: a publication of the American Association for Cancer Research, cosponsored by the American Society of Preventive. Oncology 27:1123–1132. https://doi.org/10.1158/1055-9965.epi-18-0322
Sinno AK, Saraiya M, Thompson TD, Hernandez BY, Goodman MT, Steinau M, Lynch CF, Cozen W, Saber MS, Peters ES, Wilkinson EJ, Copeland G, Hopenhayn C, Watson M, Lyu C, Unger ER (2014) Human papillomavirus genotype prevalence in invasive vaginal cancer from a registry-based population. Obstet Gynecol 123:817–821. https://doi.org/10.1097/aog.0000000000000171
Steinestel J, Al Ghazal A, Arndt A, Schnoeller TJ, Schrader AJ, Moeller P, Steinestel K (2015) The role of histologic subtype, p16(INK4a) expression, and presence of human papillomavirus DNA in penile squamous cell carcinoma. BMC Cancer 15:220. https://doi.org/10.1186/s12885-015-1268-z
Zhai JP, Wang QY, Wei D, Xu KX, Man LB (2013) Association between HPV DNA and disease specific survival in patients with penile cancer. Zhonghua Yi Xue Za Zhi 93:2719–2722
Funding
This study was supported by the grant from the Nature Science Foundation of China (No. 81872085) and Sun Yat-sen University Cancer Center Medical scientist training program (No.14zxqk08).
Author information
Authors and Affiliations
Contributions
Conceptualization: Chengbiao Chu, Kai Yao, Yun Cao; Methodology: Chengbiao Chu, Keming Chen, Xingliang Tan, Jiangli Lu, Yuanzhong Yang, YiJun Zhang; Formal analysis and investigation: Chengbiao Chu, Keming Chen; Writing, original draft preparation: Chengbiao Chu; Writing, review and editing: Chengbiao Chu, Kai Yao, Yun Cao; Funding acquisition: Kai Yao, Yun Cao; Resources: Chengbiao Chu, Xingliang Tan, Kai Yao, Yun Cao; all authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The study protocol was approved by Independent Ethics Committee of Sun Yat-sen University Cancer Center.
Data availability
The authenticity of this article has been validated by uploading the key raw data onto the Research Data Deposit public platform (www.researchdata.org.cn), with the approval RDD number as RDDB2019000656.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Quality in Pathology
Rights and permissions
About this article
Cite this article
Chu, C., Chen, K., Tan, X. et al. Prevalence of human papillomavirus and implication on survival in Chinese penile cancer. Virchows Arch 477, 667–675 (2020). https://doi.org/10.1007/s00428-020-02831-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-020-02831-7