Abstract
Sex determination in mammals is usually provided by a pair of chromosomes, XX in females and XY in males. Mole voles of the genus Ellobius are exceptions to this rule. In Ellobius tancrei, both males and females have a pair of XX chromosomes that are indistinguishable from each other in somatic cells. Nevertheless, several studies on Ellobius have reported that the two X chromosomes may have a differential organization and behavior during male meiosis. It has not yet been demonstrated if these differences also appear in female meiosis. To test this hypothesis, we have performed a comparative study of chromosome synapsis, recombination, and histone modifications during male and female meiosis in E. tancrei. We observed that synapsis between the two X chromosomes is limited to the short distal (telomeric) regions of the chromosomes in males, leaving the central region completely unsynapsed. This uneven behavior of sex chromosomes during male meiosis is accompanied by structural modifications of one of the X chromosomes, whose axial element tends to appear fragmented, accumulates the heterochromatin mark H3K9me3, and is associated with a specific nuclear body that accumulates epigenetic marks and proteins such as SUMO-1 and centromeric proteins but excludes others such as H3K4me, ubiH2A, and γH2AX. Unexpectedly, sex chromosome synapsis is delayed in female meiosis, leaving the central region unsynapsed during early pachytene. This region accumulates γH2AX up to the stage in which synapsis is completed. However, there are no structural or epigenetic differences similar to those found in males in either of the two X chromosomes. Finally, we observed that recombination in the sex chromosomes is restricted in both sexes. In males, crossover-associated MLH1 foci are located exclusively in the distal regions, indicating incipient differentiation of one of the sex chromosomes into a neo-Y. Notably, in female meiosis, the central region of the X chromosome is also devoid of MLH1 foci, revealing a lack of recombination, possibly due to insufficient homology. Overall, these results reveal new clues about the origin and evolution of sex chromosomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The X and Y chromosomes show extreme morphological and functional differentiation in mammals. Sex chromosomes are proposed to have originated from a pair of autosomes when one of the members of the pair acquired a male-determining allele and turned into a Y chromosome (Ohno 1967). This chromosome underwent a subsequent process of genetic erosion, losing most of its gene content (Graves 1995, 1998). Currently, in most mammals, the Y chromosome is very short and contains only a few genes, including the sex determining master gene SRY (sex-determining region on Y) (Cortez et al. 2014). By contrast, the X chromosome preserves most of the original gene content. The differentiation between sex chromosomes has been completed in some mammalian groups, like marsupials, in which X and Y chromosomes do not share homology anymore (Graves 1996). In eutherian mammals, the X and Y chromosomes, however, still share a small region of homology called the pseudoautosomal region (PAR) (Burgoyne 1982; Graves et al. 1998). This was provided by an autosomal translocation to both sex chromosomes occurred before the radiation of this group (Bellott et al. 2014; Cortez et al. 2014). Although this configuration is found in a wide range of mammals, there are species in which the sex chromosomes have undergone further divergence. In some species of gerbils, voles, and mice, the Y chromosome is completely differentiated and the PAR has been lost (Ashley et al. 1989; Borodin et al. 2012; Britton-Davidian et al. 2012; Carnero et al. 1991; de la Fuente et al. 2007, 2012; Page et al. 2005; Sharp 1982). In extreme cases, the entire Y chromosome has even been lost. This phenomenon has been found in bats of the genera Epomophorus and Epomops (Denys et al. 2013; Peterson and Nagorsen 1975; Primus et al. 2006), spiny rats of the genus Tokudaia (Honda et al. 1977, 1978; Kuroiwa et al. 2010), and mole voles of the genus Ellobius (Kolomiets et al. 1991; Matthey 1953; Vorontsov et al. 1980). This step in the evolution of the sex chromosomes leads to the appearance of a new sex determination system and likely requires genome reorganization, such as the movement of Y-linked genes to other chromosomes, as has been observed in Tokudaia and Ellobius (Arakawa et al. 2002; Kobayashi et al. 2007; Matveevsky et al. 2017; Mulugeta et al. 2016).
Different sex chromosome systems are known for the genus Ellobius (Mammalia: Rodentia), which is composed of five subterranean species divided into two morphologically distinct subgenera: Bramus and Ellobius. Within Bramus subgenus, the southern mole vole Ellobius fuscocapillus presents XX for females and XY for males (Lyapunova and Vorontsov 1978), while the Transcaucasian mole vole Ellobius lutescens presents a single X chromosome (X0) in both males and females (Matthey 1953). Ellobius subgenus consists of the eastern Ellobius tancrei, the northern Ellobius talpinus, and the Alay Ellobius alaicus mole voles, which show a pair of XX chromosomes in both sexes (Lyapunova and Vorontsov 1978). Loss of the Y chromosome is a derived event that occurred independently in these two subgenera, and it was preceded by a series of events related to changes in sex determination regulation (Bakloushinskaya and Matveevsky 2018). First, the TESCO (testis-specific enhancer of Sox9 core) sequence, which is responsible for the transcriptional regulation of the SOX9 (SRY-box 9) gene, was lost in a hypothetical ancestor of the whole Ellobius genus (Bagheri-Fam et al. 2012). After this event, the loss of the Y chromosome and the SRY gene and the translocation of Eif2s3y (eukaryotic translation initiation factor 2, subunit 3, structural gene Y-linked) gene occurred in the lineage leading to E. tancrei, E. talpinus, and E. alaicus and independently in Bramus lineage leading to E. lutescens, likely generating an X0/X0 system in both cases (Bakloushinskaya et al. 2019; Just et al. 1995; Matveevsky et al. 2017). Finally, in the E. tancrei, E. talpinus, and E. alaicus lineage, the single X chromosome doubled, generating an XX system in both males and females (Matveevsky et al. 2016), while in the E. lutescens lineage, an X0 system was retained in both sexes. To date, sex determination factors have not been characterized in Ellobius species (although it is known with certainty that it is not the SRY gene) (Just et al. 1995), nor is it known whether these factors are located on any of the sex chromosomes.
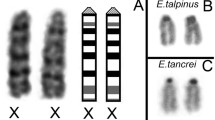
In this evolutionary scenario, the two X chromosomes present in E. tancrei, E. talpinus, and E. alaicus males and females appear to be the result of a duplication of an existing X chromosome, as they are homologous to the X chromosomes of the other Ellobius species and of other rodents (Bakloushinskaya et al. 2019, 2012; Romanenko et al. 2007). Moreover, both X chromosomes appear completely undifferentiated (isomorphic pair): their G-band pattern is identical in both sexes (Kolomiets et al. 1991; Vorontsov et al. 1980), and the little genomic data available indicate that there are no conspicuous molecular differences between them (Mulugeta et al. 2016). Nevertheless, previous studies have reported that the X chromosomes display an unexpected meiotic behavior in males (Kolomiets et al. 2010, 1991; Matveevsky et al. 2016).
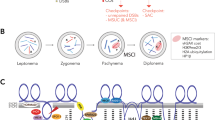
In most organisms, homologous chromosomes display very specific features during meiosis: they associate in pairs in a process called synapsis mediated by a protein structure called the synaptonemal complex (SC); recombine, leading to the formation of physical connections called chiasmata; and then segregate to different daughter cells during the first meiotic division (Zickler and Kleckner 1999). In mammalian females, the two X chromosomes behave like the rest of the autosomes, showing complete synapsis and a normal recombination schedule (Liu et al. 2004; Murdoch et al. 2013; Sung et al. 1983). By contrast, the X and Y chromosomes in males, owing to differences in size and gene content, show many modifications of the meiotic program. For instance, sex chromosome synapsis initiates later than in the rest of the chromosomes, is unstable, and usually extends heterologously beyond the region of homology (PAR) at the early pachytene stage (Goetz et al. 1984; Solari 1970, 1974). Nevertheless, the extreme size difference between the X and Y always leaves large unsynapsed chromosomal regions. The presence of these asynaptic regions leads to a specific silencing process called meiotic sex chromosome inactivation (MSCI) (Baarends et al. 2005; Mahadevaiah et al. 2001; Turner et al. 2004), a special form of a more general process called meiotic silencing of unsynapsed chromatin (MSUC) (Schimenti 2005). Recombination-related events, in essence a repair mechanism of DNA double-strand breaks (DSBs) involving the action of the proteins SPO11, RPA, RAD51, DMC1, and MLH1 (Keeney et al. 2014; Moens et al. 2002), among others, are also delayed in the sex chromosomes and may involve the action of specific SPO11 isoforms and additional or specific proteins (Acquaviva et al. 2020; Boekhout et al. 2019; Kauppi et al. 2011, 2012; Page et al. 2012; Papanikos et al. 2019).
Given the similarities in the organization and size of the X chromosomes in E. tancrei, E. talpinus, and E. alaicus, it was expected that they would behave like a pair of homomorphic chromosomes. Instead, several studies have shown that the X chromosomes in males only partially synapse, thereby triggering a MSCI phenomenon, and that recombination is restricted to regions close to the telomeres, thus resembling the behavior of highly differentiated sex chromosomes (Kolomiets et al. 2010, 1991; Matveevsky et al. 2016). This behavior indicates incipient functional differentiation of the two X chromosomes in males, which is revealed only during meiosis. In contrast, meiotic studies in E. talpinus and intraspecific E. tancrei hybrid females showed that synapsis between the X chromosomes is complete with no signs of meiotic inactivation (Kolomiets et al. 2010; Matveevsky et al. 2015). However, other aspects of meiotic behavior have not been explored. To shed light on the behavioral differences of X chromosomes in E. tancrei, and their likely incipient differentiation, we have compared sex chromosome behavior during meiosis in males and females of the 2n=34 chromosomal form of the species. We characterize the progression of sex chromosome synapsis, the appearance of epigenetic markers, and the distribution of chiasmata. Some results regarding these topics in male meiosis have been already published by some of us in previous reports (Bakloushinskaya and Matveevsky 2018; Kolomiets et al. 1991; Matveevsky et al. 2016, 2017). However, in order to make an appropriate comparison with females, some of them (mainly referring to the extent of synapsis and epigenetics marks) were presented here again. The results obtained corroborate the incipient epigenetic differentiation of the X chromosomes in males. We also report unexpected findings on sex chromosome behavior in female meiosis: synapsis between the X chromosomes is delayed and recombination is absent in the central region of these chromosomes. These findings have important consequences for our understanding of sex chromosome differentiation and reveal a more complex evolutionary scenario than initially anticipated.
Material and methods
Animals
Ellobius tancrei individuals were obtained from a long-term mole vole colony maintained at the Koltzov Institute of Developmental Biology of the Russian Academy of Sciences under standard conditions. Five males and three females of the 2n=34 chromosomal form of E. tancrei were used for this study. Animals were sacrificed, and the testes and ovaries were placed in phosphate-buffered saline (PBS) until further processed. All animal procedures were conducted under ethical permissions approved by the Ethics Committees for Animal Research of the Vavilov Institute of General Genetics and the Koltzov Institute of Developmental Biology, in accordance with the Regulations for Laboratory Practice of the Russian Federation, and by the Universidad Autónoma de Madrid (Ethics Committee Certificate CEI 55-999-A045).
Immunofluorescence
Spread spermatocytes and oocytes were obtained as previously described (Kolomiets et al. 2010; Page et al. 2003). Briefly, a cell suspension was obtained by disaggregating gonadal tissue in PBS using two tweezers. Then, the suspension was diluted in a 10-mM sucrose solution and incubated for 10 min. Afterward, a drop of the cell suspension was put onto a slide covered with 1% formaldehyde in distilled water (pH=9.5) containing 0.15% Triton X-100. Slides were placed on a flat surface in a moist chamber for 2 h. Finally, slides were washed with 0.04% Photoflo (Kodak) in distilled water, air dried, and immediately processed for immunofluorescence or stored at −80 °C. For immunofluorescence, slides were incubated overnight at 4 °C with the following primary antibodies and dilutions: mouse anti-SYCP3 (Abcam 97672) at 1:100; rabbit anti-SYCP3 (Abcam 15093) at 1:100; rabbit anti-SYCP1 (Abcam 15087) at 1:100; rabbit anti-RPA (Abcam 10359) at 1:100; rabbit anti-RAD51 (Calbiochem PC130) at 1:50; mouse anti-RNA polymerase-II (Abcam 24758-100) at 1:100; mouse anti-MLH1 (PharMingen 550838) at 1:100; mouse anti-MLH1 (Abcam 59756) at 1:50; mouse anti-histone H2AX phosphorylated at serine 139 (γH2AX) (Upstate 05-636) at 1:1000; mouse anti γH2AX (Abcam 22551) at 1:1000; rabbit anti-H3K9me3 (Abcam 8898) at 1:100; rabbit anti-H3K4me (Abcam 8895) at 1:100; mouse anti-fibrillarin (Abcam 4566) at 1:100; mouse anti-SUMO-1 (Zymed 33-2400) at 1:250; mouse anti-ubiquityl histone H2A (Millipore 05-678) at 1:400; and a human serum that recognizes centromere proteins (Antibodies Inc. 15-235) at 1:50. After incubation, slides were washed three times in PBS for 5 min each and then incubated for 1 h at room temperature with the appropriate secondary antibodies: donkey anti-rabbit, anti-mouse, or anti-human antibodies conjugated with Alexa 488, Alexa 594 (Invitrogen), or Cy3 (Jackson ImmunoResearch Laboratories). After three washes with PBS for 5 min each, slides were counterstained with 1-mM DAPI, washed in PBS, and mounted in Vectashield (Vector). For immunolabeling with more than three antibodies, we used the protocol described by Matveevsky et al. (2016). Slides were observed under an Olympus BX61 microscope equipped with an Olympus DP71 digital camera or a Zeiss Axio Imager D1 microscope equipped with a Zeiss Axiocam HRm CCD camera (Carl Zeiss) and the image-processing software AxioVision 4.6.3 (Carl Zeiss). Super resolution microscopy was performed on a Leica TCS SP8 confocal microscope equipped with a STED 3X system. Images were processed using Adobe Photoshop 7.0 and ImageJ.
Silver staining
For light microscopy, slides were treated with 2X SSC (300-mM sodium chloride and 30-mM trisodium citrate) at 60 °C for 10 min and then stained with a 50% AgNO3 solution in distilled water in a moist chamber at 60 °C for 25 min. The slides were washed three times in distilled water, air dried, and mounted with Eukitt. For electron microscopy, we followed previously described procedures (Kolomiets et al. 1991; Navarro et al. 1981). Plastic-coated slides were stained with 50% or 70% aqueous AgNO3, washed in distilled water and then air dried. Spermatocytes and oocytes were selected under the light microscope. Circles of plastic were cut from the slides with a diamond knife and then transferred to electron microscopy grids. Cells were observed under a JEM-1011 or JEM-100B electron microscope.
MLH1 foci count and distribution
MLH1 immunolocalization was used to estimate the number and distribution of chiasmata (Froenicke et al. 2002; Moens et al. 2002). The total number of MLH1 foci was recorded manually in 49 spermatocytes from two males and 30 oocytes from two females and then analyzed by a t-test with a 95% confidence interval. We only considered meiotic cells in which each bivalent showed at least one MLH1 foci. Foci distribution along the X chromosomes was analyzed in 56 spermatocytes from two males and 45 oocytes from two females. The length of the X bivalent SC in each spermatocyte or oocyte was measured using the free hand tool in ImageJ and divided into 10 equally distant intervals from the proximal to the distal telomere. Then, we measured the distances between the centromere and MLH1 foci, assigning each focus to its corresponding interval. Graph was made using Microsoft Excel.
Results
The meiotic karyotype of E. tancrei used in this study is composed of 16 autosomal bivalents and a sex bivalent, which is the largest acrocentric chromosome of the complement. This feature makes it easy to discern the sex bivalent from the other chromosomes in both male and female meiosis (Supplementary Figure 1).
Dynamics of sex chromosome synapsis in males
We first studied the dynamics of sex chromosome synapsis in males. For this purpose, spermatocytes were immunostained with SYCP3, the main component of the SC axial/lateral elements (AEs/LEs), and γH2AX, which marks the presence of DNA DSBs and in mammals accumulates on sex chromosomes during pachytene. As in other mammals, SYCP3 appears as short thin filaments at leptotene (Fig. 1a). At the beginning of zygotene, the filaments appear more elongated and, in some regions, associate in pairs (Fig. 1b). As synapsis progresses during zygotene, the AEs of homologous chromosomes become associated (Fig. 1c), and by the beginning of pachytene, all autosomes have completed synapsis (Fig. 1d). Synapsis is complete throughout pachytene (Fig. 1d–f), and at diplotene, homologous chromosomes start to separate along their length (Fig. 1g). Finally, the LEs shorten and become irregular at diakinesis (Fig. 1h). The localization of γH2AX is comparable to that of other mammals: discrete foci appear at leptotene and then extend to cover the whole nucleus during early zygotene, followed by a decrease in signal in the autosomes as synapsis progresses, with small foci associated with some bivalents remaining until early-mid pachytene (Fig. 1a–e).
Prophase I progression in E. tancrei spermatocytes. SYCP3 (green) and γH2AX (red). a–h Whole spermatocyte nuclei. c’–g’ Enlarged detail views and c”–g” schematic representations of the sex chromosomes (XX) shown in a–g. a Leptotene. Axial elements (AEs) start to form and γH2AX labeling appears as large foci scattered over the nucleus. b. Early zygotene. AEs appear as thin filaments that associate in some regions. γH2AX signal occupies the entire nucleus, although it is not homogeneously distributed: some chromosomal regions are devoid of signal, whether they are synapsed or not. c Late zygotene. Most autosomes have completed synapsis. γH2AX labeling appears as elongated foci associated with some chromosomes. Sex chromosomes (XX) are distinguishable as they present an intense γH2AX signal with an irregular outline. Enlarged detail view of the sex chromosomes (c’) and their schematic representation (c”) show that synapsis has started at both ends (arrowheads). A region of one of the X chromosomes shows a less intense γH2AX signal (arrow). d-d” Early pachytene. All autosomes have completed synapsis, and small γH2AX foci remain associated with many of them. Synapsis of sex chromosomes (enlarged in d’) is more extended than at zygotene (arrowheads). The AE of one X chromosome is continuous, while the other appears fragmented. The fragmented region is partially devoid of γH2AX (arrow). e-e” Mid pachytene. Autosomes only show a few small γH2AX foci. Sex chromosomes form a compact sex body delimited by a γH2AX signal that presents a well-defined outline. Synapsis is more extended in one of the chromosomal ends (arrowheads). As at the previous stage, one of the X chromosomes appears fragmented, and in this region, it is partially devoid of γH2AX (arrow). f-f” Late pachytene. γH2AX is not detected on the autosomes. Sex chromosomes appear synapsed only at one end (arrowhead); the opposite end is unsynapsed. As at previous stages, one AE appears fragmented and shows a region devoid of γH2AX. g-g” Early diplotene. Autosomes start to desynapse. Sex chromosome AEs are thinner and more intermingled than at previous stages. The γH2AX signal is irregular around the sex chromosomes and less intense than at previous stages. h Late diplotene. Autosomal lateral elements appear fragmented and shorter than at previous stages. Some scattered γH2AX foci appear distributed irregularly over the nucleus. At this stage, the sex chromosomes are not distinctively labeled by γH2AX and thus are not distinguishable from the autosomes. Bar represents 10 μm
The sex chromosomes become distinguishable from the rest of the chromosomes only at late zygotene (Fig. 1c’, c”). While the autosomes have mostly completed synapsis, the X chromosomes remain largely unsynapsed and conspicuously labeled with γH2AX. During pachytene (Fig. 1d’–f”), synapsis is only achieved at the chromosomal ends and is variable in length. Although there are cell-to-cell differences, synapsis tends to be at its maximum at early pachytene (Fig. 1d’–d”) and then decreases with pachytene progression (Fig. 1e’–f’). The morphology of the AEs of both X chromosomes seems different during zygotene and pachytene. Usually, one of the AEs is continuous, while the other appears clearly fragmented. In a small fraction of cells, however, we observed both AEs as fragmented (see Fig. 2) and very rarely, as continuous. At early diplotene, the sex chromosome AEs become irregular and usually fold on each other (Fig. 1g’–g”), and, by late diplotene, they are no longer distinguishable from the autosomes (Fig. 1h).
Synapsis and DNA repair in the X chromosomes of E. tancrei spermatocytes. Localization of SYCP3 (green) and SYCP1 (red) in conventional fluorescence microscopy (a, b) and STED super-resolution fluorescence microscopy (c, d). The association of sex chromosome ends is mediated by SYCP1, indicating the formation of a complete SC at both early (a, c) and late pachytene (b, d). STED does not reveal any conspicuous modification of the sex chromosome AEs. e–f Electron microscopy also does not reveal any structural modification of the AEs. g Localization of SYCP3 (green) and RPA (red). RPA localizes to both synapsed and unsynapsed AEs. h Localization of SYCP3 (green) and RAD51 (red). RAD51 also localizes to both synapsed and unsynapsed AEs. Bar represents 2 μm. Picture in e is reproduced from Matveevsky et al. (2016)
During zygotene, the γH2AX signal intensifies over both X chromosomes. At this stage, the signal is irregular at the periphery of the sex chromosomes (Fig. 1c’) but becomes more regular at pachytene (Fig. 1d’–f’), concomitant with the appearance of a typical sex body. During diplotene, the γH2AX signal starts to fade from the sex chromosomes (Fig. 1g’) and, by late diplotene, is not specifically detected on them (Fig. 1h). This pattern departs from that described for most mammals in which γH2AX remains, at least, until metaphase-I. As previously described (Matveevsky et al. 2017), a γH2AX-negative structure (occasionally two) appears associated with the sex chromosomes. This small round-like structure is detectable from zygotene to late pachytene and is typically associated with the AE of the X chromosome that often appears fragmented. Indeed, it clearly interrupts the trajectory of this chromosome’s AE (Fig. 1c’–f’).
The fact that the sex chromosomes in males do not complete synapsis does not appear to be due to structural rearrangements since the G-banding pattern is identical in both X chromosomes (Vorontsov et al. 1980). To better understand this behavior, we assessed whether there are structural modifications of the SC that could hamper synapsis progression. For this purpose, we characterized the localization of SYCP3 and SYCP1, the main component of the SC transverse filaments, using both conventional and stimulated emission depletion (STED) fluorescence microscopy. We found that the associated regions of the X chromosomes incorporate SYCP1 (Fig. 2a–b), indicating true synapsis. We did not observe structural modifications of the AEs (such as thickenings, splittings, or excrescences) in either the synapsed or unsynapsed regions (Fig. 2c–d), although small patches of SYCP1 were observed along the unsynapsed AEs during early pachytene (Fig. 2a–c). The lack of conspicuous structural modifications was corroborated by electron microscopy (Fig. 2e–f).
We also assessed whether incomplete synapsis could be due to the absence of DNA DSBs along the unsynapsed AEs by characterizing the localization of DSB repair proteins RPA (Replication protein A) and RAD51 (bacterial homologue RecA). The complete cycle of distribution of these proteins is shown in Supplementary Figures 3 and 4. Regarding sex chromosomes, the most relevant result is that both proteins appear all along the AEs of the X chromosomes (Fig. 2g–h), indicating the presence of DSBs and recombination intermediates in these chromosomal regions.
Epigenetic differentiation of X chromosomes in males
As indicated above, one of the sex chromosomes seems to be associated with one, or occasionally two, distinctive nuclear bodies. Given the structure’s intense labeling with silver staining techniques commonly used for electron microscopy (Fig. 2e–f; Fig. 3a–b), it was hypothesized to be the remains of the nucleolus and was therefore called the nucleolus-like body/structure (Kolomiets et al. 1991). To corroborate this possibility, we examined the immunolocalization of fibrillarin, a component of the nucleolar dense fibrillar component. We found that fibrillarin localizes to the ends of four autosomal bivalents bearing nucleolar organizer regions (NORs). However, no signal was found on the sex chromosomes (Fig. 3c–d), discarding a nucleolar nature for the sex chromosome-associated body.
Characterization of the nuclear body (NB) associated with sex chromosomes in E. tancrei spermatocytes. a Silver staining of a spermatocyte at pachytene. Intense staining, representing the nucleoli (Nu), is observed at the ends of four of the short acrocentric bivalents. An additional dense staining is associated with one of the X chromosomes. b Enlarged detail view of the X chromosomes. The NB is intensely stained and interrupts the trajectory of the AEs of one of the X chromosomes. c Localization of SYCP3 (green) and fibrillarin (red) in a spermatocyte at pachytene. Four fibrillarin foci are observed at the ends of the four short acrocentric bivalents, indicating the location of the nucleoli (Nu). No signal is observed associated with the sex chromosomes. d Enlarged detail view of the X chromosomes shown in c. Fibrillarin does not accumulate on either of the X chromosomes. The AE of one X chromosome is interrupted and a faint SYCP3 signal is observed, probably corresponding to the location of the NB. Bar represents 10 μm in a and c and 2 μm in b and d
More recently, some of us showed that the nuclear body contains DNA and during pachytene accumulates proteins associated with sex chromosome inactivation such as SUMO-1 (Small Ubiquitin Like Modifier 1); as a result, it was renamed as a chromatin body (ChB) (Matveevsky et al. 2017). We noticed that anti-centromere antibodies label the ChB (Fig. 4a, a’; Supplementary Figure 1). Since heterochromatic regions are frequently labeled with centromere markers, we hypothesized that this chromosomal region may be heterochromatic. To test this, we examined the localization of histone H3 trimethylated at lysine 9 (H3K9me3), a typical marker for heterochromatin (Cowell et al. 2002). We observed an accumulation of H3K9me3 at the pericentromeric regions of chromosomes, including the X chromosomes. In addition, we found an intense H3K9me3 labeling associated to one of the X chromosomes (Fig. 4b, b’). The localization of SUMO-1 (Fig. 4c, c’) and DNA (Fig. 4d, d’) in the same region indicates that this H3K9me3-rich structure corresponds to the ChB, which therefore presents a heterochromatic organization. Some additional H3K9me3 foci were observed along the same X chromosome, indicating the heterochromatinization of some additional regions of this chromosome. These marks also tend to localize to regions where the AE is fragmented, although not always.
Localization of epigenetic markers in the sex chromosomes of E. tancrei spermatocytes at pachytene. a–d Whole spermatocyte nuclei. a’–d’ Enlarged detail views of the sex chromosomes (XX) shown in a–d. a, a’ SYPC3 (green) and centromeric proteins (red). The centromeric regions of all bivalents are intensely labeled with the centromeric proteins. Additionally, a diffuse signal accumulates in the central region of one of the X chromosomes (arrow in a’). b–d’ A spermatocyte labeled for SYPC3 (green), H3K9me3 (red), SUMO-1 (pink), and DAPI (blue). b, b’ The centromeric regions of all bivalents and sex chromosomes (XX) are labeled with H3K9me3. In the sex chromosomes, in addition to the centromeric region (blue arrow in b’), a large interstitial region of one of the X chromosomes shows intense labeling (arrow in b’). Some smaller regions are also labeled along the same X chromosome (arrowheads). c, c’ SUMO-1 accumulates only in the central region of the same X chromosome (arrow in c’), coincident with the large H3K9me3 labeled region. d, d’ The sex chromosomes (XX) form a distinctive sex body. d’ DAPI staining covers the region labeled by both H3K9me3 and SUMO-1 (arrow in d’). Bar represents 10 μm in a–d, and 2 μm in a’–d’
On the other hand, we found that other epigenetic marks are partially or completely absent from the ChB. We already showed that γH2AX is excluded from this structure (Fig. 1, 5a, a’). In addition, we observed that two histone modifications, H2A ubiquitinated at lysine 119 (ubiH2A) (already reported in Matveevsky et al. (2016)) and H3 monomethylated at lysine 4 (H3K4me), are largely excluded from the ChB (Fig. 5b–c’). All three of these epigenetic marks have been implicated in the inactivation of sex chromosomes during meiosis (MSCI); therefore, their absence in the ChB might be associated with transcriptional activation of this structure. To test this, we studied the localization of RNA polymerase-II (RNA pol-II) during the first meiotic prophase (Fig. 5d, d’ and Supplementary Figure 2). We observed that this protein is already abundant in the nucleus at early pachytene (Supplementary Figure 2), contrary to what has been described in mouse in which a transcription burst seems to occur at mid-late pachytene (Page et al. 2012). However, the sex chromosomes are completely devoid of RNA pol-II signal throughout pachytene, indicating that the ChB does undergo MSCI. Unexpectedly, we observed some RNA pol-II signal on the sex chromosomes at diplotene (Supplementary Figure 2), concomitant with the decrease of γH2AX on these chromosomes (see Fig. 1h), suggesting differential regulation/reactivation of these chromosomes at this stage, which clearly departs from the pattern described in other mammals.
Localization of epigenetic markers in the sex chromosomes of E. tancrei spermatocytes at mid-late pachytene. a–d Whole spermatocyte nuclei. a’–d’ Enlarged detail views of the sex chromosomes shown in a–d. a, a’ SYPC3 (green) and γH2AX (red). As shown in Fig. 2, γH2AX covers the entire sex body, except a round structure associated with one of the X chromosomes that interrupts the trajectory of the AE (arrow in a’). b, b’ SYPC3 (green) and ubiH2A (red). The only region stained in the nucleus corresponds to the sex chromosomes. A fainter area of staining is observed in one of the chromosomes (arrow in b’). c, c’ SYPC3 (green) and H3K4me (red). A faint H3K4me signal is detected throughout the nucleus, but with a clear accumulation in the sex chromosomes. The signal is not as homogeneous as in the case of γH2AX and ubiH2A. A fainter area of staining is observed in one of the chromosomes (arrow in c’). d, d’ SYPC3 (green) and RNA polymerase-II (red). The entire nucleus is labeled, except for the sex chromosomes. Bar represents 10 μm in a–d and 2 μm in a’–d’
Sex chromosome synapsis in female meiosis
The fact that the two sex chromosomes do not complete synapsis in male meiosis raises interesting questions regarding possible differences in female meiosis. Previous studies have shown that the two X chromosomes display complete synapsis in females of the sibling species E. talpinus (Kolomiets et al. 2010) and the 49-chromosomal E. tancrei hybrid (Matveevsky et al. 2015). However, the sequence of synapsis and the presence of epigenetic modifications are not known.
To determine the sequence of synapsis during female meiosis, we examined the localization of SYCP3 and γH2AX in E. tancrei oocytes at different stages of prophase I. The leptotene stage is characterized by the presence of small SYCP3 filaments and the homogeneous labeling of the entire nucleus with γH2AX (Fig. 6a). In addition, one or more large accumulations of SYCP3 remain throughout prophase I. The γH2AX signal gradually decreases during zygotene as homologous chromosomes synapse, remaining mainly only in regions that have not yet completed synapsis (Fig. 6b). At pachytene, γH2AX is restricted to small foci associated with some SCs (Fig. 6c). We also observed one bivalent that consistently showed a delay in completing synapsis and appears intensely labeled with γH2AX. Given the morphological features and size of the bivalent and the distal position of its centromere (Fig. 6d), we were able to identify it as the sex bivalent. Synapsis of the sex chromosomes begins at both ends, although it usually appears more extended at one end than at the other (Fig. 6b’–c’). γH2AX labeling is mainly located over the asynaptic region (Fig. 6b’–d’), although it can remain also in some already synapsed regions at early pachytene. This pattern is reminiscent of that of the sex bivalent in male meiosis. However, in females, the XX chromosomes ultimately complete synapsis, forming a full-length SC by mid-late pachytene (Fig. 6e; see also Supplementary Figure 1). After synapsis completion, γH2AX does not accumulate over the sex chromosomes anymore.
Prophase I progression in E. tancrei oocytes. SYCP3 (green), γH2AX (red), and centromeres (blue). a Leptotene. AEs have formed and γH2AX labeling is observed throughout the entire nucleus. b Late zygotene. Most autosomes have completed synapsis. γH2AX labeling appears as large foci associated with a few chromosomes that have not yet completed synapsis and also the sex chromosomes (XX). Enlarged detail view of the sex chromosomes (b’) shows only two short regions of synapsis at both ends of the chromosomes, which appear mostly unsynapsed. c, c’ Early pachytene. All autosomes have completed synapsis, but faint γH2AX foci remain associated with some of them. Synapsis of the sex chromosomes (enlarged in c’) has largely progressed but is still incomplete. γH2AX intensely covers the unsynapsed regions. d, d’ Localization of the centromere marker confirms that the lagging chromosomes are undoubtedly the sex chromosomes. e Mid pachytene. All chromosomes, including the sex chromosomes, have completed synapsis. Small γH2AX foci remain associated with some bivalents. Bar represents 10 μm in a–e and 2 μm in b’–d’
We then performed an electron microscopy analysis to look for structural modifications of the AEs that could explain the delay to complete synapsis (Fig. 7). The outline of the AEs of the sex chromosomes do not show any evident deformation during zygotene (Fig. 7a, a’). At early pachytene, a slight thickening of the AEs in the unsynapsed regions can be occasionally observed (Fig. 7b–c’), as well as a sporadic interruption of one of the AEs (Fig. 7b; see also Fig. 6b); however, upon synapsis completion, the SC of the sex bivalent is completely normal (Fig. 7d, d’). We did not observe a ChB-like structure or any other nuclear bodies associated with the sex chromosomes at any stage, although nucleolar structures associated with autosomal bivalents can be clearly observed at pachytene.
Electron microscopy of E. tancrei oocytes. a–d Whole oocyte nuclei. a’–d’ Enlarged detail views of the sex chromosomes shown in a–d. a, a’ Zygotene. Most autosomes have completed synapsis, but the sex chromosomes (X) are completely unsynapsed. b–c’ Early pachytene. Synapsis is complete in the autosomes; the sex chromosomes remain largely unsynapsed. The AEs of the sex chromosomes do not show conspicuous modifications (b’), although they sometimes appear slightly thickened (c’). d, d’ By mid pachytene, the sex chromosomes have completed synapsis and are only distinguishable from the rest of the bivalents by their length and centromere position. Bar represents 5 μm
Lack of epigenetic differentiation of the sex chromosomes in females
The synaptic delay of the sex chromosomes in female meiosis prompted us to look at whether some of the epigenetic and transcriptional signatures found on the sex chromosomes in males are also present in females. We found that, in addition to γH2AX (Fig. 8a), SUMO-1 is localized to the unsynapsed segments of the sex chromosomes at early pachytene (Fig. 8b). By contrast, ubiH2A does not accumulate on either unsynapsed autosomes or sex chromosomes (Fig. 8c). In terms of histone modifications and contrary to the situation in males, both H3K9me3 (Fig. 8d) and H3K4me (Fig. 8e) were observed throughout the whole nucleus at early pachytene, with no evidence of their differential accumulation or exclusion on either of the X chromosomes before synapsis completion, revealing a lack of epigenetic differentiation between the two chromosomes. Interestingly, at early pachytene, RNA pol-II also covers the whole nucleus, including the partially synapsed sex chromosomes (Fig. 8f). This result reveals, as in males, that transcriptional activity is high in oocytes from the beginning of the pachytene stage and, more relevantly, that sex chromosomes do not apparently show a differential transcriptional status compared to autosomes.
Immunolocalization of epigenetic markers in the sex chromosomes of E. tancrei pachytene oocytes. a–f Early pachytene oocytes. Only partially synapsed sex chromosomes are observed at this stage. g–l Mid-late pachytene oocytes. Fully synapsed sex chromosomes are observed at this stage. a SYPC3 (green) and γH2AX (red). The unsynapsed regions of the sex chromosomes are marked by a γH2AX signal. b SYPC3 (green) and SUMO-1 (red) in the same cell as in a. Note that SUMO-1 localizes to the same unsynapsed region of the sex chromosomes, although it does not completely colocalize with the γH2AX signal. c SYPC3 (green) and ubiH2A (red). ubiH2A does not accumulate in the nucleus nor in either the sex chromosomes (XX) or unsynapsed autosomes (arrow). d SYPC3 (green) and H3K9me3 (red). H3K9me3 labeling is observed throughout the nucleus. The sex chromosomes (XX) do not show any specific accumulation compared with the autosomes or with each other. e SYPC3 (green) and H3K4me (red). H3K4me labeling is observed throughout the nucleus, and no specific accumulations are observed in the sex chromosomes (XX). f SYPC3 (green) and RNA polymerase-II (red). The whole nucleus, including the sex chromosomes (XX), is intensely labeled with RNA polymerase-II. g SYPC3 (green) and γH2AX (red). Faint γH2AX foci remain associated with some bivalents, but none are specific to any particular chromosome. h SYPC3 (green) and SUMO-1 (red). SUMO-1 does not specifically label any of the chromosomes at this stage. i SYPC3 (green) and ubiH2A (red). Only small nonspecific ubiH2A foci are observed scattered throughout the nucleus. j SYPC3 (green) and H3K9me3 (red). H3K9me3 labeling is observed throughout the nucleus, with some accumulation observed in specific regions that likely correspond to centromeres. k SYPC3 (green) and H3K4me (red). H3K4me labeling is observed throughout the nucleus, and no specific accumulations are observed. l SYPC3 (green) and RNA polymerase-II (red). The whole nucleus, including the sex chromosomes, is intensely labeled with RNA polymerase-II. Bar represents 10 μm
After sex chromosome synapsis completion, γH2AX and SUMO-1 no longer specifically associate with the sex chromosomes. At mid-late pachytene, large diffuse γH2AX foci remain associated with some bivalents (Fig. 8g), while SUMO-1 is no longer associated with any chromosomes (Fig. 8h). These observations suggest that the accumulation of these proteins on the sex chromosomes at early pachytene is most probably a response to asynapsis, rather than a specific mark of sex chromosomes. Similar to the pattern at early pachytene, we also did not observe any specific ubiH2A staining (Fig. 8i) or differential localization of H3K9me3 (Fig. 8j), H3K4me (Fig. 8k), or RNA pol-II (Fig. 8l) on the sex chromosomes, which, by this stage, are indistinguishable from autosomes.
Recombination distribution is restricted in both male and female meiosis
Finally, we analyzed the distribution of recombination events in both males and females on the basis of MLH1 immunolocalization, which is widely used to estimate chiasmata location (Anderson et al. 1999; Froenicke et al. 2002); cells were also immunostained with SYCP3 and centromeric proteins (Fig. 9a–j). In males, the mean number of MLH1 foci per spermatocyte was 27.29±2.39 (n=49 spermatocytes). The mean value per sex bivalent was 1.33 (n=54). Consistent with the restriction of synapsis to the distal regions of the sex chromosomes, MLH1 foci largely appear limited to the same regions (Fig. 9k). We observed three chiasmatic configurations in the sex chromosomes: the most common was a single focus in the distal end (68.52%, n=37) (Fig. 9b); followed by two foci, one at each end (31.48%, n=17) (Fig. 9c); and very rarely, a single focus in the proximal end (1.85%, n=1) (Fig. 9d). MLH1-negative sex bivalents were also found, most likely in cells already at late pachytene stages when the protein has already largely detached from many chromosomes (Fig. 9e). Differences in staging likely account for the disparity in the number of MLH1 foci found between this study and a parallel study with 2n=34 males in which fewer foci were observed (Matveevsky et al. 2020). MLH1 foci distribution, along with the unsynapsed state of the central region of the sex chromosomes, clearly indicates that this region never recombines during male meiosis.
Localization and distribution of MLH1 in E. tancrei pachytene spermatocytes (a–e) and oocytes (f–j). SYPC3 (blue), MLH1 (green), centromeres (red). a In males, large bivalents tend to present two or three MLH1 foci, while short bivalents usually present a single focus. Sex chromosomes can show a single focus located at the distal end (b), two foci, one at each end (c), or rarely, a single focus at the proximal end (d). MLH1 is not observed on the sex chromosomes at late pachytene (e). f Females present up to four MLH1 foci in the large bivalents. Loading of MLH1 to the chromosomes can take place before synapsis completion in some autosomes (arrow) and in the sex chromosomes (XX) (enlarged in g). Sex chromosomes present a single MLH1 focus on the sex bivalent, located in the proximal (h) or distal (i) region. Only occasionally are two MLH1 foci observed (j). k For the analysis of MLH1 foci distribution along the sex bivalent, the X chromosome was divided into 10 equal segments, and the position of each MLH1 focus was assigned to a discrete segment. In males (blue), MLH1 foci are only distributed at the distal regions of the chromosomes (segments 1, 2, 9, and 10). In females (red), we observed that MLH1 foci follow a bimodal distribution, with the exclusion of foci in the central region of the chromosomes (segments 5 and 6) and a low frequency in the vicinity of the centromere (segment 1). Bar represents 10 μm
The dynamics of MLH1 in females uncovered some unexpected features. First, the appearance of MLH1 foci was observed at very early pachytene. Indeed, oocytes can reach peak levels of MLH1 even before some chromosomes, particularly the sex chromosomes, have completed synapsis (Fig. 9f, g). As in males, one or two MLH1 foci were observed on the sex chromosomes in females (Fig. 9g–j); however, the mean number of foci per sex bivalent was lower than in males (1.13, n=45). The lower proportion of foci in females is not due to a lower recombination rate as the mean number of MLH1 foci, 28.83±3.09 (n=30 oocytes), was significantly higher than in males (t-test, p= 0.0133). To ascertain the distribution of MLH1 along the sex bivalent, we divided it into ten equivalent segments, and the frequency of foci in each was recorded in 45 oocytes from two females. We found that foci are not evenly distributed along the XX bivalent (Fig. 9k). Instead, they show a marked bimodal distribution, accumulating at the proximal and distal regions and being completely absent from the central region (segments 5 and 6). Although the number of cells analyzed is low, and the data need to be corroborated with more individuals, this situation indicates that recombination is highly reduced or absent in the central region of the X chromosomes, which is somewhat reminiscent of that found in males. This result cannot be attributed to strong chiasmata interference since most X bivalents only showed a single chiasma. Instead, this restriction is most likely related with the early appearance of MLH1 foci and delayed synapsis in the central region.
Discussion
Sex chromosome evolution is one of the most interesting topics in genetics (Bachtrog 2013; Waters and Ruiz-Herrera 2020). The degeneration of the Y chromosome and its putative disappearance, particularly in mammals, have fueled many heated debates (Graves 2004, 2006; Griffin 2012; Hughes et al. 2012). Yet, a handful of species have reached this evolutionary point and have managed to survive without a Y chromosome (Bakloushinskaya and Matveevsky 2018). Moreover, some Ellobius species have gone a step further and have doubled the X chromosome, giving rise to a new configuration comprising two isomorphic X chromosomes in both males and females. The implication of these chromosomes in sex determination still needs to be proven. However, several studies of meiosis have indicated that they already behave like sex chromosomes, at least in males (Kolomiets et al. 1991; Matveevsky et al. 2016, 2017). In this study, we add new evidence to this proposal and reveal unexpected sex differences that offer new ways of understanding the origin and fate of sex chromosomes in this group (Fig. 10). Moreover, our results highlight the relevance of meiotic studies to understand sex chromosome evolution (Gil-Fernández et al. 2020).
a Schematic representation of sex chromosome organization during male and female meiosis in E. tancrei. In males, sex chromosomes only synapse and recombine near the chromosomal ends. The interstitial regions remain unsynapsed. Given the lack of recombination, these regions are likely in a process of divergence. Both chromosomes are included in an inactive chromatin mass (stained in pink) as a result of MSCI. One X chromosome shows distinctive epigenetic marks, represented by the accumulation of H3K9me3 (in brown), and an association with the chromatin body (represented as a white circle). This chromosome is probably undergoing a differentiation process from the other X chromosome and has been indicated as a putative neo-Y chromosome. In females, X chromosome synapsis is complete, although delayed, and recombination occurs along the entire bivalent, except in the central segment (depicted in dark blue in one of the chromosomes). No MSCI is detected after the completion of synapsis, and no epigenetic differentiation of either sex chromosome is observed. b A hypothetical evolutionary trajectory of sex chromosomes in E. tancrei (putatively in E. talpinus and E. alaicus as well). The ancestor of all Ellobius species would have had an XX/XY sex determination system, which is still present in E. fuscocapillus. Afterward, the loss of the Y chromosome in males and one X in females would have originated an X0/X0 system, like in E. lutescens. Doubling of the X chromosome in both sexes would have originated an XX/XX system. The new X chromosome might have presented some small differences with the existing one already present. In males, the requirements for MSCI would have soon thereafter induced a restriction of synapsis and recombination, rapidly triggering the differentiation of one of the X chromosomes into a neo-Y. Currently, this differentiation has not reached the centromere, as the proximal region still recombines. In females, synapsis could have been completed soon after X chromosome doubling, owing to more relaxed meiotic controls and the absence of MSCI. Recombination between the two X chromosomes would have promoted the exchange of sequences and their homogenization, although this step is still not complete in E. tancrei. In a future step, the sex chromosomes in males could advance to a more differentiated state, with the inclusion of centromere of the neo-Y chromosome in the non-recombining region. In females, the X chromosomes would eventually be completely homologous.
Synapsis of two X chromosomes is partial in males
From the data presented here and by previous works (Kolomiets et al. 1991), it is clear that synapsis between the X chromosomes in E. tancrei males only occurs in the distal portions. Although the length of the synapsed region is variable during pachytene, the central portion of the chromosomes never shows signs of synapsis. By contrast, in E. tancrei females, the X chromosomes achieve complete (although delayed) synapsis. This difference, which may be driven by structural, genetic, and/or epigenetic factors, indicates a clear sex divergence of X chromosome behavior during meiosis (Fig. 10a).
Incomplete synapsis is one of the hallmarks of sex chromosome behavior during male meiosis in species with highly differentiated X and Y chromosomes (Solari 1970, 1974, 1993). This is due to the drastic difference in size between the two chromosomes. Synapsis is only homologous in the short PAR. Therefore, a first explanation for incomplete synapsis between X chromosomes in E. tancrei would be that, albeit similar in size and morphology, the two X chromosomes are already genetically differentiated, and they do not share enough homology to stimulate complete synapsis. Unfortunately, genetic studies do not provide to date any conclusive evidence of this differentiation (Mulugeta et al. 2016). Nevertheless, this could not be a complete explanation. In most mammalian species, synapsis between the X and Y chromosomes usually extends to the heterologous segments. In the house mouse, mole rat, and southern mole vole, for instance, synapsis occurs over almost the complete length of the Y chromosome during early pachytene (Matveevsky et al. 2018, 2017; Page et al. 2012). Therefore, the restriction of synapsis between the isomorphic X chromosomes in E. tancrei males raises further questions. This behavior would be expected in cases where there is some kind of physical or genetic impediment. Centromeres are usually one of these barriers, as it occurs in the house mouse (Page et al. 2012; Solari 1974), although there are cases in which the heterologous synapsis can extend past the centromeric region of the Y chromosome, such as in Arvicola terrestris (de la Fuente et al. 2012). In the case of E. tancrei, there is no such evident impediment since the centromere is located at the end of the chromosome. Another type of structural impairment could be caused by an inversion in one of the two chromosomes. In the house mouse, the presence of inversions has been shown to significantly delay synapsis. However, chromosomes are usually able to complete synapsis through an adjustment process that involves the formation of a loop in one of the chromosomes and, finally, the establishment of heterologous associations within the bivalent (Moses et al. 1982). In the case of E. tancrei, inversions also do not seem to be the cause of the synaptic restriction as the size and the G-band pattern of both X chromosomes in males are identical (Vorontsov et al. 1980). The formation of a synapsing loop on any of the X chromosomes, however, has not been observed during the pachytene stage, though it has been observed that one of the AEs usually appears fragmented. This does not appear to be due to a twisting of the AE but rather to the presence of the ChB. The ChB is clearly not related to the nucleolus, but the deposition of proteins and the presence of specific epigenetic modifications in this body, particularly the accumulation of H3K9me3, could be related to the fragmented appearance of one of the sex chromosomes. Thus, although the ChB usually appears in an interstitial position, far from the point of maximum extension of the SC, it is possible that the epigenetic modifications, along with the interruption of the AE, could constitute a structural impediment to synapsis extension.
In addition to structural factors, genetic or epigenetic mechanisms may be involved in the restriction of synapsis between sex chromosomes. There are examples of species in which similar phenomena have been found, mainly in insects. In the grasshopper Stethophyma grossum, several chromosomes synapse only in regions close to centromeres (Calvente et al. 2005). Synapsis restriction has been attributed to the absence of DNA break events in those regions and to delayed maturation of the assembly of the cohesin axes and AEs. A similar phenomenon has also been described in the grasshopper Paratettix meridionalis (Viera et al. 2009). Interestingly, the artificial induction of breaks is insufficient to trigger synapsis in these species (Calvente et al. 2016), indicating the existence of a different control of synapsis or additional structural requirements, such as an association of proteins that regulate chromatin conformation and SC formation. A similar though more complex control mechanism could be acting in E. tancrei, since DNA DSBs, as marked by the presence of RPA and RAD51, are observed in the asynaptic segments of the X chromosomes.
Finally, it can be argued that completion of synapsis of the X chromosome would prevent it from undergoing MSCI. In the house mouse, abolition of MCSI leads to meiotic impairment, mostly attributed to the expression of ZFY1/2 genes present on the Y chromosome (Royo et al. 2010). However, completion of synapsis of the X chromosome also leads to male meiotic arrest (Turner et al. 2006), which could be due to the expression of X-linked genes, some of which could also potentially trigger an impairment of meiotic progression. Moreover, in the case of E. tancrei, we cannot exclude the possibility that the Zfy gene has been translocated to the X chromosome, as was demonstrated for E. lutescens (Mulugeta et al. 2016). Considering this scenario, a reasonable assumption is that synapsis between the X chromosomes is conditioned by genetic and epigenetic factors acting during mole vole male meiosis.
Synapsis of X chromosomes is delayed in females
One of the most intriguing results found in this study is the delay in synapsis of the X chromosomes during female meiosis. Given the lack of any structural or epigenetic differentiation between the two X chromosomes, we hypothesize that the most likely explanation is that the interstitial segments of the X chromosomes in females are not sufficiently homologous for efficient synapsis. This is supported by the fact that, in females, MLH1 foci are not found in the central segment of the X chromosomes. However, the X chromosomes do ultimately complete synapsis in females, which may be due to the existence of more relaxed synaptic control mechanisms during female meiosis, as has been reported in other mammals (Kouznetsova et al. 2009). It could also be argued that completion of synapsis of the sex chromosomes would lead to the avoidance of an MSCI response. Contrary to males, in which MSCI is necessary for meiotic progression (Burgoyne et al. 2009; Turner et al. 2005), MSCI in females could have detrimental consequences. In this way, completion of synapsis, albeit putatively heterologous, could serve as a mechanism to ensure the outcome of meiosis. Such a scenario implies important consequences for the evolution of the X chromosomes in Ellobius (see later), as recently proposed for antelopes (Robinson et al. 2021).
Epigenetic differentiation of the sex chromosomes is found only in males
The specific accumulation of H3K9me3 along one of the X chromosomes and the presence of a ChB that accumulates SUMO-1 (Matveevsky et al. 2016) but excludes other proteins, such as γH2AX and H3K4me, is indicative of special DNA packaging and epigenetic modification processes. The heterochromatic nature of the ChB probably determines the ability to associate with other intranuclear heterochromatin-like structures and unsynapsed chromosome segments, as shown in some heterozygotes for Robertsonian translocations (Matveevsky et al. 2016, 2015, 2020). In any case, epigenetic differentiation of the XX chromosomes would be evident only in male meiosis as no specific differences have been found in somatic cells or female meiosis. The origin of these differences is not clear. Differential expression of genes between the two chromosomes can be ruled out since the absence of transcription markers such as RNA pol-II similarly affects both X chromosomes. Although the C- and G-banding patterns of XX are identical in somatic cells, the differential organization of the chromatin during meiosis, particularly the accumulation of the heterochromatin marker H3K9me3, suggests genomic differences. Nevertheless, in the absence of more conclusive karyotypic markers that could differentiate between the two X chromosomes, or genomic data about their gene content, this proposal is difficult to demonstrate. These hypothetical differences could help to explain the absence of synapsis and recombination in the central region of the X chromosomes in males (Fig. 10a). Moreover, the absence of epigenetic marks in either of the two sex chromosomes in females could indicate that the one bearing such marks represents an X chromosome version that is currently transmitted only in males. These would represent some of the first signs of the transformation of this X chromosome into a neo-Y (Fig. 10b), i.e., secondary heteromorphization, as has been previously suggested (Matveevsky et al. 2016). As an alternative interpretation, the accumulation of epigenetic markers and the ChB in one of the X chromosomes could just be a stochastic process with no relation with chromosome differentiation. Although we do not favor this interpretation, we realize it would imply an unreported and intriguing behavior of chromosomes during meiosis.
Recombination is absent in the central region of the X chromosomes in both sexes
Cessation of recombination is the hallmark of sex chromosome differentiation (Bachtrog 2006; Charlesworth and Charlesworth 2000; Charlesworth et al. 2005). It leads to the genetic isolation of the Y chromosome, which, with time, accumulates mutations and starts degenerating (Bachtrog 2013). Our results on synapsis and epigenetic modifications suggest that one of the X chromosomes in E. tancrei males is undergoing a process of differentiation. The loss of recombination in the central part of the XX chromosomes would have served as a starting point for the divergence of initially homologous sex chromosomes. The fact that the two X chromosomes in males are indistinguishable indicates that morphological differentiation has not yet occurred. This case is not an exception. In the African pygmy mouse Mus minutoides and the Okinawa spiny rat Tokudaia muenninki, recombination has stopped in a large portion of their neo-sex chromosomes without any signs of morphological differentiation (Gil-Fernández et al. 2020; Murata et al. 2015). However, the situation found in E. tancrei introduces additional factors to the standard models of sex chromosome differentiation. First, although restriction of chiasmata to the ends of the X chromosome in male meiosis could be indicative of advanced differentiation of a neo-Y (the non-recombining segment covers 80% of the chromosome; (Matveevsky et al. 2016), the presence of a chiasma in the proximal region of the X chromosome complicates this scenario. Recombination in this region, together with the distal position of the centromere, implies that, although the central region of the X chromosome behaves like a linkage group, the pericentromeric region is not linked to it and indeed the centromere could be exchanged with the X chromosome (Fig. 10b). It is possible that chromosome differentiation is occurring in the pericentromeric region, since only 30% of the X bivalents recombine in the proximal segment; however, differentiation has obviously not reached the centromere yet.
In this scenario, we expected that a recombination restriction phenomenon would occur only in males. However, we unexpectedly found that recombination is also restricted in the central region of the X chromosomes in females. Although the non-recombining region is shorter in females, this fact reveals the existence of a linkage group in the middle of the X chromosome. As mentioned above, this may be a consequence of inefficient homology between sex chromosomes in this region, which would cause a synapsis delay that, in turn, affects the resolution of recombination intermediates (Barchi et al. 2005; Costa et al. 2005; de Vries et al. 2005), particularly considering that MLH1 incorporation at crossovers in E. tancrei females can take place prior to the completion of synapsis.
An evolutionary scenario for the emergence of a new, still forming, XX/XY sex chromosome system from the XX/XX system in Ellobius
The absence of any conclusive molecular data on the constitution of the X chromosomes and the sex determination mechanisms in Ellobius makes it very difficult to draw an evolutionary trajectory of the sex chromosomes in the three species with an XX pair in both males and females. However, studies of their meiotic behavior could shed light on two specific aspects: the intermediate sex chromosome system and the evolutionary fate of the neo-Y (differentiating X) chromosome in males (Fig. 10b).
Two main scenarios are possible for the composition of the intermediate sex chromosome system. First, after losing the Y chromosome, males would have become X0, while females would have stayed XX. Although this is apparently the most parsimonious scenario, it does not seem to be the most plausible. This chromosome configuration is common in insects but not in mammals, likely due to the fact that X0 individuals usually develop as females (Fredga and Bulmer 1988). Only a few African bat species have been reported to display an XX/X0 chromosome system (Denys et al. 2013; Peterson and Nagorsen 1975); however, nothing is known about their sex determination mechanisms. The second scenario consists of an X0/X0 system. The emergence of such a system may seem odd as it would mean that both males and females would have the same chromosomal constitution. Moreover, under this system, both XX and 00 individuals would not be viable, signifying a 50% embryonic mortality rate (Lyapunova et al. 1975). However, this system has been described in the related species E. lutescens and also in the Ryukyu spiny rat Tokudaia osimensis (Fredga and Bulmer 1988; Kuroiwa et al. 2010; Vorontsov et al. 1980). Overall, our results on the meiotic behavior of sex chromosomes in E. tancrei are more aligned with the second scenario. If males in an intermediate stage were X0 and females XX, then once the X chromosome doubled, we would expect modification of sex chromosome behavior to occur only in males. However, we found that synapsis and recombination are also affected in females. A possible explanation to account for this observation is that the new X chromosome presented some subtle differences with the existing X chromosome in X0 males and females (Fig. 10b). These differences were then buffered in female meiosis, owing to its more relaxed mechanisms of synapsis control (Kouznetsova et al. 2009), gradually leading to a complete homogenization of both X chromosomes (Fig. 10b). As a result, the non-recombining region in females would be a relic of a more extended region in the hypothetical ancestor. Indeed, tracing of evolutionary chromosome rearrangements on the X chromosome of E. talpinus has revealed the presence of ancestral relics of interstitial rearrangements from the original arvicolid X chromosome (Romanenko et al. 2020).
The subsequent evolution of the X chromosomes in males could be greatly accelerated by the specific conditions of male meiosis. First, the absence of synapsis and, more relevantly, recombination in the central region of the X chromosomes would lead to the genetic isolation of one of the X chromosomes, provided it is transmitted through males only. The epigenetic marker analyses presented here and previously (Matveevsky et al. 2016) seem to support this proposal. We further hypothesize that this chromosome will experience deepening and increasing differentiation and will turn into a neo-Y as a result of secondary heteromorphization (Fig. 10b), even if it does not bear any male-determining factor, such as in the case of the Y chromosome in Drosophila. Second, the evolution and degeneration of the Y chromosome are usually accompanied by coevolution of the X chromosome gene content (Bachtrog 2020; Waters and Ruiz-Herrera 2020). As the sex chromosomes start to diverge, the appearance of unsynapsed regions will trigger MSCI, inducing the meiotic inactivation of genes on both the X and Y chromosomes. It has been shown that, during evolution, mammalian X chromosomes have been emptied of genes expressed in meiosis, which have been translocated or retrotransposed to autosomes (Khil et al. 2004; McKee and Handel 1993). In the case of the E. tancrei X chromosome, this step has already been accomplished. Indeed, the inactivation of the X chromosome could be a requisite for the successful completion of meiosis. Therefore, the putative degeneration of the neo-Y chromosome could occur more rapidly than expected, as simultaneous coevolution of the X chromosome would be no longer needed.
Overall, our analysis of the behavior of sex chromosomes during meiosis illustrates that this cell division introduces important constraints on the evolution of sex chromosomes, as it is becoming more and more clear (Bakloushinskaya and Matveevsky 2018; de la Fuente et al. 2012; Gil-Fernández et al. 2020; Page et al. 2005; Waters and Ruiz-Herrera 2020). The case of Ellobius species with an XX/XX sex chromosome system reveals that the replacement of a pair of sex chromosomes by a different chromosome pair, which is common in other vertebrates and also invertebrates (Bachtrog et al. 2014; Jeffries et al. 2018; Pennell et al. 2018), might not be as easy in mammals. The X chromosomes left behind would likely have to still accomplish meiotic requirements, like MSCI, leading to impaired synapsis and recombination. This, in turn, could lead to the recurrent differentiation of one of the members of the pair into a Y chromosome, whether it bears a sex determination factor or not. We hypothesize that meiotic requirements in other vertebrates are more relaxed, allowing for an easier turnover of sex chromosomes. For instance, meiotic studies in birds have revealed that MSCI does not operate as in eutherian mammals (Guioli et al. 2012; Pigozzi 2016). In monotreme mammals, which present an extraordinary sex chromosome system, with 5 X and 5 Y chromosomes, MSCI does not operate at all (Daish et al. 2015). Given the wide variety of sex chromosome systems, meiotic studies of other vertebrate species are critical to address the many aspects of sex chromosome evolution.
References
Acquaviva L, Boekhout M, Karasu ME, Brick K, Pratto F, Li T, van Overbeek M, Kauppi L, Camerini-Otero RD, Jasin M, Keeney S (2020) Ensuring meiotic DNA break formation in the mouse pseudoautosomal region. Nature 582:426–431. https://doi.org/10.1038/s41586-020-2327-4
Anderson LK, Reeves A, Webb LM, Ashley T (1999) Distribution of crossing over on mouse synaptonemal complexes using immunofluorescent localization of MLH1 protein. Genetics 151:1569–1579
Arakawa Y, Nishida-Umehara C, Matsuda Y, Sutou S, Suzuki H (2002) X-chromosomal localization of mammalian Y-linked genes in two XO species of the Ryukyu spiny rat. Cytogenetic and Genome Research 99:303–309
Ashley T, Jaarola M, Fredga K (1989) Absence of synapsis during pachynema of the normal sized sex chromosomes of Microtus arvalis. Hereditas 111:295–304
Baarends WM, Wassenaar E, van der Laan R, Hoogerbrugge J, Sleddens-Linkels E, Hoeijmakers JH, de Boer P, Grootegoed JA (2005) Silencing of unpaired chromatin and histone H2A ubiquitination in mammalian meiosis. Mol Cell Biol 25:1041–1053
Bachtrog D (2006) A dynamic view of sex chromosome evolution. Curr Opin Genet Dev 16:578–585
Bachtrog D (2013) Y-chromosome evolution: emerging insights into processes of Y-chromosome degeneration. Nat Rev Genet 14:113–124
Bachtrog D (2020) The Y chromosome as a battleground for intragenomic conflict. Trends Genet 36:510–522
Bachtrog D, Mank JE, Peichel CL, Kirkpatrick M, Otto SP, Ashman T-L, Hahn MW, Kitano J, Mayrose I, Ming R, Perrin N, Ross L, Valenzuela N, Vamosi JC, The Tree of Sex C (2014) Sex determination: why so many ways of doing it? PLoS Biol 12:e1001899
Bagheri-Fam S, Sreenivasan R, Bernard P, Knower K, Lovell-Badge R, Just W, Harley V (2012) Sox9 gene regulation and the loss of the XY/XX sex-determining mechanism in the mole vole Ellobius lutescens. Chromosom Res 20:191–199
Bakloushinskaya I, Matveevsky S (2018) Unusual ways to lose a Y chromosome and survive with changed autosomes: a story of mole voles Ellobius (Mammalia, Rodentia). OBM Genetics 2
Bakloushinskaya IY, Matveevsky SN, Romanenko SA, Serdukova NA, Kolomiets OL, Spangenberg VE, Lyapunova EA, Graphodatsky AS (2012) A comparative analysis of the mole vole sibling species Ellobius tancrei and E. talpinus (Cricetidae, Rodentia) through chromosome painting and examination of synaptonemal complex structures in hybrids. Cytogenet Genome Res 136:199–207
Bakloushinskaya I, Lyapunova EA, Saidov AS, Romanenko SA, O'Brien PCM, Serdyukova NA, Ferguson-Smith MA, Matveevsky S, Bogdanov AS (2019) Rapid chromosomal evolution in enigmatic mammal with XX in both sexes, the Alay mole vole Ellobius alaicus Vorontsov et al., 1969 (Mammalia, Rodentia). Comp Cytogenet 13:147–177
Barchi M, Mahadevaiah S, Di Giacomo M, Baudat F, de Rooij DG, Burgoyne PS, Jasin M, Keeney S (2005) Surveillance of different recombination defects in mouse spermatocytes yields distinct responses despite elimination at an identical developmental stage. Mol Cell Biol 25:7203–7215
Bellott DW, Hughes JF, Skaletsky H, Brown LG, Pyntikova T, Cho TJ, Koutseva N, Zaghlul S, Graves T, Rock S, Kremitzki C, Fulton RS, Dugan S, Ding Y, Morton D, Khan Z, Lewis L, Buhay C, Wang Q, Watt J, Holder M, Lee S, Nazareth L, Alfoldi J, Rozen S, Muzny DM, Warren WC, Gibbs RA, Wilson RK, Page DC (2014) Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature 508:494–499
Boekhout M, Karasu ME, Wang J, Acquaviva L, Pratto F, Brick K, Eng DY, Xu J, Camerini-Otero RD, Patel DJ, Keeney S (2019) REC114 partner ANKRD31 controls number, timing, and location of meiotic DNA breaks. Mol Cell 74:1053–1068.e1058
Borodin PM, Basheva EA, Torgasheva AA, Dashkevich OA, Golenishchev FN, Kartavtseva IV, Mekada K, Dumont BL (2012) Multiple independent evolutionary losses of XY pairing at meiosis in the grey voles. Chromosom Res 20:259–268
Britton-Davidian J, Robinson TJ, Veyrunes F (2012) Systematics and evolution of the African pygmy mice, subgenus Nannomys: A review. Acta Oecol 42:41–49
Burgoyne PS (1982) Genetic homology and crossing over in the X and Y chromosomes of mammals. Hum Genet 61:85–90
Burgoyne PS, Mahadevaiah SK, Turner JM (2009) The consequences of asynapsis for mammalian meiosis. Nat Rev Genet 10:207–216
Calvente A, Viera A, Page J, Parra MT, Gomez R, Suja JA, Rufas JS, Santos JL (2005) DNA double-strand breaks and homology search: inferences from a species with incomplete pairing and synapsis. J Cell Sci 118:2957–2963
Calvente A, Santos JL, Rufas JS (2016) Do exogenous DNA double-strand breaks change incomplete synapsis and chiasma localization in the grasshopper Stethophyma grossum? PLoS One 11:e0168499
Carnero A, Jimenez R, Burgos M, Sanchez A, Diaz de la Guardia R (1991) Achiasmatic sex chromosomes in Pitymys duodecimcostatus: mechanisms of association and segregation. Cytogenet Cell Genet 56:78–81
Charlesworth B, Charlesworth D (2000) The degeneration of Y chromosomes. Philos Trans R Soc Lond Ser B Biol Sci 355:1563–1572
Charlesworth D, Charlesworth B, Marais G (2005) Steps in the evolution of heteromorphic sex chromosomes. Heredity 95:118–128
Cortez D, Marin R, Toledo-Flores D, Froidevaux L, Liechti A, Waters PD, Grutzner F, Kaessmann H (2014) Origins and functional evolution of Y chromosomes across mammals. Nature 508:488–493
Costa Y, Speed R, Ollinger R, Alsheimer M, Semple CA, Gautier P, Maratou K, Novak I, Hoog C, Benavente R, Cooke HJ (2005) Two novel proteins recruited by synaptonemal complex protein 1 (SYCP1) are at the centre of meiosis. J Cell Sci 118:2755–2762
Cowell IG, Aucott R, Mahadevaiah SK, Burgoyne PS, Huskisson N, Bongiorni S, Prantera G, Fanti L, Pimpinelli S, Wu R, Gilbert DM, Shi W, Fundele R, Morrison H, Jeppesen P, Singh PB (2002) Heterochromatin, HP1 and methylation at lysine 9 of histone H3 in animals. Chromosoma 111:22–36
Daish TJ, Casey AE, Grutzner F (2015) Lack of sex chromosome specific meiotic silencing in platypus reveals origin of MSCI in therian mammals. BMC Biol 13:106
de la Fuente R, Parra MT, Viera A, Calvente A, Gomez R, Suja JA, Rufas JS, Page J (2007) Meiotic pairing and segregation of achiasmate sex chromosomes in eutherian mammals: the role of SYCP3 protein. PLoS Genet 3:e198
de la Fuente R, Sanchez A, Marchal JA, Viera A, Parra MT, Rufas JS, Page J (2012) A synaptonemal complex-derived mechanism for meiotic segregation precedes the evolutionary loss of homology between sex chromosomes in arvicolid mammals. Chromosoma 121:433–446
de Vries FA, de Boer E, van den Bosch M, Baarends WM, Ooms M, Yuan L, Liu JG, van Zeeland AA, Heyting C, Pastink A (2005) Mouse Sycp1 functions in synaptonemal complex assembly, meiotic recombination, and XY body formation. Genes Dev 19:1376–1389
Denys C, Kadjo B, Missoup AD, Monadjem A, Aniskine V (2013) New records of bats (Mammalia: Chiroptera) and karyotypes from Guinean Mount Nimba (West Africa). Ital J Zool 80:279–290
Fredga K, Bulmer MG (1988) Aberrant chromosomal Sex-determining mechanisms in mammals, with special reference to species with XY females [and discussion]. Philosophical Transactions of the Royal Society of London B, Biological Sciences 322:83–95
Froenicke L, Anderson LK, Wienberg J, Ashley T (2002) Male mouse recombination maps for each autosome identified by chromosome painting. Am J Hum Genet 71:1353–1368
Gil-Fernández A, Saunders PA, Martín-Ruiz M, Ribagorda M, López-Jiménez P, Jeffries DL, Parra MT, Viera A, Rufas JS, Perrin N, Veyrunes F, Page J (2020) Meiosis reveals the early steps in the evolution of a neo-XY sex chromosome pair in the African pygmy mouse Mus minutoides. PLoS Genet 16:e1008959
Goetz P, Chandley AC, Speed RM (1984) Morphological and temporal sequence of meiotic prophase development at puberty in the male mouse. J Cell Sci 65:249–263
Graves JA (1995) The origin and function of the mammalian Y chromosome and Y-borne genes--an evolving understanding. Bioessays 17:311–320
Graves JA (1996) Mammals that break the rules: genetics of marsupials and monotremes. Annu Rev Genet 30:233–260
Graves JAM (2004) The degenerate Y chromosome can conversion save it? Reprod Fertil Dev 16:527–534
Graves JAM (2006) Sex chromosome specialization and degeneration in mammals. Cell 124:901–914
Graves JA, Wakefield MJ, Toder R (1998) The origin and evolution of the pseudoautosomal regions of human sex chromosomes. Hum Mol Genet 7:1991–1996
Griffin D (2012) Is the Y chromosome disappearing?—Both sides of the argument. Chromosom Res 20:35–45
Guioli S, Lovell-Badge R, Turner JMA (2012) Error-prone ZW pairing and no evidence for meiotic sex chromosome inactivation in the chicken germ line. PLoS Genet 8:e1002560
Honda T, Suzuki H, Itoh M (1977) An unusual sex chromosome constitution found in the Amami spinous country-rat, Tokudaia osimensis osimensis. The Japanese journal of genetics 52:247–249
Honda T, Suzuki H, Itoh M, Hayashi K (1978) Karyotypical differences of the Amami spinous country-rats, Tokudaia osimensis osimensis, obtained from two neighbouring islands. The Japanese journal of genetics 53:297–299
Hughes JF, Skaletsky H, Brown LG, Pyntikova T, Graves T, Fulton RS, Dugan S, Ding Y, Buhay CJ, Kremitzki C, Wang Q, Shen H, Holder M, Villasana D, Nazareth LV, Cree A, Courtney L, Veizer J, Kotkiewicz H, Cho T-J, Koutseva N, Rozen S, Muzny DM, Warren WC, Gibbs RA, Wilson RK, Page DC (2012) Strict evolutionary conservation followed rapid gene loss on human and rhesus Y chromosomes. Nature 483:82–86
Jeffries DL, Lavanchy G, Sermier R, Sredl MJ, Miura I, Borzée A, Barrow LN, Canestrelli D, Crochet P-A, Dufresnes C, Fu J, Ma W-J, Garcia CM, Ghali K, Nicieza AG, O’Donnell RP, Rodrigues N, Romano A, Martínez-Solano Í, Stepanyan I, Zumbach S, Brelsford A, Perrin N (2018) A rapid rate of sex-chromosome turnover and non-random transitions in true frogs. Nat Commun 9:4088
Just W, Rau W, Vogel W, Akhverdian M, Fredga K, Marshall Graves JA, Lyapunova E (1995) Absence of Sry in species of the vole Ellobius. Nat Genet 11:117–118
Kauppi L, Barchi M, Baudat F, Romanienko PJ, Keeney S, Jasin M (2011) Distinct properties of the XY pseudoautosomal region crucial for male meiosis. Science 331:916–920
Kauppi L, Jasin M, Keeney S (2012) The tricky path to recombining X and Y chromosomes in meiosis. Ann N Y Acad Sci 1267:18–23
Keeney S, Lange J, Mohibullah N (2014) Self-organization of meiotic recombination initiation: general principles and molecular pathways. Annu Rev Genet 48:187–214
Khil PP, Smirnova NA, Romanienko PJ, Camerini-Otero RD (2004) The mouse X chromosome is enriched for sex-biased genes not subject to selection by meiotic sex chromosome inactivation. Nat Genet 36:642–646
Kobayashi T, Yamada F, Hashimoto T, Abe S, Matsuda Y, Kuroiwa A (2007) Exceptional minute sex-specific region in the X0 mammal, Ryukyu spiny rat. Chromosom Res 15:175–187
Kolomiets OL, Vorontsov NN, Lyapunova EA, Mazurova TF (1991) Ultrastructure, meiotic behavior, and evolution of sex chromosomes of the genus Ellobius. Genetica 84:179–189
Kolomiets OL, Matveevsky SN, Bakloushinskaya IY (2010) Sexual dimorphism in prophase I of meiosis in the Northern mole vole (Ellobius talpinus Pallas, 1770) with isomorphic (XX) chromosomes in males and females. Comparative Cytogenetics 4:55–66
Kouznetsova A, Wang H, Bellani M, Camerini-Otero RD, Jessberger R, Hoog C (2009) BRCA1-mediated chromatin silencing is limited to oocytes with a small number of asynapsed chromosomes. J Cell Sci 122:2446–2452
Kuroiwa A, Ishiguchi Y, Yamada F, Shintaro A, Matsuda Y (2010) The process of a Y-loss event in an XO/XO mammal, the Ryukyu spiny rat. Chromosoma 119:519–526
Liu L, Franco S, Spyropoulos B, Moens PB, Blasco MA, Keefe DL (2004) Irregular telomeres impair meiotic synapsis and recombination in mice. Proc Natl Acad Sci U S A 101:6496–6501
Lyapunova EA, Vorontsov NN (1978) Genetics of Ellobius (Rodentia). I Karyological characteristics of four Ellobius species Genetika 14:2012–2024
Lyapunova EA, Vorontsov NN, Zakarjan GG (1975) Zygotic mortality in Ellobius lutescens (Rodentia: Microtinae). Experientia 31:417–418
Mahadevaiah SK, Turner JM, Baudat F, Rogakou EP, de Boer P, Blanco-Rodriguez J, Jasin M, Keeney S, Bonner WM, Burgoyne PS (2001) Recombinational DNA double-strand breaks in mice precede synapsis. Nat Genet 27:271–276
Matthey R (1953) La formule chromosomique et le problème de la détermination sexuelle chez Ellobius lutescens Thomas (Rodentia-Muridae-Microtinae). Arch Klaus-Stift VererbForsch 28:65–73
Matveevsky S, Bakloushinskaya I, Tambovtseva V, Romanenko S, Kolomiets O (2015) Analysis of meiotic chromosome structure and behavior in Robertsonian heterozygotes of Ellobius tancrei (Rodentia, Cricetidae): a case of monobrachial homology. Comp Cytogenet 9:691–706
Matveevsky S, Bakloushinskaya I, Kolomiets O (2016) Unique sex chromosome systems in Ellobius: how do male XX chromosomes recombine and undergo pachytene chromatin inactivation? Sci Rep 6:29949
Matveevsky S, Kolomiets O, Bogdanov A, Hakhverdyan M, Bakloushinskaya I (2017) Chromosomal evolution in mole voles Ellobius (Cricetidae, Rodentia): bizarre sex chromosomes, variable autosomes and meiosis. Genes (Basel) 8:306
Matveevsky S, Ivanitskaya E, Spangenberg V, Bakloushinskaya I, Kolomiets O (2018) Reorganization of the Y chromosomes enhances divergence in Israeli mole rats Nannospalax ehrenbergi (Spalacidae, Rodentia): comparative analysis of meiotic and mitotic chromosomes. Genes (Basel) 9:272
Matveevsky S, Tretiakov A, Kashintsova A, Bakloushinskaya I, Kolomiets O (2020) Meiotic nuclear architecture in distinct mole vole hybrids with Robertsonian translocations: chromosome chains, stretched centromeres, and distorted recombination. Int J Mol Sci 21(20):7630
McKee BD, Handel MA (1993) Sex chromosomes, recombination, and chromatin conformation. Chromosoma 102:71–80
Moens PB, Kolas NK, Tarsounas M, Marcon E, Cohen PE, Spyropoulos B (2002) The time course and chromosomal localization of recombination-related proteins at meiosis in the mouse are compatible with models that can resolve the early DNA-DNA interactions without reciprocal recombination. J Cell Sci 115:1611–1622
Moses MJ, Poorman PA, Roderick TH, Davisson MT (1982) Synaptonemal complex analysis of mouse chromosomal rearrangements. IV. Synapsis and synaptic adjustment in two paracentric inversions. Chromosoma 84:457–474
Mulugeta E, Wassenaar E, Sleddens-Linkels E, WFJ v IJ, Heard E, Grootegoed JA, Just W, Gribnau J, Baarends WM (2016) Genomes of Ellobius species provide insight into the evolutionary dynamics of mammalian sex chromosomes. Genome Res 26:1202–1210
Murata C, Kuroki Y, Imoto I, Tsukahara M, Ikejiri N, Kuroiwa A (2015) Initiation of recombination suppression and PAR formation during the early stages of neo-sex chromosome differentiation in the Okinawa spiny rat, Tokudaia muenninki. BMC Evol Biol 15:234
Murdoch B, Owen N, Stevense M, Smith H, Nagaoka S, Hassold T, McKay M, Xu H, Fu J, Revenkova E, Jessberger R, Hunt P (2013) Altered cohesin gene dosage affects mammalian meiotic chromosome structure and behavior. PLoS Genet 9:e1003241
Navarro J, Vidal F, Guitart M, Egozcue J (1981) A method for the sequential study of synaptonemal complexes by light and electron microscopy. Hum Genet 59:419–421
Ohno S (1967) Sex chromosomes and sex linked genes. Springer, Berlin
Page J, Berrios S, Rufas JS, Parra MT, Suja JA, Heyting C, Fernandez-Donoso R (2003) The pairing of X and Y chromosomes during meiotic prophase in the marsupial species Thylamys elegans is maintained by a dense plate developed from their axial elements. J Cell Sci 116:551–560
Page J, Berrios S, Parra MT, Viera A, Suja JA, Prieto I, Barbero JL, Rufas JS, Fernandez-Donoso R (2005) The program of sex chromosome pairing in meiosis is highly conserved across marsupial species: implications for sex chromosome evolution. Genetics 170:793–799
Page J, de la Fuente R, Manterola M, Parra MT, Viera A, Berrios S, Fernandez-Donoso R, Rufas JS (2012) Inactivation or non-reactivation: what accounts better for the silence of sex chromosomes during mammalian male meiosis? Chromosoma 121:307–326
Papanikos F, Clément JAJ, Testa E, Ravindranathan R, Grey C, Dereli I, Bondarieva A, Valerio-Cabrera S, Stanzione M, Schleiffer A, Jansa P, Lustyk D, Fei J-F, Adams IR, Forejt J, Barchi M, de Massy B, Toth A (2019) Mouse ANKRD31 regulates spatiotemporal patterning of meiotic recombination initiation and ensures recombination between X and Y sex chromosomes. Mol Cell 74:1069–1085.e1011
Pennell MW, Mank JE, Peichel CL (2018) Transitions in sex determination and sex chromosomes across vertebrate species. Mol Ecol 27:3950–3963
Peterson RL, Nagorsen DW (1975) Chromosomes of fifteen species of bats (Chiroptera) from Kenya and Rhodesia. Royal Ontario Museum
Pigozzi MI (2016) The chromosomes of birds during meiosis. Cytogenetic and Genome Research 150:128–138
Primus A, Harvey J, Guimondou S, Mboumba S, Ngangui R, Baker R, Porter C (2006) Karyology and chromosomal evolution of some small mammals inhabiting the rainforest of the Rabi oilfield, Gabon. In: Alonso A, Lee ME, Campbell P, Pauwels OSG, Dallmeier F (eds) Gamba Gabon: Biodiversity of an Equatorial African Rainforest. Bulletin of the Biological Society of Washington 371-382
Robinson TJ, Cernohorska H, Kubickova S, Vozdova M, Musilova P, Ruiz-Herrera A (2021) Chromosomal evolution in Raphicerus antelope suggests divergent X chromosomes may drive speciation through females, rather than males, contrary to Haldane's rule. Sci Rep 11:3152
Romanenko SA, Sitnikova NA, Serdukova NA, Perelman PL, Rubtsova NV, Bakloushinskaya IY, Lyapunova EA, Just W, Ferguson-Smith MA, Yang F, Graphodatsky AS (2007) Chromosomal evolution of Arvicolinae (Cricetidae, Rodentia). II. The genome homology of two mole voles (genus Ellobius), the field vole and golden hamster revealed by comparative chromosome painting. Chromosom Res 15:891–897
Romanenko SA, Fedorova YE, Serdyukova NA, Zaccaroni M, Stanyon R, Graphodatsky AS (2020) Evolutionary rearrangements of X chromosomes in voles (Arvicolinae, Rodentia). Sci Rep 10:13235
Royo H, Polikiewicz G, Mahadevaiah SK, Prosser H, Mitchell M, Bradley A, de Rooij DG, Burgoyne PS, Turner JM (2010) Evidence that meiotic sex chromosome inactivation is essential for male fertility. Curr Biol 20:2117–2123
Schimenti J (2005) Synapsis or silence. Nat Genet 37:11–13
Sharp P (1982) Sex chromosome pairing during male meiosis in marsupials. Chromosoma 86:27–47
Solari AJ (1970) The spatial relationship of the X and Y chromosomes during meiotic prophase in mouse spermatocytes. Chromosoma 29:217–236
Solari AJ (1974) The behavior of the XY pair in mammals. Int Rev Cytol 38:273–317
Solari AJ (1993) Sex chromosomes and sex determination in vertebrates. CRC Press, Boca Raton
Sung WK, Komatsu M, Jagiello G (1983) A method for obtaining synaptonemal complexes of human pachytene oocytes. Caryologia 36:315–324
Turner JM, Aprelikova O, Xu X, Wang R, Kim S, Chandramouli GV, Barrett JC, Burgoyne PS, Deng CX (2004) BRCA1, histone H2AX phosphorylation, and male meiotic sex chromosome inactivation. Curr Biol 14:2135–2142
Turner JM, Mahadevaiah SK, Fernandez-Capetillo O, Nussenzweig A, Xu X, Deng CX, Burgoyne PS (2005) Silencing of unsynapsed meiotic chromosomes in the mouse. Nat Genet 37:41–47
Turner JM, Mahadevaiah SK, Ellis PJ, Mitchell MJ, Burgoyne PS (2006) Pachytene asynapsis drives meiotic sex chromosome inactivation and leads to substantial postmeiotic repression in spermatids. Dev Cell 10:521–529
Viera A, Santos JL, Rufas JS (2009) Relationship between incomplete synapsis and chiasma localization. Chromosoma 118:377–389
Vorontsov NN, Lyapunova EA, Borissov YM, Dovgal VE (1980) Variability of sex chromosomes in mammals. Genetica 52:361–372
Waters PD, Ruiz-Herrera A (2020) Meiotic executioner genes protect the Y from extinction. Trends Genet 36:728–738
Zickler D, Kleckner N (1999) Meiotic chromosomes: integrating structure and function. Annu Rev Genet 33:603–754
Acknowledgements
This work was supported by grants CGL2014-53106-P from the Ministerio de Economía y Competitividad (Spain) and the Russian Foundation for Basic Research Nos. 20-34-70027 (SM), 20-04-00618 (IB), VIGG RAS State Assignment Contract No. 0112-2019-0002 and No. 0092-2019-0007 (SM, OK), and IDB RAS State Assignment for Basic Research No. 0088-2021-0019 (IB). AG-F was supported by a predoctoral fellowship from the Ministerio de Economía y Competitividad (Spain) and the European Social Fund (European Commission). We are grateful to GN Davidovich and AG Bogdanov of the Electron Microscopy Laboratory of the Biological Faculty of Moscow State University for technical assistance and Dr. C García de la Vega for insightful discussions.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Figure 1.
a. Immunolocalization of SYPC3 (green) and centromeres (red) in a E. tancrei spermatocyte at pachytene. The X chromosomes (XX) are indicated. b. Arranged meiotic karyotype from the same spermatocyte. The XX chromosomes from a female oocyte (XX♀) are included for comparison with the male ones (XX♂). Bar represents 10 μm. (PNG 623 kb)
Supplementary Figure 2.
Immunolocalization of SYPC3 (green) and RNA polymerase-II (red) in E. tancrei spermatocytes. a. Early pachytene. RNA pol-II appears evenly distributed over the nucleus, except in the region where the sex chromosomes (XX) are located. This localization pattern is maintained through mid pachytene (b), late pachytene (c), and early diplotene (d). However, at late diplotene (e), RNA pol-II labeling extends to the sex chromosomes. By prometaphase-I (f), no RNA pol-II signal is detected. Bar represents 10 μm. (PNG 2773 kb)
Supplementary Figure 3.
Immunolocalization of SYPC3 (red) and RPA (green) in E. tancrei spermatocytes. a. Leptotene. RPA appears as discrete foci distributed over the forming AEs. During early (b) and late zygotene (c), RPA is located mainly along the chromosomal regions that have completed synapsis. d. Early pachytene. RPA is still abundantly distributed along chromosomes. Sex chromosomes (XX) present RPA foci in both synapsed and unsynapsed regions. e. Mid pachytene. Only a few RPA foci are still present along the chromosomes, while a diffuse staining is covering the unsynapsed regions of sex chromosomes (XX). f. Late pachytene. RPA foci are not detectable. A faint diffuse staining covers sex chromosomes (XX). Bar represents 10 μm. (PNG 761 kb)
Supplementary Figure 4.
Immunolocalization of SYPC3 (red) and RAD51 (green) in E. tancrei spermatocytes. a. Leptotene. RPA appears as discrete foci distributed over the forming AEs. During early (b) and late zygotene (c), RAD51 is located along both synapsed and unsynapsed AEs. d. Early pachytene. Only a few RAD51 foci are present along the autosomes, but many foci are still visible over the sex chromosomes (XX). A diffuse staining covers the unsynapsed regions of these chromosomes. This diffuse staining and a few RAD51 foci are still detectable during mid (e) and late pachytene (f). Bar represents 10 μm. (PNG 725 kb)
Rights and permissions
About this article
Cite this article
Gil-Fernández, A., Matveevsky, S., Martín-Ruiz, M. et al. Sex differences in the meiotic behavior of an XX sex chromosome pair in males and females of the mole vole Ellobius tancrei: turning an X into a Y chromosome?. Chromosoma 130, 113–131 (2021). https://doi.org/10.1007/s00412-021-00755-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-021-00755-y