Abstract
With increasing life expectancy, a growing number of individuals are being affected by Parkinson's Disease (PD), a Neurodegenerative Disease (ND). Approximately, 5–10% of PD is explained by genetic causes linked to known PD genes. With improvements in genetic testing and high-throughput technologies, more PD-associated susceptibility genes have been reported in recent years. However, a comprehensive review of the pathogenic mechanisms and physiological roles of these genes is still lacking. This article reviews novel genes with putative or confirmed pathogenic mutations in PD reported since 2019, summarizes the physiological functions and potential associations with PD. Newly reported PD-related genes include ANK2, DNAH1, STAB1, NOTCH2NLC, UQCRC1, ATP10B, TFG, CHMP1A, GIPC1, KIF21B, KIF24, SLC25A39, SPTBN1 and TOMM22. However, the evidence for pathogenic effects of many of these genes is inconclusive. A variety of novel PD-associated genes have been identified through clinical cases of PD patients and analysis of Genome-Wide Association Studies (GWAS). However, more evidence is needed in confirm the strong association of novel genes with disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
PD is the second most prominent ND after Alzheimer’s Disease (AD), involving 2–3% of the population aged above 65 years, which is mainly triggered by depletion of dopaminergic neurons located in the substantia nigra region of the basal ganglia and by the deposition of abnormal fibrillar alpha-synuclein (α-syn) in Lewy neurons [1]. The development of PD is associated with age, environmental changes, and genetic susceptibility.
Since the description of PD in 1817 by James Parkinson, scientists have been trying to decipher the pathogenesis of PD [2]. It is well known that the most hallmarks neuropathological features of PD are neuronal loss in the substantia nigra, striataum dopaminergic deficiency, and accumulation of α-syn in intraneuronal inclusion bodies [3].
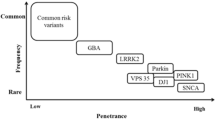
More recent studies, however, show that oxidative stress, mitochondrial dysfunction, cellular calcium imbalance, neuroinflammation, ferroptosis, Gut–brain axis, and other neurotransmitter system defects play a role in the pathogenesis of PD [4, 5]. Studies have shown that genetic factors can explain 5–10% of PD [6]. Nearly, 25 years have passed since the discovery of the first Parkinson's-related genetic gene, SNCA, mutations in which are associated with autosomal dominant PD [7]. The most frequent autosomal dominant monogenic PD is caused by mutations in the gene encoding Leucine-Rich Repeat Kinase 2 (LRRK2) [8]. Advances in genetic research methods have led to insights into how genetic background contribute to the development of PD, the use of state-of-the-art new methods in clinical practice has identified more than 100 genes or motifs associated with PD [9], and helps to predict disease risk and to predict specific clinical outcomes [10]. Figure 1 summarizes the timeline of milestone discoveries associated with major PD genes between 1997 and 2018 (Fig. 1).
However, more PD causative genes are being uncovered, which can facilitate investigation of pathogenic mechanism, and further promote the development of future diagnostic and therapeutic approaches. This study reviews relevant literature published in NCBI PubMed, Web of Science and Google Scholar from 2019 until 2022. The relevant literature was extracted from the databases with the following keyword combinations: "Parkinson's disease" OR "Novel mutation" OR "Genome-wide association studies" AND "Clinical cases" as well as free-text words. Newly reported PD-related genes include ANK2, DNAH1, STAB1, NOTCH2NLC, UQCRC1, MBNL2, GBF1, ATP10B, TFG, CHMP1A, GIPC1, IMMT, KIF21B, KIF24, MAN2C1, SLC25A39, SPTBN1, TOMM22 and ZSCAN21. However, the evidence for pathogenic effects of many of these genes is inconclusive. As additional genes are discovered to be linked to PD, studies of these genetics have revealed the complexity of PD and provided insights into its pathogenesis and etiology.
Novel genes for PD
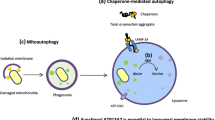
Since 2019, more than ten gene mutations have been recorded as novel causes of PD. The following sections serve as a review of the available reports on the clinical picture and genetic information. All these discoveries are recent, and it is today not proven that mutations in these genes cause PD. The presence of rare variants within these loci that may account for the increased susceptibility requires additional investigation. Table 1 summarizes the PD-associated genes identified since 2019 that commonly contribute to Early-Onset PD (EOPD) or Late-Onset PD (LOPD). Figure 2 summarizes the expression localization of these genes within the organelles (Fig. 2).
ANK2 (Ankyrin 2)
ANK2 is a gene encoding a member of the analytic family of proteins, which binds membrane proteins to the actin/myosin system in the cytoskeleton [11]. In 2019, Elisabeth Luisa Germer et al. [12] reported ANK2 mutations in 109 subjects exomes in Germany with exome sequencing. Three patients carried the ANK2 p.V3634D variant and two patients carried the ANK2 p.R3906W variant with an earliest age of onset (AAO) of 28 years. The other male patient was a compound heterozygote for the PRKN pathogenic variant (deletion 3–4 and exon 7–12 duplication) and the ANK2 p.V3634D variant. In 2017, a meta-analysis of GWAS with PD patients and controls individuals identified highly significant variants rs78738012 next to the ANK2 gene, providing independent evidence that ANK2 alterations contribute to the pathogenesis of PD [13].
ANK2 is associated with the development of several neurodevelopmental disorders, such as epilepsy, intellectual disability and autism, focal and generalized tonic–clonic seizures [14]. Studies have shown that ANK2 interacts with PINK1/Parkin-target proteins such as MIRO1 or ATP1A2, and ANK2-derived peptides are potent inhibitors of autophagy, which are both required to stabilize APC2 at microtubule branch points and to control the axonal transport of mitochondria to such points within neuronal dendrites [15]. Furthermore, in an ubiquitination analysis performed by researchers in the PARK2-mouse model, ANK2-controlled ATP1A2 was reported to be a novel neuron-specific Parkin substrates [16]. More detailed studies are needed to confirm the relationship between ANK2 and PD.
DNAH1 (Dynein Axonemal Heavy Chain 1)
DNAH1 encodes an inner dynein arm heavy chain, naturally occurring mutations are linked to primary ciliary dyskinesia and multiple morphological anomalies of the flagella that result in male infertility [17]. A woman from Poland carrying the DNAH1 p.A3639T mutation was diagnosed with PD at age 32 in 2019. Two patients with a LRRK2 p.R1441C mutation also carry a DNAH1 p.S1089G variant [12]. Currently, only the aforementioned cases have been identified with DNAH1 mutations within PD patients, more clinical data are needed to support the association between PD and DNAH1 mutation. Previous studies have shown that dysfunction of the DNAH1 gene can lead to ND. GWAS have shown that variants of DNAH1 are reported in patients with multiple sclerosis [18], which is an autoimmune-mediated ND of the Central Nervous System (CNS) characterized by inflammatory demyelination with axonal transection [19]. As of 2020, more than 30 cases of PD patients with MS have been reported and a few cases of demyelination-related secondary PD have been described. Multiple sclerosis is closely associated with PD [20]. Although mutations in DNAH1 have been reported in clinical cases leading to the development of PD, further studies are needed to determine the association.
STAB1 (Stalin 1)
Two sisters with the STAB1 p.S1089G variant in Elisabeth Luisa Germer's [12] discovery cohorts harbor a homozygous PINK1 p.Q456X variant. The patients' have an AAO of 51 and 80 years, respectively. STAB1 encodes a larger, transmembrane receptor protein that may function in angiogenesis, lymphocyte homing, cell adhesion, or receptor scavenging [21]. STAB1 has two functions in lymphocyte homing and endocytosis of ligands such as low-density lipoprotein, that would be consistent with a role for microglia in PD [22]. Additionally, Satb1-dependent Cntn5 expression in ON–OFF direction-selective ganglion cells leads to branch-specific homophilic interactions with interneurons [23]. However, there is still lack of evidence regarding the correlation between PD and STAB1 and further studies are needed.
NOTCH2NLC (Notch 2 N-Terminal Like C)
In 2019, the NOTCH2NLC gene 5’ untranslated regions (UTR) GGC repeat expansion mutations were identified as a genetic contributor of Neuronal Intranuclear Inclusion Disease (NIID) in a five-generation Chinese Han NIID family. Researchers reported three PD-affected families had GGC expansion in NOTCH2NLC [24]. NIID is a slowly progressive, ND which part of the clinical phenotypes include PD [25]. More reported cases with NOTCH2NLC GGC repeat expansion in PD patient has been discovered [26, 27].
NOTCH2NLC is located in chromosome 1q21.2 and positive regulation of the Notch signaling pathway [28], an important regulatory pathway that affects normal cell morphology and development. The principal clinical manifestations in NOTCH2NLC GGC repeat expansion include muscle weakness, cognitive impairment, PD, leukoencephalopathy, and so forth [29]. Some hypotheses have been suggested to involve in the pathogenesis of CGG expansion in NOTCH2NLC. For example, mistranslation by non-AUG upstream of extended GGC repeats may generate pathological proteins that can lead to disease [30]. Second, this repetition may produce RNA gain-of-function toxicity [31]. Third, the presence of GGC repeats may lead to aberrant methylation [24]. Researchers conducted a preliminary demonstration of the association study between NOTCH2NLC and PD, extensive experiments in cells and animal models are needed for further study.
UQCRC1 (Ubiquinol-Cytochrome C Reductase Core Protein 1)
In 2019, a novel missense mutation UQCRC1 p.Y314S was reported in the eastern Asia family with autosomal-dominant inheritance EOPD via whole-exome sequencing analysis [32]. All affected UQCRC1 heterozygotic carriers presented with asymmetrical onset tremor-predominant levodopa-responsive PD. The mean AAO of 54.2 ± 8.7 years. Another two novel variants, UQCRC1 p.I311L and an allele with concomitant splicing mutation c.70-1G > A and a frame shift insertion p.Ala25Gfs*27 in two families were detected among 699 autosomal dominant-inherited PD families from several East Asian populations [33]. However, there appear to be geographical differences in disease risk between UQCRC1 mutations and PD, with no UQCRC1 mutations detected in Caucasian PD patients in Europe [ 34, 35].
UQCRC1 is a subunit of mitochondrial ubiquinol-cytochrome c reductase and the function mainly is involved in oxidative phosphorylation and acts mitochondrial electron transport [36]. It is a biomarker of AD. Some hypotheses have been proposed to involve in the pathogenesis of UQCRC1 and PD. Researchers constructed SH-SY5Y cell lines with the UQCRC1 p.Y314S mutant, found that the mutant cell lines exhibited neuronal shortening and disrupted mitochondrial respiratory chain complex III activity in neurons [32]. Drosophila and mice with the Y314S variant both showed degeneration of dopaminergic neurons and locomotor defects. Other evidence of axonal type sensorimotor polyneuropathy in patients with UQCRC1 mutations, mutant knock-in Drosophila, and rodent models [33]. Induced pluripotent stem cells have been generated from peripheral blood mononuclear cells of a male patient with a heterozygous UQCRC1 p.Y314S mutation, and this cell model provides a platform for further studies of UQCRC1-related PD [37]. Additional UQCRC1 sequencing of PD patients from different geographic regions is recommended, as well as the construction of more UQCRC1-related animal models for the study.
ATP10B (ATPase Phospholipid Transporting 10B)
ATP10B, a component of the lysosome membrane, resides within the endoplasmic reticulum. It maintains the integrity and function of the lysosome membrane in cortical neurons. Bilateral missense and stop-gain variants of ATP10B associated with PD and dementia with Lewy bodies were reported in 2020 [38]. Firstly, compound heterozygous mutations in the ATP10B gene were reported in three patients with EOPD. Secondly, in a Belgian PD cohort reported four additional patients carrying compound heterozygous mutations, which confirming that compound heterozygous mutations in this gene are closely associated with PD. 5 of the seven mutation carriers were male. A patient with EOPD carried three ATP10B mutations simultaneously, p.G671R, p.N865K and p.V748L. 68-Year-old male patient carried both p.T161N and p.G648R mutations. The results of the cell function assay showed that impaired ATPase activity, GluCer and PC translocation activity and lysosomal function were observed in nine of the ten variants tested, leading to increased cell death, consistent with the loss of ATP10B function [38].
However, the pathogenicity of ATP10B for PD is controversial. Raquel Real et al. [39] using a publicly available case–control cohort, found no ATP10B variants that could be tested that were identified as functionally deleterious in vitro to be enriched in PD patients. Subsequent genotypic analyses of Chinese and Japanese PD patients showed no evidence that common or low-frequency genetic variants in ATP10B were associated with PD risk in the Chinese [40] and Japanese [41] populations. More clinicopathological reports are needed to support this.
TFG (Trafficking from ER to Golgi Regulator)
In 2022, a variant of the TFG gene c.1148 p.R383H was reported to be present in a Korean family and can cause PD or amyotrophic lateral sclerosis (ALS), the first time TFG was reported to be associated with PD. But the clinical phenotype of those carrying the mutated gene varied, and clinical trials have shown that of five family members, one had a PD, three had subclinical PD, and one (the proband) had ALS. In HeLa cells expressing R383H-TFG, abnormal aggregation of TDP43 were found and impairs cell viability [42].
TFG plays a role in the normal dynamic function of the endoplasmic reticulum and it was originally identified as a fusion partner leading to the formation of oncogenic products associated with multiple cancers. The discovery in 2012 that TFG mutations can cause the autosomal dominant motor and sensory neuropathies; in 2013, a homozygous mutation of TFG was reported in a family with early-onset spastic paraplegia, optic atrophy, and neuropathy; another novel mutation in TFG was discovered in 2014 as a cause of dominant axonal Charcot-Marie-Tooth disease type 2, there is increasing evidence that this gene is associated with neurological diseases [43]. These findings suggest that mutations of TFG cause ER dysfunction and neurodegeneration in ND, which is tightly associated with ER function. The development of PD is closely linked to endoplasmic reticulum stress (ERS). It was shown that TFG, a regulatory protein of endoplasmic reticulum to Golgi transport, is ubiquitinated in a NAB2-dependent manner in a cellular model of SNCA toxicity [44]. Future studies need to explore more comprehensive pathogenic mechanisms to better understand the function of TFG in PD. Finally, understanding the role of TFG proteins in PD may help discover therapeutic approaches to counteract ER dysfunction.
CHMP1A (Charged Multivesicular Body Protein 1A)
In 2021, by whole-exome data analysis, researchers reported disruptive variants of PD candidate genes in the Italian PD family, including a variant of the CHMP1A gene [45]. CHMP1A encodes a member of the CHMP protein family, which is associated with the sorting of proteins into the interior of lysosomes by multivesicular bodies. In 2019, researchers used linkage disequilibrium score regression to estimate the heritability of PD explained by genes involved in the endocytic membrane trafficking pathway, showing that the endocytic membrane transport pathway, as well as CHMP1A, contribute to PD etiology [46]. CHMP1A is closely related to neurological disorders and regulates the proliferation of neuronal progenitor cells. Loss-of-function mutations in human CHMP1A cause reduced cerebellar size (pontocerebellar hypoplasia) and reduced cerebral cortical size (microcephaly) [47]. In vivo experiments in mice show that deleting CHMP1A causes abnormalities in the brain's choroid plexus and Purkinje cell layers, as well as reduced proliferation of cerebellar granule cell precursors [48]. Moreover, knocking down Chmp1a in human iPSC-derived organoids leads to loss of progenitor cells and premature neuronal differentiation [49]. These findings offer a molecular explanation for how CHMP1A deletion in humans results in microcephaly. Interestingly, as a complex of the ESCRT-III complex, CHMP1A interacts with VPS4A in the cytoplasm to assist in the transport of ubiquitinated cargo proteins to lysosomes for degradation [50], both involved in the necroptosis and endocytosis pathways. Mutations in CHAMP2B have been linked to frontotemporal dementia and AD. Studies have shown that CHAMP2B mutation is associated with dendritic shrinkage, accumulation of ubiquitinated proteins and autophagosomes, and ultimately neuronal cell death [51]. Both CHAMP2B and CHMP1A are part of the ESCRT-III family of proteins, it is possible that the pathogenic mechanism of CHMP1A leading to PD is similar to CHAMP2B leading to NDs that needs to be addressed by future experiments.
GIPC1 (GIPC PDZ Domain Containing Family Member 1)
Short tandem repeat expansion disorders are a major cause of human neurological disease. More than 40 NDs are associated with short tandem repeat expansion disorders [52]. The repeat amplification of GIPC1 (c.875dupA) was first suggested to be associated with PD [45]. In 2022, whole-genome long read-length sequencing of 85 Chinese PD patients identified a total of 18 PD patients with GGC repeat expansion in GIPC1, with GGC repeat sizes ranging from 70 to 401. Although 8 individuals from a healthy Chinese population of 1631 were also reported to carry GIPC1 GGC repeat expansion [53]. Patients with PD usually have behavioral abnormalities. CGG Repeat Expansion of GIPC1 was also recently reported in patients with movement disorders [54]. GIPC1 is a scaffolding protein that regulates cell surface receptor expression and trafficking [55]. It has been proven that GIPC1 interacts with dopamine D2 and D3 receptors [56]. These receptors are the primary targets of antipsychotics and anti-PD drugs. GIPC1 mutation results in a reduction of DA neurons leading to locomotion defects in Drosophila [57]. In vitro and in vivo studies have shown that GIPC1 interacts with LRRK2 in dopamine cells and the ventral mesencephalon of mice [58]. CGG repeat expansions in GIPC1 has also been reported to cause oculopharyngeal myopathy, an ocular muscle disorder with dysphagia [59]. Given that patients with PD also have a clinical phenotype of involuntary ocular muscle spastic movements, future research into the mechanisms of PD and GIPC1 needs to be heavily investigated.
KIF21B (Kinesin Family Member 21B), KIF24 (Kinesin Family Member 24)
The researchers reported two members of the kinesin family both mutated in LOPD [45], KIF21B (p.A1079T and p.R211C) and KIF24 (p.I324T) mutation. Both KIF21B and KIF24 genes encode a member of the kinesin superfamily. Kinesins are ATP-dependent microtubule-based motor proteins that are involved in the intracellular transport of membranous organelles. Several kinesins are reported in a variety of ND, such as AD, ALS. KIF21B and KIF24 play an important part in neuronal function, plasticity, morphogenesis, and survival by transporting substance such as synaptic vesicle precursors and neurotransmitters [60]. The KIF21B gene also has been linked to ND. High KIF21B expression is associated with severe MS and AD pathology, and KIF21B risk SNPs can lead to ND. KIF21B expression was reported in neurons and astrocytes [61]. Interestingly, deletion of the LRRK2 gene, a strongly associated gene of PD, leads to altered expression of KIF21B [62]. In 2022, researchers reported a 9-year-old male with a p.S505R mutation in the KIF21B gene who had delayed speech, hyperactivity, poor social interaction, and autistic features. In vivo functional analysis of the mice embryonic cortex using in utero electroporation revealed that expression of the p.S505R KIF21B protein in the cerebral cortex impaired radial migration of projection neurons [63], providing further evidence that deleterious mutations in KIF21B are a cause of neurodevelopmental disorders.
KIF24 is a microtubule-dependent motor protein that acts as a negative regulator of cytogenesis, leading to restricted nucleation of cilia at centrioles. SNPs located in the KIF24 gene have been associated with Frontotemporal Lobar Degeneration [64]. Musculoskeletal disorders in PD is a complex clinically and pathophysiologically distinct problems. Recent studies have shown that missense variants of KIF24 are responsible for a wide range of skeletal ciliopathies, from fetal skeletal ciliopathies to acromegaly, and thus KIF24 may have a key role for KIF24 in musculoskeletal disorders in PD [65]. Currently, there are 45 known members of the KIF family, 38 of which are neuronally enriched, such as point mutations in the KIF5A gene has been associated with several axonopathies [66]. Understanding the function of kIF21B and KIF24 in neurons and skeletals may provide new insights into possible mechanisms for the treatment of PD.
SLC25A39 (Solute Carrier Family 25 Member 39)
The SLC25A39 p.T253N mutation has also been reported in LOPD. Immunohistochemistry experiments showed that the expression of SLC25A39 colocalized with most of the TH+ neurons in mouse mesodiencephalic dopaminergic neurons [45]. The SLC25A39 gene encodes a member of the SLC25 transporter protein or mitochondrial carrier protein family. The SLC25 mitochondrial transporter family plays a pivotal role in shaping intra-mitochondrial metabolite pools.
Mutations in members of the SLC25 family have been reported to cause severe lesions in the brains of patients. SLC25A39 and SLC25A40, located in epilepsy susceptibility loci, may cause an imbalance in metal ion homeostasis and mitochondrial dysfunction, leading to impaired synaptic transmission and neuronal survival [67]. A zebrafish model study demonstrating the role of SLC25A39 in anemia and hemoglobin synthesis, which involves intra-mitochondrial ferritin transport proteins, and glycine, a component of hemoglobin, which is thought to be transported by the solute carrier family protein SLC25A38 [68]. Though, there is no direct evidence of an interrelationship between SLC25A39 and PD, but mitochondrial function, iron ion homeostasis and reactive oxygen species production in the SLC25 family are important in the development of PD disease [69].
SPTBN1 (Spectrin Beta, Non-erythrocytic 1)
The SPTBN1 p.T2328N mutation has been documented in LOPD [45]. The SPTBN1 gene encodes spectrin protein, an actin cross-linking and molecular scaffolding protein, and plays a role in determining cell shape, arrangement of transmembrane proteins. In addition, SPTBN1 encodes beta-spectrin, an α-syn-binding protein and a component of Lewy bodies [70]. Pathological deposition of α-syn is a pathogenic factor in PD. In 2012, SPTBN1 was reported to interact specifically with α-syn in vitro and neuronal cells. Overexpression of SPTBN1 induced excessive neurite formation in dopaminergic neuronal cell bodies [71]. Exciting studies have shown that Spectrin is associated with specific neuronal structural domains and genetic links to certain inherited neurological disorders, and that mutations in the SPTAN1 gene are associated with early infantile epileptic encephalopathy, thus spectrin breakdown products could be potential biomarkers for ND [72]. The 2021 study confirmed the aforementioned facts about the close association of SPTBN1 with neurodevelopment. A rare autosomal dominant variant of SPTBN1 was reported in 29 individuals with neurological syndromes such as developmental, language, and motor delays, mental retardation, autism, and epilepsy. The researchers demonstrated that the SPTBN1 variant impaired protein stability, interfered with binding to key protein partners and changed cytoskeletal organization and dynamics [73]. The strong association of SPTBN1 with both neurodevelopment and α-syn indicates that this gene is strongly associated with PD. SPTAN1 p.R1098Q heterozygous mice exhibit progressive ataxia, as well as age-related deterioration in motor performance and muscle strength. Seizures in mice can be induced by stress, as well as their poor performance in memory tests for novel object recognition [74]. These neuropathologically relevant phenotypes are partially similar to the symptoms observed in PD patients.
TOMM22 (Translocase of Outer Mitochondrial Membrane 22)
The TOMM22 p.P13L mutant has been reported to be associated with PD [45] and TOMM22 is expressed in most TH+ neuronal cells, thus supporting the possibility that it may be cell-autonomous involvement in providing potential functional features for mdDA neurons [45]. TOMM22, the central component of the mitochondrial outer membrane translocase receptor complex, recognizes and translocates the synthesized mitochondrial precursor protein, which encodes a protein that forms a complex with TOMM20, TOMM40 and several other proteins to import cytosolic preproteins into the mitochondria [75]. Mitochondrial dysfunction leading to mitochondrial autophagy is also an essential physiological process that exacerbates PD pathogenesis, which may related to TOMM family interactions in PD pathogenesis. Studies have reported high-affinity binding of TOMM20 to oligomeric, dopamine-modified, and phosphomimetic α-syn species within the PD brain. This strong binding prevented the interaction of TOMM20 with its coreceptor, TOMM22, and impaired mitochondrial protein import, thus leading to deficient mitochondrial respiration, enhanced production of reactive oxygen species, and loss of mitochondrial membrane potential [76]. An exciting study shows that phosphorylated TOMM22 regulates mitochondrial phagocytosis, a specific type of autophagy [77]. Additional evidence is needed to demonstrate a direct link, but mitochondrial autophagy caused by TOMM22 may be an important cause of PD occurrence.
Conclusion and perspectives
An in-depth exploration of the etiological mechanisms of PD has always been pursued by researchers. Approximately 5–10% of PD is caused by genetic factors. The reported associated genes are not sufficient to explain the pathogenesis of all PD. Excitingly, many novel susceptibility genes associated with PD have recently been recorded through the mining of clinical data and GWAS data. This paper reviews genes identified since 2019 with putative or confirmed pathogenic mutations in PD ANK2, DNAH1, STAB1, NOTCH2NLC, UQCRC1, ATP10B, TFG, CHMP1A, GIPC1, KIF21B, KIF24, SLC25A39, SPTBN1 and TOMM22, and summarised the physiological roles of related genes and potential interrelationships with PD. Common risk factors between NDs, such as different pathogenic mutations in ANK2, DNAH1 or NOTCH2NLC genes can lead to multiple NDs, and this different pathogenicity of the same genes is worth exploring in the future. Although these results need more experimental data to support, they will help us to explore different pathogenic mechanisms and therapeutic approaches.
Data availability
Data availability is not applicable to this article as no new data were created or analyzed in this study.
Abbreviations
- PD:
-
Parkinson's disease
- AD:
-
Alzheimer’s disease
- α-syn:
-
Alpha-synuclein
- GWAS:
-
Genome-wide association studies
- LRRK2:
-
Leucine-rich repeat kinase 2
- AAO:
-
Age at onset
- MMAF:
-
Multiple morphological abnormalities of the flagella
- UTR:
-
Untranslated regions
- NIID:
-
Neuronal intranuclear inclusion disease
- CNS:
-
Central nervous system
- WES:
-
Whole exome sequencing
- ALS:
-
Amyotrophic lateral sclerosis
- LOPD:
-
Late-onset PD
- EOPD:
-
Early-onset PD
- ND:
-
Neurodegenerative diseases
- ERS:
-
Endoplasmic reticulum stress
References
Kumar S, Goyal L, Singh S (2022) Tremor and rigidity in patients with Parkinson’s disease: emphasis on epidemiology, pathophysiology and contributing factors. CNS Neurol Disord Drug Targets 21(7):596–609
Cherian A, Divya KP (2020) Genetics of Parkinson’s disease. Acta Neurol Belg 120(6):1297–1305
Sudhakar V, Richardson RM (2019) Gene therapy for neurodegenerative diseases. Neurotherapeutics 16(1):166–175
Mahoney-Sanchez L, Bouchaoui H, Ayton S, Devos D, Duce JA, Devedjian JC (2021) Ferroptosis and its potential role in the physiopathology of Parkinson’s Disease. Prog Neurobiol 196:101890
Bloem BR, Okun MS, Klein C (2021) Parkinson’s disease. Lancet 397(10291):2284–2303
Uwishema O, Onyeaka H, Badri R, Yucel AN, Korkusuz AK, Ajagbe AO et al (2022) The understanding of Parkinson’s disease through genetics and new therapies. Brain Behav 12(5):e2577
Polymeropoulos MH, Lavedan C, Leroy E, Ide SE, Dehejia A, Dutra A et al (1997) Mutation in the alpha-synuclein gene identified in families with Parkinson’s disease. Science 276(5321):2045–2047
Tolosa E, Vila M, Klein C, Rascol O (2020) LRRK2 in Parkinson disease: challenges of clinical trials. Nat Rev Neurol 16(2):97–107
Ye H, Robak LA, Yu M, Cykowski M, Shulman JM (2023) Genetics and Pathogenesis of Parkinson’s syndrome. Annu Rev Pathol 18:95–121
Fernandez-Santiago R, Sharma M (2022) What have we learned from genome-wide association studies (GWAS) in Parkinson’s disease? Ageing Res Rev 79:101648
Mohler PJ, Le Scouarnec S, Denjoy I, Lowe JS, Guicheney P, Caron L et al (2007) Defining the cellular phenotype of “ankyrin-B syndrome” variants: human ANK2 variants associated with clinical phenotypes display a spectrum of activities in cardiomyocytes. Circulation 115(4):432–441
Germer EL, Imhoff S, Vilarino-Guell C, Kasten M, Seibler P, Bruggemann N et al (2019) The role of rare coding variants in Parkinson’s disease GWAS loci. Front Neurol 10:1284
Chang D, Nalls MA, Hallgrimsdottir IB, Hunkapiller J, van der Brug M, Cai F et al (2017) A meta-analysis of genome-wide association studies identifies 17 new Parkinson’s disease risk loci. Nat Genet 49(10):1511–1516
Veltra D, Tilemis FN, Marinakis NM, Svingou M, Mitrakos A, Kosma K et al (2023) Combined exome analysis and exome depth assessment achieve a high diagnostic yield in an epilepsy case series, revealing significant genomic heterogeneity and novel mechanisms. Expert Rev Mol Diagn 23(1):85–103
Auburger G, Gispert S, Torres-Odio S, Jendrach M, Brehm N, Canet-Pons J et al (2019) SerThr-PhosphoProteome of brain from aged PINK1-KO+A53T-SNCA mice reveals pT1928-MAP1B and pS3781-ANK2 deficits, as hub between autophagy and synapse changes. Int J Mol Sci 20(13):3284
Key J, Mueller AK, Gispert S, Matschke L, Wittig I, Corti O et al (2019) Ubiquitylome profiling of Parkin-null brain reveals dysregulation of calcium homeostasis factors ATP1A2, Hippocalcin and GNA11, reflected by altered firing of noradrenergic neurons. Neurobiol Dis 127:114–130
Zhuang BJ, Xu SY, Dong L, Zhang PH, Zhuang BL, Huang XP et al (2022) Novel DNAH1 mutation loci lead to multiple morphological abnormalities of the sperm flagella and literature review. World J Mens Health 40(4):551
Haines J, Beecham A, McCauley J, Hadjixenofontos A, Whitehead P, Konidari I et al (2013) Whole-exome sequencing in multiplex families identifies novel rare variants in multiple sclerosis. Neurology
McGinley MP, Goldschmidt CH, Rae-Grant AD (2021) Diagnosis and treatment of multiple sclerosis: a review. JAMA 325(8):765–779
Sarin S, Wang A, Elkasaby M, Abboud H (2022) Parkinsonism in multiple sclerosis patients: a prospective observational study. Mult Scler Relat Disord 62:103796
Lin SY, Hu FF, Miao YR, Hu H, Lei Q, Zhang Q et al (2019) Identification of STAB1 in multiple datasets as a prognostic factor for cytogenetically normal AML: mechanism and drug indications. Mol Ther Nucleic Acids 18:476–484
Lecours C, Bordeleau M, Cantin L, Parent M, Paolo TD, Tremblay ME (2018) Microglial implication in Parkinson’s disease: loss of beneficial physiological roles or gain of inflammatory functions? Front Cell Neurosci 12:282
Peng YR, Tran NM, Krishnaswamy A, Kostadinov D, Martersteck EM, Sanes JR (2017) Satb1 regulates contactin 5 to pattern dendrites of a mammalian retinal ganglion cell. Neuron 95(4):869-883.e866
Tian Y, Wang JL, Huang W, Zeng S, Jiao B, Liu Z et al (2019) Expansion of human-specific GGC repeat in neuronal intranuclear inclusion disease-related disorders. Am J Hum Genet 105(1):166–176
Boivin M, Deng J, Pfister V, Grandgirard E, Oulad-Abdelghani M, Morlet B et al (2021) Translation of GGC repeat expansions into a toxic polyglycine protein in NIID defines a novel class of human genetic disorders: the polyG diseases. Neuron 109(11):1825-1835.e1825
Ma D, Tan YJ, Ng ASL, Ong HL, Sim W, Lim WK et al (2020) Association of NOTCH2NLC repeat expansions with Parkinson disease. JAMA Neurol 77(12):1559–1563
Shi CH, Fan Y, Yang J, Yuan YP, Shen S, Liu F et al (2021) NOTCH2NLC intermediate-length repeat expansions are associated with Parkinson disease. Ann Neurol 89(1):182–187
Suzuki IK, Gacquer D, Van Heurck R, Kumar D, Wojno M, Bilheu A et al (2018) Human-Specific NOTCH2NL Genes Expand Cortical Neurogenesis through Delta/Notch Regulation. Cell 173(6):1370-1384.e1316
Huang XR, Tang BS, Jin P, Guo JF (2022) The phenotypes and mechanisms of NOTCH2NLC-related GGC repeat expansion disorders: a comprehensive review. Mol Neurobiol 59(1):523–534
Sellier C, Buijsen RAM, He F, Natla S, Jung L, Tropel P et al (2017) Translation of expanded CGG repeats into FMRpolyG is pathogenic and may contribute to Fragile X tremor Ataxia syndrome. Neuron 93(2):331–347
Glineburg MR, Todd PK, Charlet-Berguerand N, Sellier C (2018) Repeat-associated non-AUG (RAN) translation and other molecular mechanisms in Fragile X tremor Ataxia syndrome. Brain Res 1693(Pt A):43–54
Lin CH, Chen PL, Tai CH, Lin HI, Chen CS, Chen ML et al (2019) A clinical and genetic study of early-onset and familial Parkinsonism in Taiwan: an integrated approach combining gene dosage analysis and next-generation sequencing. Mov Disord 34(4):506–515
Lin CH, Tsai PI, Lin HY, Hattori N, Funayama M, Jeon B et al (2020) Mitochondrial UQCRC1 mutations cause autosomal dominant parkinsonism with polyneuropathy. Brain 143(11):3352–3373
Senkevich K, Bandres-Ciga S, Gan-Or Z, Krohn L, International Parkinson’s Disease Genomics C (2021) Lack of evidence for association of UQCRC1 with Parkinson’s disease in Europeans. Neurobiol Aging 101:297.e291-297.e294
Courtin T, Tesson C, Corvol JC, Lesage S, Brice A, French Parkinson’s disease genetics g (2021) Lack of evidence for association of UQCRC1 with autosomal dominant Parkinson’s disease in Caucasian families. Neurogenetics 22(4):365–366
Unni S, Thiyagarajan S, Srinivas Bharath MM, Padmanabhan B (2019) Tryptophan oxidation in the UQCRC1 subunit of mitochondrial complex III (Ubiquinol-Cytochrome C Reductase) in a mouse model of myodegeneration causes large structural changes in the complex: a molecular dynamics simulation study. Sci Rep 9(1):10694
Ou-Yang CH, Lin HY, Huang CY, Lin CH (2020) Generation of a human induced pluripotent stem cell (iPSC) line (IBMS-iPSC-057-05) from a patient with familial Parkinsonism and polyneuropathy having a heterozygous pY314S mutation in UQCRC1 gene. Stem Cell Res 49:102031
Martin S, Smolders S, Van den Haute C, Heeman B, van Veen S, Crosiers D et al (2020) Mutated ATP10B increases Parkinson’s disease risk by compromising lysosomal glucosylceramide export. Acta Neuropathol 139(6):1001–1024
Real R, Moore A, Blauwendraat C, Morris HR, Bandres-Ciga S, International Parkinson’s Disease Genomics C (2020) ATP10B and the risk for Parkinson’s disease. Acta Neuropathol 140(3):401–402
Zhao Y, Pan H, Wang Y, Zeng Q, Fang Z, He R et al (2021) ATP10B variants in Parkinson’s disease: a large cohort study in Chinese mainland population. Acta Neuropathol 141(5):805–806
Ishiguro M, Yoshino H, Li Y, Ikeda A, Funayama M, Nishioka K et al (2021) Genetic analysis of ATP10B for Parkinson’s disease in Japan. Parkinsonism Relat Disord 88:10–12
Yoo D, Lee W, Lee SJ, Sung JJ, Jeon GS, Ban JJ et al (2022) A Novel TFG mutation in a Korean family with alpha-synucleinopathy and amyotrophic lateral sclerosis. Mov Disord 37(2):384–391
Yagi T, Ito D, Suzuki N (2016) TFG-related neurologic disorders: new insights into relationships between endoplasmic reticulum and neurodegeneration. J Neuropathol Exp Neurol 75(4):299–305
Hatstat AK, Ahrendt HD, Foster MW, Mayne L, Moseley MA, Englander SW et al (2021) Characterization of small-molecule-induced changes in Parkinson’s-related trafficking via the Nedd4 ubiquitin signaling cascade. Cell Chem Biol 28(1):14-25.e19
Gialluisi A, Reccia MG, Modugno N, Nutile T, Lombardi A, Di Giovannantonio LG et al (2021) Identification of sixteen novel candidate genes for late onset Parkinson’s disease. Mol Neurodegener 16(1):35
Bandres-Ciga S, Saez-Atienzar S, Bonet-Ponce L, Billingsley K, Vitale D, Blauwendraat C et al (2019) The endocytic membrane trafficking pathway plays a major role in the risk of Parkinson’s disease. Mov Disord 34(4):460–468
Mochida GH, Ganesh VS, de Michelena MI, Dias H, Atabay KD, Kathrein KL et al (2012) CHMP1A encodes an essential regulator of BMI1-INK4A in cerebellar development. Nat Genet 44(11):1260–1264
Coulter ME, Dorobantu CM, Lodewijk GA, Delalande F, Cianferani S, Ganesh VS et al (2018) The ESCRT-III protein CHMP1A mediates secretion of sonic hedgehog on a distinctive subtype of extracellular vesicles. Cell Rep 24(4):973-986.e978
Blanchette CR, Rodal AA (2020) Mechanisms for biogenesis and release of neuronal extracellular vesicles. Curr Opin Neurobiol 63:104–110
Stuchell-Brereton MD, Skalicky JJ, Kieffer C, Karren MA, Ghaffarian S, Sundquist WI (2007) ESCRT-III recognition by VPS4 ATPases. Nature 449(7163):740–744
Kaul Z, Chakrabarti O (2018) Endosomal sorting complexes required for ESCRTing cells toward death during neurogenesis, neurodevelopment and neurodegeneration. Traffic 19(7):485–495
Chintalaphani SR, Pineda SS, Deveson IW, Kumar KR (2021) An update on the neurological short tandem repeat expansion disorders and the emergence of long-read sequencing diagnostics. Acta Neuropathol Commun 9(1):98
Pan Y, Xue J, Chen J, Zhang X, Tu T, Xiao Q et al (2022) Assessment of GGC repeat expansion in GIPC1 in patients with Parkinson’s disease. Mov Disord 37(7):1557–1559
Fan Y, Shen S, Yang J, Yao D, Li M, Mao C et al (2022) GIPC1 CGG repeat expansion is associated with movement disorders. Ann Neurol 91(5):704–715
Lee NY, Ray B, How T, Blobe GC (2008) Endoglin promotes transforming growth factor beta-mediated Smad 1/5/8 signaling and inhibits endothelial cell migration through its association with GIPC. J Biol Chem 283(47):32527–32533
Arango-Lievano M, Sensoy O, Borie A, Corbani M, Guillon G, Sokoloff P et al (2016) A GIPC1-palmitate switch modulates dopamine Drd3 receptor trafficking and signaling. Mol Cell Biol 36(6):1019–1031
Kim J, Lee S, Ko S, Kim-Ha J (2010) dGIPC is required for the locomotive activity and longevity in Drosophila. Biochem Biophys Res Commun 402(3):565–570
Salasova A, Yokota C, Potesil D, Zdrahal Z, Bryja V, Arenas E (2017) A proteomic analysis of LRRK2 binding partners reveals interactions with multiple signaling components of the WNT/PCP pathway. Mol Neurodegener 12(1):54
Deng J, Yu J, Li P, Luan X, Cao L, Zhao J et al (2020) Expansion of GGC repeat in GIPC1 is associated with oculopharyngodistal myopathy. Am J Hum Genet 106(6):793–804
Hirokawa N, Niwa S, Tanaka Y (2010) Molecular motors in neurons: transport mechanisms and roles in brain function, development, and disease. Neuron 68(4):610–638
Kreft KL, van Meurs M, Wierenga-Wolf AF, Melief MJ, van Strien ME, Hol EM et al (2014) Abundant kif21b is associated with accelerated progression in neurodegenerative diseases. Acta Neuropathol Commun 2:144
Habig K, Walter M, Poths S, Riess O, Bonin M (2008) RNA interference of LRRK2-microarray expression analysis of a Parkinson’s disease key player. Neurogenetics 9(2):83–94
Narayanan DL, Rivera-Alvarez J, Tilly P, de Rosario MC, Bhat V, Godin JD et al (2022) Further delineation of KIF21B-related neurodevelopmental disorders. J Hum Genet 67(12):729–733
Venturelli E, Villa C, Fenoglio C, Clerici F, Marcone A, Benussi L et al (2010) Is KIF24 a genetic risk factor for frontotemporal lobar degeneration? Neurosci Lett 482(3):240–244
Reilly ML, Ain NU, Muurinen M, Tata A, Huber C, Simon M et al (2022) Biallelic KIF24 variants are responsible for a spectrum of skeletal disorders ranging from lethal skeletal ciliopathy to severe acromesomelic dysplasia. J Bone Miner Res 37(9):1642–1652
Hares K, Miners JS, Cook AJ, Rice C, Scolding N, Love S et al (2017) Overexpression of kinesin superfamily motor proteins in Alzheimer’s disease. J Alzheimers Dis 60(4):1511–1524
Slabbaert JR, Kuenen S, Swerts J, Maes I, Uytterhoeven V, Kasprowicz J et al (2016) Shawn, the drosophila homolog of SLC25A39/40, is a mitochondrial carrier that promotes neuronal survival. J Neurosci 36(6):1914–1929
Steele SL, Prykhozhij SV, Berman JN (2014) Zebrafish as a model system for mitochondrial biology and diseases. Transl Res 163(2):79–98
Ayka A, Sehirli AO (2020) The role of the SLC transporters protein in the neurodegenerative disorders. Clin Psychopharmacol Neurosci 18(2):174–187
Peuralinna T, Myllykangas L, Oinas M, Nalls MA, Keage HA, Isoviita VM et al (2015) Genome-wide association study of neocortical Lewy-related pathology. Ann Clin Transl Neurol 2(9):920–931
Lee HJ, Lee K, Im H (2012) alpha-Synuclein modulates neurite outgrowth by interacting with SPTBN1. Biochem Biophys Res Commun 424(3):497–502
Yan XX, Jeromin A, Jeromin A (2012) Spectrin breakdown products (SBDPs) as potential biomarkers for neurodegenerative diseases. Curr Transl Geriatr Exp Gerontol Rep 1(2):85–93
Cousin MA, Creighton BA, Breau KA, Spillmann RC, Torti E, Dontu S et al (2021) Pathogenic SPTBN1 variants cause an autosomal dominant neurodevelopmental syndrome. Nat Genet 53(7):1006–1021
Zalas M, Skrzymowska J, Miazek A, Miazek A (2023) Progressive ataxia, memory impairments, and seizure episodes in Spna2 R1098Q mouse variant affecting Alpha II Spectrin’s Scaffold stability. Brain Sci 13(2):261
Yano M, Hoogenraad N, Terada K, Mori M (2000) Identification and functional analysis of human Tom22 for protein import into mitochondria. Mol Cell Biol 20(19):7205–7213
Di Maio R, Barrett PJ, Hoffman EK, Barrett CW, Zharikov A, Borah A et al (2016) alpha-Synuclein binds to TOM20 and inhibits mitochondrial protein import in Parkinson’s disease. Sci Transl Med 8(342):342ra378
Kravic B, Harbauer AB, Romanello V, Simeone L, Vogtle FN, Kaiser T et al (2018) In mammalian skeletal muscle, phosphorylation of TOMM22 by protein kinase CSNK2/CK2 controls mitophagy. Autophagy 14(2):311–335
Yau WY, Sullivan R, Rocca C, Cali E, Vandrovcova J, Wood NW et al (2021) NOTCH2NLC intermediate-length repeat expansion and Parkinson’s disease in patients of European descent. Ann Neurol 89(3):633–635
Funding
National Natural Science Foundation of China, 31970574, Yuning Song, 32170543, Yuning Song.
Author information
Authors and Affiliations
Contributions
JY searched the literatures and drafted the manuscript. XW examined the literatures. YS designed the review and made a critical revision. All authors agree to be accountable for all aspects of work ensuring integrity and accuracy.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no conflict of interest.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, J., Wu, X. & Song, Y. Recent advances in novel mutation genes of Parkinson's disease. J Neurol 270, 3723–3732 (2023). https://doi.org/10.1007/s00415-023-11781-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00415-023-11781-4