Abstract
Zinc finger transcription factors of the Sp6–9 group are evolutionarily conserved in all metazoans and have important functions in, e.g., limb formation and heart development. The function of Sp6–9-related genes has been studied in a number of vertebrates and invertebrates, but data from chelicerates (spiders and allies) was lacking so far. We have isolated the ortholog of Sp6–9 from the common house spider Parasteatoda tepidariorum and the cellar spider Pholcus phalangioides. We show that the Sp6–9 gene in these spider species is expressed in the developing appendages thus suggesting a conserved role in limb formation. Indeed, RNAi with Sp6–9 in P. tepidariorum leads not only to strong limb defects, but also to the loss of body segments and head defects in more strongly affected animals. Together with a new expression domain in the early embryo, these data suggest that Sp6–9 has a dual role P. tepidariorum. The early role in head and body segment formation is not known from other arthropods, but the role in limb formation is evolutionarily highly conserved.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Sp family of transcription factors contains evolutionarily conserved zinc finger proteins which have been found in the genome of all metazoan groups including basal groups like cnidarians or placozoans (see Schaeper et al. 2010). The proteins are characterized by the presence of three consecutive zinc finger motifs and, more N-terminally, a transactivation domain called Buttonhead-box (Btd-box). The control regions of many tissue-specific as well as ubiquitously expressed genes contain binding sites for Sp-family transcription factors, indicating that Sp-family transcription factors have the potential to regulate a large number of target genes (Philipsen and Suske 1999; Suske et al. 2005). Indeed, it has been demonstrated that Sp-family genes have important functions in the embryonic development of humans and various other animal species, e.g., in processes such as cell growth, differentiation, organ development, and cancer (reviewed in Suske et al. 2005).
Previous work has shown that the Sp-family genes can be grouped into three different clades, namely Sp1–4, Sp5/Btd and Sp6–9 clade (Schaeper et al. 2010). Genes of the Sp6–9 clade have been studied in several animal species so far. In vertebrates, the Sp6–9 group comprises four closely related genes, Sp6 to Sp9, that are involved in a variety of developmental processes including the development of hair follicles, teeth, appendages, bones, the nervous system (including sensory organs and the brain), the ears, and the lungs (Hertveldt et al. 2008; Koga et al. 2005; Haro et al. 2014; Chung et al. 2014; Li et al. 2011; Kawakami et al. 2004). Arthropods, on the other hand, possess only a single gene from the Sp6–9 group. In the common fruit fly Drosophila melanogaster, the Sp6–9 ortholog is called D-Sp1, because its relationship to other Sp genes was still unclear at the time of its discovery. It is involved in leg development and the formation of mechanosensory organs (Schöck et al. 1999). In the red flour beetle Tribolium castaneum, the Sp6–9 ortholog, called Tc-Sp8, has a role in leg and antenna development (Beermann et al. 2004), and RNAi experiments in the milkweed bug Oncopeltus fasciatus have also shown that the Sp6–9 ortholog of O. fasciatus (termed Of-Sp8/9) is required for the development of the legs, antennae, and the anterior appendages of the rostrum (consisting of the labium-bearing segment, the filiform maxillae and mandibles, and a pointed labrum) (Schaeper et al. 2009). The expression pattern of Sp6–9 orthologs in basally branching insects and in the crustacean Parhyale hawaiensis (Schaeper et al. 2010) suggests that the role in appendage development is conserved among mandibulate arthropods.
While Sp6–9 genes have been studied extensively in vertebrates and in a number of mandibulate arthropods, no chelicerate (spiders and allies) representative of Arthropoda has been studied so far. We have therefore isolated the ortholog of Sp6–9 from the common house spider Parasteatoda tepidariorum, and we also supplement these data with the Sp6–9 ortholog from the cellar spider Pholcus phalangioides. We show that the Sp6–9 gene in these two spider species is expressed in the developing appendages thus suggesting a conserved role in limb formation in spiders. Gene function studies in P. tepidariorum using parental RNAi with Sp6–9 lead not only to strong limb defects, but also to the loss of body segments and head defects in more strongly affected animals. Together with a new expression domain in the early embryo of P. tepidariorum, these data suggest that Sp6–9 has a dual role in P. tepidariorum. The early role in head and body segment formation is not known from other arthropods, but the role in limb formation is evolutionarily highly conserved.
Materials and methods
P. tepidariorum and Ph. phalangioides husbandry and embryo fixation
Our P. tepidariorum and Ph. phalangioides husbandries are kept at controlled temperature (25 °C) and dark/light cycles of 14:10 h, respectively. The animals are kept separate in plastic vials sealed with styrene foam plugs. Juveniles of both spider species are fed with D. melanogaster flies. Older stages as well as adults of Ph. phalangioides and P. tepidariorum are fed with larger flies (Musca domestica and Lucilia caesar) and juvenile crickets (Acheta domesticus), respectively. Appropriate humidity levels were provided by regularly supplying water to the soil. After mating, several cocoons are produced by the females at irregular intervals. Each cocoon contains between 50 and 100 eggs (in Ph. phalangioides) and about 300 eggs (in P. tepidariorum). Freshly made cocoons were kept separately in glass test tubes until the eggs were removed for fixation. Fixation of Ph. phalangioides embryos was performed according to the fixation protocol previously published for Cupiennius salei (Prpic et al. 2008a) and with minor modifications for P. tepidariorum. Staging of the embryos was done using the staging schemes by Mittmann and Wolff (2012) and Turetzek and Prpic (2016).
Gene sequences and molecular cloning
cDNA was synthesized with the SMARTer RACE kit (Clontech Laboratories) from total RNA of mixed embryonic stages prepared with TRIzol (Invitrogen) according to the manufacturer’s instructions. We have used transcriptome sequence resources of P. tepidariorum (Posnien et al. 2014) and Ph. phalangioides (unpublished data) to identify sequences similar to the Sp6–9 ortholog from D. melanogaster (termed D-Sp1) by Basic Local Alignment Search Tool (BLAST) search. A single transcript was identified in each species and was confirmed by phylogenetic analysis to be a Sp6–9 ortholog (see the “Results” section). Fragments of these genes were cloned with the following primers: Pt_Sp6–9_forward 5′-GCA GCG TTT GAA TTC GTG GGA TAT C-3′ and Pt_Sp6–9_reverse 5′-CCG ATG GGT CCT GAG GTA ACT GC-3′ for Sp6–9 from P. tepidariorum and Pp_Sp6–9_forward 5′-GGC AGC GCAA TGC AAC AAG CTT GCG-3′ and Pp_Sp6–9_reverse 5′-CCC AAC GCG TAA TCC GAC GTC GGG-3′ for Sp6–9 from Ph. phalangioides. The sequences are available from GenBank under the accession numbers MG182359 (Sp6–9 from P. tepidariorum) and MG182360 (Sp6–9 from Ph. phalangioides). The sequence and analysis of the P. tepidariorum genes orthodenticle-1 (GenBank: AB096074.1, Akiyama-Oda and Oda 2003), Distal-less (GenBank: FM876233.2, Pechmann and Prpic 2009), dachshund1 (GenBank: LN877883.1, Turetzek et al. 2016), and hedgehog (GenBank: AB125742.1, Oda et al. 2007) have been published previously.
Phylogenetic sequence analysis
For the phylogenetic sequence analysis, we used selected protein sequences of Sp genes from diverse metazoan species. The species, species abbreviations (as used in Fig. 1), and sequence accession numbers are available in TXT format as Online Resource 1. We have used the amino acid sequences of the proteins from the BTD-box motif to the end of the third zinc finger (sequences available in TXT format as Online Resource 2) and have aligned them with Clustal Omega (Sievers et al. 2011). The alignment is available in NEXUS format as Online Resource 3. Based on this alignment, we have performed a Bayesian Markov chain Monte Carlo analysis using the parallel version of MrBayes v.3.2.5 (Ronquist and Huelsenbeck 2003). We sampled across fixed amino acid rate matrices (aamodel = mixed) and generated topological convergence (average standard deviation of split frequencies below 0.01) after 300,000 generations without Metropolis coupling.
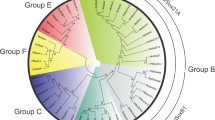
Phylogenetic analysis of Sp genes from diverse Metazoa . Unrooted 50% majority rule consensus tree after Bayesian Markov chain Monte Carlo analysis. Branch lengths in the phylogram give the expected substitutions per site. Numbers at the tree edges are clade credibility values, which are a measure of the probability of each clade in the tree. Orthologs of spider Sp genes identified in the present study are indicated by the red arrows. For sequence accession numbers, please see Online Resource 1. Am, Apis mellifera (honey bee); Dm, Drosophila melanogaster (common fruit fly); Dr, Danio rerio (zebrafish); Of, Oncopeltus fasciatus (milkweed bug); Fc, Folsomia candida (white springtail); Hs, Homo sapiens (human); Ph, Parhyale hawaiensis (Hawaiian beach-hopper, amphipod crustacean); Tc, Tribolium castaneum (red flour beetle)
Whole-mount in situ hybridization and nuclei stainings
Whole-mount in situ hybridization was performed according to the protocol by Prpic et al. (2008b). To visualize the nuclei under UV light, the embryos were treated with SYTOX-Green (Invitrogen) as previously described (Turetzek et al. 2016). Digital images were taken either with white light or with UV light under a Leica M205 FA binocular microscope equipped with a QImaging Micropublisher 5.0 RTV camera. A combination of white light and UV light was used to better depict embryo morphology in the specimens stained with blue precipitate resulting from the in situ hybridization reaction. Correction of color values and brightness of all digital images was done with Adobe Photoshop CS5 Extended for Apple Macintosh.
Cuticle preparations and imaging of P. tepidariorum larvae and nymphs
Spiders have two different principal postembryonic stages, larvae and nymphs (Foelix 2011). Larvae are the first postembryonic stage after hatching; they are largely immobile, do not feed (except from yolk remnants), and do not have hairs and bristles yet. All stages after the larval stage are called nymphs. These are fully mobile, feed from prey, and have a full complement of hairs and bristles. P. tepidariorum larvae were placed in a small drop of Voltalef H10S oil on a glass microscope slide, covered with a splinter of a glass cover slip, and fixed on a heat block at 90 °C for 2 min. After fixation, the slides can be stored in the fridge for 2–3 months. The nymphs were cleared on a microscope slide in a drop of lactic acid/Hoyers medium (1:1), covered with a glass cover slip. Before clearing, the opisthosoma of the nymphs was removed. In order to digest all proteins, the specimens were incubated at 65 °C over night until only the cuticle of the prosoma (including the appendages) remained. Images of larvae were obtained using a confocal laser scanning microscope (LSM 510, Carl Zeiss, Germany) using the autofluorescence of the cuticle. Nymphs were photographed under a Zeiss-Axioplan-2 microscope equipped with an Intas digital camera. Correction of black/white values and contrast of all digital images was done with Adobe Photoshop CS5 Extended for Apple Macintosh.
Parental RNAi in P. tepidariorum
Female spiders were mated several hours prior to injection. The double-stranded RNA (dsRNA) was injected at the lateral side of the opisthosoma of the spiders. Injection took place once per day on three consecutive days; each injection contained a volume of 4 μl dsRNA preparation with a concentration of 3 μg/μl dsRNA in injection buffer (1.4 mM NaCl, 0.07 mM Na2H PO4, 0.03 mM KH2 PO4, 4 mM KCL in water). Controls were injected with 4 μl of injection buffer. We injected eight adult females for the Sp6–9 RNAi experiment and three adult females for the negative controls. The first six cocoons of the injected females were collected, pooled and analyzed, and the obtained phenotypes were categorized into strong and weak phenotypes (explained in the “Results” section). A statistical summary of the phenotype distribution is available as Online Resource 4 (for the RNAi experiment) and Online Resource 5 (for the controls).
Results
Isolation of a Sp6–9 ortholog from P. tepidariorum and Ph. phalangioides
The Sp genes can be grouped into three principal clades, termed Sp1–4, Sp5/Btd, and Sp6–9 genes (Schaeper et al. 2010). The two sequences from P. tepidariorum and Ph. phalangioides were found after searching the transcriptome sequences using a Sp6–9 sequence via BLAST similarity search (see the “Materials and methods” section), thus suggesting that the two genes are similar in sequence to Sp6–9. Additionally, we have performed a phylogenetic sequence analysis using Bayesian Markov chain Monte Carlo inference. The resulting tree corroborates the presence of three separate monophyletic clades (Fig. 1), and the two spider sequences are included in the Sp6–9 clade with high confidence. Thus, the analysis of the evolutionary relationships of the sequences supports the initial assessment based on sequence similarity and strongly suggests that the two spider genes are indeed orthologs of Sp6–9.
Embryonic expression of Sp6–9 in P. tepidariorum and Ph. phalangioides
In P. tepidariorum, the expression of Sp6–9 is first detected at stage 5. At this stage, Sp6–9 is expressed in a broad domain along the circumference of the germ disc (Fig. 2 (a, a')). At early stage 6, this ring-shaped domain is more narrow and is now located some distance away from the rim of the germ disc (red arrow, Fig. 2 (b, b')). During the transition from radial to bilateral symmetry at late stage 6, the formerly complete ring of Sp6–9 expression is opened up along the lateral sides (i.e., the future dorsal side) (Fig. 2 (c (black arrows), c’)). Coinciding with the formation of the germ band at stage 7, Sp6–9 is now expressed in weak segmental stripes that include the region from the pedipalpal to the fourth walking-leg segment (Fig. 3a). As soon as the segmentation of the germ band is more pronounced at stage 8.1, the expression levels of the segmental stripes is elevated (Fig. 3b, c). At stage 8.2, the Sp6–9 expression is also detected in the cheliceral segment (Fig. 3d) and the formation of the ventral sulcus is initated, which divides the expression in the segmental stripes during further development (Fig. 3e). Simultaneously with the fully formed ventral sulcus, at stage 9.1, the limb buds are formed and the expression of Sp6–9 is now restricted to the forming buds of all prosomal appendages (Fig. 3e). In addition, a faint segmental expression starts at stage 8.2 in the opisthosoma (Fig. 3d) and is more clearly visible in the ventral part during stage 9.1 (Fig. 3e, arrowheads). At stage 9.2, the prosomal appendages have grown out slightly and the expression of Sp6–9 now comprises a strong distal spot and a weaker proximal ring in the walking legs and pedipalps. In the chelicerae, the expression still comprises only a distal spot (Fig. 4a). As the appendages grow further during the next stages, the bipartite expression domain of Sp6–9 remains in the legs and pedipalps; in the chelicerae, no proximal expression arises and only a strong distal domain is present (Fig. 4b, c). When the germ band is fully formed and all opisthosomal segments are present, there is also a small expression domain of Sp6–9 at the posterior end of the germ band (Fig. 4d, arrow).
Early expression of Sp6–9 in P. tepidariorum embryos. Germ discs at stage 5 (a), early stage 6 (b), and late stage 6 (c). In a'–c', fluorescent and white light microscopy has been combined to visualize the morphology of the germ disc. This shows that initially, the rim of the germ disc (black arrow) and the ring of Sp6–9-expressing cells (red arrow) coincide, but later, the ring of expressing cells is shifted towards the disc center. The arrows in c point to the opened expression ring after the break of the radial symmetry
Expression of Sp6–9 in P. tepidariorum between stage 7 and stage 9.1. a Stage 7 embryo in lateral-ventral view. b Stage 8.1 embryo in lateral-ventral view. c Stage 8.1 embryo in lateral view. d Stage 8.2 embryo in lateral-ventral view. e Stage 9.1 embryo in lateral-ventral view. Arrowheads point to expression spots in the opisthosoma. All embryos are shown with anterior to the left. ch, chelicera segment; L1–L4, walking leg segments 1 to 4; pp, pedipalp segment
Late expression of Sp6–9 in P. tepidariorum between stage 9.1 and stage 11. a Stage 9.2 embryo in lateral-ventral view. b Stage 10 embryo in lateral-ventral view. c Stage 11 embryo in ventral view. d Stage 11 embryo in frontal view. The arrow points to the expression at the end of the opisthosoma. All embryos are shown with anterior to the left, except the frontal view in d. ch, chelicera segment; L1–L4, walking leg segments 1 to 4; pp, pedipalp segment; lr, labrum
In Ph. phalangioides, the youngest stages were not available for this study. However, the remaining stages show a Sp6–9 expression pattern very similar to the expression described for P. tepidariorum (see Online Resource 6). The youngest Ph. phalangioides embryos available for this study are comparable to early stage 7 P. tepidariorum embryos and thus have finished the transition from radial to bilateral symmetry and are at the beginning of germ band formation. As observed in P. tepidariorum at the comparable stage, five weakly expressed segmental stripes in the prosomal segments from Pp to L4 are present (Online Resource 6, panel a, arrowheads and arrow). During the following approximately 5 h (comparable to stages 8.1 and 8.2 in P. tepidariorum), these stripes increase in expression level and similar to P. tepidariorum, an additional expression domain in the cheliceral segment appears (Online Resource 6, panel b) until strong stripes of Sp6–9 expression are present in all prosomal segments from the cheliceral segment to the fourth walking-leg segment (Online Resource 6, panel c). Thus, during these early germ band stages, the Sp6–9 expression does not show any spatial or temporal differences comparing haplogyne and entelegyne spider species. The only visible difference is the length of the stripes, which is caused by the differences in the shape of the germ band, which is broader and less condensed in Ph. phalangioides. This effect is gone when the germ band is fully formed and the limb primordia start to develop. Then, also, the expression of Sp6–9 in Ph. phalangioides becomes more restricted to the site of limb formation and the limb buds (Online Resource 6, panels d, e). As the appendages grow longer, they show a bipartite expression pattern with a strong spot in the tips and a weaker ring at the base, except for the chelicerae that do not show the proximal ring (Online Resource 6, panel f). Very weak spots of expression are detected in the ventral neuroectoderm at approximately 140 h AED (Online Resource 6, panel g, arrows).
Analysis of Sp6–9 function using RNAi in P. tepidariorum
We have used parental RNAi to study the function of Sp6–9 in P. tepidariorum. The most prominent feature of the resulting phenotypes included a strong malformation of the appendages. In addition, there was a second class of phenotype that lacked a number of segments. We have categorized these two phenotypes into a strong category (limb defects and missing segments) and a weak category (only limb defects). In the strong category, the most severely affected animals showed a small number of knob-like limbs (only three or four pairs of limbs), a strong malformation of the head lobes, and a normally developed opisthosoma (Fig. 5d–f; compare with wild type in Fig. 5a–c). Animals with this phenotype died as embryos before dorsal closure, because this phenotype was only observed in embryos up to inversion stages and never in older embryos, larvae, or nymphs. Less severely affected animals from the strong category showed similar prosomal limb bud deformations but did not show any defects in head lobe development (Fig. 5g–i). These animals survived until late embryogenesis (Fig. 6c, d) but were not able to hatch, because this phenotype has never been observed in larvae or nymphs. In embryos shortly before hatching, this phenotypes showed normally fused head lobes as well as a normally developed opisthosoma, but malformations in the prosomal segments with shortened chelicerae and pedipalps and only two leg-bearing segments with strongly shortened walking legs (Fig. 6c, d).
Sp6–9 RNAi embryos of P. tepidariorum. a–c Wild-type embryo during inversion viewed from lateral (a), frontal (b), and from the rear end (c). d–f A strongly affected embryo from the category “strong phenotype” during inversion viewed from lateral (d), frontal (e), and from the rear end (f). g–i) A less strongly affected embryo from the category “strong phenotype” during inversion viewed from lateral-ventral (g), frontal (h), and from the rear end (i). The arrows in d and g point to limb rudiments. The arrowheads in c, f, and i point to the posterior end of the opisthosoma. In a, d, and g, ventral is down and anterior is to the left. In b, e, and h, the embryo is shown in frontal view, facing the viewer with the (remnants of) the head lobes. In c, f, and i, the embryo is shown in ventral aspect, and the view is mainly the opisthosoma (i.e., anterior is to the left). L, limb rudiments; L1–L4, walking legs 1 to 4; lr, labrum; hl, head lobe; O4–O5, opisthosomal segments 4 to 5; pp, pedipalp
Sp6–9 RNAi embryos of P. tepidariorum. a, b Wild-type embryo after dorsal closure viewed from ventral (a) and from lateral (b). c, d A less strongly affected embryo from the category “strong phenotype” after dorsal closure viewed from ventral (c) and from lateral (d). e, f A strongly affected embryo from the category “weak phenotype” after dorsal closure viewed from ventral (e) and from lateral (f). as, anterior spinnerets; ch, chelicera; en, endite of the pedipalp; L, leg rudiments; L1–L4; walking legs 1 to 4; pp, pedipalp; ps, posterior spinnerets
In the weak category, the most severely affected animals had strongly truncated and partially thickened appendages but were otherwise normal and did not lack any body segments (Fig. 6e, f). Animals of this category with all appendages strongly shortened were not able to hatch, because this phenotype has never been observed in larvae or nymphs. However, more weakly affected animals of this category were observed at the larval and nymphal stages. Larval phenotypes included mosaic animals that had some normally developed legs, but other legs were affected to different degrees (Fig. 7). Leg defects in these animals ranged from strongly shortened legs (Fig. 7d), via truncated legs with blunt tips indicating the lack of distal tissue (Fig. 7b), to larvae where the tarsal tissue is still present but does not develop normally and thus results in aberrantly shaped tips (Fig. 7b, d). Nymphal phenotypes were even milder (Fig. 8) and included only mosaic animals with a few thickened leg segments in some legs (Fig. 8b, c).
Cuticle preparations of larvae after Sp6–9 RNAi. a Wild-type larva of P. tepidariorum. b–d Larvae of P. tepidariorum after Sp6–9 RNAi. Arrows point to the defects in tarsal morphology; arrowheads point to missing distal tissue (blunt leg tips); the asterisk marks strongly shortened leg. L1–L4, walking legs 1 to 4; pp, pedipalp
Cuticle preparations of P. tepidariorum nymphs after Sp6–9 RNAi. The opisthosoma has been removed in the preparations before bleaching. a Wild-type nymph (first instar). b, c First nymphal instars after Sp6–9 RNAi. Arrows point to defects in leg morphology. ch, chelicera; L1–L4, walking legs 1 to 4; pp, pedipalp
Gene expression in Sp6–9 RNAi animals
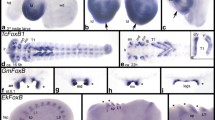
Given the dynamic expression pattern of Sp6–9 in multiple domains in the embryo and the complex phenotype affecting the head lobes, body segments, and appendages, we have assessed the effect of Sp6–9 RNAi on the expression of selected genes known to be important for head specification, body segmentation, and appendage formation. In wild-type embryos, the gene orthodenticle-1 (otd-1) is expressed along the outer circumference of the germ disc (Fig. 9a, arrowhead) and is crucial for the development of the head (Pechmann et al. 2009). In Sp6–9 RNAi animals, the expression of otd-1 is strongly affected, but not entirely absent. The expression of otd-1 is patchy and weaker than in wild-type animals (Fig. 9b–d, arrows). Also, at later developmental stages, the expression of otd-1 is strongly affected in Sp6–9 RNAi embryos. While there is a large patch of otd-1 in the head lobes in the wild type (Fig. 10a, b), this expression patch is strongly reduced in Sp6–9 embryos (Fig. 10c, d). The genes Distal-less (Dll) and dachshund (dac) are well-known for their role in appendage patterning in arthropods (reviewed in Angelini and Kaufman (2005)). In the wild type, Dll is expressed in the distal portion of all prosomal appendages and additional expression is present in the head lobes (Fig. 11a). In Sp6–9 RNAi embryos of the strong category, no expression of Dll mRNA can be detected by whole-mount in situ hybridization (Fig. 11b). The wild-type expression of dac includes a strong medial domain in the pedipalps and legs and additional expression in the nervous system (head, ventral nerve cord), heart, and the posterior end of the germ band (Fig. 11c). In strongly affected Sp6–9 RNAi embryos, the dac expression pattern is similar to wild-type embryos, except that the medial ring in the appendages is missing, and expression is only detected in the region from the anterior end of the germ band to the pedipalpal segment and the region from the third walking-leg segment to the posterior end of the germ band (Fig. 11d). Thus, there is a conspicuous gap of gene expression between anterior and posterior regions of the embryo. A comparable gap is observed in the expression pattern of the gene hedgehog (hh), a gene known for its evolutionarily conserved role in the development of the body segments (e.g., Farzana and Brown 2008; Kanayama et al. 2011; Pechmann et al. 2009; Akiyama-Oda and Oda 2010). In the wild type, hh is expressed in the posterior portion of every body segment (Fig. 11e). This is also the case in strongly affected Sp6–9 animals, except for the region between the pedipalpal segment and the third walking-leg segment, where expression of hh is absent (Fig. 11f).
The expression of otd-1 is affected after Sp6–9 RNAi in P. tepidariorum. a Wild-type embryo at the germ disc stage (stage 5). Expression of otd-1 is detected in a ring along the outer edge of the germ disc (arrowhead). b–d Stage 5 embryos after Sp6–9 RNAi. Expression of otd-1 is affected: it is located along the rim of the germ disc like in the wild type, but expression is discontinuous. Arrows point to patches of otd-1 expression
Expression of otd-1 is affected in germ band stages after Sp6–9 RNAi. a, b Wild-type embryo. c, d Embryo after Sp6–9 RNAi. In b, d, fluorescent and white light microscopy has been combined to visualize the morphology of the germ band. All embryos are at stage 10 and are shown in frontal view. The arrows point to otd-1 expression in the head lobes. ch, chelicera; pp, pedipalp; L1, L2, walking legs 1 and 2
Expression of limb and segment marker genes in P. tepidariorum embryos after Sp6–9 RNAi. a Expression of Dll in a wild-type embryo. b Expression of Dll in an embryo after Sp6–9 RNAi. c Expression of dac1 in a wild-type embryo. d Expression of dac1 in an embryo after Sp6–9 RNAi. The arrowheads denote the area of lacking gene expression. e Expression of hh in a wild-type embryo. f Expression of hh in an embryo after Sp6–9 RNAi. The arrows in e, f point to the head stripe of the hh expression pattern. ch, chelicera; pp, pedipalp; L1–L4, walking legs 1 to 4; O1–O3, opisthosomal segments 1 to 3
Discussion
The spectrum of phenotypes observed after parental RNAi corresponds well with the dynamic expression profile of Sp6–9. The early ring of expression in the germ disc (see Fig. 2) is similar to the expression of the previously described genes hunchback (hb), otd-1, hh, and odd-paired (opa) (Pechmann et al. 2009; Kanayama et al. 2011; Schwager et al. 2009). Similar to Sp6–9, these four genes are also expressed in a ring that is first located at the rim of the germ disc but is then relocated away from the rim towards the disc center. All four genes are required for head development and lead to strong head malformations after RNAi. Indeed, it has been suggested previously that this traveling gene expression from the rim towards the center is crucial for the correct patterning of the anterior head in spiders (Pechmann et al. 2009; Kanayama et al. 2011). The head lobe malformations observed in severely affected animals of the strong category of Sp6–9 RNAi (see Fig. 5d–f) support a similar role for the early ring of Sp6–9 expression. Furthermore, our results suggest that Sp6–9 might even be partially involved in the activation of one of these previously identified head genes, otd-1, because in Sp6–9 animals, otd-1 expression is disturbed (see Figs. 9 and 10).
In addition to the head defects, the previously identified genes with a dynamic expression ring in the germ disc also show a gap-like phenotype, i.e., a number of missing body segments. For example, the RNAi phenotype of hb includes a gap-like phenotype lacking the first and second walking-leg segments (Schwager et al. 2009), and a similar phenotype has been described for Dll (Pechmann et al. 2011). Intriguingly, the gap-like phenotype of Sp6–9 RNAi animals in the strong category also includes two missing leg-bearing segments (see Fig. 6d), and our results with hh staining in these embryos strongly suggest that the two missing segments are indeed the first and second walking-leg segments, too (see Fig. 11f). In summary, the similar expression dynamics and RNAi phenotypes of Sp6–9 and genes like otd-1, hb, or Dll suggest that they might all belong to the same gene network that governs the development of the head and anterior body segments.
Apart from the malformed head and the lacking body segments, the deformation of the appendages is the third prominent aspect of the Sp6–9 RNAi phenotypes. The leg defects are similar in embryos with and without the gap-like phenotype (compare Fig. 6d with Fig. 6f), indicating that these two roles of Sp6–9 are independently regulated. Indeed, the expression of Sp6–9 in the appendages appears later and independently from the early expression ring. We therefore assume that the strong-category phenotypes result when the RNAi effect sets in early and thus affects both the early expression ring in the germ disc and also the later expression in the legs. The weak-category phenotypes then result if the RNAi effect sets in later (or weaker) and therefore leaves the early expression unaffected and just affects the expression in the appendages. These limb defects indicate that Sp6–9 expression in the limbs is required for the growth of all prosomal appendages. The lack of dac1 and Dll expressions in the appendages (see Fig. 10b, d) also suggests that normal proximal-distal pattern formation in the limbs is disrupted.
From an evolutionary perspective, the role in appendage development is highly conserved in the arthropods. The D. melanogaster ortholog of Sp6–9, called D-Sp1, has a very strong leg phenotype that includes severe leg shortening and bloated leg segments (Cordoba et al. 2016) that is very similar to the leg phenotypes observed in Sp6–9 RNAi animals in the present study. Strongly shortened legs after the loss of Sp6–9 function were also reported in the beetle T. castaneum (Beermann et al. 2004) and the milkweed bug O. fasciatus (Schaeper et al. 2009). A role in head formation or segment formation, however, has not been described for an arthropod Sp6–9 ortholog so far. Interestingly, however, a crucial role in head segmentation has been described for members of the Sp5/Btd gene family (e.g., the D. melanogaster gene buttonhead (btd)) (Cohen and Jürgens 1990; Wimmer et al. 1993), suggesting that the spider Sp6–9 ortholog might (partially) combine the function of Sp5 and Sp6–9 genes in other arthropods.
References
Akiyama-Oda Y, Oda H (2003) Early patterning of the spider embryo: a cluster of mesenchymal cells at the cumulus produces Dpp signals received by germ disc epithelial cells. Development 130:1735–1747
Akiyama-Oda Y, Oda H (2010) Cell migration that orients the dorsoventral axis is coordinated with anteroposterior patterning mediated by Hedgehog signaling in the early spider embryo. Development 137:1263–1273
Angelini DR, Kaufman TC (2005) Insect appendages and comparative ontogenetics. Dev Biol 286:57–77
Beermann A, Aranda M, Schröder R (2004) The Sp8 zinc-finger transcription factor is involved in allometric growth of the limbs in the beetle Tribolium castaneum. Development 131:733–742
Chung HA, Medina-Ruiz S, Harland RM (2014) Sp8 regulates inner ear development. Proc Natl Acad Sci U S A 111:6329–6334
Cohen SM, Jürgens G (1990) Mediation of Drosophila head development by gap-like segmentation genes. Nature 346:482–485
Cordoba S, Requena D, Jory A, Saiz A, Estella C (2016) The evolutionarily conserved transcription factor Sp1 controls appendage growth through Notch signaling. Development 143:3623–3631
Farzana L, Brown SJ (2008) Hedgehog signaling pathway function conserved in Tribolium segmentation. Dev Genes Evol 218:181–192
Foelix R (2011) The biology of spiders, 3rd edn. Oxford University Press, Oxford, pp 270–272
Haro E, Delgado I, Junco M, Yamada Y, Mansouri A, Oberg KC, Ros MA (2014) Sp6 and Sp8 transcription factors control AER formation and dorsal-ventral patterning in limb development. PLoS Genet 10:e1004468
Hertveldt V, Louryan S, van Reeth T, Drèze P, van Vooren P, Szpirer J, Szpirer C (2008) The development of several organs and appendages is impaired in mice lacking Sp6. Dev Dyn 237:883–892
Kanayama M, Akiyama-Oda Y, Nishimura O, Tarui H, Agata K, Oda H (2011) Travelling and splitting of a wave of hedgehog expression involved in spider-head segmentation. Nat Commun 2:500
Kawakami Y, Esteban CR, Matsui T, Rodríguez-León J, Kato S, Izpisúa Belmonte JC (2004) Sp8 and Sp9, two closely related buttonhead-like transcription factors, regulate Fgf8 expression and limb outgrowth in vertebrate embryos. Development 131:4763–4774
Koga T, Matsui Y, Asagiri M, Kodama T, de Crombrugghe B, Nakashima K, Takayanagi H (2005) NFAT and Osterix cooperatively regulate bone formation. Nat Med 11:880–885
Li X, Sun C, Lin C, Ma T, Madhavan MC, Campbell K, Yang Z (2011) The transcription factor Sp8 is required for the production of parvalbumin-expressing interneurons in the olfactory bulb. J Neurosci 31:8450–8455
Mittmann B, Wolff C (2012) Embryonic development and staging of the cobweb spider Parasteatoda tepidariorum C. L. Koch, 1841 (syn.: Achaearanea tepidariorum; Araneomorphae; Theridiidae). Dev Genes Evol 222:189–216
Oda H, Nishimura O, Hirao Y, Tarui H, Agata K, Akiyama-Oda Y (2007) Progressive activation of Delta-Notch signaling from around the blastopore is required to set up a functional caudal lobe in the spider Achaearanea tepidariorum. Development 134:2195–2205
Pechmann M, McGregor AP, Schwager EE, Feitosa NM, Damen WG (2009) Dynamic gene expression is required for anterior regionalization in a spider. Proc Natl Acad Sci U S A 106:1468–1472
Pechmann M, Prpic NM (2009) Appendage patterning in the South American bird spider Acanthoscurria geniculata (Araneae: Mygalomorphae). Dev Genes Evol 219:189–198
Pechmann M, Khadjeh S, Turetzek N, McGregor AP, Damen WG, Prpic NM (2011) Novel function of Distal-less as a gap gene during spider segmentation. PLoS Genet 7:e1002342
Philipsen S, Suske G (1999) A tale of three fingers: the family of mammalian Sp/XKLF transcription factors. Nucleic Acids Res 27:2991–3000
Posnien N, Zeng V, Schwager EE, Pechmann M, Hilbrant M, Keefe JD, Damen WGM, Prpic NM, McGregor A, Extavour CG (2014) A comprehensive reference transcriptome resource for the common house spider Parasteatoda tepidariorum. PLoS One 9:e104885
Prpic NM, Schoppmeier M, Damen WGM (2008a) Collection and fixation of spider embryos. Cold Spring Harb Protoc 3:930–932. https://doi.org/10.1101/pdb.prot5067
Prpic NM, Schoppmeier M, Damen WGM (2008b) Whole-mount in situ hybridization of spider embryos. Cold Spring Harb Protoc 3:933–936. https://doi.org/10.1101/pdb.prot5068
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Schaeper ND, Prpic NM, Wimmer EA (2009) A conserved function of the zinc finger transcription factor Sp8/9 in allometric appendage growth in the milkweed bug Oncopeltus fasciatus. Dev Genes Evol 219:427–435
Schaeper ND, Prpic NM, Wimmer EA (2010) A clustered set of three Sp-family genes is ancestral in the Metazoa: evidence from sequence analysis, protein domain structure, developmental expression patterns and chromosomal location. BMC Evol Biol 10:88
Schöck F, Purnell BA, Wimmer EA, Jäckle H (1999) Common and diverged functions of the Drosophila gene pair D-Sp1 and buttonhead. Mech Dev 89:125–132
Schwager EE, Pechmann M, Feitosa NM, McGregor AP, Damen WG (2009) hunchback functions as a segmentation gene in the spider Achaearanea tepidariorum. Curr Biol 19:1333–1340
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins DG (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539
Suske G, Bruford E, Philipsen S (2005) Mammalian SP/KLF transcription factors: bring in the family. Genomics 85:551–556
Turetzek N, Pechmann M, Schomburg C, Schneider J, Prpic NM (2016) Neofunctionalization of a duplicate dachshund gene underlies the evolution of a novel leg segment in arachnids. Mol Biol Evol 33:109–121
Turetzek N, Prpic NM (2016) Observations on germ band development in the cellar spider Pholcus phalangioides. Dev Genes Evol 226:413–422
Wimmer EA, Jäckle H, Pfeifle C, Cohen SM (1993) A Drosophila homologue of human Sp1 is a head-specific segmentation gene. Nature 366:690–694
Acknowledgements
We thank Beate Preitz for the help with microscopy. We also thank all members of the department for comments that helped to improve the manuscript.
Funding
This work was supported by the Deutsche Forschungsgemeinschaft (grant numbers PR 1109/4-1 and PR 1109/6-1 to N.M.P. and PE 2075/1-1 and PE 2075/1-2 to M.P.). Additional financial backing has been received from the Göttingen Graduate School for Neurosciences, Biophysics and Molecular Biosciences (GGNB), the Göttingen Center for Molecular Biosciences (GZMB), and the University of Göttingen (GAU). N.T. has been supported by a Christiane-Nüsslein-Volhard-Foundation fellowship and a “Women in Science” award by L’Oréal Deutschland and the Deutsche UNESCO-Kommission. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
N.M.P. is a member of the editorial board of Development Genes and Evolution serving as communicating editor for the topical collection “Size & Shape.”
Additional information
Communicated by Volker Hartenstein
Electronic supplementary material
Online Resource 1
(TXT 1 kb)
Online Resource 2
(TXT 4 kb)
Online Resource 3
(NEX 7 kb)
Online Resource 4
(JPEG 329 kb)
Online Resource 5
(JPEG 281 kb)
Online Resource 6
(JPEG 638 kb)
Rights and permissions
About this article
Cite this article
Königsmann, T., Turetzek, N., Pechmann, M. et al. Expression and function of the zinc finger transcription factor Sp6–9 in the spider Parasteatoda tepidariorum . Dev Genes Evol 227, 389–400 (2017). https://doi.org/10.1007/s00427-017-0595-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-017-0595-2