Abstract
Purpose
The present study aimed to evaluate the effect of chia flour associated with a high fat diet on intestinal health in female ovariectomized Wistar rats.
Methods
The study was conducted with 32 adult female ovariectomized Wistar rats, which were separated into four groups: standard diet (ST), standard diet with chia (STC), high fat diet (HF) and high fat diet with chia (HFC) for 18 weeks. Cecum content pH, short chain fatty acid content, brush border membrane functionality and morphology and the gut microbiota were evaluated.
Results
This study demonstrated that the consumption of chia flour increased the production of acetic and butyric acids, the longitudinal and circular muscle layers and crypt thickness. It also improved the expression of aminopeptidase (AP) and sucrose-isomaltase (SI) and decreased the cecum content pH. Further, the consumption of chia improved richness and decreased diversity of the microbiota. Operational Taxonomic Units (OTUs) clustering indicated difference between the ST and STC groups. In the linear discriminant analysis effect size (LEfSe) analysis, the Bacteroides genus and members of the Muribaculaceae and Lachnospiraceae families were enriched in the STC treatment group. The STC group demonstrated the enrichment of Kyoto Encyclopedia of Genes and Genomes (KEGG) metabolic pathways related to peptidoglycan and coenzyme A biosynthesis.
Conclusion
Our results suggest that chia flour, which is rich in dietary fiber and phenolic compounds, presented potential properties to improve intestinal health.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The intestine is the largest vital epithelial organ. To perform its functions, the intestinal epithelium needs to form a barrier, which depends on a well-balanced cellular homeostasis, orchestrated by an interaction and balance between differentiation, renewal, proliferation, and the intestinal ecosystem. Disruption of the balance in intestinal homeostasis is associated with a wide range of pathological changes, including metabolic disorders and inflammatory and autoimmune diseases [1].
Excessive consumption of a western diet, high in fat and low in vegetables and dietary fiber, is one of the main risk factors for various metabolic and chronic inflammatory disorders involving different organs. In addition to obesity and metabolic syndrome, many diseases that affect multiple organs are induced or worsened by the chronic consumption of a high-fat diet (HFD) [2,3,4]. Saturated fat, in addition to triggering various harmful metabolic responses to the body, is harmful to the intestine, since it increases the permeability of the organ, which leads to deleterious effects on the intestinal microbial ecosystem. These effects cause intestinal dysbiosis and impair the health status of individuals [5].
Growing interest in the dynamic role of microbiome disorders in health has shown that the gut microbiome is still linked to menopause in women [6, 7]. Estrogen and its receptors, in addition to reproductive functions, play a multifactorial role in inflammation, autoimmunity and in the physiology of the gastrointestinal tract, contributing to the prevention and/or progression of various conditions in this tissue such as inflammatory bowel diseases. At low estrogen levels (as seen after menopause) pro-inflammatory pathways are stimulated [8]. These hormonal changes are also related to greater body adiposity, which can be aggravated by the consumption of a high-fat diet [9] and be an aggravating factor in inflammation. The post-menopausal period can affect the abundance of some genera, microbial diversity, Firmicute/Bacteroidete ratio, permeability and intestinal motility in humans. Thus, a relationship between the gut microbiome and systemic health is suggested, which may be negatively affected by menopausal changes and inflammation [6, 7].
The Salvia genus, native to southern Mexico and northern Guatemala, comprises approximately 900 seed species of the Lamiaceae family, 61 of which are cultivated in Brazil. Among the species of the Salvia genus, chia (Salvia hispanica L.) is an herbaceous plant with high nutritional and functional value [10]. It is a source of protein, dietary fiber, polyunsaturated fatty acids, phenolic compounds, vitamins and minerals [11]. In relation to the content of dietary fiber, it is important to highlight that about 30% of the dietary fiber is composed of insoluble fiber and 3% of soluble fiber [11]. Moreover, chia has a substantial amount of phytochemicals, such as rosmarinic acid, rosmarinyl glucoside and ferulic acid [12], vitamin E, flavones and flavanones [11]. Chia consumption has proven antihyperlipidemic [13], anti-inflammatory [14], hypoglycemic [15], and cardioprotective in vivo activity [16]. Some studies with humans demonstrated that chia consumption can induce short-term satiety in healthy individuals [17] and have a beneficial effect on some markers of the lipid and fatty acid profile [18]. Besides, the literature reveals that the concentration of dietary fiber present in this food is able to improve intestinal morphology and health when associated with a standard diet, which indicates that chia is capable of inducing beneficial changes in the microbial composition [15].
Despite the available knowledge about the effect of chia consumption on health parameters and intestinal morphology, the effects of this food on intestinal health after menopause changes are still undetermined. We hypothesized that chia consumption may improve intestinal health parameters in animals that have undergone the characteristic changes of menopause and fed a high-fat diet. Therefore, the present study aimed to evaluate the effect of chia flour associated with a high fat diet on intestinal health in female ovariectomized Wistar rats. Our primary outcome was to assess the effects of chia consumption on parameters of measurements of intestinal health (brush border membrane functionality, microbiota composition, intestinal functionality and histomorphometric measurement) during the post menopause, the secondary outcome was analysing the biometric and intake measurements and correlation analysis.
Material and methods
Sample material
Chia seeds (Salvia hispanica L.) grown in the state of Rio Grande do Sul (Brazil) were used in the study. The seeds were ground up in three replicates, using a knife mill (Marconi Equipment, Brazil) to a particle size of 850 µm to obtain flour. Subsequently, chia flour was packed in polyethylene aluminum bags and stored in a freezer (– 18 ± 1 °C) until the time for analysis.
Animals and diets
Thirty-two 21-day-old female rats (Rattus norvegicus, Wistar, albinus variation), newly weaned, from the Central Animal Facility of the Center for Biological Sciences and Health at the Federal University of Viçosa, Minas Gerais, Brazil, were systematically subdivided into 4 groups (n = 8). Randomization of animals was performed based on the body weight of the animals, using the WinPepi Program version 11.65, to obtain the same weight average at the beginning of the study and to maintain homogeneity between groups. So that the average weight did not exceed 5 g per group. To calculate the estimated number of animals per group it was used descriptive studies, as detailed by Fontelles et al. [19]. The animals were distributed into individual stainless-steel cages, one rat per cage in a controlled temperature environment (22 °C) and automatically controlled light and dark cycles of 12 h. The animals received deionized water ad libitum and their respective experimental diets was offered in feeders and was available ad libitum to the animals.
The experimental diets were based either on the standard diet [20] or high-fat diet (Research Diets, New Brunswick, NJ), with modifications. Chia was added to the diets (STC and HFC) to replace a 100% of oil and dietary fiber (Table 1). The standard diet was composed of 19% protein, 17% fat and 64% carbohydrate. The high fat diet was prepared in the following proportions: 51% fat, 15% protein and 34% carbohydrate. The amount of chia offered in the diets was based on its composition: lipids (32.16 g.100 g−1), proteins (18.18 g 0.100 g−1), total dietary fiber (33.37 g. 100 g−1), soluble fiber (2.89 g. 100 g−1), insoluble fiber (30.47 g. 100 g−1) and carbohydrates (4.59 g. 100 g−1) [11]. The other ingredients were added in sufficient quantities to provide the planned amounts of lipids, proteins, carbohydrates, fiber and calories (Table 1).
During the first 7 weeks, the animals received standard AIN-93 M diet [20] (n = 16), or high fat diet (n = 16), according to their group. After that period, the rats (n = 32) were subjected to ovariectomy (OVX). The rats maintained their respective diets for 3 weeks to recover from surgery. At 10 weeks, the OVX animals were redistributed into 4 groups (n = 8), and each group received one of the following diets for 8 weeks: standard diet (ST); standard diet + chia (STC); high-fat diet (HF); high-fat diet + chia (HFC). Each single animal was considered an experimental unit.
Body weight and food intake were monitored weekly during the experimental period to determine the feed efficiency ratio (FER) [15]. On the 126th day, after 12 h of fasting, the animals were anesthetized with isoflurane (Isoforine, Cristália®). The cecum weight was measured, its content was collected in a sterile microtube, immediately frozen in liquid nitrogen and stored at – 80 °C for analysis. The colon segment was collected, part of which was flushed with phosphate buffered saline solution, fixed in formaline for 24 h and kept in ethanol 70% for histological analysis, while other part was stored at − 80 °C before analysis. The team was blinded during the conduct of the experiment and during euthanasia. The order of the analyzes was done randomly. The person who performed the analysis of the results was not blind, but the results were analyzed by more than one person. All the experimental procedures with animals were performed in accordance with the Directive 86/609/EEC of November 24, 1986, in compliance with the ethical principles for animal experimentation. The study protocol was approved by the Ethics Committee of the Federal University of Viçosa (Protocol 20/2017; date of approval: July 13th, 2017).
Cecum content pH
About 1 g of cecum content was homogenized in 10 mL of distilled water, with the aid of vortex glass spheres, for cecum content pH analysis. Subsequently, the glass electrode of the pHmeter (Bel, Italy) was inserted. The measurements were performed in duplicate [21].
IgA quantification
For the determination of sIgA, 1:5 (w/v) fecal homogenates were prepared according to standard procedures. The fecal samples were defrosted on ice. Suspensions were prepared by adding 200 mg feces to 800µL of phosphate-buffered saline and homogenizing with the aid of a vortex. The mucosal immunity was evaluated based on the fecal sIgA concentration, which was measured using an Immunochron enzymelinked immunosorbent assay (ELISA) [22]. Total protein in the sample homogenate was quantified by the Bradford method [23]. The results were expressed in ng of IgA per milligram of protein.
Short chain fatty acids (SCFA) content
Cecum content samples (~ 500 mg) were homogenized in 1 mL of Milli-Q water for analysis, with the aid of vortex, and centrifuged at 12,000 g, for 10 min. The supernatant was removed, and the other steps were performed as described by Siegfried, Ruckemann and Stumpf (1984) [24]. Subsequently, the samples were analyzed by high performance liquid chromatography (HPLC), using a Dionex Ultimate 3000 Dual detector HPLC apparatus (Dionex Corporation, Sunnyvale, CA, USA) equipped with a refractive index detector Shodex RI-101, using a Bio-Rad HPX-87H column, 300 × 4.6 mm maintained, at 45 °C. The analyses were performed isocratically under the following conditions: mobile phase sulfuric acid 5 mmol l–1, flow rate 0.7 ml min–1 and injection volume 20 μl. Stock solutions of the standards were prepared using the acetic, propionic and butyric acid. All SCFA were prepared with a final concentration of 10 mmol/L.
mRNA extraction from intestinal tissue and cDNA synthesis
The colon was macerated in liquid nitrogen under RNAse free conditions, and the samples were aliquoted for total RNA extraction. Total RNA was extracted with the TRIzol Reagent (Invitrogen, Carlsbad, CA, USA). The extracted mRNA was used to synthesize the cDNA with the M-MLV reverse transcription kit (Invitrogen Corp., Grand Island, NY) [25].
Determination of gene expression of proteins involved in intestinal health by quantitative reverse transcriptase polymerase chain reaction (RT-qPCR)
mRNA expression levels of genes in the intestinal tissue that are involved in intestinal health were analyzed by RT-qPCR. The SYBR Green PCR master mix from Applied Biosystems (Foster City, CA) was employed, and the analyses were performed on the StepOne™ Real-Time PCR System (Thermo Fisher Scientific) by means of the measurement system involving SYBR-Green Fluorescence and Primer Express software (Applied Biosystems, Foster City, CA). Sense and antisense primer sequences were ordered (Choma Biotechnologies) to amplify AP (ID: 301368687), SI (ID: 301368688), PepT1 (ID: 301368693) and SGLT1 (ID: 301368686). The relative expression levels of mRNA were normalized to the endogenous control (beta-actin; Table 2).
Colon histomorphometric analysis
Semi-serialized histological colon fragments, 3 μm-thick, were obtained on a semi-automated rotating microtome (Leica, Brazil) and stained by hematoxylin and eosin technique. Slides were examined under a AX70 photomicroscope (Olympus, Japan). Twenty random fields per animal were selected to measure crypt depth, crypt thickness and thickness of the circular and longitudinal muscle layers. Only crypts with definite and visible connective epithelium were used, and the images were processed using the ImagePro-Plus® software version 4.5 (Media Cybernetics, Rockville USA) [15].
DNA extraction and sequencing
Total genomic DNA was extracted from the cecum content samples, following a mechanical disruption and phenol/chloroform extraction protocol [26]. PCR amplicon libraries targeting the hypervariable V4-region of the 16S rRNA gene were produced using the primers 515F (5’GTGYCAGCMGCCGCGGTAA3’) and 806R (GGACTACNVGGGTWTCTAAT3’) and a barcoded primer set adapted for the Illumina MiSeq plataform (Illumina, San Diego, California, USA) [27, 28]. The samples were loaded onto an Illumina flow cell for paired-end sequencing reactions, using the Illumina MiSeq platform in the Environmental Sample Preparation and Sequencing Facility (ESPSF), at the Argonne National Laboratory (Lemont, Illinois, USA).
The sequences obtained for all samples in the present study were submitted to Sequence Read Archive (SRA), at the National Center for Biotechnology Information (NCBI), under the accession number PRJNA805268.
Data processing and analysis were performed using Mothur software v.1.40.0 [29]. In summary, the R1 and R2 paired-end reads were joined, and sequences smaller than 150 or greater than 300 bp were removed. Chimera sequences were detected and removed using UCHIME [30]. After cleaning, the sequences were aligned with the 16S rRNA gene, using the SILVA database v.138 [31].
The Operational Taxonomic Units (OTUs) were grouped with a 97% sequence similarity cutoff. To correct sampling bias due to unequal amplicon library sizes, the samples were normalized for the lowest number of sequences produced from any sample. Alpha diversity was estimated by using Chao1, Shannon and Simpson indices, and beta diversity was assessed by Principal Coordinate Analysis (PCoA) based on the Jaccard dissimilarity index [32]. The taxonomy was evaluated in each experimental group and the Firmicutes/Bacteroidetes ratio was calculated. The final analysis investigation of relative abundances at all taxonomic levels was carried out using the linear discriminant analysis effect size (LEfSe) method to investigate bacterial biomarkers that could represent differences in the gut microbiota of the experimental treatments. The PICRUSt2 software system was used to carry out metagenome functional predictive analysis to investigate any alterations in the metagenomic potential of the gut microbiota. The Greengenes database was 363 used to identify the 16S rRNA gene copy number to normalize feature abundance, and KEGG ortholog prediction was calculated.
Statistical analysis
The biological data were initially submitted to a Kolmogorov–Smirnov normality test. Then, a one-way analysis of variance (ANOVA) was applied, followed by the post-hoc of Newman-Keuls to compare all experimental groups to verify the effect of standard and high fat diets on intestinal health. Further, the t-test was used to compare the pairs ST × STC, and HF × HFC (the same diet with or without chia) to verify the effect of chia on intestinal health. The correlations between biological and gut microbial markers were assessed by the Pearson's correlation test. The inflammatory and stress oxidative markers have already been carried out and published [14]. The analysis was performed in Graphpad Prism version 9.0.
For the microbiome results, Chao1, Shannon and Simpson indices were used to calculate bacterial richness within the samples, while differences between the groups were analyzed by ANOVA. Differences between Beta-diversity indices were analyzed by the Pairwise PERMANOVA test. Statistically significant p values associated with microbial clades and functions were corrected for multiple comparisons using the Benjamini Hochberg false discovery rate (FDR) correction. Statistical analysis was performed using SPSS version 20.0. The level of significance of all analyses was established at p < 0.05.
Results
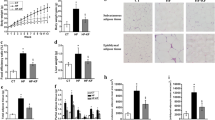
The weight gain (Fig. 1A) and feed efficiency ratio (Fig. 1C) in groups fed with high fat diet (HF and HFC) were higher (p < 0.05) than in groups fed with standard diets (ST and STC). However, the food intake was lower (p < 0.05) in animals that consumed higher amounts of fat, regardless of the presence of chia (Fig. 1B). Besides, chia consumption did not change (p > 0.05) the weight gain, food intake or feed efficiency ratio of the animals.
A Weight gain, B Food intake and C FER. ST standard diet, STC standard diet + chia, HF: high fat diet, HFC high fat diet + chia, FER feed efficiency ratio. Data expressed as mean ± standard deviation (n = 8/group). Means followed by the same lowercase letter do not differ significantly according to the Newman-Keuls test at the 5% threshold of probability. Means followed by the same capital letter do not differ from each other, by the t test at 5% probability, compared to the groups that received the same diet, either with or without chia
The cecum content pH was lower (p < 0.05) in groups fed with chia. The cecum weight and the IgA concentration did not differ (p > 0.05) among the experimental groups (Table 3). Among the organic acids analyzed, acetic acid increased (p < 0.05) in the cecal content of the STC group, compared to the ST group, and no difference (p > 0.05) was observed in the HF group in relation to the ST group. However, HFC presented lower concentration of acetic acid in relation to the STC group. Butyric acid increased in groups fed with chia (STC and HFC) in relation to the control group (ST). In addition, the comparison between ST and STC revealed that chia consumption was able to increase (p < 0.05) butyric acid, which was also observed in high fat diet groups. The propionic acid cecal concentration increased (p < 0.05) among the groups that were fed a high fat diet (HF and HFC) in relation to standard diet (ST and STC) (Table 3).
The crypt depth was lower (p < 0.05) in the STC group. Chia seed consumption increase the crypt thickness. The circular muscle layer was increased by the consumption of chia in animals that were fed a standard diet and did not differ among the high fat groups (p > 0.05). Chia increased the longitudinal muscle layer in the group fed with a standard diet. However, HFC group present a reduction (p < 0.05) on longitudinal muscle layer in comparison to the HF group, but the value did not differ in relation to STC group. Among the groups that fed a standard diet, those who fed STC presented higher longitudinal muscle layer compared to ST (Table 3).
SGLT1, AP and SI mRNA gene expression were higher in the group fed with STC. The consumption of a high fat diet (HF and HFC) reduced the SGLT1 gene expression, compared to the consumption of a standard diet (ST and STC). Chia consumption increased the mRNA gene expression of AP and SI and decreased the expression of PepT1, when consumed in a standard diet. When consumed in a high fat diet, the mRNA gene expression of SGLT1 and SI was decreased (Fig. 2).
Effect of chia consumption (standard diet and high fat diet) on the gene expression of proteins in the intestinal tissue. RT-qPCR analysis. A SGLT1 expression, B PepT1 expression, C AP expression and D SI expression. ST standard diet, STC standard diet + chia, HF high fat diet, HFC high fat diet + chia, SGLT1 sodium-glucose transport protein 1, PepT1 peptide transporter 1, AP amino peptidase, SI sucrose isomaltase. Data expressed as mean ± standard deviation (n = 8/group). Means followed by the same lowercase letter did not differ significantly according to the Newman-Keuls test, at 5% probability. Means followed by the same capital letter do not differ from each other, according to the t test, at 5% probability, in the comparison of the groups that received the same diet, either with or without chia
The 16S rRNA gene sequencing from the cecal content generated 569.661 raw sequences. After filtering and cleaning, 430.127 good quality sequences were obtained. The Good's coverage estimator was > 99% across samples, which indicates that the current sequencing depth could represent most of the bacterial community in the experimental groups (Supplementary Table).
Alpha diversity was assessed by using the Chao, Shannon, and Simpson indexes. There was no difference in the richness and diversity indexes among the experimental treatments (p > 0.05) (Fig. 3A–C). However, after comparing the pairs, considering the type of diet (ST × STC, and HF × HFC), Chao1 indicated an increased richness in the STC group, compared to the ST group (p < 0.05) (Fig. 3A), and the Simpson index revealed decreased diversity (p < 0.05) in the STC group, compared to the ST control group (Fig. 3C).
Alpha-diversity metrics of bacterial communities in the cecum faces. Data expressed as median; bars = minimum and maximum value. n = 6/group (ST, STC and HF groups), n = 5/group (HFC group). ST standard diet, STC standard diet + chia, HF high fat diet, HFC high fat diet + chia. Treatment groups indicated by the same letter are not significantly different (p < 0.05). *Indicates differences between the groups ST and STC (paired t test, p < 0.05)
Principal Coordinate Analysis (PCoA) was used to assess beta diversity. Based on the Jaccard dissimilarity index and PERMANOVA, we observed some significant results (p < 0.05) in the clustering of samples, for OTUs and genuses (Fig. 4A and C). The OTUs clustering of the four treatments indicated difference between the ST and HF groups (p ≤ 0.05), but no difference was verified between the other experimental groups (p > 0.05). Accordingly, the pairwise comparison revealed difference between the ST and STC treatments (p = 0.02), which indicates a potential effect of the chia flour on the gut microbiome taxa (Fig. 4A). Spatial ordination at phylum level (Fig. 4B) indicated no significant changes between all treatment groups and in pairwise clustering (p > 0.05). However, at genus level (Fig. 4C), we observed difference in the distance metrics among the experimental groups (p < 0.001), with significant distinction between the ST and HFC groups (p = 0.02). After a pairwise clustering according to the type of diet consumed, there was significant difference by PERMANOVA (p < 0.05). However, after the correction of the data by FDR, this difference was lost (p > 0.05).
Changes in the β-diversity of the cecal microbiome of adult Wistar rats according to each treatment group. A Principal Coordinate Analysis (PCoA) based on the Jaccard distance at OUT level. B PCoA based on Jaccard distance at the Phylum level. C PCoA based on Jaccard distance at Genus level. n = 6/group (ST, STC and HF groups), n = 5/group (HFC group). ST standard diet, STC standard diet + chia, HF high fat diet, HFC high fat diet + chia
The taxonomic classification of the samples presented 17 phyla, 25 classes, 61 orders, 100 families and 199 genera. The stratification of the phyla that comprised more than 0.2% of relative abundance, after FDR correction, is exhibited in Fig. 5A. There was no statistical difference at the level of phylum and genus when all treatment groups were compared nor after a pairwise comparison (p > 0.05). However, the Firmicutes to Bacteroidetes ratio was lower in the ST group, compared to the STC group (p < 0.05) (Fig. 5B). The sequencing revealed that the four groups were mainly dominated by the Firmicutes (76–80% of relative abundance) and Bacteroidetes (11–14% of relative abundance) phyla. At genus level (Fig. 5C), we observed a dominancy of genera from Muribaculaceae (5–7% of relative abundance) and Lachnospiraceae (3–5% of relative abundance) families, and Oscillibacter and Bacteroides (1–2% of relative abundance) (Fig. 5C), identified after the treatment period.
Relative abundances of bacterial microbiota composition at phylum and genera level of adult female Wistar rats. A Relative abundance of each identified phylum; B Firmicutes/Bacteroidetes ratio; C Genera samples displayed according to each experimental group (ST, STC, HF and HFC). Data expressed as median; bars = minimum and maximum value. n = 6/group (ST, STC and HF groups), n = 5/group (HFC group). ST standard diet, STC standard diet with chia, HF high fat diet, HFC high fat diet with chia. Only phyla with abundance > 0.2% and genera with abundance > 1% in at least one group were displayed. Data were analyzed with an FDR correction. *Indicates differences between the groups ST and STC (paired t test, p < 0.05)
The analysis performed without FDR correction revealed differences related to the abundance of Bacteroidetes (p = 0.03), which were found in large numbers in the cecal content of the animals (> 10%). In addition, Lachnospiraceae_NK4A136_group exhibited difference between the groups (p = 0.01), when the FDR correction was disregarded. However, all differences disappeared after FDR correction (p > 0.05).
The linear discriminant analysis effect size (LEfSe) showed that the Bacteroides genus and members of the Muribaculaceae and Lachnospiraceae families were significantly enriched (p < 0.05) in the STC treatment group, compared to the ST control group (Fig. 6A). In the HFC group, Faecalibaculum and Ruminococcus genera and members of the Lachnospiraceae family were significantly enriched (p < 0.05), compared to the HF control group (Fig. 6B).
LEfSe method used to compute Linear discriminant analysis (LDA) scores of the relative abundance difference between the experimental groups. A LDA scores of the relative abundance difference between the STC group and the ST control group; B Computed LDA scores of the relative abundance difference between the HFC group and the HF control group. ST standard diet, STC standard diet with chia, HF high fat diet, HFC high fat diet with chia. n = 6/group (ST, STC and HF groups), n = 5/group (HFC group)
The STC group demonstrated significant enrichment (p < 0.05) of KEGG metabolic pathways related to peptidoglycan and coenzyme A biosynthesis, compared to the ST control group. However, there was no difference (p > 0.05) in the other KEGG metabolic pathways or in the pathways between the HF and HFC groups (Fig. 7A and B).
Difference in the relative abundance of the most abundant KEGG microbial metabolic pathways in the microbiota of adult Wistar rats. A Enriched pathways between the ST and STC treatment groups; B Enriched pathways between the HF and HFC treatment groups. Extended error bar plot was performed by bioinformatic software (STAMP) with FDR correction. Welch’s two-sided test was used and Welch’s inverted was 0.95. ST standard diet, STC standard diet with chia, HF high fat diet, HFC high fat diet with chia. n = 6/group (ST, STC and HF groups), n = 5/group (HFC group). Data express in 95% confidence interval
The Pearson correlation analysis was used to assess the relationship between changes in gut microbial abundance, intestinal health parameters and inflammatory and oxidative stress markers. Changes in the Lachnospiraceae family were negatively correlated with the Muribaculaceae family (r = – 0.667) and Bacteroides (r = – 0.429), and positively correlated with butyric acid (r = 0.536) and catalase (CAT) (r = 0.469). The Muribaculaceae family was positively correlated with Ruminococcus (r = 0.638) and total antioxidant capacity of plasma (TAC) (r = 0.505), and negatively correlated with propionic acid (r = – 0.571) and superoxide dismutase (SOD) (r = – 0.487). Oscillibacter was negatively correlated with propionic acid (r = – 0.575) and positively correlated with TAC (r = 0.601). Faecalibaculum was negatively correlated with cecum content pH (r = – 0.468); Bacteroides was negatively correlated with butyric acid (r = – 0.576). Ruminococcus was negatively correlated with tumor necrosis factor alpha (TNF-α) (r = – 0.543) and SOD (r = – 0.530) and positively correlated with TAC (r = 0.520). Acetic acid was positively correlated with butyric acid (r = 0.559); propionic acid was negatively correlated with TAC (r = – 0.743) and positively correlated with SOD (r = 0.568) and thiobarbituric acid reactive substances (TBARS) (r = 0.619); butyric acid was negatively correlated with cecum content pH (r = – 0.649) and positively correlated with IgA (r = 0.577). Cecum content pH was negatively correlated with CAT (r = – 0.546). TNF was positively correlated with Il-1β (r = 0.6705), SOD (r = 0.493) and TBARS (r = 0.546). Il-1β was negatively correlated with TAC (r = – 0.457) and positively correlated with TBARS (r = 0.657). TAC was negatively correlated with TBARS (r = – 0.395); and SOD was positively correlated with TBARS (r = 0.794). (Fig. 8).
Heatmap of Pearson correlation analysis. IgA immunoglobulin, TNF-α tumor necrosis factor alpha, Il-1β Interleukin 1β, TAC_Plasma total antioxidant capacity of plasma, CAT catalase, SOD superoxide dismutase, TBARS thiobarbituric acid reactive substances. *Indicates statistically significant difference (p < 0.05). n = 6/group (ST, STC and HF groups), n = 5/group (HFC group)
Discussion
Few studies in the literature have evaluated the impact of chia consumption on intestinal parameters. However, due to chia chemical composition, mainly the amount of dietary fiber, polyunsaturated fatty acids, phenolic compounds and other bioactive compounds, it is suggested that this food may have a positive effect on the microbiota and intestinal health. Thus, the present study demonstrated that the consumption of chia flour associated with standard diet increases of acetic and butyric acids content, decreases the cecum pH, increases longitudinal muscle layer, circular muscle layer and crypt thickness and improves the mRNA gene expression of AP and SI. In addition, chia flour promotes some compositional changes in the gut microbiota.
The groups fed with high fat diet presented lower food intake (in grams per week) and higher weight gain, which corroborates previous reports that the higher amounts of fat and so, calories, can promote greater satiety and reduce consumption, while increasing the accumulation of body fat [14]. On the other hand, chia consumption improved some intestinal parameters, such as cecum content pH, increased the production of short-chain fatty acids and the expression of intestinal functionality biomarkers, which could be attributed to the higher amount of dietary fibers present in the food. The dietary fibers present in chia seeds are composed of soluble and insoluble fractions and can benefit intestinal health, since they are able to be fermented by colonic bacteria, thus producing short-chain fatty acids and consequently reducing intestinal pH, which may increase mineral solubility and therefore absorption [12]. In addition, the acidification of the intracolonic environment is considered important for the suppression of the production and the activity of a range of bacterial metabolites implicated in colonic diseases [33].
In this study, an increase was observed in crypt thickness, longitudinal muscle layer and circular muscle layer in the groups fed with chia in a standard diet. In Gallus gallus feeding, the digestive and absorptive capabilities of the brush border membrane may be directly related with morphometric parameters, such as villi height, crypt depth and the ratio between villi height and crypt depth [34]. These functional and morphological effects can be explained by the potential increased proliferation of intestinal cells due to the increased motility of the digestive tract by the soluble fiber, which leads to hyperplasia and/or hypertrophy of intestinal muscle cells [15].
The gene expressions of aminopeptidase (AP), sodium-glucose transport protein 1 (SGLT1), sucrase isomaltase (SI) and peptide transporter 1 (PepT1) are used as biomarkers of brush border membrane (BBM) digestive and absorptive functions [12]. In the current study, the expression of BBM functional genes AP and SI was upregulated by chia consumption (STC) compared to the standard diet group (ST), and chia consumption (HFC) did not affect the gene expression of AP and PepT1 and decreased the expression of SGLT1 and SI in HFC group relative to the HF group. Furthermore, we observed an increase in longitudinal muscle layer and in crypt thickness in STC group compared to ST group. Thus, in diet normal conditions chia increased the intestinal BBM functionality, and morphology; however, it not happened in HFD condition. These results are benefic since AP is an exopeptidase that cleaves amino acids from the N-terminus of peptides, and SI is the major disaccharidase located on the duodenal BBM, which hydrolyzes disaccharides or oligosaccharides into monosaccharides for absorption [34]. Others studies observed that the intra-amniotic administration of different concentrations of chia soluble extract (5, 10, 25 and 50 mg/mL) [12], and black, carioca and white bean extracts (50 mg/mL) [35] did not affect the BBM functional gene expression. In the study of Wang et al. [36] the wheat bran extract showed significant increases in the villus heights, goblet cell diameter and numbers. Thus, chia compared to other types of dietary fibers is a good source of this nutrient, which can improve the intestinal health.
Phytochemicals have been descripted as agents that exert effects similar to prebiotics. Phenolics compounds have been shown to be positive modulators of intestinal microbiota [37, 38]. This interaction entails microbial degradation of polyphenols and modulation of gut microbiota by polyphenols and their metabolites, which inhibits pathogenic bacteria and stimulates beneficial bacteria [39]. Chia phenolics demonstrated beneficial effects in gene expression modulation of gluconeogenic and glycolytic enzymes [40] and antioxidant and anti-inflammatory activity [37, 38]. Other important nutrient present in chia is alpha linolenic acid [11, 40, 41]. The alpha linolenic acid exerts effects on the intestinal microbiota, the host-microbiome interaction, and interactions between the host immune system and gut microbiota. Accordingly, the gut microbiota modulates the absorption and metabolism of omega-3 PUFAs and directly or indirectly modulates subsequent physiological and immune responses in the host [42]. Diets rich in alpha linolenic acid reduced the colonic abundance of potentially pathogenic bacteria and increased bacteria with beneficial potential. This eventually inhibits the host inflammatory responses [43]. Further, brown and golden flaxseeds with high content of insoluble dietary fiber and alpha-linolenic acid demonstrated reduced intestinal permeability and endotoxemia in perimenopausal overweight women [44]. Thereby, the combination of dietary fiber, lipids, and polyphenol from chia in our study may explain some beneficial effects of chia on intestinal health.
In this study, the current sequencing depth could represent most of the bacterial community in the experimental groups. In alpha diversity, we found that the consumption of chia with a standard diet (STC × ST) increased the Chao index, improved richness, raised the number of different species and decreased the Simpson index, which reduced the dominance and indicated a greater number of different individuals in the community. Beta diversity analysis by the OTU’s clustering showed difference between the ST and HF groups and between the ST and STC groups, which indicates a potential effect of the chia flour on the gut microbiome taxa.
Both the menopausal change and obesity were noted to enhance dysbiosis by reducing microbiome diversity and increasing the Firmicutes to Bacteroidetes ratio [6, 7]. This ratio has been used as a marker for intestinal homeostasis in animal studies [45, 46]. In our study, no difference was observed at the level of phylum and genus, but the Firmicutes to Bacteroidetes ratio was lower in the ST group, compared to the STC group. Findings related to the alterations in this ratio are still inconsistent, which highlights the complexity of the gut microbial ecosystem. The Firmicutes to Bacteroidetes ratio does not necessarily reflect a common pattern of all genera belonging to these phyla, since multiple genera of the same phylum can be found in higher or lower proportions [47]. Although we did not detect significant modifications in the taxonomy of the cecal microbiota, upregulation in the SCFA-producing bacteria in the group that fed chia was observed through LEfSe analysis.
LEfSe determines the features (organisms, clades, operational taxonomic units, genes or functions) most likely to explain differences between classes. Our LEfSe analysis suggested that Bacteroides genus and members of the Muribaculaceae and Lachnospiraceae families were the main bacteria that caused the difference in the structure of intestinal microbiota in the group fed with standard diet (STC × ST). Bacteroides genus includes species that produce acetate and propionate and promote goblet cell differentiation and the expression of mucin-related genes [48]. The Muribaculaceae family can degrade dietary carbohydrates and ferment polysaccharides into SCFAs [49,50,51]. The family Lachnospiraceae is also a SCFA producer, and its increased relative abundance is associated with increased production of SCFAs [52]. In the group HFC, the organisms most likely to explain differences, besides the Lachnospiraceae family, were Faecalibaculum and Ruminococcus genera, both producers of butyrate [48, 53].
The elevation in SCFA-producing bacterial populations justifies the increased SCFA concentration in the lumen. These SCFAs may lead to intestinal cellular proliferation, and this connection could explain the increased crypt thickness in chia fed group. Then, by favoring bacterial fermentation, chia seed affected the intestinal luminal pH [51]. In agreement with the Pearson correlation analysis we performed, the Lachnospiraceae family was positively correlated with butyric acid, and the concentration of butyric acid was negatively correlated with the pH of the cecal content. The study conducted by Guo et al., 2022 [52] demonstrated that inulin supplementation has beneficial effects on the intestinal microbiota, with an increased abundance of butyrate-producing bacteria from the Ruminococcaceae and Lachnospiraceae families, thus leading to increased production of SCFAs, which suggests that the beneficial effects of inulin on glucose and lipid metabolism can be partially attributed to the modulation of the gut microbiota and the signaling of SCFAs.
The gut microbiome data demonstrates how the specific microbial profile within a treatment group is linked to the physiological status of the host. The results reveal that chia consumption with a standard diet upregulated the pathways related to peptidoglycan and coenzyme A biosynthesis. As a limitation of our study, we can highlight the absence of a group without ovariectomy (SHAM group—sham surgery), so we could not attribute effects to ovariectomy compared to non-ovariectomized animals on intestinal health.
Conclusions
The consumption of chia by female ovariectomized Wistar rats positively affected gut health by increasing the production of short chain fatty acids, decreasing the pH of cecum content, improving the intestinal functionality and morphology, and promoting some compositional changes in the gut microbiota, mainly with the consumption of a standard diet. Thus, it was observed that chia, which is rich in fiber and phenolic compounds, demonstrated potential properties to improve intestinal health. Further studies are still needed to clarify the effects of chia on the intestinal health and microbiome and to make clearer what can be the implications to human population, whereas there is a just a few studies evaluating chia consumption and impacts on intestinal human health.
References
Sun X, Zhu M-J (2017) AMP-activated protein kinase: a herapeutic target in intestinal diseases. Open Biol. https://doi.org/10.1098/rsob.170104
Tessitore A, Mastroiaco V, Vetuschi A et al (2017) Development of hepatocellular cancer induced by long term low fat-high carbohydrate diet in a NAFLD/NASH mouse model. Oncotarget 8:53482–53494. https://doi.org/10.18632/oncotarget.18585
Wali JA, Jarzebska N, Raubenheimer D et al (2020) Cardio-metabolic effects of high-fat diets and their underlying mechanisms — a narrative review. Nutrients 12:1–18
Sferra R, Pompili S, Cappariello A et al (2021) Prolonged chronic consumption of a high fat with sucrose diet alters the morphology of the small intestine. Int J Mol Sci 22:7280. https://doi.org/10.3390/ijms22147280
Abulizi N, Quin C, Brown K et al (2019) Gut mucosal proteins and bacteriome are shaped by the saturation index of dietary lipids. Nutrients 11:1–24. https://doi.org/10.3390/nu11020418
Shieh A, Epeldegui M, Karlamangla AS, Greendale GA (2020) Gut permeability, inflammation, and bone density across the menopause transition. JCI Insight 5:1–12. https://doi.org/10.1172/jci.insight.134092
Schreurs MPH, de Romano PJ, V van SA, Werner HMJ, (2021) How the gut microbiome links to menopause and obesity, with possible implications for endometrial cancer development. J Clin Med. https://doi.org/10.3390/jcm10132916
Garcia-Villatoro EL, Allred CD (2021) Estrogen receptor actions in colitis. Essays Biochem 65:1003–1013. https://doi.org/10.1042/EBC20210010
Mishima MDV, Pereira B, Lopes RC et al (2020) Bioavailability of calcium from chia (Salvia hispanica L.) in ovariectomized rats fed a high fat diet. J Am Coll Nutr. https://doi.org/10.1080/07315724.2020.1790441
Kulczynski B, Kobus-Cisowska J, Taczanowski M et al (2019) The chemical composition and nutritional value of chia seeds – current state of knowledge. Nutrients 115:1–16
Da Silva BP, Anunciação PC, da Silva Matyelka JC et al (2017) Chemical composition of Brazilian chia seeds grown in different places. Food Chem 221:1709–1716. https://doi.org/10.1016/j.foodchem.2016.10.115
Da Silva BP, Kolba N, Martino HSD et al (2019) Soluble extracts from chia seed (Salvia hispanica L.) affect brush border membrane functionality, morphology and intestinal bacterial populations in vivo (Gallus gallus). Nutrients. https://doi.org/10.3390/nu11102457
Da Silva BP, Toledo RCL, Grancieri M et al (2019) Effects of chia (Salvia hispanica L.) on calcium bioavailability and inflammation in Wistar rats. Food Res Int 116:592–599. https://doi.org/10.1016/j.foodres.2018.08.078
da Silva BP, Toledo RCL, Mishima MDV et al (2019) Effects of chia (Salvia hispanica L.) on oxidative stress and inflammation in ovariectomized adult female Wistar rats. Food Funct. https://doi.org/10.1039/C9FO00862D
da Silva BP, Dias DM, de Castro Moreira ME et al (2016) Chia seed shows good protein quality, hypoglycemic effect and improves the lipid profile and liver and intestinal morphology of Wistar rats. Plant Foods Hum Nutr 71:225–230. https://doi.org/10.1007/s11130-016-0543-8
Mishima MDV, Ladeira LCM, da Silva BP et al (2021) Cardioprotective action of chia (Salvia hispanica L.) in ovariectomized rats fed a high fat diet. Food Funct 12:3069–3082. https://doi.org/10.1039/d0fo03206a
Ayaz A, Akyol A, Inan-eroglu E et al (2017) Chia seeds added yogurt reduces short-term and increased satiety. Nutr Res Pract 11:412–418
Silva LDA, Verneque BJF, Mota APL, Duarte CK (2021) Chia seed (Salvia hispanica L.) consumption and lipid profile: a systematic review and meta-analysis. Food Funct 12:8835–8849. https://doi.org/10.1039/d1fo01287h
Fontelles MJ, Simões MG, Almeida JCFR (2010) Metodologia da pesquisa: diretrizes para o cálculo do tamanho da amostra. Rev Paran Med 24:57–64
Reeves PG, Nielsen FH, Fahey GC (1993) AIN-93 purified diets for laboratory rodents: final report of the American institute of nutrition ad hoc writing committee on the reformulation of the AIN-76A Rodent diet. J Nutr 123:1939–1951. https://doi.org/10.1093/jn/123.11.1939
Grancieri M, Costa NMB, das Graças Vaz Tostes M et al (2017) Yacon flour (Smallanthus sonchifolius) attenuates intestinal morbidity in rats with colon cancer. J Funct Foods 37:666–675. https://doi.org/10.1016/j.jff.2017.08.039
da Graças Vaz Tostes M, Viana ML, Grancieri M et al (2014) Yacon effects in immune response and nutritional status of iron and zinc in preschool children. Nutrition 30:666–672. https://doi.org/10.1016/j.nut.2013.10.016
Bradford MM (1976) A rapid and sensitive method for the quantitation microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Siegfried VR, Ruckemmann H, Stumpf G (1984) Method for the determination of organic acids in silage by high performance liquid chromatography. Landwirtsch Forsch 37:298–304
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Stevenson DM, Weimer PJ (2007) Dominance of Prevotella and low abundance of classical ruminal bacterial species in the bovine rumen revealed by relative quantification real-time PCR. Appl Microbiol Biotechnol 75:165–174. https://doi.org/10.1007/s00253-006-0802-y
Caporaso JG, Lauber CL, Walters WA et al (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci U S A 108:4516–4522. https://doi.org/10.1073/pnas.1000080107
Caporaso JG, Lauber CL, Walters WA et al (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624. https://doi.org/10.1038/ismej.2012.8
Schloss PD, Westcott SL, Ryabin T et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. https://doi.org/10.1128/AEM.01541-09
Edgar RC, Haas BJ, Clemente JC et al (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Quast C, Pruesse E, Yilmaz P et al (2013) The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res 41:590–596. https://doi.org/10.1093/nar/gks1219
Lozupone C, Knight R (2005) UniFrac: A new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71:8228–8235. https://doi.org/10.1128/AEM.71.12.8228-8235.2005
Tousen Y, Matsumoto Y, Nagahata Y et al (2019) Resistant starch attenuates bone loss in ovariectomised mice by regulating the intestinal microbiota and bone-marrow inflammation. Nutrients. https://doi.org/10.3390/nu11020297
Hou T, Kolba N, Glahn RP, Tako E (2017) Intra-amniotic administration (Gallus gallus) of Cicer arietinum and Lens culinaris prebiotics extracts and duck egg white peptides affects calcium status and intestinal functionality. Nutrients 9:975. https://doi.org/10.3390/nu9070785
Dias DM, Kolba N, Hart JJ et al (2019) Soluble extracts from carioca beans (Phaseolus vulgaris L.) affect the gut microbiota and iron related brush border membrane protein expression in vivo (Gallus gallus). Food Res Int 123:172–180. https://doi.org/10.1016/j.foodres.2019.04.060
Wang X, Kolba N, Liang J, Tako E (2019) Alterations in gut microflora populations and brush border functionality following intra-amniotic administration (Gallus gallus) of wheat bran prebiotic extracts. Food Funct 10:4834–4843. https://doi.org/10.1039/c9fo00836e
Plamada D, Vodnar DC (2022) Polyphenols—gut microbiota interrelationship: a transition to a new generation of prebiotics. Nutrients 14:1–27. https://doi.org/10.3390/nu14010137
Santos D, Frota EG, Vargas BK et al (2022) What is the role of phenolic compounds of yerba mate (Ilex paraguariensis) in gut microbiota? Phytochemistry 203:113341. https://doi.org/10.1016/j.phytochem.2022.113341
Corrêa TAF, Rogero MM, Hassimotto NMA, Lajolo FM (2019) The two-way polyphenols-microbiota interactions and their effects on obesity and related metabolic diseases. Front Nutr 6:1–15. https://doi.org/10.3389/fnut.2019.00188
Enes BN, de Paula Dias Moreira L, Toledo RCL et al (2020) Effect of different fractions of chia (Salvia hispanica L.) on glucose metabolism, in vivo and in vitro. J Funct Foods 71:157–164. https://doi.org/10.1016/j.jff.2020.104026
de Paula Dias Moreira L, Enes BN, de São José VPB et al (2022) Chia (Salvia hispanica L.) flour and oil ameliorate metabolic disorders in the liver of rats fed a high-fat and high fructose diet. Foods. https://doi.org/10.3390/foods11030285
Fu Y, Wang Y, Gao H et al (2021) Associations among dietary omega-3 polyunsaturated fatty acids, the gut microbiota, and intestinal immunity. Mediators Inflamm. https://doi.org/10.1155/2021/8879227
Kumar M, Pal N, Sharma P et al (2022) Omega-3 fatty acids and their interaction with the gut microbiome in the prevention and amelioration of type-2 diabetes. Nutrients. https://doi.org/10.3390/nu14091723
Sant’ Ana CT, de Amorim AD, Gava AP et al (2022) Brown and golden flaxseed reduce intestinal permeability and endotoxemia, and improve the lipid profile in perimenopausal overweight women. Int J Food Sci Nutr 73:829–840
Rinninella E, Raoul P, Cintoni M et al (2019) What is the healthy gut microbiota composition? A changing ecosystem across age, environment, diet, and diseases. Microorganisms. https://doi.org/10.3390/microorganisms7010014
Abokor AA, McDaniel GH, Golonka RM et al (2021) Immunoglobulin A, an active liaison for host-microbiota homeostasis. Microorganisms 9:1–31. https://doi.org/10.3390/microorganisms9102117
Pinart M, Dötsch A, Schlicht K et al (2022) Gut microbiome composition in obese and non-obese persons: a systematic review and meta-analysis. Nutrients 14:1–41
Makki K, Deehan EC, Walter J, Bäckhed F (2018) The impact of dietary fiber on gut microbiota in host health and disease. Cell Host Microbe 23:705–715. https://doi.org/10.1016/j.chom.2018.05.012
Lagkouvardos I, Lesker TR, Hitch TCA et al (2019) Sequence and cultivation study of Muribaculaceae reveals novel species, host preference, and functional potential of this yet undescribed family. Microbiome 7:1–15. https://doi.org/10.1186/s40168-019-0637-2
Smith BJ, Miller RA, Ericsson AC et al (2019) Changes in the gut microbiome and fermentation products concurrent with enhanced longevity in acarbose-treated mice. BMC Microbiol 19:1–16. https://doi.org/10.1186/s12866-019-1494-7
Gomes MJC, Kolba N, Agarwal N et al (2021) Modifications in the intestinal functionality, morphology and microbiome following intra-amniotic administration (Gallus gallus) of grape (Vitis vinifera) stilbenes (resveratrol and pterostilbene). Nutrients. https://doi.org/10.3390/nu13093247
Guo J, Zhang M, Wang H et al (2022) Gut microbiota and short chain fatty acids partially mediate the beneficial effects of inulin on metabolic disorders in obese ob/ob mice. J Food Biochem. https://doi.org/10.1111/jfbc.14063
de SG Olivares P, Pacheco ABF, Aranha LN et al (2021) Gut microbiota of adults with different metabolic phenotypes. Nutrition. https://doi.org/10.1016/j.nut.2021.111293
Acknowledgements
The authors are thankful to the Foundation for Research Support of Minas Gerais (FAPEMIG, Brazil, number: APQ-02183-17), for the financial support for the research; we are also grateful to the Coordination for the Improvement of Higher Education Personnel (CAPES, Brazil, grant number 88887.599144/2021-00), the National Counsel of Technological and Scientific Development (CNPq, Brazil, number: 406517/2018-5, Research Productivity’s fellowships [PQ2—grant number 310910/2020-0]), and Foundation for Research an Innovation Support of Espírito Santo (FAPES, Brazil, PRONEX—CNPq/FAPES, Public Notice 24/2018—TO 567/2018).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mishima, M.D.V., da Silva, B.P., Gomes, M.J.C. et al. Effect of chia flour associated with high fat diet on intestinal health in female ovariectomized Wistar rats. Eur J Nutr 62, 905–919 (2023). https://doi.org/10.1007/s00394-022-03043-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00394-022-03043-2