Abstract
Main conclusion
This review focuses on HATs and HDACs that modify non-histone proteins, summarizes functional mechanisms of non-histone acetylation as well as the roles of HATs and HDACs in rice and Arabidopsis.
Abstract
The growth and development of plants, as well as their responses to biotic and abiotic stresses, are governed by intricate gene and protein regulatory networks, in which epigenetic modifying enzymes play a crucial role. Histone lysine acetylation levels, modulated by histone acetyltransferases (HATs) and histone deacetylases (HDACs), are well-studied in the realm of transcriptional regulation. However, the advent of advanced proteomics has unveiled that non-histone proteins also undergo acetylation, with its underlying mechanisms now being clarified. Indeed, non-histone acetylation influences protein functionality through diverse pathways, such as modulating protein stability, adjusting enzymatic activity, steering subcellular localization, influencing interactions with other post-translational modifications, and managing protein–protein and protein–DNA interactions. This review delves into the recent insights into the functional mechanisms of non-histone acetylation in plants. We also provide a summary of the roles of HATs and HDACs in rice and Arabidopsis, and explore their potential involvement in the regulation of non-histone proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Post-translational modifications (PTMs) are a diverse array of covalent alterations to a protein’s structure that take place after its synthesis by the ribosome. These modifications are integral to the regulation of protein function and are crucial for a myriad of cellular processes (Verdin and Ott 2015). In recent years, the term PTMs has become more synonymous with the covalent addition of small molecules to proteins, particularly the enzymatic attachment of groups such as acetyl, methyl, or phosphate groups to amino acid side chains (Suskiewicz 2024). This specificity allows PTMs to modulate protein activity with a precision that surpasses other regulatory mechanisms. The initial discovery in the realm of PTMs was the identification of reversible protein phosphorylation, which serves as a key regulatory mechanism for enzyme function in humans (Sutherland and Wosilait 1955). Since then, a spectrum of PTMs, including ubiquitination, acetylation, and methylation, has been unveiled (Gough and Sadanandom 2021). Among these, lysine acetylation (LysAc) stands out as a highly conserved and reversible PTM across both prokaryotic and eukaryotic organisms, affecting not only histones but also a wide array of non-histone proteins (Zhang et al. 2009; Rao et al. 2014; Kumar et al. 2021).
Acetyl-coenzyme A (acetyl-CoA), a key molecule in cellular metabolism, serves a dual role as both the acetyl donor for protein acetylation and a precursor for the biosynthesis of various phytochemicals (Chen et al. 2017). Protein LysAc involves the transfer of an acetyl group from acetyl-CoA to the ε-amino position of a lysine residue within a protein. This process neutralizes the positive charge of the lysine side chain (Ali et al. 2018). LysAc is catalyzed by lysine acetyltransferases (KATs, also named as HATs), which are the ‘writers’ that transfer acetyl groups from acetyl-CoA to lysine residues, while lysine deacetylases (KDACs, also named as HDACs) remove these groups, thus acting as the ‘erasers’ of the epigenetic code (Choudhary et al. 2014; Narita et al. 2019; Tyagi et al. 2021). The interplay between these ‘writers’ and ‘erasers’ is meticulously balanced, as evidenced by the genome-wide identification of HAT and HDAC genes in plants such as Arabidopsis and rice (Hou et al. 2021; Chen et al. 2020; Pandey et al. 2002; Liu et al. 2012). These enzymes through interacting with histone octamer contribute to the dynamic regulation of histone acetylation, which in turn influences chromatin structure and gene expression.
In addition to histones, the mammalian p53 protein was the first non-histone protein recognized for its ability to undergo acetylation and deacetylation, highlighting the broad impact of these modifications on cellular function (Gu and Roeder 1997). By 2019, acetylation had been identified on over a hundred types of proteins in mammalian cells, including enzymes, transcription factors, and nuclear receptors (Narita et al. 2019). In plants, the discovery of Lys-acetylated sites on 74 proteins across different Arabidopsis organs in 2011 marked a significant step forward in understanding the role of LysAc in non-histone proteins (Finkemeier et al. 2011). With the advent of acetylated proteomics, a wealth of non-histone lysine acetylation modifications in plants has been revealed, implicating their involvement in diverse biological processes such as catalysis and binding (Fang et al. 2015).
While much attention has been given to the acetylation of histone proteins and its epigenetic implications for plant phenotypes, the non-histone acetylation/deacetylation landscape in plants remains less explored. This review aims to bridge this gap by providing a comprehensive overview of the role of HDACs and HATs in the post-translational regulation of non-histone proteins. We broaden the scope of research into post-translational modifications and explore the biochemical outcomes of these processes, including the influence on protein stability, changes in enzymatic activity, adjustments in cellular localization, and the modulation of interactions between protein–protein and protein–DNA. These processes are intricately linked to various signaling pathways and metabolic processes, ultimately shaping plant growth, development, and stress responses.
Functional mechanisms of non-histone acetylation modifications
Regulation of enzyme activity
Non-histone acetylation modification has been identified as a regulator of enzyme activity. In Arabidopsis, it has been discovered that histone deacetylase 6 (HDA6) can bind to the negative regulatory factor BRASSINOSTEROID INSENSITIVE 2 (BIN2), which plays a significant role in plant development, and deacetylate the lysine 189 (K189) site, resulting in a reduction of its kinase activity (Hao et al. 2016). Similarly, in rice, OsHDAC1 deactivates OsGSK2, thereby preventing the degradation of the downstream transcriptional factor OsBZR1 and controlling lateral root formation (Hou et al. 2022). However, deacetylation can also activate the enzyme activity of target proteins. For instance, empirical evidence has identified that the cytoplasmic histone deacetylase FolSir2 in tomato can bind to the kinase FolGSK3 and deacetylate the lysine 271 (K271) site, thereby activating FolGSK3-kinase activity and promoting pathogenic fungal infection (Zhang et al. 2023). Many metabolic enzymes have also been shown to be regulated by acetylation/deacetylation. For example, in Arabidopsis, lysine deacetylation significantly increases the activity of the Rubisco large subunit as well as other central metabolic enzymes, such as the Calvin cycle enzyme phosphoglycerate kinase, the glycolytic enzyme glyceraldehyde 3-phosphate dehydrogenase, and the tricarboxylic acid (TCA) cycle enzyme malate dehydrogenase (Finkemeier et al. 2011). Plant mitochondria are crucial for plant growth and development. All of the TCA cycle enzymes in mitochondria have been identified to be lysine-acetylated, including most components of the pyruvate dehydrogenase complex (König et al. 2014). In Pisum sativum mitochondria, a minimum of 358 acetyl-proteins have been identified in pea seedlings, with 93 of these proteins found to be acetylated, including NDP-kinase, AAA + , LMW-Hsp, cytochrome b5 reductase, and formate dehydrogenase (Smith-Hammond et al. 2014). These findings indicate that histone deacetylases can act as dual regulators, capable of both activating and inhibiting the activity of target proteins, and play a pivotal role in signaling and metabolism pathways (Fig. 1A).
Functional mechanisms of non-histone protein acetylation. A Regulation of enzyme activity. Acetylation acts as a regulator within signaling pathways, influencing the catalytic activities of numerous metabolic enzymes. B Regulation of protein degradation. In plants, acetylation-dependent proteins can prevent protein ubiquitination, thereby influencing degradation process of proteins. C Regulation of protein synthesis. The enzyme OsHDA714 can remove acetyl groups from ribosomal proteins, thereby facilitating the translation process. D Regulation of protein–protein interactions. The acetylation of certain non-histone proteins can modulate their interactions with other proteins. E Regulation of protein–DNA interactions. Deacetylation by HDACs on non-histone proteins can diminish their DNA binding capacity. F Regulation of subcellular localization. Deacetylation or acetylation of non-histones may induce the translocation of these proteins within the cell. G Crosstalk with other PTMs. Acetylation can interact with other PTMs, such as succinylation and phosphorylation, indicating a complex regulatory network
Regulation of protein degradation and synthesis
In mammalian cells, it has been observed that LysAc plays a dual role in regulating protein degradation, affecting both proteasome-dependent and -independent pathways (Narita et al. 2019). In plants, acetylation appears to prevent ubiquitination of proteins, thus inhibiting proteasome-mediated degradation (Fig. 1B). For instance, in rice, the histone deacetylase OsHDA706 has been shown to reduce the ubiquitination level of the key enzyme OsLOX14 in the jasmonic acid (JA) biosynthesis pathway by deacetylating specific lysine residues (K204, K216, K310, and K317). This action prevents OsLOX14 from being degraded by the 26S proteasome, enhancing its stability and contributing to the accumulation of JA, which in turn improves the plant’s broad-spectrum antiviral defense. However, the precise mechanism by which OsHDA706 deacetylates OsLOX14 to reduce its ubiquitination level remains to be elucidated (Yang et al. 2024). Similarly, another study in rice has identified that the histone deacetylase OsHDA716 can bind to and reduce the acetylation level at specific lysine residues (K263, K281, and K294) of the transcription factor OsbZIP46. This reduction in acetylation promotes the ubiquitination and subsequent degradation of OsbZIP46 by the 26S proteasome complex, which negatively impacts the cold tolerance of rice (Sun et al. 2024).
In Arabidopsis, AtMBP-1, a transcriptional repressor involved in stress tolerance and glycolytic gene expression, is subject to LysAc, making it susceptible to proteasome degradation. However, the deacetylase AtSRT1 can remove this acetylation, significantly enhancing AtMBP-1’s stability in vivo (Liu et al. 2017). Additionally, OsHDA714 has been found to interact with ribosomal proteins, reducing their acetylation levels and thus preventing modifications by ubiquitination complexes. This action increases the abundance of ribosomal proteins and facilitates the translation process, thereby controlling protein homeostasis and coordinating stress responses, metabolism, and ribosome function (Xu et al. 2021) (Fig. 1C). These findings indicate that the modulation of acetylation levels on different proteins can either be beneficial or detrimental to ubiquitination, impacting protein stability and potentially influencing protein synthesis.
Regulation of protein–protein and protein–DNA interactions
Acetylation of non-histone proteins plays a crucial role in modulating their interactions with other protein factors (Fig. 1D). In Arabidopsis, the NINJA-TPL complex serves as a pivotal negative regulator within JA signaling pathways. HDA6-mediated acetylation of TPL diminishes its interaction with the NINJA protein, whereas acetylation by GCN5 counteracts this effect, restoring TPL’s binding capacity. This dynamic regulation is essential for orchestrating a genome-wide transcriptional program that underpins plant immunity and adaptive growth (An et al. 2022). In rice, the interaction between OsGSK2 and OsBZR1 leads to an increase in OsBZR1’s phosphorylation levels within the nucleus, promoting its subsequent ubiquitination and degradation. Conversely, deacetylation of OsGSK2 by OsHDAC1 precludes its binding to OsBZR1 (Hou et al. 2022). This finding uncovers a dual mechanism by which OsHDAC1 not only suppresses enzymatic activity but also modulates the binding capabilities of this protein. Similarly, protein acetylation can influence protein–DNA interactions (Fig. 1E). In Arabidopsis has demonstrated that the deacetylase HDA9 interacts with the transcription factor WRKY53, deacetylating it and thereby inhibiting its binding to the W-Box element. This results in a reduced trans-activation function, affecting the expression of downstream target genes such as WRKY13, WRKY15, WRKY18, WRKY22, WRKY29, and WRKY62, and also activating WRKY6 and WRKY42, which in turn repress Arabidopsis’s tolerance to salt stress (Zheng et al. 2020). Furthermore, the deacetylation of OsbZIP46, which is involved in the regulation of cold tolerance, is controlled by OsHDA716. This modification not only mediates the ubiquitination and subsequent degradation of OsbZIP46 but also impacts its DNA binding capacity due to the acetylation site located within the DNA binding domain. Consequently, this leads to a decrease in its transcriptional activity (Sun et al. 2024).
Regulation of subcellular localization
Numerous studies of plant proteins have shown that that post-translational modifications, such as phosphorylation and ubiquitination, can significantly influence the intracellular distribution of proteins. These modifications can enhance nuclear and cytoplasmic localization, facilitate the release of plasma membrane proteins into the cell interior, and regulate the export of cytoplasmic proteins (Zhang and Zeng 2020). Recent studies have highlighted the impact of acetylation on protein localization within plant cells. For instance, the histone deacetyltransferase FolSir2 mediates the deacetylation of FolSrpk1, reducing its acetylation level and causing its dissociation from the cytoplasmic FolAha1 protein. This process leads to the nuclear translocation of FolSrpk1, uncovering a conserved mechanism where FolSir2-mediated cytoplasmic protein deacetylation plays a role in plant-fungal pathogenicity (Zhang et al. 2023, 2024). Additionally, the protein FolSvp1 directly binds to and translocates the tomato pathogenesis-related protein SlPR1 from the apoplast, the space outside the plasma membrane, to the host nucleus. This translocation is facilitated by the nuclear localization signal present in FolSvp1. The activity of FolSvp1 is contingent upon covalent acetylation modification, which is essential for its full virulence in host tomato tissues. This acetylation is catalyzed by the lysine acetyltransferase FolArd1 prior to the secretion of FolSvp1 (Li et al. 2022). These findings suggest that acetylation not only modulates the subcellular localization of proteins but also has a profound influence on the disease resistance mechanisms in plants (Fig. 1F and Table 1).
Crosstalk between acetylation and other PTMs
Non-histone proteins are often subjected to multiple PTMs, which can interact in a process known as PTM crosstalk (Soufi et al. 2012). The interaction between LysAc and other PTMs facilitates the integration of various biological signals to regulate life processes under diverse conditions. In this context, the focus is on the crosstalk between LysAc and two other PTMs: succinylation and phosphorylation (Fig. 1G).
Acetyl-CoA and succinyl-CoA, key metabolites in cellular energy production, are associated with enzymes that are modified by both LysAc and succinylation, indicating a functional link between these two PTMs (He et al. 2016). In rice, 699 acetylated sites across 389 proteins and 665 succinylated sites across 261 proteins were identified, with 133 sites on 78 proteins being dually modified. These modifications are crucial for seed germination and are predominantly found in ribosomal and proteasome proteins, hinting at roles in protein synthesis, repair, and degradation post-infection (Zhou et al. 2018).
Phosphorylation is another PTM that interacts with LysAc. In Arabidopsis, the diurnal changes in acetylated and phosphorylated proteins regulate circadian rhythms and light responses, with acetylation mainly affecting metabolic and stress response proteins, and phosphorylation impacting RNA processing and cell cycle proteins (Uhrig et al. 2017, 2019). A total of 909 acetylated and 2549 phosphorylated proteins were identified, with 134 proteins involved in core cellular processes showing dual modifications, suggesting a synergistic control over these processes (Uhrig et al. 2019). In wheat, both LysAc and phosphorylation are present in two key enzymes including sucrose synthase (SuSy) and ADP glucose pyrophosphorylase (AGPase) for starch biosynthesis, indicating a role in yield formation under drought stress (Zhu et al. 2018). In rice anthers, 189 proteins with dual lysine acetylation and phosphorylation modifications were identified, with 103 preferentially expressed in meiocytes, highlighting the regulation of the diurnal cycle, stress responses, and pollen development (Li et al. 2018).
To sum up, the crosstalk between LysAc and succinylation or phosphorylation, is widespread in plants and is integral to metabolic processes, seed development, stress responses, and the diurnal cycle (Xia et al. 2021). These interactions likely play a coordinated role in the regulation of the plant life cycle.
Regulation of plant phenotypes and biological functions by HATs and HDACs
Extensive research has shown HATs and HDACs are involved in various plant developmental processes, including root growth, leaf development, flowering time regulation, and seed germination by regulating the transcription of target genes through the acetylation or deacetylation of histones. In addition to histones, lysine acetylation and deacetylation also play roles in the function of certain non-histone proteins in plants, as previously discussed. This section further elaborates on the functions of HATs and HDACs and explores their potential in modifying non-histone proteins (Supplementary Table 1).
Class I type-HDACs
In Arabidopsis, there are six Class I type-HDACs, including HDA6, HDA9, HDA19, HDA7, HDA17, and HDA10. Among these, HDA6, HDA19, and HDA9 have been extensively characterized and studied. For instance, HDA6 has been reported to regulate leaf development, flowering, and salt stress response by repressing KNOX, FLC, FT, and ABIs, respectively (Yu et al. 2011, 2017; Luo et al. 2012a, b). Interestingly, HDA6 also targets the non-histone protein BIN2, which is involved in flowering and salt stress response (Kim et al. 2023; Ju et al. 2023; Zhu et al. 2023), suggesting that HDA6 can control these traits through BIN2 as well. Similarly, HDA9 promotes leaf development and flowering, enhances salt tolerance, and weakens heat tolerance by repressing GI, HsfA2, and WRKY33, respectively (Yu et al. 2011; Suzuki et al. 2018; Park et al. 2019; Mayer et al. 2019; Niu et al. 2022; Yang et al. 2023). Recently, another non-histone target, WRKY53, was reported to function in regulating leaf development (Andrade Galan et al. 2024; Wang et al. 2023a, b), suggesting another potential connection between WRKY53 and HDA9 in leaf development.
HDA19, another widely studied Arabidopsis histone deacetylase, is reported to be involved in root and flower development, seed maturation, dormancy, oxidative stress tolerance, and pathogen defense (Niu et al. 2019; Ning et al. 2019; Chen et al. 2019; Gao et al. 2015; Zhou et al. 2013; Wang et al. 2013; Krogan et al. 2012; Zheng et al. 2023). A large number of downstream deacetylation genes involved in these phenotypes have been identified. However, the role of HDA9 in non-histone proteins remains unclear.
In addition, HDA7 has been identified as being required for female gametophyte and embryo development (Cigliano et al. 2013). HDA6, HDA9, and HDA19 were reported to be localized in the nucleus (Alinsug et al. 2012), suggesting that Class I members of HDACs might jointly regulate phenotypes in the nucleus by modifying both histones and non-histones.
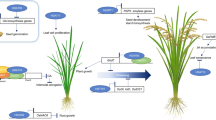
Unlike Arabidopsis Class I members, which are located in the nucleus, rice Class I members HDA701, HDA702, and HDA710 are found in both the nucleus and cytoplasm (Hou et al. 2024; Ullah et al. 2020). Functional studies have found that HDA701, HDA702, and HDA705 negatively regulate rice broad-spectrum disease resistance, and HDA710 negatively regulates salt tolerance by mediating histone modification levels (Chen et al. 2021, 2022; Hou et al. 2022; Zhao et al. 2016; Ullah et al. 2020). Currently, the non-histone target OsGSK2 of HDA702, which controls root growth, has been identified in both the nucleus and cytoplasm (Hou et al. 2022). OsGSK2 also controls disease resistance (He et al. 2020; Meng et al. 2024), suggesting that HDA702 might regulate rice disease resistance through the GSK2 pathway. However, the non-histone targets of HDA701, HDA705, and HDA710 in the cytoplasm remain unclear. Furthermore, HDA703 is located in the nucleus and has been found to regulate rice growth and heading date through repression of Ghd7 (Wang et al. 2023a, b, 2021). The functions of other Class I members, including HDA707 and HDA709, have not yet been revealed.
Class II-type HDACs
Class II members of HDACs including HDA5, HDA8, and HDA14 have been localized to the cytoplasm (Alinsug et al. 2012), while HDA15 and HDA18 exhibit localization in both the nucleus and cytoplasm (Alinsug et al. 2012; Liu et al. 2013), implying that Class II-HDACs may act as key regulators of non-histone protein modifications. In line with this hypothesis, HDA14 has been found to deacetylate tubulin (Tran et al. 2012). Similarly, HDA5 has been shown to deacetylate the stress-responsive transcription factor GT2L and the dehydration-related protein ERD7, thereby regulating the response to salt stress (Tilak et al. 2023). Furthermore, it has been observed that white light can facilitate the translocation of HDA15 from the cytoplasm into the nucleus, where it represses hypocotyl growth by modulating histone acetylation (Liu et al. 2013; Zhao et al. 2019). When located in the nucleus, HDA15 can repress leaf senescence, flowering, and enhance resistance to salt stress through the action of WRKYs, LOX2, LARP1C, and NCED3 (Huang et al. 2022; Truong et al. 2021; Lee and Seo 2019). HDA18 is known to influence Arabidopsis root development by modulating histone acetylation at four kinase genes (Liu et al. 2013). The non-histone targets of HDA8, HDA15, and HDA18 remain unidentified.
In rice, there are five Class II-type HDAC members, including HDA704, HDA712, HDA713, HDA714, and HDA716. HDA714 has been found in the cytoplasm, and HDA716, which is localized in the nucleus, has been shown to deacetylate RPS6 and bZIP46 (Sun et al. 2024). Additionally, HDA704 has been reported to enhance salt tolerance by reducing the acetylation level of ABI5 (Guo et al. 2023). The non-histone targets of HDA712 and HDA713, however, remain unclear.
Class III (SIR2) and class IV-type HDACs
In humans, previous reports have showed that seven SIR2 HDACs require NAD+ as a cofactor for their enzymatic activity, and these members were located in the nucleolus, cytosol, and mitochondria (Seto and Yoshida 2014). This multi-pattern localization reveals that they function in deacetylating numerous non-histone and histone proteins (Seto and Yoshida 2014). Unlike humans, only two SIR2-type HDACs have been identified in Arabidopsis and rice (Hou et al. 2021). Among them, Arabidopsis SRT1 and SRT2 can control the development of hypocotyls in the nucleus through histone modification pathways (Zhang et al. 2018). Similarly, rice SRT701 is a nuclear-localized protein (Chung et al. 2009), which can control rice seed development and leaf senescence through the histone modification pathway (Zhang et al. 2016). However, the non-histone targets within the nucleus of AtSRT1, AtSRT2, and OsSRT701 remain unknown.
Unlike AtSRT1, AtSRT2, and OsSRT701, a previous report showed that SRT702 is a chloroplast-localized protein (Chung et al. 2009), implying that it serves as the main regulator of non-histone modifications. However, recent studies have found that SRT702 is also localized in the nucleus and can negatively regulate rice broad-spectrum disease resistance through histone modification pathways (Chen et al. 2024), suggesting that SRT702 is functionally similar to human SIRTs. The role of SRT702 in non-histone protein deacetylation requires further identification.
HD2-type HDACs
Unlike mammalian cells and yeast, HD2-type HDACs are uniquely identified in plants. In Arabidopsis, there are four HD2 members: HD2A, HD2B, HD2C, and HD2D, which are predominantly located in the nucleolus (Zhou et al. 2004). Multiple studies have demonstrated that HD2A, HD2B, and HD2C regulate the balance between embryonic, seed, and root development, as well as drought stress, by controlling rRNA processing (Pontes et al. 2007; Kim et al. 2014; Chen et al. 2018; Luo et al. 2022; Tahir and Tian 2021; Han et al. 2023a, b). This suggests that HD2-type HDACs may play crucial roles in rRNA production. However, in contrast to Arabidopsis HD2-type HDACs, the rice HDT701 has been found to be involved in flowering and resistance to fungal and bacterial pathogens by modulating the expression of OsIDS1 or pathogenesis-related proteins (PRs) (Ding et al. 2012; Cho et al. 2018; Li et al. 2021).
GNAT, MYST, CBP and TAF250-type HATs
Although research on plant HATs has been less extensive compared to HDACs, recent studies have shed more light on their roles. For instance, the Arabidopsis AtGCN5, located in the nucleus, modulates JA signaling to mediate plant defense through reversible acetylation of TOPLESS (An et al. 2022). Additionally, AtGCN5 is implicated in regulating salicylic acid (SA) homeostasis by adjusting histone acetylation levels (Kim et al. 2020). Given that SA is a key player in bolstering plant disease resistance, AtGCN5 is likely to exert control over this resistance via both histone and non-histone protein pathways. Similarly, the rice GCN5 has been identified to influence crown root development by deacetylating non-histone ADA2 and histone H3 (Zhou et al. 2017; Yu et al. 2024). Furthermore, Arabidopsis HATs HAG3, HAC1, and HAF1 have been found to impact UV-B responses negatively by regulating the expression of DNA repair enzymes and sunscreen content (Fina and Casati 2015; Fina et al. 2017). HAC1 also promotes leaf senescence and modulates the expression of ERF022 (Hinckley et al. 2019). The specific functions of other HATs and their non-histone targets remain largely unknown. Apart from the rice GCN5, the roles of HATs are yet to be fully elucidated. However, subcellular localization studies (Liu et al. 2012), have indicated that OsHAC701, OsHAG703, and OsHAG704 are present in both the nucleus and cytosol, suggesting a potential role for these HATs in non-histone acetylation within the cytosol.
Future perspectives
The exploration of acetylation modifications in non-histone proteins has significantly broadened our comprehension of lysine acetylation. This modification is not confined to the histones within the nucleus; it also plays a more extensive role in the regulation of non-histone proteins across various metabolic pathways and functions within subcellular organelles such as chloroplasts and mitochondria, as well as in the cytoplasm. The acetylation of non-histone proteins modulates their functionality, which in turn influences plant growth, development, and stress responses.
While proteomic analyses have shed light on the subcellular distribution of acetylated proteins and identified numerous lysine acetylation sites through gene ontology (GO) term annotations (Xiong et al. 2016), the specific roles of various proteins acetylated by HATs are still largely unexplored. The mechanisms by which individual HATs and HDACs select their targets remain largely unknown. A key question is what guides the specificity of HATs or HDACs towards non-histone proteins. Answering these questions will likely necessitate a biochemical characterization of enzyme complexes and their subcellular localization. It is worth noting that, although HDACs are primarily nuclear, some, such as HDAC6 (Hubbert et al. 2002), have also been detected in the cytoplasm.
The interaction mechanisms of these PTMs remain unclear due to the limitations of current detection technologies. There is an urgent need to enhance the temporal and spatial resolution of mass spectrometry-based assays to enable high-throughput detection of multiple PTMs. Enzyme-catalyzed proximity labeling is an emerging method for studying the spatial and interaction characteristics of proteins within living cells. By expressing TurboID in cells, fused with target proteins of interest, the enzyme catalyzes the formation of biotin-5’-AMP from biotin, facilitated by ATP and biotin. This compound diffuses from the active site and binds to lysines of nearby endogenous proteins, leading to biotinylation of these proteins and thus facilitating the identification of neighboring proteins in physical space (Branon et al. 2018). This technique may prove valuable for large-scale identification of PTM crosstalk in non-histone proteins.
Analyzing molecular and genetic evidence of the interactions between multiple PTMs will yield new insights into the regulatory mechanisms governing plant growth, development, and adaptation to environmental changes. Elucidating the specific pathways of acetylation regulation in non-histone proteins and the crosstalk among PTMs opens up new avenues for genetic improvement and plant breeding. Techniques such as site-directed mutagenesis or CRISPR/Cas technology could be employed to modify lysine acetylation sites, altering enzyme activity, and potentially enhancing or reducing phenotypic plasticity. This approach could lead to the emergence of new plant variants, although the establishment of such modifications still requires further research and validation.
Data availability
No data was used for the research described in the article.
References
Ali I, Conrad RJ, Verdin E, Ott M (2018) Lysine acetylation goes global: from epigenetics to metabolism and therapeutics. Chem Rev 118(3):1216–1252. https://doi.org/10.1021/acs.chemrev.7b00181
Alinsug MV, Chen FF, Luo M, Tai R, Jiang L, Wu K (2012) Subcellular localization of class II HDAs in Arabidopsis thaliana: nucleocytoplasmic shuttling of HDA15 is driven by light. PLoS One 7(2):e30846. https://doi.org/10.1371/journal.pone.0030846
An C, Deng L, Zhai H, You Y, Wu F, Zhai Q, Goossens A, Li C (2022) Regulation of jasmonate signaling by reversible acetylation of TOPLESS in Arabidopsis. Mol Plant 15(8):1329–1346. https://doi.org/10.1016/j.molp.2022.06.014
Andrade Galan AG, Doll J, Faiß N, Weber P, Zentgraf U (2024) Complex formation between the transcription factor WRKY53 and antioxidative enzymes leads to reciprocal inhibition. Antioxidants (Basel) 13(3):315. https://doi.org/10.3390/antiox13030315
Branon TC, Bosch JA, Sanchez AD, Udeshi ND, Svinkina T, Carr SA, Feldman JL, Perrimon N, Ting AY (2018) Efficient proximity labeling in living cells and organisms with TurboID. Nat Biotechnol 36(9):880–887. https://doi.org/10.1038/nbt.4201
Chen C, Li C, Wang Y (2017) Cytosolic acetyl-CoA promotes histone acetylation predominantly at H3K27 in Arabidopsis. Nature Plants 3:814–824. https://doi.org/10.1038/s41477-017-0023-7
Chen X, Lu L, Qian S, Scalf M, Smith LM, Zhong X (2018) Canonical and noncanonical actions of Arabidopsis histone deacetylases in ribosomal RNA processing. Plant Cell 30(1):134–152. https://doi.org/10.1105/tpc.17.00626
Chen WQ, Drapek C, Li DX, Xu ZH, Benfey PN, Bai SN (2019) Histone deacetylase HDA19 affects root cortical cell fate by interacting with SCARECROW. Plant Physiol 180(1):276–288. https://doi.org/10.1104/pp.19.00056
Chen X, Ding AB, Zhong X (2020) Functions and mechanisms of plant histone deacetylases. Sci China Life Sci 63(2):206–216. https://doi.org/10.1007/s11427-019-1587-x
Chen X, Xu Q, Duan Y, Liu H, Chen X, Huang J, Luo C, Zhou DX, Zheng L (2021) Ustilaginoidea virens modulates lysine 2-hydroxyisobutyrylation in rice flowers during infection. J Integr Plant Biol 63(10):1801–1814. https://doi.org/10.1111/jipb.13149
Chen X, Duan Y, Qiao F, Liu H, Huang J, Luo C, Chen X, Li G, Xie K, Hsiang T, Zheng L (2022) A secreted fungal effector suppresses rice immunity through host histone hypoacetylation. New Phytol 235(5):1977–1994. https://doi.org/10.1093/plphys/kiac334
Chen X, Liu C, Wang H, Liu Q, Yue Y, Duan Y, Wang Z, Zheng L, Chen X, Wang Y, Huang J, Xu Q, Pan Y (2024) Ustilaginoidea virens-secreted effector Uv1809 suppresses rice immunity by enhancing OsSRT2-mediated histone deacetylation. Plant Biotechnol J 22(1):148–164. https://doi.org/10.1111/pbi.14174
Cho LH, Yoon J, Wai AH, An G (2018) Histone deacetylase 701 (HDT701) induces flowering in rice by modulating expression of OsIDS1. Mol Cells 41(7):665–675. https://doi.org/10.14348/molcells.2018.0148
Choudhary C, Weinert B, Nishida Y (2014) The growing landscape of lysine acetylation links metabolism and cell signalling. Nat Rev Mol Cell Biol 15:536–550. https://doi.org/10.1038/nrm3841
Chung PJ, Kim YS, Park SH, Nahm BH, Kim JK (2009) Subcellular localization of rice histone deacetylases in organelles. FEBS Lett 583(13):2249–2254. https://doi.org/10.1016/j.febslet.2009.06.003
Cigliano RA, Cremona G, Paparo R, Termolino P, Perrella G, Gutzat R, Consiglio MF, Conicella C (2013) Histone deacetylase AtHDA7 is required for female gametophyte and embryo development in Arabidopsis. Plant Physiol 163(1):431–440. https://doi.org/10.1104/pp.113.221713
Ding B, Bellizzi Mdel R, Ning Y, Meyers BC, Wang GL (2012) HDT701, a histone H4 deacetylase, negatively regulates plant innate immunity by modulating histone H4 acetylation of defense-related genes in rice. Plant Cell 24(9):3783–3794. https://doi.org/10.1105/tpc.112.101972
Fang X, Chen W, Zhao Y, Ruan S, Zhang H, Yan C, Jin L, Cao L, Zhu J, Ma H, Cheng Z (2015) Global analysis of lysine acetylation in strawberry leaves. Front Plant Sci 6:739. https://doi.org/10.3389/fpls.2015.00739
Fina JP, Casati P (2015) HAG3, a histone acetyltransferase, affects UV-B responses by negatively regulating the expression of DNA repair enzymes and sunscreen content in Arabidopsis thaliana. Plant Cell Physiol 56(7):1388–1400. https://doi.org/10.1093/pcp/pcv054
Fina JP, Masotti F, Rius SP, Crevacuore F, Casati P (2017) HAC1 and HAF1 histone acetyltransferases have different roles in UV-B responses in Arabidopsis. Front Plant Sci 8:1179. https://doi.org/10.3389/fpls.2017.01179
Finkemeier I, Laxa M, Miguet L, Howden AJ, Sweetlove LJ (2011) Proteins of diverse function and subcellular location are lysine acetylated in Arabidopsis. Plant Physiol 155(4):1779–1790. https://doi.org/10.1104/pp.110.171595
Gao MJ, Li X, Huang J, Gropp GM, Gjetvaj B, Lindsay DL, Wei S, Coutu C, Chen Z, Wan XC, Hannoufa A, Lydiate DJ, Gruber MY, Chen ZJ, Hegedus DD (2015) SCARECROW-LIKE15 interacts with HISTONE DEACETYLASE19 and is essential for repressing the seed maturation programme. Nat Commun 6:7243. https://doi.org/10.1038/ncomms8243
Gough C, Sadanandom A (2021) Understanding and exploiting post-translational modifications for plant disease resistance. Biomolecules 11(8):1122. https://doi.org/10.3390/biom11081122
Gu W, Roeder RG (1997) Activation of p53 sequence-specific DNA binding by acetylation of the p53 C-terminal domain. Cell 90(4):595–606. https://doi.org/10.1016/S0092-8674(00)80521-8
Guo Y, Tan Y, Qu M, Hong K, Zeng L, Wang L, Zhuang C, Qian Q, Hu J, Xiong G (2023) OsWR2 recruits HDA704 to regulate the deacetylation of H4K8ac in the promoter of OsABI5 in response to drought stress. J Integr Plant Biol 65(7):1651–1669. https://doi.org/10.1111/jipb.13481
Han Y, Georgii E, Priego-Cubero S, Wurm CJ, Hüther P, Huber G, Koller R, Becker C, Durner J, Lindermayr C (2023a) Arabidopsis histone deacetylase HD2A and HD2B regulate seed dormancy by repressing DELAY OF GERMINATION 1. Front Plant Sci 14:1124899. https://doi.org/10.3389/fpls.2023.1124899
Han Y, Haouel A, Georgii E, Priego-Cubero S, Wurm CJ, Hemmler D, Schmitt-Kopplin P, Becker C, Durner J, Lindermayr C (2023b) Histone deacetylases HD2A and HD2B undergo feedback regulation by ABA and modulate drought tolerance via mediating ABA-induced transcriptional repression. Genes (Basel) 14(6):1199. https://doi.org/10.3390/genes14061199
Hao Y, Wang H, Qiao S, Leng L, Wang X (2016) Histone deacetylase HDA6 enhances brassinosteroid signaling by inhibiting the BIN2 kinase. Proc Natl Acad Sci USA 113(37):10418–10423. https://doi.org/10.1073/pnas.1521363113
He D, Wang Q, Li M, Damaris RN, Yi X, Cheng Z, Yang P (2016) Global proteome analyses of lysine acetylation and succinylation reveal the widespread involvement of both modification in metabolism in the embryo of germinating rice seed. J Proteome Res 15(3):879–890. https://doi.org/10.1021/acs.jproteome.5b00805
He Y, Hong G, Zhang H, Tan X, Li L, Kong Y, Sang T, Xie K, Wei J, Li J, Yan F, Wang P, Tong H, Chu C, Chen J, Sun Z (2020) The OsGSK2 kinase integrates brassinosteroid and jasmonic acid signaling by interacting with OsJAZ4. Plant Cell 32(9):2806–2822. https://doi.org/10.1105/tpc.19.00499
Hinckley WE, Keymanesh K, Cordova JA, Brusslan JA (2019) The HAC1 histone acetyltransferase promotes leaf senescence and regulates the expression of ERF022. Plant Direct 3(8):e00159. https://doi.org/10.1002/pld3.159
Hou J, Ren R, Xiao H, Chen Z, Yu J, Zhang H, Shi Q, Hou H, He S, Li L (2021) Characteristic and evolution of HAT and HDAC genes in Gramineae genomes and their expression analysis under diverse stress in Oryza sativa. Planta 253(3):72. https://doi.org/10.1007/s00425-021-03589-1
Hou J, Zheng X, Ren R, Shi Q, Xiao H, Chen Z, Yue M, Wu Y, Hou H, Li L (2022) The histone deacetylase 1/GSK3/SHAGGY-like kinase 2/BRASSINAZOLE-RESISTANT 1 module controls lateral root formation in rice. Plant Physiol 189(2):858–873. https://doi.org/10.1093/plphys/kiac015
Hou J, Xiao H, Yao P, Ma X, Shi Q, Yang J, Hou H, Li L (2024) Unveiling the mechanism of broad-spectrum blast resistance in rice: the collaborative role of transcription factor OsGRAS30 and histone deacetylase OsHDAC1. Plant Biotechnol J 22(6):1740–1756. https://doi.org/10.1111/pbi.14299
Huang D, Lan W, Ma W, Huang R, Lin W, Li M, Chen CY, Wu K, Miao Y (2022) WHIRLY1 recruits the histone deacetylase HDA15 repressing leaf senescence and flowering in Arabidopsis. J Integr Plant Biol 64(7):1411–1429. https://doi.org/10.1111/jipb.13272
Hubbert C, Guardiola A, Shao R (2002) HDAC6 is a microtubule-associated deacetylase. Nature 417(6887):455–458. https://doi.org/10.1038/417455a
Ju L, Dong H, Yang R, Jing Y, Zhang Y, Liu L, Zhu Y, Chen KM, Ping J, Sun J (2023) BIN2 phosphorylates the Thr280 of CO to restrict its function in promoting Arabidopsis flowering. Front Plant Sci 14:1068949. https://doi.org/10.3389/fpls.2023.1068949
Kim YK, Kim S, Shin YJ, Hur YS, Kim WY, Lee MS, Cheon CI, Verma DP (2014) Ribosomal protein S6, a target of rapamycin, is involved in the regulation of rRNA genes by possible epigenetic changes in Arabidopsis. J Biol Chem 289(7):3901–3912. https://doi.org/10.1074/jbc.M113.515015
Kim S, Piquerez SJM, Ramirez-Prado JS, Mastorakis E, Veluchamy A, Latrasse D, Manza-Mianza D, Brik-Chaouche R, Huang Y, Rodriguez-Granados NY, Concia L, Blein T, Citerne S, Bendahmane A, Bergounioux C, Crespi M, Mahfouz MM, Raynaud C, Hirt H, Ntoukakis V, Benhamed M (2020) GCN5 modulates salicylic acid homeostasis by regulating H3K14ac levels at the 5’ and 3’ ends of its target genes. Nucleic Acids Res 48(11):5953–5966. https://doi.org/10.1093/nar/gkaa369
Kim TW, Park CH, Hsu CC, Kim YW, Ko YW, Zhang Z, Zhu JY, Hsiao YC, Branon T, Kaasik K, Saldivar E, Li K, Pasha A, Provart NJ, Burlingame AL, Xu SL, Ting AY, Wang ZY (2023) Mapping the signaling network of BIN2 kinase using TurboID-mediated biotin labeling and phosphoproteomics. Plant Cell 35(3):975–993. https://doi.org/10.1093/plcell/koad013
König AC, Hartl M, Pham PA, Laxa M, Boersema PJ, Orwat A, Kalitventseva I, Plöchinger M, Braun HP, Leister D, Mann M, Wachter A, Fernie AR, Finkemeier I (2014) The Arabidopsis class II sirtuin is a lysine deacetylase and interacts with mitochondrial energy metabolism. Plant Physiol 164(3):1401–1414. https://doi.org/10.1104/pp.113.232496
Krogan NT, Hogan K, Long JA (2012) APETALA2 negatively regulates multiple floral organ identity genes in Arabidopsis by recruiting the co-repressor TOPLESS and the histone deacetylase HDA19. Development 139(22):4180–4190. https://doi.org/10.1242/dev.085407
Kumar V, Thakur JK, Prasad M (2021) Histone acetylation dynamics regulating plant development and stress responses. Cell Mol Life Sci 78(10):4467–4486. https://doi.org/10.1007/s00018-021-03794-x
Lee HG, Seo PJ (2019) MYB96 recruits the HDA15 protein to suppress negative regulators of ABA signaling in Arabidopsis. Nat Commun 10(1):1713. https://doi.org/10.1038/s41467-019-09417-1
Li X, Ye J, Ma H, Lu P (2018) Proteomic analysis of lysine acetylation provides strong evidence for involvement of acetylated proteins in plant meiosis and tapetum function. Plant J 93(1):142–154. https://doi.org/10.1111/tpj.13766
Li W, Xiong Y, Lai LB, Zhang K, Li Z, Kang H, Dai L, Gopalan V, Wang GL, Liu W (2021) The rice RNase P protein subunit Rpp30 confers broad-spectrum resistance to fungal and bacterial pathogens. Plant Biotechnol J 19(10):1988–1999. https://doi.org/10.1111/pbi.13612
Li J, Ma X, Wang C, Liu S, Yu G, Gao M, Qian H, Liu M, Luisi BF, Gabriel DW, Liang W (2022) Acetylation of a fungal effector that translocates host PR1 facilitates virulence. Elife 11:e82628. https://doi.org/10.7554/eLife.82628
Liu X, Luo M, Zhang W (2012) Histone acetyltransferases in rice (Oryza sativa L.): phylogenetic analysis, subcellular localization and expression. BMC Plant Biol 12:145. https://doi.org/10.1186/1471-2229-12-145
Liu C, Li LC, Chen WQ, Chen X, Xu ZH, Bai SN (2013) HDA18 affects cell fate in Arabidopsis root epidermis via histone acetylation at four kinase genes. Plant Cell 25(1):257–269. https://doi.org/10.1105/tpc.112.107045
Liu X, Wei W, Zhu W, Su L, Xiong Z, Zhou M, Zheng Y, Zhou DX (2017) Histone deacetylase AtSRT1 links metabolic flux and stress response in Arabidopsis. Mol Plant 10(12):1510–1522. https://doi.org/10.1016/j.molp.2017.10.010
Luo M, Wang YY, Liu X, Yang S, Lu Q, Cui Y, Wu K (2012a) HD2C interacts with HDA6 and is involved in ABA and salt stress response in Arabidopsis. J Exp Bot 63(8):3297–3306. https://doi.org/10.1093/jxb/ers059
Luo M, Yu CW, Chen FF, Zhao L, Tian G, Liu X, Cui Y, Yang JY, Wu K (2012b) Histone deacetylase HDA6 is functionally associated with AS1 in repression of KNOX genes in arabidopsis. PLoS Genet 8(12):e1003114. https://doi.org/10.1371/journal.pgen.1003114
Luo Y, Shi DQ, Jia PF, Bao Y, Li HJ, Yang WC (2022) Nucleolar histone deacetylases HDT1, HDT2, and HDT3 regulate plant reproductive development. J Genet Genomics 49(1):30–39. https://doi.org/10.1016/j.jgg.2021.10.002
Mayer KS, Chen X, Sanders D, Chen J, Jiang J, Nguyen P, Scalf M, Smith LM, Zhong X (2019) HDA9-PWR-HOS15 is a core histone deacetylase complex regulating transcription and development. Plant Physiol 180(1):342–355. https://doi.org/10.1104/pp.18.01156
Meng F, Zheng X, Wang J, Qiu T, Yang Q, Fang K, Bhadauria V, Peng YL, Zhao W (2024) The GRAS protein OsDLA involves in brassinosteroid signalling and positively regulates blast resistance by forming a module with GSK2 and OsWRKY53 in rice. Plant Biotechnol J 22(2):363–378. https://doi.org/10.1111/pbi.14190
Narita T, Weinert BT, Choudhary C (2019) Functions and mechanisms of non-histone protein acetylation. Nat Rev Mol Cell Biol 20(3):156–174. https://doi.org/10.1038/s41580-018-0081-3
Ning YQ, Chen Q, Lin RN, Li YQ, Li L, Chen S, He XJ (2019) The HDA19 histone deacetylase complex is involved in the regulation of flowering time in a photoperiod-dependent manner. Plant J 98(3):448–464. https://doi.org/10.1111/tpj.14229
Niu D, Lin XL, Kong X, Qu GP, Cai B, Lee J, Jin JB (2019) SIZ1-mediated SUMOylation of TPR1 suppresses plant immunity in Arabidopsis. Mol Plant 12(2):215–228. https://doi.org/10.1016/j.molp.2018.12.002
Niu Y, Bai J, Liu X, Zhang H, Bao J, Zhao W, Hou Y, Deng X, Yang C, Guo L, Geng Z, Xie H, Wu H, Shen M, Lou X, Tang W, Liu X, Sun D, Cao X, Zheng S (2022) HISTONE DEACETYLASE 9 transduces heat signal in plant cells. Proc Natl Acad Sci USA 119(45):e2206846119. https://doi.org/10.1073/pnas.2206846119
Pandey R, Müller A, Napoli CA, Selinger DA, Pikaard CS, Richards EJ, Bender J, Mount DW, Jorgensen RA (2002) Analysis of histone acetyltransferase and histone deacetylase families of Arabidopsis thaliana suggests functional diversification of chromatin modification among multicellular eukaryotes. Nucleic Acids Res 30(23):5036–5055. https://doi.org/10.1093/nar/gkf660
Park HJ, Baek D, Cha JY, Liao X, Kang SH, McClung CR, Lee SY, Yun DJ, Kim WY (2019) HOS15 interacts with the histone deacetylase HDA9 and the evening complex to epigenetically regulate the floral activator GIGANTEA. Plant Cell 31(1):37–51. https://doi.org/10.1105/tpc.18.00721
Pontes O, Lawrence RJ, Silva M, Preuss S, Costa-Nunes P, Earley K, Neves N, Viegas W, Pikaard CS (2007) Postembryonic establishment of megabase-scale gene silencing in nucleolar dominance. PLoS One 2(11):e1157. https://doi.org/10.1371/journal.pone.0001157
Rao RS, Thelen JJ, Miernyk JA (2014) Is Lys-Nɛ-acetylation the next big thing in post-translational modifications? Trends Plant Sci 19(9):550–553. https://doi.org/10.1016/j.tplants.2014.05.001
Seto E, Yoshida M (2014) Erasers of histone acetylation: the histone deacetylase enzymes. Cold Spring Harb Perspect Biol 6(4):a018713. https://doi.org/10.1101/cshperspect.a018713
Smith-Hammond CL, Hoyos E, Miernyk JA (2014) The pea seedling mitochondrial Nε-lysine acetylome. Mitochondrion 19 Pt B:154–165. https://doi.org/10.1016/j.mito.2014.04.012
Soufi B, Soares NC, Ravikumar V, Macek B (2012) Proteomics reveals evidence of cross-talk between protein modifications in bacteria: focus on acetylation and phosphorylation. Curr Opin Microbiol 15(3):357–363. https://doi.org/10.1016/j.mib.2012.05.003
Sun Y, Xie Z, Jin L, Qin T, Zhan C, Huang J (2024) Histone deacetylase OsHDA716 represses rice chilling tolerance by deacetylating OsbZIP46 to reduce its transactivation function and protein stability. Plant Cell 36(5):1913–1936. https://doi.org/10.1093/plcell/koae010
Suskiewicz MJ (2024) The logic of protein post-translational modifications (PTMs): chemistry, mechanisms and evolution of protein regulation through covalent attachments. BioEssays 46(3):e2300178. https://doi.org/10.1002/bies.202300178
Sutherland EW, Wosilait WD (1955) Inactivation and activation of liver phosphorylase. Nature 175(4447):169–170. https://doi.org/10.1038/175169a0
Suzuki M, Shinozuka N, Hirakata T, Nakata MT, Demura T, Tsukaya H, Horiguchi G (2018) OLIGOCELLULA1/HIGH EXPRESSION OF OSMOTICALLY RESPONSIVE GENES15 promotes cell proliferation with HISTONE DEACETYLASE9 and POWERDRESS during leaf development in Arabidopsis thaliana. Front Plant Sci 9:580. https://doi.org/10.3389/fpls.2018.00580
Tahir MS, Tian L (2021) HD2-type histone deacetylases: unique regulators of plant development and stress responses. Plant Cell Rep 40(9):1603–1615. https://doi.org/10.1007/s00299-021-02688-3
Tilak P, Kotnik F, Née G, Seidel J, Sindlinger J, Heinkow P, Eirich J, Schwarzer D, Finkemeier I (2023) Proteome-wide lysine acetylation profiling to investigate the involvement of histone deacetylase HDA5 in the salt stress response of Arabidopsis leaves. Plant J 115(1):275–292. https://doi.org/10.1111/tpj.16206
Tran HT, Nimick M, Uhrig RG, Templeton G, Morrice N, Gourlay R, DeLong A, Moorhead GB (2012) Arabidopsis thaliana histone deacetylase 14 (HDA14) is an α-tubulin deacetylase that associates with PP2A and enriches in the microtubule fraction with the putative histone acetyltransferase ELP3. Plant J 71(2):263–272. https://doi.org/10.1111/j.1365-313x.2012.04984.x
Truong HA, Lee S, Trịnh CS, Lee WJ, Chung EH, Hong SW, Lee H (2021) Overexpression of the HDA15 gene confers resistance to salt stress by the induction of NCED3, an ABA biosynthesis enzyme. Front Plant Sci 12:640443. https://doi.org/10.3389/fpls.2021.640443
Tyagi SC, Stanisic D, Singh M (2021) Epigenetic memory: gene writer, eraser and homocysteine. Mol Cell Biochem 476(2):507–512. https://doi.org/10.1007/s11010-020-03895-4
Uhrig RG, Schläpfer P, Mehta D, Hirsch-Hoffmann M, Gruissem W (2017) Genome-scale analysis of regulatory protein acetylation enzymes from photosynthetic eukaryotes. BMC Genomics 18(1):514. https://doi.org/10.1186/s12864-017-3894-0
Uhrig RG, Schläpfer P, Roschitzki B, Hirsch-Hoffmann M, Gruissem W (2019) Diurnal changes in concerted plant protein phosphorylation and acetylation in Arabidopsis organs and seedlings. Plant J 99(1):176–194. https://doi.org/10.1111/tpj.14315
Ullah F, Xu Q, Zhao Y, Zhou DX (2020) Histone deacetylase HDA710 controls salt tolerance by regulating ABA signaling in rice. J Integr Plant Biol. https://doi.org/10.1111/jipb.13042. (online ahead of print)
Verdin E, Ott M (2015) 50 years of protein acetylation: from gene regulation to epigenetics, metabolism and beyond. Nat Rev Mol Cell Biol 16:258–264. https://doi.org/10.1038/nrm3931
Wang Z, Cao H, Sun Y, Li X, Chen F, Carles A, Li Y, Ding M, Zhang C, Deng X, Soppe WJ, Liu YX (2013) Arabidopsis paired amphipathic helix proteins SNL1 and SNL2 redundantly regulate primary seed dormancy via abscisic acid-ethylene antagonism mediated by histone deacetylation. Plant Cell 25(1):149–166. https://doi.org/10.1105/tpc.112.108191
Wang J, Liu C, Chen Y, Zhao Y, Ma Z (2021) Protein acetylation and deacetylation in plant-pathogen interactions. Environ Microbiol 23(9):4841–4855. https://doi.org/10.1111/1462-2920.15725
Wang H, Jiao X, Zhang X, Zhang M, Liu Y, Chen X, Fang R, Yan Y (2023a) Ammonium protects rice against rice stripe virus by activating HDA703/OsBZR1-mediated BR signaling. Plant Sci 326:111504. https://doi.org/10.1016/j.plantsci.2022.111504
Wang Q, Li X, Guo C, Wen L, Deng Z, Zhang Z, Li W, Liu T, Guo Y (2023b) Senescence-related receptor kinase 1 functions downstream of WRKY53 in regulating leaf senescence in Arabidopsis. J Exp Bot 74(17):5140–5152. https://doi.org/10.1093/jxb/erad240
Xia L, Kong X, Song H, Han Q, Zhang S (2021) Advances in proteome-wide analysis of plant lysine acetylation. Plant Commun 3(1):100266. https://doi.org/10.1016/j.xplc.2021.100266
Xiong Y, Peng X, Cheng Z, Liu W, Wang GL (2016) A comprehensive catalog of the lysine-acetylation targets in rice (Oryza sativa) based on proteomic analyses. J Proteomics 138:20–29. https://doi.org/10.1016/j.jprot.2016.01.019
Xu Q, Liu Q, Chen Z, Yue Y, Liu Y, Zhao Y, Zhou DX (2021) Histone deacetylases control lysine acetylation of ribosomal proteins in rice. Nucleic Acids Res 49(8):4613–4628. https://doi.org/10.1093/nar/gkab244
Yang J, Qu X, Li T, Gao Y, Du H, Zheng L, Ji M, Zhang P, Zhang Y, Hu J, Liu L, Lu Z, Yang Z, Zhang H, Yang J, Jiao Y, Zheng X (2023) HY5-HDA9 orchestrates the transcription of HsfA2 to modulate salt stress response in Arabidopsis. J Integr Plant Biol 65(1):45–63. https://doi.org/10.1111/jipb.13372
Yang Z, Du J, Tan X, Zhang H, Li L, Li Y, Wei Z, Xu Z, Lu Y, Chen J, Sun Z (2024) Histone deacetylase OsHDA706 orchestrates rice broad-spectrum antiviral immunity and is impeded by a viral effector. Cell Rep 43(3):113838. https://doi.org/10.1016/j.celrep.2024.113838
Yu CW, Liu X, Luo M, Chen C, Lin X, Tian G, Lu Q, Cui Y, Wu K (2011) HISTONE DEACETYLASE6 interacts with FLOWERING LOCUS D and regulates flowering in Arabidopsis. Plant Physiol 156(1):173–184. https://doi.org/10.1104/pp.111.174417
Yu CW, Tai R, Wang SC, Yang P, Luo M, Yang S, Cheng K, Wang WC, Cheng YS, Wu K (2017) HISTONE DEACETYLASE6 acts in concert with histone methyltransferases SUVH4, SUVH5, and SUVH6 to regulate transposon silencing. Plant Cell 29(8):1970–1983. https://doi.org/10.1105/tpc.16.00570
Yu Y, Zhao F, Yue Y, Zhao Y, Zhou DX (2024) Lysine acetylation of histone acetyltransferase adaptor protein ADA2 is a mechanism of metabolic control of chromatin modification in plants. Nat Plants 10(3):439–452. https://doi.org/10.1038/s41477-024-01623-0
Zhang Y, Zeng L (2020) Crosstalk between ubiquitination and other post-translational protein modifications in plant Immunity. Plant Commun 1(4):100041. https://doi.org/10.1016/j.xplc.2020.100041
Zhang J, Sprung R, Pei J, Tan X, Kim S, Zhu H, Liu CF, Grishin NV, Zhao Y (2009) Lysine acetylation is a highly abundant and evolutionarily conserved modification in Escherichia coli. Mol Cell Proteomics 8(2):215–225. https://doi.org/10.1074/mcp.m800187-mcp200
Zhang H, Lu Y, Zhao Y, Zhou DX (2016) OsSRT1 is involved in rice seed development through regulation of starch metabolism gene expression. Plant Sci 248:28–36. https://doi.org/10.1016/j.plantsci.2016.04.004
Zhang F, Wang L, Ko EE, Shao K, Qiao H (2018) Histone deacetylases SRT1 and SRT2 interact with ENAP1 to mediate ethylene-induced transcriptional repression. Plant Cell 30(1):153–166. https://doi.org/10.1105/tpc.17.00671
Zhang N, Lv F, Qiu F, Han D, Xu Y, Liang W (2023) Pathogenic fungi neutralize plant-derived ROS via Srpk1 deacetylation. EMBO J 42(9):e112634. https://doi.org/10.15252/embj.2022112634
Zhang N, Hu J, Liu Z, Liang W, Song L (2024) Sir2-mediated cytoplasmic deacetylation facilitates pathogenic fungi infection in host plants. New Phytol 241(4):1732–1746. https://doi.org/10.1111/nph.19438
Zhao J, Li M, Gu D, Liu X, Zhang J, Wu K, Zhang X, Teixeira da Silva JA, Duan J (2016) Involvement of rice histone deacetylase HDA705 in seed germination and in response to ABA and abiotic stresses. Biochem Biophys Res Commun 470(2):439–444. https://doi.org/10.1016/j.bbrc.2016.01.016
Zhao L, Peng T, Chen CY, Ji R, Gu D, Li T, Zhang D, Tu YT, Wu K, Liu X (2019) HY5 interacts with the histone deacetylase HDA15 to repress hypocotyl cell elongation in photomorphogenesis. Plant Physiol 180(3):1450–1466. https://doi.org/10.1104/pp.19.00055
Zheng Y, Ge J, Bao C, Chang W, Liu J, Shao J, Liu X, Su L, Pan L, Zhou DX (2020) Histone deacetylase HDA9 and WRKY53 transcription factor are mutual antagonists in regulation of plant stress response. Mol Plant 13(4):598–611. https://doi.org/10.1016/j.molp.2019.12.011
Zheng Y, Li Z, Cui X, Yang Z, Bao C, Pan L, Liu X, Chatel-Innocenti G, Vanacker H, Noctor G, Dard A, Reichheld JP, Issakidis-Bourguet E, Zhou DX (2023) S-Nitrosylation of the histone deacetylase HDA19 stimulates its activity to enhance plant stress tolerance in Arabidopsis. Plant J 114(4):836–854. https://doi.org/10.1111/tpj.16174
Zhou C, Labbe H, Sridha S, Wang L, Tian L, Latoszek-Green M, Yang Z, Brown D, Miki B, Wu K (2004) Expression and function of HD2-type histone deacetylases in Arabidopsis development. Plant J 38(5):715–724. https://doi.org/10.1111/j.1365-313X.2004.02083.x
Zhou Y, Tan B, Luo M, Li Y, Liu C, Chen C, Yu CW, Yang S, Dong S, Ruan J, Yuan L, Zhang Z, Zhao L, Li C, Chen H, Cui Y, Wu K, Huang S (2013) HISTONE DEACETYLASE19 interacts with HSL1 and participates in the repression of seed maturation genes in Arabidopsis seedlings. Plant Cell 25(1):134–148. https://doi.org/10.1105/tpc.112.096313
Zhou S, Jiang W, Long F, Cheng S, Yang W, Zhao Y, Zhou DX (2017) Rice homeodomain protein WOX11 recruits a histone acetyltransferase complex to establish programs of cell proliferation of crown root meristem. Plant Cell 29(5):1088–1104. https://doi.org/10.1105/tpc.16.00908
Zhou H, Finkemeier I, Guan W, Tossounian MA, Wei B, Young D, Huang J, Messens J, Yang X, Zhu J, Wilson MH, Shen W, Xie Y, Foyer CH (2018) Oxidative stress-triggered interactions between the succinyl- and acetyl-proteomes of rice leaves. Plant Cell Environ 41(5):1139–1153. https://doi.org/10.1111/pce.13100
Zhu GR, Yan X, Zhu D, Deng X, Wu JS, Xia J, Yan YM (2018) Lysine acetylproteome profiling under water deficit reveals key acetylated proteins involved in wheat grain development and starch biosynthesis. J Proteomics 185:8–24. https://doi.org/10.1016/j.jprot.2018.06.019
Zhu T, Li B, Chen Y, Jing Y, Wang S, Li W, Gao N, Liao C, Wang L, Xiao F, Li T (2023) BRASSINOSTEROID-INSENSITIVE 2 regulates salt stress tolerance in Arabidopsis by promoting AGL16 activity. Biochem Biophys Res Commun 678:17–23. https://doi.org/10.1016/j.bbrc.2023.08.031
Funding
This research was supported by grants from the National Natural Science Foundation of China (No. 31871238).
Author information
Authors and Affiliations
Contributions
J.H. and L.L. conceived the ideas. Z.Z. performed data curation and original draft. Y.Z. performed data curation, formal analysis, methodology and validation. J.H. and L.L. supervised the research and revised the article.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no competing interests.
Ethical approval and consent to participate
Not applicable.
Additional information
Communicated by Gerhard Leubner.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Z., Zeng, Y., Hou, J. et al. Advances in understanding the roles of plant HAT and HDAC in non-histone protein acetylation and deacetylation. Planta 260, 93 (2024). https://doi.org/10.1007/s00425-024-04518-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-024-04518-8