Abstract
Sex chromosome differentiation is subject to independent evolutionary processes among different lineages. The accumulation of repetitive DNAs and consequent crossing-over restriction guide the origin of the heteromorphic sex chromosome region. Several Neotropical fish species have emerged as interesting models for understanding evolution and genome diversity, although knowledge of their genomes is scarce. Here, we investigate the content of repetitive DNAs between males and females of Apareiodon sp. based on large-scale genomic data focusing on W sex chromosome differentiation. In Apareiodon, females are the heterogametic sex (ZW) and males are the homogametic sex (ZZ). The genome size estimate for Apareiodon was 1.2 Gb (with ~ 42× and ~ 47× coverage for males and females, respectively). In Apareiodon sp., approximately 36% of the genome was composed of repetitive DNAs and transposable elements (TEs) were the most abundant class. Read coverage analysis revealed different amounts of repetitive DNAs in males and females. The female-enriched clusters were located on the W sex chromosome and were mostly composed of microsatellite expansions and DNA transposons. Landscape analysis of TE contents demonstrated two major waves of invasions of TEs in the Apareiodon genome. Estimation of TE insertion times correlated with in situ locations permitted the inference that helitron, Tc1-mariner, and CMC EnSpm DNA transposons accumulated repeated copies during W chromosome differentiation between 20 and 12 million years ago. DNA transposons and microsatellite expansions appeared to be major players in W chromosome differentiation and to guide modifications in the genome content of the heteromorphic sex chromosomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sex can be determined by heteromorphic sex chromosomes that differ in their size, morphology, and gene content (Charlesworth 1991; Bachtrog 2013). Ancient sex-specific W or Y chromosomes (proto-sex chromosomes), such as those found in fish evolutionary lineages, have usually lost most functional genes owing to a loss of recombination with their former homologs, the Z or X chromosomes (Charlesworth et al. 2005; Steinemann and Steinemann 2005). The first step in chromosome differentiation is recombination suppression, which most likely evolved to prevent recombination between sex-determining genes, but the mechanisms of and reasons why this process occurs remain unclear (Charlesworth 1991; Bergero and Charlesworth 2009).
At the origin of heteromorphic sex chromosomes, mutations that are beneficial to one sex but detrimental to the other provide the selective force to suppress recombination between nascent sex chromosomes (Charlesworth 1996; Bachtrog 2013). The heterogametic sex tends to establish sexually antagonistic mutations in a population due to their beneficial effects, and the restriction of recombination between the proto-X (or Z) and proto-Y (or W) chromosomes may be a consequence of selection favoring genetic linkage between sexually antagonistic mutations and the sex-determining region (for a review, see Bachtrog 2013).
Recent “omics” studies have demonstrated that several genes distributed across the whole genome contribute to male and female differentiation (Teaniniuraitemoana et al. 2014; Chalopin et al. 2015; Chen et al. 2015). Moreover, the evolutionary forces that connect sexual dimorphism and the sex chromosomes are far more complicated than current theory allows. In the Mexican axolotl Ambystoma mexicanum, there is a miniscule difference between the sex chromosomes resulting from W-specific sequences (Keinath et al. 2018). The corresponding region contains a duplicated copy of the ATRX gene that appears to be a strong candidate for the primary sex determining locus (Keinath et al. 2018). The subsequent steps of chromosomal differentiation can involve mechanisms, such as chromosomal rearrangements and the accumulation of repetitive DNA sequences, that can trigger genetic degeneration (Charlesworth et al. 2005; Steinemann and Steinemann 2005; Bachtrog 2013). Y- or W-originated chromosomes present non-recombining portions corresponding to male- or female-specific regions, respectively, and the recombinant portion is referred to as the “pseudo-autosomal region” (PAR) (Bergero and Charlesworth 2009).
Repetitive DNAs are sequences that have been studied in more detail in recent decades (Marino-Ramirez et al. 2005; Feschotte 2008; Kapusta et al. 2013; Schemberger et al. 2016). They constitute the major fraction of genomes and can be classified as being dispersed (transposable elements—TEs—repetitions that occur at random positions in the genome) or in tandem (repetitions that are directly adjacent to each other) (Charlesworth et al. 1994). Several studies have described the role of repetitive DNAs in the structure, function, and evolution of genomes (Marino-Ramirez et al. 2005; Feschotte 2008; Kapusta et al. 2013) and in triggering the origination of a non-recombinant region between sex chromosomes (Steinemann and Steinemann 2005). In medaka fish, the insertion of a TE into the regulatory region of the dmrt1bY gene on the sex chromosome rewired the gene regulatory network cascades of sex determination (Herpin et al. 2010).
An interesting model for investigating sex chromosome origins and evolution is provided by the Parodontidae Neotropical fish group, composed of 32 species arranged into three genera, Parodon, Apareiodon, and Saccodon. Some of these species are cytogenetically characterized by heteromorphic W sex chromosomes in different stages of evolutionary differentiation (Bellafronte et al. 2011; Schemberger et al. 2011). There are species without evidences for sex chromosome occurrence, species presenting proto-sex chromosomes, species possessing a ZZ/ZW sex chromosome system ranging from small to large W chromosomes (e.g., Apareiodon sp., an undescribed species related to Apareiodon ibitiensis), and species presenting a multiple ZZ/ZW1W2 sex chromosome system (Vicari et al. 2006; Schemberger et al. 2011; do Nascimento et al. 2018).
Chromosomal studies in Paradontidae have focused on the mapping of repetitive DNAs in the karyotype and demonstrated enrichment of repetitive elements on the W chromosome (Schemberger et al. 2014, 2016; Ziemniczak et al. 2014). The recent advances in large-scale DNA sequencing have allowed comparative genomic analyses of repeated DNAs in ways that were never considered before (Treangen and Salzberg 2012). Therefore, we conducted whole-genome repetitive DNA analysis between males and females of Apareiodon sp. (ZW) to reveal the composition and evolutionary history of the W sex chromosome. Our study aimed to integrate genomic analyses and cytogenetic data to elucidate the role of repetitive DNAs in W sex chromosome differentiation.
Materials and methods
DNA extraction, next-generation sequencing, and genome assembly
DNA was isolated from the livers of two males (ZZ chromosomes) and two females (ZW chromosomes) of Apareiodon sp. (Verde River, Paraná State, Brazil; − 25°04′35″ and − 50°04′03″) using illustra tissue and a cell genomicPrep Mini Spin Kit (GE Healthcare, Chicago, Illinois, USA) following the manufacturer’s protocol. Two independent DNA libraries were sequenced on the Illumina HiSeq platform (100-base paired-end reads) for each sex (male and female) and filtered using FASTX-Toolkit software. Reads showing ≥ 90% of sequenced bases with Phred scores over 30 were selected. The genome was assembled using Velvet software (Zerbino and Birney 2008) due to its good performance for short-read datasets, and a 1.2 Gb total assembly length was generated (as expected for the Parodontidae genome). The statistics of genome assembly (scaffold and contig evaluations) were generated using the Perl script assemblathon_stats.pl (Bradnam et al. 2013). Additionally, genome data quality control was carried out with BUSCO v.3; the source code is available through the GitLab project, https://gitlab.com/ezlab/busco (Waterhouse et al. 2017).
Repeat identification, landscape pipelines, and TE insertion time estimation
The repetitive sequences of males and females were annotated using RepeatMasker (Smit et al. 2013–2015). For repetitive annotation, a custom repeat library was obtained with RepeatModeler (Smit and Hubley 2008–2015) using the default parameters. The custom library was merged with Repbase Update 20181026. RepeatMasker was run with the “slow” (-s), “library” (-lib), and “align” (-a) parameters. To summarize the RepeatMasker results, we used buildSummary.pl. output files from RepeatMasker as the input to generate calcdivergencefromalign.pl and repeatlandscape.pl files to calculate Kimura divergence values and plot the repeat landscape.
The nucleotide distances between all copies of each TE measured using the Kimura two-parameter method according to RepeatMasker (previously described) were compared to estimate insertion age. An average substitution rate (r) of 10−8 for substitutions per synonymous site per year and insertion time (T), as estimated for TEs (Arkhipova and Meselson 2005), was applied using the formula T = k/2r (Li et al. 2019).
Repetitive DNA analyses using RepeatExplorer
RepeatExplorer (Novak et al. 2010) was used to comparatively analyze repetitive DNAs between males and females. This software performs graph-based clustering of raw Illumina reads and uses a small sample of sequenced reads (0.1–0.5× coverage) (Novak et al. 2013) as the input, providing a fast and accurate way to compare two or more datasets. The reads were quality filtered based on the default parameters of paired_fastq_filtering. R, and a random sample of 5 million reads for each genome (~ 0.45× genome coverage) was selected. Finally, graph-based clustering was applied for de novo repeat identification and comparative analysis using the RepeatExplorer pipeline with the default parameters according to the developer’s recommendations (https://repeatexplorer-elixir.cerit-sc.cz/; Novak et al. 2010). The results from clustering were visually inspected with respect to their graph composition, which indicated the type of repeat and proportion of reads from each genome. The clusters showing differential contents in comparison of the female and male read datasets were selected for analysis. Blastn 2.2.31+ software was applied to search for the presence of Sat1WP satellite DNA (Schemberger et al. 2014) in the obtained clusters.
Chromosome preparation and fluorescence in situ hybridization (FISH)
Chromosome preparations were obtained from anterior kidney cells using an air-drying method (Bertollo et al. 1978), and C-banding was conducted according to Sumner (1972). The procedures employed in this study were in agreement with the Committee on the Ethics of the Use of Animals of the State University of Ponta Grossa (protocol 29/2016). Several DNA sequences were used as probes for FISH mapping, which are as follows: (a) the microdissected W chromosome; (b) the Cot-1 DNA fraction; (c) the satellite DNA Sat-1WP; (d) TEs; and (e) microsatellite sequences. The heterochromatic fraction (C-banding) of the Apareiodon sp. W chromosome was obtained by microdissection and DOP-PCR (referred to as Wm) and was subsequently synthetized as probe using digoxigenin 11-dUTP (Roche Applied Science, Mannheim, Germany) according to the method described by Vicari et al. (2010). The repetitive Cot-1 DNA fraction and the satellite DNA Sat-1WP were obtained from previous studies and labeled with digoxigenin 11-dUTP (Vicari et al. 2010; Schemberger et al. 2014).
Abundant TEs obtained from RepeatMasker annotated sequences and RepeatExplorer clusters were used for primer design and subjected to PCR amplification and probe labeling (Table S1—Electronic supplementary material). The PCR amplification of each TE was performed using 1× Taq Reaction buffer (200 mM Tris, pH 8.4, 500 mM KCl), 40 μM dNTPs, 2 mM MgCl2, 2 U of Taq DNA polymerase (Invitrogen, Carlsbad, California, USA), and each primer set at 0.2 μM. The amplicons produced by conventional PCR were labeled with digoxigenin 11-dUTP through nick translation (Dig Nick Translation Mix Roche Applied Science, Mannheim, Germany).
The abundant microsatellites resulting from the RepeatMasker and RepeatExplorer analysis of Apareiodon sp. genomic data—(GATA)n, (GA)n, (CA)n, (CAA)n, (CAT)n, (CG)n, (GT)n, (GAC)n, (CGG)n, (TAA)n, (GAA)n, (GAG)n, (GACA)n, (TA)n, (CAC)n, and (CAG)n—were labeled with tetramethylrhodamine or biotin at the 5′ end during synthesis (Sigma-Aldrich, St. Louis, Missouri, USA) and were used as probes. The general protocol for FISH followed the protocol described by Pinkel et al. (1986) under ~ 80% stringency conditions (2.5 ng/μl probe, 50% formamide, 2× SSC, 10% dextran sulfate, 42 °C for 16 h). Post-hybridization washes were performed at high stringency (50% formamide at 42 °C for 20 min, 0.1× SSC at 60 °C for 15 min, and 4× SSC 0.05% Tween at room temperature for 10 min). Streptavidin Alexa Fluor 488 (Molecular Probes, Eugene, Oregon, USA) and anti-digoxigenin rhodamine fab fragment (Roche Applied Science) antibodies were used for probe detection. The chromosomes were stained with DAPI (0.2 μg/ml) in Vectashield mounting medium (Vector, Burlingame, CA, USA) and analyzed by epifluorescence microscopy.
Results
Genome sequencing and assembly
A total of 508,168,176 male reads and 563,850,145 female reads (~ 100 nt) were obtained after filtering, generating ~ 42× and ~ 47× coverage, respectively. The genome assembly yielded 660,652 scaffolds for males and 501,976 scaffolds for females, for a total of 1.2 Gb (available at http://sacibase.ibb.unesp.br/jbrowse/JBrowse-1.12.1/index.html?data=data-apareiodon-sp). The N50 value was 14,634 for males and 22,307 bp for females (Table S2—Electronic supplementary material). Other quality metrics are shown in Table S2. Using BUSCO (searching 978 metazoan orthologous groups), 82.5 and 82.3% of the genes were found in the male and female genomes, respectively, most of which were intact in both genomes (Fig. S1—Electronic supplementary material). The same analysis using 4584 orthologous Actinopterygii groups identified 69.3 and 73.9% of the genes in the male and female genomes, respectively, most of which were intact in both genomes (Fig. S1—Electronic supplementary material).
Repetitive genomic composition
RepeatMasker analysis of the Apareiodon sp. assemblies showed that 36.2 and 35.3% of the genome was composed of repeat elements in males and females, respectively (Fig. 1a). A total of 56 superfamilies of TEs were identified, including 23 DNA transposon superfamilies and 33 retrotransposon superfamilies (Tables S3 and S4—Electronic supplementary material). DNA transposons presented a higher percentage (~ 38%) than other repetitive elements in the genome (Fig. 1a). Tc1/mariner and Hat transposons were most frequent within this category (Fig. 1b; Tables S3 and S4—Electronic supplementary material). The retrotransposons presented a 26.9% repetitive composition in males and females (Fig. 1b; Tables S3 and S4—Electronic supplementary material). The LTR retrotransposon Gypsy was predominant within the TE category (Fig. 1b; Tables S3 and S4—Electronic supplementary material). L2 and Rex-Babar were the most frequent among LINE retrotransposons (Fig. 1b; Tables S3 and S4—Electronic supplementary material). The SINE derived from transfer RNA (tRNA) was most abundant in this TE order (Fig. 1b; Tables S3 and S4—Electronic supplementary material). CMC/EnSpm, LTR/ERV, LTR/Gypsy, and LTR/Pao presented the largest fragments of ~ 5000 bp (Tables S3 and S4—Electronic supplementary material). The shortest fragment was 10 bp for several TEs (Tables S3 and S4—Electronic supplementary material). A total of 24.6% of the elements for males and 23.8% for females were unclassified elements in the genome (Fig. 1a).
Repetitive DNA composition and abundance in the Apareiodon sp. genome. a Percentage of repetitive classes based on whole-genome analysis of males and females conducted with RepeatMasker. DNA transposons and retrotransposon/LINEs were the most abundant. b Percentages of more frequent TEs in the genomes of males (dark color) and females (light color). On the y axis, 100% indicates the sum of the total length of the DNA transposon analyzed, and the x axis indicates the TE superfamily. c Percentages of more frequent tandem repeats in the genomes of males (dark color) and females (light color). On the y axis, 100% indicates the sum of the total length of tandem repeats analyzed, and the x axis indicates the repetition unit
A total of 20 abundant microsatellites were identified in the male and female genomes of Apareiodon sp. (Fig. 1c; Table S5—Electronic supplementary material). The microsatellites accounted for 8.9% of the male genome and 9.9% of the female genome (Fig. 1a; Table S5—Electronic supplementary material), and AC/GT, CA/GT, TC/GA, and CT/GT were the most abundant repeats in the genome (Fig. 1c; Table S5—Electronic supplementary material).
Repetitive divergence analyses and TE insertion ages
The relative ages of the different TE families were estimated by calculating Kimura-2 parameter value distances between individual copies and their consensus sequences (Smit and Hubley 2015). For each TE family/superfamily, the consensus sequence provides an approximation of the sequence of the ancestral TE. The landscape graphs were organized into two categories—DNA transposons and retrotransposons (Fig. 2 and interactive Electronic supplementary material Figs. S2, S3, S4, S5). Two evident DNA transposon invasion phases were detected in the DNA transposons of Apareiodon sp. males and females (Fig. 2a,b). The ancient peak of DNA transposon invasion showed higher copy numbers of DNA/helitron, DNA/Tc1-mariner, and DNA/CMC EnSpm (Fig. 2a,b and Electronic supplementary material Figs. S2, S3). The recent DNA transposon invasion phase mainly involved DNA/Hat, DNA/Harbinger, and DNA/Crypton and a second round of DNA/Tc1-mariner invasion (Fig. 2a,b and Electronic supplementary material Figs. S2, S3).
Repeat landscapes of male and female Apareiodon sp. genomes. The graphs show, for each element, the sequence divergence from their consensus sequence (x-axis) in relation to their number of copies in the genome (y-axis). Peaks represent waves of insertion (yellow arrows) of specific elements in the genome. Elements exhibiting older waves of insertion are present on the right side of the graph, while recent waves of insertions are depicted on the left side. Different colors indicate the distinct element types described on the right side. a Landscape of DNA transposons in the male genome (increases in Tc1-mariner, helitron, CMC EnSpm, and Hat occur in the ~ 40 to ~ 20 Kimura distance range in the ancient peak and at ~ 10 to 4 in the recent peak). b Landscape of DNA transposons in the female genome (increases in Tc1-mariner, helitron, CMC EnSpm, and Hat occur in the ~ 40 to ~ 20 Kimura distance range in the ancient peak and in the ~ 10 to 4 Kimura distance range in the recent peak). c Landscape of retrotransposons in the male genome (increases in SINE/tRNA and occur in the LINE/L2 ~ 29 to ~ 5 Kimura distance range in the recent peak; SINE/MIR and LINE/Rex-Babar increase in ~ 14 to ~ 4 Kimura distance range). d Landscape of retrotransposons in the female genome (increases in SINE/tRNA and occur in the LINE/L2 ~ 29 to ~ 5 Kimura distance range in the recent peak; SINE/MIR and LINE/Rex-Babar increase in ~ 14 to ~ 4 Kimura distance range). For a detailed interactive version of the graph, please refer to Figs. S1, S2, S3, and S4—Electronic supplementary material
Retrotransposons also presented two principal genomic invasion phases in Apareiodon sp. (yellow arrows in Fig. 2c,d). The ancient peak involved LTR/Pao, LINE/L1, and LTR/Gypsy as the most representative retrotransposons (Fig. 2c,d and Electronic supplementary material Figs. S4, S5). The recent peak was represented by SINE/MIR, SINE/tRNA, LINE/Rex-Babar, and LINE/L2 (Fig. 2c,d and Electronic supplementary material Figs. S4, S5).
According to neutral nucleotide substitution rates, we estimated the insertion time of the major TEs in the Apareiodon sp. genome. The putative mean age of the TE insertions in the genome ranged from ~ 25 million years ago (my) to ~ 1.5 my (Fig. 3). The ancient invasion phase demonstrated in landscape analysis involved LINE/L1, LTR/Gypsy, and LTR/Pao bursts in the genome between ~ 25 and 20 my, followed by DNA/helitron (~ 20–15 my) and Tc1-mariner and CMC EnSpm bursts between ~ 20 and 12 my (Fig. 3). The accumulation of helitron TEs was visibly higher in the female genome between this lapse time (Fig. 3). In the most recent TE invasion phase of the landscape analysis, SINE/MIR, LINE/Rex-Babar, and LINE/L2 element bursts were observed between ~ 7.5 and 2.5 my and DNA/Hat and a second Tc1-mariner invasion peak between ~ 3 and 1.5 my (Fig. 3).
Insertion time in millions of years (my) for the most enriched TEs in the Apareiodon sp. genome (a–c, g–j retrotransposons; d–f DNA transposons). The evolutionary age estimation indicates that the sex chromosome started to experience recombination suppression caused by the burst of TE insertion, as demonstrated in d–f
Comparative analysis of male and female repetitive sequences
To obtain the repetitive sequences accumulated on the W chromosome, comparative analysis between the sexes was performed using RepeatExplorer. The analyses resulted in 69 clusters (CL) with different graphing patterns (Fig. S6—Electronic supplementary material). A total of 67.9% of the reads were included in clusters, while 31.1% were singlets. CLs 6, 24, and 40 exhibited a higher proportion of female repeats and were selected for analyses (Fig. 4, Table 1). The cluster with the highest proportion of female reads was CL6. This cluster was enriched with different classes of microsatellites, and those presenting major differences between the sexes were selected for FISH mapping (Table 1). DNA/CMC EnSpm, DNA/Hat, DNA/helitron, DNA/Tc1-mariner, and LTR/Gypsy transposons were also enriched in CL6 of the female genomes and were subjected to FISH mapping. Blastn searches identified a fragment of Sat1WP (Schemberger et al. 2014) in CL6. CL24 exhibited enrichment of Maui/LINE-L2, and CL40 showed SSU-rRNA_Hsa enrichment in females. In parallel, CLs 4, 15, 48, and 49 presented higher proportions of male repeats (Fig. 4). These clusters exhibited a slightly higher percentage of penta- and hexanucleotide expansions of the DNA transposon Kolobok and LTR/Gypsy (Table S6—Electronic supplementary material).
Comparative repeat analysis of male and female genomes using RepeatExplorer software. Each column represents 100% of the reads of a cluster. The read proportions for the male and female genomes are indicated in blue and red, respectively. Clusters with a higher proportion of reads in females are expanded in the lower part of the graph
Karyotype organization and repetitive physical mapping validation on the sex chromosomes
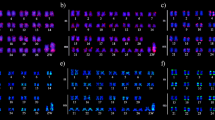
Apareiodon sp. possesses 2n = 54 chromosomes arranged in 48 meta/submetacentric and six subtelocentric chromosomes in males and in 47 meta/submetacentric and seven subtelocentric chromosomes in females (Fig. 5; Fig. S7—Electronic supplementary material). This difference is due to a ZZ/ZW heteromorphic sex chromosome system (Figs. 5, 6; Fig. S7—Electronic supplementary material). The W chromosome of Apareiodon sp. is almost totally heterochromatic and enriched with repetitive sequences, which was demonstrated by heterochromatin detection (C-banding) and FISH mapping of Wm heterochromatin and Cot-1 probes (Fig. 6; Fig. S7—Electronic supplementary material). The Sat1WP satellite is present in the interstitial region of the long arm on of the W chromosome (Fig. 6; Fig. S7—Electronic supplementary material).
The Z sex chromosome of Apareiodon sp. (metacentric chromosome) and W chromosome repetitive composition. a C-banding (CB) and FISH mapping with Wm, Cot-1, and Sat1WP probes. b FISH mapping of microsatellites selected via RepeatExplorer analyses ((GATA)n, (GA)n, (CA)n, (CAA)n, (CAT)n, and (GC)n). c FISH mapping of DNA transposons helitron, Tc1-mariner, and CMC EnSpm selected by RepeatExplorer. The scale bar corresponds to 5 μm
The sequences with the greatest difference scores related to females were used as probes to validate elements enriched on the W chromosome. FISH analysis revealed a hybridization signal mostly on the long arm of the W chromosome (Fig. S7—Electronic supplementary material). The helitron element possesses notable accumulation on the W long arm and a small site in the interstitial region of the Z short arm (Fig. 5a). The Tc1-mariner clusters were located in the interstitial region of the W long arm and the subterminal region of the Z short arm (Fig. 5b). The CMC EnSpm transposon accumulated in the proximal region of the W long arm and at a small site in the proximal region of the Z short arm (Fig. 5c). Gypsy and Hat were dispersed in the genome without W chromosome accumulation (data not shown).
The (GATA)n, (GA)n, (CA)n, (CAA)n, (CAT)n, and (CG)n clusters were located in the interstitial region of the W long arm (Fig. 6; Fig. S7—Electronic supplementary material). (GA)n, (CAT)n, and (CG)n accumulated in the terminal region of the Z short arm, while (GATA)n, (CA)n, and (CAA)n accumulated in the proximal region of the Z short arm (Fig. 6; Fig. S7—Electronic supplementary material).
Discussion
Repetitive DNA fraction in Apareiodon
The first step in understanding sex chromosome differentiation is to obtain knowledge of the repetitive genomic composition associated with physical mapping methodology (Mongue et al. 2017). Most of the sequenced fish genomes are from species with XX/XY chromosomal determination (e.g., Medaka) or without heteromorphic sex chromosomes (e.g., Danio rerio), and only one ZZ/ZW species genome is available, for tongue sole, Cynoglossus semilaevis (for a review, see Chalopin et al. 2015). This is the first genome assembly study in a Neotropical fish in which females are the heteromorphic sex. We set out to generate 1.2 Gb of genome data as expected for Apareiodon sp. This genome is almost 3× smaller than the human genome and shows a similar length to that of Danio rerio (1.4 Gb). On the basis of genome quality assessments comparing orthologous genes, it is possible to conclude that the Apareiodon sp. female genome is slightly better assembled than the male genome, as demonstrated by BUSCO analysis (Waterhouse et al. 2017).
The RepeatMasker analyses of the Apareiodon sp. assembly showed that ~ 36% of the genome matched diverse classes of repetitive DNAs, with a majority of DNA transposon classes. In teleosteans, the repetitive fraction ranges from 7% in Takifugu rubripes, with a majority of LINE retrotransposons, to 55% in zebrafish, with a majority of DNA transposons (Chalopin et al. 2015). Studies in mammals using the same approach have demonstrated a 31 to 49% repetitive element composition (Kirkness et al. 2003; de Koning et al. 2011). Mammals present less retrotransposon superfamily diversity than fish species (Volff et al. 2003). However, it is important to note that next-generation sequencing, even when producing high genome coverage, results in short reads that can lead to ambiguity in repetitive DNA assembly (Treangen and Salzberg 2012).
Microsatellite expansions were abundant in the Apareiodon sp. genome while classical satellites were underrepresented. Di-, tri-, and tetranucleotide expansions were more abundant. On the other hand, as demonstrated by FISH analysis (Schemberger et al. 2011), the satellite DNA pPh2004 described in Parodon hilarii (Vicente et al. 2003) was not detected (by Blastn) in the Apareiodon sp. genome. Our data show that Apareiodon sp. microsatellites can hitchhike within mobile elements, promoting chromosome site dispersion (Coates et al. 2010; Milani and Cabral-de-Mello 2014; Pucci et al. 2016) and increases in copy number via replication slippage (Kelkar et al. 2011). Furthermore, microsatellite genesis can also occur in degenerate TE segments (Wilder and Hollocher 2001).
Many families of satellite DNAs have been derived from transposable elements or include a major component that is related to part of a mobile element (Heikkinen et al. 1995; Kapitonov et al. 1998). We compared CL 48, which presented more TE LTR/Gypsy copies in male genome than in the female genome, contrary to the situation in CL 6. The graph structure of CL 48 is characterized by the presence of long multiple parallel lines that form a (more or less) much more linearly organized cluster, which is characteristic of dispersed and less degenerated sequences (Fig. S5) according to Novak et al. (2013). On the other hand, the graph structure of CL 6 formed a star circular-like structure that is characteristic of tandem repeats (Fig. S5) (see Novak et al. 2013). CL 6 included TEs (especially helitron, Tc1-mariner, and EnSpm TEs) that occur in the W sex chromosome-specific region. The helitron and EnSpm TEs exhibited in situ locations organizing extensive sites on the Apareiodon sp. chromosomes. The higher degeneracy level of these TEs and their in situ locations characteristic of tandem repeats indicate a probable origin from satellite DNAs during the evolutionary history of the origin of W chromosome differentiation.
Apareiodon sp. showed great diversity of TEs, and the DNA transposon class accounted for the majority of repetitive DNA present in the genome. Similarly, Schemberger et al. (2016) demonstrated accumulation of the DNA transposon Tc1-mariner in Parodontidae species. DNA transposons and retrotransposons presented different expansion histories as demonstrated by landscape analysis. TE activity invasion waves showed two evident time points (one ancient and another recent) with different levels of substitution rates demonstrated by Kimura analysis.
More recent activity of the TEs in the Apareiodon sp. genome can be observed between ~ 7.5 and 1.5 my with high copy numbers of Tc1-mariner, DNA/Harbinger, DNA/Hat, DNA/P, SINE/MIR, SINE/tRNA, LINE/Rex-Babar, and LINE/L2. The ancient invasion of the TEs was verified by a large increase in the copy number of LTR/Pao, LINE/L1, LTR/Gypsy, DNA/helitron, Tc1-mariner, and DNA/CMC EnSpm around ~ 25–12 my. Inactivated transposons can accumulate mutations at neutral rates until losing their molecular identity, as represented in this study by high Kimura distance values associated with copy numbers. In Apareiodon sp., the different levels of molecular deterioration suggested the occurrence of senescent TEs with deteriorated element identity, neutral TEs or TEs coopted by the genome, and a small copy number of autonomous TEs, as suggested for the “TE cycle life” (Kidwell and Lisch 2001; Fernández-Medina et al. 2012).
Sex chromosome origin and differentiation
The accumulation of repetitive DNAs is one of the key features of genomes and sex chromosome differentiation (Charlesworth et al. 1994) and can be cytogenetically characterized by chromosomal rearrangements and heterochromatinization (Gazoni et al. 2018; Komissarov et al. 2018; de Oliveira et al. 2018; Xin et al. 2018). The repetitive DNA data for Apareiodon sp. obtained via RepeatMasking and RepeatExplorer analyses were used in this study to screen the W chromosome repeat DNA composition. Interestingly, the different repetitive DNAs located on the W chromosome of Apareiodon were involved in TE invasions between ~ 20 and 12 my (helitron, Tc1-mariner, and EnSpm) detected by landscape analysis, and they exhibited major coverage in female genome clusters detected by RepeatExplorer. Other TEs, such as DNA/Hat and LTR/Gypsy, expanded in the genome before 20 my or after 12 my. These TEs showed no consistent signals on sex chromosomes and no participation in the origin of the W chromosome.
The repetitive sequences detected by RepeatExplorer presented more coverage in the W genome (68.8%) compared with the ZZ genome (67%), which was also demonstrated by the microdissected W and Cot-1 probes (Schemberger et al. 2014). In CL 6, (GACA)n, (GATA)n, (CAA)n, (CAT)n, (AC)n, (CAG)n, (CA)n, (GA)n, DNA/helitron, DNA/Tc1-mariner, and DNA/CMC EnSpm, DNA repeats showed a greater number of reads in females compared to the male genome, especially DNA/helitron repeats. This result was validated by FISH, which detected prominent interstitial signals of DNA/helitron, DNA/Tc1-mariner, DNA/CMC EnSpm, (GATA)n, (GA)n, (CAA)n, (CAT)n, (CA)n, and (CG)n on the W long arm.
Repetitive sequences are found at a high proportion on such heterochromatic sex chromosomes, and the evolution and emergence of sex chromosomes has been connected to the dynamics of repeats and transposable elements (Chalopin et al. 2015). Our TE insertion age estimates for Apareiodon sp. demonstrated that the three DNA transposons, especially the helitron transposon, with a high copy number insertion on the W chromosome, probably caused recombination suppression between the Z and W chromosomes 20 my. After helitron invasion, the W chromosome accumulated Tc1-mariner and EnSpm elements, which also exhibited a high copy number on the Z chromosome. TEs amplifications from not only the specific Y or W sex chromosome but also the corresponding regions of the X and Z chromosomes have also been observed in other fish species (Chalopin et al. 2015).
The genomic organization of the sex chromosomes has been well studied in other vertebrate model organisms with relatively old sex chromosomes, such as mammals (up to 166 my) and birds (approximately 200 my) (Bachtrog 2013). In fish species, Chalopin et al. (2015) proposed an early origin for sex chromosomes when compared to mammals and birds. The Y chromosome of a stickleback lineage originated 10 my (White et al. 2015), and the W chromosome of the tongue sole originated 30 my (Chen et al. 2014). However, the Japan sea lineage of the three-spine stickleback possesses a neo-Y chromosome dated to 1.5–2 my (Chalopin et al. 2015). Our data demonstrated that the Apareiodon sp. sex chromosome system shows a compatible emergence time to those of other teleosteans. These data corroborate the hypothesis that teleost fish exhibit multiple independent incidences of sex chromosome evolution, in a group that displays a wide variety of sex chromosome systems and diverse XY or ZW types with very young ages and various stages of differentiation (Chalopin et al. 2015).
Our study also identified and mapped several repetitive sequences constituting the Wm probe and further elucidated the theory of the evolution of sex chromosomes predicted by Schemberger et al. (2011). This study suggests that the ancestral proto-sex chromosome underwent an inversion event in its terminal region that rearranged the W chromosome repetitive DNAs to the proximal region. Here, we propose that the DNA helitron element (located in the proximal region of the Z short arm and in the W long arm) was involved in one important step of W differentiation, the crossing-over restriction that occurred during the W chromosome’s evolutionary history, according to the theory of sex chromosome gene erosion and the formation of the W chromosome specific region mediated by TE invasion (Charlesworth et al. 2005).
After ZW non-recombinant region origin, bursts of TEs and simple repeat accumulation occurred around young W sex-specific chromosome regions. We found that the interstitial W region was enriched with (GATA)n, (CA)n, (CAA)n, (CAT)n, and (CG)n microsatellites, indicating microsatellite sequences together with helitron, Tc1-mariner, and CMC EnSpm TEs participated in W chromosome rearrangement. The accumulation of microsatellites and TEs with different numbers of reads and signals in clusters in both sexes corroborates the hypothesis regarding the derivation of the W chromosome in Parodontidae. The W PAR was predicted by previous studies with the pPh2004 satellite in Parodon species, which is shared by the Z long arm and W short arm (Schemberger et al. 2011; Ziemniczak et al. 2014). The pPh2004 satellite is lacking in Apareiodon species with differentiated W chromosomes, making it difficult to clarify Z chromosome identity (Schemberger et al. 2011; Traldi et al. 2016). Based on previous data and the new data obtained here, it is possible to infer that the PAR of the ZW chromosomes in Apareiodon sp. is euchromatic and is organized by the long arm of the Z chromosome and the short arm of the W chromosome.
Usually, the heteromorphic sex chromosome accumulates TE DNA and exhibits a partial shift of euchromatic to heterochromatic regions (Charlesworth et al. 2005; Steinemann and Steinemann 2005). It has been proposed that gene silencing could potentially be achieved through the recruitment of TEs to the heteromorphic sex chromosome region, resulting in transcriptionally inactive heterochromatin, or through the accumulation of mutations in regulatory regions (Bachtrog 2013). The genomic and in situ location data showed that bursts of helitron, Tc1-mariner, and CMC EnSpm TEs and simple repeat accumulation around the Apareiodon sp. W chromosome-specific region could transform chromatin to a condensed region.
Regarding DNA structure, inverted polypurine/polypyrimidine DNA motifs tend to form triplex structures (triple helix or H-DNA, see Frank-Kamenetskii and Mirkin 1995). This type of repetitive DNA is an essential part of heterochromatic regions and is important for the maintenance of chromosome structure and the formation of heterochromatic W bodies in chickens (Saitoh et al. 1991; Komissarov et al. 2018). The W chromosome-specific region in Apareiodon sp. is heterochromatic and rich in microsatellite expansions, and the role of these expansions may be associated with chromatin nucleation and heterochromatic W body formation. In contrast, the major concentration of (GATA)n on the sex chromosomes was related to the activity of chromatin decondensation (Singh et al. 1994; Viger et al. 1998; Priyadarshini et al. 2003).
In previous experiments involving the in situ localization of the (GATA)n expansion, several accumulation sites were detected on the autosomes and W chromosome of Parodontidae species (Ziemniczak et al. 2014). Similarly, the W chromosome of chickens is rich in the (GGAAA)n expansion, and the involvement of this satellite in upstream regions of avian gonad-specific protein-coding sequences has been discussed (Komissarov et al. 2018). Interestingly, GATA-4 is a transcription factor that is specifically expressed in Sertoli cells in the testis and granulosa cells in the ovaries, where GATA sequences are used as transcription factor binding sites (Viger et al. 1998). In addition, GATA sequences are present in the promotor region of doublesex and mab-3-related transcription factor 1 (Dmrt-1), a gene involved in postnatal testis differentiation (Raymond et al. 2000).
According to Bachtrog (2013), the repetitive DNA structure of the sex chromosomes continues to cause challenges in deciphering their genomic sequences. Here, we have made progress in the elucidation of the repetitive DNA organization on the sex chromosomes of a Neotropical fish genome, and advances in the understanding of ZW sex determination mechanisms are expected in the future.
Conclusion
Our data provide a detailed view of the characterization and identification of repetitive elements in the Apareiodon sp. genome. Most TEs show high levels of nucleotide substitution, leading to defective, neutral, or coopted copies. Two different waves of TE invasion occurred in the evolutionary history of the Apareiodon sp. lineage, resulting in DNA transposon and retrotransposon accumulation. An ancient invasion of helitron DNA TE appeared during the Z/W recombination suppression event approximately 20 my. The W-specific genomic region expanded with accumulation of microsatellites and satellite DNA originating from TEs, increasing the diversity of repetitive DNAs in the sex chromosome composition and Z/W differentiation.
References
Arkhipova I, Meselson M (2005) Deleterious transposable elements and the extinction of asexuals. Bioessays 27:76–85. https://doi.org/10.1002/bies.20159
Bachtrog D (2013) Y-chromosome evolution: emerging insights into processes of Y-chromosome degeneration. Nat Rev Genet 14:113–124. https://doi.org/10.1038/nrg3366
Bellafronte E, Schemberger MO, Moreira-Filho O, Almeida MC, Artoni RF, Margarido VP, Vicari MR (2011) Chromosomal markers in Parodontidae: an analysis of new and reviewed data with phylogenetic inferences. Rev Fish Biol Fish 12:559–570. https://doi.org/10.1007/s11160-010-9177-3
Bergero R, Charlesworth D (2009) The evolution of restricted recombination in sex chromosomes. Trends Ecol Evol 24:94–102. https://doi.org/10.1016/j.tree.2008.09.010
Bertollo LAC, Takahashi CS, Moreira-Filho O (1978) Cytotaxonomic considerations on Hoplias lacerdae (Pisces, Erythrinidae). Braz J Genet 1:103–120
Bradnam KR, Fass JN, Alexandrov A, Baranay P, Bechner M, Birol I, Boisvert S, Chapman JA, Chapuis G, Chikhi R, Chitsaz H, Chou WC, Corbeil J, del Fabbro C, Docking TR, Durbin R, Earl D, Emrich S, Fedotov P, Fonseca NA, Ganapathy G, Gibbs RA, Gnerre S, Godzaridis É, Goldstein S, Haimel M, Hall G, Haussler D, Hiatt JB, Ho IY, Howard J, Hunt M, Jackman SD, Jaffe DB, Jarvis ED, Jiang H, Kazakov S, Kersey PJ, Kitzman JO, Knight JR, Koren S, Lam TW, Lavenier D, Laviolette F, Li Y, Li Z, Liu B, Liu Y, Luo R, MacCallum I, MacManes MD, Maillet N, Melnikov S, Naquin D, Ning Z, Otto TD, Paten B, Paulo OS, Phillippy AM, Pina-Martins F, Place M, Przybylski D, Qin X, Qu C, Ribeiro FJ, Richards S, Rokhsar DS, Ruby JG, Scalabrin S, Schatz MC, Schwartz DC, Sergushichev A, Sharpe T, Shaw TI, Shendure J, Shi Y, Simpson JT, Song H, Tsarev F, Vezzi F, Vicedomini R, Vieira BM, Wang J, Worley KC, Yin S, Yiu SM, Yuan J, Zhang G, Zhang H, Zhou S, Korf IF (2013) Assemblathon 2: evaluating de novo methods of genome assembly in three vertebrate species. GigaScience 2(10). https://doi.org/10.1186/2047-217X-2-10
Chalopin D, Volff JN, Galiana D, Anderson JL, Schartl M (2015) Transposable elements and early evolution of sex chromosomes in fish. Chromosom Res 23:545–560. https://doi.org/10.1007/s10577-015-9490-8
Charlesworth B (1991) The evolution of sex chromosomes. Science 251:1030–1033. https://doi.org/10.1126/science.1998119
Charlesworth B (1996) The evolution of chromosomal sex determination and dosage compensation. Curr Biol 6:149–162. https://doi.org/10.1016/S0960-9822(02)00448-7
Charlesworth B, Sniegowski P, Stephan W (1994) The evolutionary dynamics of repetitive DNA in eukaryotes. Nature 371:215–220. https://doi.org/10.1038/371215a0
Charlesworth D, Charlesworth B, Marais G (2005) Steps in the evolution of heteromorphic sex chromosomes. Heredity 95:118–128. https://doi.org/10.1038/sj.hdy.6800697
Chen S, Zhang G, Shao C, Huang Q, Liu G, Zhang P, Song W, An N, Chalopin D, Volff JN, Hong Y, Li Q, Sha Z, Zhou H, Xie M, Yu Q, Liu Y, Xiang H, Wang N, Wu K, Yang C, Zhou Q, Liao X, Yang L, Hu Q, Zhang J, Meng L, Jin L, Tian Y, Lian J, Yang J, Miao G, Liu S, Liang Z, Yan F, Li Y, Sun B, Zhang H, Zhang J, Zhu Y, du M, Zhao Y, Schartl M, Tang Q, Wang J (2014) Whole-genome sequence of a flatfish provides insights into ZW sex chromosome evolution and adaptation to a benthic lifestyle. Nat Genet 46:253–260. https://doi.org/10.1038/ng.2890
Chen X, Mei J, Wu J, Jing J, Ma W, Zhang J, Gui JF (2015) A comprehensive transcriptome provides candidate genes for sex determination/differentiation and SSR/SNP markers in yellow catfish. Mar Biotechnol 17:190–198. https://doi.org/10.1007/s10126-014-9607-7
Coates BS, Sumerford DV, Hellmich RL, Lewis LC (2010) Helitron-like transposon superfamily from Lepidoptera disrupts (GAAA)n microsatellites and is responsible for flanking sequence similarity within a microsatellite family. J Mol Evol 70:275–288. https://doi.org/10.1007/s00239-010-9330-6
de Koning AJ, Gu W, Castoe TA, Batzer MA, Pollock DD (2011) Repetitive elements may comprise over two-thirds of the human genome. PLoS Genet 71:e1002384. https://doi.org/10.1371/journal.pgen.1002384
de Oliveira EA, Sember A, Bertollo LAC, Yano CF, Ezaz T, Moreira-Filho O, Hatanaka T, Trifonov V, Liehr T, al-Rikabi ABH, Ráb P, Pains H, Cioffi MB (2018) Tracking the evolutionary pathway of sex chromosomes among fishes: characterizing the unique XX/XY1Y2 system in Hoplias malabaricus (Teleostei, Characiformes). Chromosoma 127:115–128. https://doi.org/10.1007/s00412-017-0648-3
do Nascimento VD, Coelho KA, Nogaroto V, Almeida RB, Ziemniczak K, Centofante L, Pavanelli CS, Torres RA, Moreira-Filho O, Vicari MR (2018) Do multiple karyomorphs and population genetics of freshwater darter characines (Apareiodon affinis) indicate chromosomal speciation? Zool Anz 272:93–103. https://doi.org/10.1016/j.jcz.2017.12.006
Fernández-Medina RD, Ribeiro JMC, Carareto CMA, Velasque L, Struchiner CJ (2012) Losing identity: structural diversity of transposable elements belonging to different classes in the genome of Anopheles gambiae. BMC Genomics 13:272. https://doi.org/10.1186/1471-2164-13-272
Feschotte C (2008) Transposable elements and the evolution of regulatory networks. Nat Rev Genet 9:397–405. https://doi.org/10.1038/nrg2337
Frank-Kamenetskii MD, Mirkin SM (1995) Triplex DNA structures. Annu Rev Biochem 64:65–95. https://doi.org/10.1146/annurev.bi.64.070195.000433
Gazoni T, Haddad CFB, Narimatsu H, Cabral-de-Mello DC, Lyra ML, Parise-Maltempi PP (2018) More sex chromosomes than autosomes in the Amazonian frog Leptodactylus pentadactylus. Chromosoma 127:269–278. https://doi.org/10.1007/s00412-018-0663-z
Heikkinen E, Launonen V, Muller E, Bachmann L (1995) The pvB370 BamIII satellite DNA family of the Drosophila virilis group and its evolutionary relation to mobile dispersed genetic pDv elements. J Mol Evol 41:604–614. https://doi.org/10.1007/BF00175819
Herpin A, Braasch I, Kraeussling M, Schmidt C, Thoma EC, Nakamura S, Tanaka M, Schartl M (2010) Transcriptional rewiring of the sex determining Dmrt1 gene duplicate by transposable elements. PLoS Genet 6:e1000844. https://doi.org/10.1371/journal.pgen.1000844
Kapitonov VV, Holmquist GP, Jurka J (1998) L1 repeat is a basic unit of heterochromatin satellites in Cetaceans. Mol Biol Evol 15:611–612
Kapusta A, Kronenberg Z, Lynch VJ, Zhuo X, Ramsay L, Bourque G, Yandell M, Feschotte C (2013) Transposable elements are major contributors to the origin, diversification, and regulation of vertebrate long noncoding RNAs. PLoS Genet 9:e1003470. https://doi.org/10.1371/journal.pgen.1003470
Keinath MC, Timoshevskaya N, Timoshevskiy VA, Voss SR, Smith JJ (2018) Miniscule differences between the sex chromosomes in the giant genome of a salamander, Ambystoma mexicanum. Sci Rep 8:17882. https://doi.org/10.1038/s41598-018-36209-2
Kelkar YD, Eckert KA, Chiaromonte F, Makova KD (2011) A matter of life or death: how microsatellites emerge in and vanish from the human genome. Genome Res 21:2038–2048 http://www.genome.org/cgi/doi/10.1101/gr.122937.111
Kidwell MG, Lisch DR (2001) Perspective: transposable elements, parasitic DNA, and genome evolution. Evolution 55:1–24. https://doi.org/10.1111/j.0014-3820.2001.tb01268.x
Kirkness EF, Bafna V, Halpern AL, Levy S, Remington K, Rusch DB, Delcher AL, Pop M, Wang W, Fraser CM, Venter JC (2003) The dog genome: survey sequencing and comparative analysis. Science 301:1898–1903. https://doi.org/10.1126/science.1086432
Komissarov AS, Galkina SA, Koshel EI, Kulak MM, Dyomin AG, O’Brien SJ, Gaginskaya ER, Saifitdinova AF (2018) New high copy tandem repeat in the content of the chicken W chromosome. Chromosoma 127:73–83. https://doi.org/10.1007/s00412-017-0646-5
Li SF, Guo YJ, Li JR, Zhang DX, Wang BX, Li N, Deng CL, Gao WJ (2019) The landscape of transposable elements and satellite DNAs in the genome of a dioecious plant spinach (Spinacia oleracea L.). Mob DNA 10(3). https://doi.org/10.1186/s13100-019-0147-6
Marino-Ramirez L, Lewis KC, Landsman D, Jordan IK (2005) Transposable elements donate lineage-specific regulatory sequences to host genomes. Cytogenet Genome Res 110:333–341. https://doi.org/10.1159/000084965
Milani D, Cabral-de-Mello DC (2014) Microsatellite organization in the grasshopper Abracris flavolineata (Orthoptera: Acrididae) revealed by FISH mapping: remarkable spreading in the A and B chromosomes. PLoS One 9:e97956. https://doi.org/10.1371/journal.pone.0097956
Mongue AJ, Nguyen P, Volenikova A, Walters JR (2017) Neo-sex chromosomes in the Monarch butterfly, Danaus plexippus. G3 (Bethesda) 7:3281–3294. https://doi.org/10.1534/g3.117.300187
Novak P, Neumann P, Macas J (2010) Graph-based clustering and characterization of repetitive sequences in 592 next-generation sequencing data. BMC Bioinformatics 11:378. https://doi.org/10.1186/1471-2105-11-378
Novak P, Neumann P, Pech J, Steinhaisl J, Macas J (2013) RepeatExplorer: a galaxy-based web server for genome-wide characterization of eukaryotic repetitive elements from next-generation sequence reads. Bioinformatics 29:792–793. https://doi.org/10.1093/bioinformatics/btt054
Pinkel D, Straume T, Gray JW (1986) Cytogenetic analysis using quantitative, high-sensitivity, fluorescence hybridization. Proc Natl Acad Sci U S A 83:2934–2938. https://doi.org/10.1073/pnas.83.9.2934
Priyadarshini P, Murthy BS, Nagaraju J, Singh L (2003) GATA-binding protein expressed predominantly in the pupal ovary of the silkworm, Bombyx mori. Insect Biochem Mol Biol 33:185–195. https://doi.org/10.1016/S0965-1748(02)00190-X
Pucci MB, Barbosa P, Nogaroto V, Almeida MC, Artoni RF, Pansonato-Alves JC, Scacchetti PC, Foresti F, Moreira-Filho O, Vicari MR (2016) Chromosomal spreading of microsatellites and (TTAGGG)n sequences in the Characidium zebra and C. gomesi genomes (Characiformes: Crenuchidae). Cytogenet Genome Res 149:182–190. https://doi.org/10.1159/000447959
Raymond CS, Murphy MW, O'Sullivan MG, Bardwell VJ, Zarkower D (2000) Dmrt1, a gene related to worm and fly sexual regulators, is required for mammalian testis differentiation. Genes Dev 14:2587–2595. https://doi.org/10.1101/gad.834100
Saitoh Y, Saitoh H, Ohtomo K, Mizuno S (1991) Occupancy of the majority of DNA in the chicken W chromosome by bent-repetitive sequences. Chromosoma 101:32–40. https://doi.org/10.1007/BF00360684
Schemberger MO, Bellafronte E, Nogaroto V, Almeida MC, Schühli GS, Artoni RF, Moreira-Filho O, Vicari MR (2011) Differentiation of repetitive DNA sites and sex chromosome systems reveal closely related group in Parodontidae (Actinopterygii: Characiformes). Genetica 139:1499–1508. https://doi.org/10.1007/s10709-012-9649-6
Schemberger MO, Oliveira JIN, Nogaroto V, Almeida MC, Artoni RF, Cestari MM, Moreira-Filho O, Vicari MR (2014) Construction and characterization of a repetitive DNA library in Parodontidae (Actinopterygii: Characiformes): a genomic and evolutionary approach to the degeneration of the W sex chromosome. Zebrafish 11:518–527. https://doi.org/10.1089/zeb.2014.1013
Schemberger MO, Nogaroto V, Almeida MC, Artoni RF, Valente GT, Martins C, Moreira-Filho O, Cestari MM, Vicari MR (2016) Sequence analyses and chromosomal distribution of the Tc1/Mariner element in Parodontidae fish (Teleostei: Characiformes). Gene 593:308–314. https://doi.org/10.1016/j.gene.2016.08.034
Singh L, Wadhwan R, Naidu S, Nagaraj R, Ganesan M (1994) Sex- and tissue-specific Bkm (GATA)-binding protein in the germ cells of heterogametic sex. J Biol Chem 269:25321–25327
Smit AFA, Hubley R (2008–2015) RepeatModeler Open—1.0 Available from http://www.repeatmasker.org. Accessed 26 Nov 2018
Smit AFA, Hubley R, Green P (2013–2015) RepeatMasker Open—4.0. Available at: http://www.repeatmasker.org. Accessed 26 Nov 2018
Steinemann S, Steinemann M (2005) Retroelements: tools for sex chromosome evolution. Cytogenet Genome Res 110:134–143. https://doi.org/10.1159/000084945
Sumner AT (1972) A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res 75:304–306. https://doi.org/10.1016/0014-4827(72)90558-7
Teaniniuraitemoana V, Huvet A, Levy P, Klopp C, Lhuillier E, Gaertner-Mazouni N, Le Moullac G (2014) Gonad transcriptome analysis of pearl oyster Pinctada margaritifera: identification of potential sex differentiation and sex determining genes. BMC Genomics 15:491. https://doi.org/10.1186/1471-2164-15-491
Traldi JB, Vicari MR, Martinez JDF, Blanco DR, Lui RL, Moreira-Filho O (2016) Chromosome analyses of Apareiodon argenteus and Apareiodon davisi (Characiformes, Parodontidae): an extensive chromosomal polymorphism of 45S and 5S ribosomal DNAs. Zebrafish 13:19–25. https://doi.org/10.1089/zeb.2015.1124
Treangen TJ, Salzberg SL (2012) Repetitive DNA and next-generation sequencing: computational challenges and solutions. Nat Rev Genet 13:36–46. https://doi.org/10.1038/nrg3117
Vicari MR, Moreira-Filho O, Artoni RF, Bertollo LAC (2006) ZZ/ZW sex chromosome system in an undescribed species of the genus Apareiodon Characiformes, Parodontidae. Cytogenet Genome Res 114:163–168. https://doi.org/10.1159/000093333
Vicari MR, Nogaroto V, Noleto RB, Cestari MM, Cioffi MB, Almeida MC, Moreira-Filho O, Bertollo LAC, Artoni RF (2010) Satellite DNA and chromosomes in Neotropical fishes: methods, applications and perspectives. J Fish Biol 76:1094–1116. https://doi.org/10.1111/j.1095-8649.2010.02564.x
Vicente VE, Bertollo LA, Valentini SR, Moreira-Filho O (2003) Origin and differentiation of sex chromosome system in Parodon hilarii (Pisces, Parodontidae). Satellite DNA, G and C banding. Genetica 119:115–120. https://doi.org/10.1023/A:1026082904672
Viger RS, Mertineit C, Trasler JM, Nemer M (1998) Transcription factor GATA 4 is expressed in a sexually dimorphic pattern during mouse gonadal development and is a potent activator of the MIS promoter. Development 125:2665–2675
Volff JN, Bouneau L, Ozouf-Costaz C, Fischer C (2003) Diversity of retrotransposable elements in compact pufferfish genomes. Trends Genet 19:674–678. https://doi.org/10.1016/j.tig.2003.10.006
Waterhouse RM, Seppey M, Simão FA, Manni M, Ioannidis P, Klioutchnikov G, Kriventseva EV, Zdobnov EM (2017) BUSCO applications from quality assessments to gene prediction and phylogenomics. Mol Biol Evol 35:543–548. https://doi.org/10.1093/molbev/msx319
White MA, Kitano J, Peichel CL (2015) Purifying selection maintains dosage-sensitive genes during degeneration of the threespine stickleback Y chromosome. Mol Biol Evol 32:1981–1995. https://doi.org/10.1093/molbev/msv078
Wilder J, Hollocher H (2001) Mobile elements and the genesis of microsatellites in dipterans. Mol Biol Evol 18:384–392. https://doi.org/10.1093/oxfordjournals.molbev.a003814
Xin H, Zhang T, Han Y, Wu Y, Shi J, Xi M, Jiang J (2018) Chromosome painting and comparative physical mapping of the sex chromosomes in Populus tomentosa and Populus deltoides. Chromosoma 127:313–321. https://doi.org/10.1007/s00412-018-0664-y
Zerbino D, Birney E (2008) Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res gr-074492. https://doi.org/10.1101/gr.074492.107
Ziemniczak K, Traldi JB, Nogaroto V, Almeida MC, Artoni RF, Moreira-Filho O, Vicari MR (2014) In situ localization of (GATA)n and (TTAGGG)n repeated DNAs and W sex chromosome differentiation in Parodontidae (Actinopterygii: Characiformes). Cytogenet Genome Res 144:325–332. https://doi.org/10.1159/000370297
Acknowledgments
The authors are grateful to ICMBio (Instituto Chico Mendes de Conservação da Biodiversidade) (license number 10538-1 to collect specimens). This study was supported by the Fundação Araucária (Fundação Araucária de Apoio ao Desenvolvimento Científico e Tecnológico do Estado do Paraná), FAPESP (Fundação de Amparo à Pesquisa do Estado de São Paulo, grant number: 2015/16661-1), CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior, Finance Code 001), and CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico, grant number: 304166/2016-2).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 448 kb)
ESM 2
(PDF 2809 kb)
Supplementary figure 2
Repeat landscape of DNA transposons in the male Apareiodon sp. genome. The graphs show, for each element, the sequence divergence from their consensus sequence (x-axis) in relation to their number of copies in the genome (y-axis). Peaks represent waves of insertion (yellow arrows) of specific elements in the genome. Elements with older waves of insertion are presented on the right side of the graph, while recent waves of insertions are depicted on the left side. Different colors indicate the distinct element types described on the right side. (HTML 26 kb)
Supplementary figure 3
Repeat landscape of DNA transposons in the female Apareiodon sp. genome. The graphs show, for each element, the sequence divergence from their consensus sequence (x-axis) in relation to their number of copies in the genome (y-axis). Peaks represent waves of insertion (yellow arrows) of specific elements in the genome. Elements with older waves of insertion are presented on the right side of the graph, while recent waves of insertions are depicted on the left side. Different colors indicate distinct element types described on the right side. (HTML 25 kb)
Supplementary figure 4
Repeat landscape of retrotransposons in the male Apareiodon sp. genome. The graphs show, for each element, the sequence divergence from their consensus sequence (x-axis) in relation to their number of copies in the genome (y-axis). Peaks represent waves of insertion (yellow arrows) of specific elements in the genome. Elements with older waves of insertion are presented on the right side of the graph, while recent waves of insertions are depicted on the left side. Different colors indicate distinct element types described on the right. (HTML 40 kb)
Supplementary figure 5
Repeat landscape of retrotransposons in the male Apareiodon sp. genome. The graphs show, for each element, the sequence divergence from their consensus sequence (x-axis) in relation to their number of copies in the genome (y-axis). Peaks represent waves of insertion (yellow arrows) of specific elements in the genome. Elements with older waves of insertion are presented on the right side of the graph, while recent waves of insertions are depicted on the left side. Different colors indicate distinct element types described on the right side. (HTML 40 kb)
Rights and permissions
About this article
Cite this article
Schemberger, M.O., Nascimento, V.D., Coan, R. et al. DNA transposon invasion and microsatellite accumulation guide W chromosome differentiation in a Neotropical fish genome. Chromosoma 128, 547–560 (2019). https://doi.org/10.1007/s00412-019-00721-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-019-00721-9