Abstract
Nematodes of the genus Strongyloides are intestinal parasites of vertebrates including man. Currently, Strongyloides and its sister genus Parastrongyloides are being developed as models for translational and basic biological research. Strongyloides spp. alternate between parthenogenetic parasitic and single free-living sexual generations, with the latter giving rise to all female parasitic progeny. Parastrongyloides trichosuri always reproduces sexually and may form many consecutive free-living generations. Although the free-living adults of both these species share a superficial similarity in overall appearance when compared to Caenorhabditis elegans, there are dramatic differences between them, in particular with respect to the organization of the germline. Here we address two such differences, which have puzzled investigators for several generations. First, we characterize a population of non-dividing giant nuclei in the distal gonad, the region that in C. elegans is populated by mitotically dividing germline stem cells and early meiotic cells. We show that in these nuclei, autosomes are present in higher copy numbers than X chromosomes. Consistently, autosomal genes are expressed at higher levels than X chromosomal ones, suggesting that these worms use differential chromatin amplification for controlling gene expression. Second, we address the lack of males in the progeny of free-living Strongyloides spp. We find that male-determining (nullo-X) sperm are present in P. trichosuri, a species known to produce male progeny, and absent in Strongyloides papillosus, which is consistent for a species that does not. Surprisingly, nullo-X sperm appears to be present in Strongyloides ratti, even though this species does not produce male progeny. This suggests that different species of Strongyloides employ various strategies to prevent the formation of males in the all-parasitic progeny of the free-living generation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The nematode Strongyloides stercoralis is one of the most prevalent parasitic round worms in humans. Strongyloidiasis is considered a neglected tropical disease (Olsen et al. 2009). The rat parasite Strongyloides ratti and the sheep parasite Strongyloides papillosus are two other, more experimentally accessible members of the nematode genus Strongyloides, which consists of small-intestinal parasites of numerous vertebrates (Viney and Lok 2007; Dorris et al. 2002; Speare 1989). S. ratti can be maintained in the laboratory in their natural host while S. papillosus can be reared in rabbits. The easy access to the free-living adults (see below) and a number of recently developed resources for working with Strongyloides spp. and their relative Parastrongyloides trichosuri, a facultative parasite of Australian possums (Grant et al. 2006a; 2006b; Shao et al. 2012; Eberhardt et al. 2007; Nemetschke et al. 2010b; Viney et al. 2002), render this group of parasites highly attractive not only for parasitological research of medical and veterinary interest but also for the study of basic biological questions like host parasite interactions (Bleay et al. 2007; Crook and Viney 2005; Viney et al. 2006) and evolution (Fenton et al. 2004; Gemmill et al. 2000; Streit 2014).
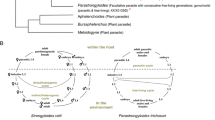
The life cycles of S. ratti and S. papillosus (Fig. 1a) have been reviewed in several places (Streit 2008; Viney and Lok 2007). In brief, the parasitic worms are all female and live in the small intestines of their respective hosts. They give rise to both female and male offspring by mitotic parthenogenesis. The young females either develop directly into infective third-stage larvae (L3i) and search for a new host (termed homogonic or direct development) or, together with all the males, give rise to a facultative free-living generation that reproduces sexually (termed heterogonic or indirect development). In the genus Strongyloides, offspring of free-living adults are all female and bound to develop into L3i, with very few known exceptions (Streit 2008; Yamada et al. 1991). Parastrongyloides with its best-studied representative P. trichosuri is a genus closely related to Strongyloides (Dorris et al. 2002). Parastrongyloides spp., like Strongyloides spp., also form parasitic and free-living generations of reproducing adults. However, the life history (Fig. 1b) and reproductive modes of this genus differ in interesting ways from those of Strongyloides spp. (Grant et al. 2006b). The distinguishing feature that led to the installation of the new genus Parastrongyloides was the presence of males in the parasitic generation (Mackerras 1959). Linked to this, a major difference from Strongyloides spp. is that free-living Parastrongyloides spp. produce progeny of both sexes. For P. trichosuri, it was confirmed genetically that reproduction in both generations is indeed sexual (Grant et al. 2006b; Kulkarni et al. 2013). In addition, P. trichosuri can undergo an apparently unlimited number of consecutive free-living generations (Grant et al. 2006b). It is therefore a facultative parasite.
Life cycle and introduction to the Strongyloides gonad a The generalized life cycle of Strongyloides species. b The life cycle of Parastrongyloides trichosuri. c Representative examples of dissected DAPI-stained gonads from Strongyloides spp. females (top) and males (bottom) showing the gonadal organization with giant nuclei occupying the entire distal arm, followed by a region of small compact nuclei. Scale bar 20 μm
P. trichosuri employs XX/XO chromosomal sex determination with 2n = 6 in females and 2n = 5 in males. There is no indication of an environmental influence on sex determination in this species (Grant et al. 2006b; Kulkarni et al. 2013). However, in S. ratti and in S. papillosus, as in all species of Strongyloides investigated thus far, the sex ratio in the progeny of parthenogenetic parasitic females is influenced by the immune status of the host (reviewed in Streit 2008) such that an increasing immune response of the host against the worms leads to a higher proportion of males. Nevertheless, in S. ratti and in S. papillosus, males and females differ in their chromosomal complement. In S. ratti, females have a pair of X chromosomes along with two pairs of autosomes while males have only one X chromosome. Hence, they employ an environmentally controlled XX/XO sex determination with 2n = 6 in females and 2n = 5 in males (Harvey and Viney 2001; Nigon and Roman 1952). In S. papillosus, the genetic material homologous to autosome I and to the X chromosome of S. ratti is combined into one chromosome (Nemetschke et al. 2010a). Additionally for this species, oocytes that give rise to males undergo a sex-specific chromatin diminution event that creates a hemizygous region corresponding in sequence to the X chromosome in S. ratti, presumably functionally restoring the ancestral XX/XO sex-determining system (Albertson et al. 1979; Nemetschke et al. 2010a; Kulkarni et al. 2013). In this process, an internal portion of one chromosome is eliminated and both ends are retained as separate chromosomes. This leads to karyotypes of 2n = 5 in males and 2n = 4 in females. Given that Strongyloides males are heterogametic, it is puzzling that they sire only female (homogametic) progeny. There are multiple, not mutually exclusive, hypothetical explanations for this: (i) male-determining mature sperm (nullo-X sperm in the case of S. ratti and sperm lacking the region undergoing chromatin diminution in S. papillosus, for simplicity referred to from here on as nullo-X sperm) may be rare or never formed at all; (ii) nullo-X sperm may be incapable or highly inefficient at fertilizing oocytes; or (iii) genetically male zygotes may be unviable. It is currently unknown as to which of these explanations hold true in Strongyloides spp. For S. papillosus, absence of nullo-X sperm was postulated based on genetic experiments (Nemetschke et al. 2010a), while for S. stercoralis, measurements of DNA content of sperm based on DNA binding dyes indicated that nullo-X sperm may be present (Hammond and Robinson 1994).
In the model nematode Caenorhabditis elegans, as in many other nematodes (Rudel et al. 2005), the gonads in both sexes are essentially tubes. In the hermaphrodites, the gonad has two arms, one extending anteriorly and one posteriorly but both terminating in a central vulva. The male gonad has just one arm with a posterior opening. These arms contain a germ cell production and differentiation line (Hubbard and Greenstein 2005). Although the overall morphology of the gonad arms is very similar to that of C. elegans, the organization and appearance of the germ cells is very different in Strongyloides and Parastrongyloides spp. (Hammond and Robinson 1994; Triantaphyllou and Moncol 1977). In these genera, the distal region contains giant nuclei that take on various shapes (Fig. 1c). No nuclear divisions have ever been reported in this region of the gonad in adult free-living stages. These nuclei have a DNA content of up to several hundred C, where one C is the DNA content of a haploid set of chromosomes (Hammond and Robinson 1994). Interestingly, these authors noted that the DNA contents they observed in different nuclei and among individuals were not full multiples of the entire genome, suggesting that different portions of the genome are amplified to various extents. The region with these giant nuclei is followed proximally by a band of very small, compact nuclei. At the end of this band, nuclei with condensed and presumably meiotic chromosomes can be observed. Further down in the gonad, depending on sex, differentiated oocytes or sperm are present, very similar to C. elegans.
Here, we elaborate on the chromatin and chromosome complements in germ cells of Strongyloididae. Based on quantitative sequencing approaches, we show that the X chromosome (or the X-derived region in the case of S. papillosus) is underrepresented in comparison to the autosomes in the giant germline nuclei of both sexes in two species of Strongyloides and in P. trichosuri. Differential chromatin amplification likely serves as a way of controlling gene expression since X-encoded genes are, on average, expressed at a much lower level than autosomal genes in the distal gonad of S. ratti females. Additionally, based on quantitative sequencing of isolated mature sperm, we confirm the absence of nullo-X sperm in S. papillosus but, surprisingly, its presence in S. ratti. For this species, we found evidence for the presence of nullo-X sperm and unviable early embryos, suggesting that the two species of Strongyloides employ different strategies to avoid the formation of males in the progeny of the free-living generation.
Materials and methods
Culturing and manipulating nematodes
S. ratti ED321 and S. papillosus isolate LIN were maintained as described (Eberhardt et al. 2007; Nemetschke et al. 2010b; Viney et al. 1992). All animal experimentation was done according to national and international guidelines. The required permits were granted by the local authorities. P. trichosuri was cultured in continuous free-living cycles (Grant et al. 2006b) at 20 °C on NGM plates seeded with E. coli OP 50 bacteria (Stiernagle 1999) supplemented with a piece of autoclaved rabbit feces.
DAPI staining
Adult worms (of the desired age) were fixed with ice cold 100 % methanol and directly mounted (without a rehydration series) on polylysine-coated glass slides in 10 μL of Vectashield containing 1 μg mL−1 4′,6-diamidino-2-phenylindole (DAPI).
DNA extractions from dissected gonads for Illumina sequencing
About 500 distal arms of the gonads per biological replicate were manually dissected from adult worms (both sexes) of all three species. Samples were frozen in liquid nitrogen and stored at −20 °C for DNA extraction using the Illumina Epicenter Masterpure™ DNA purification kit. The samples were measured for their DNA content using the Qubit High Sensitivity DNA measuring kit and then used for making libraries according to the Illumina platform. Libraries were made using the NEXTflix™ ChIP-Seq10ng kit (Bioo Scientific) and then sequenced.
RNA extraction from dissected gonads
For each of the four independent replicates, approximately 100 distal gonad arms were dissected from adult free-living female worms of S. ratti and samples were immediately put on dry ice. Five hundred microliters of TRIzol was added to the samples, and they were then frozen in liquid nitrogen and stored at −80 °C until RNA extraction. RNA was extracted using the RNA micro kit (Invitrogen) and quantified using the Qubit High Sensitivity RNA measuring kit. cDNA was prepared using SMARTer Ultra Low Input RNA for Illumina Sequencing (Clontech Laboratories). Libraries were made using the Low Input Library Prep Kit (Clontech Laboratories) and then Illumina sequenced.
DNA extraction from sperm
Adult males of all three species were incubated in a solution of 0.5 μg/L (0.2 M) levamisole at room temperature for 45 min as described for S. papillosus (Nemetschke et al. 2010a). This causes muscle contraction and as a consequence the release of sperm. Then the ejaculated mature sperm were collected using a mouth pipette and transferred into sterile Eppendorf tubes. DNA extraction was done using the Illumina Epicenter Masterpure™ DNA purification kit. Libraries were made according to the Low Input Library Prep Kit (Clontech) and Illumina sequenced.
Analysis of sequencing data
Draft genome assemblies including chromosome information and gene annotations for S. ratti, S. papillosus, and P. trichosuri were provided by the Sanger Institute. We used version 0.5.9-r16 of BWA software (Li and Durbin 2009) to align raw reads of all three species to their respective genome assemblies. Read counts per 2-kb window were calculated from the resulting alignment files using custom Perl scripts. To ensure comparability across samples, read counts were normalized to one million aligned reads.
For transcriptome analysis of the S. ratti samples, we employed version 2.0.3 of TopHat aligner to map RNA-seq reads to the S. ratti genome and used version 2.0.1 of Cufflinks software to quantify expression levels (Trapnell et al. 2012). Tests for higher expression levels on autosomes with respect to the X chromosomes were done using a Wilcoxon ranksum test, as implemented in R.
Results
Appearance and number of giant nuclei
We observed that in the species studied, the giant nuclei in the distal gonad arms can take on various shapes depending on age. In general, they went from being relatively small and round or oval in young virgins (Fig. 2a) to becoming highly elongated, irregular, and large (>15 μm) in older females that are becoming infertile (Fig. 2b). This change in size and shape appeared to be more extreme in P. trichosuri than in the two species of Strongyloides (Fig. 2d). In these nuclei, frequently the nucleoli were clearly visible by DIC microscopy (Fig. 2e), as expected for nuclei in interphase. In all three species, the number of giant nuclei per gonad arm decreased significantly with age at least in females (Fig. 2f) but individual giant nuclei appeared to incorporate more DAPI in older worms than in younger ones, raising the question if their DNA content is higher. The number of giant nuclei per gonad arm ranged from around 20 to 50. The two arms within a single female tended to have similar but rarely equal numbers of giant nuclei. For example, among 33 S. papillosus females, only three had the same number in both arms while the maximum difference observed was 14. However, 25 (75 %) of the females fell within a range of 2–7 giant nuclei more in one arm than in the other. In 17 worms, the anterior arm had the higher number; in 13, this was the posterior arm.
Morphology and number of germline nuclei. DAPI-stained virgin (a) and old S. papillosus female (b), respectively. Arrows point to giant nuclei in the distal gonad arm. Note the change in size, number, and morphology of giant nuclei. The asterisks mark the position of the band of small nuclei. Note the change in position accompanied by an increase in number of small nuclei from virgin to old females. c DAPI-stained free-living S. papillosus male worms. Arrows point to the band of small nuclei. Note the change in position in males from L4 to adult. The band moves anteriorly over time reducing the space occupied by the giant nuclei in the gonad. Asterisks mark the anterior end of the worms. d DAPI-stained female P. trichosuri showing the extreme irregular giant nuclei (asterisk) in older infertile females. Giant nuclei insets show that often multiple giant nuclei are seen clumping together, and sometimes a thin line is seen within a nucleus (asterisks, See Suppl Movies 1 and 2). e DIC image showing the giant nuclei (outlined in white) in the distal gonad arm (outlined in yellow) showing a clearly visible nucleolus (outlined in orange, arrow heads). f The changes in the numbers of giant nuclei over age in females of all three species. For each species, there is a significant reduction in giant nuclei number over time
In males, a band of small presumably early spermatogonic nuclei (Triantaphyllou and Moncol 1977) migrates anteriorly with age, thereby increasing the volume filled with mature sperm and decreasing the volume containing giant nuclei (Fig 2c).
Differential DNA amplification in the giant nuclei
The giant nuclei in the distal arm have been postulated to be a result of repeated replication of the chromosomes in the absence of intervening cell divisions (Hammond and Robinson 1994). Although these authors had already noted that it was unlikely that the DNA content of the giant nuclei was the product of uniform full genome amplifications, no information about what sequences are amplified was available. We isolated distal arms of gonads containing giant nuclei from free-living adults of S. ratti, S. papillosus, and P. trichosuri and subjected them to quantitative sequencing (Figs. 3 and 4). In all species, it appeared that the portion of the genome that is present in only one copy in males (the X chromosome and the region undergoing chromatin diminution, respectively) was underrepresented in both sexes. More precisely, in XX/XO-based sex determination, one would expect a two times higher coverage of autosomes as compared to the X chromosome and the region undergoing chromatin diminution respectively in males, but equal coverage in females. Instead, we find a four- to sixfold increase in median coverage for autosomal regions in both sexes of S. ratti (Fig. 3a) and P. trichosuri (Fig. 4a) with no obvious difference between the sexes. Although it is difficult to analyze this type of data statistically, it appears that in S. papillosus (Fig. 4b), which does not have a free X chromosome, the difference is smaller (only about two- to threefold) and the underrepresentation might be slightly more pronounced in males than in females.
DNA and RNA sequencing of S. ratti giant nuclei a Graphs indicate the genome-wide distribution of coverage (non-overlapping 2-kb windows) obtained from sequencing giant nuclei in S. ratti females and males, respectively (f1, f2, and f3 indicate female biological replicates while m1 and m2 are male biological replicates). The higher coverage peak corresponds to the autosomes (blue) and the lower coverage peak is that of the X chromosome (red). Md indicates median values. b Chromosome-wide analysis of one female (left) and one male (right) replicate showing a relatively uniform amplification of DNA across the length of a chromosome (See Suppl Fig. 1a for the other replicates). The slight increase in coverage towards the left end of the X may or may not be real (see text). c Quantitative RNA sequencing from the giant nuclei in S. ratti female replicates shows that autosomal genes show a strong trend towards higher expression (p < 0.001, Wilcoxon ranksum test), consistent with the underrepresentation of the X chromosomal genes compared with the autosomal ones
DNA sequencing from P. trichosuri and S. papillosus giant nuclei. a, b Graphs plotting the genome-wide distribution of coverage (non-overlapping 2-kb windows) obtained from giant nuclei sequencing in P. trichosuri and S. papillosus, respectively (f1, f2, and f3 indicate female biological replicates while m1, m2, and m3 are male biological replicates). The higher coverage peak corresponds to the autosomes (blue) and the lower coverage peak is that of the X chromosome (red). The panel iL3 (bottom left) shows the equal coverage of autosomal and X chromosomal sequences obtained from sequencing infective larvae of S. papillosus, which are all female and lack giant nuclei in their gonads. This experiment serves as a control demonstrating that underexpression of the X chromosomes compared to the autosomes in the panels above is not a consequence of some feature of the X chromosome rendering it inefficient for sequencing. Md indicates median values
The high quality of the draft genome sequence available for S. ratti (Strongyloides sequencing consortium, submitted for publication) allowed us to analyze the chromatin amplification along the individual chromosomes (Fig 3b, Suppl Fig. 1a). While we cannot exclude slight differences, we found no clear indication of differential DNA amplification among different regions of chromosomes. In Fig. 3b, it appears as if sequences at the left end of chromosome X are present in somewhat higher copy number than the rest of the chromosome. However, the quality of the assembly of the X chromosome is not as good as for autosomes, and for the moment, we cannot tell if the left end is indeed amplified more than the rest of the X or if this slight increase is artificially caused due to genome assembly problems. To determine whether the lower copy number of the X chromosomes is also reflected in the levels of gene products, we also isolated and sequenced the RNA of the distal portion of the gonads from S. ratti females. Indeed, X-derived mRNAs are on average much less abundant than transcripts encoded on autosomes (Fig. 3c, Suppl Fig. 1b) while there is no difference between the two autosomes.
Presence or absence of male determining sperm
As explained in the “Introduction,” reproduction in the free-living generation of the two species of Strongyloides in question is sexual but, unlike in P. trichosuri, produces only females. In order to address if genetically male-determining (nullo-X) sperm exists, we isolated mature sperm from free-living males of both species of Strongyloides and from P. trichosuri and we quantitatively sequenced the sperm genomes (Fig. 5a). As expected, in P. trichosuri, which produces both sexes, X chromosomal sequences are covered by only about half as many reads as autosomal regions indicating that X-bearing and nullo-X sperm are present in about equal numbers (Fig. 5a, top left panel). In S. papillosus, on the other hand, we did not observe an underrepresentation of the X-derived sequences, confirming the genetic findings (Fig. 5a, top right panel). Surprisingly, X chromosomal sequences were underrepresented in mature sperm in S. ratti, indicating that nullo-X sperm are present (Fig. 5a, bottom panels). It must, however, be noted that the two independent experiments, shown in Fig. 5a bottom panels, differed rather strongly. While in one experiment (Fig. 5a, bottom right) the under representation of the X chromosome was very clear and the difference was very close to the expected 50 %, in the second one, the difference was less than expected (Fig. 5a, bottom left). This might indicate that the proportion of nullo-X sperm formed might be variable over time or cultures. Alternatively, it may be a consequence of stochastic fluctuations, which have to be taken into account given the very small amount of starting material available. However, there is additional evidence for the existence of nullo-X sperm in S. ratti. In contrast to S. papillosus (Nemetschke et al. 2010a), we observed two different karyotypes among the very young embryos of S. ratti, namely with five or with six chromosomes, which correspond to the diploid number of chromosomes for males and females, respectively (Fig. 5b, Suppl Movies 3, 4, and 5). In addition, within the uteri of S. ratti, but not S. papillosus females, we observed dying embryos among normally developing ones (Fig. 5c). Dying embryos were observed at a frequency of 13 % (n = 116). Although we do not know when exactly the embryos die and this number is therefore an underestimate, it is considerably less than the 50 % that would be expected if half of the sperm were nullo-X leading to non-viable embryos (see “Discussion”). For comparison with S. papillosus, 0 out of 55 embryos were scored to be developing abnormally.
X chromosomes in sperm of P. trichosuri, S. papillosus, and S. ratti. a DNA sequencing of mature sperm showing genome-wide distribution of coverage (non-overlapping 2-kb windows) of the autosomes (blue) compared with the X chromosome (red) of P. trichosuri (m1, top left), S. papillosus (m1, top right), and S. ratti (bottom panels m1 and m2), respectively. The X chromosome is underrepresented in P. trichosuri and S. ratti indicating presence of nullo-X sperm in these species. S. ratti (bottom panels, m1 and m2) shows variability in the amount of nullo-X sperm between replicates. Md indicates median values. b The two distinct karyotypes seen in early embryos of free-living S. ratti females, with 2n = 5 the expected male karyotype (top) and 2n = 6 the female karyotype (bottom). Also see Suppl Movies 3, 4, and 5. c Dying or abnormal embryos as seen in DAPI-stained free-living S. ratti females (top), marked with green asterisks, amid normally developing ones. In contrast no such abnormal embryos are observed in S. papillosus (bottom)
Discussion
Giant non-dividing nuclei had been noticed and described in the distal gonads of Strongyloides spp. by multiple authors over the years (see, for example, Basir 1950; Hammond and Robinson 1994; Triantaphyllou and Moncol 1977). For S. stercoralis, it had also been proposed that the DNA content of these nuclei is as high as several hundred C and that the exact DNA amount per nucleus could not have resulted from a succession of consecutive full genome duplications (Hammond and Robinson 1994). This suggests that different regions of the genome are amplified to variable extents. All these earlier studies were based on cytological observations using DNA binding dyes, and no information about the genomic regions amplified was available. Here we show that in S. ratti, S. papillosus and P. trichosuri X chromosomal regions (in S. papillosus the evolutionarily X chromosome-derived portion of the larger chromosome) are present in lower copy numbers than autosomal regions. Interestingly, for S. papillosus, in which the X chromosome is fused with an autosome (Kulkarni et al. 2013; Nemetschke et al. 2010a), the difference is smaller. While in males this difference can be partially explained by the lower dose of the X due to XX/XO sex determination, in females, it must be caused solely by differential amplification. Quantitative RNA sequencing consistently confirmed that X-linked genes are expressed at lower levels on average than autosomal genes (shown for S. ratti females), suggesting that the differential DNA amplification contributes to the control of gene expression in the germline. Underexpression of X chromosomal sequences compared with autosomal ones also occurs in the gonads of the model nematode C. elegans, and this phenomenon appears to be widespread among nematodes (Kelly et al. 2002). In C. elegans, the expression differential appears to be due to differential chromatin modifications and consequentially differential transcription from equal copy numbers. By contrast, in Strongyloides spp., an increase in autosomal copies might be an important determinant for the higher expression of autosomal genes. However, the about five to six times lower copy number of X chromosomal genes, compared with autosomal ones, does not fully explain the more than ten times lower median expression of X-linked genes. There must be additional gene-specific and/or chromosome-wide control mechanisms at work.
In general, endoreplication in the germline seems to be less common than in other tissues, with the exception of amplification of ribosomal RNA genes, which can be construed as an adaptation for rapid oogenesis. Incidentally, the giant nuclei have previously been proposed to act as nurse cells supporting the development of oocytes (Hammond and Robinson 1994), a role that in C. elegans is assumed by early meiotic cells in pachytene (Hubbard and Greenstein 2005). In this regard, it is interesting to note that one of the best-characterized nurse cells, those found in the egg chambers of the fruit fly, also amplify their genome (Bastock and St Johnston 2008; Lee et al. 2009). On closer inspection of our RNA data, we observed high expression of mRNAs encoding ribosomal components, chaperones, and proteasome components, among others. These signs of high gene expression activity further support that the giant nuclei of Strongyloides and Parastrongyloides act as nurse cells. An interesting though somewhat puzzling observation is the reduction in number of the giant nuclei in females with age. One possible explanation would be that some of them are broken up into small, diploid nuclei, thereby replenishing the population of germ cells available for oogenesis. Such a process has been proposed for certain snakes (Becak et al. 2003), and it was shown that some endopolyploid tumor cells use a meiosis-related mechanism to revert to normal diploidy (Erenpreisa et al. 2009; Ianzini et al. 2009). In fact, we noticed a significant (p < 0.01 in a t test) increase in small nuclei number from around 78 (±24.4 [standard deviation]) per arm in young S. papillosus females (n = 26 arms) to about 97 (±17.2) at peak reproductive age (n = 36 arms). The corresponding numbers for S. ratti were 56 (±16.0) in young (n = 18) and 83 (±19.1) in older (n = 20) worms. Nevertheless, we failed to observe any mitotic figures or condensed mitotic chromosomes in this region, a somewhat puzzling observation, which has however been reported before (Triantaphyllou and Moncol 1977). Alternatively, some giant nuclei may undergo apoptosis or fuse with each other, as suggested by the elongated shape of these nuclei (Fig. 2d giant nuclei insets, Suppl Movies 1 and 2). However, for the moment, these hypothetical explanations remain speculative and the actual dynamics of the giant nuclei in Strongyloides spp. will need to be investigated in live worms. To this end, GFP-tagged histone proteins expressed from transgenes, as has been established in C. elegans (Praitis et al. 2001), will be advantageous. For the moment, although transgenic techniques for Strongyloides spp. have been developed (Lok 2013), no germline-expressed promoters are available.
Most species of Strongyloides do not produce males in the progeny of the free-living generations (Streit 2008). Based on genetic arguments for S. papillosus (Nemetschke et al. 2010a), it was proposed that Strongyloides spp. males do not produce male-determining (nullo-X) sperm. However, DNA quantification using DNA binding dyes in S. stercoralis (Hammond and Robinson 1994) provided evidence that some species of Strongyloides might produce such sperm. While our quantitative sequencing of mature sperm confirmed absence of male-determining sperm for S. papillosus, we did find evidence for the presence of nullo-X sperm in S. ratti. Additionally, we also found early embryos with a male karyotype in this species. This might suggest that these two species use different strategies to prevent males among the infective larvae, either by avoidance of male-determining sperm as in S. papillosus or by inviability of male embryos as in S. ratti. In addition, nullo-X sperm might be less successful in fertilizing oocytes, reducing the number of “wasted” non-viable embryos. However, the difference might also be only quantitative and it might depend as much on the isolate as on the species. A very small proportion of nullo-X sperm (more precisely, sperm not containing a copy of the region undergoing chromatin diminution) in S. papillosus would probably have gone unnoticed in the earlier studies as our observations indicate that the number of nullo-X sperm in S. ratti might vary among experiments. A mechanism for producing an (variable) excess of X-bearing sperm has been described for the nematode Rhabditis sp. SB347 (Shakes et al. 2011). Therefore, it is also possible that both mechanisms are at work in both species but to variable extents.
The results presented here, on one hand, enhance our understanding of the reproductive biology of a fascinating group of parasitic nematodes. On the other hand, they also illustrate the usefulness of the Strongyloides/Parastrongyloides system for comparative evolutionary studies over very different phylogenetic distances. It will be highly revealing to further study the interesting differences (representing evolutionary changes) within the group and in comparison to other nematode model systems in evolutionary biology, in particular Caenorhabditis spp. and Pristionchus spp. (Sommer and Bumbarger 2012).
References
Albertson DG, Nwaorgu OC, Sulston JE (1979) Chromatin diminution and a chromosomal mechanism of sexual differentiation in Strongyloides papillosus. Chromosoma 75(1):75–87
Basir MA (1950) The morphology and development of the sheep nematode, Strongyloides papillosus (Wedl, 1856). Can J Res 28d:173–196
Bastock R, St Johnston D (2008) Drosophila oogenesis. Curr Biol 18(23):R1082–R1087. doi:10.1016/j.cub.2008.09.011
Becak ML, Becak W, Pereira A (2003) Somatic pairing, endomitosis and chromosome aberrations in snakes (Viperidae and Colubridae). An Acad Bras Cienc 75(3):285–300
Bleay C, Wilkes CP, Paterson S, Viney ME (2007) Density-dependent immune responses against the gastrointestinal nematode Strongyloides ratti. Int J Parasitol 37(13):1501–1509
Crook M, Viney ME (2005) The effect of non-immune stresses on the development of Strongyloides ratti. Parasitol 131(Pt 3):383–392
Dorris M, Viney ME, Blaxter ML (2002) Molecular phylogenetic analysis of the genus Strongyloides and related nematodes. Int J Parasitol 32(12):1507–1517
Eberhardt AG, Mayer WE, Streit A (2007) The free-living generation of the nematode Strongyloides papillosus undergoes sexual reproduction. Int J Parasitol 37:989–1000
Erenpreisa J, Cragg MS, Salmina K, Hausmann M, Scherthan H (2009) The role of meiotic cohesin REC8 in chromosome segregation in gamma irradiation-induced endopolyploid tumour cells. Exp Cell Res 315(15):2593–2603. doi:10.1016/j.yexcr.2009.05.011
Fenton A, Paterson S, Viney ME, Gardner MP (2004) Determining the optimal developmental route of Strongyloides ratti: an evolutionarily stable strategy approach. Evol Int J Org Evol 58(5):989–1000
Gemmill AW, Viney ME, Read AF (2000) The evolutionary ecology of host-specificity: experimental studies with Strongyloides ratti. Parasitol 120(Pt 4):429–437
Grant WN, Skinner SJ, Howes JN, Grant K, Shuttleworth G, Heath DD, Shoemaker CB (2006a) Heritable transgenesis of Parastrongyloides trichosuri: a nematode parasite of mammals. Int J Parasitol 36(4):475–483
Grant WN, Stasiuk S, Newton-Howes J, Ralston M, Bisset SA, Heath DD, Shoemaker CB (2006b) Parastrongyloides trichosuri, a nematode parasite of mammals that is uniquely suited to genetic analysis. Int J Parasitol 36(4):453–466
Hammond MP, Robinson RD (1994) Endoreplication in the ovary, testis, and intestine of Strongyloides stercoralis. J Parasitol 80(6):905–910
Harvey SC, Viney ME (2001) Sex determination in the parasitic nematode Strongyloides ratti. Genetics 158(4):1527–1533
Hubbard EJ, Greenstein D (2005) Introduction to the germ line. Wormbook : the online review of C elegans Biology:1–4. Doi:10.1895/wormbook.1.18.1, http://www.wormbook.org
Ianzini F, Kosmacek EA, Nelson ES, Napoli E, Erenpreisa J, Kalejs M, Mackey MA (2009) Activation of meiosis-specific genes is associated with depolyploidization of human tumor cells following radiation-induced mitotic catastrophe. Cancer Res 69(6):2296–2304. doi:10.1158/0008-5472.CAN-08-3364
Kelly WG, Schaner CE, Dernburg AF, Lee MH, Kim SK, Villeneuve AM, Reinke V (2002) X-chromosome silencing in the germline of C. elegans. Development 129(2):479–492
Kulkarni A, Dyka A, Nemetschke L, Grant WN, Streit A (2013) Parastrongyloides trichosuri suggests that XX/XO sex determination is ancestral in Strongyloididae (Nematoda). Parasitol 140:1822–1830. doi:10.1017/S0031182013001315
Lee HO, Davidson JM, Duronio RJ (2009) Endoreplication: polyploidy with purpose. Genes Dev 23(21):2461–2477. doi:10.1101/gad.1829209
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25(14):1754–1760. doi:10.1093/bioinformatics/btp324
Lok J (2013) piggyBac: a vehicle for integrative DNA transformation of parasitic nematodes. Mob Gen Elem 3(2):e24417. doi:10.4161/mge.24417
Mackerras MJ (1959) Strongyloides and Parastrongyloides (Nematoda: Rhabdiasoidea) in Australian marsupials. Aust J Zool 7(2):87–104
Nemetschke L, Eberhardt AG, Hertzberg H, Streit A (2010a) Genetics, chromatin diminution, and sex chromosome evolution in the parasitic nematode genus Strongyloides. Curr Biol 20(19):1687–1696
Nemetschke L, Eberhardt AG, Viney ME, Streit A (2010b) A genetic map of the animal-parasitic nematode Strongyloides ratti. Mol Biochem Parasitol 169(2):124–127
Nigon V, Roman E (1952) Le déterminisme du sexe et le development cyclique de Strongyloides ratti. Bull Biol Fr Belg 86:404–448
Olsen A, van Lieshout L, Marti H, Polderman T, Polman K, Steinmann P, Stothard R, Thybo S, Verweij JJ, Magnussen P (2009) Strongyloidiasis—the most neglected of the neglected tropical diseases? Trans R Soc Trop Med Hyg 103(10):967–972
Praitis V, Casey E, Collar D, Austin J (2001) Creation of low-copy integrated transgenic lines in Caenorhabditis elegans. Genetics 157(3):1217–1226
Rudel D, Riebesell M, Sommer RJ (2005) Gonadogenesis in Pristionchus pacificus and organ evolution: development, adult morphology and cell-cell interactions in the hermaphrodite gonad. Dev Biol 277(1):200–221
Shakes DC, Neva BJ, Huynh H, Chaudhuri J, Pires-daSilva A (2011) Asymmetric spermatocyte division as a mechanism for controlling sex ratios. Nat Commun 2:157. doi:10.1038/ncomms1160
Shao H, Li X, Nolan TJ, Massey HC Jr, Pearce EJ, Lok JB (2012) Transposon-mediated chromosomal integration of transgenes in the parasitic nematode Strongyloides ratti and establishment of stable transgenic lines. PLoS Pathog 8(8):e1002871. doi:10.1371/journal.ppat.1002871
Sommer RJ, Bumbarger DJ (2012) Nematode model systems in evolution and development. Wiley Interdiscip Rev Dev biol 1(3):389–400. doi:10.1002/wdev.33
Speare R (1989) Identification of species of Strongyloides. In: Grove DI (ed) Strongyloidiasis: a major roundworm infection of man. Taylor and Francis, London, pp 11–83
Stiernagle T (1999) Maintenance of C. elegans. In: Hope IA (ed) C. elegans a practical approach. The practical approach series. Oxford University Press, Oxford, pp 51–67
Streit A (2008) Reproduction in Strongyloides (Nematoda): a life between sex and parthenogenesis. Parasitol 135(3):285–294
Streit A (2014) How to become a parasite without sex chromosomes: a hypothesis for the evolution of Strongyloides spp. and related nematodes. Parasitol 141:1244–1254. doi:10.1017/S003118201400064X
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7(3):562–578. doi:10.1038/nprot.2012.016
Triantaphyllou AC, Moncol DJ (1977) Cytology, reproduction, and sex determination of Strongyloides ransomi and S. papillosus. J Parasitol 63(6):961–973
Viney ME, Lok JB (2007) Strongyloides spp. In: Community TCeR (ed) Wormbook wormbook DOI: 10.1895/wormbook.1.141.1, http://www.wormbook.org
Viney ME, Matthews BE, Walliker D (1992) On the biological and biochemical nature of cloned populations of Strongyloides ratti. J Helminthol 66(1):45–52
Viney ME, Green LD, Brooks JA, Grant WN (2002) Chemical mutagenesis of the parasitic nematode Strongyloides ratti to isolate ivermectin resistant mutants. Int J Parasitol 32(14):1677–1682
Viney ME, Steer MD, Wilkes CP (2006) The reversibility of constraints on size and fecundity in the parasitic nematode Strongyloides ratti. Parasitol 133(Pt 4):477–483
Yamada M, Matsuda S, Nakazawa M, Arizono N (1991) Species-specific differences in heterogonic development of serially transferred free-living generations of Strongyloides planiceps and Strongyloides stercoralis. J Parasitol 77(4):592–594
Acknowledgments
We thank James Lightfoot for critically reading early versions of this manuscript and anonymous reviewers for the valuable suggestions.
Compliance with ethical standards
ᅟ
Funding
This work was funded by the Max Planck Society.
Conflict of interest
The authors declare that they have no competing interests.
Animal experiments
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Suppl Fig. 1
a. Chromosome wide analysis of 2 female (f2 and f3) and 1 male (m2) replicate showing a relatively uniform amplification of DNA across the length of a chromosome. b. Biological replicates used for RNA sequencing from the distal arm containing the giant nuclei in S. ratti females show high reproducibility. (GIF 258 kb)
(GIF 322 kb)
Suppl Movies 1 and 2
3D reconstruction of giant nuclei (MOV 2062 kb)
Suppl Movies 3, 4, and 5
Movies 3 and 4 show male S. ratti embryonic karyotype and Movie 5 shows the female karyotype respectively. (AVI 315 kb)
Rights and permissions
About this article
Cite this article
Kulkarni, A., Holz, A., Rödelsperger, C. et al. Differential chromatin amplification and chromosome complements in the germline of Strongyloididae (Nematoda). Chromosoma 125, 125–136 (2016). https://doi.org/10.1007/s00412-015-0532-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-015-0532-y