Abstract
Introduction
Classification of variants of unknown significance (VUS) in the breast cancer genes BRCA1 and BRCA2 changes with accumulating evidence for clinical relevance. In most cases down-staging towards neutral variants without clinical significance is possible.
Methods
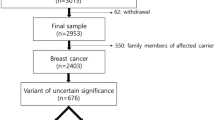
We searched the database of the German Consortium for Hereditary Breast and Ovarian Cancer (GC-HBOC) for changes in classification of genetic variants as an update to our earlier publication on genetic variants in the Centre of Dresden. Changes between 2015 and 2017 were recorded.
Results
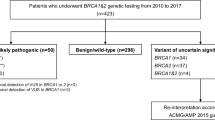
In the group of variants of unclassified significance (VUS, Class 3, uncertain), only changes of classification towards neutral genetic variants were noted. In BRCA1, 25% of the Class 3 variants (n = 2/8) changed to Class 2 (likely benign) and Class 1 (benign). In BRCA2, in 50% of the Class 3 variants (n = 16/32), a change to Class 2 (n = 10/16) or Class 1 (n = 6/16) was observed. No change in classification was noted in Class 4 (likely pathogenic) and Class 5 (pathogenic) genetic variants in both genes. No up-staging from Class 1, Class 2 or Class 3 to more clinical significance was observed.
Conclusion
All variants with a change in classification in our cohort were down-staged towards no clinical significance by a panel of experts of the German Consortium for Hereditary Breast and Ovarian Cancer (GC-HBOC). Prevention in families with Class 3 variants should be based on pedigree based risks and should not be guided by the presence of a VUS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
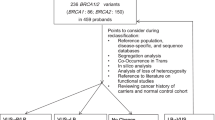
Classification of genetic variants of the genes BRCA1 and BRCA2 is an ongoing process. About 14% of all variants that are found in these genes are variants of unclassified significance (VUS, Class 3, uncertain) as reported earlier [1]. The German Consortium for Hereditary Breast and Ovarian Cancer (GC-HBOC) updates the classification of all genetic variants in close cooperation with the ENIGMA Consortium (Evidence-based Network for the Interpretation of Germline Mutant Alleles), which bases its classification on the ACMG guidelines [2]. This work is done by a panel of experts. According to Plon et al., variants are classified into Classes 1–5 with Class 1 being benign without any clinical relevance and Class 5 being a pathogenic mutation [3]. In our work, on genetic BRCA1/2 variants in the Centre of Dresden, we listed VUS Class 3 (uncertain) for BRCA1 and for BRCA2, but due to the ongoing classification process, some of them are currently classified differently. In all of the cases, the variants were down-staged to less important variants of Class 2 (likely benign) or Class 1 (benign). In BRCA1 25% of the Class 3 variants (n = 2/8) changed to Class 2 (likely benign) and Class 1 (benign). In BRCA2, in 50% of the Class 3 variants (n = 16/32), a change to Class 2 (n = 10/16) or Class 1 (n = 6/16) was observed. No change in classification was noted in Class 4 (likely pathogenic) and Class 5 (pathogenic) genetic variants in both genes. No up-staging from Class 1, Class 2 or Class 3 to more clinical significance was observed. Altogether, in BRCA1, 16 index patients and, in BRCA2, 57 index patients were affected by the change in classification. Instead of the initially reported VUS rate of 14%, we now report a VUS rate of 6%. The changes are the following: BRCA1: c.591C>T, Class 2; c.1486C>T, Class 1; BRCA2: c.5199C>T, Class 2; c.4068G>A, Class 2; c.10095delCins11, Class 2; c.7347T>C, Class 2; c.6322C>T, Class 1; c.3516G>A, Class 2; c.68-7T>A, Class 2; c.6540G>C, Class 2; c.4054G>T, Class 2; c.7017G>C, Class 1; c.10234A>G, Class 1; c.9730G>A, Class 1; c.198A>G, Class 2; c.1395A>C, Class 2; c.3264T>C, Class 1; c.5418A>G, Class 1; c.8694G>A, Class 2; c.10110G>A, Class 2. The GC-HBOC informs all its centres via a recall system on revisions in the classification of genetic variants in its joint database [5]. Based on the changes from Class 3 to Class 2 or Class 1, no modification of clinical recommendations is needed. Since most VUS Class 3 will result in benign effects, intensified screening is only justified in families with high risks according to pedigree-based calculation models as Cyrillic or BOADICEA [2]. This is also the case in families without any abnormal finding in the molecular analysis of the breast and ovarian cancer genes. Because of lacking clinical consequences, some have already suggested not to disclose the result of a Class 3 variant to patients [4]. However, segregation analysis on research basis, where appropriate according to pedigree information, is an important aspect for the classification of variants and collection of blood from other family members would not be possible without basic information about the VUS. For good clinical practice, it is important to state that intensified surveillance or prophylactic operations are not justified by VUS. Patients with analysis results with VUS outside the GC-HBOC are recommended to contact their geneticist for an update on classification after a few years.
References
Meisel C, Sadowski CE, Kohlstedt D, Keller K, Staritz F, Grubling N, Becker K, Mackenroth L, Rump A, Schrock E et al (2017) Spectrum of genetic variants of BRCA1 and BRCA2 in a German single center study. Arch Gynecol Obstet 295(5):1227–1238
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, Grody WW, Hegde M, Lyon E, Spector E et al (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med Off J Am Coll Med Genet 17(5):405–424
Plon SE, Eccles DM, Easton D, Foulkes WD, Genuardi M, Greenblatt MS, Hogervorst FB, Hoogerbrugge N, Spurdle AB, Tavtigian SV (2008) Sequence variant classification and reporting: recommendations for improving the interpretation of cancer susceptibility genetic test results. Hum Mutat 29(11):1282–1291
Eccles DM, Mitchell G, Monteiro AN, Schmutzler R, Couch FJ, Spurdle AB, Gomez-Garcia EB (2015) BRCA1 and BRCA2 genetic testing-pitfalls and recommendations for managing variants of uncertain clinical significance. Ann Oncol Off J Eur Soc Med Oncol ESMO 26(10):2057–2065
Waha A, Versmold B, Kast K, Kiechle M, Ditsch N, Meindl A, Niederacher D, Hahnen E, Arnold N, Mundhenke C, Horvath J, Auber B, Dikow N, Hauke J, Wappenschmidt B, Riess O, Schott S, Speiser D, Faust U, Sutter C, Rhiem K, Schmutzler RK (2017) Konsensusempfehlung des Deutschen Konsortiums Familiärer Brust- und Eierstockkrebs zum Umgang mit Ergebnissen der Multigenanalyse. Geburtshilfe Frauenheilkd 77:733–739. https://doi.org/10.1055/s-0043-108531
Funding
German Cancer Aid Grant No. 110837 for GC-HBOC.
Author information
Authors and Affiliations
Contributions
KK: conception of the work, analysis and interpretation of data, critical revision of the manuscript for important intellectual content, and final approval of the version to be published. PW: critical revision of the manuscript for important intellectual content, and final approval of publication. NA: interpretation of data, critical revision of the manuscript for important intellectual content, and final approval of publication.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional research committee (No. EK 162072007) and with the 1064 Helsinki declaration and its later amendments or comparable ethical standards. This article does not contain any studies with animals performed by any of the authors.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Rights and permissions
About this article
Cite this article
Kast, K., Wimberger, P. & Arnold, N. Changes in classification of genetic variants in BRCA1 and BRCA2. Arch Gynecol Obstet 297, 279–280 (2018). https://doi.org/10.1007/s00404-017-4631-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00404-017-4631-2