Abstract
Crinoids are considered as the most basal extant echinoderms. They retain aboral nervous system with a nerve center, which has been degraded in the eleutherozoan echinoderms. To investigate the evolution of patterning of the nervous systems in crinoids, we examined temporal and spatial expression patterns of three neural patterning-related homeobox genes, six3, pax6, and otx, throughout the development of a feather star Anneissia japonica. These genes were involved in the patterning of endomesodermal tissues instead of the ectodermal neural tissues in the early planktonic stages. In the stages after larval attachment, the expression of these genes was mainly observed in the podia and the oral nervous systems instead of the aboral nerve center. Our results indicate the involvement of these three genes in the formation of oral nervous system in the common ancestor of the echinoderms and suggest that the aboral nerve center is not evolutionally related to the brain of other bilaterians.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
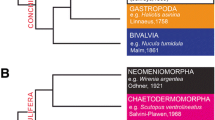
Crinoids are considered as the most basal group of the extant echinoderms due to the fossil records (Paul and Smith 1984; Bottjer et al. 2006) and the molecular analyses (Scouras and Smith 2006; Janies et al. 2011). They are largely divided into two groups by their adult morphology, stalked sea lilies and stalkless feather stars. Feather stars are more dominant than sea lilies in the present seas, which have over 500 species and inhabit in both shallow and deep seas (Clark and Clark 1967). Morphological and molecular analyses indicate that feather stars are derived from ancestral sea lilies (Hyman 1955; Cohen et al. 2004; Rouse et al. 2013); however, many of the “basal” body organizations are conserved, such as larval stalk, unique water vascular system that adapts to the suspension feeding, and well-developed aboral nervous system with a ganglion (Hyman 1955). For these reasons, feather stars can be considered as the important group to reveal evolutionary origin of the unique features of the echinoderm.
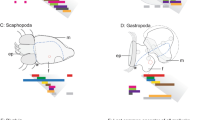
Many bilaterians (protostomes and deuterostomes) have a brain or a nerve center in their heads. In contrast, echinoderms usually do not have a clear head structure nor a brain. The main nervous system of the echinoderms consists of a unit of one nerve ring and five radial nerves (Hyman 1955; Brusca et al. 2016). Two such units exist in eleutherozoan echinoderms (starfish, brittle stars, sea urchins, and sea cucumbers): the predominant oral (ectoneural) nervous system and the inconspicuous deeper oral (hyponeural) nervous system (Supplementary Fig. 1a). The last group of the extant echinoderms, crinoids, also possesses these two nervous systems; however, these two systems do not play principal role. Instead, they have the third system, the aboral (entoneural) nervous system, which serves both motor and sensory functions in crinoids (Supplementary Fig. 1b). In addition, the aboral nervous system of the crinoids possesses a relatively large ganglion in the aboral center of their body, which works as the nerve center at least to integrate the motion of the arms (Marshall 1884).
The molecular mechanism of the anterior-posterior (A-P) regionalization of brain or central nervous system (CNS) is basically conserved in bilaterians (Reichert 2009), although morphology of CNS varies among phyla or classes. In the basal deuterostomes, hemichordates retain this molecular mechanism in their ectoderm (Lowe et al. 2003; Pani et al. 2012); however, no such mechanism was observed in echinoderms. A recent study on a stalked crinoid Metacrinus rotundus indicated the alteration of the expression and function of three brain regionalization–related homeobox genes six3, pax6, and otx; the three genes are involved in the A-P patterning of the larval endomesoderm instead of the ectodermal nervous systems (Omori et al. 2011). The study yet did not analyzed gene expression in the adult structures; thus, the expression and the function of these genes in the adult structures of the crinoids were still obscure.
Anneissia japonica is a feather star crinoid which inhabits the shallow seas of the Japanese temperate coast. Because of the ease of collection, this species has been chosen for the model to confirm the developmental process of feather stars from the fertilized egg through to the adult (Dan and Dan 1941; Kubota 1969, 1970; Shibata et al. 2008). The developmental process of A. japonica after the larval hatch is summarized in Fig. 1. This species only took doliolaria-type planktonic larva (Fig. 1a, b), instead of taking both dipleurula and doliolaria-type larva as observed in those eleutherozoan echinoderms and a stalked crinoid (Garstang 1894; Nakano et al. 2003). Attachment of the planktonic larva with an anterior ventral surface and coincident rotation of the endomesodermal structures results in the change of body axis and relating tissue arrangement (Fig. 1c, d). The rearrangement of the body axis is widely known in the development of echinoderms, which makes it difficult to simply compare the body axis of echinoderms and that of other bilaterians to reveal the evolution of body patterning. A. japonica did not make a rudiment; thus, it is relatively easy to trace each tissue throughout the axis rearrangement.
Normal development of Anneissia japonica after larval hatch. Bright-field pictures (a–f) and schematic illustrations of the inner structures (a’–f’) are shown for the six developmental stages. Structures in same colors in a’–f’ have developmentally same origins. a, a’ Semi-doliolaria larva. Ventral view. Two somatocoels were formed beside enterohydrocoel (e.hc). b, b’ Doliolaria larva. Para-ventral (b) or ventral (b’) view. An adhesive pit (ap), which is identical to the future attachment point (red asterisks), was formed on the anterior ventral surface, and the anterior part of the enterohydrocoel (e.hc) was separated to be a hydrocoel (hc). Relative positions of the endomesodermal tissues began to rotate. c, c’ Attachment stage. An attachment point (red asterisks) is arranged to down. (gray arrows). Rotation of the endomesodermal tissues continued (gray arrows). d, d’ Cystidean stage. Lateral view with the attachment point to the bottom. Rotation of the endomesodermal tissues finished, which resulted to arrange the tissue order from top to bottom as stomodeum (std), hydrocoel (hc), and enteric sac (es). e, e’ Pentacrinoid stage. Lateral view with the mouth opening to the top. Primary podia (pp) were formed from the hydrocoel and the stomodeum of the earlier stages. The aboral nerve center (a.nc) was also formed in the calyx (ca). f, f’ Juvenile. Oral (ventral) view (f) or lateral view with the oral side up. Abbreviations: ag, axial gland; an, anus; a.nc, aboral nerve center; a.rdn, aboral radial nerve; ca, calyx; ci, cirrus; e.hc, enterohydrocoel; es, enteric sac; hc, hydrocoel; l.sc, left somatocoel; m, mouth; pi, pinnule; pp., primary podia; r.sc, right somatocoel; std., stomodeum. Scale bars represent 100 μm (a–d), 1 mm (e), or 1 cm (f)
To reveal the evolution of patterning of the nervous systems in crinoids, including the evolutionally origin of the aboral nerve center of crinoids, we cloned homologs of six3, pax6, and otx from A. japonica and examined temporal and spatial expression patterns of these genes. We improved in situ hybridization methods in the feather star and revealed the expression patterns of three homeobox genes from early planktonic stage to juvenile.
Materials and methods
Collection and larval culture of Anneissia japonica
Collection, keeping of adult specimens in the aquarium and/or in the sea, and culturing of larvae were conducted as described in Shibata et al. (2008).
RNA extraction and cloning of the genes
Total RNA was isolated from different developmental stages of A. japonica using TRIzol reagent (Invitrogen). To remove any polysaccharides, we added final concentration of 1 M of ammonium acetate at the isopropanol precipitation. After the 70% ethanol washing, residual DNA was digested with DNase I (TaKaRa). cDNA was prepared from 500 ng total RNA using random hexamers in a 20-μL reaction as described in the instructions of SuperScript III Reverse Transcriptase (Invitrogen). The cDNA was dissolved in 80 μL TE buffer (10 mM Tris–HCl (pH 7.5), 1 mM EDTA).
Fragments of six3, pax6, and otx were isolated by PCR using sets of degenerated primers. The primer sequences are listed in Supplementary Table 1. After the PCR cycles, the products were subjected to gel-electrophoresis. DNA fragments which were extracted from the bands of the appropriate size were cloned into the EcoRV site of the pBluescript II SK (+) vector. Gene-specific primers for 5′ and 3′ rapid amplification of cDNA ends (RACE) analyses were then designed in each fragment. RACE analyses were performed using the GeneRacer kit (Invitrogen) with total RNAs which were extracted from planktonic or settled larvae of A. japonica as templates. The resulted sequences were confirmed by comparing conserved domains of each gene to those of other animals (see Supplementary Information for detail).
Q-PCR
Primers were designed in the coding region of the target genes to obtain 102–115 base pair products (Supplementary Table 1). Mitochondrial cytochrome oxidase subunit I gene (cox1) was used as a reference of the Q-PCR. The Q-PCR reaction was carried out using LightCycler (Roche) following the instructions of GoTaq qPCR Master Mix (Promega). All Q-PCR experiments were performed twice using two groups of cDNA prepared separately. Relative concentrations of mRNA were normalized to the cox1 values.
WISH
Digoxigenin (DIG)-labeled antisense riboprobes were prepared using plasmids containing 472–892 bp fragments of the genes examined. The primers used for generating the probes are listed in Supplementary Table 1.
Fixation and storing of the samples were performed as described in Omori et al. (2011). The fixed specimens were rehydrated with a graded series of ethanol in PBST (phosphate-buffered saline containing 0.1% Tween 20) and treated with 2~10 μg/ml proteinase K (ProK) in PBST at 37 °C for 20 min. The optimal concentration of ProK was 2 μg/ml for the planktonic larvae, 5 μg/ml for the settled larvae, and 10 μg/ml for the juvenile. The ProK-treated specimens were washed with PBST three times, re-fixed in 4% paraformaldehyde in PBST at 4 °C for 30 min, washed with PBST three times, and then incubated in hybridization buffer (50% formamide, 5× SSC, 100 μg/mL yeast RNA, 50 μg/mL heparin, 1% Tween 20) at 50 °C for 4–6 h. The hybridization reaction was carried out in hybridization buffer containing 0.2 μg/ml (for the larvae) or 0.5 μg/ml (for the juveniles) DIG-labeled riboprobes at 50 °C for 5–10 days. Sample washing and immunodetection were carried out as described in Omori et al. (2011). Stained specimens were immersed in PBST and observed under the light microscopes. After the observation of whole specimens, the stained samples were dehydrated by acetone (5 × 30 min), embedded in Technovit 8100 (Heraeus Kulzer), and cut into 7–10-μm-thick slices. The dissected WISH samples were counterstained with Nuclear Fast Red and observed under light microscopes.
Results
Temporal gene expression profiling of six3, pax6, and otx
Temporal expression profiles revealed by the quantitative real-time PCR (Q-PCR) analysis (Fig. 2) showed that almost no maternal six3 and pax6 mRNAs existed in the unfertilized egg. six3 was highly expressed in the semi-doliolaria and the doliolaria. The expression of six3 declined after the larval attachment, following slight increase of the expression in the late cystidean and gradual decrease through to the juvenile.
Temporal expression profiles of six3, pax6, and otx. Values are represented as the relative RNA copy numbers when the cox1 expression is considered as 100. The developmental stages we investigated are unfertilized egg (EGG), semi-doliolaria (SD), early and late doliolaria (ED, LD), attachment stage (AT), early and late cystidean (EC, LC), early and late pentacrinoid (EP, LP), and juvenile (JUV)
pax6 expression became detectable in the semi-doliolaria, although the expression level of pax6 was under 1/10 of that of six3 and otx. The expression level of pax6 declined to almost 0 in doliolaria and gradually increased after the larval attachment. The peak of the expression was detected in the late pentacrinoid, and the expression slightly declined in the juvenile stage.
otx mRNA substantially existed in the unfertilized egg. The high-level expression of otx was also observed in the semi-doliolaria; however, the expression level was decreased in the early doliolaria. otx expression continued to decline by the early cystidean, which resulted the 1/10 of the expression level compared to that of the semi-doliolaria. The expression level of otx slightly increased in the late cystidean and gradually declined through to the juvenile.
Taken together, six3 and otx had two peaks of gene expression (semi-doliolaria and cystidean-pentacrinoid). pax6 showed low expression level in the planktonic stage and relatively high expression level after attachment.
Spatial expression patterns of six3, pax6, and otx in the planktonic and settled larvae
In the semi-doliolaria, six3 was expressed in the anterior ectoderm in a belt which horizontally surrounds the larva (Fig. 3a, double-arrowheads), and in the anterior region of the enterohydrocoel (Fig. 3a, arrow). The ectodermal expression in the banded region decreased slightly in the late doliolaria (Fig. 3b, double-arrowheads), while weak expression was also observed in the entire region of the anterior ectoderm. The ectodermal expression completely disappeared after the larval attachment. The six3 expression was also observed in the hydrocoel in the late doliolaria (Fig. 3b, arrow), which derives from the anterior region of the enterohydrocoel (Dan et al. 1988). The endomesodermal expression in the hydrocoel also disappeared after the larval attachment. In the early attached larva, six3 was expressed in a part of inner cell masses which was formed from invaginated stomodeum (Fig. 3c, arrowheads). These expression patterns were maintained in the early cystidean stage (Fig. 3d, arrowheads). The six3-positive cell masses developed into the epithelium which covers the oral surface of the podia (Fig. 3e, arrowheads).
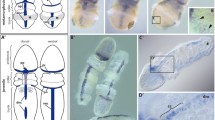
Spatial expression patterns of six3 (a–e), pax6 (f–j), and otx (k–o) throughout the larval development of A. japonica. Expression of the genes is represented by the sections of the WISH samples, which were counterstained with Nuclear Fast Red. The planktonic larvae (a, b, f, g, k, l) are viewed ventrally with the vegetal pole down. The settled larvae (c–e, h–j, m–o) are oriented with the attachment point to down. Red asterisks on c, h, i, m, n, o indicate non-specific signals which often appear around the attachment point. asix3 expression in an early doliolaria. Expression in the anterior enterohydrocoel is indicated by an arrow. The ectodermal expression is represented by double-arrowheads. bsix3 expression in a doliolaria. Expression in the hydrocoel is indicated by an arrow. The ectodermal expression is represented by double-arrowheads. csix3 expression in an early attached larva. Arrowheads represent the expression in the stomodeum. dsix3 expression in an early cystidean. Expression in the stomodeum derivative cells is represented by arrowheads. esix3 expression in a late cystidean. The expression detected in the oral surface of podia is indicated by arrowheads. fpax6 expression in an early doliolaria. The expression in the anterior enterohydrocoel is indicated by an arrow. gpax6 expression in a doliolaria. The expression in the hydrocoel is indicated by an arrow. hpax6 expression in an early attached larva. An arrowhead represents the expression in the stomodeum. ipax6 expression in an early cystidean. The expression in the stomodeum-derivative cells is represented by arrowheads. jpax6 expression in a late cystidean. The expression in the outer layers of all podia is indicated by arrowheads. kotx expression in a early doliolaria. Asterisks depict the point of strong expression of the ectoderm. The expression in the posterior enterohydrocoel is indicated by an arrow. lotx expression in a doliolaria. Asterisks depict the point of ciliary bands, where the strong expression is observed. The expression in the enteric sac is indicated by an arrow. motx expression in an early attached larva. An arrowhead represents the expression in the stomodeum. notx expression in an early cystidean. The expression in the stomodeum derivative cells is represented by arrowheads. ootx expression in a late cystidean. Arrowheads depict the expression in the oral ring nerve. Scale bars represent 100 μm
pax6 expression was observed in the anterior region of the enterohydrocoel in the early doliolaria (Fig. 3f, arrow) and in the hydrocoel in the late doliolaria (Fig. 3g, arrow). The expression pattern of pax6 in the endomesoderm was the same as that of six3 (Fig. 3a, b, arrows). After the larval attachment, the endomesodermal expression of pax6 decreased to a non-detectable level, and the expression of pax6 started in the inner cell mass derived from invaginated stomodeum (Fig. 3h, arrowhead). The expression area of pax6 expanded in the cell mass in the early cystidean (Fig. 3i, arrowheads). The pax6 expressing cell mass developed into the outer layer of the podia in the mouth-opening stage (Fig. 3j, arrowheads).
otx was broadly expressed in the ectoderm of the early doliolaria (Fig. 3k). The expression was especially stimulated in four circumferential belts and the most posterior wall (Fig. 3k, asterisks). The otx-negative cells were located in the positions of future ciliary bands. otx was also expressed in the posterior part of the enterohydrocoel (Fig. 3k, arrow), which is the precursor of the enteric sac. The expression in the ectoderm and the endomesoderm continued until the late doliolaria stage. otx was strongly expressed in the ciliary bands in the late doliolaria (Fig. 3l, asterisks) and in the enteric sac (Fig. 3l, arrow). After the larval attachment, the expression of otx observed in the planktonic larvae was diminished to a non-detectable level, and the expression of otx in the wall of the invaginating stomodeum was newly observed (Fig. 3m, arrowhead). The expression of otx in the stomodeum continued in the cell mass exhibiting a doughnut shape which derives from the wall of the stomodeum in the early cystidean (Fig. 3n, arrowheads). In the mouth-opening stage larva, otx was expressed in a ring surrounding the mouth (Fig. 3o, arrowheads).
Spatial expression patterns of six3, pax6, and otx in the juvenile tissues
In the juvenile of A. japonica, six3 expression was observed in the basal part of the podia (Fig. 4a, arrows; b) and the basal lamina of the food grooves which includes oral radial nerves (Fig. 4a, b, arrowheads). The oral ring nerve was also six3 positive (Supplementary Fig. 4a). The six3 expression in the podia was not observed in the podia around the mouth (Fig. 4a). In the calyx, six3 was expressed in the axial gland (Fig. 4c, arrow). Faint expression of six3 was also observed in the aboral ring nerve (Fig. 4c, arrowheads). The deep stain in the filament-like structures in the chambered organ (Fig. 4c, asterisks) was an artifact, which was also observed in the negative control sample (Fig. 4j, asterisks).
Spatial expression patterns of six3 (a–c), pax6 (d–f), and otx (g–i) in A. japonica juvenile. Ventral views of the WISH samples (a, d, g) or the dissected WISH samples which represent sagittal sections of the arms (b, e, h) and horizontal sections of the calyx (c, f, i, j). For the dissected arm samples (b, e, h), distal side of the sample is arranged to left, and ventral (oral) side is arranged to up. a A whole-mount image of the expression of six3 in the juvenile. six3 expression was detected in the basal part of the podia (arrows) and the basal lamina of the ambulacral grooves (arrowheads). bsix3 expression in the arm. Basal cells of the almost all podia (pod) on the arm were six3 positive. An arrowhead represents six3 expression in the basal lamina. csix3 expression in the calyx. An arrow represents six3 expression in the axial gland. Arrowheads depict expression of six3 in the aboral ring nerve. d A whole-mount image of the expression of pax6 in the juvenile. All podia represented the strong expression of pax6. epax6 expression in the arm. The strong pax6 expression in the podia was limited in the outer layer. fpax6 expression in the calyx. An arrow represents pax6 expression in the axial gland. g A whole-mount image of the expression of otx in the juvenile. Arrows indicate the otx expression in the distal podia. hotx expression in the arm. Arrows indicate the otx-positive podia on the distal part of the arm. Arrowheads indicate the otx-negative podia on the proximal part of the arm. iotx expression in the calyx. No significant signal was observed. j A calyx of the no probe negative sample. Red asterisks on c, f, i, and j indicate the non-specific signals. Abbreviations: ag, axial gland; a.rn, aboral ring nerve; m, mouth; pod, podia. Scale bars represent 1 mm (a, d, g), or 100 μm (b, c, e, f, h, i, j)
pax6 was expressed in the outer layer of all the podia (Fig. 4d, e). In the calyx, pax6 was expressed in the axial gland (Fig. 4f, arrow). No significant expression of pax6 was observed in other tissues in the calyx.
otx was expressed in the podia in the distal tip of the arms (Fig. 4g, h, arrows). Podia in the proximal part of the arms did not express otx (Fig. 4h, arrowheads). No significant expression of otx was observed in the calyx (Fig. 4i).
Discussion
Involvement of the three homeobox genes in the patterning of larval endomesoderm
In the planktonic larvae of A. japonica, six3 and pax6 were expressed in the anterior enterohydrocoel of early doliolaria and the hydrocoel of doliolaria, while otx was expressed in the posterior enterohydrocoel of early doliolaria and the enteric sac of doliolaria (Fig. 5a, b). As the enterohydrocoel is the term for the closed archenteron of feather stars (Hyman 1955; Kubota 1969), these results suggest that six3, pax6, and otx are involved in the patterning of the archenteron of A. japonica planktonic larvae. In the planktonic larvae of a sea lily crinoid Metacrinus rotundus, six3, pax6, otx, and hox genes are involved in A-P patterning of the archenteron and its derivatives which include axohydrocoel, enteric sac, and left and right somatocoels (Hara et al. 2006; Omori et al. 2011). While the precursor of the somatocoels has not been clearly detected in A. japonica, somatocoels of other feather star Antedon adriatica are known to be derived from posterior enterohydrocoel, which is separated by the anterior enterohydrocoel by a constriction that emerges on the middle of the enterohydrocoel (Hyman 1955). Our results suggested that the involvement of six3, pax6, otx, and hox genes in A-P patterning of the endomesoderm is a basal feature in crinoids, although external shapes of the early larvae are not similar between a sea lily and feather stars (Hyman 1955; Nakano et al. 2003),
Schematic illustrations of the expression patterns of six3, pax6, and otx in the larvae and juvenile of A. japonica. Semi-doliolaria (a) and doliolaria (b) are the ventral view with the posterior side down. Thus, the left side of the figures corresponds to the right side of the larvae. Attachment stage (c), cystidean (d), pentacrinoid (e), and juvenile (f) represent the lateral view with the attachment point to the bottom. Green-, pink-, and blue-colored areas represent the gene-expressing area of six3, pax6, and otx, respectively. Stripes of green and pink, green and blue, or pink and blue exhibit the area where 2 different genes are co-expressed. Pale green- and pale pink-colored areas in f represent the weak expressions of six3 and pax6, respectively. Abbreviations: cb, ciliary bands; e.hc, enterohydrocoel; es, enteric sac; hc, hydrocoel; l.nv, larval nerve; o.rn, oral ring nerve; std, stomodeum
otx expression in the endodermal tissues is widely observed in deuterostomes (Ang et al. 1994; Gan et al. 1995; Wada et al. 1996; Williams and Holland 1996; Mitsunaga-Nakatsubo et al. 1998; Tomsa and Langeland 1999; Harada et al. 2000; Shoguchi et al. 2000; Nielsen et al. 2003; Lowe et al. 2003; Hinman et al. 2003). In the embryos and larvae of an amphioxus Branchiostoma floridae, six3 and pax6 are expressed in the anterior tip of the endoderm (Glardon et al. 1998; Kozmik et al. 2007), which is separated by the more posterior otx-expressing endodermal area within the progress of its development (Williams and Holland 1996). The expression of these three genes in the mesendoderm is also reported in some hemichordates (Lowe et al. 2003; Gonzalez et al. 2017). The similarity of the expression patterns of six3, pax6, and otx among crinoids, hemichordates, and amphioxus implies the involvement of these genes in A-P patterning of the endomesoderm in the basal deuterostomes. However, pax6 is only expressed in the ectodermal tissues in ascidians (Glardon et al. 1997; Irvine et al. 2008), hemichordates (Lowe et al. 2003; Gonzalez et al. 2017), and a basal metazoan xenacoelomorph (Hejnol and Martindale 2008). Further studies on more species of deuterostomes and basal metazoans will reveal the basal function of these genes in body patterning.
Ectodermal otx expression in the planktonic larvae and its putative function
In the doliolaria larva of A. japonica, otx was expressed in the circumferential ciliated bands. A doliolaria-type larva with some circumferential ciliated bands is known in crinoids, brittle stars, and sea cucumbers (Hyman 1955; Nakano et al. 2003). A sea cucumber Psolus chitinoides, which develops into doliolaria directly without the auricularia stage, expresses otx in all three ciliary bands of the doliolaria (Lowe and Wray 1997). In contrast, a different species of sea cucumber Apostichopus japonicus, which develops into both auricularia and following doliolaria, does not express otx in the ciliary bands of doliolaria (Shoguchi et al. 2000). Different properties of otx expression in ciliary bands in the doliolaria stage of different species suggest that otx is not responsible for the maintenance of circumferential ciliary bands. In the hatching stage of A. japonica, the entire surface of the larva is uniformly covered with cilia. The larva rearranges the ciliated cells to form the ciliary bands afterward. In the semi-doliolaria larva, the expression of otx is restricted to a prospective non-ciliary-band region. Taking the expression patterns of otx in the semi-doliolaria and the doliolaria together, it is suggested that otx is involved in the formation of ciliary bands by inducing cell migration of the ciliary cells. otx expression in the ciliary cells has been also observed in the polychaete protostomes (Arendt et al. 2001; Arenas-Mena and Wong 2007) and in the more basal metazoans like a sea anemone Nematostella vectensis (Mazza et al. 2007). The widely conserved otx expression pattern associated with the ciliary cells suggests an ancestral role of otx in forming the ciliary cells and/or regionalization of the ciliary band.
Switching of the expression status of three homeobox genes after the larval attachment
After attachment of the larva, the expression of six3, pax6, and otx completely ceased in the regions where these genes expressed in the planktonic larvae. In parallel, these genes start to be expressed in the wall of stomodeum (Fig. 5c). The temporal expression profiles of three genes also showed the change of expression phase after the larval attachment (Fig. 2). It has been reported that the adult nervous system begins to form at least 1 day after the larval attachment without any connection to the larval nerves (Nakano et al. 2009). These imply that the larval attachment triggers the transition of gene expression pattern to form adult structures.
The expressions of six3, pax6, and otx continued in the wall of the stomodeum in the cystidean (Fig. 5d), and the expression area of these genes shifted to the podia in the later stage (Fig. 5e). A previous immunohistochemical observation using sea star synaptotagmin antibody has suggested that primary oral nerves exist in the podia or in their precursors (Nakano et al. 2009). The pax6 expression pattern in the podia is similar to the patterns of synaptotagmin immuno-reactivity, and six3-expressing cells on the surface of the podia were also located in the synaptotagmin-positive area (Figs. 3e, j and 4b, e, Supplementary figure 1c). In addition, six3 expression was also observed in the oral nerves of the juvenile (Fig. 4a; supplementary Fig. 4a). These suggest that six3 and pax6 are involved in the formation of the oral nervous systems. It was also reported that otx is expressed in the oral ring nerve and the primary podia in sea urchins (Nielsen et al. 2003; Morris et al. 2004). In our results in A. japonica, otx was expressed in the ringed area around the mouth in the pentacrinoid (Fig. 3o, arrowheads), where the intense synaptotagmin immuno-reactivity was reported (Nakano et al. 2009). Taken together, six3, pax6, and otx are possibly involved in the formation of the oral nervous system in the post-attachment stages of A. japonica, and this property was probably acquired in the common ancestor of the echinoderms.
Gene expression in the juvenile tissues and its evolutionally relationship among deuterostomes
otx was expressed in the outer layer of the podia in the juvenile of A. japonica (Fig. 4g, h). otx expression in the outer layer of the podia has been also observed in some eleutherozoan echinoderms (Lowe and Wray 1997; Morris et al. 2004), suggesting that this expression pattern was acquired in the common ancestor of extant echinoderms. However, the otx expression in the podia was restricted to the distal tip of the arms, in contrast to the uniform expression in the entire podia observed in the eleutherozoans (Lowe and Wray 1997). It has been reported that pax6 is mainly expressed in the inner layer of podia in sea urchins (Czerny and Busslinger 1995; Ullrich-Luter et al. 2011), while pax6 is expressed in the outer layer of podia in A. japonica (Fig. 4d, e). These differences may reflect the morphological difference of the podia. The podia of the crinoids contain many sensory papillae on their surface, which are not observed in the podia of the eleutherozoans. As pax6 and otx are involved in the formation of optic sensory organs collaborating with six3 in various phyla (Callaerts et al. 1997; Gehring and Ikeo 1999; Arendt 2003; Zuber et al. 2003; Martínez-Morales et al. 2004; Stierwald et al. 2004; Kozmik 2005), our result implies the existence of some optic sensory organs on the podia of A. japonica. Further morphological and molecular studies will confirm the existence of podial sensory organs.
Crinoids have three nervous systems: oral, deeper oral, and aboral systems (Hyman 1955). Disruption of the aboral nerve center results in the disorganization of the arm movement, while removal of the oral and the deeper oral nervous systems does not affect its behavior (Marshall 1884). Thus, the aboral nervous system is considered as the main nervous system in crinoids (Hyman 1955). This evokes an idea that the aboral nerve center is evolutionally similar to the central nervous system of other animals. However, in the present study, homologs of the brain-patterning genes pax6 and otx were not expressed in the aboral nerve center of A. japonica (Figs. 4f, i and 5f). Other brain-patterning gene homolog six3 was weakly expressed in the whole aboral ring nerve (Figs. 4c and 5f); however, the expression was not biased in any axes. These imply that the aboral nervous system of crinoids is not homologous to the central nervous system of other animals. The arms of the crinoids play important roles in their feeding and movement. The aboral nerve center may originally develop in the basal echinoderm clade to function as the center for motor neuron which organizes the arm movement. Further morphological and molecular analyses will reveal the origin of the aboral nervous system.
References
Ang S-L, Conlon RA, Jin O, Rossant J (1994) Positive and negative signals from mesoderm regulate the expression of mouse Otx2 in ectoderm explant. Development 120:2979–2989
Arenas-Mena C, Wong KS-Y (2007) HeOtx expression in an indirectly developing polychaete correlates with gastrulation by invagination. Dev Genes Evol 217:373–384. https://doi.org/10.1007/s00427-007-0150-7
Arendt D (2003) Evolution of eyes and photoreceptor cell types. Int J Dev Biol 47:563–571
Arendt D, Technau U, Wittbrodt J (2001) Evolution of the bilaterian larval foregut. Nature 409:81–85. https://doi.org/10.1038/35051075
Bottjer DJ, Davidson EH, Peterson KJ, Cameron RA (2006) Paleogenomics of echinoderms. Science 314(80):956–960. https://doi.org/10.1126/science.1132310
Brusca RC, Moore W, Shuster SM (2016) Invertebrates, 3rd edn. Sinauer, Sunderland
Callaerts P, Halder G, Gehring WJ (1997) Pax-6 in development and evolution. Annu Rev Neurosci 20:483–532
Clark AH, Clark AM (1967) A monograph of the existing crinoids 1(5). Bull US Nat Mus 82:1–860
Cohen BL, Améziane N, Eleaume M, de Forges BR (2004) Crinoid phylogeny: a preliminary analysis (Echinodermata: Crinoidea). Mar Biol 144:605–617. https://doi.org/10.1007/s00227-003-1212-7
Czerny T, Busslinger M (1995) DNA-binding and transactivation properties of Pax-6: three amino acids in the paired domain are responsible for the different sequence recognition of Pax-6 and BSAP (Pax-5). Mol Cell Biol 15:2858–2571
Dan JC, Dan K (1941) Early development of Comanthus japonicus. Japanese J Zool 9:565–574
Dan K, Ando Y, Sekiguchi K, Watanabe H (eds) (1988) Development of Invertebrates, Vol 2. Baifukan, Tokyo. [In Japanese]
Gan L, Mao C-A, Wikramanayake AH, Angerer LM, Angerer RC, Klein WH (1995) An orthodenticle-related protein from Strongylocentrotus purpuratus. Dev Biol 167:517–528
Garstang W (1894) Preliminary note on a new theory of the phylogeny of the Chordata. Zool Anzeiger 27:122–125
Gehring WJ, Ikeo K (1999) Pax 6: mastering eye morphogenesis and eye evolution. Trends Genet 15:371–377
Glardon S, Callaerts P, Halder G, Gehring WJ (1997) Conservation of Pax-6 in a lower chordate, the ascidian Phallusia mammillata. Development 124:817–825
Glardon S, Holland LZ, Gehring WJ, Holland ND (1998) Isolation and developmental expression of the amphioxus Pax-6 gene (AmphiPax-6): insights into eye and photoreceptor evolution. Development 125:2701–2710
Gonzalez P, Uhlinger KR, Lowe CJ (2017) The adult body plan of indirect developing hemichordates develops by adding a Hox-patterned trunk to an anterior larval territory. Curr Biol 27:87–95. https://doi.org/10.1016/j.cub.2016.10.047
Hara Y, Yamaguchi M, Akasaka K, Nakano H, Nonaka M, Amemiya S (2006) Expression patterns of Hox genes in larvae of the sea lily Metacrinus rotundus. Dev Genes Evol 216:797–809. https://doi.org/10.1007/s00427-006-0108-1
Harada Y, Okai N, Taguchi S, Tagawa K, Humphreys T, Satoh N (2000) Developmental expression of the hemichordate otx ortholog. Mech Dev 91:337–339
Hejnol A, Martindale MQ (2008) Acoel development supports a simple planula-like urbilaterian. Phil Trans R Soc B 363:1493–1501. https://doi.org/10.1098/rstb.2007.2239
Hinman VF, Nguyen AT, Davidson EH (2003) Expression and function of a starfish Otx ortholog, AmOtx: a conserved role for Otx proteins in endoderm development that predates divergence of the eleutherozoa. Mech Dev 120:1165–1176. https://doi.org/10.1016/j.mod.2003.08.002
Hyman LH (1955) The invertebrates, vol IV, Echinodermata. McGraw-Hill, New York
Irvine SQ, Fonseca VC, Zompa MA, Antony R (2008) Cis-regulatory organization of the Pax6 gene in the ascidian Ciona intestinalis. Dev Biol 317:649–659. https://doi.org/10.1016/j.ydbio.2008.01.036
Janies DA, Voight JR, Daly M (2011) Echinoderm phylogeny including Xyloplax, a progenetic asteroid. Syst Biol 60:420–438. https://doi.org/10.1093/sysbio/syr044
Kozmik Z (2005) Pax genes in eye development and evolution. Curr Opin Genet Dev 15:430–438. https://doi.org/10.1016/j.gde.2005.05.001
Kozmik Z, Holland ND, Kreslova J, Oliveri D, Schubert M, Jonasova K, Holland LZ, Pestarino M, Benes V, Candiani S (2007) Pax-Six-Eya-Dach network during amphioxus development: conservation in vitro but context specificity in vivo. Dev Biol 306:143–159. https://doi.org/10.1016/j.ydbio.2007.03.009
Kubota H (1969) Development of Comanthus japonica I. From spawning to attachment. Jpn J Dev Biol 23:92–93 (in Japanese)
Kubota H (1970) Development of Comanthus japonica II. After setting. Jpn J Dev Biol 24:40–41 (in Japanese)
Lowe CJ, Wray GA (1997) Radical alterations in the roles of homeobox genes during echinoderm evolution. Nature 389:718–721. https://doi.org/10.1038/39580
Lowe CJ, Wu M, Salic A, Evans L, Lander ES, Stange-Thomann N, Gruber CE, Gerhart J, Kirschner M (2003) Anteroposterior patterning in hemichordates and the origins of the chordate nervous system. Cell 113:853–865
Marshall AM (1884) On the nervous system of Antedon rosaceus. Q J Microsc Sci 24:507–548
Martínez-Morales JR, Rodrigo I, Bovolenta P (2004) Eye development: a view from the retina pigmented epithelium. BioEssays 26:766–777. https://doi.org/10.1002/bies.20064
Mazza ME, Pang K, Martindale MQ, Finnerty JR (2007) Genomic organization, gene structure, and developmental expression of three clustered otx genes in the sea anemone Nematostella vectensis. J Exp Zool Part B Mol Dev Evol 308B:494–506. https://doi.org/10.1002/jez.b.21158
Mitsunaga-Nakatsubo K, Akasaka K, Sakamoto N, Takata K, Matsumura Y, Kitajima T, Kusunoki S, Shimada H (1998) Differential expression of sea urchin Otx isoform (HpOtxE and HpOtxL) mRNAs during early development. Int J Dev Biol 42:645–651
Morris VB, Zhao J-T, Shearman DCA, Byrne M, Frommer M (2004) Expression of an Otx gene in the adult rudiment and the developing central nervous system in the vestibula larva of the sea urchin Holopneustes purpurescens. Int J Dev Biol 48:17–22
Nakano H, Hibino T, Oji T, Hara Y, Amemiya S (2003) Larval stages of a living sea lily (stalked crinoid echinoderm). Nature 421:158–160. https://doi.org/10.1038/nature01236.1
Nakano H, Nakajima Y, Amemiya S (2009) Nervous system development of two crinoid species, the sea lily Metacrinus rotundus and the feather star Oxycomanthus japonicus. Dev Genes Evol 219:565–576. https://doi.org/10.1007/s00427-010-0317-5
Nielsen MG, Popodi EM, Minsuk S, Raff RA (2003) Evolutionary convergence in Otx expression in the pentameral adult rudiment in direct-developing sea urchins. Dev Genes Evol 213:73–82. https://doi.org/10.1007/s00427-003-0299-7
Omori A, Akasaka K, Kurokawa D, Amemiya S (2011) Gene expression analysis of Six3, Pax6, and Otx in the early development of the stalked crinoid Metacrinus rotundus. Gene Expr Patterns 11:48–56. https://doi.org/10.1016/j.gep.2010.09.002
Pani AM, Mullarkey EE, Aronowicz J, Assimacopoulos S, Grove EA, Lowe CJ (2012) Ancient deuterostome origins of vertebrate brain signalling centres. Nature 483:289–294. https://doi.org/10.1038/nature10838
Paul CRC, Smith AB (1984) The early radiation and phylogeny of echinoderms. Biol Rev 59:443–481
Reichert H (2009) Evolutionary conservation of mechanisms for neural regionalization, proliferation and interconnection in brain development. Biol Lett 5:112–116. https://doi.org/10.1098/rsbl.2008.0337
Rouse GW, Jermiin LS, Wilson NG, Eeckhaut I, Lanterbecq D, Oji T, Young CM, Browning T, Cisternas P, Helgen LE, Stuckey M, Messing CG (2013) Fixed, free, and fixed: the fickle phylogeny of extant Crinoidea (Echinodermata) and their Permian-Triassic origin. Mol Phylogenet Evol 66:161–181. https://doi.org/10.1016/j.ympev.2012.09.018
Scouras A, Smith MJ (2006) The complete mitochondrial genomes of the sea lily Gymnocrinus richeri and the feather star Phanogenia gracilis: signature nucleotide bias and unique nad4L gene rearrangement within crinoids. Mol Phylogenet Evol 39:323–334. https://doi.org/10.1016/j.ympev.2005.11.004
Shibata TF, Sato A, Oji T, Akasaka K (2008) Development and growth of the feather star Oxycomanthus japonicus to sexual maturity. Zool Sci 25:1075–1083. https://doi.org/10.2108/zsj.25.1075
Shoguchi E, Harada Y, Numakunai T, Satoh N (2000) Expression of the otx gene in the ciliary bands during sea cucumber embryogenesis. genesis 27:58–63
Stierwald M, Yanze N, Bamert RP, Kammermeier L, Schmid V (2004) The Sine oculis/Six class family of homeobox genes in jellyfish with and without eyes: development and eye regeneration. Dev Biol 274:70–81. https://doi.org/10.1016/j.ydbio.2004.06.018
Tomsa JM, Langeland JA (1999) Otx expression during lamprey embryogenesis provides insights into the evolution of the vertebrate head and jaw. Dev Biol 207:26–37. https://doi.org/10.1006/dbio.1998.9163
Ullrich-Luter EM, Dupont S, Arboleda E, Hausen H, Arnone MI (2011) Unique system of photoreceptors in sea urchin tube feet. Proc Natl Acad Sci 108:8367–8372. https://doi.org/10.1073/pnas.1018495108
Wada S, Katsuyama Y, Sato Y, Itoh C, Saiga H (1996) Hroth, an orthodenticle-related homeobox gene of the ascidian, Halocynthia roretzi: its expression and putative roles in the axis formation during embryogenesis. Mech Dev 60:59–71
Williams NA, Holland PWH (1996) Old head on young shoulders. Nature 383:490
Zuber ME, Gestri G, Viczian AS, Barsacchi G, Harris WA (2003) Specification of the vertebrate eye by a network of eye field transcription factors. Development 130:5155–5167. https://doi.org/10.1242/dev.00723
Acknowledgments
We are grateful to Ms. Toko Tsurugaya (Urawa University) for her support in culturing and handling A. japonica. We also appreciate Mr. Minoru Sekimoto, Mr. Mamoru Sekifuji, Mr. Hisanori Kohtsuka, and Ms. Natsuko Sugii (Misaki Marine Biological Station) for their help in collecting and culturing of A. japonica, and we are obliged to Dr. Yoko Nakajima (Keio University), Dr. Daisuke Kurokawa, and Dr. Mariko Kondo (Misaki Marine Biological Station) for their valuable comments.
Funding
This work was supported in part by the Global COE Program (Integrative Life Science Based on the Study of Biosignaling Mechanisms), MEXT, Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants performed by any of the authors. All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Additional information
Communicated by Mark Q. Martindale
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Omori, A., Shibata, T.F. & Akasaka, K. Gene expression analysis of three homeobox genes throughout early and late development of a feather star Anneissia japonica. Dev Genes Evol 230, 305–314 (2020). https://doi.org/10.1007/s00427-020-00665-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-020-00665-6