Abstract
The clinical diagnosis of Alzheimer’s disease (AD) is a probabilistic formulation that may lack accuracy particularly at early stages of the dementing process. Abnormalities in amyloid-beta precursor protein (APP) metabolism and in the level of APP secretases have been demonstrated in platelets, and to a lesser extent in leukocytes, of AD patients, with conflicting results. The aim of the present study was to compare the protein level of the APP secretases A-disintegrin and metalloprotease 10 (ADAM10), Beta-site APP-cleaving enzyme 1 (BACE1), and presenilin-1 (PSEN1) in platelets and leukocytes from 20 non-medicated older adults with AD and 20 healthy elders, and to determine the potential use of these biomarkers to discriminate cases of AD from controls. The protein levels of all APP secretases were significantly higher in platelets compared to leukocytes. We found statistically a significant decrease in ADAM10 (52.5%, p < 0.0001) and PSEN1 (32%, p = 0.02) in platelets from AD patients compared to controls, but not in leukocytes. Combining all three secretases to generate receiver-operating characteristic (ROC) curves, we found a good discriminatory effect (AD vs. controls) when using platelets (the area under the curve—AUC—0.90, sensitivity 88.9%, specificity 66.7%, p = 0.003), but not in leukocytes (AUC 0.65, sensitivity 77.8%, specificity 50.0%, p = 0.2). Our findings indicate that platelets represent a better biological matrix than leukocytes to address the peripheral level of APP secretases. In addition, combining the protein level of ADAM10, BACE1, and PSEN1 in platelets, yielded a good accuracy to discriminate AD from controls.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The clinical diagnosis of Alzheimer’s disease (AD) is a probabilistic definition that takes into account the patient’s complaints, the objective characterization of cognitive impairment, family history of dementia, neuropsychiatric examination, laboratory tests, and brain imaging [1, 2]. New technologies have raised the possibility of using biological markers to reinforce the diagnosis of probable AD [3,4,5], addressing in vivo molecules related to core pathogenic mechanisms of AD and to surrogate markers of the neurodegenerative process, i.e., respectively related to the formation of neuritic plaques and neurofibrillary tangles [6,7,8].
The beta-amyloid peptide (Aβ) is the major component of plaques in AD, which are characteristically found in neocortical areas and in the hippocampus [9]. Amyloid-related biomarkers are commonly referred to as ‘molecular’ (core) markers of the disease process, e.g., cerebrospinal fluid (CSF) Aβ1−42 and amyloid-PET, as opposed to downstream markers of the neurodegenerative process (e.g., CSF Tau and phosphorylated Tau, hippocampal atrophy) [10, 11]. As compared to CSF-based biomarkers, which undoubtedly bear a closer relationship with the abnormalities that occur in the brain [12], the search for peripheral biomarkers of AD is justified by its better accessibility and tolerability, i.e., samples can be obtained by less invasive procedures [3]. In addition, blood-based biomarkers may be more adequate for longitudinal studies that require multiple sampling [13]. However, the accuracy of these methods critically relies on the development of technologies sensitive enough to ascertain changes that occur at much lower concentrations compared to the CSF, and that may be subject to the interference of multiple unspecific proteins expressed in the same biological matrix [12]. Recent studies presented promising results on plasma concentrations of neurofilament light (NFL) and Tau protein [14], but, to date, the search of amyloid-related biomarkers in the peripheral blood—largely represented by plasma concentrations of Aβ peptides—has yielded inconsistent results [3, 12].
The amyloid-β precursor protein (APP) is an integral type-I transmembrane protein present in several cell types [15,16,17,18,19,20]. It is a key element in the amyloid cascade, being the precursor of Aβ peptide, which is released by sequential cleavage of APP by β- and γ-secretase (amyloidogenic pathway). Alternatively, in the non-amyloidogenic pathway, APP cleavage by α- and γ-secretases precludes Aβ formation [21]. Beta-site APP-cleaving enzyme 1 (BACE1) has been reported to exert β-secretase activity, and A-disintegrin and metalloprotease 10 (ADAM10), a member of the ADAM family, has been associated with α-secretase activity [22,23,24]. Finally, presenilin-1 (PSEN1) comprises one of the four proteins of γ-secretase complex [25, 26]. Both central and peripheral biomarkers of the APP metabolism may provide important insights into the neurobiology of AD [27, 28], some of which with diagnostic implications [29,30,31].
Human platelets are largest source of circulating APP, and second most important source of APP beside the brain [32, 33]. In addition, platelets mainly express two of the three major APP variants (APP751 and APP770) [34,35,36,37] and bear the enzymatic machinery required for APP metabolism [16, 38, 39]. Peripheral blood leukocytes also contain the enzymatic machinery for APP processing and, as well as platelets, exhibit the isoforms APP751 and APP770 [40,41,42,43,44].
Considering that abnormalities in APP metabolism and in level of APP secretases have been consistently demonstrated in platelets of AD [27, 45,46,47,48,49,50], and to a lesser extent in leukocytes [51,52,53], sometimes with conflicting results, the aim of the present study is to compare the level of the APP secretases ADAM10, BACE1, and PSEN1 in platelets and leukocytes simultaneously drawn from AD patients and healthy controls, using the same methodology.
Methods
Subjects and assessment
The present study was conducted at the Laboratory of Neuroscience (LIM-27), Institute of Psychiatry, University of Sao Paulo, Brazil, according to the tenets of the Helsinki declaration. The protocol was approved by the local Ethics Committee (CAPPesq-HCFMUSP—No. 982.945/2015) and designed to include a cross-sectional assessment of the whole sample at baseline.
Forty physically healthy older adults were enrolled to the present study, being 20 patients with mild or moderate AD, whose diagnosis was established according to the Diagnostic and Statistical Manual of Mental Disorders—version IV (DSM-IV) criteria [54], and the specification of probable AD was made according to the National Institute of Neurological and Communicative Disorders and Stroke—Alzheimer’s Disease and Related Disorders Association (NINCDS-ADRDA) diagnostic criteria [55], and 20 age-matched healthy elderly adults with no evidence of cognitive impairment or psychiatric disorder at the clinical and neuropsychological assessment. Patients and controls were assessed by a geriatric psychiatrist and a neurologist using the Brazilian version of the Cambridge Examination for Mental Disorders of the Elderly (CAMDEX) structured interview [56], which includes a cognitive screening schedule Cambridge Cognition Examination (CAMCOG), from which the Mini-Mental State Examination (MMSE) scores can be drawn [57]. Cases with neuropsychiatric comorbidities or other neurodegenerative diseases were ruled out. Eligible participants were included in the study only if not under current treatment with medications with potential effect on platelet function, such as non-steroidal anti-inflammatory drugs (NSAIDs), aspirin, and clopidogrel. In addition, AD patients were not receiving antidementia treatment with acetylcholinesterase inhibitors or memantine at baseline. A 6-month wash-out period was required for participants with previous use of these or other drugs with relevant systemic and/or CNS effects (e.g., antidiabetic agents, lithium, neuroleptics, atypical antipsychotics, antidepressants, and anticonvulsants).
All participants (or respective caregivers in case of dementia) signed a written informed consent prior to enrollment.
There were no statistically significant differences in gender distribution (χ2 = 0.125; p = 0.7) and in age (t = − 0.71; Z = 2.22; p = 0.5) between diagnostic groups. AD patients were less educated (Mann–Whitney U test; Z = − 3.94; p < 0.0001) and had lower scores on cognitive tests (Mann–Whitney U test for both; MMSE, Z = − 3.78, p = 0.004; and CAMCOG, Z = − 5.05, p = 0.005) compared to healthy controls (Table 1).
Preparation of platelets and leukocytes
Platelets and leukocytes were used to determine the protein level of BACE1, ADAM10, and PSEN1 for patients and controls. Samples of peripheral blood (8 ml) were collected by venipuncture of the median cubital vein into 0.1-M sodium citrate-coated tubes (S-Monovett, Sarsted). This procedure was performed in the morning (between 8 and 10 a.m.) and after a 10–12-h fasting. Blood samples were homogenized with 200 µl of 0.09M ethylenediaminetetraacetic acid (EDTA) and centrifuged at 200×g for 10 min at room temperature. Platelet-rich plasma was separated from blood cells, and platelets were collected by centrifugation at 1159×g for 15 min at room temperature. Pellets were washed with 5 ml of 10-mM Tris, pH 7.4, and resuspended in lysis buffer containing 10-mM Tris–HCl, pH 7.4, 1-mM ethylene glycol tetra acetic acid (EGTA), 100-mM phenylmethanesulphonyl fluoride (PMSF), and protease inhibitors. Platelet homogenates were stored at − 80 °C. Regarding leukocytes, isolation was performed by lysing Red Blood Cell (RBC) (pH 7.6) with 0.01-M sodium chloride, 0.01-M Tris–HCl, pH 7.4, 5-mM magnesium chloride, and 0.001-M EDTA, vortexed for 10 min and centrifuged at 965×g for 15 min at 4 °C. Then, leukocyte pellet was resuspended twice in RBC lysis buffer, vortexed for 10 min, and centrifuged at 425×g for 10 min at 4 °C. Finally, pellet was resuspended in 1 mL of RBC lysis buffer and 10-µL PMSF and stored at − 80 °C. To perform the extraction of leukocyte proteins, the samples were thawed on ice and immediately spun down by centrifugation at 239×g for 15 min at 4 °C. The supernatants were discarded and the pellets resuspended with radio-immunoprecipitation assay (RIPA) buffer. After resuspension, samples were centrifuged again at 239×g for 15 min at 4 °C and the supernatant was kept for experimentation.
Determination of platelet and leukocyte protein levels of BACE1, ADAM10, and PSEN1
All procedures described below were similar for both platelets and leukocytes. Protein concentrations were determined in all aliquots by a modified Lowry method [58] using Bio-Rad DC protein kit assay (Bio-Rad Hercules) before immunoblotting. Aliquots holding 25 µg of total protein obtained from each of these samples were submitted to electrophoresis in 8–10% polyacrylamide gels, and transferred to nitrocellulose membranes by Western blotting. After blocking unspecific bindings, membranes were incubated with primary antibodies as follows: (i) 1 h with anti-BACE1 (Abcam, ab108394); (ii) overnight with anti-ADAM10 (Abcam, ab124695); (iii) overnight for platelet and 1 h for leukocyte with anti-PSEN1 (Abcam, ab76083). All membranes were then incubated for 1 h with anti-rabbit IgG, peroxidase-conjugated, secondary antibody (GE Healthcare Life Sciences, NA934V). Next, an enhanced chemoluminescence assay was performed using Amersham™ ECL™ Select Western Blotting Detection Reagent (GE Healthcare Life Sciences). Imaging was performed in Alliance 4.7 equipment (Uvitec Cambridge), which captures the chemiluminescent light emission from the reaction of the peroxidase conjugate with the ECL reagent. Light emission was captured for 5 min and the densitometry of distinct bands was performed with specific software tools. The densitometries of 56-, 65-, and 53-kDa bands were used to estimate the levels of BACE1, ADAM10, and PSEN1 respectively for both matrices (Fig. 1). Each sample was analyzed in duplicate, and additional blots of the same samples were performed whenever the reading between replicates displayed a variation > 15%. For normalization of leukocyte assay values, we used β-actin as loading (internal) control (IC), and membranes were incubated for 1 h with anti-beta-actin antibody (Abcam, ab6276) and then for 1 h with anti-mouse IgG, peroxidase-conjugated, secondary antibody (Sigma, A9044). For normalization of platelet assays, we used an internal standard (IS), which consisted of an in-house prepared suspension of platelets, generated from a pool of platelets drawn from young healthy volunteers. This IS was added to each gel to control for analytical differences between blots performed in different days (inter-assay variation). The IS readings in proportion to the 65, 56, and 53 kDa was then calculated, respectively yielding the normalized estimate of the ADAM10, BACE1, and PSEN1. The optical density of the secretases in leukocytes and in platelets was normalized on the optical density of the IC or IS, respectively.
Data analysis
All statistical analyses were performed with the aid of the software Statistical Package for the Social Sciences (SPSS)—version 14 [59]. Differences in sociodemographic characteristics, cognitive performance, leukocytes, and platelet secretases’ concentrations at baseline were analyzed by parametric (t test) or non-parametric tests (Mann–Whitney U test) between groups according to the data distribution. Differences in gender distribution were analyzed by Chi-squared tests. For MMSE and CAMCOG scores, a one-way analysis covariance (ANCOVA) was conducted with years of education used as covariate. Finally, the Wilcoxon signed test was used to compare the protein level between the two matrices (platelets and leukocytes) and a scatter plot was used for representation of the values for secretases in platelets and leukocytes.
Receiver-operating characteristic (ROC) curves were built to determine the diagnostic accuracy (sensitivity and specificity) and area under the curve (AUC) that best discriminates cases of AD from controls, using the level of APP secretases in leukocytes and platelets. Then, we evaluated the cut-off APP secretases densitometry values by Youden’s index. Statistical significance of the differences between groups was set at 5%.
Results
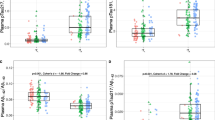
We found a statistically significant decrease in platelet level of ADAM10 (52.5% mean reduction) and PSEN1 (32% mean reduction) in AD patients compared to healthy subjects (Fig. 2a); with respect to the platelet level of BACE1, although 30% higher in AD patients, such difference was not statistically significant. Please refer to Table 1 for means and statistical test values.
Levels of BACE1, ADAM10, and PSEN1 in platelets and leukocytes. Scatter plot representation by Graph Pad: of BACE1, ADAM10, and PSEN1 levels in platelets (a) and leukocytes (b), according diagnosis; and of protein levels according to matrices irrespective of diagnostic group (c). BACE1 β-site APP-cleaving enzyme 1, ADAM10 A-Desintegrin and Metallopeptidase 10, PSEN1 presenilin 1, AD Alzheimer’s disease
As opposed to that, in leukocytes, we found a statistically significant reduction (40%) in the level of BACE1 in AD compared to controls, along with a significant reduction (40%) in PSEN1 in AD. No significant differences were found in the level of ADAM10 (p = 1.0) (Fig. 2b).
Comparing the level of APP secretases in these two biological matrices irrespective of diagnostic groups, we observed consistent and statistically significant differences in the level of BACE1 (sevenfold higher; Wilcoxon signed test, p < 0.0001), PSEN1 (threefold higher; Wilcoxon signed test, p < 0.0001), and ADAM10 (30% higher; Wilcoxon signed test, p = 0.05) in platelets as compared to leukocytes (Fig. 2c).
Next, we used Youden´s index to establish the best cut-off values based on the expression of APP secretases to determine sensitivity (Se) and specificity (Sp) parameters. ROC curves were built to determine the contribution of the level of APP secretases in platelets or leukocytes to classify cases (AD) and controls (Table 2). With respect to the level of BACE1, no statistically significant differences were observed comparing the AUC (AD vs. controls) in platelets (0.67; p = 0.15; 95% CI 0.5–0.9) or in leukocytes (0.72; p = 0.012; 95% CI 0.5–0.9) (Fig. 3a). Nonetheless, when evaluating the ROC curves obtained with the level of ADAM10 and PSEN1, the use of platelets proved more advantageous than leukocytes to discriminate the two groups. In platelets, the level of ADAM10 in AD vs. controls (Fig. 3b) yielded a significantly higher AUC (0.90, p = 0.001; 95% CI 0.8–1.0) and better sensitivity/specificity profile (Se 88.9%; Sp78.9%), compared to these parameters in leukocytes (AUC = 0.56, p = 0.64; 95% CI 0.3–0.8; Se 77.8%; Sp 36.8%). The level of PSEN1 in platelets yielded results in a similar trend (AUC = 0.80, p = 0.01; 95% CI 0.7–1.0; Se 77.8%; Sp 83.3%), as opposed to non-significant values in leukocytes (AUC = 0.69, p = 0.1; 95% CI 0.5–0.9; Se 77.8%; Sp 50.0%) (Fig. 3c). The combined analysis of values obtained with the three secretases indicated again that the level of ADAM10, BACE1, and PSEN1 in platelets (AUC = 0.86, p = 0.003; 95% CI 0.7–1.0; Se 88.9%; Sp 66.7%), but not in leukocytes (AUC = 0.65, p = 0.3; 95% CI 0.4–0.9; Se 77.8%; Sp 50.0%), may contribute to the classification of cases and controls (Fig. 3d).
Receiver-operating characteristic (ROC) curve analyses of the protein level of APP secretases in AD patients vs. controls a BACE1; b ADAM10; c PSEN1; d Combined analysis of all APP secretases. Platelet data represented with a continuous line, and leukocyte data with a dashed line. The reference line is represented in bold. BACE1 β-site APP-cleaving enzyme 1, ADAM10 A-Desintegrin and Metallopeptidase 10, PSEN1 presenilin 1
Discussion
In this study, we aimed to compare two blood-based peripheral cells, i.e., platelet and leukocyte, with respect to the protein level of the APP secretases ADAM10, BACE1, and PSEN1. In addition, we also intended to determine the diagnostic accuracy of these biomarkers to differentiate cases of AD from controls.
Our overall findings indicate that the level of all secretases, but particularly of BACE1 and PSEN1, is significantly higher in platelets compared to leukocytes. In general, our results in platelets are in line with the literature, reinforcing the notion of decreased level/activity of ADAM10 in AD [27, 46, 48, 50], along with less consensual findings about the level of BACE1 [49, 50] and PSEN1 [60].
Regarding the level of BACE1, which was not significantly different in platelets from AD vs. controls in the present sample, such levels in leukocytes were 40% decreased in AD. The level of BACE1 in leukocytes has been subject of fewer investigations [52]. BACE1 is widely expressed in the brain and is less present in other organs, locating predominantly in acidic intracellular compartments [61,62,63]. One study has already reported no significant differences in the mRNA level of BACE1 in lymphocytes from AD patients compared to controls [64, 65]. In contrast, Delvaux et al. [53] showed decreased BACE1 levels in leukocytes when compared to the levels of this protein in brain tissue corroborating our findings. Presumably, the level of BACE1 in leukocytes does not reflect the increased levels of this secretase in the AD cortex [66,67,68,69] or the increased activity/level of this enzyme in the CSF [70,71,72,73,74]. The fact which we found different BACE1 protein levels in leukocytes among diagnostic groups analyzed may be due to this protein has other physiological functions, since BACE1 has substrates besides APP, such transmembrane proteins, many of which function in cell signaling, immune, or inflammatory responses, and which suggests a role for BACE1 in this ability [75]. Regarding platelets, the increased protein level [45] and activity [23] of BACE1 have already been reported. A previous study of our group did not identify difference in levels of this protein when comparing AD with healthy control at baseline [50], as well as the present study. Furthermore, Sarno et al. [50] identified that long-term treatment with donepezil was associated with a decreased level of BACE1 in platelets, suggesting that the role of BACE1 in platelets is directly linked to the amyloidogenic pathway of proteolytic processing of APP.
We are unaware of other studies that addressed ADAM10 in AD leukocytes. In our study, the protein level of ADAM10 in leukocytes was similar in AD as compared to controls, although significantly reduced in AD platelets, as stated above. ADAM10 has played an important role in both the development and normal functioning of the brain [76]. Moreover, APP cleavage by ADAM10 liberates APP-alpha, which is involved in the regulation of synaptic plasticity and learning and memory (reviewed in [77]). Nonetheless, as BACE1, ADAM10 also presents other substrates and is involved in different pathophysiological processes besides AD such as embryonic development, cell adhesion, signal transduction, immune system, and cancer [78, 79]. Considering that ADAM10 has multiple functions and that, in our studies, we did not find a difference in their leukocyte protein levels between diagnostic groups, we can suggest that this matrix does not reflect the pathophysiological processes of AD, although it contains the necessary machinery for Aβ formation.
Similarly, the level PSEN1 in leukocytes was not significantly different in the two groups, whereas platelet PSEN1 levels were lower in the present sample of AD patients. Increased PSEN1 level has been reported in platelets from patients with AD [60], and studies in the post-mortem brain have shown conflicting results, i.e., similar level in AD and controls [80], increased levels in AD [81, 82], and decreased levels of PSEN1 in AD [83, 84]. Such divergent results may be due to PSEN1 is involved in both the pathways of APP, amyloidogenic and non-amyloidogenic [85], and that little is known about other pathophysiological functions of this protein [86].
The analysis of ROC curves using the level of each APP secretase separately indicated a good diagnostic accuracy (AUC > 0.80) to discriminate AD from controls when using platelet ADAM10 or platelet PSEN1; even better results were obtained with the combination of data of the protein level of all APP secretases in platelets to build the ROC curves. The high sensitivity (88.9%) attained with this method suggests the potential use of platelet (but not leukocyte) APP biomarkers in the diagnostic screening of AD.
Establishing the best matrix is extremely important for biomarker research. In this regard, platelets have proven a good biological matrix for the study of AD biomarkers, given their biochemical similarities with neurons, and the pathological changes that may mirror abnormalities that are central to many neuropsychiatric disorders [87,88,89,90,91]. In the present study, we provide evidence that those platelets are superior to leukocytes for this purpose. Nonetheless, we must acknowledge that the relatively small sample size and the distinct distribution of patients and controls with respect to the education level represent limitations of the present study, which we understand that may be attenuated by the fact that all participants were medication-free. With respect to the lower education level in the AD group compared to controls, this is a persistent finding in studies conducted in developing Countries and certainly in other Brazilian studies. This discrepancy can be possibly explained by the fact that low education is a well-established risk factor for cognitive decline and dementia. In the present study, analysis of covariance did not suggest any interfering effect of education level on the values obtained for protein expression in platelets or leukocytes.
Conclusions
In this study, we propose that platelets represent a better biological matrix compared to leukocytes to address AD-related changes in the regulation of APP metabolism. Although the protein level of ADAM10, as well as the combination of secretases (ADAM10, BACE1, and PSEN1) in platelets, may have a potential use as a peripheral biomarker of AD, further studies on this subject are necessary, so that the data obtained in platelets can be validated by other methodologies and reproduced by multicenter studies with a larger sample size.
References
Barthel H, Gertz H-J, Dresel S et al (2011) Cerebral amyloid-β PET with florbetaben (18F) in patients with Alzheimer’s disease and healthy controls: a multicentre phase 2 diagnostic study. Lancet Neurol 10:424–435. https://doi.org/10.1016/S1474-4422(11)70077-1
McKhann GM, Knopman DS, Chertkow H et al (2011) The diagnosis of dementia due to Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimer’s Dement 7:263–269. https://doi.org/10.1016/j.jalz.2011.03.005
Henriksen K, O’Bryant SE, Hampel H et al (2014) The future of blood-based biomarkers for Alzheimer’s disease. Alzheimer’s Dement 10:115–131. https://doi.org/10.1016/j.jalz.2013.01.013
O’Bryant SE (2016) Introduction to special issue on Advances in blood-based biomarkers of Alzheimer’s disease. Alzheimer’s Dement (Amst Neth) 3:110–112. https://doi.org/10.1016/j.dadm.2016.06.003
O’Bryant SE, Mielke MM, Rissman RA et al (2017) Blood-based biomarkers in Alzheimer disease: current state of the science and a novel collaborative paradigm for advancing from discovery to clinic. Alzheimer’s Dement 13:45–58. https://doi.org/10.1016/j.jalz.2016.09.014
Bussière T, Giannakopoulos P, Bouras C et al (2003) Progressive degeneration of nonphosphorylated neurofilament protein-enriched pyramidal neurons predicts cognitive impairment in Alzheimer’s disease: stereologic analysis of prefrontal cortex area 9. J Comp Neurol 463:281–302. https://doi.org/10.1002/cne.10760
Rodgers A (2008) Alzheimer’s disease: unraveling the mystery. Alzheimer’s Disease Education and Referral (ADEAR) Center, Silver Spring
Qin W, Ho L, Wang J et al (2009) S100A7, a novel Alzheimer’s disease biomarker with non-amyloidogenic α-secretase activity acts via selective promotion of ADAM-10. PLoS One 4:e4183. https://doi.org/10.1371/journal.pone.0004183
Yoon S-S, Jo SA (2012) Mechanisms of amyloid-β peptide clearance: potential therapeutic targets for Alzheimer’s disease. Biomol Ther (Seoul) 20:245–255. https://doi.org/10.4062/biomolther.2012.20.3.245
Jack CR, Knopman DS, Jagust WJ et al (2010) Hypothetical model of dynamic biomarkers of the Alzheimer’s pathological cascade. Lancet Neurol 9:119–128. https://doi.org/10.1016/S1474-4422(09)70299-6
Blennow K, Dubois B, Fagan AM et al (2015) Clinical utility of cerebrospinal fluid biomarkers in the diagnosis of early Alzheimer’s disease. Alzheimer’s Dement 11:58–69. https://doi.org/10.1016/j.jalz.2014.02.004
Blennow K (2017) A review of fluid biomarkers for Alzheimer’s disease: moving from CSF to blood. Neurol Ther 6:15–24. https://doi.org/10.1007/s40120-017-0073-9
Galasko D, Golde TE (2013) Biomarkers for Alzheimer’s disease in plasma, serum and blood—conceptual and practical problems. Alzheimers Res Ther 5:10. https://doi.org/10.1186/alzrt164
Gisslén M, Price RW, Andreasson U et al (2016) Plasma Concentration of the neurofilament light protein (NFL) is a biomarker of CNS injury in HIV infection: a cross-sectional study. EBioMedicine 3:135–140. https://doi.org/10.1016/j.ebiom.2015.11.036
Bayer T, Cappai R, Masters C et al (1999) It all sticks together—the APP-related family of proteins and Alzheimer’s disease. Mol Psychiatry 4:524–528. https://doi.org/10.1038/sj.mp.4000552
Di Luca M, Colciaghi F, Pastorino L et al (2000) Platelets as a peripheral district where to study pathogenetic mechanisms of Alzheimer disease: the case of amyloid precursor protein. Eur J Pharmacol 405:277–283
Skovronsky DM, Lee VM-Y, Praticò D (2001) Amyloid precursor protein and amyloid β peptide in human platelets. J Biol Chem 276:17036–17043. https://doi.org/10.1074/jbc.M006285200
Racchi M, Govoni S (2003) The pharmacology of amyloid precursor protein processing. Exp Gerontol 38:145–157
Haass C (2004) Take five–BACE and the gamma-secretase quartet conduct Alzheimer’s amyloid beta-peptide generation. EMBO J 23:483–488. https://doi.org/10.1038/sj.emboj.7600061
Zainaghi IA, Forlenza OV, Gattaz WF (2007) Abnormal APP processing in platelets of patients with Alzheimer’s disease: correlations with membrane fluidity and cognitive decline. Psychopharmacology 192:547–553. https://doi.org/10.1007/s00213-007-0748-5
Cole SL, Vassar R (2008) BACE1 structure and function in health and Alzheimer’s disease. Curr Alzheimer Res 5:100–120. https://doi.org/10.2174/156720508783954758
Seals DF, Courtneidge SA (2003) The ADAMs family of metalloproteases: multidomain proteins with multiple functions. Genes Dev 17:7–30. https://doi.org/10.1101/gad.1039703
Chang C, Werb Z (2001) The many faces of metalloproteases: cell growth, invasion, angiogenesis and metastasis. Trends Cell Biol 11:S37-43
Vingtdeux V, Marambaud P (2012) Identification and biology of α-secretase. J Neurochem 120:34–45. https://doi.org/10.1111/j.1471-4159.2011.07477.x
Schellenberg G, Bird T, Wijsman E et al (1992) Genetic linkage evidence for a familial Alzheimer’s disease locus on chromosome 14. Science 258:668–671. https://doi.org/10.1126/science.1411576
De Strooper B (2003) Aph-1, Pen-2, and nicastrin with Presenilin generate an active γ-secretase complex. Neuron 38:9–12
Tang K, Hynan LS, Baskin F, Rosenberg RN (2006) Platelet amyloid precursor protein processing: a bio-marker for Alzheimer’s disease. J Neurol Sci 240:53–58. https://doi.org/10.1016/j.jns.2005.09.002
Zainaghi IA, Talib LL, Diniz BS et al (2012) Reduced platelet amyloid precursor protein ratio (APP ratio) predicts conversion from mild cognitive impairment to Alzheimer’s disease. J Neural Transm 119:815–819. https://doi.org/10.1007/s00702-012-0807-x
Diniz BSO, Pinto JA, Forlenza OV (2008) Do CSF total tau, phosphorylated tau, and β-amyloid 42 help to predict progression of mild cognitive impairment to Alzheimer’s disease? A systematic review and meta-analysis of the literature. World J Biol Psychiatry 9:172–182. https://doi.org/10.1080/15622970701535502
Forlenza OV, Diniz BS, Teixeira AL et al (2015) Lower cerebrospinal fluid concentration of brain-derived neurotrophic factor predicts progression from mild cognitive impairment to Alzheimer’s disease. NeuroMol Med 17:326–332. https://doi.org/10.1007/s12017-015-8361-y
Forlenza OV, Radanovic M, Talib LL et al (2015) Cerebrospinal fluid biomarkers in Alzheimer’s disease: diagnostic accuracy and prediction of dementia. Alzheimer’s dement diagnosis. Assess Dis Monit 1:455–463. https://doi.org/10.1016/j.dadm.2015.09.003
Gardella JE, Gorgone GA, Newman P et al (1992) Characterization of Alzheimer amyloid precursor protein transcripts in platelets and megakarocytes. Neurosci Lett 138:229–232. https://doi.org/10.1016/0304-3940(92)90921-S
Veitinger M, Varga B, Guterres SB, Zellner M (2014) Platelets, a reliable source for peripheral Alzheimer’s disease biomarkers? Acta Neuropathol Commun 2:65. https://doi.org/10.1186/2051-5960-2-65
Rosenberg RN, Baskin F, Fosmire JA et al (1997) Altered amyloid protein processing in platelets of patients with Alzheimer disease. Arch Neurol 54:139–144
Bush AI, Tanzi RE (1998) Alzheimer-disease-related abnormalities of amyloid β precursor protein isoforms in the platelet—the brains delegate in the periphery. Arch Neurol 55:1179–1180. https://doi.org/10.1001/archneur.55.9.1179
Di Luca M (1998) Differential level of platelet amyloid beta precursor protein isoforms: an early marker for Alzheimer disease. Arch Neurol 55:1195–1200. https://doi.org/10.1001/archneur.55.9.1195
Li Q, Whyte S, Tanner J et al (1998) Secretion of Alzheimer’s disease Abeta amyloid peptide by activated human platelets. Lab Investig 78:461–469
Li QX, Evin G, Small DH et al (1995) Proteolytic processing of Alzheimer’s disease ?βA4 amyloid precursor protein in human platelets. J Biol Chem 270:14140–14147. https://doi.org/10.1074/jbc.270.23.14140
Evin G, Zhu A, Holsinger RMD et al (2003) Proteolytic processing of the Alzheimer’s disease amyloid precursor protein in brain and platelets. J Neurosci Res 74:386–392. https://doi.org/10.1002/jnr.10745
Golde TE, Estus S, Usiak M et al (1990) Expression of beta amyloid protein precursor mRNAs: recognition of a novel alternatively spliced form and quantitation in alzheimer’s disease using PCR. Neuron 4:253–267. https://doi.org/10.1016/0896-6273(90)90100-T
Mönning U, König G, Banati RB et al (1992) Alzheimer beta A4-amyloid protein precursor in immunocompetent cells. J Biol Chem 267:23950–23956
Li QX, Berndt MC, Bush AI et al (1994) Membrane-associated forms of the beta A4 amyloid protein precursor of Alzheimer’s disease in human platelet and brain: surface expression on the activated human platelet. Blood 84:133–142
Delvaux E, Bentley K, Stubbs V et al (2013) Differential processing of amyloid precursor protein in brain and in peripheral blood leukocytes. Neurobiol Aging 34:1680–1686. https://doi.org/10.1016/j.neurobiolaging.2012.12.004
Schuck F, Wolf D, Fellgiebel A, Endres K (2016) Increase of α-secretase ADAM10 in platelets along cognitively healthy aging. J Alzheimer’s Dis 50:817–826. https://doi.org/10.3233/JAD-150737
Vidal R, Ghiso J, Wisniewski T, Frangione B (1996) Alzheimer’s presenilin 1 gene expression in platelets and megakaryocytes. FEBS Lett 393:19–23. https://doi.org/10.1016/0014-5793(96)00845-9
Colciaghi F, Borroni B, Pastorino L et al (2002) [alpha]-Secretase ADAM10 as well as [alpha]APPs is reduced in platelets and CSF of Alzheimer disease patients. Mol Med 8:67–74
Decourt B, Walker A, Gonzales A et al (2013) Can platelet BACE1 levels be used as a biomarker for Alzheimer’s disease? Proof-of-concept study. Platelets 24:235–238. https://doi.org/10.3109/09537104.2012.688899
Manzine PR, de França Bram JM, Barham EJ et al (2013) ADAM10 as a biomarker for Alzheimer’s disease: a Study with Brazilian Elderly. Dement Geriatr Cogn Disord 35:58–66. https://doi.org/10.1159/000345983
Marksteiner J, Humpel C (2013) Platelet-derived secreted amyloid-precursor protein-β as a marker for diagnosing Alzheimer’s disease. Curr Neurovasc Res 10:297–303
Sarno TA, Talib LL, Joaquim HPG et al (2017) Protein expression of BACE1 is downregulated by Donepezil in Alzheimer’s disease platelets. J Alzheimers Dis 55:1445–1451. https://doi.org/10.3233/JAD-160813
Rogaev EI, Sherrington R, Wu C et al (1997) Analysis of the 5? Sequence, genomic structure, and alternative splicing of thepresenilin-1gene (psen1) associated with early onset Alzheimer disease. Genomics 40:415–424. https://doi.org/10.1006/geno.1996.4523
Decourt B, Sabbagh MN (2011) BACE1 as a potential biomarker for Alzheimer’s disease. J Alzheimers Dis 24(Suppl 2):53–59. https://doi.org/10.3233/JAD-2011-110017
Delvaux E, Bentley K, Stubbs V et al Differential processing of amyloid precursor protein in brain and in peripheral blood leukocytes. https://doi.org/10.1016/j.neurobiolaging.2012.12.004
American Psychiatric Association (2000) Diagnostic and statistical manual of mental disorders, 4th edn. American Psychiatric Association, Washington, DC
McKhann G, Drachman D, Folstein M et al (1984) Clinical diagnosis of Alzheimer’s disease: report of the NINCDS-ADRDA Work Group* under the auspices of Department of Health and Human Services Task Force on Alzheimer’s Disease. Neurology 34:939–939. https://doi.org/10.1212/WNL.34.7.939
Roth M, Tym E, Mountjoy CQ (1986) CAMDEX. A standardised instrument for the diagnosis of mental disorder in the elderly with special reference to the early detection of dementia. Br J Psychiatry 149:698–709. https://doi.org/10.1192/bjp.149.6.698
Folstein MF, Folstein SE, McHugh PR (1975) “Mini-mental state”. A practical method for grading the cognitive state of patients for the clinician. J Psychiatr Res 12:189–198. https://doi.org/10.1016/0022-3956(75)90026-6
Lowry O, Rosebrough N, Farr A, Randall R (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193:265–275
SPSS Incorporation (2006) Statistical Package for Social Science (SPSS). Release version 14 Chicago (IL). In: SPSS Inc. http://www.scielo.br/scielo.php?script=sci_nlinks&ref=000114&pid=S0365-0596201000010000300018&lng=pt. Accessed 12 Mar 2018
Bermejo-Bescós P, Martín-Aragón S, Jiménez-Aliaga K et al (2013) Processing of the platelet amyloid precursor protein in the mild cognitive impairment (MCI). Neurochem Res 38:1415–1423. https://doi.org/10.1007/s11064-013-1039-7
Vassar R (1999) Beta-secretase cleavage of Alzheimer’s amyloid precursor protein by the transmembrane aspartic protease BACE. Science 286:735–741. https://doi.org/10.1126/science.286.5440.735
Marcinkiewicz M, Seidah NG (2000) Coordinated expression of β-amyloid precursor protein and the putative β-secretase BACE and α-secretase ADAM10 in mouse and human brain. J Neurochem 75:2133–2143. https://doi.org/10.1046/j.1471-4159.2000.0752133.x
Yan R, Vassar R (2014) Targeting the β secretase BACE1 for Alzheimer’s disease therapy. Lancet Neurol 13:319–329. https://doi.org/10.1016/S1474-4422(13)70276-X
Herrera-Rivero M, Soto-Cid A, Hernández ME, Aranda-Abreu GE (2013) Tau, APP, NCT and BACE1 in lymphocytes through cognitively normal ageing and neuropathology. An Acad Bras Cienc 85:1489–1496. https://doi.org/10.1590/0001-376520130013
Herrera-Rivero M, Elena Hernández-Aguilar M, Emiliano Aranda-Abreu G (2015) A strategy focused on MAPT, APP, NCSTN and BACE1 to build blood classifiers for Alzheimer’s disease. J Theor Biol 376:32–38. https://doi.org/10.1016/j.jtbi.2015.03.039
Holsinger RMD, McLean CA, Beyreuther K et al (2002) Increased expression of the amyloid precursor β-secretase in Alzheimer’s disease. Ann Neurol 51:783–786. https://doi.org/10.1002/ana.10208
Yang L-B, Lindholm K, Yan R et al (2003) Elevated β-secretase expression and enzymatic activity detected in sporadic Alzheimer disease. Nat Med 9:3–4. https://doi.org/10.1038/nm0103-3
Fukumoto H (2002) β-secretase protein and activity are increased in the neocortex in Alzheimer disease. Arch Neurol 59:1381. https://doi.org/10.1001/archneur.59.9.1381
Ahmed RR, Holler CJ, Webb RL et al (2010) BACE1 and BACE2 enzymatic activities in Alzheimer’s disease. J Neurochem 112:1045–1053. https://doi.org/10.1111/j.1471-4159.2009.06528.x
Holsinger RMD, Lee JS, Boyd A et al (2006) CSF BACE1 activity is increased in CJD and Alzheimer disease versus other dementias. Neurology 67:710–712. https://doi.org/10.1212/01.wnl.0000229925.52203.4c
Verheijen JH, Huisman LGM, Van Lent N et al (2006) Detection of a soluble form of BACE-1 in human cerebrospinal fluid by a sensitive activity assay. Clin Chem 52:1168–1174. https://doi.org/10.1373/clinchem.2006.066720
Zhong Z, Ewers M, Teipel S et al (2007) Levels of β-secretase (BACE1) in cerebrospinal fluid as a predictor of risk in mild cognitive impairment. Arch Gen Psychiatry 64:718. https://doi.org/10.1001/archpsyc.64.6.718
Zetterberg H, Andreasson U, Hansson O et al (2008) Elevated Cerebrospinal Fluid BACE1 Activity in Incipient Alzheimer Disease. Arch Neurol 65:1102–1107. https://doi.org/10.1001/archneur.65.8.1102
Mulder SD, van der Flier WM, Verheijen JH et al (2010) BACE1 activity in cerebrospinal fluid and its relation to markers of AD pathology. J Alzheimer’s Dis 20:253–260. https://doi.org/10.3233/JAD-2010-1367
Vassar R, Kandalepas PC (2011) The β-secretase enzyme BACE1 as a therapeutic target for Alzheimer’s disease. Alzheimers Res Ther 3:20. https://doi.org/10.1186/alzrt82
Endres K, Deller T (2017) Regulation of alpha-secretase ADAM10 in vitro and in vivo: genetic, epigenetic, and protein-based mechanisms. Front Mol Neurosci 10:56. https://doi.org/10.3389/fnmol.2017.00056
Kögel D, Deller T, Behl C (2012) Roles of amyloid precursor protein family members in neuroprotection, stress signaling and aging. Exp Brain Res 217:471–479. https://doi.org/10.1007/s00221-011-2932-4
Edwards DR, Handsley MM, Pennington CJ (2009) The ADAM metalloproteinases. Mol Aspects Med 29:258–289
Reiss K, Saftig P (2009) The A Disintegrin And Metalloprotease (ADAM) family of sheddases: physiological and cellular functions. Semin Cell Dev Biol 20:126–137. https://doi.org/10.1016/j.semcdb.2008.11.002
Mathews PM, Cataldo AM, Kao BH et al (2000) Brain expression of presenilins in sporadic and early-onset, familial Alzheimer’s disease. Mol Med 6:878–891
Borghi R, Piccini A, Barini E et al (2010) Upregulation of presenilin 1 in brains of sporadic, late-onset Alzheimer’s disease. J Alzheimer’s Dis 22:771–775. https://doi.org/10.3233/JAD-2010-100729
Kakuda N, Shoji M, Arai H et al (2012) Altered γ-secretase activity in mild cognitive impairment and Alzheimer’s disease. EMBO Mol Med 4:344–352. https://doi.org/10.1002/emmm.201200214
Davidsson P, Bogdanovic N, Lannfelt L, Blennow K (2001) Reduced expression of amyloid precursor protein, presenilin-1 and rab3a in cortical brain regions in Alzheimer’s disease. Dement Geriatr Cogn Disord 12:243–250
Verdile G, Gnjec A, Miklossy J et al (2004) Protein markers for Alzheimer disease in the frontal cortex and cerebellum. Neurology 63:1385–1392. https://doi.org/10.1212/01.WNL.0000141848.45315.A6
Capell A, Beher D, Prokop S et al (2004) γ-secretase complex assembly within the early secretory pathway*. https://doi.org/10.1074/jbc.M409106200
Bustos V, Pulina MV, Bispo A et al (2017) Phosphorylated Presenilin 1 decreases β-amyloid by facilitating autophagosome–lysosome fusion. Proc Natl Acad Sci USA 114:7148–7153. https://doi.org/10.1073/pnas.1705240114
Pletscher A, Laubscher A (1980) Blood platelets as models for neurons: uses and limitations. J Neural Transm. https://doi.org/10.1007/978-3-7091-8582-7_2
Freson K, Labarque V, Thys C et al (2007) What’s new in using platelet research? To unravel thrombopathies and other human disorders. Eur J Pediatr 166:1203–1210. https://doi.org/10.1007/s00431-007-0543-7
Catricala S, Torti M, Ricevuti G (2012) Alzheimer disease and platelets: how’s that relevant. Immun Ageing 9:20. https://doi.org/10.1186/1742-4933-9-20
Talib LL, Joaquim HP, Forlenza OV (2012) Platelet biomarkers in Alzheimer’s disease. World J psychiatry 2:95–101. https://doi.org/10.5498/wjp.v2.i6.95
Behari M, Shrivastava M (2013) Role of platelets in neurodegenerative diseases: a universal pathophysiology. Int J Neurosci 123:287–299. https://doi.org/10.3109/00207454.2012.751534
Acknowledgements
Financial support: FAPESP (Fundação de Amparo à Pesquisa de São Paulo, Grant nos 2013/20695-3 and 2009/52825-8), CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico, Project 554535/2005-0), and ABADHS (Associação Beneficente Alzira Denise Hertzog da Silva).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None.
Rights and permissions
About this article
Cite this article
Bram, J.M.F., Talib, L.L., Joaquim, H.P.G. et al. Protein levels of ADAM10, BACE1, and PSEN1 in platelets and leukocytes of Alzheimer’s disease patients. Eur Arch Psychiatry Clin Neurosci 269, 963–972 (2019). https://doi.org/10.1007/s00406-018-0905-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00406-018-0905-3