Abstract
Purpose
Inulin is a soluble dietary fiber that has been implicated in regulating the intestinal health. Here, we describe a synergetic response of intestinal microbiota and epithelial functions to increased intake of inulin in a porcine model.
Methods
Twenty growing-pigs were randomly allocated to two groups (n = 10) and fed with a basal diet (BD) or BD containing 0.5% inulin (INU) for 21 days.
Results
We show that INU supplementation not only elevated villus height and the abundance of zonula occludens-1 (ZO-1), but also increased acetate and butyrate concentrations in cecum (P < 0.05). Moreover, INU decreased IL-6 and TNFα secretion, and reduced intestinal epithelial cell apoptosis in ileum and cecum (P < 0.05). Interestingly, we observed an elevated 16S rRNA gene copies in cecum after INU ingestion (P < 0.05). INU had no influence on overall diversity, but acutely altered the abundance of specific bacteria. INU decreased the abundance of phylum Proteobacteria in ileum, but increased the phylum Bacteroidetes in the ileum and cecum (P < 0.05). INU significantly elevated the Lactobacillus spp. and Bacteroides spp. in the ileum and cecum, respectively. Importantly, INU elevated the expression levels of GPR43, GLP-2, and ZO-1, but decreased the expression levels of histone deacetylase 1 (HDAC1) and TNFα in the ileum and cecum mucosa (P < 0.05). Moreover, INU also elevated the expression levels of GPR109A and angiopoietin-4 (ANG-4) in the cecum mucosa (P < 0.05).
Conclusions
This study indicated how the intestinal microbiome and epithelium adapt to inulin ingestion, and furthered our understanding of the mechanisms behind the dietary fiber-modulated intestinal microbiota and health.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, dietary fibers have attracted considerable research interest because of their roles in maintaining the gut health and regulating nutrient metabolism [1, 2]. Dietary fibers may act as a prebiotic by being resistant to gastric acidity and enzymes such as maltase, isomaltase, and sucrose, and then undergoing fermentation by a wide variety of microorganisms in the large intestine to SCFA that promote the growth of beneficial microorganisms such as the Bifidobacterium and Lactobacillus species [3]. The fiber-derived SCFA has also been implicated in regulating the growth of intestinal epithelial mucosa. For instance, cecal infusion of butyrate was found to increase intestinal cell proliferation in piglets [4]. In addition to their beneficial effects on intestinal health, dietary fiber-derived SCFAs also participate in the regulation of metabolism. Mice fed a butyrate-enriched high-fat diet have increased thermogenesis and energy expenditure and are resistant to obesity [5]. In the same manner, oral acetate gavage in an obese and diabetic strain of rats reduced weight gain and improved glucose tolerance [6]. More recently, the butyrate and propionate were found to serve as the substrates for intestinal gluconeogenesis, which ultimately contribute to blood glucose homeostasis [7].
Inulin is an important dietary fiber consisting of a mixture of linear fructose polymers or fructans of two to sixty units each linked by unique β(2–1) bonds with a glucose unit linked by an α(1–2) bond at the end of each chain [8]. This unique structure prevents it from being hydrolyzed by salivary or pancreatic enzymes, and, therefore, inulin barely has prebiotic effects for monogastric animals [8, 9]. Inulins are present in many vegetables, and can also be extracted and isolated from chicory roots, to be used as food ingredients [10]. A number of studies have indicated that dietary inulin supplementation has beneficial effects on growth performance and intestinal health in a variety of animal species [11,12,13]. For instance, inulin supplementation was found to module the immune response and restore gut morphology in malnourished mice [13]. Moreover, specific changes were observed in the human colon microbiota after ingestion of inulin-type fructans [14]. However, little is known about the synergetic responses of intestinal microbiota and epithelium barrier function to this unique dietary non-starch carbohydrate.
In the present study, we describe the synergetic responses of intestinal microbiota and epithelium to dietary inulin supplementation in a porcine model. Pig (Sus scrofa) is an excellent model species used in biomedical researches, as they are closely related to humans in terms of anatomy, genetics and physiology [15]. Moreover, both species are omnivorous and their organs generally share common functional features. The digesta samples from ileum and cecum were collected from the growing pigs fed with a basal diet or basal diet containing 0.5% inulin. The bacterial community in the intestine was analyzed using 16S rRNA sequencing. The ileum and cecum samples were used, as previous study indicated that the distal of small intestine (ileum) and cecum are major degradation sites of ingested inulin in young pigs [16]. This study will facilitate understanding of the mechanisms behind the dietary non-starch carbohydrate modulated gut health in monogastric mammals.
Methods
Studies involving animals were conducted according to the Regulations for the Administration of Affairs Concerning Experimental Animals (Ministry of Science and Technology, China, revised in June 2004). Sample collection was approved by the Institutional Animal Care and Use Committee of Sichuan Agricultural University, Sichuan, China (no. 20180901).
Animals housing and sample collection
Twenty crossbred (Duroc × Landrace × Yorkshire) growing pigs with an average initial body weight of 22.27 ± 0.14 kg, were randomly allocated to two groups (n = 10). Pigs were kept individually and fed with a fiber-free basal diet (BD) or BD containing with 0.5% inulin (INU). The diets (Table S1) were formulated to meet the nutrient recommendations of the National Research Council 2012 [17], and the chemical composition of the diet was analyzed using the AOAC method [18]. Pigs were fed ad libitum and given free access to water. After 21 days, pigs were sacrificed by exsanguination under deep anesthesia via intravenous injection of pentobarbital sodium (59 mg/kg body weight), and the intestinal tissues and mucosa samples were collected immediately. Additionally, approximately 4 g digesta from the middle section of the ileum and cecum was transferred into sterile tubes and immediately frozen at – 80 ∘C for analysis of the short-chain fatty acid (SCFA) concentration and the bacterial community.
Analysis of intestinal mucosa morphology
The ileal and cecal samples were douched with physiologic saline and stored in 4% paraformaldehyde solution. The preserved segments were prepared after staining with hematoxylin and eosin (HE) solution suing standard paraffin embedding procedures [19]. At least ten intact, well-oriented crypt-villus units were selected in triplicate as sources of each pig intestine cross section. Morphometric variables, including villus height and crypt depth were measured with an image processing and analysis system (Image Pro Plus, Media Cybernetics, Bethesda, MD, USA).
Immunofluorescence analysis
The localization of ZO-1 protein was determined by using immunofluorescence. 4% paraformaldehyde-fixed samples were rinsed in PBS and then incubated with ethylene diamine tetraacetic acid (EDTA, 1 mol/L, pH 9.0) for antigen retrieval. Tissue sections were blocked with 3% bovine serum albumin prior to incubation with rabbit anti-ZO-1 polyclonal antibody (1:250; Abcam Plc., Cambridge, UK) overnight at 4 °C. Slides were then washed three times with PBS and incubated with goat anti-rabbit IgG-FITC secondary antibody (Gooddbio Technology Co., Ltd., Wuhan, China) for 1 h at room temperature in the dark. Finally, slides were washed three times with PBS, and the nuclei were stained with 4′-6-diamidino-2-phenylindole (DAPI) for 10 min at room temperature in the dark. All slides were examined for fluorescence using a laser scanning confocal microscope (FV1000; Olympus Corporation, Tokyo, Japan).
Determination of intestinal epithelial cell apoptosis
Ileal and cecal epithelial cells were isolated and the percentage of apoptotic cells was analyzed using flow cytometry with a PE Annexin V Apoptosis Detection Kit (Becton, Dickinson and Company, BD Biosciences, San Joes, CA, USA). The sample preparation was based on the manual provided by the manufacturer, and the apoptotic cells were examined by flow cytometry (CytoFlex, Beckman Coulter, Inc., Brea, CA, USA) within 1 h.
Measurements of mucosal inflammatory cytokines
The concentrations of interleukin-1 (IL-1), IL-6, and tumour necrosis factor-α (TNFα) in the intestinal mucosa were determined using the corresponding ELISA kits (Beijing Winter Song Boye Biotechnology Co., Ltd). All determinations were carried out in triplicated, and absorbance was measured using a multi-mode microplate reader (SpectraMax M2, Molecular Devices, Sunnyvale, CA, USA).
Analysis of SCFA concentrations
The SCFA (acetic acid, propionic acid, and butyric acid) concentrations were determined using a gas chromatograph system (VARIAN CP-3800, Varian, Palo Alto. CA. USA; capillary column 30 m × 0.32 mm × 0.25 µm film thickness) following previous method [20]. After vortex, the digesta was centrifuged at 4 °C for 10 min (12,000 g), and the supernatant (1 ml) was then transferred into an Eppendorf tube (2 ml) and mixed with 0.2 ml metaphosphoric acid. After 30 min incubation at 4 °C, the tubes were centrifuged at 4 °C for 10 min (12, 000 g) and aliquots of the supernatant (1 µl) were analyzed using the GC with a flame ionization detector and an oven temperature of 100–150 °C. The polyethylene glycol column was operated with highly purified N2 as the carrier gas at 1.8 ml/min.
Determination of the digesta pH values
Immediately after the pigs were killed, approximately 5 g digesta was collected into ice-bathed sterile centrifugal tube, and then the pH value of each sample was determined using a PHS-3C pH meter (Shanghai, China).
Analysis of the bacterial community
Nucleic acids were extracted from 0.5 g of digesta sample using the Stool DNA kit (Omega Bio-Tech, Doraville, CA, USA). The diversity and composition of the bacterial community in each of these samples was determined using the protocol described in Caporase et al. [21]. For amplification of the V4 region of the 16S rRNA gene, the 520F/802R primer set (520F: CCATCTCATCCCTGCGTGTCTCCGAC; 802R: CCTCTCTATGGGCAGTCGGT-GAT) was used [22]. The amplification procedures were based on previously published protocol [23]. Briefly, the PCR conditions used were 5 min at 95 °C, 35 cycles of 30 s at 94 °C, 30 s at 55 °C and 90 s at 72 °C, followed by 10 min at 72 °C. Amplification was carried out using a Verity Thermocycler (Applied Biosystems). The PCR products derived from amplification of specific 16S rRNA gene hypervariable regions were purified by electrophoretic separation on an 1.5% agarose gel and the use of a Wizard SV Gen PCR Clean-Up System (Promega), followed by a further purification step involving the Agencourt AMPure XP DNA purification beads (Beckman Coulter Genomics GmbH, Bernried, Germany) to remove primer dimers. The sequencing was conducted on an Illumina MiSeq 2000 platform (Personal Biotechnology, Shanghai).
Pairs of reads from the original DNA fragments are merged using FLASH [24], a very fast and accurate software tool, which is designed to merge pairs of reads when the original DNA fragments are shorter than twice the length of reads. Sequencing reads was assigned to each sample according to the unique barcode of each sample. Sequences were analyzed with the QIIME software package and UPARSE pipeline [25], in addition to custom Perl scripts to analyze alpha (within sample) and beta (between sample) diversity. UPARSE was used to pick operational taxonomic units (OTUs). Sequences were assigned to OTUs using 97% species-level sequence identity. The first sequence assigned to each OTU was used as the reference sequence for that OTU field. RDP classifier was used to assign taxonomic data to each representative sequence [26]. Rarefaction curves were generated using QIIME, and this software was also used to calculate both weighted and unweighted UniFrac for principal coordinate analysis (PCoA) and unweighted pair group method with arithmetic mean (UPGMA) clustering.
RNA isolation, reverse transcription, and qPCR
Total RNA was isolated from intestinal mucosa using the RNAiso Plus (Takara Biotechnology Co., Ltd., Dalian, China). The concentration and quality of total RNA were assessed using a spectrophotometer (NanoDrop 2000, Thermo Fisher Scientific, Inc., Waltham, MA, USA). About 1 µg total RNA was used to synthesize cDNA, based on the protocol of PrimeScript™ RT reagent kit (Takara Biotechnology Co., Ltd). The G protein-coupled receptor 41 (GPR41), GPR43, GPR109A, Sodium-couple monocarboxylate transporter 1 (SLC5A8), Histone deacetylase 1 (HDAC1), HDAC3, HDAC4, IL-1, IL-10, IL-18, and TNFα mRNA levels in the intestinal mucosa were quantified using qPCR. In brief, the specific primers were designed using Primer Express 3.0 software (Applied Biosystems, Foster City, CA, USA) and synthesized by Sangon Biotech Co., Ltd. (Shanghai, China). All qPCR reactions were performed in triplicate on a QaunStudoTM 6 Flex Real-Time PCR System (Applied Biosystems), using SYBR Premix Ex Taq™ II (Tli RNaseH Plus) (Takara Biotechnology Co., Ltd). Amplification was performed in a final volume of 10 µl, which consisted of 5 µl of SYBR Premix Ex Taq™ II, 0.2 µl ROX Reference Dye II, 0.4 µl forward primer (10 µmol/l), 0.4 µl reverse primer (10 µmol/l), 1 µl cDNA and 3 µl diethylpyrocarbonate-treated water, under the following cycling conditions: 95 °C for 30 s, followed by 40 cycles; at 95 °C for 5 s and 60 °C for 34 s. After the amplification, a melt curve was generated at 95 °C for 15 s, 60 °C for 1 min and 95 °C for 15 s. Porcine glyceraldehyde-3-phosphate dehydrogenase (GAPDH) gene was used as the housekeeping gene, and the relative gene expression was calculated using the comparative CT method [27].
Statistical analysis
Differences in the intestinal parameters, gene copy numbers, and concentrations of SCFAs were analyzed by Student’s t test of SAS 9.4 statistic software (SAS Institute Inc., Cary, NC). The results were expressed as mean ± S.D. Statistical significance and a tendency towards difference were considered as P < 0.05 and P < 0.10, respectively.
Results
Influences of dietary inulin supplementation on intestinal morphology and apoptosis of intestinal epithelial cells
As shown in Fig. 1a and Table 1, dietary INU supplementation had no significant influence on the mucosa morphology in the ileum (P > 0.05). However, dietary INU supplementation significantly elevated the villus height in the cecum (P < 0.05). INU supplementation had no significant influence on the localization of ZO-1 protein in the ileal mucosa. However, INU supplementation improved the localization and abundance of the ZO-1 protein in the cecal mucosa (Fig. 1b). Importantly, dietary INU supplementation significantly decreased the total number of apoptotic cells in the ileum and cecum (Fig. 1c).
Effect of INU on mucosa morphology, tight-junction protein distribution, and apoptosis of intestinal epithelial cells. a H&E staining of the ileal and cecal samples. b Immunofluorescence assays of the ZO-1 protein (red) localization. c Cell apoptosis was analyzed using flow cytometry. 30,000 cells were used in each acquisition reading. Frames were divided into 4 quadrants: Q1-UL represents necrotic cells; Q1-UR represents late apoptotic and early necrotic cells; Q1-LR represents early apoptotic cells; and Q1-LL represents normal cells, respectively. Pigs were fed with basal diet (Control) or inulin containing diet (Inulin), n = 10; * means P < 0.05, ** means P < 0.01
Influences of dietary inulin supplementation on secretion of mucosal inflammatory cytokines
We show that dietary INU supplementation had no significant influence on the content of IL-1 in the intestinal mucosa (Table 1). However, INU significantly decreased the contents of inflammatory cytokines such as the IL-6 and TNFα in the ileal and cecal mucosa (P < 0.05).
Influences of dietary inulin supplementation on microbial metabolites
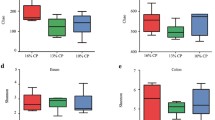
As shown in Table 1, dietary INU supplementation tended to increase the acetate content in the ileal digesta (P = 0.09). Interestingly, both the acetate and butyrate contents in the cecal digesta were significant higher in the INU group than in the control group (P < 0.05). Moreover, INU tended to decrease the pH value in the cecal digesta (P = 0.08).
Influence of dietary inulin supplementation on total 16S rRNA gene copies in the ileum and cecum
The 16S rRNA gene copies were determined using quantitative PCR, and equal volumes of the purified DNA of all samples were used in the assay. We show that the average copy number in the cecal samples were higher than in the ileal samples (Fig. 2). Dietary INU supplementation had no significant influence on the total 16S rRNA gene copies in the ileal digesta. However, INU significantly elevated the total 16S rRNA gene copies in the cecal digesta (P < 0.05).
Influence of inulin on bacterial community structures
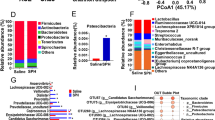
Digesta samples were compared based on the proportions of bacterial lineages in each specimen using 16S rRNA sequencing. Equal volumes of the purified DNA from each sample were used. The DNA samples were amplified by PCR using bar-coded primers flanking the V4 region of the 16S rRNA gene, and samples were sequenced using the Illumina method. All sequencing information has been deposited in the National Center for Biotechnology Information (NCBI) and can be accessed in the Short Read Archie (SRA) under the accession number PRJNA559763. We generated a total of 1,285,591 sequences with an average of 65,394 sequences per ileal sample and 63,166 per cecal sample after post-quality filtering (Table S2). These sequences had an average length of 220–230 base pairs. The qualified sequence (> 0.0001%) were clustered into 1051 bacterial OTUs with UPARSE software [25]. As expected, more OTUs were identified in the cecum than in the ileum. Interestingly, 164 OTUs were only found in the ileum of INU group, whereas 98 OTUs were specifically identified in the ileum of the control group. The two groups shared 432 OTUs in the ileal samples (Fig. 3). Additionally, more than half of the identified OTUs in the cecal samples were shared by the two groups.
Number of identified OTUs in various comparisons. a Venn diagram shows various comparisons of ileal OTUs at the genus level; b Venn diagram shows various comparisons of cecal OTUs at the genus level. Digesta samples were collected from pigs fed with the basal diet (Control) or inulin containing diet (Inulin). For 16S rRNA analysis, two digesta samples in each group were pooled (n = 5)
The overall bacterial diversity in the ileal and cecal digesta was estimated for each group. The alpha diversity metric, Abundance Coverage Estimator (ACE) was used to assess species richness and the Simpsons Index was used to assess species evenness [28, 29]. In addition to those specifically identified OTUs in each group, no significant difference was observed in overall bacterial diversity between the two groups (Table S3). The beta diversity also did not show significant difference between the two groups (Fig. S1). However, dietary INU supplementation significantly altered the relative abundance of intestinal bacteria at phylum level (Fig. 4). In this study, all the qualified seque3nces from ileal and cecal samples were assigned to 12 and 14 known phyla, respectively. Two phyla, Firmicutes (75.32%) and Proteobacteria (22.03%) were predominantly identified in the ileum of the control group. Interestingly, INU supplementation significantly decreased the abundance of the Proteobacteria (P < 0.01), but elevated the abundance of Bacteroidetes in the ileal samples (Fig. 4 and Table S4). In the cecal samples, Firmicutes (52.25%) and Bacteroidetes (43.58%) are the two predominant bacteria phyla identified in the control group. However, dietary INU supplementation tended to increase the relative abundance of the Bacteroidetes (P = 0.09).
Bar graph shows the phylum level composition of bacteria. Color-coded bar plot shows the relative abundance of bacterial phyla across the different samples. Digesta samples were collected from pigs fed with the basal diet (Control) or inulin containing diet (Inulin). For 16S rRNA analysis, two digesta samples in each group were pooled (n = 5)
At genus level, the sequences from the ileal and cecal samples were affiliated into 35 known genera (Fig. 5 and Table S5). Clostridiales (51.56%), Lactobacillus (4.31%), Actinobacillus (7.49%), and Enterobacteraceae (13.76%) were predominant genera identified in the ileal samples of the control group. However, INU supplementation significantly elevated the abundance of Lactobacillus (P < 0.05), and decreased the abundance of Actinobacillus and Enterobacteraceae (P < 0.05) in the ileal samples. As compared to the ileal samples, the distribution of the major genera identified in the cecal samples differed considerably (Fig. 5 and Table S5). The Actinobacillus and Enterobacteraceae, which were the two predominant genus in the ileal samples (more than 7%) were presented in extremely low abundance in the cecal samples of the control group. In contrast, the Prevotellaceae (16.56%) and Alloprevotella (14.58%) were two predominant genus identified in the cecal samples. Interestingly, INU supplementation decreased the abundance of Faecalibacterium, but significantly elevated the abundance of Campylobacter and Bacteroides in the cecal samples.
Heatmap distribution of OTUs. OTUs were arranged in rows and are clustered on the vertical axis. Samples are arranged vertically and are on the horizontal axis. Different colors indicate the relative abundance of taxons. Digesta samples were collected from pigs fed with the basal diet (Control) or inulin containing diet (Inulin). For 16S rRNA analysis, two digesta samples in each group were pooled (n = 5)
Influences of inulin on expression profiling of GPRs, HDACs, and inflammatory cytokines
As shown in Fig. 6, dietary INU supplementation significantly elevated the expression levels of GPR43, but decreased the expression levels of HDAC1 in the ileal and cecal mucosa (P < 0.05). Moreover, INU supplementation acutely elevated the expression level of GPR109A in the cecal mucosa (P < 0.01). The expression of TNFα was significantly down-regulated by INU in the ileal and cecal mucosa (P < 0.01). Interestingly, INU supplementation significantly elevated the expression level of IL-10 in the ileal mucosa (P < 0.05).
Influences of inulin on expression levels of critical genes related to intestinal barrier functions
Dietary INU supplementation significantly increased the expression levels of tight-junction protein ZO-1 both in the ileum and cecum (Fig. 7). However, INU had no significant influences on the expression levels of occluding and claudin-1. Dietary INU supplementation not only elevated the expression level of GLP-2 in the ileum, but also significantly elevated the expression levels of GLP-2 and ANG-4 in the cecum (P < 0.05).
Discussion
Gut microbiota provides the host with a broad range of functions such as digestion of complex dietary macronutrients, production of vitamins, defense against pathogens, and maintenance of the immune system [30]. Previous studies indicated that dysbiosis of the gut microbiota composition is associated with a variety of diseases, including metabolic disorders and inflammatory bowel disorder [30, 31]. Dietary fibers have long been looked as a stimulator of gastrointestinal development for mammalian animals. Recent studies on a variety animal species indicated that dietary fibers had dramatic influences on the gut microbiota [1, 32, 33]. In this study, we describe a synergetic response of the intestinal microbiota and epithelium to the increased intake of dietary fibers (inulin) in a porcine model. We show that dietary INU supplementation not only elevated the villus height and abundance of ZO-1 in the cecum, but also significantly reduced the number of apoptotic cells in the ileal and cecal mucosa. This is probably due in part to the decreased secretion of inflammatory cytokines (IL-6 and TNFα) in the intestinal mucosa. TNFα has been classified as homotrimeric transmembrane protein with a prominent role in systemic inflammation, which can participate in apoptosis through activating caspases [34]. The IL-6 was found to promote protein degradation via up-regulation of the E3 ubiquitin ligases [35]. In the present study, both the contents of TNFα and IL-6 in the ileal and cecal mucosa were significantly reduced after INU ingestion, indicating a beneficial role of INU in suppressing the mucosal inflammation.
As a soluble fiber, INU cannot be hydrolyzed by digestive enzymes in the small intestine, but at least partially hydrolyzed and fermented by intestinal microbes [36]. Evidence is accumulating to show that one of the mechanisms in which gut microbiota affects human health is its ability to produce a variety of beneficial metabolites that protect against diseases. The major products from the microbial fermentation in the gut are short chain fatty acids (SCFAs) such as the acetate, propionate, and butyrate [37]. However, when the fermentable fibers are in short supply, microbes switch to energetically less favorable sources for growth such as amino acids from dietary or endogenous proteins, or dietary fats, resulting in decreased activity of microbial fermentation and SCFAs production [38, 39]. In this study, INU supplementation significantly increased the acetate and butyrate contents in the cecal digesta, indicating an elevated fermentative activity of the microbiota. Moreover, the pH value of the cecal digesta tended to be decreased which may contribute to inhibition of harmful microbial fermentation in the intestine.
Previous study indicated that diets enriched in complex carbohydrates favor increased diversity of the gut microbiota [40]. In contrast, long-term intake of high-fat and high-sucrose diet can lead to extinction of several taxa of the gut microbiota [41]. In this study, we found that the cecal digesta had more 16S rRNA gene copy numbers and higher bacterial diversity than the ileal digesta, suggesting that the large intestine is the main site of microbial fermentation. This is also consistent with a well-known fact that dietary fibers are resistant to digestion in the small intestine, allowing passage largely intact into the large intestine. The Firmicutes and Proteobacteria are two dominant phyla identified in the ileum. The Proteobacteria are a major phylum of Gram-negative bacteria, which may increase upon ingestion of high-fat and high-protein diets [42]. Importantly, the phylum involves a number of pathogenic bacteria such as the Salmonella and Neisseriales. In this study, INU supplementation significantly decreased the abundance of the Proteobacteria in the ileum. In cecum, Firmicutes and Bacteroidetes are the two predominant bacteria phyla, and INU ingestion tended to increase the abundance of the Bacteroidetes. This is in agreement with the fact that most bacteria of this phylum have genes encoding hydrolytic enzymes to degrade non-starch carbohydrates [43]. Moreover, the result is also consistent with previous report that children consuming high-fiber diet showed a higher number of bacteria belonging to Bacteroidetes and Prevotella [44].
Lactobacillus (belonging to the Firmicutes phylum) has long been looked as a beneficial genus of the gut microbiota, since it serves as the major group of lactic acid producing bacteria [45]. In contrast, the Actinobacillus is a genus of Gram-negative bacteria, which has been found characteristically as commensals on the alimentary, respiratory, and genital mucosa. Importantly, some occur only as pathogens, while the commensal species are also found in a variety of lesions [46]. In the present study, INU ingestion significantly elevated the abundance of Lactobacillus in the ileum, but decreased the abundance of Actinobacillus in the ileum and cecum, indicating a beneficial role of INU in regulating of the intestinal microbiota. The Bacteroides represent a major constituent of the microflora in the large intestine, which contribute to complex polysaccharide breakdown and are one of the predominant producers of short chain fatty acids [47]. In the present study, INU ingestion significantly increased the abundance of Bacteroides spp., which may contribute to the elevated production of acetate and butyrate in the cecum.
Previous studies indicated that SCFAs regulated the intestinal health through two major signaling pathways [48, 49]. The first pathway involves epigenetic modulation. SCFAs, in particular the butyrate, has been identified as an intrinsic inhibitor of histone deacetylases (HDACs), promoting gene expression via the inhibition of the HDACs-induced deacetylation of lysine residues within histones [48]. Another pathway is signaling through the G-protein-coupled receptors (GPRs). For instance, activation of GPR43 and GPR109A activates the NLRP3 inflammasome, which is critical for intestinal homeostasis [49]. To explore the synergetic responses of intestinal epithelium to INU supplementation, we explored the expression profiles of critical GPRs and HDACs in the mucosa. We observed that INU ingestion elevated the expression levels of GPR43 in the ileum and cecum. This is agreement with the elevated acetate content, since the GPR43 has been looked as a major receptor for acetate [50]. Similarly, the expression level of butyrate-specific receptor, GPR109A was elevated in the cecum, which may contribute to the suppressed inflammatory responses in the cecum (i.e., reduced TNFα expression) [49]. Interestingly, INU ingestion significantly decreased the expression levels of HDAC1 in the ileal and cecal mucosa. HDAC1 is responsible for epigenetic alterations leading to modulation of chromatin structure and transcriptional regulation across lysine residues deacetylation on the N-terminal part of the core histones [51]. The decreased expression of HDAC1 was reported to be associated with the butyrate-suppressed inflammation [52]. Moreover, both the HDAC1 and HDAC2 were found to restrain the intestinal inflammatory response by regulating intestinal epithelial cell proliferation and differentiation [53].
Finally, we explored the expression levels of several critical genes related to intestinal barrier functions. Tight junction proteins, such as ZO-1, Occludin, and Claudin-1 form a complex structure in the paracellular space between two adjacent epithelial cells, which act as a selectively permeable barrier to prevent the invasion of intestinal bacteria and other antigens [54]. In the present study, the expression of ZO-1 was significantly upregulated in the ileal and cecal mucosa. Moreover, INU upregulated the expressions of GLP-2 and ANG-4 in the cecal mucosa. GLP-2 is an intestinotrophic hormone that promotes intestinal growth and proliferation though a series of downstream mediators [55]. ANG-4 participates in the formation of blood vessels and contributes to the development of a variety of tissues and organs [56]. Both these results suggested a beneficial role of dietary fiber supplementation on the intestinal barrier functions.
In summary, our results indicate a synergetic response of intestinal microbiota and epithelium to dietary INU supplementation. The SCFA-mediated GPR and HDAC co-regulation networks in the intestinal epithelium not only allows the animals to receive signaling from their resident microbiota, but also regulate a variety of physiological processes in the epithelium, especially growth and metabolism, and play critical roles in maintaining the epithelial integrity and suppressing intestinal inflammation. Our results will also facilitate understanding of the mechanisms behind the dietary fiber modulated gut health in monogastric animals.
Availability of data and material
All sequencing information has been deposited in the National Center for Biotechnology Information (NCBI) and can be accessed in the Short Read Archie (SRA) under the accession number PRJNA559763.
References
Hamaker BR, Tuncil YE (2014) A perspective on the complexity of dietary fiber structures and their potential effect on the gut microbiota. J Mol Biol 426:3838–3850
Kovatcheva-Datchary P, Nilsson A, Akrami R, Lee YS, De Vader F, Arora T, Hallen A, Martens E, Bjorck I, Backhed F (2015) Dietary fiber-induced improvement in glucose metabolism is associated with increased abundance of prevotella. Cell Metab 22:971–982
Florowska A, Krygier K, Florowski T, Dluzewska E (2016) Prebiotics as functional food ingredients preventing diet-related diseases. Food Funct 18:2147–2155
Kien CL, Blauwiekel R, Bunn JY, Jetton TL, Frankel WL, Holst JJ (2007) Cecal infusion of bytyrate increases intestinal cell proliferation in piglets. J Nutr 137:916–922
Gao Z, Yin J, Zhang J, Ward RE, Martin RJ, Lefevre M, Cefalu WT, Ye J (2009) Butyrate improves insulin sensitivity and increases energy expenditure in mice. Diabetes 58:1509–1517
Yamashita H, Fujisawa K, Ito E, Idei S, Kawaguchi N, Kimoto M, Hiemori M, Tsuji H (2007) Improvement of obesity and glucose tolerance by acetate in Type 2 diabetic Otsuka Long-Evans Tokushima fatty (OLETF) rats. Biosci Biochem 71:1236–1243
De Vadder F, Kovatcheva-Datchary P, Goncalves D, Vinera J, Zitoun C, Duchampt A, Backhed F, Mithieux G (2014) Microbiota-generated metabolites promote metabolic benefits via gut-brain neural circuits. Cell 156:84–96
Mensink MA, Frijlink HW, van der Voort MK (2015) Inulin, a flexible oligosaccharide Ӏ: review of its physicochemical characteristics. Carbohydr Polym 130:405–419
Niness KR (1406S) Inulin and oligofructose: what are they? J Nutr 129(Suppl. 7):1402S–1406S
Kalala G, Kambashi B, Everaert N, Beckers Y, Richel A, Pachikian B, Neyrinck AM, Delzenne NM, Bindelle J (2017) Characterization of frucatans and dietary fiber profiles in raw and steamed vegetables. Int J Food Sci Nutr 18:1–8
Femia AP, Luceri C, Dolara P, Giannini A, Biggeri A, Salvadori M, Clune Y, Collins KJ, Paglierani M, Caderni G (2002) Antitumorigenic activity of the prebiotic inulin enriched with oligofructose in combination with the probiotics Lactobacillus rhamnosus and Bifidobacterium lactis on azoxymethane-induced colon carcinogenesis in rats. Carcinogenesis 23:1953–1960
Delzenne NM, Daubioul C, Neyrinck A, Lasa M, Taper HS (2002) Inulin and oligofructose modulate lipid metabolism in animals: review of biochemical events and future prospects. Br J Nutr 87(Suppl. 2):S255–S259
Shukla G, Bhatia R, Sharma A (2016) Prebiotic inulin supplementation modulates the immune response and restores gut morphology in Giardia duodenalis-infected malnourished mice. Parasitol Res 115:4189–4198
Vandeputte D, Falony G, Vieira-Silva S, Wang J, Sailer M, Theis S, Verbeke K, Raes J (2017) Prebiotic inulin-type fructans induce specific changes in the human gut microbiota. Gut 66:1968–1974
Meurens F, Summerfield A, Nauwynck H, Saif L, Gerdts V (2012) The pig: a model for human infectious diseases. Trends Microbiol 20:50–57
Yasuda K, Maiorano R, Welch RM, Miller DD, Lei XG (2007) Cecum is the major degradation site of ingested inulin in young pigs. J Nutr 137:2399–2404
National Research Council (2012) Nutrient requirements for swine, 11th edn. National Academy Press, Washington, DC
AOAC (1990) Official methods of analysis, 15th edn. Association of Official Analytical Chemists, Washington, DC
Lillie RD (1969) Histopathologic technic and practical histochemistry, 3rd edn. McGraw-Hill Boock Co., New York
Franklin M, Mathew A, Vickers J, Clift R (2002) Characterization of microbial populations and volatile fatty acid concentrations in the jejunum, ileum, and cecum of pigs weaned at 17 vs 24 days of age. J Anim Sci 80:2904–2910
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Goron JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Loupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Bates ST, Berg-Lyons D, Caporaso JG, Walters WA, Knight R, Fierer N (2010) Examining the global distribution of dominant archaeal populations in soil. ISME J 5:908–917
Caporaso JG, Lauber CL, Walters WA, Berg-lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R, Gordon JI (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA 108(Suppl. 1):4516–4522
Magoc T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963
Edgar RC (2013) UPARSE: highly accurate OUT sequences from microbial amplicon reads. Nat Methods 10:996–998
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microb 73:5261–5267
Livak KJ, Schmittgen TD (2012) Analysis of relative gene expression data using real-time quantitative PCR and the 2 (− Delta Delta C (T)) method. Methods 25:402–408
Pitta DW, Pinchak E, Dowd SE, Osterstock J, Gontcharova V, Youn E, Dorton K, Yoon I, Min BR, Fulford JD, Wickersham TA, Malinowski DP (2010) Rumen bacterial diversity dynamics associated with changing from Bermuda grass hay to grazed winter wheat diets. Microb Ecol 59:511–522
Mahaffee WF, Kloepper JW (1997) Temporal changes in the bacterial communities of soil, rhizosphere, and endorhiza associated with filed-grown cucumber (Cucumis Sativus L.). Microb Ecol 34:210–223
Lattimer JM, Haub MD (2010) Effect of dietary fiber and its components on metabolic health. Nutrients 2:1266–1289
Kamada N, Seo SU, Chen GY, Nunez G (2013) Role of the gut microbiota in immunity and inflammatory disease. Nat Rev Immunol 13:321–335
Van den Abbeele P, Gerard P, Rabot S, Bruneau A, Aidy SEI, Derrien M, Kleerebezem M, Zoetendal EG, Smidt H, Verstrete W, Van de Wiele T, Possemiers S (2011) Arabinoxylans and inulin differentially modulate the mucosal and luminal gut microbiota and mucin degradation in humanized rats. Environ Microbiol 13:2667–2680
Tian G, Wu XY, Chen DW, Yu B, He J (2017) Adaptation of gut microbiome to different dietary nonstarch polysaccharide fractions in a porcine model. Mol Nutr Food Res 61:1700012
Gunther C, Neumann H, Neurath MF, Becker C (2013) Apoptosis, necrosis and necroptosis: cell death regulation in the intestinal epithelium. Gut 62:1062–1071
Argiles JM, Lopez-Soriamo FJ (1999) The role of cytokines in cancer cachexia. Med Res Rev 19:223–248
Roberfroid MB, Van Loo JA, Gibson GR (1998) The bifidogenic nature of chicory inulin and its hydrolysis products. J Nutr 128:11–19
Cummings JH, Pomare EW, Branch WJ, Naylor CP, Macfarlane GT (1987) Short chain fatty acids in human large intestine, portal, hepatic and venous blood. Gut 28:1221–1227
Wall R, Ross RP, Shanahan F, O’Mahony L, O’Mahony C, Coakley M, Hart O, Lawlor P, Quigley EM, Kiely B, Fitzgerald G, Stanton C (2009) Metabolic activity of the enteric microbiota influences the fatty acid composition of murine and porcine liver and adipose tissues. Am J Clin Nutr 89:1393–1401
Russel WR, Gratz SW, Duncan SH, Holtrop G, Ince J, Scobbie L, Duncan G, Johnstone AM, Lobley GE, Wallace RJ, Duthie GG, Flint HJ (2011) High-protein, reduced-carbohydrate weight-loss diets promote metabolite profiles likely to be detrimental to colonic health. Am J Clin Nutr 93:1062–1072
Sonnenburg ED, Smits SA, Tikhonov M, Higginbottom SK, Wingreen NS, Sonnenburg JL (2016) Diet-induced extinctions in the gut microbiota compound over generations. Nature 529:212–215
Schnorr SL, Candela M, Rampelli S, Centanni M, Consolandi BG, Turroni S, Biagi E, Peano C, Severgnini M, Fiori J, Gotti R, Bellis GD, Luiselli D, Brigidi P, Mabulla A, Marlowe F, Henry AG, Crittenden AN (2014) Gut microbiome of the Hadza hunter-gatherers. Nat Commun 5:3654
Madigan M, Martinko J (2005) Brock biology of microorganisms, 11th edn. Prentice Hall, New York
Dodd D, Mackie RI, Cann IKO (2011) Xylan degradation, a metabolic property shared by rumen and human colonic Bacteroidetes. Mol Microbiol 79:292–304
De Filippo C, Cavalieri D, Di Paola M, Poullet JB, Massart S, Collini S, Pieraccini G, Lionetti P (2010) Impact of diet in shaping gut microbiota revealed by a comparative study in children form Europe and rural Africa. Proc Natl Acad Sci USA 107:14691–14696
Khan M, Raoult D, Richet H, Lepidi H, La Scola B (2007) Growth promoting effects of single-dose intragastrically administered probiotics in chickens. Br Poult Sci 48:732–735
Guierrez CB, Rodriguez Barbosa JI, Suarez J, Gonzalez OR, Tascon RI, Rodriguez Ferri EF (1995) Efficacy of a variety of disinfectants against Actinobacillus pleuropneumoniae serotype 1. Am J Vet Res 56:1025–1029
Rey FE, Faith JJ, Bain J, Muehlbauer MJ, Stevens RD, Newgard CB, Gordon JI (2010) Dissecting the in vivo metabolic potential of two human gut acetogens. J Biol Chem 285:22082–22090
MacDonald VE, Howe LJ (2009) Histone acetylation: where to go and how to get there. Epigenetics 4:39–43
Macia L, Tan J, Vieira AT, Leach K, Stanley D, Luong S, Maruya M, Mkenzi CI, Hijikata A, Wong C, Binge L, Thorburn AN, Chevalier N, Ang C, Marino E, Robert R, Offermanns S, Teixeira MM, Moore RJ, Flavell RA, Fagarasan S, Mackay CR (2015) Metabolite-sensing receptors GPR43 and GPR109A facilitate dietary fibre-induced gut homeostasis through regulation of the inflammasome. Nat Commun 6:6734
Brown AJ, Goldsworthy SM, Bames AA, Eilert MM, Tcheang L, aniels D, Muir AI, Wigglesworth MJ, inghorn I, Fraser N, Pike NB, Strum JC, Steplewski KM, Murdock PR, Holder JC, Marshall FH, Szkeres PG, Wilson S, Ignar DM, Foord SM, Wise A, Dowell SJ (2003) The orphan G protein-coupled receptors GPR41 and GPR43 are activated by propionate and other short chain carboxylic acids. J Biol Chem 278: 11312–11319
Dovey OM, Foster CT, Cowley SM (2010) Histone deacetylase 1 (HDAC1), but not HDAC2, controls embryonic stem cell differentiation. Proc Natl Acad Sci USA 107:8242–8247
Zimmerman MA, Singh N, Martin PM, Thangaraju M, Ganapathy V, Waller JL, Shi H, Robertson K, Munn DH, Liu K (2012) Butyrate suppresses colonic inflammation through HDAC1-dependent Fas upregulation and Fas-mediated apoptosis of T cells. Am J Physiol Gastrointest Liver Physiol 302:1405–1415
Turgeon N, Blais M, Gaqne JM, Tardif V, Boudreau F, Asselin PN (2013) HDAC1 and HDAC2 restrain the intestinal inflammatory response by regulating intestinal epithelial cell differentiation. PLoS ONE 6:e73785
Ulluwishewa D, Anderson RC, McNabb WC, Moughan PJ, Wells JM, Roy NC (2011) Regulation of tight junction permeability by intestinal bacteria and dietary components. J Nutr 141:769–776
Walsh NA, Yusta B, DaCambra MP, Anini Y, Drucker DJ, Brubaker PL (2003) Glucagon-like peptide-2 receptor activation in the rat intestinal mucosa. Endocrinology 144:4385–4392
Kesler CT, Pereira ER, Cui CH, Nelson GM, Masuck DJ, Baish JW, Padera TP (2015) Aniopoietin-4 increases permeability of blood vessels and promotes lymphatic dilation. FASEB J 29:3668–3677
Acknowledgements
We thank Huifen Wang for technical help with the biochemical analysis. This study was supported by the National Natural Science Foundation of China (31972599) and the Development program of Sichuan Province (2018NZDZX0005).
Author information
Authors and Affiliations
Contributions
JH designed the experiments; WW and HX performed the animal trial and wrote the manuscript; PZ, JY, ZH, JL, and YL participated the biochemical assays; HY revised the manuscript. DC, BY, and XM conceived the experiment. All authors have read and approved the final draft.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethics approval
Studies involving animals were conducted according to the Regulations for the Administration of Affairs Concerning Experimental Animals (Ministry of Science and Technology, China, revised in June 2004). Sample collection was approved by the Institutional Animal Care and Use Committee of Sichuan Agricultural University, Sichuan, China (no. 20180901).
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
He, J., Xie, H., Chen, D. et al. Synergetic responses of intestinal microbiota and epithelium to dietary inulin supplementation in pigs. Eur J Nutr 60, 715–727 (2021). https://doi.org/10.1007/s00394-020-02284-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00394-020-02284-3