Abstract
Main conclusion
The analysis of meiotic pairing affinities and genomic formulae in species and hybrids of Zea allowed us to speculate an evolutionary model to recreate the ancient polyploidization of maize and allied species.
Abstract
The meiotic pairing affinities and the genomic formulae analysis in Zea species and hybrids obtained in new and previous crosses, together with the molecular data known in the genus, allowed us to speculate an evolutionary model to attempt to recreate the ancient polyploidization process of Zea species. We propose that x = 5 semispecies are the ancestors of all modern species of the genus. The complex evolutionary process that originated the different taxa could be included hybridization between sympatric diploid ancestral semispecies (2n = 10) and recurrent duplication of the hybrid chromosome number, resulting in distinct auto- and allopolyploids. After the merger and doubling of independent genomes would have undergone cytological and genetical diploidization, implying revolutionary changes in genome organization and genic balance processes. Based on the meiotic behaviour of the 2n = 30 hybrids, that showed homoeology between the A subgenomes of all parental species, we propose that this subgenome A would be pivotal in all the species and would have conserved the rDNA sequences and the pairing regulator locus (PrZ). In the hypothetical model postulated here, the ancestral semispecies with the pivotal subgenome A would have had a wide geographic distribution, co-occurring and hybridizing with the semispecies harbouring B subgenomes, thus enabling sympatric speciation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Zea (Tribe Maydeae, Poaceae) is composed of the sections Zea and Luxuriantes. The Zea section consists of the well-established taxonomic entity Zea mays L., which includes four subspecies: Zea mays subsp. mays, Zea mays subsp. mexicana, Zea mays subsp. parviglumis and Zea mays subsp. huehuetenanguensis (Doebley 1990), all of which have 2n = 20 chromosomes. The Luxuriantes section (Doebley & Iltis) comprises the perennials Z. diploperennis (Iltis, Doebley & Guzman) (2n = 20) and Z. perennis (Hitchc.) Reeves & Mangelsdorf (2n = 40), as well as the annual Z. luxurians (Durieu & Ascherson) Bird (2n = 20).
Genetic studies showed that the maize genome is composed of two chromosome subgenomes, implying whole-genome duplication (Moore et al. 1995; Gale and Devos 1998). Gaut and Doebley (1997), who analyzed the pattern of sequence divergence among 14 pairs of duplicated genes, proposed a segmental allotetraploid origin of maize. Moreover, they estimated that the divergence of the genomes of the two maize progenitors, together with that of the Sorghum, occurred about 20.5 million years ago (Mya) and that the allotetraploid event occurred at about 11.4 Mya. Swigoñová et al. (2004) also supported a tetraploid origin for maize after analyzing the sequences of 11 orthologous genes.
The association of homologous or homeologous during meiosis, forming univalents, bivalents and multivalents, reveals the relative affinities between the parental genomes of hybrids and polyploid species (Sybenga 1975, 1996, 1999; Jenkins and Rees 1991; Sybenga et al. 1994; rev. in Soares et al. 2021). The meiotic behaviour of Zea species and their artificial hybrids revealed a relative affinity between homologous or homoeologous chromosomes and detected chromosomal rearrangements acting as reproductive isolation mechanisms. These studies allowed to postulate that maize and its allied species are polyploids with a basic number x = 5 (Naranjo et al. 1990; Poggio et al. 2005; González et al. 2006). These researchers assigned genomic formulae to different Zea species, and arbitrarily named AA and BB the two ancestral chromosomal subgenomes involved in the hybridization and polyploidy processes. The cytogenetic studies have strongly supported an allopolyploid origin for maize and its allied species (Poggio et al. 2005; González and Poggio 2015; Poggio and González 2018). Also based on cytogenetic analysis, Zafar Iqbal et al. (2018) proposed the origin of only two Zea species: maize and Z. perennis, which would have arisen from the crossing of two autopolyploids.
Modern maize is functionally diploid, with chromosomes normally pairing to form 10 bivalents during meiosis I. Cytological diploidization is essential for understanding polyploid speciation and is closely related to the process of meiotic stabilization (Poggio and González 2018). The restriction of chromosome associations to homologous/homoeologous chromosomes (i.e. diploid-like meiotic behaviour) has been traditionally explained by the action of Pairing homoeologous genes (Ph), which ensures correct chromosome segregation and overcomes reduction in fertility due to meiotic irregularities in autopolyploid, allopolyploid, and allo-autopolyploid species (rev. in Blasio et al. 2022). Poggio and González (2018) hypothesized that cytological diploidization in Zea species occurred by restriction of pairing and/or genetic divergence between homoeologous chromosomes. These authors proposed the existence of a paring regulator locus (PrZ) whose expression is inhibited by colchicine. After tetraploidization, many genes are lost in a return to genetic diploidy in a nonrandom mode depending on function (Freeling and Thomas 2006; Birchler and Veitia 2007, 2010, 2012). These authors state that the whole genome duplication do not disturb gene dosage balance and the multiple interacting components of a complex are preserved through the genetic diploidization. During the rebalancing allopolyploid, the shift between genomes can fixed different genetic variants on different chromosomes (Birchler and Veitia 2012).
In the present work, we analyse pairing affinities in Zea species and hybrids obtained in new and previous crosses. These studies, together with the molecular data known in the genus, allowed us to propose an evolutionary model to explain the polyploid speciation of maize and its wild relatives.
Materials and methods
Plant material
Zea mays subsp. mays Palomero Toluqueño race was provided by CIMMYT (Centro Internacional de Mejoramiento de Maíz y Trigo). Zea luxurians from Guatemala, Zea diploperennis from San Miguel (Ciudad Guzmán, Jalisco, Mexico) and Zea perennis from Piedra Ancha (San Gabriel, Jalisco, Mexico) were provided by Dra. C. Prywed from Colegio de Postgraduados, Montecillos, México. Z. mays subsp. mexicana from Chalco-Amecameca (Mesa Central, México) and Zea mays subsp. parviglumis from Balsas valley (Guerrero, México) were provided by Dr. T. A. Kato Yamakake from Colegio de Postgraduados, Montecillos, México.
New intra- and interspecific artificial crossings were carried out to obtain the F1 hybrid plants. About 20 plants per species were hand-pollinated with a bulk of pollen from 5 male plants. The species and hybrids were cultivated in the greenhouse of the Facultad de Agronomía (FA), Universidad de Buenos Aires (UBA). The material from species and hybrids was deposited at the seed bank of the Laboratory of Genetic Resources N. I. Vavilov at the FA, UBA.
Meiotic analysis
Young panicles from Zea species and F1 hybrids were fixed in a 3:1 solution of absolute ethanol:acetic acid (v/v) and squashed in a drop of 2% acetic haematoxylin. The frequencies of each pairing configurations were determined at diakinesis-metaphase I. The meiotic behaviour was analyzed in 200 to 300 meiocyte from 3–5 individuals per studied species and hybrids.
Normal (stained) and aborted (unstained) pollen grains were differentiated using Alexander's stain (Alexander 1969) and the percentage viability was estimated in at least 250 pollen grains from 5 individuals for each hybrid combination. The percentage of viable seeds was estimated in at least 200 seeds from 3–5 individuals for each hybrid combination.
Results
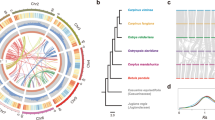
We analyzed the meiotic behaviour of species and artificial inter- and intrasectional hybrids obtained in the present study. Figure 1 shows the most frequent meiotic configurations and the genomic formulae proposed for the species and hybrids analyzed in this and previous studies. The progenitor species, with 2n = 20, formed 10 bivalents (II), while Z. perennis, with 2n = 40, had the most frequent meiotic configuration of 5 quadrivalents (IV) + 10 bivalents (II) (Figs. 1, 2A, B). Figure 2 illustrates the genomic origin (subgenomes A or B) of the univalents (I), bivalents (II), trivalents (III) and quadrivalents (IV), according to the genomic formulae previously proposed for species and hybrids.
Zea intra- and interspecific hybrids. Zea section enclosed in orange squares and Luxuriantes section in blue squares. Red arrows indicate the hybrids studied in the present work and black arrows the hybrids analyzed in Naranjo et al. (1990), Poggio et al. (2005) and Poggio and González (2018). In the genomic formulae of Zea species and hybrids, the subgenomes are arbitrarily named as AA and BB and the black arcs show the meiotic associations more frequently observed among homologous and homoeologous chromosomes. Arrows show the more frequent meiotic configurations of hybrids. Ref.: I: univalents, II: bivalents, III: trivalents, IV: quadrivalents
A Ten bivalents from Z. m. subsp. mays at metaphase I. The homologous pairings Am-Am and Bm-Bm are exemplified in 2 bivalents. B Diakinesis of Z. perennis with the most frequent configuration 5IV + 10II. The IV are formed by chromosomes of subgenomes A (ApApAp´Ap´) and the II are formed by chromosomes of subgenomes B (Bp1Bp1 and Bp2Bp2). C 9II + 2I at metaphase I of Z. diploperennis x Z. m. subsp. mays. Homoeologous pairing Am-Ad on one bivalent and Bm and Bd univalents are indicated. D 5III + 5II + 5I of Z. perennis x Z. m. subsp. parviglumis at metaphase I. The III formed by the pairing between subgenomes A (ApAp´Apar), the II formed by the subgenomes B from Z. perennis (Bp1Bp2) and the I by the subgenomes B from Z. m. subsp. parviglumis (Bpar). Letters and suffixes were assigned by convention and correspond to the genomic formulae proposed in Fig. 1. Bars = 10 µm
In the hybrids involving Z. luxurians (2n = 20) as one of the progenitors and Z. mays subsp. parviglumis, Z. mays subsp. mexicana or Z. diploperennis as the other, the most frequent configuration was 9II + 2I, with a range from 7 to 10 bivalents. All these hybrids showed several meiotic abnormalities, such as (i) heterozygosity in the knob regions at pachytene (Fig. 3A); (ii) heteromorphic bivalents or univalents of different size at diplotene-diakinesis and/or metaphase I (Fig. 3B, G); (iii) up to three bridges and three fragments at anaphase I (Fig. 3C); (iv) up to 10 laggard chromosomes at anaphase I (Fig. 3D) and (v) meiotic asynchrony of two groups of five bivalents each at diplotene-metaphase I (Fig. 3F–H). These hybrids exhibited high pollen and seed sterility (90% and 99%, respectively).
A Pachitene of Z. luxurians x Z. diploperennis. White arrows indicate the large knobs of Z. luxurians and black arrows the small knobs of Z. diploperennis. B Metaphase I of Z. luxurians x Z. m. subsp. mexicana with 9II + 2I. The black arrow points to 2 univalents of different sizes and the white arrow a heteromorphic bivalent. Inner squares enclose heteromorphic bivalents in diakinesis. C Anaphase I of Z. luxurians x Z. m. subsp. mexicana with one bridge (black arrow) and one fragment (white arrow). D Anaphase I of Z. luxurians x Z. diploperennis with laggard chromosomes. E Metaphase I of Z. perennis x Z. m. subsp. parviglumis showing 5III + 5II + 5I. F Two asynchronous groups of 5II each in diakinesis of Z. diploperennis x Z. m. subsp. mays. G Metaphase I of Z. luxurians x Z. m. subsp. parviglumis showing two separate groups, one comprising 5 bivalents and the other 4 bivalents and 2 univalents of different size (black arrows). H Metaphase I of Z. diploperennis x Z. m. subsp. parviglumis hybrid with two asynchronous groups of 5II each. Bar = 10 µm
The hybrids Z. diploperennis x maize and Z. diploperennis x Z. m. subsp. parviglumis (2n = 20) exhibited regular meiosis, 10II or 9II + 2I were the most frequent configurations at prophase and metaphase I (Fig. 2C). Almost 50% of the studied cells contained two asynchronous groups of 5 IIs each, spatially separated at diakinesis and metaphase I (Fig. 3F). These hybrids were highly sterile, having a seed fertility of only 9.7%.
The hybrids Z. perennis x Z. m. subsp. parviglumis (2n = 30) were totally sterile and showed 5III + 5II + 5I as the most frequent chromosome associations and had up to 8III (Figs. 1, 2D, 3E).
Discussion
In this study, we speculate the evolutionary steps leading to the actual species of Zea based on the analysis of the meiotic behaviour observed in species and hybrids. Since 1990, much of our work has addressed the relative meiotic pairing affinity among Zea species. These early studies suggested a basic number of x = 5 as well as the polyploid nature the genus (Naranjo et al. 1990; Poggio et al. 1999, 2005). Both in previous research and in this study, the presence of two asynchronous groups of five bivalents each in diplotene-metaphase I has been reported in species and hybrids with 2n = 20. This phenomenon was considered to be a consequence of the spatial and temporal separation of the two relict diploid genomes, composed of 5II each, arbitrarily named AA and BB (Poggio et al. 2005; González and Poggio 2011). The Zea taxa with 2n = 20 are characterized by strict bivalent formation, raising the question of whether these species should be considered as genomic allopolyploids (sensu Stebbins 1971). And, if so, how does this fit with molecular evidence that maize is a segmental allopolyploid? (Moore et al. 1995; Gaut and Doebley 1997; Schnable et al. 2011). These questions were answered in the next section by proposing that such species underwent processes of cytological diploidization, without losing sight of the extensively studied processes of genetic diploidization (rev. in Birchler and Veitia 2012).
Cytological diploidization in Zea
In polyploids, diploid-like meiosis ensures correct chromosome segregation and overcomes fertility reduction due to meiotic irregularities. The development of a diploidization genetic system (Ph locus) confers stabilization to all polyploid wheat species because it inhibits the appearance of multivalent chromosomes with deleterious intergenomic exchange (Blasio et al. 2022). In this regard, a pairing regulator locus (PrZ) was reported in Zea, with its expression being suppressed by colchicine (Poggio et al. 1990; Poggio and González 2018). The treatment with colchicine in maize, Z. m. subsp. mexicana, Z. luxurians and Z. m. subsp. parviglumis led to the formation of up to 5IV, revealing pairing between chromosomes of different homoeologous subgenomes (A and B). These results suggested that these species should be considered segmental allotetraploids.
Poggio and González (2018) showed that colchicine treatment did not result in any homoeology between subgenomes A and B in the tetraploid Z. diploperennis and proposed that their diploidized meiotic behaviour would have been acquired by the divergence of homoeologous chromosomes belonging to these subgenomes. On this basis, this specie could be considered genomic allopolyploid (sensu Stebbins 1971).
The colchicine-treated Z. perennis showed up to 10IV and high affinity within subgenomes A and B, but no homology between them. In Z. perennis, the PrZ-pairing regulator locus was assumed to affect the A and B subgenomes independently. In conclusion, this species appeared to be the result of a combination between an intervarietal autotetraploid (ApAp´ subgenomes) and a segmental allopolyploid (Bp1Bp2 subgenomes) in the same nucleus (Poggio and González 2018).
In addition to the process of cytological diploidization caused by the effect of pairing regulator genes, it is worthwhile to consider the fact that genomes from two diverged species arising by hybridization and polyploidy are usually unstable and experience massive genetic changes (deletions, inversions, translocations, homoeologous exchanges and epigenetic modifications, among others) (rev. in Blasio et al. 2022). Since gene loss occurs following allotetraploid events, the spectrum of preserved genes is not random (Birchler and Veitia 2010). Studies on retaining selected duplicate genes revealed that signal transduction and transcription factors as well regulatory modifiers and quantitative trait determinants are preferentially maintained in a dosage-sensitive relationship (Birchler et al. 2001; Birchler and Veitia 2007). These duplicates are preserved due to their balance and selection against deleting one member that prevents their rapid loss (Birchler et al. 2005; Freeling and Thomas 2006). On the contrary, genes lacking an interacting balance relationship would be randomly deleted over evolutionary time, returning to the diploid level (Birchler and Veitia 2007). In an allopolyploid, genetic diploidization not only causes a reduction in gene expression which is comparable to the level of its diploid progenitors, but also promotes the divergence between subgenomes (e.g. bias fractionation) (Soltis et al. 1993; Soltis and Soltis 1999; Schnable et al. 2011; Schnable and Freeling 2011; Renny-Byfield et al. 2017). Genetic diploidization is also associated with neofunctionalization, subfunctionalization and genome downsizing (Leitch and Bennett 2004). This process would maintain the selected duplicate pairs in the evolutionary lineage (Birchler and Veitia 2010).
Polyploid speciation in Zea
To attempt to recreate the ancient allopolyploidization process of the Zea species, we propose the existence of two ancestral x = 5 populations (arbitrarily named AA and BB). The genomes of these ancestors would have diverged 20.5 Mya (Gaut and Doebley 1997). We suggest that they gave rise to various population systems at intermediate stages of divergence and reproductive isolation, connected by a reduced amount of interbreeding and gene flow. We propose that these populations would be considered semispecies sensu Grant (1985). These semispecies, which likely differ in genomic composition (A1A1, A2A2, A3A3, B1B1, B2B2, BxBx, ByBy, BzBz) would be the ancestors of all modern species of the genus Zea (Fig. 4).
Hypothetical model of the allopolyploid speciation in the genus Zea, based on cytogenetic studies. A and B ancestral subgenomes. Subscripts represent the variability of ancestral semispecies. Homology is higher between subgenomes bearing numerical subscripts than between subgenomes bearing letter subscripts. Hybrid and polyploid combinations resulting from different crosses are shown. Genomic formulae of extant species are highlighted in bold. The dashed line represents our alternative model for the origin of modern maize
We hypothetize that the evolutionary process that originated the different Zea species included: (a) hybridization between sympatric diploid ancestral semispecies (2n = 10) harbouring genomes with different levels of homoeology (A1B1, A2B2, A1Bx, A2A3, ByBz), and (b) recurrent duplication of the hybrids, giving rise to distinct auto- and allopolyploids (A1A1B1B1, A2A2B2B2, A1A1BxBx, A2A2A3A3, ByByBzBz; Fig. 4). Gaut and Doebley (1997) estimated that hybridization and polyploidy events in Zea occurred about 11.4 Mya. At this point, the merger and doubling of independent genomes induced changes in genome organization, gene expression and molecular interactions. In this regard, it is pertinent to mention the gene balance hypothesis stating that the stoichiometry of members of multisubunit complexes can affect the amount of functional complete product, which affects patterns of gene expression of the regulatory complex and ultimately, the phenotype and evolutionary fitness (Birchler et al. 2005; Birchler and Veitia 2007, 2010, 2012). Feldman and Levy (2012) added that revolutionary changes occur suddenly after hybridization and at early stages of allopolyploidy and are cumulative throughout the polyploid evolution. This genome remodelling could have led to genome downsizing, as reported for Z. perennis (González et al. 2006), or to the loss or inactivation of the rDNA loci towards a diploid-like number, as is the case for Nicotiana allopolyploids (Kovarik et al. 2008; Renny-Byfield et al. 2013). The latter phenomenon may have occurred in Zea because all species with 2n = 20 have a single chromosome pair bearing the 45 s rDNA sequence, while in Z. perennis it is present in two chromosome pairs, instead of the two and four pairs expected, respectively (G. E. González and L. Poggio, unpublished data). It is clear that the hypotheses proposed here, based on cytogenetic analyses, while not disagreeing with molecular studies, are speculative. Further whole-genome sequencing of Zea species, in which these data are not yet available, will be needed to support or modify them.
The hybrids between taxa of the Zea section showed regular meiosis, with 10II at prophase-metaphase I and high pollen viability, thus implying high pairing affinity between the parental species, which do not experience post-zygotic reproductive isolation (Fig. 1; Poggio et al. 2005; González and Poggio 2011). To hypothesize the origin of the modern taxa of the Zea section (Fig. 4), we postulate that Z. m. subsp. parviglumis and Z. m. subsp. mexicana would have arisen from a polyploid ancestor whose genomic formula would be A1A1B1B1. Genetic studies have suggested that maize was domesticated from Z. m. subsp. parviglumis between 12,000 and 9000 years before the present (Matsuoka et al. 2002). The cultivated maize would have spread throughout Mexico, reaching the highland about 6500 years before present, aided by adaptive introgression from Z. m. subsp. mexicana (Wang et al. 2017; Calfee et al. 2021; Tenaillon et al. 2023; Yang et al. 2023). Cytogenetical data also led us to speculate another hypothesis about the origin of maize, suggesting that the primitives Z. m. subsp. mexicana, Z. m. subsp. parviglumis and maize evolved from a common ancestor (A1A1B1B1). Modern maize would have been domesticated from the latter, thus implying that maize and its sister subspecies probably derived directly from the same ancestral polyploid (Fig. 4). This hypothesis appears plausible based on patterns of shared variants studied by Ross-Ibarra et al. (2009), which suggest that the subspecies of the Zea section arose nearly simultaneously, on the order of 100,000–300,000 years, and diverged 30–60,000 years ago. The studies of maize origin are hindered by the effects of hybridization and introgression. Recent whole-genome analyses uncovered a substantial role for Z. m. subsp. mexicana and Z. m. subsp. parviglumis in making the modern maize (Yang et al. 2023).
The hybrids of Z. luxurians and subspecies of Z. mays and Z. diploperennis showed 9II + 2I, heteromorphic bivalents and univalents of different size at prophase and metaphase I. Zea luxurians has the highest DNA and heterochromatin content, as well as more and larger heterochromatic knobs than other 2n = 20 Zea taxa (Tito et al. 1991; González et al. 2013). Therefore, the heteromorphic bivalents are likely the result of homoeologous pairing between the chromosomes of Z. luxurians and those of the other species involved in these crosses. The hybrids also showed up to three bridges and fragments at anaphase I, indicating that the parental species differ in at least three paracentric inversions. Such meiotic abnormalities may explain the high hybrid pollen sterility, revealing the postzygotic reproductive isolation between Z. luxurians and the remaining Zea taxa. The patterns of shared variants by Z. luxurians and Z. m. subsp. mays prompted Ross-Ibarra et al. (2009) to rule out any ancient divergence between these species. González and Poggio (2015), and Poggio and González (2018), studying the meiotic behaviour and the genomic affinities at repetitive sequences level of hybrids, suggested that AL and BL subgenomes from Z. luxurians are homoeologous to each other but somewhat different from those of the Zea section.
Hybrids between Z. diploperennis and taxa of the Zea section and Z. luxurians showed regular meiosis and low pollen fertility, indicating postzygotic reproductive isolation between the parental species. The absence of quadrivalents in colchicine-treated materials suggested a considerable divergence between the Ad and Bd genomes of Z. diploperennis (Molina et al. 2013; González and Poggio 2015). This meiotic behaviour led us to postulate that Z. diploperennis is a typical allopolyploid (Fig. 4).
Z. perennis showed 5IV + 10II at prophase I. Poggio and González (2018) indicated that this species is alloautooctoploid with four subgenomes somewhat divergent from one another, where the A (Ap and Ap´) subgenomes are homoeologous to each other and constitute the quadrivalents, while pairing between B (Bp1 and Bp2) subgenomes can only occur when the PrZ genes are inhibited. Colchicine-treated cells also showed high divergence between the A and B subgenomes of Z. perennis, as evidenced by the absence of octovalents (Poggio and González 2018; present work). The meiotic analyses of the 2n = 30 hybrids showed 5III + 5II + 5I (Fig. 1), and molecular cytogenetic studies revealed that the III were formed by the A (Ap and Ap´) subgenomes of Z. perennis and the 2n = 20 parental subgenome (Ax); the bivalents belonged to the homoeologous subgenomes (Bp1 and Bp2) of Z. perennis and the univalents belonged to the 2n = 20 parental subgenome (Bx) (Poggio et al. 1999; González et al. 2006; Poggio and González 2018). The fact that five chromosomes of the 2n = 20 parent form the trivalents suggests that these chromosomes retained homologous/homoeologous sequences of one of the Z. perennis ancestral subgenomes. It was concluded that Z. perennis might have resulted from the combination of an intervarietal autotetraploid (A subgenomes) and a segmental allopolyploid (B subgenomes) in the same nucleus. Hence, we speculate that modern Z. perennis would have originated from a cross between an autopolyploid (A2A2A3A3) and an allopolyploid (ByByBzBz). This would be followed by another round of polyploidy giving rise to the octoploid A2A2A3A3ByByBzBz from which modern Z. perennis might have derived (Fig. 4). Since the meiotic behaviour of the 2n = 30 hybrids showed homoeology between the A subgenomes of all the parental species (Poggio et al. 1999, 2005; González et al. 2006; present work), this subgenome could be considered pivotal (“pivotal genome” sensu Stebbins 1971). We postulate that the rDNA sequences were conserved in the pivotal subgenome A since in the 2n = 30 hybrids these sequences were detected by FISH in a trivalent formed by the A subgenomes from the parents (Poggio et al. 1999; González et al. 2006). We also suggest that the pairing regulator gene (PrZ) was provided by the subgenome A, since all the extant taxa show a PrZ locus (Poggio and González 2018). In the model proposed here, the ancestral semispecies with the pivotal subgenome A would have had a wide geographic distribution, co-occurring and hybridizing with the semispecies harbouring the B subgenomes, thus enabling sympatric speciation.
Data availability
All data generated or analyzed during this study are provided in this published article or it will be provided from the corresponding author upon a reasonable request.
Abbreviations
- Mya:
-
Million years ago
- PrZ:
-
Pairing regulator locus
References
Alexander MP (1969) Differential staining of aborted and nonaborted pollen. Stain Technol 44:117–122
Birchler JA, Veitia RA (2007) The gene balance hypothesis: from classical genetics to modern genomics. Plant Cell 19:395–402
Birchler JA, Veitia RA (2010) The gene balance hypothesis: implications for gene regulation, quantitative traits and evolution. New Phytol 186:54–62
Birchler JA, Veitia RA (2012) Gene balance hypothesis: connecting issues of dosage sensitivity across biological disciplines. Proc Natl Acad Sci USA 109:14746–14753
Birchler JA, Bhadra U, Bhadra MP, Auger DL (2001) Dosage-dependent gene regulation in multicellular eukaryotes: implications for dosage compensation, aneuploid syndromes, and quantitative traits. Dev Biol 234:275–288
Birchler JA, Riddle NC, Auger DL, Veitia RA (2005) Dosage balance in gene regulation: biological implications. Trends Genet 21:219–226
Blasio F, Prieto P, Pradillo M, Naranjo T (2022) Genomic and meiotic changes accompanying polyploidization. Plants 11:125. https://doi.org/10.3390/plants11010125
Calfee E, Gates D, Lorant A, Perkins MT, Coop G, Ross-Ibarra J (2021) Selective sorting of ancestral introgression in maize and teosinte along an elevational cline. PLoS Genet 17(10):e1009810. https://doi.org/10.1371/journal.pgen.1009810
Doebley JF (1990) Molecular systematics of Zea (Gramineae). Maydica 35:143–150
Feldman M, Levy AA (2012) Genome evolution due to allopolyploidization in wheat. Genetics 192:763–774
Freeling M, Thomas BC (2006) Gene-balanced duplications, like tetraploidy, provide predictable drive to increase morphological complexity. Annu Rev Plant Biol 60:433–453
Gale MD, Devos KM (1998) Comparative genetics in the grasses. Proc Natl Acad Sci USA 95:1971–1974
Gaut BS, Doebley JF (1997) DNA sequence evidence for the segmental allotetraploid origin of maize. Proc Natl Acad Sci USA 94:6809–6814
González GE, Poggio L (2011) Karyotype of Zea luxurians and Z. mays subsp. mays using FISH/DAPI and analysis of meiotic behavior of hybrids. Genome 54:26–32
González GE, Poggio L (2015) Genomic affinities revealed by GISH suggests intergenomic restructuring between parental genomes of the paleopolyploid genus Zea. Genome 58:433–439
González GE, Comas C, Confalonieri VA, Naranjo CA, Poggio L (2006) Genomic affinities between maize and Zea perennis using classical and molecular cytogenetic (GISH-FISH). Chrom Res 14:629–635
González GE, Fourastié MF, Poggio L (2013) Número y composición de secuencias de los knobs (DAPI-FISH) y su utilidad en la caracterización de accesiones de maíz y teocintle. Rev Fitotec Mex 36:127–135
Grant V (1985) The evolutionary process: a critical review of evolutionary theory. Columbia University Press, San Francisco
Jenkins G, Rees H (1991) Strategies of bivalent formation in allopolyploid plants. Proc R Soc Biol Sci 243:209–214
Kovarik A, Dadejova M, Lim UK, Chase MW, Clarkson JJ, Knapp S, Leitch AR (2008) Evolution of rDNA in Nicotiana allopolyploids: a potential link between rDNA homogenization and epigenetics. Ann Bot 101:815–823
Leitch IJ, Bennett MD (2004) Genome downsizing in polyploid plants. Biol J Linn Soc 82:651–663
Matsuoka Y, Vigouroux Y, Goodman M, Sánchez J, Buckler E, Doebley JF (2002) A single domestication for maize shown by multilocus microsatellite genotyping. Proc Natl Acad Sci USA 99:6080–6084
Molina MC, López CG, Staltari S, Chorzempa SE, Moreno Ferrero V (2013) Cryptic homoeology analysis in species and hybrids of genus Zea. Biol Plant 57:449–456
Moore G, Devos KM, Wang Z, Gale MD (1995) Cereal genome evolution. Grasses, line up and form a circle. Curr Biol 5:737–739
Naranjo CA, Molina MC, Poggio L (1990) Evidencias de un número básico x=5 en el género Zea y su importancia en estudios del origen del maíz. Academia Nacional de Ciencias Exactas, Físicas y Naturales de Buenos Aires 5:43–53
Poggio L, González GE (2018) Cytological diploidization of paleopolyploid genus Zea: divergence between homoeologous chromosomes or activity of pairing regulator genes? PLoS ONE 13(1):e0189644. https://doi.org/10.1371/journal.pone.0189644
Poggio L, Molina MC, Naranjo CA (1990) Cytogenetic studies in the genus Zea: colchicine-induced multivalents. Theor Appl Genet 79:461–464
Poggio L, Confalonieri VA, Comas C, González GE, Naranjo CA (1999) Genomic affinities among Zea luxurians, Zea perennis and Zea diploperennis: meiotic behaviour in the F1 and genomic in situ hybridization (GISH). Genome 42:993–1000
Poggio L, González GE, Confalonieri VA, Comas C, Naranjo CA (2005) The genome organization and diversification of maize and its allied species revisited: evidences from classical and FISH-GISH cytogenetics analysis. Cytogenet Genome Res 109:259–267
Renny-Byfield S, Kovarik A, Kelly LJ et al (2013) Diploidization and genome size change in allopolyploids is associated with differential dynamics of low- and high-copy sequences. Plant J 74:829–839
Renny-Byfield S, Rodgers-Melnick E, Ross-Ibarra J (2017) Gene fractionation and function in the ancient subgenomes of maize. Mol Biol Evol 34:1825–1832
Ross-Ibarra J, Tenaillon M, Gaut BS (2009) Historical divergence and gene flow in the genus Zea. Genetics 181:1399–1413
Schnable JC, Freeling M (2011) Genes identified by visible mutant phenotypes show increased bias toward one of two subgenomes of maize. PLoS ONE 6(3):e17855. https://doi.org/10.1371/journal.pone.0017855
Schnable JC, Springer NM, Freeling M (2011) Differentiation of the maize subgenomes by genome dominance and both ancient and ongoing gene loss. Proc Natl Acad Sci USA 108:4069–4074
Soares NR, Mollinari M, Oliveira GK, Pereira GS, Vieira MLC (2021) Meiosis in polyploids and implications for genetic mapping: a review. Genes 12:1517
Soltis DE, Soltis PS (1999) Polyploidy: recurrent formation and genome evolution. Trend Ecol Evol 14:48–352
Soltis DE, Soltis PS, Rieseberg LH (1993) Molecular data and the dynamic nature of polyploidy. Crit Rev Plant Sci 12:243–273
Stebbins GL (1971) Chromosomal evolution in higher plants. Edward Arnold Ltd., London
Swigoñová Z, Lai J, Ma J, Ramakrishna W, Llaca V, Bennetzen JL, Messing J (2004) On the tetraploid origin of the maize genome. Comp Funct Genomics 5:281–284
Sybenga J (1975) Meiotic configurations. Springer, Berlin, Heidelberg, New York
Sybenga J (1996) Chromosome pairing affinity and quadrivalent formation in polyploids. Do segmental allopolyploids exist? Genome 39:1176–1184
Sybenga J (1999) What makes homologous chromosomes find each other in meiosis? A review and a hypothesis. Chromosoma 108:209–219
Sybenga J, Schabbink E, van Eden J, de Jong JH (1994) Pachytene pairing and metaphase I configurations in a tetraploid somatic Lycopersicon esculentum × L. peruvianum hybrid. Genome 37:54–60
Tenaillon MI, Burban E, Huynh S et al (2023) Crop domestication as a step toward reproductive isolation. Am J Bot 110:e16173
Tito C, Poggio L, Naranjo CA (1991) Cytogenetics studies in the genus Zea: DNA content and heterochromatin in species and hybrids. Theor Appl Genet 83:58–64
Wang L, Beissinger TM, Lorant A, Ross-Ibarra C, Ross-Ibarra J, Hufford MB (2017) The interplay of demography and selection during maize domestication and expansion. Genome Biol 18:215
Yang N, Wang Y, Liu X et al (2023) Two teosintes made modern maize. Science 382:eadg8940
Zafar Iqbal M, Cheng M, Zhao Y et al (2018) Mysterious meiotic behavior of autopolyploid and allopolyploid maize. Comp Cytogen 12(2):247–265
Acknowledgements
The authors would like to thank the Comisión Nacional de Investigaciones Científicas y Técnicas-CONICET (PIP 2115CO) and Universidad de Buenos Aires (UBACYT-20020170100614BA).
Author information
Authors and Affiliations
Contributions
GG and LP conceived and designed research, conducted experiments, analyzed data and wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest associated with the article.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
González, G.E., Poggio, L. Polyploid speciation in Zea (Poaceae): cytogenetic insights. Planta 259, 67 (2024). https://doi.org/10.1007/s00425-024-04345-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-024-04345-x