Abstract
Stem cell division contributes to the generation of various cell types during animal development, especially a diverse pool of neural cells in the nervous system. One example is reiterated unequal stem cell divisions, in which a large stem cell undergoes a series of oriented unequal divisions to produce a chain of small daughter cells that differentiate. We show that reiterated unequal stem cell divisions are involved in the formation of the brain in simple chordate appendicularians (larvaceans). Two large neuroblasts in the anterior and middle of the brain-forming region of hatched larvae were observed. They produced at least 30 neural cells out of 96 total brain cells before completion of brain formation at 10 hours after fertilization by reiterated unequal stem cell divisions. The daughter cells of the anterior neuroblast were postmitotic, and the number was at least 19. The neuroblast produced small daughter neural cells posteriorly every 20 min. The neural cells first moved toward the dorsal side, turned in the anterior direction, aligned in a single line according to their birth order, and showed collective movement to accumulate in the anterior part of the brain. The anterior neuroblast originated from the right-anterior blastomeres of the eight-cell embryos and the right a222 blastomere of the 64-cell embryo. The posterior neuroblast also showed reiterated unequal stem cell divisions, and generated at least 11 neural cells. Sequential unequal stem cell divisions without stem cell growth have been observed in protostomes, such as insects and annelids. The results provide the first examples of this kind of stem cell division during brain formation in non-vertebrate deuterostomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stem cell division contributes to the generation of various cell types during developmental processes and homeostasis in adult organisms. Stem cells possess self-renewal abilities and produce daughter cells committed to differentiation (Doe 2008; Homem and Knoblich 2012; Kang and Reichert 2015; Uzquiano et al. 2018). One well-analyzed stem cell division involves cell divisions that are asymmetric in cell fate and unequal in cell size. A large stem cell undergoes a series of oriented unequal divisions to produce a chain of small daughter cells that differentiate. Such examples have been extensively analyzed in insects and leeches. In Drosophila neurogenesis, larval neuroblasts in the ventral neurogenic region generate a chain of small ganglion mother cells (GMCs) by 3 to 18 sequential unequal stem cell divisions. A GMC typically divides just once into a specific pair of postmitotic neurons or glial cells depending on the position of the neuroblast in the segment and birth order of the GMC (Doe and Goodman 1985; Doe 2008; Kang and Reichert 2015). In leech embryogenesis, blastomeres called M, N, O/P, and Q teloblasts generate a chain of several dozen blast cells by highly unequal stem cell divisions. One blast cell derived from M and O/P teloblasts gives rise to descendants that distribute to a single segment showing a segmentally reiterated pattern. N and Q teloblasts alternatively generate two blast cells with distinct fates, and the pair contributes to one segment width (Weisblat et al. 1988; Weisblat and Kuo 2014).
Stem cell divisions are also involved in the formation of the cerebral cortex of the brain in mammals (Lui et al. 2011; Taverna et al. 2014; Uzquiano et al. 2018). In this case, two daughter cells, self-renewed stem cell and differentiating cell, are equal in size because the stem cells grow during each cell cycle. Apical radial glial cells (aRGs, also called ventricular radial glial cells) in the ventricular zone proliferate through symmetric divisions. They also undergo asymmetric stem cell divisions to produce cells committed to differentiation. In the latter case, aRGs produce both of postmitotic neurons and neural progenitors, such as outer radial glial cells and intermediate progenitors. They further divide and contribute to an increase in the net neural cell number. At a later stage, aRGs produce astrocytes.
The brain (cerebral ganglion) of the larvacean Oikopleura dioica is located in the rostral region of the trunk (Fig. 1A, B). Here, the trunk means a body part except for the tail (Fig. 1A, B). In the present study, we focused on brain formation during larval development. O. dioica is a marine invertebrate belonging to the subphylum Tunicata (also known as Urochordata), which forms a sister clade with vertebrates in the phylum Chordata. It demonstrates several advantages for investigating organ formation processes using live imaging at the single-cell level (Kishi et al. 2014, 2017; Morita et al. 2020). The embryos and larvae are transparent and small (~ 100 μm). They consist of a small number of cells (~ 4000 cells in adults). During embryogenesis, cell lineages are invariant between individuals (Delsman 1910; Stach et al. 2008; Stach and Anselmi 2015). This animal is also characterized by rapid development. Organ formation proceeds within seven hours, from the hatching of larvae at three hours after fertilization, to the completion of organ formation in fully functional juveniles at ten hours. Juveniles start feeding and are miniatures of the adult form. At this stage, most somatic cells are postmitotic, and each cell increases in size to grow into a sexually mature adult on the fifth day (Nishida 2008). Thus, larvaceans are a promising model organism for the developmental studies of chordates.
Stem cell division of a large brain neuroblast. (A) Schematic drawing of a newly hatched larva at three hours post fertilization (hpf). The brain-forming region (pink) locates in the anterior-dorsal region of the trunk. A green cell is the neuroblast. (B) Schematic drawing of a 10-hpf juvenile. Organ formation has completed by the tail shift ventrally. Green cells are the neuroblast and small neural cells generated. (C) The entire trunk of a hatched larva of O. dioica. Dorsal view. The tail was removed to immobilize the larvae. The rectangular region is enlarged in D. Differential interference contrast. (D) Closer view of the brain. Snapshots of the time-lapse movie. The time after hatching is represented as hh: mm. A large neuroblast and its small daughter cell are encircled by broken lines, and corresponding nuclei are indicated by red and green dots, respectively. The small daughter cells are out of focus just after the cell division (from 1:21 to 1:31). They have been focused on at the end (1:35). Otolith cell is visible as a landmark in the brain (Holmberg 1984). A, anterior. P, posterior. Scale bars, 10 μm. See Supplementary Movie S1 for the repetitive sequence of stem cell division
It has been reported that sequential unequal stem cell divisions are involved in the formation of the ventral sensory organ (VSO) and the Nasse region in the trunk epidermal layer during O. dioica larval development. The VSO embedded in the ventral trunk epidermis consists of approximately 30 cells with sensory cilia and axons, and the cells are aligned in a single line perpendicular to the ventral midline (Bollner et al. 1986). The two large bilateral cells located apart from the ventral midline show reiterated unequal cell division, producing a chain of small VSO cells toward the ventral midline (Kishi et al. 2017). The Nasse region consists of very small cells aligned in three rows containing approximately 30 cells each. The Nasse region of the trunk epidermis is important for the secretion of the fibers of the food-trapping filter of the house in which the animal resides (Flood 1991). Some of the ventral edge cells in the Nasse region are generated through stem cell divisions, whereby a large mother cell generates a chain of dorsally positioned small daughter cells (Kishi et al. 2017). These are the first examples of sequential unequal stem cell division in deuterostomes. In this study, we found that sequential unequal stem cell divisions also contribute to the formation of the brain in this animal.
The structure of the brain of O. dioica has been described (Olsson et al. 1990; Bone 1998; Cañestro et al. 2005; Søviknes and Glover 2007; Nishida et al. 2021). Three-dimensional (3D) maps of the brain with the positions of all nuclei were obtained with hatching larvae (three hours after fertilization) and functional juveniles (ten hours) (Nishida et al. 2021). The brain-forming region of the hatching larva consists of 28 cells, and the brain of the functional juvenile comprises 96 cells. In this study, we used live imaging to elucidate brain formation during development of O. dioica.
Materials and methods
Animals
O. dioica has been cultured in our laboratory for hundreds of generations (Bouquet et al. 2009; Omotezako et al. 2013; Martí-Solans et al. 2015). O. dioica is a dioecious organism with a life cycle of five days at 20 °C (Fenaux 1998a; Nishida 2008). Naturally spawned eggs were fertilized with sperm. Larvae were hatched three hours after fertilization. They complete organogenesis by seven hours after hatching (hph), become functional, and initiate feeding.
Mounting of larvae for live imaging
To immobilize swimming larvae for time-lapse movies, the tail was cut off immediately after hatching with a fine knife made of tungsten wire. In other experiments, notochord development in the tail was suppressed by Brachyury DNAi. Double-stranded PCR products (50 ng/μL) of Brachyury (PCR-Bra-248 bp, Omotezako et al. 2015) were injected into the ovarian cytoplasm. Brachyury DNAi did not affect trunk development.
Three Dimensional time-lapse imaging
Live imaging of nuclei and chromosomes was performed using fluorescent proteins at 20 °C. 2 μg/μL Histone 2B (H2B)-EGFP mRNA (Kishi et al. 2014; Morita et al. 2020) was injected into the ovary syncytium of females on day five (approximately half a day before spawning) (Omotezako et al. 2013). After spawning and fertilization, embryos with the highest fluorescence intensity were selected and imaged after hatching. Larval development was recorded using a wide-field microscope equipped with DeltaVision (GE Healthcare, Buckinghamshire, UK). Twenty optical z-sections at a resolution of 896 × 896 pixels were recorded at 2 μm slice intervals through the trunk, in most cases. The time-lapse interval was one or two minutes. To visualize the cell membrane, 5 ng/μL FM-1-43 FX (Invitrogen, Carlsbad, CA, USA) was used in seawater (Betz et al. 1996; Kishi et al. 2014). Larvae were transferred into FM seawater for at least 15 min before imaging.
Cell labeling using photo-convertible nls-Kaede
nls-Kaede mRNA (2 μg/μL) (Ando et al. 2002; Kwan et al. 2012; Kishi et al. 2017) was injected into the ovary syncytium. To label the animal or vegetal hemisphere, eight-cell embryos with vitelline membranes were attached to a poly L-lysine-coated glass-bottomed dish so that the animal-vegetal axis was parallel to the bottom. To convert nls-Kaede fluorescence from green to red, the animal or vegetal cells were exposed to ultraviolet (UV) light for three seconds under an Olympus BX-61 microscope with a LUMPlanApo 40 x/0.80 water lens (Kishi et al. 2017; Onuma et al. 2020; Morita et al. 2020). For labeling the left or right halves, one blastomere of the two-cell embryo was exposed to UV light.
Image processing
Images and movies were processed using the ImageJ software (NIH). Focused slices from hyperstacks were selected to generate movies demonstrating relevant cell behaviors using the Hyper with Stacks function of ImageJ.
Serial block face scanning electron microscopy (SBF-SEM)
SBF-SEM was performed as previously described (Morita et al. 2020; Nishida et al. 2021). Larvae were fixed within ten minutes after hatching. SBF-SEM observations were performed using a Merlin scanning electron microscope (Carl Zeiss Microscopy, Jena, Germany) equipped with a 3View in-chamber ultramicrotome system (Gatan, Pleasanton, CA, USA). The serial slice thickness was 70 nm. The datasets of SBF-SEM serial section images are available in the Dryad repository as “Images of the larva” (36 GB): https://doi.org/10.5061/dryad.dbrv15dzh.
SIMI BioCell
4D time-lapse image datasets of O. dioica embryos observed using DIC optics are available. Image data have been recorded and reported by Stach et al. (2008, 2015) (deposited in the MorphDBase repository, https://www.morphdbase.de/?T_Stach_20151112-M-45.1). In this study, the image data of Scan_20 was used to trace the neuroblast lineage during embryogenesis with SIMI BioCell Viewer software (software for tracing cell lineages, Schnabel et al. 1997, http://www.simi.com/en/products/cell-research/simi-biocell.html). We focused on the neuroblast precursor cells in the 3D space during embryogenesis and developed Supplementary Movie S9.
Results
Primary evidences for stem cell divisions of a large neuroblast in the brain
Tadpole shaped tailbud-larvae hatch at 3 hpf at 20 °C. Just after hatching, there is no obvious structure within the trunk (Fig. 1A). A rudimentary central nervous system with a smooth outline is barely recognizable in the serial sections (Nishida et al. 2021). Organ formation in the trunk occurs during 7 h after the hatching stage. It is completed and fully functional juveniles are formed at 10 h after fertilization (Fig. 1B). The juveniles start feeding and are considered to be miniature adult forms (see Supplementary Movie 5) (Fenaux 1998a, 1998b; Nishida 2008; Nishida et al. 2021).
We first noticed that a relatively large cell in the brain repeatedly divided, showing the disappearance and reappearance of the nucleus during larval development after hatching (three to ten hours after fertilization) in 4D (3D + time) time-lapse movies that were recorded using differential interference contrast optics (two specimens with good orientation) (Fig. 1, see Supplementary Movie 1 for repetitive sequence). The large cell seemed to produce a dozen of smaller daughter cells by sequential unequal cell divisions. Furthermore, the size of the mother cell gradually reduced at each cell division. Measured cell area was reduced to 53% from the beginning to the end of Supplementary Movie 1 in both specimens, namely, the cell volume was reduced to 38%. This is the putative third example of sequential unequal stem cell division in this organism, in addition to those observed in the epidermis layer reported by Kishi et al. (2017).
To confirm that the cell divided unequally, the cell membrane was fluorescently stained with FM dye, and a live recording was performed. It was difficult to obtain good specimens with a good orientation to show both larger and smaller daughter cells in a single focus. One specimen with a good orientation is shown in Fig. 2 and Supplementary Movie 2. The large cell repeated unequal cell divisions, producing smaller daughter cells posteriorly, gradually reducing cell size. Measured cell area was reduced to 75% during 8 rounds of cell division in Supplementary Movie 2, namely, the cell volume was reduced to 66%. This means that the cell volume was reduced by 4.2% at each cell division on average. More compelling evidences of the stem cell divisions are provided in the flowing sections.
Stem cell division of a large brain neuroblast visualized by membrane staining. The cell membrane was stained with FM 1-43. (A) The entire trunk of a hatched larva. The rectangular region is enlarged in B. (B) Snapshots of the time-lapse movie. The time after the start of recording is shown as hh: mm. A large neuroblast and its small daughter cell are encircled by broken lines. A, anterior. P, posterior. Scale bars, 10 μm. See Supplementary Movie S2 for repetitive sequence of the stem cell division
The large mother cell is in the brain; therefore, we call it a neuroblast. We termed smaller daughter cells neural cells, a term which encompasses neurons and glial cells, in this report, because we are unsure whether all of these cells could be characterized as neurons or whether some of them were glial cells. It has been reported that the brain of O. dioica comprises neurons (approximately two-thirds of the total population) and ‘glial’ cells (one-third) (Søviknes and Glover 2007). The glial cells do not form continuous sheaths and are therefore referred to as support cells, rather than glial cells. Søviknes et al. (2005) describe a difference in nuclear shape between neurons and support cells in this animal. The authors stained DNA and observed the specimens with confocal microscope and described that support cells have very intense and dense nuclear staining while neurons have nuclei with a less dense and patchier staining. However, with the methods we used it was difficult to distinguish neurons and glial cells unambiguously.
Descendant neural cells moved anteriorly and settled in the anterior region of the brain
It was difficult to trace the location of daughter cells during larval development in the aforementioned samples; therefore, nuclei were fluorescently labeled with histone 2B-EGFP fusion protein (H2B-EGFP). It was difficult to discern the dorsal-ventral axis of the living larvae immediately after hatching. Therefore, in the H2B-EGFP-expressing specimens, live recordings were started one-to-two hours after hatching. Specimens mounted in good orientation, namely lateral view, in which the neuroblast and its descendant neural cells stayed in the same focal plane, were selected and monitored (two specimens; Fig. 3 and Supplementary Movies 3 and 4). The nucleus of the neuroblast was relatively large and showed weaker fluorescence. The neuroblast entered metaphase showing condensed chromosomes and then divided unequally to generate a neural cell with a small nucleus (Fig. 3B), confirming this mode of stem cell division. The neuroblast always produced neural cells in the posterior direction. Neural cells first moved toward the dorsal side and then turned in the anterior direction. They composed a single line according to their birth order and showed collective movement. Some putative neural daughter cells that had been produced and aligned in a line were already recognized at the beginning of the recording. The final destination was the anterior part of the brain, as shown in Fig. 4 and Supplementary Movie 5. In Supplementary Movie 5, a single neural cell is continuously focused on until the end of brain formation showing how this single neural cell moved first dorsally and then anteriorly, and eventually stayed within the brain. Similarly, other daughter neural cells successively accumulated in the anterior-most region. It took approximately 1 h 50 min for the daughter neural cells to reach the anterior tip of the brain after birth. We did not observe cell division in the neural cells. Therefore, it is likely that each neural cell produced from the neuroblast would differentiate into neuron or glia without further cell division. Other cells in the brain that were already present in hatched larva divided, but rarely, except for another putative neuroblast in the posterior region, which we describe in a later section.
Stem cell division of a neuroblast visualized by nuclear labeling. Nuclei and chromosomes were visualized with Histone 2B-EGFP fusion protein. (A) The entire trunk of a larva. Left view. The tail formation was disrupted by the knockdown of the Brachyury gene to immobilize the larvae. The rectangular region is enlarged in B. (B) The time after the start of recording is shown as hh: mm. The first three images represent interphase when the nucleus of the neuroblast is visible (yellow arrow), metaphase with condensed chromosomes, and anaphase when chromosomes were separated into the neuroblast (yellow arrow) and its small sister cell (red dot). The neuroblast generated the small sister cell posteriorly. The next three images show the movement of daughter cells from 42 min to 1 h 21 min. Five daughter cells are sequentially marked by red, blue, yellow, purple, and orange dots according to their birth order. They were aligned in a single line and moved anteriorly along the dorsal roof of the brain. (C) Another example. (D) Closer views. Seven daughter cells are traced in this case (additionally marked by pink and green dots). See Supplementary Movies 3 and 4 for the repetitive sequence of stem cell division
Daughter neural cells stayed within the brain and settled in the anterior region. Nuclei and chromosomes were visualized with Histone 2B-EGFP. The movie was merged with that acquired after light transmission to show the brain border (green broken line). (A) The entire trunk of a larva. Left view. The rectangular region is enlarged in B. (B) Snapshots of the time-lapse movie. A single neural daughter cell is marked in pink and traced to the final destination using fluorescent images. It stayed within the brain border without further cell division. The time after hatching is shown as hh: mm: ss. See also Supplementary Movie S5
Frequency and number of stem cell divisions
The neuroblast had a large nucleus that showed weaker fluorescence than the surrounding brain cells in the H2B-EGFP-expressing specimens. When larval cells were labeled with Kaede fluorescent protein with a nuclear localization signal (nls-Kaede), the large nucleus of the neuroblast showed high labeling intensity, appearing bright. In addition, nls-Kaede was dispersed into the cytoplasm during the breakdown of the nuclear membrane when the cells approached metaphase. Hence, it was especially useful to monitor the cell cycles of the neuroblast by observing the appearance and disappearance of nucleus in time-lapse movies (two specimens, Fig. 5 and Supplementary Movie 6). The interval between stem cell divisions was monitored, and the number of cell divisions was counted during larval development. Four representative cases presented in Supplementary Movies 1, 5, and 6 are shown in Fig. 6. Each graph is centered using the time when oral gland cells migrated to their final destination on both sides of the pharynx (shown by orange arrowheads). Oral gland cells migrate from the posterior region of the trunk and settle on both sides of the pharynx; this is easily noticeable as a temporal landmark during larval development (Kishi et al. 2014). For instance, the migration was evident in the two specimens, as shown in Supplementary Movie 6.
Cell cycle imaging using nls-Kaede. Kaede fluorescent protein with nuclear-localization signal (nls) was expressed in the entire larvae. It was concentrated into nuclei during interphase, then dispersed into the cytoplasm during nuclear membrane breakdown when cells approached metaphase. (A) The entire trunk of a larva. The rectangular region is enlarged in B. (B) Snapshots of the time-lapse movie. The time after hatching is shown as hh: mm: ss. The large nucleus of the neuroblast is indicated by white arrows, and it disappeared in two minutes (black arrows). This sequence was repeated twice here (19 times in Supplementary Movie S6). (C) Another example. (D) Closer views
Frequency and number of cell divisions of the neuroblast. Vertical blue bars indicate the timing of the stem cell divisions (precisely, nuclear membrane breakdown) of the neuroblast after hatching (0 min) with numbers of cell divisions indicated over the bars. The duration of each cell cycle is indicated between the bars. The timings of recording initiation and oral gland cell migration are indicated by arrowheads. Each graph is centered using the timing of oral gland cell migration. It is plausible that room temperature was different between A/B and C/D. (A) The data was obtained from the right specimen that is shown in Supplementary Movie 6 (nls-Kaede labeling). (B) The left specimen that is shown in Supplementary Movie 6. At the beginning of the recording, the neuroblast already approached metaphase without the nuclear membrane; hence, we placed the first bar just before the start of the recording. (C) The data was obtained from the right specimen of Supplementary Movie 1 (differential interference optics). Probably, they had experienced one more cell division before the start of the recording, considering the recording was started 20 min after hatching. These data suggest that the neuroblast divided at least 19 times after hatching. (D) The data was obtained using a larva shown in Supplementary Movie 5 (Histone 2B-EGFP)
Stem cell divisions could be observed immediately after hatching, although we do not know whether they had already started before hatching. The average cell cycle duration in each specimen was 20 min in Fig. 6A and B, 18 min in C, and 12 min in D. The averages most likely varied depending on room temperature, even though we set air conditioners of laboratory rooms to 20 °C. However, we cannot rule out the possibility that there may be some intrinsic variation among individuals in cell division rate. Cell cycle duration tended to increase as development proceeded. The specimens in Fig. 6A and B show mostly complete recordings. The recording was slightly delayed at the start in Fig. 6C. Stem cell division was likely terminated at 6–6.5 h after hatching. The number of cell divisions after hatching was at least 19, and the cells divided 11 times before oral gland cell migration. However, after the last cell division, we lost the nucleus of the neuroblast in the 3D space as the nucleus became increasingly smaller. We did not observe any signs of cell division in the relevant area.
Origin of neuroblast
The origin of the neuroblast during embryogenesis was also investigated. First, we labeled left or right blastomeres at the two-cell stage using photo-convertible nls-Kaede protein and changing its fluorescent color from green to red under UV illumination (Kishi et al. 2017; Onuma et al. 2020; Morita et al. 2020). In this experiment, converted red fluorescence was inherited by the descendant cells, while green fluorescence was observed in every cell at later stages as the translation of non-converted green Kaede from injected mRNA continued even after UV illumination. Z-stack images of the larvae were acquired five to seven hours after hatching (Fig. 7A and B). The anterior region of the brain showed red fluorescence when the right blastomere was labeled, indicating that the neuroblast was derived from the right blastomere of the two-cell embryo.
Embryonic origin of the neuroblast. (A, B) The left or right hemisphere of the two-cell embryos was exposed to UV light to convert nls-Kaede fluorescence from green to red. The descendants were detected at 4-7 h after hatching. The brain margin is indicated by the blue line. The asterisk indicates the position of the otolith cell in the sensory vesicle. (A) Left blastomeres are labeled by red fluorescence. Nuclei in the anterior brain region (arrows) were not labeled. (B) Right blastomeres are labeled. Nuclei in the anterior brain region were labeled. A, anterior. P, posterior. (C-F) Cells of the animal (C, D) or vegetal hemisphere (E, F) were labeled in red at the eight-cell stage, and their descendants were observed at 30-60 min after hatching. The results show that the neuroblast (arrows) is derived from animal blastomeres. Larvae shown in C-F are the same specimens as those depicted in Fig. 5. See Supplementary Movie 7 for the time-lapse movie
The brain-forming region of pre-hatching embryos originates from a-line animal blastomeres and A-line vegetal blastomeres (Stach et al. 2008). The animal or vegetal hemisphere at the 8-cell stage was then labeled with red nls-Kaede. In O. dioica, the animal and vegetal hemispheres are first separated by the third cleavage plane at the eight-cell stage (Fujii et al. 2008). Time-lapse movies of larval development were recorded (Fig. 7C-F, Supplementary Movie 7). The results clearly showed that the neuroblast had an animal-hemisphere origin.
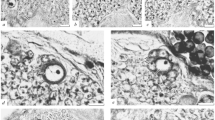
Finally, SBF-SEM data of the hatched larvae (Nishida et al. 2021) and cell lineage data (Stach et al. 2008) were used to trace the embryonic origin of the neuroblast. A cell with a relatively large nucleus and huge amount of cytoplasm has been recognized within the brain-forming region, which demonstrates a clear boundary between the surrounding epidermal and endodermal cells just after hatching (Nishida et al. 2021). The position is shown in Fig. 8A (white line), and the sections of the cell are shown in Fig. 8B and Supplementary Movie 8. This cell can be considered a neuroblast cell. It protruded into the epidermis layer on the dorsal side. The margin of brain-forming region, which is apparent in most parts, becomes ambiguous at the anterior of this cell in hatched larvae (Nishida et al. 2021). This is consistent with the current finding indicating that the neuroblast form the anterior part of the brain after hatching. The maximal cell size in the serial section was 14.7 μm, and the length along the anterior-posterior axis was 21.0 μm. A nucleus was observed in the anterior region of the cell with a size of 5.2 μm. We did not observe any remarkable differences in the cytoplasmic architecture between neuroblast and the surrounding cells.
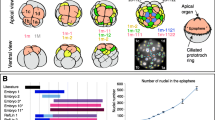
Cell lineage of the neuroblast. (A) Three-dimensional reconstruction of serial SBF-SEM sections of a larva at ten minutes after hatching. Left view. The full trunk and base of the tail are shown. The epidermis layer is removed. The central nervous system (CNS) is shown in pink with light blue nuclei. The shape of the CNS is drawn in the inset. B, brain. TG, tail ganglion. The outline of the neuroblast is indicated by a white line. Note that the size and shape of nuclei do not reflect those in the sample and are represented by spheres with arbitrary sizes to show the position and number of nuclei. See Supplementary Movie S3 in Nishida et al. (2021) for a 3D rotation of this. (B) Three sections of a larva are shown in A at the level of the neuroblast in the anterior to posterior order. Anterior view. The dorsal side is shown in the upward direction. The cytoplasm of the neuroblast cell is shown in pink, and its nucleus is highlighted in blue. Blue line indicates the margin of the nervous system. Red line indicates the basal border of the epidermis layer. N, nucleus. (C) (Left) Position of the neuroblast of an embryo just before hatching in the SIMI BioCell data (Stach et al. 2008, Scan_20). The neuroblast nucleus is marked by an asterisk. The cell had a relatively large nucleus and the cell protruded to the epidermis layer as shown in B. Blue line indicates the margin of the nervous system. (Middle) Virtual embryo showing positions of the nuclei of embryonic cells using the SIMI BioCell data in the same orientation as the left image. Dorsal view. The neuroblast nucleus is marked in red and indicated by a red arrow. Epidermis, nervous system, muscle, notochord, endoderm, endodermal strand, and germ cells are shown in light blue, yellow, pink, dark green, gray, light green, and dark blue, respectively. (Right). The virtual embryo is shown in another orientation. Left view. (D) Lineage tree showing the neuroblast origin (red line). Two red circles indicate the timing of nuclear membrane breakdown; however, the daughter cells were not precisely identified. D, dorsal. V, ventral. A, anterior. P, posterior. See also Supplementary Movies 8 and 9. Figure 8A is modified from Nishida et al. (2021) with permission
The cell was identified in Z-stack optical images of an embryo that was used to trace the cell lineages during embryogenesis (Scan_20, Stach et al. 2008), based on the position and features observed in the SBF-SEM images. Its position in an optical image and in the virtual embryos are shown in Fig. 8C. In addition, time-lapse movies of real and virtual embryogenesis were generated using SIMI BioCell software (Supplementary Movie 9), in which precursor cells of the neuroblast are continuously focused on or marked in red, respectively. The origin of the neuroblast is shown in Fig. 8D. These analyses indicate that the neuroblast was derived from the right-animal blastomere of the eight-cell embryo, which is consistent with the cell labeling results, and from the right a222 blastomere of the 64-cell embryo (the underbar implies blastomere in the right half of embryos). a222 was further divided four times. Interestingly, the cell showed two rounds of the breakdown of the nuclear membrane before hatching (Fig. 8D, two red circles at the bottom). However, we were not able to identify the daughter cells in the Z-stack image after each cell division, probably because the cell division plane was perpendicular to the focal plane, and the daughter cells were small. Hence, stem cell division may have already been initiated before hatching, and the number of cell divisions is two.
The presence of another neuroblast in the middle region of the brain
During the analysis of the aforementioned neuroblast, we noticed another cell that showed similar stem cell divisions. The presence of the cell was confirmed in four specimens. The cell was clearly recognized in the nls-Kaede-expressing larvae when the animal hemisphere was labeled in red (Supplementary Movie 10, yellow arrows). We characterized this cell as the posterior neuroblast and the aforementioned neuroblast as the anterior neuroblast. The number of neuroblasts in the brain likely is just two because we did not observe such neuroblast-like behaviors in other regions of the brain in 4D recordings. The nucleus of the posterior neuroblast was slightly smaller than that of the anterior neuroblast. It divided with a similar duration of the cell cycle as shown by the anterior one. In contrast, the posterior neuroblast ceased cell division at five hours after hatching (1-1.5 h earlier than the anterior neuroblast). The number of cell divisions was counted, and it was 11 times in the best specimen so far recorded (Supplementary Movie 10). We were not able to identify posterior neuroblast in the SBF-SEM images and SIMI BioCell data. It was also difficult to determine where the daughter neural cells were accumulated within the brain because they were not aligned in a single line. Therefore, we mostly focused on anterior neuroblast in this analysis.
Discussion
Stem cell divisions of neuroblasts
The number of cells in the brain of functional juveniles at 10 h is 96 (Nishida et al. 2021). In the adult brain, the number of cells is in the order of 80-90 (Søviknes et al. 2005). Therefore, the brain formation is completed by 10 h after fertilization. In 10 h juvenile, 35 cells are present in the region anterior to the sensory vesicle with an otolith/statocyte in it. Among these, at least 19 were derived from anterior neuroblast. At hatching, more than 10 cells that were not the anterior neuroblast were already present at the similar anteroposterior level to the anterior neuroblast (Supplementary Movie 3 in Nishida et al. 2021). These numbers are consistent with our observation that daughter neural cells generated by anterior neuroblast are postmitotic. This reflects the small number of cells in the brain of larvaceans. The findings are in contrast with the observations noted in insects, leeches, and mammals, where most of the daughter cells of stem cells further divide after birth.
The brain-forming region of the 3 h hatched larva comprises 28 cells (Nishida et al. 2021). The entire brain of juvenile comprises 96 cells, of which at least 30 cells are generated by anterior and posterior neuroblasts (19+11). The number of cells other than stem cells and their progeny can be estimated to be 26 (= 28-2) in the hatched larva and 64 (= 96-30-2) in juvenile. Therefore, they divide 1.3 times on average. This is consistent with the current observation that brain cells other than neuroblasts scarcely divide during larval development when compared to neuroblasts.
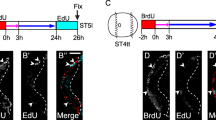
Sequential unequal stem cell divisions of anterior neuroblasts were already observed just after hatching. It was likely that such cell divisions would occur twice before hatching. Looking at the cell lineage tree during embryogenesis (Stach et al. 2008), there is no sign of reiterated stem cell divisions in generation of the central nervous system. Therefore, brain-lineage cells that are already present at hatching are likely to be generated by ordinary cell divisions, except for this case. Unfortunately, it is difficult to continue live imaging during hatching events because larvae move when escaping from the vitelline membranes. Stem cell divisions of the anterior neuroblasts continued at least 19 times for 6.5 h after hatching. Posterior neuroblasts were divided at least 11 times and ceased dividing at 5 h. These results are consistent with those of Søviknes and Glover (2007), who investigated bromodeoxyuridine (BrdU) incorporation during larval development. They were not aware of the stem cell divisions of the neuroblasts because BrdU incorporation was examined in fixed samples at 9 h of development (the developmental speed seemed slower by 1.2–1.4 fold in their experiments compared to that in our experiment). Most brain cells are postmitotic by 6 h, except for cells in the anterodorsal region. The anterodorsal cells that incorporate BrdU are neural cells generated by anterior neuroblast.
The positions of all brain cells have been described in juveniles with completed organ formation (Nishida et al. 2021). However, the types and functions of each brain cell remain elusive, with the exception of several cells. For instance, a pair of sensory neurons in the anterior brain extends dendrites to receive information from secondary sensory cells (coronal cells and dorsal sensory organ cells around the mouth). These two brain cells regulate the reversal of ciliary movements in gill opening to expel inadequate food from the mouth (Olsson et al. 1990; Morita et al. 2020). In addition, approximately seven GABAergic neurons are present in fixed locations in 15 h juvenile, although their function is unknown (Bollner et al. 1991; Søviknes et al. 2005). The neuroblasts of O. dioica might sequentially generate different types of neural cells in a stereotypical order, as shown in Drosophila embryos. Drosophila neuroblasts sequentially express four transcription factors in a fixed order, and the transcription factor that is expressed at birth of differentiating progeny specifies neuronal cell types (Isshiki et al. 2001; Kohwi and Doe 2013; Toma et al. 2016; Gallaud et al. 2017). It is also unclear whether there are substantial functional differences between brain cells that may or may not be derived from neuroblasts in O. dioica.
Evolutionary considerations
Sequential unequal stem cell divisions without stem cell growth have been observed in protostomes, such as insects and annelids. Sequential unequal stem cell divisions observed in O. dioica are the first examples of this type of stem cell division in deuterostomes (Kishi et al. 2017; present study). It is unknown whether this type of stem cell division is retained up to chordates or lost in deuterostome lineages and regained in appendicularians. It is worth exploring the presence of sequential unequal stem cell divisions in other deuterostomes. In ascidians, which constitute tunicates together with appendicularians, the formation and cell lineages of the larval brain have been well described (Cole and Meinertzhagen 2004; Taniguchi and Nishida 2004; Esposito et al. 2017; Hudson 2020; Oonuma et al. 2016; Oonuma and Kusakabe 2021). The larvae of Ciona intestinalis have 177 neurons in the central nervous system (Ryan and Meinertzhagen 2019). To date, no evidence of stem cell division has been obtained. However, stem cell divisions may occur during the formation of the adult brain (cranial ganglion) during metamorphosis.
In insects and vertebrates, stem cell division plays a crucial role in the formation of a large number of neurons and glial cells in the brain (Doe and Goodman 1985; Lui et al. 2011; Taverna et al. 2014; Uzquiano et al. 2018). In this study, we showed that stem cell division also contributes to the formation of the brain with a small cell number. Therefore, the formation of the brain by stem cell divisions is a widely conserved feature of brain-forming processes in various animals, which are independent of the size of the brain. The formation of the vertebrate brain involves stem cell division. However, asymmetric cell divisions of the stem cells are equal in size. This is because stem cells grow during each cell cycle with a nutrient supply from a large reserve of yolk in eggs or the placenta. In contrast, the development of O. dioica and Drosophila is fast, and no room is available for the growth of neuroblasts. Hence, neuroblasts are large and divide unequally in size to generate small progeny. Molecular mechanisms of stem cell division during nervous system development have been extensively investigated in Drosophila and mice (Matsuzaki 2000; Shitamukai and Matsuzaki 2012; Homem and Knoblich 2012). It will be interesting to study the functions of similar molecular players in stem cell divisions of O. dioica to reveal the conservation and deep homology in stem cell divisions and mechanisms of brain formation.
Data availability
The datasets of 4D time-lapse images supporting the conclusions of this article are available in the Dryad repository at https://datadryad.org/stash/dataset/doi:10.5061/dryad.5tb2rbp8c.
References
Ando R, Hama H, Yamamoto-Hino M, Mizuno H, Miyawaki A (2002) An optical marker based on the UV-induced green-to-red photoconversion of a fluorescent protein. Proc Natl Acad Sci U S A 99:12651–12656
Betz WJ, Mao F, Smith CB (1996) Imaging exocytosis and endocytosis. Curr Opin Neurobiol 6:365–371
Bollner T, Holmberg K, Olsson R (1986) A rostral sensory mechanism in Oikopleura dioica (Appendicularia). Acta Zool 67:235–241
Bollner T, Storm-Mathisen J, Ottersen OP (1991) GABA-Like Immunoreactivity in the Nervous System of Oikopleura dioica (Appendicularia). Biol Bull 180:119–124
Bone Q (1998) Nervous system, sense organs, and excitable epithelia. In The Biology of Pelagic Tunicates (ed. Bone, Q), pp. 55–80. Oxford University Press.
Bouquet JM, Spriet E, Troedsson C, Otterå H, Chourrout D, Thompson EM (2009) Culture optimization for the emergent zooplanktonic model organism Oikopleura dioica. J Plankton Res 31:359–370
Cañestro C, Bassham S, Postlethwait J (2005) Development of the central nervous system in the larvacean Oikopleura dioica and the evolution of the chordate brain. Dev Biol 285:298–315. https://doi.org/10.1016/j.ydbio.2005.06.039
Cole AG, Meinertzhagen IA (2004) The central nervous system of the ascidian larva: mitotic history of cells forming the neural tube in late embryonic Ciona intestinalis. Dev Biol 271:239–262
Doe CQ (2008) Neural stem cells: balancing self-renewal with differentiation. Development 135:1575–1587
Delsman HC (1910) Beiträge zur Entwicklungsgeschichte von Oikopleura dioica. Verh Rijksinst Onderz Zee 3:3–24
Doe CQ, Goodman CS (1985) Early events in insect neurogenesis. I. Development and segmental differences in the pattern of neuronal precursor cells. Dev Biol 111:193–205. https://doi.org/10.1016/0012-1606(85)90445-2
Esposito R, Yasuo H, Sirour C, Palladino A, Spagnuolo A, Hudson C (2017) Patterning of brain precursors in ascidian embryos. Development 144:258–264. https://doi.org/10.1242/dev.142307
Fenaux R (1998a) Life history of the Appendicularia. In: Bone Q (ed) The biology of pelagic tunicates. Oxford University Press, New York, pp 151–159
Fenaux R (1998b) Anatomy and functional morphology of the Appendicularia. In: Bone Q (ed) The biology of pelagic tunicates. Oxford University Press, New York, pp 25–34
Flood PR (1991) Architecture of, and water circulation and flow rate in, the house of the planktonic tunicate Oikopleura labradoriensis. Mar Biol 111:95–111
Fujii S, Nishio T, Nishida H (2008) Cleavage pattern, gastrulation, and neurulation in the appendicularian, Oikopleura dioica. Dev Genes Evol 218:69–79
Gallaud E, Pham T, Cabernard C (2017) Drosophila melanogaster neuroblasts: A model for asymmetric stem cell divisions. Results Probl Cell Differ 61:183–210. https://doi.org/10.1007/978-3-319-53150-2_8
Holmberg K (1984) A transmission electron microscopic investigation of the sensory vesicle in the brain of Oikopleura dioica (Appendicularia). Zoomorphology 104:298–303
Homem CC, Knoblich JA (2012) Drosophila neuroblasts: a model for stem cell biology. Development 139:4297–4310. https://doi.org/10.1242/dev.080515
Hudson C (2020) A simple method to identify ascidian brain lineage cells at neural plate stages following in situ hybridization. Methods Mol Biol 2047:325–345. https://doi.org/10.1007/978-1-4939-9732-9_18
Isshiki T, Pearson B, Holbrook S, Doe CQ (2001) Drosophila neuroblasts sequentially express transcription factors which specify the temporal identity of their neuronal progeny. Cell 106:511–521. https://doi.org/10.1016/s0092-8674(01)00465-2
Kang KH, Reichert H (2015) Control of neural stem cell self-renewal and differentiation in Drosophila. Cell Tissue Res 359:33–45. https://doi.org/10.1007/s00441-014-1914-9
Kishi K, Onuma TA, Nishida H (2014) Long-distance cell migration during larval development in the appendicularian, Oikopleura dioica. Dev Biol 395:299–306
Kishi K, Hayashi M, Onuma TA, Nishida H (2017) Patterning and morphogenesis of the intricate but stereotyped oikoplastic epidermis of the appendicularian, Oikopleura dioica. Dev Biol 428:245–257
Kohwi M, Doe CQ (2013) Temporal fate specification and neural progenitor competence during development. Nat Rev Neurosci 14:823–838. https://doi.org/10.1038/nrn3618
Kwan KM, Otsuna H, Kidokoro H, Carney KR, Saijoh Y, Chien CB (2012) A complex choreography of cell movements shapes the vertebrate eye. Development 139:359–372
Lui JH, Hansen DV, Kriegstein AR (2011) Development and evolution of the human neocortex. Cell 146:18–36. https://doi.org/10.1016/j.cell.2011.06.030
Martí-Solans J, Ferrández-Roldán A, Godoy-Marín H, Badia-Ramentol J, Torres-Aguila NP, Rodríguez-Marí A, Bouquet JM, Chourrout D, Thompson EM, Albalat R, Cañestro C (2015) Oikopleura dioica culturing made easy: a low-cost facility for an emerging animal model in EvoDevo. Genesis 53:183–193
Matsuzaki F (2000) Asymmetric division of Drosophila neural stem cells: a basis for neural diversity. Curr Opin Neurobiol 10:38–44. https://doi.org/10.1016/s0959-4388(99)00052-5
Morita R, Onuma TA, Manni L, Ohno N, Nishida H (2020) Mouth opening is mediated by separation of dorsal and ventral daughter cells of the lip precursor cells in the larvacean, Oikopleura dioica. Dev Genes Evol 230:315–327. https://doi.org/10.1007/s00427-020-00667-4
Nishida H (2008) Development of the appendicularian Oikopleura dioica: Culture, genome and cell lineages. Dev Growth Differ 50(Suppl 1):S239–S256
Nishida H, Ohno N, Caicci F, Manni L (2021) 3D reconstruction of structures of hatched larva and young juvenile of the larvacean Oikopleura dioica using SBF-SEM. Sci Rep 11:4833. https://doi.org/10.1038/s41598-021-83706-y
Olsson R, Holmberg K, Lilliemarck Y (1990) Fine structure of the brain and brain nerves of Oikopleura dioica (Urochordata, Appendicularia). Zoomorphology 110:1–7
Omotezako T, Nishino A, Onuma TA, Nishida H (2013) RNA interference in the appendicularian Oikopleura dioica reveals the function of the Brachyury gene. Dev Genes Evol 223:261–267
Omotezako T, Onuma TA, Nishida H (2015) DNA interference: DNA-induced gene silencing in the appendicularian Oikopleura dioica. Proc R Soc B 22:282
Onuma TA, Hayashi M, Gyoja F, Kishi K, Wang K, Nishida H (2020) A chordate species lacking Nodal utilizes calcium oscillation and Bmp for left-right patterning. Proc Natl Acad Sci U S A 117:4188–4198. https://doi.org/10.1073/pnas.1916858117
Oonuma K, Tanaka M, Nishitsuji K, Kato Y, Shimai K, Kusakabe TG (2016) Revised lineage of larval photoreceptor cells in Ciona reveals archetypal collaboration between neural tube and neural crest in sensory organ formation. Dev Biol 420:178–185. https://doi.org/10.1016/j.ydbio.2016.10.014
Oonuma K, Kusakabe TG (2021) The complete cell lineage and MAPK- and Otx-dependent specification of the dopaminergic cells in the Ciona brain. Development 148:dev198754. https://doi.org/10.1242/dev.198754
Ryan K, Meinertzhagen IA (2019) Neuronal identity: the neuron types of a simple chordate sibling, the tadpole larva of Ciona intestinalis. Curr Opin Neurobiol 56:47–60. https://doi.org/10.1016/j.conb.2018.10.015
Schnabel R, Hutter H, Moerman D, Schnabel H (1997) Assessing normal embryogenesis in Caenorhabditis elegans using a 4D microscope: variability of development and regional specification. Dev Biol 184:234–265
Shitamukai A, Matsuzaki F (2012) Control of asymmetric cell division of mammalian neural progenitors. Dev Growth Differ 54:277–286. https://doi.org/10.1111/j.1440-169X.2012.01345.x
Søviknes AM, Chourrout D, Glover JC (2005) Development of putative GABAergic neurons in the appendicularian urochordate Oikopleura dioica. J Comp Neurol 490:12–28. https://doi.org/10.1002/cne.20629
Søviknes AM, Glover JC (2007) Spatiotemporal patterns of neurogenesis in the appendicularian Oikopleura dioica. Dev Biol 311:264–275. https://doi.org/10.1016/j.ydbio.2007.08.058
Stach T, Anselmi C (2015) High-precision morphology: bifocal 4D-microscopy enables the comparison of detailed cell lineages of two chordate species separated for more than 525 million years. BMC Biol 13:113
Stach T, Winter J, Bouquet JM, Chourrout D, Schnabel R (2008) Embryology of a planktonic tunicate reveals traces of sessility. Proc Natl Acad Sci U S A 105:7229–7234
Taniguchi K, Nishida H (2004) Tracing cell fate in brain formation during embryogenesis of the ascidian Halocynthia roretzi. Dev Growth Differ 46:163
Taverna E, Götz M, Huttner WB (2014) The cell biology of neurogenesis: toward an understanding of the development and evolution of the neocortex. Annu Rev Cell Dev Biol 30:465–502. https://doi.org/10.1146/annurev-cellbio-101011-155801
Toma K, Wang TC, Hanashima C (2016) Encoding and decoding time in neural development. Dev Growth Differ 58:59–72. https://doi.org/10.1111/dgd.12257
Uzquiano A, Gladwyn-Ng I, Nguyen L, Reiner O, Götz M, Matsuzaki F, Francis F (2018) Cortical progenitor biology: key features mediating proliferation versus differentiation. J Neurochem 146:500–525. https://doi.org/10.1111/jnc.14338
Weisblat DA, Price DJ, Wedeen CJ (1988) Segmentation in leech development. Development 104:161–168
Weisblat DA, Kuo DH (2014) Developmental biology of the leech Helobdella. Int J Dev Biol 58:429–443. https://doi.org/10.1387/ijdb.140132dw
Acknowledgments
We thank Ms. Chiho Takeuchi, Ms. Keiko Kayahara, and the students in our laboratory for their assistance with the laboratory culture of O. dioica.
Funding
This work was supported by Grants-in-Aid for Scientific Research (KAKENHI) from JSPS and MEXT Japan to H.N. [17KT0023, 19H03234] and T.A.O. [18K06256, 20H05946]. The SBF-SEM observation was supported by the JSPS KAKENHI [JP16H06280] and “Advanced Bioimaging Support (ABiS)”.
Author information
Authors and Affiliations
Contributions
H.N. conceived the study and H.N., A.Y., and T.A.O. directed the study. K.K. and R.M. performed primary experiments. M.K. and H.N. analyzed the data. M.K. and H.N. prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Hiroki Nishida
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Movie S1. Stem cell division of a large brain neuroblast. Related to Fig. 1. The entire trunk of hatched larvae of O. dioica. Dorsal and ventral views of the two larvae. The tail was removed to immobilize the larvae. The time after hatching is represented as hh: mm: ss. The time-lapse recording was carried out at intervals of two minutes. The movie runs at 5 frames/s, which is 600 times faster than in real-time. The nucleus of a large neuroblast is indicated by an arrow when it appears after each cell division (18 times in both specimens). The scale bar is shown in Fig. 1A. (AVI 10296 kb)
Movie S2. Stem cell division of a large brain neuroblast visualized by membrane staining. Related to Fig. 2. The entire trunk of a hatched larva. Dorsal view. The cell membrane was stained with FM 1-43. The time after recording initiation is shown as hh: mm. The time-lapse interval was two minutes. It is 600 times faster than in real-time. Small daughter cells are indicated by white dots when they appear after each cell division. Stem cell division was repeated eight times in this movie. Note that the neuroblast gradually decreased in size. Sensory vesicle expanded at the end of the movie. The scale bar is shown in Fig. 2A. (AVI 711 kb)
Movie S3. The stem cell division of a large brain neuroblast is visualized by nuclear labeling. Related to Fig. 3. Nuclei and chromosomes were visualized using histone 2B-EGFP fusion protein. Two specimens show the cranial region. Left view. The tail formation was disrupted by the knockdown of Brachyury gene to immobilize the larvae. The time after the start of the recording is shown as hh: mm. The time-lapse recording was performed at intervals of one minute. The movie runs at 5 frames/s, which is 300 times faster than in real-time. The neuroblast generated small daughter cells posteriorly. The daughter cells are sequentially marked by red, blue, yellow, purple, orange, pink, and green dots, according to their birth order. They were aligned in a single line and moved anteriorly along the dorsal margin of the brain. Several cells had already been generated at the beginning of the recording, and they were aligned in a single line in the dorsal margin of the brain anteriorly to the red-labeled cell in both specimens. Note that daughter neural cells do not divide and accumulate in the anterior region of the brain. Most cells other than the neuroblast did not divide in the brain region. (AVI 1047 kb)
Movie S4. Stem cell division of a large brain neuroblast visualized by nuclear labeling. Related to Fig. 3. This movie is the same as that in Supplementary Movie S3. However, it is four times faster, that is, 1200 times faster than in real-time so that the anterior movement of daughter neural cells can be easily recognized. (AVI 1047 kb)
Movie S5. Daughter neural cells stayed within the brain and settled in the anterior region. Related to Fig. 4. Nuclei and chromosomes were visualized using histone 2B-EGFP. The movie was merged with that acquired using transmitted light to show the brain border. The time after hatching is represented as hh: mm: ss. The time-lapse interval was two minutes. The movie plays 600 times faster than in real-time. A single neural daughter cell is marked with a pink dot and traced to the final destination. It remained within the brain border without further cell division. (AVI 17922 kb)
Movie S6. Cell cycle imaging using nls-Kaede. Related to Fig. 5. Kaede fluorescent protein with a nuclear localization signal (nls-Kaede) was expressed in entire larvae. It was concentrated in the nuclei during interphase and then dispersed into the cytoplasm at the nuclear membrane break down when the cells approached metaphase. The time after hatching is expressed as hh: mm: ss. The time-lapse interval was two minutes. The movie plays 1200 times faster than in real-time. A large nucleus of the neuroblast is visible, and then it disappears (arrows). This sequence was repeated 19 times for both specimens. Migrating oral gland cells can be observed, and they arrive at the anterior-ventral region at approximately 3 h 30 min. (AVI 10216 kb)
Movie S7. Embryonic origin of the neuroblast. Related to Fig. 7. The animal (left panel) and vegetal (right panel) hemispheres of eight-cell embryos were exposed to UV light to convert nls-Kaede fluorescence from green to red. Both are the same specimens as those depicted in Supplementary Movie S6. The time-lapse interval was two minutes. The movie plays 1200 times faster than in real-time. The time after hatching is indicated as hh: mm: ss. The large nucleus of the neuroblast appears and disappears (arrows), and this sequence is repeated. The neuroblast and its descendant neural cells in the anterior region of the brain were red when the animal hemisphere was labeled. (AVI 17139 kb)
Movie S8. Serial block face scanning electron microscopy (SBF-SEM) images of the neuroblast. Related to Fig. 8B. The movie shows serial sections of a larva 10 min after hatching, which were obtained using SBF-SEM at 210 nm intervals (Nishida et al. 2021). Anterior view. The dorsal side is shown in the upward direction. The movie starts from the anterior side. The cytoplasm of the neuroblast cell is shown in pink, and its nucleus is highlighted in blue. The nucleus is located on the anterior side of the large cytoplasm of the cell. The dorsal side of the cell protrudes towards the epidermis layer. The scale is shown in Fig. 8B. (AVI 19834 kb)
Movie S9. Cell lineage of the neuroblast. Related to Fig. 8C. (Left) Time-lapse movie was generated using SIMI BioCell data (Stach et al. 2008, Scan_20). It starts at the four-cell stage and continues to hatch from the vitelline membrane, corresponding to 2 h 30 min at 20 °C. The red cross indicates the position of the nucleus in the neuroblast lineage cell. Note that the orientation was rotated by 90° in Fig. 8C. (Middle) Virtual embryogenesis showing the positions of embryonic cell nuclei using SIMI BioCell data. This is shown in the same orientation as in the movie on the left. At the beginning (4-cell stage), the animal pole faces upwards, and the posterior side is shown in the front. A dorsal view of the tailbud embryo is shown at the end of the movie. Neuroblast lineage cell is marked in red. The epidermis, nervous system, muscle, notochord, endoderm, endodermal strand, and germ cells are shown in light blue, yellow, pink, dark green, gray, light green, and dark blue, respectively, after the 32-cell stage. (Right) Virtual embryogenesis that is shown in a different orientation. In the beginning, the anterior pole is depicted downward, and the vegetal pole faces in the front. The left view is shown at the end of the movie. (AVI 16783 kb)
Movie S10. Stem cell divisions of the posterior neuroblast. Posterior neuroblast also showed reiteration of the appearance and disappearance of the nucleus in the middle region of the brain. The cell divided eleven times. The time-lapse interval was two minutes. The movie plays 1200 times faster than in real-time. The time after hatching is indicated as hh: mm. The larva is the same specimen as that depicted in Supplementary Movie S7; however, the posterior neuroblast has been focused on continuously. The anterior and posterior neuroblasts are indicated by red and yellow arrows, respectively. This movie also shows that the posterior neuroblast had an animal hemisphere origin. (AVI 15076 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Konishi, M., Kishi, K., Morita, R. et al. Formation of the brain by stem cell divisions of large neuroblasts in Oikopleura dioica, a simple chordate. Dev Genes Evol 233, 35–47 (2023). https://doi.org/10.1007/s00427-023-00704-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-023-00704-y