Abstract
Main conclusion
EgPHI-1 is a member of PHI-1/EXO/EXL protein family. Its overexpression in tobacco resulted in changes in biomass partitioning, xylem fiber length, secondary cell wall thickening and composition, and lignification.
Abstract
Here, we report the functional characterization of a PHOSPHATE-INDUCED PROTEIN 1 homologue showing differential expression in xylem cells from Eucalyptus species of contrasting phenotypes for wood quality and growth traits. Our results indicated that this gene is a member of the PHI-1/EXO/EXL family. Analysis of the promoter cis-acting regulatory elements and expression responses to different treatments revealed that the Eucalyptus globulus PHI-1 (EgPHI-1) is transcriptionally regulated by auxin, cytokinin, wounding and drought. EgPHI-1 overexpression in transgenic tobacco changed the partitioning of biomass, favoring its allocation to shoots in detriment of roots. The stem of the transgenic plants showed longer xylem fibers and reduced cellulose content, while the leaf xylem had enhanced secondary cell wall thickness. UV microspectrophotometry of individual cell wall layers of fibers and vessels has shown that the transgenic plants exhibit differences in the lignification of S2 layer in both cell types. Taken together, the results suggest that EgPHI-1 mediates the elongation of secondary xylem fibers, secondary cell wall thickening and composition, and lignification, making it an attractive target for biotechnological applications in forestry and biofuel crops.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fibers, tracheary elements and vessels are specialized cell types that compose the wood or secondary xylem (Turner et al. 2007). Xylem cells form thick secondary cell walls (SCW) composed mainly by cellulose, hemicellulose and lignin (Boerjan et al. 2003). The deposition of these compounds on the SCW determines the dimensions of xylem cells, while the properties of these cells depend on the chemical and mechanical properties of SCW (Meents et al. 2018). SCW have multiple layers (S1–S2) that differ in microfibril organization and in the ratios of cellulose to matrix (lignin, hemicellulose, and pectin) components (Mellerowicz et al. 2008).
Lignification is the last step of SCW’s biosynthesis. Lignin accounts for about 30% of the biomass in woody tissues, conferring rigidity to SCW and being fundamental for structural support and impermeability required for water and nutrient transport over long distances (Leplé et al. 2007; Meents et al. 2018). Lignin is also essential in determining the quality of timber, mainly for pulp and paper industries. The presence of lignin is a limiting factor for pulp and paper production. It needs to be extracted from the wood by thermochemical methods that are expensive both economically and environmentally, reducing pulp yield (Peter et al. 2007).
The deposition of lignin in SCW requires transcriptional regulators to coordinate the expression of dozens of genes participating in this process (Andersson-Gunneras et al. 2006; Zhong et al. 2010). Several efforts have been dedicated to develop genetically engineered trees with emphasis on either reduced lignin levels or altered lignin composition to improve the efficiency of wood pulp production (Hu et al. 1999; Pilate et al. 2002; Quang et al. 2012; Nuoendagula et al. 2018). Although the roles of several genes involved in secondary xylem formation have been elucidated, our knowledge about SCW biosynthesis and its integration into plant metabolism and connection with wood quality is still fragmentary. Recent studies have used multi-omics approaches to obtain transcriptomic, proteomic and metabolomic data in attempts to improve and understand the interactions of stress response pathways and other complex processes related to the growth and production of biomass (Nakahama et al. 2018; Obudulu et al. 2018).
To gain a more in-depth knowledge, we have carried out a comparative analysis between the xylem transcriptomes of Eucalyptus grandis and E. globulus using high density microarrays (Pasquali et al. 2005). These two species display contrasting characteristics of growth and wood quality. E. grandis is a species native to tropical and subtropical regions that exhibits rapid growth but produces a low-density wood; whereas, E. globulus is a species from temperate climates that produces high basic wood density and good fibre and handsheet properties, but exhibits slow growth (Bernard 2003; Myburg et al. 2007).
One of the differentially expressed genes found in our previous study encodes a polypeptide with high similarity to PHOSPHATE-INDUCED PROTEIN 1 (PHI-1) from tobacco and to EXORDIUM (EXO) and EXORDIUM-like (EXL) proteins from Arabidopsis. This gene was observed to be 7.5 X more expressed in the xylem of E. globulus than in E. grandis. PHI-1/EXO/EXL comprises an emerging and widely distributed family of proteins involved in signaling pathways that control cell division and differentiation in response to hormonal and environmental signals (Sano et al. 1999; Sano and Nagata 2002; Farrar et al. 2003; Coll-Garcia et al. 2004; Schröder et al. 2009, 2011, 2012). PHI-1 was originally identified as a protein involved in the phosphate-induced cell cycle activity in cell cultures of tobacco (Sano et al. 1999; Sano and Nagata 2002). The gene identified by T-DNA mutagenesis in Arabidopsis and named EXO was found to encode a protein structurally related to PHI-1 of tobacco. EXO was preferentially abundant in embryo cells in division, apical meristems and young leaves, suggesting its role in the maintenance of meristematic cells (Farrar et al. 2003). The extracellular EXO protein of Arabidopsis was further identified as a mediator of cell expansion and shoot and root growth promoted by brassinosteroids (Coll-Garcia et al. 2004; Schröder et al. 2009). Later, the brassinosteroids-regulated EXL gene from Arabidopsis (EXL1) was shown to be required for adaptation to carbon- and energy-limiting growth conditions (Schröder et al. 2011, 2012; Lisso et al. 2013).
Although PHI-1/EXO/EXL does not show similarities to any protein with known function, they are a widely distributed class of proteins, present in coniferous, monocotyledonous and dicotyledonous plants, as well as in mosses and soil bacteria (Sano et al. 1999; Dellagi et al. 2000; Schröder et al. 2009). In higher plants, PHI-1 domain-encoding genes have been reported to be differentially expressed during secondary growth (Ko et al. 2004), in suspension cultures of dedifferentiated cells (Iwase et al. 2005), in response to fungal infection (Mustafa et al. 2009; Wu et al. 2010) and insect damage (De-Vos and Jander 2009), during growth in contaminated soils (Norton et al. 2008), in association with weeds (Dita et al. 2009), in response to drought stress (Fox et al. 2018), hypoxia and darkness (van Veen et al. 2016), Botrytis cinerea infection and abiotic stress treatments (Sham et al. 2014; Frey et al. 2020) and root treatment with mycorrhiza helper bacteria such as Streptomyces sp. (Kurth et al. 2015). EXL2 and EXL4 were proposed to act as components of a gene regulatory network in bud dormancy in woody and herbaceous species, potentially involved in sugar sensing (Tarancón et al. 2017). Determination of the EXL7 histone epigenetic marks in response to B. cinerea (Sham et al. 2014) revealed this gene as a potential biomarker for the early detection of this fungus in Arabidopsis and tomato plants (Crespo-Salvador et al. 2018).
In this study, we report the functional characterization of an Eucalyptus homologue of PHI-1, named Eucalyptus globulus PHI-1 (EgPHI-1). Our results show that EgPHI-1 overexpression in transgenic tobacco modifies plant growth, biomass partitioning and the SCW properties. EgPHI-1 is the first reported member of the PHI-1/EXO/EXL family associated with secondary xylem differentiation and cell wall properties in plants.
Materials and methods
Plant material
Wild-type (WT) and three EgPHI-1-overexpressing transgenic tobacco lines (L1, L2 and L3) were used in this study. The EgPHI-1 transgenic lines represent distinct events of Agrobacterium-mediated genetic transformation that were previously generated (Sousa et al. 2014) and selected on the basis of their increased EgPHI-1 mRNA expression and protein levels as compared to the WT plants (Suppl. Fig. S1). Ninety- or 60-day-old WT and transgenic (T2 generation) plants, cultivated in 20-L pots filled with soil and washed sand (2:1 ratio) under standardized greenhouse conditions (70–80% relative humidity and 25–30 o C) and cultivation practices, were used in the analyses.
Hormonal and stress-inducing treatments were applied to 4-month-old E. grandis plants grown in vitro in Murashige and Skoog medium (MS, Invitrogen) solidified with 0.7% (w/v) Phytoagar (Duchefa, Haarlem, The Netherlands) at 26 °C and a photoperiod of 16:8 h light:dark. Groups of 3–4 plantlets were separately sprayed three times with 0.2 mg L−1 kinetin (Sigma-Aldrich, St. Louis, MO, USA), 2 mg L−1 1-naphthaleneacetic acid (NAA; Sigma-Aldrich) or water as a control. Other groups of 3–4 plantlets were subjected to drought stress or mechanical wounding by, respectively, leaving the culture flasks opened overnight in the growth chamber or clenching leaves and stems with a flat forceps followed by water sprays and overnight incubation in the growth chamber. All plants were harvested 24 h after treatments, immediately frozen and stored in liquid nitrogen. The control group of plants was frozen in liquid nitrogen without any treatment.
EgPHI-1 characteristics and phylogenetic analysis
The Expasy platform tools (http://expasy.org/tools/) were used to calculate/deduce the different nucleotide and amino acid sequence properties of EgPHI-1. The 1.5-kb 5′-upstream promoter region of EgPHI-1 (locus Eucgr.F00356), available in the Eucalyptus reference genome database (http://www.phytozome.net/eucalyptus.php), was used to analyze putative regulatory cis-elements, employing the PlantCare software (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/), and predicted transcription factors binding sites using the Plant Promoter Analysis Navigator (PlantPAN; http://PlantPAN.itps.ncku.edu.tw/). Comparative analyses were performed between the EgPHI-1-deduced protein and 43 amino acid sequences with the PHI-1/EXO conserved domain (Schröder et al. 2009) by multiple sequence alignment using the default parameters of ClustalW (Thompson et al. 1994). Phylogenetic analysis was performed using the maximum likelihood method based on the JTT matrix-based model (Jones et al. 1992) of MEGA5 software (Tamura et al. 2011). The tree with the highest log likelihood was chosen and shown. The bootstrap test was conducted with 1,000 replicates and statistical values were presented as percentages in the branches.
RNA isolation and reverse transcription-quantitative PCR (RT-qPCR)
Total RNA was isolated from E. grandis seedlings using the PureLinkTM Plant RNA Reagent (Invitrogen), according to the manufacturer’s instructions. The purity and concentration of total RNA samples were determined spectrophotometrically and by agarose gel electrophoresis. Total RNA samples were treated with RNase-free DNase I (Fermentas) to remove any contaminating genomic DNA. Total RNA treated with DNase I was used in cDNA synthesis with the M-MLV Reverse Transcriptase (Promega, Madison, WI, USA), according to manufacturer’s instructions. The steady-state EgPHI-1 mRNA levels were estimated using the ABI 7500 Real-Time PCR System (Applied Biosystems, Foster City, CA, USA) and the SYBR Green PCR Master Mix (Applied Biosystems), according to manufacturer’s recommendations. Quantitative, real-time PCRs were performed using three biological and three technical replicates. The gene-encoding ribosomal protein L23A (RibL23A) (de Oliveira et al. 2012) was used as internal reference gene for normalizing EgPHI-1 expression. A negative control without cDNA template was included for each primer pair. Primer sequences employed for RibL23A were 5′-AAGGACCCTGAAGAAGGACA-3′and 5′-CCTCAATCTTCTTCATCGCA-3′, and for EgPHI-1 were 5′-GCTCTGACCAGTTTCCGAAAA-3′ and 5′CCGGAACAGAGGTTAGGTAAGCT-3′. The relative expression values were calculated using the 2-∆∆Ct method (Livak et al. 2001).
Analysis of plant growth
Plant height, leaf number per plant, leaf area, root length and root system volume were analyzed in WT and transgenic tobacco plants grown for 90 days under greenhouse conditions. Total leaf area was determined using the LI-3100 area meter (Li-Cor). Root volume was measured using the intact root system to displace the water column in a graduated cylinder. Leaves, stems and roots were separated and kept in an air-circulation stove at 75 °C until a constant weight was obtained. These materials were used to determine the dry biomass of the different plant parts.
Xylem anatomy
Stem samples (third internode from the apex) were collected from 90-day-old WT and transgenic tobacco plants and fixed in 70% ethanol for anatomical analysis of the xylem. Free-hand stem cross sections were cleared and stained with 1% (v/v) astra blue and 1% (v/v) safranin. Stem samples were also used to obtain isolated fibers. These samples were maintained in a solution of glacial acetic acid and hydrogen peroxide (1:1, v/v) for 48 h at 60 °C. The resulting fibrous complex was washed and stained with 1% (v/v) safranin. Microscopic slides were prepared with the double staining cross sections, as well as with isolated staining fibers. Slides were viewed under a light microscope (Olympus - CX41) and photographed with a digital camera (Olympus - C7070) attached to the equipment. Xylem anatomy was also observed at cross sections of the base of the petioles (third fully expanded leaves from the apex). Lignin autofluorescence of vessels was detected using an epifluorescence microscope (Leica DMRA2) equipped with a camera (Leica MPS60) at 340–380-nm excitation wavelength and 400-nm barrier filters. Images were analyzed employing the ImageJ 1.44 software (http://imagej.nih.gov.ij) and 135 measurements of fiber length were performed. Three biological replicates (plants) were used, each containing three slides.
Determination of stem chemical composition
Stems of 60-day-old WT and transgenic plants were harvested and kept in an air-circulation stove at 75 °C until a constant weight was obtained. Three grams of dry stems was milled to pass through a 0.84-mm screen. Milled samples were extracted with 95% ethanol for 6 h in a Soxhlet apparatus. The percentage of extractives was determined on the basis of the dry weight of the extracted and non-extracted milled samples. The hydrolysis of ethanol-extracted samples was performed with 72% sulfuric acid at 30 °C for 1 h as previously described (Ferraz et al. 2000). The resulting acid solutions were diluted with water and the mixture was heated at 121 °C/1 atm for 1 h. Thereafter, residual materials were cooled and filtered (glass filter number 3). Solids were dried to a constant weight at 105 °C and determined as insoluble lignin. The absorbance of the filtrated at 205 nm was used to determine soluble lignin concentration. Monomeric sugar concentrations in the soluble fraction were determined by HPLC (HPX87H column; Bio-Rad, Hercules, CA, USA) at 45 °C and an elution rate of 0.6 mL min−1 with 5 mmol L−1 sulfuric acid. Temperature-controlled refractive index detector at 35 °C was used to detect sugars. Xylose, mannose and galactose were eluted at the same time under these conditions, and appeared as single peaks. Glucose, xylose, arabinose and acetic acid were used as external calibration standards. No corrections were performed because of sugar-degradation reactions that take place during acid hydrolysis. The factors used to convert sugar monomers to anhydromonomers were 0.90 for glucose to glucan and 0.88 for xylose to xylan and arabinose to arabinan. Acetyl content was calculated as the acetic acid content multiplied by 0.72. This procedure was conducted in triplicate. Glucose was reported as cellulose after correction by the hydrolysis factor, and the other sugars and acetic acid were used to calculate the hemicellulose content in samples.
Sample preparations and cellular scanning by UV-microspectrophotometry
Samples of the stem base taken from 60-day-old WT and transgenic plants were dissected from the xylem of each sample. Small blocks (approximately 1 × 1 × 5 mm) were dehydrated in a graded series of acetone and embedded in Spurr’s (Spurr 1969) epoxy resin formulated to the standard hardness of the blocks (Spurr Low-Viscosity Embedding Kit, EM0300, Sigma-Aldrich). Sections (1 µm) were prepared from the samples with a LEICA EM-UC7 ultramicrotome fitted with a diamond knife (4 mm-Histo, Diatome, Biel, Switzerland). Sections were transferred to quartz microscope slides. UV-microspectrophotometry analyses were performed with a ZEISS Axio Imager/J&M microspectrophotometer (Zeiss, Jena, Germany/J&M Analytik, Essingen, Germany) equipped with a scanning stage, allowing the determination of image profiles. The S2 layers of SCW from fibers and vessels in stem xylem of WT and transgenic plants were analyzed by photometric point measurements (1 μm2 spot size) in a wavelength range between 240 and 400 nm using the TIDASDAQ software (J&M Analytik). For the assessment of UV absorbance spectra, 20 and 10 point measurements from three biological replicates of each plant were recorded for fibers and vessels, respectively. Individual fibers of WT and L2 were selected and scanned for the UV-image profiles at constant wavelength of 280 nm (absorbance maximum of lignin). UV-image profiles showed the lignin distribution in SCW of selected fibers of stem xylem. The scan program digitizes rectangular fields with a local geometrical resolution of 1 μm2. A 255 color scale varying from light green to red was used to represent the range of 0.0–0.5 absorbance units (4-D Map software from J&M Analytik). The colors in the images are proportional to the absorption intensities, ranging from light green to red for low to intense absorptions, respectively. Scans were depicted, without repetitions or statistics, as histograms for semi-quantitative evaluation of lignin occurrence into cell layers. The histograms illustrate the pixel frequency of images at a 1-µm2 geometrical resolution.
Statistical analysis
Statistical significance between WT and transgenic plants was determined by analysis of variance (ANOVA) followed by Student’s t test. A probability (P) of 0.05 or less was taken to indicate statistical significance. Results are expressed as mean ± SE for (n) experiments.
Results
Identification and sequence analysis of EgPHI-1
The EgPHI-1 cDNA sequence characterized in this study was 100% identical to coding sequence of the locus Eucgr.F00356 in the E. grandis genome sequence database (http://www.phytozome.net/eucalyptus.php). Therefore, we assumed that EgPHI-1 corresponds to this locus in the reference genome sequence available for Eucalyptus spp. EgPHI-1 is 1,234 bp in length and contains an open reading frame (ORF) of 951 bp encoding a deduced protein of 316 amino acid residues with a theoretical isoelectric point of 8.06 and a calculated molecular weight of 33.1 kDa. A signal peptide possibly involved in the secretory pathway targeting to the endoplasmic reticulum was identified in the N-terminal portion of the EgPHI-1-deduced amino acid sequence, with a probable cleavage site between residues G27 and S28 (Fig. 1a). Potential sites of glycosylation at T257 and T264, as well as potential phosphorylation sites at serine (S57, S70, S120, S147 and S310), threonine (T65, T84 and T213) and tyrosine phosphorylation (Y248) were identified in the deduced amino acid sequence of EgPHI-1 along with potential protein kinase C phosphorylation sites (PKC, T104) (Fig. 1a).
Characterization of the EgPHI-1 sequence and expression. a Nucleotide sequence of the EgPHI-1 coding sequence (top line, left numbered) and deduced amino acid residues (bottom line, right numbered). Sequence underlined in blue represents a putative signal peptide. Red diamond indicates the probable cleavage point of the signal peptide. Sequences underlined in black, orange and green represent the possible phosphorylation sites for serine, threonine and tyrosine, respectively. A protein kinase C (PKC) putative phosphorylation site is indicated by a purple square. Amino acid residues in red represent possible glycosylation sites. The sequence highlighted in gray represents the PHI-1 functional domain (Pfam04674). b Cis-acting regulatory elements and motifs in the EgPHI-1 promoter region. Potential regulatory sequences identified in the 1.5-kb 5′-upstream promoter region of EgPHI-1 are represented as follows: closed circles, Sp1; black box, GAG-motif; gray arrow to right, TCT-motif; solar circle, circadian; open boxes, I-Box; inverted black triangles, G-Box; open diamond, box II; open triangles, ABRE; gray diamonds, ARE; black triangle, GA-motif; gray triangle, HSE; black arrow to left, MBS; open diamond, ARF; gray arrow to left, TCA-element; black arrows to right, MRE; gray boxes, GATA-motif; open circle, ARR-B; gray circles, as-2-box; closed diamond, TATA. c Expression of EgPHI-1 in E. grandis seedlings subjected to different treatments. Total RNA was isolated from E. grandis seedlings treated (wound, NAA, kinetin and drought) and untreated. Relative gene expression was assessed by RT-qPCR in three biological replicates and three technical replicates per plant. Expression was calculated by the 2-∆∆Ct method and values were normalized against RibL23A. Bars represent the mean ± SE of three replicates (n = 3). Statistically significant differences at P ≤ 0.05 (*), P ≤ 0.01 (**) and P ≤ 0.001 (***) between treated and untreated E. grandis seedlings are indicated
Promoter analysis and expression of EgPHI-1 in Eucalyptus grandis
Analysis of the 1.5 kb 5′-upstream promoter region of EgPHI-1 indicated the presence of several putative cis-acting regulatory elements (Fig. 1b). They include those involved in the response to light, such as Sp1 [CC(G/A)CCC], GAG-motif [AGAGATG], Box II [ACACGTTGT], I-box [GATATGG/CCTTATCCT], TCT-motif [TCTTAC], GA-motif [ATAGATAA], GATA-motif [(AAG)GATA(GGA/AGG)] and G-box [(TAA/C)ACGT(G/T)]. In addition, two as-2-box cis-elements [GATAATGATG] involved in shoot-specific expression were also identified. The promoter region of EgPHI-1 also has one element that regulates the circadian rhythm (Circadian [CAANNNNATC]), two binding sites of MYB (MRE [AACCTAA] and MBS [CGGTCA]), two response elements to abscisic acid (ABA; ABRE [(GCA/C)ACGTG(TC)]), three response elements to auxin (ARF [TGTCG]), one response element to cytokinin (ARR-B [GGGCATG]) and two response elements to methyl jasmonate (MeJA; CGTCA-motif [CGTCA] and TGACG-motif [TGACG]). Other cis-acting regulatory elements found in our analysis were as follows: two ARE elements [TGGTTT], which are essential for anaerobic induction; one HSE element [AGAAAATTCG], which responds to heat stress; and one TCA-element [TCAGAAGAGG], which is important for the responses induced by salicylic acid (Fig. 1b). The biological significance of the following identified motifs as transcription factor-binding sites (TFBS) was confirmed with the PlantPAN database: Box II, I-box, G-box, MYB, ABRE, ARF, ARR-B and CGTCA-motif.
To examine some of the conditions that affect EgPHI-1 expression, E. grandis seedlings were grown in vitro and subjected to four different treatments. Expression analysis showed that EgPHI-1 transcription was induced by mechanical wounding, NAA, kinetin and drought (Fig. 1c). E. grandis seedlings treated with NAA showed the highest EgPHI-1 expression compared with other treatments. EgPHI-1 expression in the kinetin-, mechanical wounding- and drought-treated seedlings was nearly twice as high as that of untreated seedlings (P ≤ 0.05; Fig. 1c).
EgPHI-1 protein properties and phylogenetic analysis
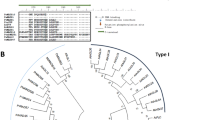
The major structural feature of EgPHI-1 is a region of 260-amino acid residues (Y44–S314) harboring the PHI-1 domain [Pfam 04674] (Fig. 1a). Alignment of EgPHI-1 and 43 other previously described (Schröder et al. 2009) homologous amino acid sequences indicated that EgPHI-1 belongs to a large orthologous group of proteins that includes the previously characterized tobacco PHI-1 and Arabidopsis EXO, EXL1, EXL2 and EXL4 (Sano et al. 1999; Farrar et al. 2003) (Fig. 2a; Suppl. Fig. S2). EgPHI-1 grouped into a specific sub-clade containing proteins from Oryza sativa and Vitis vinifera (Fig. 2a). EgPHI-1 shares 64% similarity with AAM08535 of O. sativa and 75% similarity with CAO61694 of V. vinifera. A high degree of conservation among these sequences is visualized in Fig. 2b.
Phylogenetic relationships and primary structure conservation of EgPHI-1. a Phylogenetic tree based on EgPHI-1 deduced amino acid sequence and other PHI-1/EXO/EXL homologous sequences previously described (Schröder et al. 2009) and represented as following: ARATH (Arabidopsis thaliana), POPTR (Populus trichocarpa), TOBAC (Nicotiana tabacum), ORYSA (Oryza sativa Japonica Group), SOLTU (Solanum tuberosum), VITVI (Vitis vinifera), POPTR (Populus trichocarpa x Populus deltóides), PICSI (Picea sitchensis), PENCI (Cenchrus ciliaris), PHYPA (Physcomitrium patens), SORBI (Sorghum bicolor) and SOLUS (Candidatus Solibacter usitatus). The EgPHI-1 sequence is marked by a green diamond. Tobacco PHI-1 sequence is marked by a red diamond. EXO/EXL sequences are marked by blue diamonds. The numbers on the branches indicate bootstrap values (%). The black dot indicates the mid-point rooting. b Conservation between EgPHI-1 amino acid residues and the two nearest amino acid sequences (O. sativa AAM08535 and V. vinifera CAO61694). Minimum and maximum conservation between amino acid residues are represented by color range from brown to yellow and by numbers, respectively. *Indicates omissions to regions with low conservation. Lateral numbers represent the size of the aligned fragment. Non-colored amino acid residues represent variations in the sequence
EgPHI-1 overexpression changes growth and biomass partitioning in transgenic plants
The EgPHI-1-overexpressing transgenic plants were, in general, taller (Fig. 3a) and had more leaves (Fig. 3b), with smaller leaf areas (Fig. 3c), than the WT plants. EgPHI-1 overexpression also caused significant changes in the allocation of biomass. All transgenic plants showed a significant increase in leaf biomass (Fig. 3d), while only L3 exhibited a significant increase (P ≤ 0.05) in stem biomass (Fig. 3e). On the other hand, the root system biomass was significantly reduced in all transgenic plants (Fig. 3f), while the root length was not affected, except for L1 (Fig. 3g). Conversely, the root systems of all transgenic plants showed a higher volume than WT roots (Fig. 3h). Significant changes were observed in the proportion of dry biomass among the different organs of transgenic plants (Fig. 3i), although their total biomass did not show significant variation (P ≤ 0.05) when compared to WT plants (data not shown).
Growth of transgenic tobacco expressing EgPHI-1. Transgenic (L1, L2 and L3) and WT plants grown for 90 days under greenhouse conditions were evaluated as to height (a), leaf number per plant (b), specific leaf area (c), leaf dry biomass (d), stem dry biomass (e), root system dry biomass (f), root length (g), root system volume (h) and total dry biomass (i). Bars represent the mean ± SE of three replicates (n = 3). Statistically significant differences at P ≤ 0.05 (*), P ≤ 0.01 (**) and P ≤ 0.001 (***) between transgenic and WT plants are indicated
EgPHI-1 overexpression changes the anatomy of stem and leaf xylem
Cross sections of the third internode from the apex of WT and transgenic plants were prepared and analyzed under light microscopy. No consistent changes in the anatomy of the secondary stem xylem were observed by analyzing the double-stained images (Fig. 4a; Suppl. Fig. S3). However, EgPHI-1 overexpression produced significant changes in the morphology of stem fibers, as it increased the fiber length in all transgenic plants (Fig. 4b).
Stem anatomy of transgenic tobacco expressing EgPHI-1. a Stem cross section of transgenic (L1, L2 and L3) and WT plants grown for 90 days under greenhouse conditions. The cross sections were stained with 1% (v/v) astra blue and 1% (v/v) safranin, specific dyes for cellulose and lignin, respectively. The double staining cross sections were viewed under a light microscope. VC, vascular cambium; V, vessels; X, xylem; F, fibers; P, phloem. Magnification bars represent 100 µm. b Fiber length. Fibers were obtained from stem samples (third internode from the apex) of plants grown for 90 days in a greenhouse by the chemical maceration method. Macerated material was washed and stained with 1% (v/v) safranin. Fiber slides were viewed under a light microscope. Bars represent the mean ± SE of the three technical replicates (n = 135). Bars represent the mean ± SE of the three replicates (n = 30). Statistically significant difference at P ≤ 0.01 (**) and P ≤ 0.001 (***) between the transgenic and WT plants are indicated
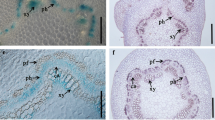
Cross sections of the base of the petioles of WT and transgenic plants were observed by epifluorescence microscopy to evidence the lignified structures in leaf xylem. In general, the lignified xylem of transgenic lines was more organized and closer to each other, resulting in a thicker and more compact structure (Fig. 5a). Observations of electronically modified images (black and white color) to highlight lignified areas (blue fluorescence) demonstrated an increased intensity of fluorescence in all transgenic plants (Fig. 5b).
Lignin autofluorescence in vessels of transgenic tobacco expressing EgPHI-1. Lignin autofluorescence in vessels of the base of petioles of transgenic (L1, L2 and L3) and WT plants. Cross sections were performed at the petioles of the third fully expanded leaf from the apex of plants grown for 90 days under greenhouse conditions. The autofluorescence was detected using 340–380-nm excitation wavelength and 400-nm barrier filters. a To highlight lignified areas (blue fluorescence), images were electronically modified (black and white color). b Magnification bars represent 100 µm
EgPHI-1 overexpression reduces the cellulose content in stems
WT and transgenic plants were evaluated according to their overall lignin, cellulose and hemicellulose contents in stems. All transgenic lines showed a significant reduction in cellulose content as compared with WT plants (Table 1). The percentages of decrease in cellulose contents were of ~ 4.7%, 5% and 2.5%, respectively, for L1, L2 and L3 as compared with WT. Significant alterations in lignin content were not observed in most transgenic lines, except for L2 (Table 1).
EgPHI-1 overexpression hardly affected the relative proportions of the hemicellulose components (Suppl. Table S1). A slight increase in the content of xylan was observed only for L1, while only L2 showed a minor decrease in the levels of this component that was apparently compensated by a proportional increase in the levels of arabinosyl. Conversely, the contents of extractives were significantly affected in the transgenic plants, with two (L2 and L3) out of the three transgenic lines analyzed showing a significant increase in comparison with WT (Suppl. Fig. S4).
EgPHI-1 overexpression changes lignification of individual cell wall layers of fibers and vessels
UV absorbance point spectra analysis was used to evaluate the lignin content in SCW layers of fibers and vessels present in the xylem. Profiles of lignified S2-layers of fibers (Fig. 6a) and vessels (Fig. 6b) were generated for WT and transgenic plants. The S2-layers of fibers and vessels from all samples presented typical and similar UV spectra of lignin moieties with a major band close at 280 nm (Fig. 6a and b, respectively). Fibers and vessels of L2 and L3 showed significantly lower UV absorptions at 280 nm as compared with the WT (Fig. 6c and d, respectively). Conversely, fibers of L1 and the WT exhibited similar UV absorbance at 280 nm (Fig. 6a, c); whereas, the vessels of L1 displayed a significantly higher UV absorbance at 280 nm than the WT (Fig. 6b, d). These data indicate that the EgPHI-1 overexpression changes the lignification of individual cell wall S2 of fibers and vessels.
UV absorbance spectra of the lignified cell walls of fibers (a) and vessels (b) of WT and transgenic (L1, L2 and L3) tobacco plants expressing EgPHI-1. At least 20 spectra were recorded from fibers and 10 from vessels, from three biological replicates of each WT and transgenic line. The spectra average is shown in this figure. Standard deviations, calculated from the absorbance values measured at wavelength of 280 nm, were 5% and 4% for fiber and vessel cells, respectively. Statistically significant differences at P ≤ 0.05 (*) and P ≤ 0.01 (**) between the transgenic and WT, at the wavelength of 280 nm, are indicated to fibers (c) e vessels (d)
Topochemical distribution of lignin in fibers
UV scanning profiles of lignin distributions in fibers were compared between WT (Fig. 7a) and L2 (Fig. 7b) transgenic plants. The scanning UV images of these cells indicated that WT and L2 fibers exhibited different absorbance levels, with the most intense absorbance seen in cell corners and compound middle lamella for both plants (Figs. 7c, d). The S2 layer of WT fiber showed a more uniform lignin distribution, with higher absorbance intensity (Fig. 7c, e), differing from a diffuse lignin distribution in the S2 layers of L2 (Fig. 7d, f). The average absorbance values for each image were calculated from the frequency histograms. The fibers of WT and L2 showed average absorptions at 280 nm of 0.20 and 0.16, respectively (Figs. 7e, f, respectively).
Fibers of stem xylem of WT and transgenic tobacco plants expressing EgPHI-1. Fiber selected from a control WT plant (a) and an L2 transgenic line (b). UV scanning profiles of lignin distribution in individual fibers of stem xylem of WT (c) and L2 transgenic plants (d). Histograms of lignin distribution in fibers of stem xylem of WT (e) and L2 transgenic plants (f) according to absorbance intensity values at 280 nm. Red boxes illustrate the spot size (l μm2) used to record one of the UV spectra presented in Fig. 6. The differently colored pixels (green to red) mark the absorbance intensity at 280 nm from 0.0 for light green to 0.5 for dark red. Magnification bars represent 10 µm. CC, cell corners; CML, compound middle lamella; S2, layer of secondary cell wall
Discussion
EgPHI-1 was shown to encode a predicted amino acid sequence with homology to previously described PHI-1/EXO/EXL proteins (Sano et al. 1999; Sano and Nagata 2002; Farrar et al. 2003; Coll-Garcia et al. 2004; Schröder et al. 2009, 2011, 2012). Analysis of its deduced amino acid sequence indicated the absence of hydrophobic regions and the presence of a signal peptide for secretion (Fig. 1a), suggesting that EgPHI-1 is a soluble extracellular protein, possibly localized in the apoplast. Arabidopsis EXO:GFP and EXO:HA fusion proteins were detected in the apoplast and cell wall, respectively, in accordance with the predictions of sequence analysis tools such as TargetP and Predotar (Schröder et al. 2009). Proteomic analyses have also identified EXO, EXL1, EXL2, EXL3, and EXL4 proteins as part of the cell wall proteome (Borderies et al. 2003; Bayer et al. 2006; Feiz et al. 2006; Jamet et al. 2006). Moreover, EXL2 expression was detected in both xylem and phloem of Arabidopsis (Tarancón et al. 2017). The different post-translational modification sites, including the predicted glycosylation and phosphorylation sites (Fig. 1a), suggest that EgPHI-1 activity may be regulated by phosphorylation/dephosphorylation promoted by kinases and specific phosphatases, as reported for the tobacco PHI-1 (Sano et al. 1999).
The identification of known cis-acting regulatory elements in the promoter region of EgPHI-1 (Fig. 1b) provided important clues about its regulation and function. These results suggest that EgPHI-1 is involved in ABRE-dependent ABA perception and signaling (Xiong and Zhu 2003), as reported for tobacco PHI-1 (Sano and Nagata 2002) and Arabidopsis EXO (Lisso et al. 2013) and EXL2 (Tarancón et al. 2017), and auxin and cytokinin signaling pathways via ARF (Roosjen et al. 2018) and ARR-B (Argyros et al. 2008), respectively. The presence of MBS and MRE also suggests that EgPHI-1 is regulated by transcription factors of the MYB family (Dubos et al. 2010), which have been implicated in regulating the secondary growth in Arabidopsis and the lignification process and flavonoid biosynthesis in Antirrhinum and other plant species (Tamagnone et al. 1998; Goicoechea et al. 2005; Zhong et al. 2008; Nakano et al. 2015; Guo et al. 2017). Other cis-acting regulatory elements identified suggest that the expression of EgPHI-1 may be under light and circadian control (Saibo et al. 2009) and induced under anaerobic (Shinozaki et al. 2003) and heat conditions (Haralampidis et al. 2002), injuries caused by insects and fungal pathogens (De-Vos and Jander 2009; Wang et al. 2011) and salicylic acid (Ohtake et al. 2000). The presence of cis-elements regulated by light, circadian cycle and anaerobic conditions was also reported in the promoter region of EXL1, which was found to be expressed especially under conditions of low irradiance, prolonged darkness and anoxia (Schröder et al. 2011, 2012).
EgPHI-1 expression was upregulated by NAA, kinetin, wound and drought (Fig. 1c). These results are consistent with the presence of the cis-elements ARF, ARR-B and ABRE found in its promoter region (Fig. 1b). Auxin and cytokinin are considered essential for early xylem differentiation during both normal development and wounding (Ye 2002; Pesquet et al. 2005; Turner et al. 2007). Wounding treatment further confirmed that EgPHI-1 is a wound-inducible gene. Upregulation of EgPHI-1 by drought also suggests its involvement in mechanisms of abiotic stress response and tolerance, likely via an ABRE-dependent signaling pathway (Yamaguchi-Shinozaki and Shinozaki 2006; Gómez-Porras et al. 2007; Fujita et al. 2011). Such hypothesis has been further confirmed by Sousa et al. (2014), using the same EgPHI-1-overexpressing transgenic tobacco lines, which exhibited enhanced tolerance to osmotic stresses induced by NaCl, PEG and mannitol. Besides, the induction of BiP gene expression observed in the EgPHI-1-overexpressing transgenic plants also evidences that this gene may modulate plant stress response via the unfolded protein response (UPR) (Sousa et al. 2014).
The PHI-1 domain present in EgPHI-1 is a long and structurally conserved region that identifies the PHI-1/EXO/EXL protein family and whose function and mechanism of action have not been clarified yet. However, members of this family have been implicated in processes of cell expansion and proliferation (Farrar et al. 2003; Schröder et al. 2009, 2011, 2012). EgPHI-1 has significant similarity to tobacco PHI-1 (Sano and Nagata 2002) and Arabidopsis EXO/EXL (Schröder et al. 2009), and our phylogenetic analysis showed that it belongs to the large orthologous group of proteins that includes these characterized tobacco and Arabidopsis PHI-1/EXO/EXL proteins and close orthologues of grape and rice (Fig. 2a). Thus, EgPHI-1 represents an Eucalyptus member of PHI-1/EXO/EXL protein family with functions or mode of action that may be similar, at least in part, from the previously characterized members of this family.
EgPHI-1 expression in tobacco promoted significant changes in the growth of transgenic plants, which included alterations in height and leaf number, area and biomass, and root system volume and biomass (Fig. 3). Although we are not able to rule out completely that uidA gene expression may at least partially account for these observed phenotypes, since endogenous plant beta-glucuronidase (GUS) expression and activity have been associated with cell growth (Sudan et al. 2006; Eudes et al. 2008), to our knowledge and experience, this seems to be unlikely because we have not found any evidence of such an association between cell growth and uidA expression in the literature, nor was it observed in any of our other experiments with transgenic plants overexpressing uidA. Our results are consistent with the previously reported role of PHI-1/EXO/EXL in controlling cell division, differentiation and expansion and shoot and root growth (Sano et al. 1999; Sano and Nagata 2002; Farrar et al. 2003; Coll-Garcia et al. 2004; Schröder et al. 2009).
Measurements of dry weight indicated that EgPHI-1 expression changed the biomass partitioning (Fig. 3i), which was preferentially allocated to shoots at the expense of root allocation. These results contrast with those observed for EXO of Arabidopsis, in which its overexpression promoted both shoot and root growth (Schröder et al. 2009). EgPHI-1 expression enhanced leaf biomass and number of leaves per plant, but reduced leaf area, suggesting an increase in leaf density. This is further supported by the thicker xylem SCWs observed in leaves of the transgenic plants (Fig. 5). The higher root system volume exhibited by transgenic plants also indicates the presence of thicker roots.
The observed changes in the anatomy of stem and leaf xylem (Figs. 4b and 5) caused by EgPHI-1 expression indicate an involvement of this gene in xylem fiber length and SCW properties. Long fibers are desirable in the production of strong paper. This is the first report describing the involvement of a member of the PHI-1/EXO/EXL protein family in such processes. However, the expression of genes encoding cell wall-modifying proteins such as KCS1, Exp5, AGP4, δ-TIP and WAK-1 was reported to be EXO dependent (Coll-Garcia et al. 2004; Schröder et al. 2009). It is believed that the EXO protein promotes cell expansion via the modulation of the expression of this set of genes (Farrar et al. 2003; Coll-Garcia et al. 2004). KCS1 is a protein involved in wax biosynthesis (Todd et al. 1999), while Exp5 is an expansin involved in cell wall loosening and break of non-covalent bonding between cellulose and hemicellulose (Cosgrove 2005). AGP4 is an extracellular arabinogalactan protein that has been implicated in many plant growth and developmental processes (Steinmacher et al. 2012). On the other hand, δ-TIP protein is a plant aquaporin that changes the membrane water permeability (Forrest and Bhave 2008). In turn, WAK1 is a transmembrane protein containing a cytoplasmic Ser/Thr kinase domain and an extracellular domain that interacts with cell wall pectins, playing a role in development and cell expansion (Decreux and Messiaen 2005). The relationship between EgPHI-1 and these cell wall-modifying proteins, if any, remains to be examined in future investigation.
The cell wall is composed mainly of lignin, cellulose and hemicellulose, and the proportions of these compounds are considered to be critical to SCW formation and determination of the properties of plant fibers (Cook et al. 2012; Meents et al. 2018). Our data show that the EgPHI-1 overexpression reduces the cellulose content in stems (Table 1). However, the proportions of cellulose/lignin and hemicellulose/cellulose were not significantly affected in the transgenic plants, possibly as the result of a compensatory mechanism to maintain the appropriate proportion among these compounds and the cell wall integrity, as previously proposed (Hu et al. 1999). For instance, L1 transgenic plants showed a decreased cellulose/lignin proportion in comparison with WT, but a compensatory increase in the hemicellulose/cellulose ratio (Table 1). Variations in the hemicellulose composition (xylan and arabinosyl groups), but not in its content, were also observed in some transgenic lines (Suppl. Table S1). Xylan is the main SCW hemicellulose and its level influences pulping properties, since it affects the delignification process (Bindschedler et al. 2007). On the other hand, arabinosyl increases the flexibility of cell walls (Jones et al. 2003). The increased extractive contents in stems of the transgenic lines may be the result of an increase in the content of waxy materials and low molar mass aromatics (Aharoni et al. 2004). It has been proposed that the lignin/cellulose reduction may cause a change in carbon flow that is directed towards the biosynthesis of other constituents of primary or SCWs (Ye 2002).
UV microspectrophotometry enabled us to determine the lignin content and distribution within individual cell wall layers of specific cell types. Two transgenic lines showed lower UV absorbance on S2 layers of fiber and vessel SCWs at 280 nm (Fig. 6). The UV spectrum from the SCW of the vessels from L3 presented a shoulder at 315 nm, suggesting the presence of alpha–beta unsaturated structures in the lignin, usually assigned to the presence of non-polymerized residues of the lignin precursors (Fig. 6b). In grasses, this absorption band has been assigned to the presence of hydroxycinnamic acids linked to hemicellulose and/or lignin (Siqueira et al. 2011). Softwood lignin, like that found in tobacco, presents absorption maximum at 280 nm and the reduction in absorbance values observed in transgenic lines suggests that the expression of EgPHI-1 may modify the lignification in both cell types.
UV scanning profiles revealed differences in the topochemical distribution of lignin in fibers of the WT and L2 transgenic plants. Although the areas of higher and lower lignin contents were virtually the same in both plants, with compound middle lamella and cell corners exhibiting the highest absorbances and SCW the lowest, the lignification distribution seems to be distinct between WT and L2 (Fig. 7). WT plants showed an apparently uniform pattern of lignin deposition, whereas a more diffuse lignification pattern was observed in L2. The compound middle lamella has been observed to show much higher lignin concentrations than the S2 layer. For instance, in angiosperms, the lignin concentration in compound middle lamella comprises approximately 40% as compared with 19% and 25% of the S2 layers of fibers and vessel elements, respectively (Donaldson et al. 2001).
Conclusion
EgPHI-1 represents a member in the PHI-1/EXO/EXL protein family that plays a role in partitioning of biomass, elongation of secondary xylem fibers, cell wall thickening and composition, and lignification. EgPHI-1 may act on signaling pathways that control cell division and differentiation in response to endogenous (i.e., auxin and cytokine) and environmental (i.e., wound and drought) signals. The modulation of complex traits such as shoot growth and wood formation is of special interest for biotechnological applications in forestry.
Author contribution statement
Conceived and designed the experiments: AOS, AF and MGCC. Performed the experiments: AOS, ETCMA, NSL and GOS. Analyzed the data: AOS, AF and MGCC. Contributed reagents/materials/analysis tools: AF, DCS, RPK, GP and MGCC. Wrote the paper: AOS, LRC and MGCC. All the authors read and approved the final manuscript.
Abbreviations
- EXO:
-
EXORDIUM
- EXL:
-
EXORDIUM-Like
- SCW:
-
Secondary cell wall
- PHI-1:
-
PHOSPHATE-INDUCED PROTEIN 1
- EgPHI-1:
-
Eucalyptus globulus PHI-1
- S2:
-
Layer of secondary cell wall
References
Aharoni A, Dixit S, Jetter R, Thoenes E, van Arkel G, Pereira A (2004) The SHINE clade of AP2 domain transcription factors activates wax biosynthesis, alters cuticle properties, and confers drought tolerance when overexpressed in Arabidopsis. Plant Cell 16:2463–2480. https://doi.org/10.1105/tpc.104.022897
Andersson-Gunneras S, Mellerowicz EJ, Love J, Segerman B, Ohmiya Y, Coutinho PM, Nilsson P, Henrissat B, Moritz T, Sundberg B (2006) Biosynthesis of cellulose-enriched tension wood in Populus: global analysis of transcripts and metabolites identifies biochemical and developmental regulators in secondary wall biosynthesis. Plant J 45:144–165. https://doi.org/10.1111/j.1365-313X.2005.02584.x
Argyros RD, Mathews DE, Chiang Y-H, Palmer CM, Thibault DM, Etheridge N, Argyros DA, Mason MG, Kieber JJ, Schaller GE (2008) Type B response regulators of Arabidopsis play key roles in cytokinin signaling and plant development. Plant Cell 20:2102–2116. https://doi.org/10.1105/tpc.108.059584
Bayer EM, Bottrill AR, Walshaw J, Vigouroux M, Naldrett MJ, Thomas CL, Maule Dr AJ (2006) Arabidopsis cell wall proteome defined using multidimensional protein identification technology. Proteomics 6:301–311. https://doi.org/10.1002/pmic.200500046
Bernard M (2003) Eucalyptus: a strategic forest tree. In: Wei R-P, Xu D (eds) Eucalyptus plantations: research, management and development. World Scientific Publishing, Singapore, pp 3–18
Bindschedler LV, Tuerck J, Maunders M, Ruel K, Petit-Conil M, Danoun S, Boudet A-M, Joseleau J-P, Bolwell GP (2007) Modification of hemicellulose content by antisense down-regulation of UDP-glucuronate decarboxylase in tobacco and its consequences for pulping properties. Phytochemistry 68:2635–2648. https://doi.org/10.1016/j.phytochem.2007.08.029
Boerjan W, Ralph J, Baucher M (2003) Lignin biosynthesis. Annu Rev Plant Biol 54:519–546. https://doi.org/10.1146/annurev.arplant.54.031902.134938
Borderies G, Jamet E, Lafitte C, Rossignol M, Jauneau A, Boudart G, Monsarrat B, Esquerré-Tugayé M-T, Boudet A, Pont-Lezica R (2003) Proteomics of loosely bound cell wall proteins of Arabidopsis thaliana cell suspension cultures: a critical analysis. Electrophoresis 24:3421–3432. https://doi.org/10.1002/elps.200305608
Coll-Garcia D, Mazuch J, Altmann T, Müssig C (2004) EXORDIUM regulates brassinosteroid-responsive genes. FEBS Lett 563:82–86. https://doi.org/10.1016/S0014-5793(04)00255-8
Cook CM, Daudi A, Millar DJ, Bindschedler LV, Khan S, Bolwell GP, Devoto A (2012) Transcriptional changes related to secondary wall formation in xylem of transgenic lines of tobacco altered for lignin or xylan content which show improved saccharification. Phytochemistry 74:79–89. https://doi.org/10.1016/j.phytochem.2011.10.009
Cosgrove DJ (2005) Growth of the plant cell wall. Nat Rev Mol Cell Biol 6:850–861. https://doi.org/10.1038/nrm1746
Crespo-Salvador Ó, Escamilla-Aguilar M, López-Cruz J, López-Rodas G, González-Bosch C (2018) Determination of histone epigenetic marks in Arabidopsis and tomato genes in the early response to Botrytis cinerea. Plant Cell Rep 37:153–166. https://doi.org/10.1007/s00299-017-2218-9
de Oliveira LA, Breton MC, Bastolla FM, Camargo SS, Margis R, Frazzon J, Pasquali G (2012) Reference genes for the normalization of gene expression in Eucalyptus species. Plant Cell Physiol 53:405–422. https://doi.org/10.1093/pcp/pcr187
De Vos M, Jander G (2009) Myzus persicae (green peach aphid) salivary components induce defence responses in Arabidopsis thaliana. Plant, Cell Environ 32:1548–1560. https://doi.org/10.1111/j.1365-3040.2009.02019.x
Decreux A, Messiaen J (2005) Wall-associated kinase WAK1 interacts with cell wall pectins in a calcium-induced conformation. Plant Cell Physiol 46:268–278. https://doi.org/10.1093/pcp/pci026
Dellagi A, Birch PRJ, Heilbronn J, Avrova AO, Montesano M, Palva ET, Lyon GD (2000) A potato gene, erg-1, is rapidly induced by Erwinia carotovora ssp. atroseptica, Phytophthora infestans, ethylene and salicylic acid. J Plant Physiol 157:201–205. https://doi.org/10.1016/S0176-1617(00)80191-1
Dita MA, Die JV, Ronàn B, Krajinski F, KüSter H, Moreno MT, Cubero JI, Rubiales D (2009) Gene expression profiling of Medicago truncatula roots in response to the parasitic plant Orobanche crenata. Weed Res 49:66–80. https://doi.org/10.1111/j.1365-3180.2009.00746.x
Donaldson L, Hague J, Snell R (2001) Lignin distribution in coppice poplar, linseed and wheat straw. Holzforschung 55:379–385. https://doi.org/10.1515/HF.2001.063
Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, Lepiniec L (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15:573–581. https://doi.org/10.1016/j.tplants.2010.06.005
Eudes A, Mouille G, Thévenin J, Goyallon A, Minic Z, Jouanin L (2008) Purification, cloning and functional characterization of an endogenous beta-glucuronidase in Arabidopsis thaliana. Plant Cell Physiol 49:1331–1341. https://doi.org/10.1093/pcp/pcn108
Farrar K, Evans IM, Topping JF, Souter MA, Nielsen JE, Lindsey K (2003) EXORDIUM - a gene expressed in proliferating cells and with a role in meristem function, identified by promoter trapping in Arabidopsis. Plant J 33:61–73. https://doi.org/10.1046/j.1365-313X.2003.01608.x
Feiz L, Irshad M, Pont-Lezica RF, Canut H, Jamet E (2006) Evaluation of cell wall preparations for proteomics: a new procedure for purifying cell walls from Arabidopsis hypocotyls. Plant Methods 2:10. https://doi.org/10.1186/1746-4811-2-10
Ferraz A, Baeza J, Rodriguez J, Freer J (2000) Estimating the chemical composition of biodegraded pine and eucalypt wood by DRIFT spectroscopy and multivariate analysis. Bioresource Technol 74:201–212. https://doi.org/10.1016/S0960-8524(00)00024-9
Forrest KL, Bhave M (2008) The PIP and TIP aquaporins in wheat form a large and diverse family with unique gene structures and functionally important features. Funct Integr Genomics 8:115–133. https://doi.org/10.1007/s10142-007-0065-4
Fox H, Doron-Faigenboim A, Kelly G, Bourstein R, Attia Z, Zhou J, Moshe Y, Moshelion M, David-Schwartz R (2018) Transcriptome analysis of Pinus halepensis under drought stress and during recovery. Tree Physiol 38:423–441. https://doi.org/10.1093/treephys/tpx137
Frey FP, Pitz M, Schön C, Hochholdinger F (2020) Transcriptomic diversity in seedling roots of European flint maize in response to cold. BMC Genomics 21:300. https://doi.org/10.1186/s12864-020-6682-1
Fujita Y, Fujita M, Shinozaki K, Yamaguchi-Shinozaki K (2011) ABA-mediated transcriptional regulation in response to osmotic stress in plants. J Plant Res 124:509–525. https://doi.org/10.1007/s10265-011-0412-3
Goicoechea M, Lacombe E, Legay S, Mihaljevic S, Rech P, Jauneau A, Lapierre C, Pollet B, Verhaegen D, Chaubet-Gigot N, Grima-Pettenati J (2005) EgMYB2, a new transcriptional activator from Eucalyptus xylem, regulates secondary cell wall formation and lignin biosynthesis. Plant J 43:553–567. https://doi.org/10.1111/j.1365-313X.2005.02480.x
Gómez-Porras JL, Riaño-Pachón DM, Dreyer I, Mayer JE, Mueller-Roeber B (2007) Genome-wide analysis of ABA-responsive elements ABRE and CE3 reveals divergent patterns in Arabidopsis and rice. BMC Genomics 8:260. https://doi.org/10.1186/1471-2164-8-260
Guo H, Wang Y, Wang L, Hu P, Wang Y, Jia Y, Zhang C, Yu Z, Zhang Y, Wang C, Yang C (2017) Expression of the MYB transcription factor gene BplMYB46 affects abiotic stress tolerance and secondary cell wall deposition in Betula platyphylla. Plant Biotechnol J 15:107–121. https://doi.org/10.1111/pbi.12595
Haralampidis K, Milioni D, Rigas S, Hatzopoulos P (2002) Combinatorial interaction of cis elements specifies the expression of the Arabidopsis AtHsp90-1 gene. Plant Physiol 129:1138–1149. https://doi.org/10.1104/pp.004044
Hu W-J, Harding SA, Lung J, Popko JL, Ralph J, Stokke DD, Tsai C-J, Chiang VL (1999) Repression of lignin biosynthesis promotes cellulose accumulation and growth in transgenic trees. Nat Biotechnol 17:808–812. https://doi.org/10.1038/11758
Iwase A, Ishii H, Aoyagi H, Ohme-Takagi M, Tanaka H (2005) Comparative analyses of the gene expression profiles of Arabidopsis intact plant and cultured cells. Biotechnol Lett 27:1097–1103. https://doi.org/10.1007/s10529-005-8456-x
Jamet E, Canut H, Boudart G, Pont-Lezica RF (2006) Cell wall proteins: a new insight through proteomics. Trends Plant Sci 11:33–39. https://doi.org/10.1016/j.tplants.2005.11.006
Jones DT, Taylor WR, Thornton JM (1992) The rapid generation of mutation data matrices from protein sequences. Bioinformatics 8:275–282. https://doi.org/10.1093/bioinformatics/8.3.275
Jones L, Milne JL, Ashford D, McQueen-Mason SJ (2003) Cell wall arabinan is essential for guard cell function. Proc Natl Acad Sci USA 100:11783–11788. https://doi.org/10.1073/pnas.1832434100
Ko JH, Han KH, Park S, Yang J (2004) Plant body weight-induced secondary growth in Arabidopsis and its transcription phenotype revealed by whole-transcriptome profiling. Plant Physiol 135:1069–1083. https://doi.org/10.1104/pp.104.038844
Kurth F, Feldhahn L, Bönn M, Herrmann S, Buscot F, Tarkka MT (2015) Large scale transcriptome analysis reveals interplay between development of forest trees and a beneficial mycorrhiza helper bacterium. BMC Genomics 16:658. https://doi.org/10.1186/s12864-015-1856-y
Leplé J-C, Dauwe R, Morreel K et al (2007) Downregulation of cinnamoyl-coenzyme A reductase in poplar: multiple-level phenotyping reveals effects on cell wall polymer metabolism and structure. Plant Cell 19:3669–3691. https://doi.org/10.1105/tpc.107.054148
Lisso J, Schröder F, Müssig C (2013) EXO modifies sucrose and trehalose responses and connects the extracellular carbon status to growth. Front Plant Sci 4:219. https://doi.org/10.3389/fpls.2013.00219
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Meents MJ, Watanabe Y, Samuels AL (2018) The cell biology of secondary cell wall biosynthesis. Ann Bot 121:1107–1125. https://doi.org/10.1093/aob/mcy005
Mellerowicz E, Sundberg B (2008) Wood cell walls: biosynthesis, developmental dynamics and their implications for wood properties. Curr Opin Plant Biol 11:293–300. https://doi.org/10.1016/j.pbi.2008.03.003
Mustafa BM, Coram TE, Pang ECK, Taylor PWJ, Ford R (2009) A cDNA microarray approach to decipher lentil (Lens culinaris) responses to Ascochyta lentis. Australas Plant Path 38:617–631. https://doi.org/10.1071/AP09048
Myburg AA, Potts BM, Marques CMP, Kirst M, Gion J-M, Grattapaglia D, Grima-Pettenati J (2007) Eucalyptus. In: Kole CR (ed) Genome mapping and molecular breeding in plants. Springer, Berlin, pp 115–160
Nakahama K, Urata N, Shinya T, Hayashi K, Nanto K, Rosa AC, Kawaoka A (2018) RNA-seq analysis of lignocellulose-related genes in hybrid Eucalyptus with contrasting wood basic density. BMC Plant Biol 18:156. https://doi.org/10.1186/s12870-018-1371-9
Nakano Y, Yamaguchi M, Endo H, Rejab NA, Ohtani M (2015) NAC-MYB-based transcriptional regulation of secondary cell wall biosynthesis in land plants. Front Plant Sci 6:288. https://doi.org/10.3389/fpls.2015.00288
Norton GJ, Nigar M, Williams PN, Dasgupta T, Meharg AA, Price AH (2008) Rice-arsenate interactions in hydroponics: a three-gene model for tolerance. J Exp Bot 59:2277–2284. https://doi.org/10.1093/jxb/ern098
Nuoendagula Tsuji Y, Takata N, Sakamoto S, Nakagawa-Izumi A, Taniguchi T, Ralph J, Mitsuda N, Kajita S (2018) Change in lignin structure, but not in lignin content, in transgenic poplar overexpressing the rice master regulator of secondary cell wall biosynthesis. Physiol Plant 163:170–182. https://doi.org/10.1111/ppl.12684
Obudulu O, Mähler N, Skotare T, Bygdell J, Abreu IN, Ahnlund M, Gandla ML, Petterle A, Moritz T, Hvidsten TR, Jönsson LJ, Wingsle G, Trygg J, Tuominen H (2018) A multi-omics approach reveals function of secretory carrier-associated membrane proteins in wood formation of Populus trees. BMC Genomics 19:11. https://doi.org/10.1186/s12864-017-4411-1
Ohtake Y, Takahashi T, Komeda Y (2000) Salicylic acid induces the expression of a number of receptor-like kinase genes in Arabidopsis thaliana. Plant Cell Physiol 41:1038–1044. https://doi.org/10.1093/pcp/pcd028
Pasquali G, Bastolla FM, Pazzini F, Kirch RP, Pizzoli G, Carazzole MF, Brondani RV, Coelho ASG, Grattapaglia D, Brommonschenkel SH, Pappas Jr GJ, Pereira GAG, Cascardo JCDM (2005) Sequencing and differential expression of xylem specific genes from two Eucalyptus species with highly contrasting wood properties. In: Institute FAAB (ed) IUFRO Tree Biotechnol 2005, Pretoria, South Africa, 2005. Programme and Abstracts of the IUFRO Tree Biotechnol, pp 22
Pesquet E, Ranocha P, Legay S, Digonnet C, Barbier O, Pichon M, Goffner D (2005) Novel markers of xylogenesis in zinnia are differentially regulated by auxin and cytokinin. Plant Physiol 139:1821–1839. https://doi.org/10.1104/pp.105.064337
Peter GF, White DE, Torre RDL, Singh R, Newman D (2007) The value of forest biotechnology: a cost modelling study with loblolly pine and kraft linerboard in the southeastern USA. Int J Biotechnol 9:415–435. https://doi.org/10.1504/IJBT.2007.014269
Pilate G, Guiney E, Holt K, Petit-Conil M, Lapierre C, Leple J-C, Pollet B, Mila I, Webster EA, Marstorp HG, Hopkins DW, Jouanin L, Boerjan W, Schuch W, Cornu D, Halpin C (2002) Field and pulping performances of transgenic trees with altered lignification. Nat Biotechnol 20:607–612. https://doi.org/10.1038/nbt0602-607
Quang T, Hallingbäck H, Gyllenstrand N, Arnold S, Clapham D (2012) Expression of genes of cellulose and lignin synthesis in Eucalyptus urophylla and its relation to some economic traits. Trees 26:893–901. https://doi.org/10.1007/s00468-011-0664-5
Roosjen M, Paque S, Weijers D (2018) Auxin Response Factors: output control in auxin biology. J Exp Bot 69:179–188. https://doi.org/10.1093/jxb/erx237
Saibo NJM, Lourenco T, Oliveira MM (2009) Transcription factors and regulation of photosynthetic and related metabolism under environmental stresses. Ann Bot 103:609–623. https://doi.org/10.1093/aob/mcn227
Sano T, Nagata T (2002) The possible involvement of a phosphate-induced transcription factor encoded by Phi-2 gene from tobacco in ABA-signaling pathways. Plant Cell Physiol 43:12–20. https://doi.org/10.1093/pcp/pcf002
Sano T, Kuraya Y, Amino S-I, Nagata T (1999) Phosphate as a limiting factor for the cell division of tobacco BY-2 cells. Plant Cell Physiol 40:1–16. https://doi.org/10.1093/oxfordjournals.pcp.a029464
Schröder F, Lisso J, Lange P, Müssig C (2009) The extracellular EXO protein mediates cell expansion in Arabidopsis leaves. BMC Plant Biol 9:20. https://doi.org/10.1186/1471-2229-9-20
Schröder F, Lisso J, Müssig C (2011) EXORDIUM-LIKE1 promotes growth during low carbon availability in Arabidopsis. Plant Physiol 156:1620–1630. https://doi.org/10.1104/pp.111.177204
Schröder F, Lisso J, Müssig C (2012) Expression pattern and putative function of EXL1 and homologous genes in Arabidopsis. Plant Signal Behav 7:22–27. https://doi.org/10.4161/psb.7.1.18369
Sham A, Al-Azzawi A, Al-Ameri S, Al-Mahmoud B, Awwad F, Al-Rawashdeh A, Iratni R, AbuQamar S (2014) Transcriptome analysis reveals genes commonly induced by Botrytis cinerea infection, cold, drought and oxidative stresses in Arabidopsis. PLoS ONE 9:e113718. https://doi.org/10.1371/journal.pone.0113718
Shinozaki K, Yamaguchi-Shinozaki K, Seki M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant Biol 6:410–417. https://doi.org/10.1016/S1369-5266(03)00092-X
Siqueira G, Milagres AM, Carvalho W, Koch G, Ferraz A (2011) Topochemical distribution of lignin and hydroxycinnamic acids in sugar-cane cell walls and its correlation with the enzymatic hydrolysis of polysaccharides. Biotechnol Biofuels 4:7. https://doi.org/10.1186/1754-6834-4-7
Sousa AO, Assis ETCM, Pirovani CP, Alvim FC, Costa MGC (2014) Phosphate-induced-1 gene from Eucalyptus (EgPHI-1) enhances osmotic stress tolerance in transgenic tobacco. Genet Mol Res 13:1579–1588. https://doi.org/10.4238/2014.March.12.10
Spurr AR (1969) A low-viscosity epoxy resin embedding medium for electron microscopy. J Ultrastruct Res 26:31–43
Steinmacher DA, Saare-Surminski K, Lieberei R (2012) Arabinogalactan proteins and the extracellular matrix surface network during peach palm somatic embryogenesis. Physiol Plant 146:336–349. https://doi.org/10.1111/j.1399-3054.2012.01642.x
Sudan C, Prakash S, Bhomkar P, Jain S, Bhalla-Sarin N (2006) Ubiquitous presence of β-glucuronidase (GUS) in plants and its regulation in some model plants. Planta 224:853–864. https://doi.org/10.1007/s00425-006-0276-2
Tamagnone L, Merida A, Parr A, Mackay S, Culianez-Macia FA, Roberts K, Martin C (1998) The AmMYB308 and AmMYB330 transcription factors from Antirrhinum regulate phenylpropanoid and lignin biosynthesis in transgenic tobacco. Plant Cell 10:135–154. https://doi.org/10.1105/tpc.10.2.135
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. https://doi.org/10.1093/molbev/msr121
Tarancón C, González-Grandío E, Oliveros JC, Nicolas M, Cubas P (2017) A conserved carbon starvation response underlies bud dormancy in woody and herbaceous species. Front Plant Sci 8:788. https://doi.org/10.3389/fpls.2017.00788
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680. https://doi.org/10.1093/nar/22.22.4673
Todd J, Post-Beittenmiller D, Jaworski JG (1999) KCS1 encodes a fatty acid elongase 3-ketoacyl-CoA synthase affecting wax biosynthesis in Arabidopsis thaliana. Plant J 17:119–130. https://doi.org/10.1046/j.1365-313X.1999.00352.x
Turner S, Gallois P, Brown D (2007) Tracheary element differentiation. Annu Rev Plant Biol 58:407–433. https://doi.org/10.1146/annurev.arplant.57.032905.105236
van Veen H, Vashisht D, Akman M, Girke T, Mustroph A, Reinen E, Hartman S, Kooiker M, van Tienderen P, Schranz ME, Bailey-Serres J, Voesenek LACJ, Sasidharan R (2016) Transcriptomes of eight Arabidopsis thaliana accessions reveal core conserved, genotype- and organ-specific responses to flooding stress. Plant Physiol 1761:668–689. https://doi.org/10.1104/pp.16.00472
Wang Q, Han C, Ferreira AO et al (2011) Transcriptional programming and functional interactions within the Phytophthora sojae RXLR effector repertoire. Plant Cell 23:2064–2086. https://doi.org/10.1105/tpc.111.086082
Wu J, Zhang Y, Zhang H, Huang H, Folta KM, Lu J (2010) Whole genome wide expression profiles of Vitis amurensis grape responding to downy mildew by using Solexa sequencing technology. BMC Plant Biol 10:234. https://doi.org/10.1186/1471-2229-10-234
Xiong L, Zhu J-K (2003) Regulation of abscisic acid biosynthesis. Plant Physiol 133:29–36. https://doi.org/10.1104/pp.103.025395
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803. https://doi.org/10.1146/annurev.arplant.57.032905.105444
Ye Z-H (2002) Vascular tissue differentiation and pattern formation in plants. Annu Rev Plant Biol 53:183–202. https://doi.org/10.1146/annurev.arplant.53.100301.135245
Zhong R, Lee C, Zhou J, McCarthy RL, Ye Z-H (2008) A battery of transcription factors involved in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell 20:2763–2782. https://doi.org/10.1105/tpc.108.061325
Zhong R, Lee C, Ye Z-H (2010) Evolutionary conservation of the transcriptional network regulating secondary cell wall biosynthesis. Trends Plant Sci 15:625–632. https://doi.org/10.1016/j.tplants.2010.08.007
Acknowledgements
The authors are grateful to José Moreira da Silva (School of Engineering of Lorena, University of São Paulo, São Paulo, Brazil) for the technical assistance in chemical and microscopic analyses. We are also thankful to Dr. Fernando Masarin for helping in discussions. This work was supported by ‘Financiadora de Estudos e Projetos’ (FINEP, Brazilian Ministry of Science, Technology, Innovations and Communications—MCTIC) [Grant No. 2101063500], ‘Conselho Nacional de Desenvolvimento Científico e Tecnológico’ (CNPq, MCT) [Grant Nos. 506348/04-0, 578632/08-0, 305549/16-2], ‘Fundação de Amparo à Pesquisa do Estado da Bahia’ (FAPESB, Secretary of Science, Technology and Innovation—SECTI) [Grant No. BOL1569/2010] and ‘Fundação de Amparo à Pesquisa do Estado de São Paulo’ (FAPESP) [Grant No. 11/50535-2].
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sousa, A.O., Camillo, L.R., Assis, E.T.C.M. et al. EgPHI-1, a PHOSPHATE-INDUCED-1 gene from Eucalyptus globulus, is involved in shoot growth, xylem fiber length and secondary cell wall properties. Planta 252, 45 (2020). https://doi.org/10.1007/s00425-020-03450-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-020-03450-x