Abstract
Purpose
Irritable bowel syndrome (IBS) is a functional bowel disorder. This study aimed to assess the effect of a probiotic product (containing Lactobacillus casei Zhang, Lactobacillus plantarum P-8, and Bifdobacterium animalis subsp. lactis V9) as an adjunct to a routine regimen in IBS management.
Methods
Forty-five patients with IBS were randomized into the probiotic (n = 24) and control (n = 21) groups, receiving the routine regimen with or without probiotics for 28 days, respectively. Serum and fecal samples were collected and analyzed.
Results
The IBS-symptom severity score (P < 0.01), serum levels of IL-6 (P < 0.01) and TNF-α (P < 0.001) were significantly lower in the probiotic group than the control group at day 28. The probiotic adjunctive treatment resulted in significant decreases in some bacterial genera that worsen IBS, such as Bacteroides (P < 0.01), Escherichia (P < 0.05), and Citrobacter (P < 0.05), significant decreases were also observed in some beneficial genera in the control group, including Bifidobacterium (P < 0.05), Eubacterium (P < 0.05), Dorea (P < 0.01), and Butyricicoccus (P < 0.05). Furthermore, significant correlations were found between some monitored parameters and compositional changes in the fecal microbiota, suggesting that the clinical improvement of IBS was likely associated with gut microbiota modulation. The enterotype analysis revealed that the initial fecal microbiota composition could influence clinical outcomes.
Conclusions
The adjunctive use of probiotics with a routine regimen showed additional clinical effectiveness compared to the routine regimen alone in managing IBS. A pretreatment gut microbiome analysis might help tailor a personalized probiotic regimen to optimize treatment effects.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Irritable bowel syndrome (IBS) is a functional bowel disorder that affects 10–25% of the world’s population [1]. The disorder imposes a huge economic burden on healthcare systems and severely compromises patients' quality of life [2].
The etiology of IBS is not completely understood; however, some recent clinical evidence has revealed a link between IBS and gut dysbiosis [3]. The gut microbiomes in patients with IBS differ from those in healthy individuals. Although there have been conflicting data on the gut microbial composition of IBS patients [4], a significantly higher Firmicutes to Bacteroidetes ratio has been consistently reported in patients with IBS [5]. Small intestinal bacteria overgrowth (SIBO) is a potential underlying cause of IBS, as its reversion ameliorates IBS-related symptoms [6]. Another study found that the severity of IBS was negatively associated with microbial richness, an abundance of methanogens, and enterotypes enriched with Clostridiales or Prevotella species [7]. Moreover, some commonly employed IBS management strategies, such as rifaximin, medical food serum-derived bovine immunoglobulins, prebiotics, probiotics, and administration of low fermentable oligosaccharides, disaccharides, monosaccharides, and polyols (FODMAP) diet, would induce variable changes in patients' gut microbiota, intestinal epithelial integrity, and/or colonic environment [8]. Thus, gut microbiota modulation has been proposed as a strategy for IBS management [9].
Probiotics are increasingly used as food ingredients, dietary supplements, and dairy starter cultures. Probiotic administration has been reported to be beneficial for patients with IBS [10, 11]. Additionally, the low cost, potential high efficacy, and low risk of probiotic intake make it a practical and attractive approach for IBS management. However, contrasting effects have been reported, which may relate to the specificity of probiotic strain and variation in host factors [12]. Since both the prevalence and clinical presentation of IBS vary greatly between race and ethnicity, differences in gut microbiota between individuals might also play a role in IBS [13]. Although gut dysbiosis is a possible mechanism that drives IBS development, many previous works have only focused on studying the clinical outcome but not treatment-specific gut microbiota modulation. Thus, extensive clinical research is still required to clarify the role of gut microbiota in IBS development and to design personalized treatment for patients.
This study aimed to assess the effect of adjunctive use of Probio-Fit® with the routine regimen in treating Chinese patients suffering from IBS. Probio-Fit® is a multi-strain probiotic formulation composed of Lactobacillus (L.) casei Zhang, L. plantarum P-8, and Bifidobacterium (B.) animalis subsp. lactis V9, which is an excellent probiotic product supported by previous studies [14, 15]. The primary outcomes were measured by changes in the IBS-symptom severity score (IBS-SSS), IBS-quality of life (IBS-QoL) score, and five selected serum inflammatory markers. Since this study hypothesized that gut microbiota modulation was related to the improvement of IBS symptoms, the single-molecule real-time (SMRT) technology was used to sequence full-length 16S rRNA genes to describe changes in patients’ fecal microbiota at the species level as the secondary outcome.
Materials and methods
Ethics statement
This study was approved by the Ethical Committee of the Affiliated Hospital of Inner Mongolia Medical University. All participants provided written informed consents before the trial started. The Ethics Committee of Inner Mongolia Agricultural University and the Navy General Hospital of PLA approved the procedures and protocols. The trial was registered in the Chinese Clinical Trial Registry (Identifier number: ChiCTR2000035339).
Subjects
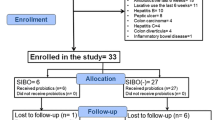
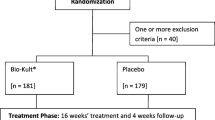
The current clinical trial was a sub-study under a clinical trial organized by the Affiliated Hospital of Inner Mongolia Medical University, and it was performed at the Navy General Hospital of PLA, Beijing. All recruited patients (aged = 37.0 ± 15.1 years; weight = 64.93 ± 11.93 kg; male: female = 10:14 in the probiotic group, male: female = 13:11 in the control group, Supplementary Table 1) fulfilled the Rome III diagnostic standard for IBS (entailing chronic abdominal pain or discomfort at least 3 days/month), and they were randomly assigned to probiotic (n = 24) and control (n = 24) groups, respectively. All patients were assessed for hematuria, liver and kidney dysfunction. Routine abdominal ultrasound, chest radiographs, and colonoscopy were also performed. Patients suffering from other gastrointestinal diseases, obesity (BMI ≥ 30), autoimmune diseases, cardiac and renal insufficiency, malignant tumors, diabetes mellitus, or having previous abdominal surgery history were excluded. Moreover, pregnant/lactating women and participants who had taken medications (e.g., antibiotics, traditional Chinese medicine, or treatments that could affect the interpretation of results) within 4 weeks prior to this trial were excluded.
Trial design
All patients kept their normal diet and living habits during the intervention period, but they were requested to avoid from taking foods that could cause allergic reactions and gas-production, including spicy, greasy, and irritating diets. All patients received standard therapy at the beginning of the trial regardless of their symptoms. The control group received routine regimen including lactulose oral solution (3 × 15 ml/day) and antidiarrheal drug (3 × 3 g of montmorillonite powder/day). Three daily oral doses (0.1 g each time) of trimebutine maleate were given to patients with abdominal pain and discomfort, while the sedative drug flupentixol (3 × 0.1 g/day) was prescribed to patients with anxiety and insomnia when required. The probiotic group received both the regular regimen and two grams of probiotic powder. The probiotic powder used in this study was Probio-Fit®, which was packed in individually sealed sachets (Beijing Scitop Bio-tech Co., Ltd., Beijing, China), consisting of three different bacterial strains (namely L. casei Zhang 3 × 109 CFU/g; B. animalis subsp. lactis V9, 4 × 109 CFU/g; and L. plantarum P-8, 3 × 109 CFU/g). Patients took one sachet per day after a meal; the content of the sachet was dissolved in warm water or milk before consumption. The treatments continued for 28 days. The primary outcome was a clinical improvement, and the secondary outcome was the changes in fecal microbiota structure and composition. Serum samples (days 0 and 28) were collected at the hospital. Fecal samples were collected at home by the participants (days 0, 7, and 28); collected fecal samples were temporarily stored in a domestic freezer and were transported to the laboratory in an ice box at 4 °C within 24 h. All samples were stored at − 80 °C until analysis.
Genomic DNA extraction, polymerase chain reaction (PCR), SMRT sequencing
Fecal DNA was extracted with the QIAGEN DNA Stool Mini-Kit (QIAGEN, Hilden, Germany) following the manufacturers’ instructions. The quality of the extracted DNA was assured by agarose gel electrophoresis and spectrophotometry (final DNA concentration > 100 ng/μL; 260 nm/280 nm ratio between 1.8 and 2.0).
The full-length 16S rRNA genes were amplified from the extracted genomic DNA by PCR using the forward 27F (5′-AGAGTTTGATCMTGGCTCAG-3′) and reverse 1492R (5′-ACCTTGTTACGACTT-3′) primers, incorporating with a set of 16-base barcodes for each DNA sample [16]. The PCR program: 95 °C for 4 min; 30 cycles of 95 °C for 30 s, 58 °C for 30 s, and 72 °C for 30 s with a final extension at 72 °C for 5 min [17].
For SMRT sequencing, DNA libraries were constructed with the barcoded-amplicons using the Pacific Biosciences SMRT bell™ template prep kit 1.0 [16]. The sequencing reaction was achieved with P6-C4 chemistry on a PacBio RS II instrument following the manufacturers' guidelines (Pacific BioSciences of California, Inc., USA). Raw sequences were deposited to the MG-RAST database (project number mgp92156).
Gut microbiota bioinformatics analysis
The protocol RS_ReadsOfinsert.1 of the SMRT Portal (version 2.3) was used for sequence extraction from the raw data. Raw sequences were restrictively filtered with four criteria: (i) minimum full passes of up to 5; (ii) minimum predicted accuracy of 90; (iii) minimum insert read length of 1400; and (iv) maximum insert read length of 1800. The filtered sequences were then barcode-sorted into different samples, followed by trimming of barcode and primer sequences.
The remaining high-quality sequences were then analyzed by the Quantitative Insights Into Microbial Ecology (QIIME) package (version 1.7) [18]. Briefly, the most abundant sequence from each cluster was chosen as the representative sequence to be aligned by PyNAST [19] and UCLUST [20] under 100% clustering of sequence identity. The representative and unique sequences were classified into operational taxonomic units (OTUs) under the threshold of 97% identity by UCLUST. Chimeric sequences were removed with ChimeraSlayer [21]. The remaining OTUs were assigned to the lowest taxonomic level using the Ribosomal Database Project II (Release 11.5) and the Greengenes database (version 13.8) [22, 23] at a minimum bootstrap threshold of 80% [24]. The de novo taxonomic tree constructed by the representative chimera-checked OTU set using FastTree [25] was used to assess the alpha- and beta-diversity.
To compare the alpha-diversity between samples, the sampling depth was normalized by the multiple_rarefactions.py program in the QIIME pipeline. The alpha-diversity and sequencing depth were evaluated with the Shannon–Wiener index and Chao 1 index. The beta-diversity and microbiota community structure were assessed by principle coordinate analysis (PCoA) of the weighted UniFrac distances derived from the phylogenetic tree [26]. Permutational multivariate analysis (PERMANOVA) was used to evaluate differences between groups.
Measurement of serum levels of cytokines, d-lactate, and lipopolysaccharides (LPS)
The serum interleukin (IL)-6, IL-8, tumor necrosis factor (TNF)-α, d-lactate, and LPS levels were determined at days 0 and 28 by enzyme-linked immunosorbent assays (Beijing Yiming Renaissance Technology Co., Ltd., China) with replicate measurements.
Assessment of clinical outcomes
The clinical outcomes were assessed by IBS-SSS [27] and IBS-QoL score [28]. The overall IBS-SSS ranged from 0 to 500, with a higher score representing worse conditions. Scoring < 175, 175–300, and > 300 represented mild, moderate, and severe IBS, respectively [27]. The IBS-QoL scale ranged from 0 to 100, and a higher score represented a better quality of life [28].
Statistical analysis
The statistical power analysis was calculated with PASS11, a routine regimen effect was estimated at 55–65%, and the probiotics as adjunctive routine regimen effect was estimated to be 90–95%. The sample size was estimated to be approximately 20–24 participants per group with a power of 0.80 and an alpha error of 0.05. Statistical analyses were performed using R packages (http://www.r-project.org/). Mann–Whitney tests were performed to evaluate intragroup differences in the levels of serum components, IBS-QoL score, and IBS-SSS between day 0 and day 28, as well as intergroup differences in these parameters at day 0 and day 28, respectively. Mann–Whitney tests were also applied to evaluate intragroup differences in the relative abundances of different taxa between the two-time points, as well as the intergroup differences at the same time point. P values < 0.05 were considered statistically significant [29]. Correlations between gut mucosal bacteria and the monitored parameters were calculated using the Spearman's rank correlation coefficient in R package. The principal component analysis for enterotypes was determined based on similarities of 16S rRNA-based phylogenetic profiles [17].
Results
Changes in serum and clinical parameters
A total of forty-eight patients with IBS met the inclusion criteria (age = 37.0 ± 15.1 years; weight = 64.93 ± 11.93 kg; BMI < 30), and they were randomly assigned to the probiotic (n = 24) and the control (n = 24) groups, respectively. However, as three patients from the control group failed to provide the last stool samples, only 21 control subjects finished the trial. No obvious organic, pathological changes and adverse reactions were identified in all participated patients during the trial. Changes in clinical and serum parameters are shown in Fig. 1. At day 28, the IBS-QoL score increased while the IBS-SSS decreased for both groups. However, the changes in IBS-QoL score and IBS-SSS of the probiotic group was greater than the control group over time; and at day 28, the IBS-SSS was significantly lower in the probiotic group than the control group (P < 0.01). Significant decreases were also observed in the serum levels of IL-8, TNF-α, and LPS for both groups from day 0 to day 28 (P < 0.0001). On the other hand, significant decreases in the levels of IL-6 (P < 0.0001) and d-lactate (P < 0.0001) were only seen in the probiotic group but not the control group. Moreover, the serum levels of IL-6 (P < 0.01) and TNF-α (P < 0.001) of the probiotic group were significantly lower than the control group on day 28.
Effects of treatments on clinical and serum parameters. Serum levels of a interleukin-8, b interleukin-6, c tumor necrosis factor-α, d d-lactate, e lipopolysacchride, f IBS-Quality of Life score, and g IBS-Symptom Severity Score. The horizontal line of the boxplot represents the mean value, while the whisker represents standard error. Error bars in the line charts represent standard error. **P < 0.01, ***P < 0.001, and ****P < 0.0001; Mann–Whitney test
Changes in alpha-diversity and structure of fecal microbiota during treatment
The diversity and richness of the gut microbiota of the patients were evaluated by the Shannon index and Chao1 index, respectively (Fig. 2a, b). Although no significant difference (P > 0.05) was observed in the Shannon index between the control and probiotic groups at the end of the trial, the Shannon index significantly decreased in the control group from the day 0 to day 7 (P < 0.05), contrasting to an uptrend observed in the probiotic group during the same period. At day 7, the Shannon index of the probiotic group was non-significantly higher than that of the control group (P = 0.06), suggesting that the intake of the routine drug might reduce the gut microbiota diversity and administering probiotics together with the routine drug could help lower such effect in some subjects (Fig. 2a). Moreover, the gut microbial richness continuously increased in the probiotic group over time, but such an increase only lasted until day 7 in the control group, followed by a slight decline afterwards (Fig. 2b). These results might represent the specific effect of using probiotics as an adjunctive treatment on the gut microbial diversity and richness.
Effects of treatments on alpha- and beta-diversity of irritable bowel syndrome (IBS) gut microbiota. Changes in a Shannon index; b Chao1 index. Error bars represent standard error. *P < 0.05; Mann–Whitney test. Principal coordinates analysis (PCoA; weighted UniFrac distance) of the gut microbiota of the two groups of patients with IBS at c day 0; d day 7; and e day 28. F-value and P-value on the PCoA score plots represent the difference between groups calculated by permutational multivariate analysis of variance (PERMANOVA)
To visualize the effect of the adjunctive probiotic treatment on the structure of patients’ fecal microbiota, a PCoA was performed based on the weighted UniFrac distances at different time points (Fig. 2c–e). No obvious clustering pattern was observed on the PCoA score plot between the two groups at the baseline level (Fig. 2c), but obvious clustering pattern was seen on the PCoA score plots at day 7 (P = 0.004, F = 4.094; PERMANOVA) and day 28 (P = 0.079, F = 1.857; PERMANOVA) (Fig. 2d, e), suggesting that the probiotic application caused obvious changes in the gut microbiota community. Interestingly, the microbiota structure exhibited a higher divergence between the control and the probiotic groups at day 7 compared to day 28, which was consistent with the changes observed in alpha-diversity.
Probiotic-directed modulation of patients’ fecal microbiota
To investigate probiotic-directed changes in the gut microbiota composition in patients with IBS, the fecal microbial taxonomic profiles at different time points (days 0, 7, 28) were analyzed. A more dramatic shift was observed in the gut microbiota in the control group after 7 day of routine drug treatment. At the genus level (Fig. 3a, b), the relative abundance of Prevotella first decreased and then increased in the probiotic group over time, whereas the control group showed an opposite trend. The relative abundance of Bacteroides of the control group increased significantly at day 7 (P < 0.0001), followed by a decline at day 28; and significantly more Bacteroides was found in the control group compared with the probiotic group at day 7 (P < 0.01). Significantly less Bifidobacterium (P < 0.05) and Eubacterium (P < 0.05) were observed in the control group at day 7 compared with the baseline level (Fig. 3b). Significantly less Dorea (P < 0.01) and Butyricicoccus (P < 0.05) were observed in the control group after the 28-day intervention, whereas only mild fluctuations were shown in these taxa for the probiotic group over time. Meanwhile, significantly less Citrobacter (P < 0.05), Clostridium (P < 0.05), and Escherichia (P < 0.05) were detected in the probiotic group after the 28-day treatment compared with day 0, whereas most participants in the control group showed only small variation in these taxa over time (Fig. 3b). It is interesting to note that among the few species that showed significant changes in their relative abundances during the intervention, some of them exhibited an opposite trend of changes between the two groups, suggesting a differential modulation effect of the adjunctive probiotic treatment towards the gut microbiota.
Effects of treatments on irritable bowel syndrome (IBS) gut microbiota composition. a Changes in the relative abundance of dominant genera; b Changes in the relative abundance of significant differential abundant genera identified between the two groups; c Spearman’s correlation heatmap of clinical and serum parameters and differential abundant genera. Error bars represent standard error. *P < 0.05, **P < 0.01, and ***P < 0.001; Mann–Whitney test
At day 28, interesting correlations were observed between some of the modulated genera and the monitored clinical and serum parameters (Fig. 3c). The IBS-SSS correlated significantly and negatively with most of the significantly diminished bacterial genera of the control group, such as Bifidobacterium (P < 0.05), Butyricicoccus (P < 0.05), Eubacterium (P < 0.01), and Prevotella (P < 0.05), while correlated positively with Escherichia (P < 0.05). Moreover, significantly less Escherichia was found in the probiotic group after 28-day treatment (P < 0.05). The IBS-QoL score showed a negative correlation with Clostridium (P < 0.05). The cytokine TNF-α showed a negative correlation with Bifidobacterium (P < 0.05) and Butyricicoccus (P < 0.05), but correlated positively with Citrobacter (P < 0.05). The serum IL-6 level correlated positively with Bacteroides (P < 0.05) and Escherichia (P < 0.05). The serum IL-8 level correlated positively with Escherichia (P < 0.05) and Citrobacter (P < 0.05), but associated negatively with Butyricicoccus (P < 0.05), Eubacterium (P < 0.05), and Prevotella (P < 0.01). These results suggested that the relief of IBS-associated symptoms might be associated with changes in the gut microbiota composition.
Treatment-induced changes in enterotype
To evaluate changes in the global microbial community, the gut microbiota enterotype of all samples were determined based on the similarity of 16S rRNA-based phylogenetic profile. Three distinct enterotypes could be identified at day 0. Five subjects of the control group and two subjects of the probiotic group belonged to enterotype 1, characterized by a high proportion of Eubacterium (28.61%) and Bacteroides (27.03%); seven subjects of the control group and 12 subjects of the probiotic group belonged to enterotype 2, which had a high proportion of Bacteroides (22.73%), Ruminococcus (22.73%), and Eubacterium (4.18%); seven subjects of the control group and ten subjects of the probiotic group belonged to enterotype 3, which was rich in Bacteroides (46.70%) (Fig. 4a–c).
Changes in the enterotype of subjects and microbial composition. a Principal component analysis (PCA) score plot showing enterotype clusters at day 0. b Boxplots showing the distribution of three most representative genera (Eubacterium, Bacteroides, and Prevotella) of enterotypes 1–3 at day 0. c Distribution of subjects' enterotype at day 0. d PCA score plot showing enterotype clusters at day 28. e Boxplots showing the distribution of Eubacterium, Bacteroides, and Prevotella of enterotypes 5 and 6 at day 28. f Distribution of subjects’ enterotype at day 28. g Distribution of identified differential abundant species in enterotype 5 (ET5) and enterotype 6 (ET6) at different time points; the mean relative abundance is represented in log scale. Error bars in the line charts represent standard error. *P < 0.05, **P < 0.01, and ***P < 0.001; Mann–Whitney test
At day 28, a drastic shift in enterotype was observed. All subjects were reclassified into two new enterotypes. Five subjects of the probiotic group belonged to enterotype 5 (enriched in Prevotella; 39.92%), while most subjects shifted to enterotype 6 (enriched in Bacteroides; 43.54%) (Fig. 4d–f). In contrast, all control subjects shifted to enterotype 5 on day 28. The fact that only the probiotic group but not the control group comprised two different enterotypes at day 28 suggested that the gut microbiota of the participants who received the adjunctive probiotic therapy responded divergently.
To further explore the divergent response within the probiotic group, differences in the initial microbial composition of subjects belonging to enterotypes 5 and 6 was compared. At day 0, enterotype 5 comprised significantly more Akkermansia muciniphila, Alistipes putredinis, Prevotella copri, and Ruminococcus bromii sequences, and significantly less Megamonas rupellensis, Clostridium saccharogumia, and Bacteroides uniformis than enterotype 6. Additionally, the number of differential abundant species increased at day 7 and then decreased at day 28. However, some of these differential abundant species (including Prevotella copri, Ruminococcus bromii, Alistipes putredinis) showed significant differences in their relative abundances persistently until day 28 (Fig. 4g). These results support that the species-level gut microbiota of the subjects of the probiotic group responded divergently toward the probiotic treatment and that the divergent response could be dependent on the initial gut microbiota composition of the individuals.
Initial gut microbiota composition influenced the level of inflammation towards probiotic treatment
The decrease in the magnitude of the serum LPS was significantly greater in subjects belonging to enterotype 5 than those belonging to enterotype 6 (P = 0.036), although such enterotype-specific difference was not as obvious for the decrease in the magnitude of serum TNF-α (P = 0.076) (Fig. 5a). Such results suggested that the host response towards the probiotic treatment was enterotype-specific.
Correlation between clinical and serum parameters with differential abundant gut taxa. a Reduction (concentration at day 0 minus concentration at day 28) in serum lipopolysaccharide (LPS) and tumor necrosis factor (TNF)-α levels of subjects belonging to enterotypes 5 and 6, respectively. Error bars in the line charts represent standard error. *P < 0.05; Mann–Whitney test. b Spearman's correlation network plot of clinical markers and identified differential abundant taxa. The green diamonds and red circles represent significantly modulated bacteria and clinical markers, respectively. Significant correlations between bacterial species and clinical markers are connected by straight lines. The line color represents the correlation strength as illustrated by the color scale of Spearman’s rho, ranging between 0.40 and − 0.56. Positive correlation is represented by a value greater than zero, while negative correlation is represented by a value smaller than zero
A correlation network was then constructed to identify associations between changes in the relative abundances of differential abundant microbes and clinical improvement (Fig. 5b). The data showed that the treatment outcomes were linked with the initial gut microbiota composition of the patients. For example, IBS-SSS correlated negatively with Ruminococcus gnavus, Prevotella copri, Alistipes indistinctus, and Alistipes indistinctus (most species were enriched in enterotype 5 at day 0); IBS-QoL correlated positively with Akkermansia muciniphila, but associated negatively with Megamonas rupellensis (both species were enriched in enterotype 6 at day 0). Lipopolysaccharide showed significant negative correlations with a number of bacteria, such as Akkermansia muciniphila, Oscillibacter valericigenes, Bacteroides uniformis, and Ruminococcus gnavus; most of which were enriched in enterotype 5 at day 0. The level of TNF-α correlated negatively with Clostridium saccharogumia, Oscillibacter valericigenes, Ruminococcus gnavus, Bacteroides uniformis; IL-6 correlated negatively with Akkermansia muciniphila and Coprobacter fastidiosus; IL-8 correlated negatively with Eubacterium xylanophilum and Eubacterium coprostanoligenes; d-lactate correlated positively with Megasphaera elsdenii and Clostridium saccharogumia.
Discussion
Irritable bowel syndrome is a common but difficult to treat medical condition, and the alteration of the intestinal microbiota has been proposed as one possible cause of IBS [30]. Previous studies reported different effects of a number of probiotic strains on alleviating clinical symptoms of IBS. For example, L. acidophilus DDS-1 and B. lactis UABla-12 could reduce the severity of abdominal pain and other IBS-associated symptoms [31]. Lactobacillus acidophilus CL1285, L. casei LBC80R, and L. rhamnosus CLR2 could improve the quality-of-life and IBS symptoms [32], but B. longum NCC3001 showed only weak clinical effect to improve depression in patients with IBS [33]. Hence, the clinical efficacy of probiotics on IBS is likely strain-specific. This work investigated the effect of adjunctive treatment with a multi-species probiotic powder product, Probio-Fit®, on the clinical outcomes of patients with IBS. Probio-Fit® contains three bacterial strains, namely L. casei Zhang, L. plantarum P-8, and B. animalis subsp. lactis V9. All three strains could improve the human colonic environment and modulate gut microbiota composition [34,35,36,37] (and unpublished data).
The severity of IBS was assessed with the overall IBS-QoL, IBS-SSS, and five serum factors (IL-6, IL-8, TNF-α, d-lactate, and LPS). Both the control and probiotic groups showed an increased IBS-QoL score and a decreased IBS-SSS at day 28; however, the IBS-SSS declined significantly more for the probiotic group, suggesting the adjunctive treatment improved the clinical effectiveness in managing IBS when compared to routine regimen alone. The increase in serum pro-inflammatory cytokines, such as IL-6, IL-8, and TNF-α, is thought to be a factor that triggers IBS development [38]. The abdominal discomfort associated with IBS is related to intestinal barrier dysfunction, causing increased intestinal permeability and low-grade inflammation [39]. d-lactate and LPS are biomarkers for intestinal permeability [40]. The serum levels of IL-8, TNF-α, and LPS showed a numerical decrease after four weeks of routine treatment with or without probiotic supplementation, suggesting that IBS-related inflammation was reduced in both groups. However, the serum IL-6 and d-lactate levels decreased significantly more in the probiotic group, and the serum IL-6 (P < 0.01) and TNF-α (P < 0.001) levels in the probiotic group were significantly lower than the control group at day 28. d-lactate is a metabolic product of bacterial fermentation, which is released into the blood circulation when the intestinal mucosa is destroyed [41]. Hence, our results suggest that the probiotic treatment strengthened the anti-inflammatory effect of routine therapy via mechanisms relating to the maintenance of gut-barrier integrity.
Our data showed that although routine regimen did not change the gut microbiota diversity significantly, it did cause temporary fluctuations in the Shannon diversity index. One interesting effect of administering probiotics in combination with routine regimen was the stabilization of the microbiota community during the course of treatment. Meanwhile, the adjunctive treatment also increased the gut microbiota richness in the participants. Such effects were not seen in the control group. In agreement with the changes in α-diversity, the weighted UniFrac distances of the control and probiotic groups did not differ significantly initially until day 7. Although such difference narrowed down at day 28, obvious clustering pattern could still be observed on the PCoA plot. The different patterns of change in the gut microbial diversity between the two groups suggested that there was probiotic-dependent differential modulation.
The different patterns of microbial compositional change in response to the intervention between the two sample groups also support the observation of probiotic-dependent differential modulation. Obviously less Bacteroides was found in the probiotic adjunctive treatment group on day 7. Patients with IBS are known to have a high level of gut Bacteroides [42], and some enterotoxigenic strains of Bacteroides fragilis could cause high-grade inflammation [43]. Prevotella was another differential abundant genus found in this work. The average relative abundance of Prevotella of the probiotic group increased from day 7, but an opposite trend was observed in the control group. A previous work reported that more Prevotella was found in the gut microbiota of healthy subjects than patients with IBS [7].
Bifidobacterium is an important genus in a healthy intestinal tract of a human. Members of this genus play key roles in degrading complex carbohydrates and inducing the maturation of the host immune system. It has been reported that the bifidobacterial population diminished significantly in the gut microbiota of IBS sufferers [44]. Although the probiotic adjunctive treatment did not increase the proportion of Bifidobacterium, less Bifidobacterium was detected in the control group at day 7, suggesting that the routine drug treatment did cause an adverse effect on the gut microbiota at least in the early phase of the treatment, and such unfavorable effect was diminished by the adjunctive probiotic treatment. Contradictory results have been obtained regarding the abundance of gut Lactobacillus in patients with IBS. Some studies found no apparent change in the level of gut Lactobacillus in patients with IBS, while other studies observed an increase in certain species, such as Lactobacillus salivarius [41]. Our study did not find any obvious change in the relative abundance of Lactobacillus in patients with IBS. In addition, the relative abundance of some gut commensals (e.g., Dorea, Butyricicoccus, and Eubacterium) [45] declined significantly in the control group as the treatment continued. On the other hand, the relative abundance of some potentially harmful genera, e.g., Citrobacter, Clostridium, and Escherichia, significantly decreased in the probiotic group but not the control group after the 28-day treatment. Citrobacter and Escherichia belong to the Enterobacteriaceae family, which is implicated in post-infectious IBS [46]. Some Clostridium species, such as Clostridium perfringens and Clostridium difficile, are harmful bacteria relating to serious gut inflammatory diseases like ulcerative colitis and Crohn's disease [47]. A previous human clinical trial revealed that the consumption of L. plantarum P-8, one of the strains taken by subjects in this work, could reduce the amount of Escherichia [34].
The improvement in some of the monitored clinical and serum parameters correlated with several probiotic-modulated genera. For example, the relative abundance of Bifidobacterium, Butyricicoccus, and Prevotella correlated negatively with IBS-SSS, and TNF-α, and IL-8, while Escherichia, Clostridium, and Citrobacter (P < 0.05) correlated positively with the decreases in IBS-SSS and serum parameters (IL-8, IL-6, and TNF-α). These results are in line with a previous study showing fermented milk consumption alleviated the symptoms of IBS accompanied by mild gut microbiota modulation involving few metagenomic species [48]. A previous report found that administering B. infantis improved the overall IBS-associated symptoms along with the modulation of IL-10/IL-12 ratio and Th-1 pro-inflammatory cytokines [49]. These observations support that the improvement of IBS-associated symptoms by probiotic supplementation was via host immune modulation.
The use of probiotics as adjunctive therapy in managing IBS also showed an interesting effect in shifting the gut microbiota of subjects to either a Bacteroides-dominated or a Prevotella-dominated enterotype, whereas all subjects from the control group adopted the Bacteroides-dominated enterotype. Meanwhile, the extent of decrease in serum LPS and TNF-α was lower for subjects adopted the Prevotella-enterotype after the adjunctive probiotic treatment. A previous study reported that there was more Prevotella in the gut microbiota of healthy individuals than patients with IBS, and thus these bacteria might help protect from IBS development [7]. It is worth noting that the initial gut microbiome of subjects adopted the Prevotella-enterotype was enriched in several species, including Akkermansia muciniphila, Alistipes putredinis, Prevotella copri, and Ruminococcus bromii; and the relative abundance of some of these bacteria correlated positively and significantly with the decrease in inflammatory markers like LPS and TNF-α. These results suggested that the initial microbiota composition could affect the clinical outcome of the adjunctive probiotic treatment, which was previously reported in a study that applied the probiotic product BIO-25 in IBS management [50]. The mechanism of how the initial gut microbiome directed the clinical outcome awaits further elucidation. One reason could be that certain microbes were more responsive to environmental changes, and if these taxa comprised a relatively high proportion of the initial gut microbiota, the overall microbiota structure might be modulated or reshaped more readily by treatments like probiotic or drug administration.
In conclusion, the adjunctive use of probiotics with a routine regimen showed additional clinical effectiveness when compared to routine regimen alone in managing IBS. In addition, our results clearly demonstrated that the probiotic product did not work equally well for all participants of this trial; and the initial gut microbiota composition might affect the clinical efficacy of the probiotic treatment. Such data would be helpful in designing probiotic-based regimen at a personalized level to achieve maximum benefit for IBS sufferers.
Abbreviations
- IBS:
-
Irritable bowel syndrome
- IL:
-
Interleukin
- TNF:
-
Tumor necrosis factor
- LPS:
-
Lipopolysaccharide
- SIBO:
-
Small intestinal bacteria overgrowth
- FODMAP:
-
Fermentable oligosaccharides, Disaccharides, monosaccharides, and polyols
- L.:
-
Lactobacillus
- B.:
-
Bifidobacterium
- SMRT:
-
Single-molecule real-time
- PCR:
-
Polymerase chain reaction
- QIIME:
-
Quantitative insights into microbial ecology
- PCoA:
-
Principle coordinate analysis
- PERMANOVA:
-
Permutational multivariate analysis of variance
- SSS:
-
Symptom severity score
- QoL:
-
Quality of life score
References
Defrees DN, Bailey J (2017) Irritable bowel syndrome: epidemiology, pathophysiology, diagnosis, and treatment. Prim Care 44(4):655–671. https://doi.org/10.1016/j.pop.2017.07.009
Gwee KA, Ghoshal UC, Chen MH (2018) Irritable bowel syndrome in Asia: pathogenesis, natural history, epidemiology, and management. J Gastroen Hepatol 33(1):99–110. https://doi.org/10.1111/jgh.13987
Miyoshi J, Chang EB (2017) The gut microbiota and inflammatory bowel diseases. Transl Res 179:38–48. https://doi.org/10.1016/j.trsl.2016.06.002
Quigley EMM (2018) The gut-brain axis and the microbiome: clues to pathophysiology and opportunities for novel management strategies in irritable bowel syndrome (IBS). J Clin Med. https://doi.org/10.3390/jcm7010006
Labus JS, Hollister EB, Jacobs J, Kirbach K, Oezguen N, Gupta A, Acosta J, Luna RA, Aagaard K, Versalovic J, Savidge T, Hsiao E, Tillisch K, Mayer EA (2017) Differences in gut microbial composition correlate with regional brain volumes in irritable bowel syndrome. Microbiome 5:49. https://doi.org/10.1186/s40168-017-0260-z
Pyleris E, Giamarellos-Bourboulis EJ, Tzivras D, Koussoulas V, Barbatzas C, Pimentel M (2012) The prevalence of overgrowth by aerobic bacteria in the small intestine by small bowel culture: relationship with irritable bowel syndrome. Digest Dis Sci 57(5):1321–1329. https://doi.org/10.1007/s10620-012-2033-7
Tap J, Derrien M, Tornblom H, Brazeilles R, Cools-Portier S, Dore J, Storsrud S, Le Neve B, Ohman L, Simren M (2017) Identification of an intestinal microbiota signature associated with severity of irritable bowel syndrome. Gastroenterology 152(1):111. https://doi.org/10.1053/j.gastro.2016.09.049
Stern EK, Brenner DM (2018) Gut microbiota-based therapies for irritable bowel syndrome. Clin Transl Gastroen 9:e134. https://doi.org/10.1038/ctg.2018.2
Harris LA, Baffy N (2017) Modulation of the gut microbiota: a focus on treatments for irritable bowel syndrome. Postgrad Med 129(8):872–888. https://doi.org/10.1080/00325481.2017.1383819
Ishaque SM, Khosruzzaman SM, Ahmed DS, Sah MP (2018) A randomized placebo-controlled clinical trial of a multi-strain probiotic formulation (Bio-Kult (R)) in the management of diarrhea- predominant irritable bowel syndrome. Bmc Gastroenterol. https://doi.org/10.1186/s12876-018-0788-9
Shin SP, Choi YM, Kim WH, Hong SP, Park JM, Kim J, Kwon O, Lee EH, Hahm KB (2018) A double blind, placebo-controlled, randomized clinical trial that breast milk derived Lactobacillus gasseri BNR17 mitigated diarrhea-dominant irritable bowel syndrome. J Clin Biochem Nutr 62(2):179–186. https://doi.org/10.3164/jcbn.17-73
Whelan K (2011) Probiotics and prebiotics in the management of irritable bowel syndrome: a review of recent clinical trials and systematic reviews. Curr Opin Clin Nutr 14(6):581–587. https://doi.org/10.1097/MCO.0b013e32834b8082
El-Salhy M, Patcharatrakul T, Hatlebakk JG, Hausken T, Gilja OH, Gonlachanvit S (2017) Chromogranin A cell density in the large intestine of Asian and European patients with irritable bowel syndrome. Scand J Gastroentero 52(6–7):691–697. https://doi.org/10.1080/00365521.2017.1305123
Xu H, Huang W, Hou Q, Kwok LY, Laga W, Wang Y, Ma H, Sun Z, Zhang H (2019) Oral administration of compound probiotics improved canine feed intake, weight gain immunity and intestinal microbiota. Front Immunol 10:666. https://doi.org/10.3389/fimmu.2019.00666
Zhang J, Zhao J, Jin H, Lv R, Shi H, De G, Yang B, Sun Z, Zhang H (2020) Probiotics maintain the intestinal microbiome homeostasis of the sailors during a long sea voyage. Gut microbes. https://doi.org/10.1080/19490976.2020.1722054
Mosher JJ, Bernberg EL, Shevchenko O, Kan J, Kaplan LA (2013) Efficacy of a 3rd generation high-throughput sequencing platform for analyses of 16S rRNA genes from environmental samples. J Microbiol Meth 95(2):175–181. https://doi.org/10.1016/j.mimet.2013.08.009
Liu WJ, Zheng Y, Kwok LY, Sun ZH, Zhang JC, Guo Z, Hou QC, Menhe B, Zhang HP (2015) High-throughput sequencing for the detection of the bacterial and fungal diversity in Mongolian naturally fermented cow's milk in Russia. Bmc Microbiol 15:45. https://doi.org/10.1186/s12866-015-0385-9
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Tumbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–336. https://doi.org/10.1038/nmeth.f.303
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R (2010) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26(2):266–267
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26(19):2460–2461
Haas BJ, Gevers D, Earl AM, Feldgarden M, Ward DV, Giannoukos G, Ciulla D, Tabbaa D, Highlander SK, Sodergren E, Methe B, DeSantis TZ, Petrosino JF, Knight R, Birren BW, Consortium HM (2011) Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res 21(3):494–504
Cole JR, Chai B, Farris RJ, Wang Q, Kulam-Syed-Mohideen AS, McGarrell DM, Bandela AM, Cardenas E, Garrity GM, Tiedje JM (2007) The ribosomal database project (RDP-II): introducing myRDP space and quality controlled public data. Nucleic Acids Res 35:D169–D172
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microb 72(7):5069–5072. https://doi.org/10.1128/Aem.03006-05
Hou Q, Xu H, Zheng Y, Xi X, Kwok LY, Sun Z, Zhang H, Zhang WJJoDS (2015) Evaluation of bacterial contamination in raw milk, ultra-high temperature milk and infant formula using single molecule, real-time sequencing technology. J Diary Sci 98(12):8464–8472
Price MN, Dehal PS, Arkin AP (2009) FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol Biol Evol 26(7):1641–1650
Lozupone C, Knight R (2005) UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microb 71(12):8228–8235
Francis CY, Morris J, Whorwell PJ (2003) The irritable bowel severity scoring system: a simple method of monitoring irritable bowel syndrome and its progress. Aliment Pharmacol Ther 11(2):395–402. https://doi.org/10.1046/j.1365-2036.1997.142318000.x
Patrick DL, Drossman DA, Frederick IO, Dicesare J, Puder KL (1998) Sciences quality of life in persons with Irritable bowel syndrome (development and validation of a new measure). Dig Dis Sci 43(2):400–411. https://doi.org/10.1023/A:1018831127942
Benjamini Y, Yekutieli D (2001) The control of the false discovery rate in multiple testing under dependency. Ann Stat 29(4):1165–1188. https://doi.org/10.1214/aos/1013699998
Ringel Y (2017) The gut microbiome in irritable bowel syndrome and other functional bowel disorders. Gastroenterol Clin N 46(1):91. https://doi.org/10.1016/j.gtc.2016.09.014
Martoni CJ, Srivastava S, Leyer GJ (2020) Lactobacillus acidophilus DDS-1 and Bifidobacterium lactis UABla-12 improve abdominal pain severity and symptomology in irritable bowel syndrome: randomized controlled trial. Nutrients. https://doi.org/10.3390/nu12020363
Preston K, Krumian R, Hattner J, de Montigny D, Stewart M, Gaddam S (2018) Lactobacillus acidophilus CL1285, Lactobacillus casei LBC80R and Lactobacillus rhamnosus CLR2 improve quality-of-life and IBS symptoms: a double-blind, randomised, placebo-controlled study. Benef Microbes 9(5):697–706. https://doi.org/10.3920/BM2017.0105
Meyer C, Vassar M (2018) The fragility of probiotic bifidobacterium longum NCC3001 use for depression in patients with irritable bowel syndrome. Gastroenterology 154(3):764. https://doi.org/10.1053/j.gastro.2017.09.055
Kwok LY, Guo Z, Zhang J, Wang L, Qiao J, Hou Q, Zheng Y, Zhang H (2015) The impact of oral consumption of Lactobacillus plantarum P-8 on faecal bacteria revealed by pyrosequencing. Benef Microbes 6(4):405–413. https://doi.org/10.3920/Bm2014.0063
Kwok LY, Wang L, Zhang J, Guo Z, Zhang H (2014) A pilot study on the effect of Lactobacillus casei Zhang on intestinal microbiota parameters in Chinese subjects of different age. Benef Microbes 5(3):295–304. https://doi.org/10.3920/Bm2013.0047
Wang L, Zhang J, Guo Z, Kwok L, Ma C, Zhang W, Lv Q, Huang W, Zhang HJN (2014) Effect of oral consumption of probiotic Lactobacillus planatarum P-8 on fecal microbiota, SIgA, SCFAs, and TBAs of adults of different ages. Nutrition 30(7–8):776–783. https://doi.org/10.1016/j.nut.2013.11.018
Zhang J, Wang L, Guo Z, Sun Z, Gesudu Q, Kwok L, Menghebilige, Zhang HP (2014) 454 pyrosequencing reveals changes in the faecal microbiota of adults consuming Lactobacillus casei Zhang. FEMS Microbiol Ecol 88(3):612–622. https://doi.org/10.1111/1574-6941.12328
Seyedmirzaee S, Hayatbakhsh MM, Ahmadi B, Baniasadi N, Rafsanjani AMB, Nikpoor AR, Mohammadi M (2016) Serum immune biomarkers in irritable bowel syndrome. Clin Res Hepatol Gas 40(5):631–637. https://doi.org/10.1016/j.clinre.2015.12.013
Camilleri M, Lasch K, Zhou W (2012) Irritable bowel syndrome: methods, mechanisms, and pathophysiology. The confluence of increased permeability, inflammation, and pain in irritable bowel syndrome. Am J Physiol-Gastr L 303(7):G775–G785. https://doi.org/10.1152/ajpgi.00155.2012
Bischoff SC, Barbara G, Buurman W, Ockhuizen T, Schulzke JD, Serino M, Tilg H, Watson A, Wells JM (2014) Intestinal permeability—a new target for disease prevention and therapy. Bmc Gastroenterol 14:189. https://doi.org/10.1186/s12876-014-0189-7
Sun XQ, Fu XB, Zhang R, Lu Y, Deng Q, Jiang XG, Sheng ZY (2001) Relationship between plasma D(-)-lactate and intestinal damage after severe injuries in rats. World J Gastroenterol 7(4):555–558. https://doi.org/10.3748/wjg.v7.i4.555
Liu Y, Zhang L, Wang X, Wang F, Zhang J, Jiang R, Wang X, Wang K, Liu Z, Xia ZJ, Nie Y, Lv XL, Wu XL, Zhu HQ, Duan LP (2016) Similar fecal microbiota signatures in patients with diarrhea-predominant Irritable bowel syndrome and patients with depression. Clin Gastroenterol Hepatol 4(11):1602–1611. https://doi.org/10.1016/j.cgh.2016.05.033
Macfarlane S, Woodmansey EJ, Macfarlane GT (2005) Colonization of mucin by human intestinal bacteria and establishment of biofilm communities in a two-stage continuous culture system. Appl Environ Microb 71(11):7483–7492. https://doi.org/10.1128/Aem.71.11.7483-7492.2005
Pittayanon R, Lau JT, Yuan YH, Leontiadis GI, Tse F, Surette M, Moayyedi P (2019) Gut microbiota in patients with irritable bowel syndrome—a systematic review. Gastroenterology 157(1):97–108. https://doi.org/10.1053/j.gastro.2019.03.049
Qin JJ, Li RQ, Raes J, Arumugam M, Burgdorf KS, Manichanh C, Nielsen T, Pons N, Levenez F, Yamada T, Mende DR, Li JH, Xu JM, Li SC, Li DF, Cao JJ, Wang B, Liang HQ, Zheng HS, Xie YL, Tap J, Lepage P, Bertalan M, Batto JM, Hansen T, Le Paslier D, Linneberg A, Nielsen HB, Pelletier E, Renault P, Sicheritz-Ponten T, Turner K, Zhu HM, Yu C, Li ST, Jian M, Zhou Y, Li YR, Zhang XQ, Li SG, Qin N, Yang HM, Wang J, Brunak S, Dore J, Guarner F, Kristiansen K, Pedersen O, Parkhill J, Weissenbach J, Bork P, Ehrlich SD, Wang J, Consortium M (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464(7285):59-U70. https://doi.org/10.1038/nature08821
Rajilic-Stojanovic M, Biagi E, Heilig HGHJ, Kajander K, Kekkonen RA, Tims S, de Vos WM (2011) Global and deep molecular analysis of microbiota signatures in fecal samples from patients with irritable bowel syndrome. Gastroenterology 141(5):1792–1801. https://doi.org/10.1053/j.gastro.2011.07.043
Azimirad M, Yadegar A, Aghdaei HA, Kelly CR (2019) Enterotoxigenic clostridium perfringens infection as an adverse event after faecal microbiota transplantation in two patients with ulcerative colitis and recurrent clostridium difficile infection: a neglected agent in donor screening. J Crohns Colitis 13(7):960–961. https://doi.org/10.1093/ecco-jcc/jjz006
Veiga P, Pons N, Agrawal A, Oozeer R, Guyonnet D, Brazeilles R, Faurie JM, van Hylckama Vlieg JE, Houghton LA, Whorwell PJ, Ehrlich SD, Kennedy SP (2014) Changes of the human gut microbiome induced by a fermented milk product. Sci Rep 4:6328. https://doi.org/10.1038/srep06328
Whorwell PJ, Altringer L, Morel J, Bond Y, Charbonneau D, O’Mahony L, Kiely B, Shanahan F, Quigley EMM (2006) Efficacy of an encapsulated probiotic Bifidobacterium infantis 35624 in women with irritable bowel syndrome. Am J Gastroenterol 101(7):1581–1590. https://doi.org/10.1111/j.1572-0241.2006.00734.x
Hod K, Dekel R, Cohen NA, Sperber A, Ron Y, Boaz M, Berliner S, Maharshak N (2018) The effect of a multispecies probiotic on microbiota composition in a clinical trial of patients with diarrhea-predominant irritable bowel syndrome. Neurogastroent Motil 30(12):e13456. https://doi.org/10.1111/nmo.13456
Acknowledgements
We sincerely thank all the volunteers for their participation. This research was supported by the National Natural Science Foundation of China (Grant Nos. 31720103911, 31972083) and Inner Mongolia Science & Technology Major Projects (Grant No. ZDZX2018018).
Author information
Authors and Affiliations
Contributions
The authors’ responsibilities were as follows: HX analyzed the data. CM, HX, FZ, YL, PC, LC conducted research. HX and LYK wrote the paper. HZ, ZS, LC designed research. HZ and LYK had primary responsibility for the final content, and all authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xu, H., Ma, C., Zhao, F. et al. Adjunctive treatment with probiotics partially alleviates symptoms and reduces inflammation in patients with irritable bowel syndrome. Eur J Nutr 60, 2553–2565 (2021). https://doi.org/10.1007/s00394-020-02437-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00394-020-02437-4