Abstract
Information on microbial biomass carbon (MBC) is crucial to assess their stocks and role for plant nutrient release in soil. Next to fumigation-extraction, molecular methods are routinely used to estimate the contribution of fungi, bacteria, and archaea to the soil microbial community. However, more information on the links between these different indices would deepen the understanding of microbial processes. The current study is based on 11 datasets, which contain MBC and MBN data obtained by fumigation-extraction and information on bacterial, archaeal, and fungal gene abundance, totalling 765 data points from agricultural, forest, and rangeland soils. Some of these datasets additionally provide information on double-stranded deoxyribonucleic acid (dsDNA) and fungal ergosterol. MBC varied around the median of 206 µg g−1 soil. MBN followed with a median MB-C/N ratio of 4.1. Median microbial gene abundance declined from bacteria (96 × 108) to archaea (4.4 × 108) to fungi (1.8 × 108). The median ratio of MBC/dsDNA was 15.8 and that of bacteria/dsDNA was 5.8 × 108 µg−1. The relationships between MBC and dsDNA as well as between bacterial gene abundance and dsDNA were both negatively affected by soil pH and positively by clay content. The median ergosterol/MBC and fungi/ergosterol ratios were 0.20% and 4.7 (n × 108 µg−1), respectively. The relationship between fungal gene abundance and ergosterol was negatively affected by soil pH and clay content. Our study suggests that combining fumigation-extraction with molecular tools allows more precise insights on the physiological interactions of soil microorganisms with their surrounding environment.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Soil microorganisms are key drivers of biogeochemical processes in soil and considered the active fraction of soil organic matter (SOM). The soil microbial biomass comprises C as well as macro- and micronutrients stored in living microorganisms (Hemkemeyer et al. 2021). Fungi and bacteria are the two dominant groups of the soil microbial biomass (Joergensen and Wichern 2008), but less abundant groups like archaea and protists are also important for soil functioning (Gattinger et al. 2002; Geisen et al. 2023). Most microorganisms in soil are dormant and do not grow as energy is limited (Jenkinson 1988; Joergensen and Wichern 2018). For this reason, the composition of the microbial groups in most soil is rather stable, comparing different soils (Joergensen and Wichern 2008) or seasons (Birgander et al. 2014). Changes in composition of the microbial groups require death and regrowth of microorganisms, decomposing their dead neighbours, which reduced the microbial biomass by 60–85% in comparison with the original level (Harden et al. 1993; Joergensen et al. 1990).
Microbial growth is nearly exclusively provided at the hot spots of energy availability, i.e., the rhizosphere around roots, the detritussphere around freshly incorporated plant residues and to some extent also the drilosphere around earthworm burrows (Banfield et al. 2018; Kuzyakov and Blagodatskaya 2015). Approximately 2 to 3% of soil microorganisms live in these areas where plenty of energy is available. However, most soil microorganisms inhabit areas of energy limitation and therefore starve, which results in dormancy. Nevertheless, dormant microorganisms also have important functions (Joergensen and Wichern 2018). They catabolize nutrient-containing organic components, leading to nitrogen, sulphur, and phosphorus mineralization. In addition, dormant microorganisms are an important reservoir for the maintenance of microbial biodiversity in soils.
Microbial biomass carbon (MBC) is a defined SOM fraction, which is relatively easy to quantify and to understand and can therefore be used in quantitative biogeochemical models. MBC data allow turnover calculations, which draws a quantitative relationship between MBC, microbial basal respiration, and soil organic C (SOC) (Anderson and Domsch 1989, 2010). Different approaches have been developed to estimate MBC, such as microscopic methods, substrate-induced activities, specific cell components, and fumigation methods (Jenkinson 1988; Joergensen and Wichern 2008; Kaiser et al. 1992), which all have specific advantages and disadvantages as discussed elsewhere (Martens 1995). The most common method of measuring MBC and microbial biomass nitrogen (MBN) is nowadays the fumigation-extraction (FE) method (Brookes et al. 1985; Vance et al. 1987), which gives access to the cytoplasm through cell-lysis. The advantages of this method are the clear separation between living and dead organisms, no discrimination of any microbial group, and the possibility to measure virtually all elements rendered extractable after fumigation (Khan et al. 2009; Schwalb et al. 2023a, b). The main disadvantage of the FE method is an insufficient difference in C concentration between fumigated and non-fumigated in low SOM environments, e.g., subsoils (Jörgensen et al. 2002) or desert soils (Wichern and Joergensen 2009).
Specific cell components are an important approach for measuring microbial biomass (Joergensen and Emmerling 2006; Joergensen and Wichern 2008), often with the advantage of giving additional information on major microbial groups, such as fungi, bacteria, and archaea. Cell-membrane components, such as PLFA and ergosterol (Joergensen 2022; Meyer et al. 2021), provide mainly information on the cell envelope but their quantity is affected by the cell volume and the number of organelles in eukaryotic fungi. Ergosterol is highly specific for the fungal phyla Ascomycota and Basidiomycota but has the advantage not to occur in plants or other organisms (Baldrian et al. 2013; Joergensen and Wichern 2008; Weete et al. 2010). Double-stranded deoxyribonucleic acid (dsDNA) gives information on the genomic core of a cell and is the basis for further molecular-genetic tools such as quantitative real-time polymerase chain reaction (qPCR), elucidating, inter alia, microbial diversity in soil (Hemkemeyer et al. 2014).
The main problem of utilising specific cell components for estimating soil microbial biomass is the absence of appropriate conversion values to biomass (Joergensen and Emmerling 2006). This is partly caused by their accumulation in dead organisms for a certain time (Joergensen and Wichern 2008) and their high variation in the biomass (Djajakirana et al. 1996; Jenkinson 1988). This is also true for molecular genetic tools to measure bacterial and archaeal (Stoddard et al. 2015) as well as fungal gene abundance (Baldrian et al. 2013; Heidrich and Beule 2022), which are increasingly used as routine approaches to estimate the contribution of these different groups to the soil microbial community (Meyer et al. 2021). The problem of many studies on the diversity of the soil microbiome is the determination of the relative and not of the absolute abundances by qPCR.
For unifying the view on soil microorganisms considering quantitative (MBC) and qualitative (microbial domains or diversity) data, the current study uses 11 datasets provided by the Sustainable Food Systems Research Centre at Rhine-Waal University of Applied Sciences in Kleve, Germany. All datasets contain MBC and MBN values obtained via FE and information on bacterial, archaeal, and fungal gene abundance. Some of these datasets additionally provide information on dsDNA and fungal ergosterol. With this set of data, we investigated the following three research questions: (1) What are the relationships between indices for total microbial biomass, i.e., MBC and dsDNA? (2) What are the relationships between dsDNA and the indices for prokaryotic microorganisms, i.e., bacterial and archaeal gene abundance? (3) What are the relationships between fungal indices, i.e., fungal gene abundance and ergosterol?
Materials and methods
Data sources
Data were acquired from 11 recent soil datasets: six derived from agricultural soils without additions: (1) Cover crops (Germany): experimental soil was taken at 0–30 cm in March 2016 from two fields: Neulouisendorf (52 m asl, 51°42′16″ N, 6°18′01″ E) and Bedburg-Hau (17 m asl, 51°45′53″ N, 6°11′17″ E). In both soils, ten different cover crops and six mixtures with six plants per pot were grown for 60 d at 70% water holding capacity in a greenhouse at an average of 22 °C (Hemkemeyer et al. unpublished data). (2) Neulouisendorf (Germany) soil samples were taken at 0–30 cm from 2016 until 2018 from a field experiment in Neulouisendorf (50 m asl, 51°42′07″ N, 6°18′18″ E) from two parts of the same field, which were in the crop rotational stage of carrying cover crops in 2016 and 2017, respectively. Besides fallows, there were seven species and five mixtures in 2016 and four species and two mixtures in 2017. Sampling took place in October/November of the same year and in March of the following year, with the latter differing in plots, which had either been harvested in autumn or mulched in winter (Hemkemeyer et al. unpublished data). (3) Pfalzdorf-2019 (Germany) soil samples, were taken at 0–30 cm from the Pfalzdorf (26 m asl, 51°42′15″ N, 6°09′41″ E) long-term intensive potato trial after six full three-year crop rotations in October 2019 (Hemkemeyer et al. 2024). (4) For Pfalzdorf-2015 (Germany) partly the same, partly different plots were already sampled in February 2015 at different stages in the crop rotation (Schwalb 2016, unpublished BSc thesis; partly published in Hemkemeyer et al. 2024).
(5) DOK (Switzerland) soil samples were collected at 0–20 cm depth in July and August 2019 from the long-term fertilization trial on arable land (Schwalb et al. 2023a), close to Therwil (307 m asl, 47°30′09.3″ N 7°32′21.6″ E), Switzerland, established in 1978 (Mäder et al. 2002). (6) Askov (Denmark) soil samples were taken at 0–20 cm depth in October 2019 from field B5 of the long-term experiment on arable land (Schwalb et al. 2024), established in 1894 and located at the Askov Experimental Station (63 m asl, 55°28’ N, 09°07’ E) in South Jutland, Denmark (Christensen et al. 2022).
Two datasets derived from non-agricultural soils: (7) Issyk-Kul (Kyrgyzstan) soil samples were taken in late September 2021 from natural vegetation at 0–30 cm depth around the Issyk-Kul lake (Iskakova et al. unpublished results), Kyrgyzstan, with sea buckthorn (Hippophae rhamnoides L.) and barberry (Berberis vulgaris L.) vegetation. Sampling sites were the eastern shore near Karakol (1726 m asl, 42°29′39″ N, 78°24′9″ E), the southern shore near Ton (1619 m asl, 42°33′59″ N, 78°16′49″ E), the northern shore near Korumdu (1629 m asl, 42°40′49″ N, 77°19′42″ E), and the western shore near Balykchy (1611 m asl, 42°27′35″ N, 76°14′2″ E). (8) Jalal-Abad (Kyrgyzstan) soil samples were collected at 0–30 and 30–60 cm depth in October 2019 close to Charbak (1000 m asl, 41°51′11″ N, 73°00′29″ E), Kyzyl-Unkur (1300 m asl, 41°23′31″, 73°03′40″ E), and Jay-Terek (1600 m asl, 41°17′16″ N, 72°53′03″ E) from natural, partly-managed walnut (Juglans regia L.) forests (Oskonbaeva et al. 2023).
The soils Pfalzdorf-2019, DOK, Askov, Issyk-Kul, and Jalal-Abad had been incubated at 22 °C for 14 or 28 d in the dark at 50% water holding capacity prior to analyses. Basic soil characteristics of all 11 soil datasets are presented in Table 1.
Finally, three datasets are derived from incubation experiments with agricultural soils, which received amendments in comparison with controls without amendments: (9) Salinity (Bangladesh) soil samples were taken after an incubation experiment with NaCl, rice (Oryza sativa L.) straw, and manure application treatments to investigate the salinity effects on C and N mineralization (Wichern et al. 2020). The paddy rice field soils were sampled in 2015 at 0–15 cm depth in Mymensingh (23 m asl) and Nalitabari (32 m asl), Bangladesh. (10) Frass (Germany) soil samples received two types of frass from black soldier fly (Hermetia illucens) larvae, differing in their feeding regime (Rummel et al. 2021). The arable soil was collected at Rotthalmünster, Germany (360 m asl, 48°21′39″ N, 13°11′33″ E). (11) Microplastics (Germany) soil samples were obtained from an incubation study with low density polyethylene and polypropylene additions, using soil samples collected at 0–15 cm depth from two arable fields in Kleve, (20 m asl, 50°08’ N, 6°02’ E), North Rhine-Westphalia, Germany (Blöcker et al. 2020).
Soil characteristics
Soils were sieved (< 2 mm) and air-dried before analysis. Soil pH was measured in a soil suspension with H2O (1:5 w/v) or 0.01 M CaCl2 (1:2 w/v), stirred in 5 min intervals and measured after 30 min. Soil pH-CaCl2 was converted to pH-H2O using the following equation: pH-H2O = (pH-CaCl2 + 0.373) / 0.923 according to Ahern et al. (1995). Total C and N were measured after milling by dry combustion at 900 °C using an elemental analyzer (Vario Max Cube CHN, Elementar, Langenselbold, Germany). Carbonate was measured gas-volumetrically after adding 10% HCl, using a Scheibler apparatus. SOC was calculated as the difference between total C and carbonate C.
Microbial biomass
MBC and MBN were determined in moist 20 g subsamples using the FE method (Brookes et al. 1985; Vance et al. 1987). Briefly, one 10 g portion at 50% water holding capacity was fumigated for 24 h at 25 °C with ethanol-free CHCl3. After CHCl3 removal, samples were extracted with 40 ml 0.5 M K2SO4 for 30 min by horizontal shaking at 200 rev min−1 and filtered (VWR 305, particle retention 2–3 μm). The other 10 g portion of non-fumigated soil was extracted similarly. Organic C and total N in the extracts were measured after combustion at 850 °C using an automated Multi N/C 2100S analyzer (Analytic Jena, Germany). MBC was EC/kEC, where EC = (organic C extracted from fumigated soils) − (organic C extracted from non-fumigated soils) and kEC = 0.45 (Joergensen 1996; Wu et al. 1990). MBN was EN/kEC, where EN = (total N extracted from fumigated soils)—(total N extracted from non-fumigated soils) and kEN = 0.54 (Brookes et al. 1985; Joergensen and Mueller 1996).
Ergosterol
The fungal cell-membrane component ergosterol was extracted for 30 min with 100 ml ethanol (96%) from a 2 g moist soil according to Djajakirana et al. (1996). Ergosterol was determined by reversed-phase HPLC (1260 Infinity, Agilent, Santa Clara, USA), using a C18 column and HPLC-grade methanol (100%) as liquid phase at a detection wavelength of 282 nm.
Double-stranded DNA
As a microbial biomass index (Bardelli et al. 2017), dsDNA was extracted from frozen soil samples using the FastDNA Spin Kit for Soil (MP Biomedicals, Santa Ana, USA) with modifications according to Hemkemeyer et al. (2014), which included bead-beating for 2 × 45 s at 6.5 m s−1, an additional washing step with 5.5 M guanidine thiocyanate to remove contaminants, and re-use of the eluate for a second elution step. Subsequently, dsDNA was quantified using the intercalating-dye system QuantiFluor (Promega, Mannheim, Germany) at 485 nm excitation and 520 nm emission in the microplate reader FLUOstar Omega (BMG Labtech, Ortenberg, Germany). Considering DNA loss during the extraction procedure, obtained dsDNA content and abundances of marker genes (see below) were corrected by dividing the data by the extraction efficiency as described in Hemkemeyer et al. (2024).
Microbial gene abundance
Quantification of microbial domains/kingdoms was done via quantitative real-time PCR (qPCR) targeting the Internal Transcribed Spacer 1 region (ITS1) for fungal quantification and the 16S rRNA gene for quantification of bacteria and archaea using the Light Cycler 480 SYBR Green I Master and Light Cycler 480 Probes Master, respectively, in a Light Cycler 480 Instrument II (Roche Diagnostics, Mannheim, Germany). Primers were NSI1 and 58A2R for fungi (Martin and Rygiewicz 2005), BAC338F and BAC805R in combination with the probe BAC516F for bacteria, and ARC787F and ARC1059R with probe ARC915F for archaea (Yu et al. 2005). Reaction mixtures and cycling conditions have been described by Wichern et al. (2020). Cloning fragments for qPCR standards originated from the following species Fusarium graminearum (fungi), Bacillus subtilis (bacteria), and Methanobacterium oryzae (archaea) and were used in serial dilutions: 107 – 101 copies µl−1 for fungi and archaea and 108 – 102 copies µl−1 for bacteria. Preparation of the standards has been described by Rummel et al. (2021).
Statistical analyses
The results presented in tables and figures are expressed on an oven-dry basis (about 24 h at 105 °C). Outliers of replicated data were removed according to the test proposed by Doerffel (1984). Normality was tested by the Shapiro–Wilk test and equal variance by the Levene test. All microbial data were log-transformed to normalize the distribution. The significance of differences between the datasets was tested by the Kruskal–Wallis One-way Analysis of Variance on Ranks, followed by Dunn’s pairwise multiple comparison procedure. Multiple regression models were calculated between MBC, bacterial gene abundance, and fungal gene abundance as dependent variables and dsDNA, ergosterol, clay content, and soil pH-H2O as independent variables. All regression models were tested for normality (Shapiro–Wilk), homogeneity of variance, absence of correlation between the residuals (Durban-Watson statistics) and absence of multi-collinearity, calculating the variance inflation factor (VIF), which never exceeded 4.0. All statistical analyses were performed using SigmaPlot 13.0 (Systat, San José, USA).
Results
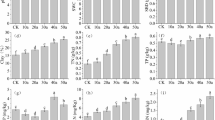
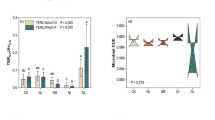
MBC varied in a sixfold range around the median of 206 µg g−1 soil across the 11 soil datasets (Table 2). MBN followed MBC with a median MB-C/N ratio of 4.1, ranging from 3.3 to 7.3. Median microbial gene abundance declined from bacteria (96 × 108), archaea (4.4 × 108) to fungi (1.8 × 108). The range between minimum and maximum number increased in the same order from 9-, 12- to 15-fold, respectively. In contrast to the MB-C/N ratio, the median bacteria/archaea (27) and especially the bacteria/fungi gene abundance ratio (57), showed similar or even larger variation.
The median dsDNA content was 15.7 µg g−1 soil in the 5 datasets (Table 3). The median ratio of MBC/dsDNA ratio was 15.8 and that of bacterial gene abundance-to-dsDNA was 5.8 × 108 µg−1 soil. The relationships between MBC and dsDNA (Table 4) as well as between bacterial gene abundance and dsDNA were both negatively affected by soil pH and positively by clay content (Table 4). This means that the ratios of MBC and bacterial gene abundance to dsDNA declined with increasing soil pH and decreasing clay content. Both soil properties were implemented into the multiple non-linear regression analysis to improve the prediction of MBC (Fig. 1a) and bacterial gene abundance (Fig. 1b) from dsDNA.
Relationships between measured and predicted (a) MBC and (b) bacterial gene abundance according to multiple regression analysis of dsDNA, soil pH, and clay content (Table 4), separated according to 5 datasets
The median ergosterol content was 0.42 µg g−1 soil in 7 datasets (Table 5), the median ergosterol/MBC and fungi/ergosterol ratios were 0.20% and 4.7 × 108 µg−1, respectively. The non-linear regression between fungal gene abundance and ergosterol was negatively affected by soil pH and clay content (Table 4, Fig. 2). This means that the ratios of fungal gene abundance-to-ergosterol declined with increasing soil pH and clay content. The correlation coefficients of ergosterol (r = 0.22) and fungal gene abundance (r = 0.26) with dsDNA were less close than those with archaeal (r = 0.52) and bacterial gene abundance (r = 0.65).
Relationship between measured and predicted fungal gene copies according to multiple regression analysis of ergosterol, soil pH, and clay content (Table 4), separated according to 7 datasets
Discussion
Total microbial biomass indices
In our study, MBC and MBN covered a quantitative range known for many soils, such as from arable soils (Kaiser et al. 1992; Nieder et al. 2008; Wardle 1998), rangeland (Joergensen 2010), and forest soils (Anderson and Joergensen 1997) under humid temperate but also under tropical climatic conditions (Joergensen 2010). MBC depends on the C input by root and shoot residues or litter fall but also on the environmental turnover conditions, i.e., temperature (Conant et al. 2011; Kätterer et al. 1998; Kirschbaum 2006), soil moisture (Faust et al. 2019; Lakshmi et al. 2003; Moyano et al. 2013), soil pH (Anderson and Domsch 1993; Lauber et al. 2008; Rousk et al. 2009), and clay content (Müller and Höper 2004; Wentzel et al. 2015).
The significant relationship between MBC and dsDNA (Fig. 1a) had marked deviations in specific datasets, e.g., caused by the low dsDNA content in the clayey and carbonaceous Kyrgyzstan soils. Clay content and soil pH have strong effects on the adsorption of organic compounds (Baldock and Skjemstad 2000; Schweizer et al. 2021), particularly phosphorus (P) containing compounds (Gérard 2016) such as MBP (Brookes et al. 1982) or adenosine triphosphate (ATP) (Jenkinson 1988). Consequently, high clay content and high soil pH might have lowered the extraction efficiency of dsDNA in these soils, emphasizing the need for extraction protocols adapted to specific soil conditions (Guerra et al. 2020). However, clay content and soil pH certainly also affect composition of microbial groups and their physiological status.
Conversion of dsDNA to MBC
The median MBC/dsDNA ratio of 15.8 observed in the current study falls within the range observed in other studies (Table 6). The weighted mean of these datasets from the literature is 15, clearly above the weighted mean of 6 proposed by Joergensen and Emmerling (2006). Also, the median MBC/dsDNA ratios observed in the current five datasets, ranging from 7.6 to 43.7, are like the means presented in Table 6, ranging from 2.2 to 38. Čapek et al. (2023) stated that the lower and upper slope bounds of the linear relationship between MBC and dsDNA can be expected within 3.5 to 22. The current relationship between MBC and dsDNA was significantly affected by clay content and soil pH. These two soil properties are the dominant factors, controlling the composition of microbial groups and their physiological status.
An increasing clay content promotes bacteria (Wentzel et al. 2015) and reduces microbial maintenance requirements (Filip et al. 1972; Höper and Kleefisch 2001). As the turnover rate is the product of maintenance × yield coefficient, lower maintenance requirements slow down the microbial turnover in soil (Joergensen and Wichern 2018; van Veen et al. 1984). Similarly, also a higher soil pH lowers the microbial demand for maintenance energy (Anderson and Domsch 1993, 2010). In addition, a high soil pH has positive effects on bacteria (Strickland and Rousk 2010) in agreement with the current results. These relationships largely explain clay and pH effects on the link between MBC and dsDNA, particularly as bacteria are the dominant and highly variable source of dsDNA (see discussion below).
The conversion of ATP (Jenkinson 1988) and substrate-induced respiration (SIR) (Anderson and Domsch 1978) to MBC are affected by the physiological activity of soil microorganisms. For this reason, ATP and SIR require a pre-incubation period after sieving (Jenkinson 1988). It is unknown, whether dsDNA measurements need a similar pretreatment if the data should be used as microbial biomass index.
Extracellular relic DNA (eDNA) from dead microorganisms is abundant in soil (Ascher et al. 2009; Levy-Booth et al. 2007) and contributes to the blurring of the relationship between MBC and dsDNA. Carini et al. (2016) examined eDNA in a wide range of soils, using the PCR viability based on the photoreactive DNA intercalating dye propidium monoazide. In their study, they found that on average 40% of both prokaryotic and fungal DNA was extracellular or from cells that were no longer intact. This value is in line with Gómez-Brandón et al. (2017), who observed that approximately 25% of dsDNA were not part of the microbial biomass. However, propidium monoazide may not work in every soil and provided sometimes only qualitative assessments of eDNA (Wang et al. 2021).
A largely unknown percentage of bias is introduced by the soil conditions at sampling time and the sample handling, e.g., dry matter determination, weighing, and pipetting. Also, different DNA extraction kits always showed significant differences (Fredricks et al. 2005; Zielińska et al. 2017). However, most studies (Table 6) used the FastDNA Spin Kit for Soil as extraction tool without systematic bias on the MBC/dsDNA ratios in comparison with other approaches. The datasets used in our study all implemented an additional washing step according to Hemkemeyer et al. (2014) to remove contaminants and considered DNA losses during the extraction as described by Hemkemeyer et al. (2024). Most studies used PicoGreen as dye (Table 6), again without apparent systematic differences to QuantiFluor, used in the current datasets, and Hoechst 33258.
MBC can be calculated from dsDNA, in sand and silt loams, using a conversion factor of 15, based on the current datasets and the weighted mean obtained by the literature survey (Table 6). Due to the large variation between the current datasets, we recommend checking this relationship for unknown soils by measuring both indices. In clayey and tropical soils, particularly with high contents of iron oxides (Huang et al. 2016), the calculated MBC data should be related to SOC as plausibility check.
Contribution of the main taxonomic groups to gene abundance and dsDNA
Bacteria contributed approximately 94% to the microbial marker gene abundance, archaea 4%, and fungi only 2%. These percentages agree with several other publications (e.g., Beule et al. 2019; Hartmann et al. 2015; Meyer et al. 2021; Tamez-Hidalgo et al. 2016). The close relationship between dsDNA and bacterial marker genes indicates that soil dsDNA is mainly of bacterial origin. Different reasons for the low contribution of fungi to the microbial gene abundance are known. The most important reason is the lower DNA concentration in the biomass of eukaryotic fungi compared to prokaryotic bacteria.
In the cultured marine species Cycloclasticus oligotrophus, Button and Robertson (2001) measured a bacterial biomass C-to-bacterial DNA ratio of 3.0 (µg µg−1), assuming 48% C in bacterial biomass dry weight. This ratio was considerably lower in starving bacteria in their natural environment. Starvation has been reported to decrease the DNA content of marine bacteria from 30 to 1 fg per cell (Moyer and Morita 1989), which finally leads to cytoplasm-less ghost particles (Hessenberger et al. 1996). This generally means the lower the gene copy numbers, the lower the microbial activity (Stoddard et al. 2015). In contrast to bacteria, Tellenbach et al. (2010) obtained a fungal biomass C-to-fungal DNA ratio (µg µg−1) for endophytic symbionts colonizing Norway spruce roots, which varied at least between 4,400 and 8,200, assuming 48% C in fungal biomass dry weight. Similarly, Baldrian et al. (2013) measured a mean fungal biomass C-to-fungal DNA ratio (µg µg−1) of 5,300, varying from 2,100 to 13,300 for different soil and litter colonizing fungi.
One reason for this strong variation is that fungi can contain up to several hundred copies of rRNA genes, which are interspaced by ITS sequences (Heidrich and Beule 2022) and vary by orders of magnitude across different fungal species (Lofgren et al. 2019). However, even among isolates of a single fungal species, copy numbers of 18S rRNA genes per genome can vary largely (Herrera et al. 2009; Lofgren et al. 2019; Zhao and Gibbons 2018). Nonetheless, the ratios of cell surface, cell volume, and genome are most likely more variable in laboratory cultures than in soil organisms, which has been shown for ATP (Contin et al. 2001; Dyckmans et al. 2003; Jenkinson 1988) and ergosterol (Djajakirana et al. 1996; Joergensen and Wichern 2008).
The thick and complex cell walls of fungi may result in poor release of DNA from the cells during extraction and after cell death (Fredricks et al. 2005; Tellenbach et al. 2010; Starke et al. 2019). In addition, the fungal DNA is densely packed in protein complexes in the nucleus (Galliano et al. 2021). However, according to the manufacturer, the FastDNA Spin Kit for Soil is also able to destroy even spores and, thus, presumably also fungal cell walls. Also, the friction by soil particles during bead beating may affect fungal mycelia stronger than prokaryotic cells.
Another reason for the low contribution of fungi to microbial gene abundance is that the primers NSI1 and 58A2R targeting ITS, used in the current datasets, are designed for Dikarya and are, thus, not representative for many fungal species from other phyla, especially Mucoromycota (Bonfante and Venice 2020). For example, arbuscular mycorrhizal fungi belonging to the Glomeromycota (Bodenhausen et al. 2021; Lekberg et al. 2018; Řezáčová et al. 2016; Victorino et al. 2020) can contribute 30% or more to the soil fungal biomass (Faust et al. 2017).
Based on amino sugar measurements, the contribution of fungal biomass to total soil microbial biomass was in most soils 75% and that of bacteria 25% (Joergensen and Wichern 2008), whereas archaea are not covered by this type of measurement. From the mean bacterial-to-archaeal gene abundance ratio, the biomass ratios of these prokaryotic microorganisms can be estimated. However, as the number of 16S rRNA gene copies per cell is 6.5 times lower for archaea (majority 1, ranging from 1–5) than for bacteria (majority 6–7, ranging from 1–21) according to the database rrnDB 5.8 (Stoddard et al. 2015), the bacterial-to-archaeal biomass ratio would decline from 23.5 to 3.6. In this case, fungi contribute on average approximately 70%, bacteria 23% and archaea 7% to the soil microbial biomass. This contribution of archaea would be markedly above the 2% proposed by Joergensen and Emmerling (2006), solely based on just one study by Gattinger et al. (2002), measuring phospholipid ether-lipids in soil.
Relationship between fungal indices
A median fungal gene abundance-to-ergosterol ratio of 4.7 × 108 µg−1 is close to the lower range Meyer et al. (2021) found in their soil (5.3 × 108 µg−1), but much higher compared to findings from sources other than soil (Table 7). These different comparisons of genome markers with the fungal cell membrane component ergosterol reveal that fungal cells are much larger in energy-rich litter, liquid cultures, and faeces as compared to C- and energy-limited soil ecosystems (Meyer et al. 2021).
The high ratio of fungal gene abundance-to-ergosterol after rice straw addition (Salinity, Wichern et al. 2020), indicates that the fungal cells remain small, although the organic amendment promoted fungal activity. The combination of low ergosterol-to-MBC ratio (Sardinha et al. 2003; Wichern et al. 2006) and high fungal gene-abundance-to-ergosterol indicates unfavorable conditions for soil fungi. In contrast, N-rich black soldier fly larvae frass application caused a stronger increase in ergosterol than in fungal gene abundance (Frass, Rummel et al. 2021), indicating that this organic amendment led to larger fungal cells. However, a higher number of comparisons between fungal gene abundance and ergosterol would help to strengthen this view, which is based on the current dataset.
Conclusions
Microbial biomass carbon (MBC) and double-stranded desoxyribonucleic acid (dsDNA) are closely related, but their ratio is not 42 and declined with increasing soil pH and decreasing clay content. MBC can be calculated from dsDNA, in sand and silt loams, using a conversion factor of 15. However, in clayey and tropical soils, the calculated MBC data should be related to soil organic C as plausibility check. Bacteria contribute 94%, to the total microbial gene abundance and are, thus, the main but highly variable source of dsDNA. The fungal gene abundance is significantly related to the fungal cell membrane component ergosterol. A low fungal gene abundance/ergosterol ratio indicates large fungal cells, and a high ratio the reverse. Due to group specific difference in gene concentration within the biomass, bacteria contribute 23%, archaea 7% and fungi 70% to MBC. The reasons for the variation between the different microbial indices and their respective ratios require further and stronger experimental explanations, which would allow more precise insights on the physiological changes of soil microorganisms in response to their surrounding environment. We, thus, recommend combining fumigation-extraction with cell-membrane components and molecular genetic tools to deepen our understanding of soil microbial communities and their involvement in biogeochemical cycles.
Data availability
The datasets analyzed in the current study are available from the corresponding author on reasonable request.
References
Agnelli A, Ascher J, Corti G, Ceccherini MT, Nannipieri P, Pietramellara G (2004) Distribution of microbial communities in a forest soil profile investigated by microbial biomass, soil respiration and DGGE of total and extracellular DNA. Soil Biol Biochem 36:859–868. https://doi.org/10.1016/j.soilbio.2004.02.004
Ahern CR, Baker DE, Aitken RL (1995) Models for relating pH measurements in water and calcium chloride for a wide range of pH, soil types and depths. Plant Soil 171:47–52. https://doi.org/10.1007/BF00009563
Anderson JPE, Domsch KH (1978) A physiological method for the quantitative measurement of microbial biomass in soil. Soil Biol Biochem 10:215–221. https://doi.org/10.1016/0038-0717(78)90099-8
Anderson TH, Domsch KH (1989) Ratios of microbial biomass carbon to total organic carbon in arable soils. Soil Biol Biochem 21:471–479. https://doi.org/10.1016/0038-0717(89)90117-X
Anderson TH, Domsch KH (1993) The metabolic quotient for CO2 (qCO2) as a specific activity parameter to assess the effects of environmental conditions, such as pH, on the microbial biomass of forest soils. Soil Biol Biochem 25:393–395. https://doi.org/10.1016/0038-0717(93)90140-7
Anderson TH, Domsch KH (2010) Soil microbial biomass: the eco-physiological approach. Soil Biol Biochem 42:2039–2043. https://doi.org/10.1016/j.soilbio.2010.06.026
Anderson TH, Joergensen RG (1997) Relationship between SIR and FE estimates of microbial biomass C in deciduous forest soils at different pH. Soil Biol Biochem 29:1033–1042. https://doi.org/10.1016/S0038-0717(97)00011-4
Anderson TH, Martens R (2013) DNA determinations during growth of soil microbial biomasses. Soil Biol Biochem 57:487–495. https://doi.org/10.1016/j.soilbio.2012.09.031
Ascher J, Ceccherini MT, Pantani OL, Agnelli A, Borgogni F, Guerri G, Nannipieri P, Pietramellara G (2009) Sequential extraction and genetic fingerprinting of a forest soil metagenome. Appl Soil Ecol 42:176–181. https://doi.org/10.1016/j.apsoil.2009.03.005
Baldock JA, Skjemstad JO (2000) Role of the soil matrix and minerals in protecting natural organic materials against biological attack. Org Geochem 31:697–710. https://doi.org/10.1016/S0146-6380(00)00049-8
Baldrian P, Vĕtrovský T, Cajthaml T, Dobišásová P, Petránková M, Šnajdr J, Eichlerová I (2013) Estimation of fungal biomass in forest litter and soil. Fungal Ecol 6:1–11. https://doi.org/10.1016/j.funeco.2012.10.002
Banfield CC, Pausch J, Kuzyakov Y, Dippold MA (2018) Microbial processing of plant residues in the subsoil – the role of biopores. Soil Biol Biochem 125:309–318. https://doi.org/10.1016/j.soilbio.2018.08.004
Bardelli T, Gomez-Brandon M, Ascher-Jenull J, Fornasier F, Arfaioli P, Francioli D, Egli M, Sartori G, Insam H, Pietramellara G (2017) Effects of slope exposure on soil physico-chemical and microbiological properties along an altitudinal climosequence in the Italian Alps. Sci Total Environ 575:1041–1055. https://doi.org/10.1016/j.scitotenv.2016.09.176
Bardelli T, Ascher-Jenull J, Stocker EB, Fornasier F, Arfaioli P, Fravolini G, Alves Medeiros LR, Egli M, Pietramellara G, Insam H, Gómez-Brandón M (2018) Impact of slope exposure on chemical and microbiological properties of Norway spruce deadwood and underlying soil during early stages of decomposition in the Italian Alps. Catena 167:100–115. https://doi.org/10.1016/j.catena.2018.04.031
Beule L, Corre MD, Schmidt M, Gobel L, Veldkamp E, Karlovsky P (2019) Conversion of monoculture cropland and open grassland to agroforestry alters the abundance of soil bacteria, fungi and soil-N-cycling genes. PLoS ONE 14:e0218779. https://doi.org/10.1371/journal.pone.0220713
Bhople P, Djukic I, Keiblinger K, Zehetner F, Liu D, Bierbaumer M, Zechmeister-Boltenstern S, Joergensen RG, Murugan R (2019) Variations in soil and microbial biomass C, N and fungal biomass ergosterol along elevation and depth gradients in Alpine ecosystems. Geoderma 345:93–103. https://doi.org/10.1016/j.geoderma.2019.03.022
Birgander J, Rousk J, Olsson PA (2014) Comparison of fertility and seasonal effects on grassland microbial communities. Soil Biol Biochem 76:80–89. https://doi.org/10.1016/j.soilbio.2014.05.007
Blagodatskaya EV, Blagodatsky SA, Anderson TH (2003) Quantitative isolation of microbial DNA from different types of soil of natural and agricultural ecosystems. Microbiol 72:744–749. https://doi.org/10.1023/B:MICI.0000008379.63620.7b
Blöcker L, Watson C, Wichern F (2020) Living in the plastic age - Different short-term microbial response to microplastics addition to arable soils with contrasting soil organic matter content and farm management legacy. Environ Poll 267:115468. https://doi.org/10.1016/j.envpol.2020.115468
Bodenhausen N, Deslandes-Herold G, Waelchli J, Held A, van der Heijden MGA, Schlaeppi K (2021) Relative qPCR to quantify colonization of plant roots by arbuscular mycorrhizal fungi. Mycorrhiza 31:137–148. https://doi.org/10.1007/s00572-020-01014-1
Bonfante P, Venice F (2020) Mucoromycota: going to the roots of plant-interacting fungi. Fungal Biol Rev 34:100–113. https://doi.org/10.1016/j.fbr.2019.12.003
Bragato G, Fornasier F, Brus DJ (2016) Characterization of soil fertility and soil biodiversity with dsDNA as a covariate in a regression estimator for mean microbial biomass C. Eur J Soil Sci 67:827–834. https://doi.org/10.1111/ejss.12387
Brookes PC, Powlson DS, Jenkinson DS (1982) Measurement of microbial biomass phosphorus in soil. Soil Biol Biochem 14:319–329. https://doi.org/10.1016/0038-0717(82)90001-3
Brookes PC, Landman A, Pruden G, Jenkinson DS (1985) Chloroform fumigation and the release of soil nitrogen: a rapid direct extraction method to measure microbial biomass nitrogen in soil. Soil Biol Biochem 17:837–842. https://doi.org/10.1016/0038-0717(85)90144-0
Button DK, Robertson BR (2001) Determination of DNA content of aquatic bacteria by flow cytometry. Appl Environ Microbiol 67:1636–1645. https://doi.org/10.1128/AEM.67.4.1636-1645.2001
Čapek P, Choma M, Kaštovská E, Tahovská K, Glanville HC, Šantrůčková H (2023) Revisiting soil microbial biomass: considering changes in composition with growth rate. Soil Biol Biochem 184:109103. https://doi.org/10.1016/j.soilbio.2023.109103
Carini P, Marsden PJ, Leff JW, Morgan EE, Strickland MS, Fierer N (2016) Relic DNA is abundant in soil and obscures estimates of soil microbial diversity. Nat Microbiol 2:16242. https://doi.org/10.1038/nmicrobiol.2016.242
Chernysheva EV, Fornasier F, Borisov AV (2023) Factors for conversion of the content of double-stranded DNA to carbon of soil microbial biomass. Eurasian Soil Sci 56:672–681. https://doi.org/10.1134/S1064229323600021
Christensen BT, Thomsen IK, Eriksen J (2022) The Askov long-term field experiment (1894–2021) represents a unique research platform. J Plant Nutr Soil Sci 185:187–201. https://doi.org/10.1002/jpln.202100354
Conant RT, Ryan MG, Ågren GI, Birge HE, Davidson EA, Eliasson PE, Evans SE, Frey SD, Giardina CP, Hopkins FM, Hyvönen R, Kirschbaum MUF, Lavallee JM, Leifeld J, Parton WJ, Steinweg JM, Wallenstein MD, Wetterstedt JAM, Bradford MA (2011) Temperature and soil organic matter decomposition rates - synthesis of current knowledge and a way forward. Glob Chang Biol 17:3392–3404. https://doi.org/10.1111/j.1365-2486.2011.02496.x
Contin M, Todd A, Brookes PC (2001) The ATP concentration in the soil microbial biomass. Soil Biol Biochem 33:701–704. https://doi.org/10.1016/0038-0717(79)90012-9
Djajakirana G, Joergensen RG, Meyer B (1996) Ergosterol and microbial biomass relationship in soil. Biol Fertil Soils 22:299–304. https://doi.org/10.1007/BF00334573
Doerffel K (1984) Statistik in der analytischen chemie, 3rd edn. Verlag Chemie, Weinheim
Dyckmans J, Chander K, Joergensen RG, Priess J, Raubuch M, Sehy U (2003) Adenylates as an estimate of microbial biomass C in different soil groups. Soil Biol Biochem 35:1485–1491. https://doi.org/10.1016/S0038-0717(03)00245-1
Faust S, Heinze S, Ngosong C, Sradnick A, Oltmanns M, Raupp J, Geisseler D, Joergensen RG (2017) Effect of biodynamic soil amendments on microbial communities in comparison with inorganic fertilization. Appl Soil Ecol 114:82–89. https://doi.org/10.1016/j.apsoil.2017.03.006
Faust S, Koch HJ, Joergensen RG (2019) Respiration response to different tillage intensities in transplanted soil columns. Geoderma 352:289–297. https://doi.org/10.1016/j.geoderma.2019.05.023
Filip Z, Haider K, Martin JP (1972) Influence of clay minerals on growth and metabolic activity of Epicoccum nigrum and Stachybotrys chartarum. Soil Biol Biochem 4:135–145. https://doi.org/10.1016/0038-0717(72)90004-1
Fornasier F, Ascher J, Ceccherini MT, Tomat E, Pietramellara G (2014) A simplified rapid, low-cost and versatile DNA-based assessment of soil microbial biomass. Ecol Indic 45:75–82. https://doi.org/10.1016/j.ecolind.2014.03.028
Fredricks DN, Smith C, Meier A (2005) Comparison of six DNA extraction methods for recovery of fungal DNA as assessed by quantitative PCR. J Clin Microbiol 43:5122–5128. https://doi.org/10.1128/JCM.43.10.5122-5128.2005
Galliano I, Dapr V, Zaniol E, Alliaudi C, Graziano E, Montanari P, Calvi C, Bergallo M (2021) Comparison of methods for isolating fungal DNA. Pract Lab Medic 25:e00221. https://doi.org/10.1016/j.plabm.2021.e00221
Gangneux C, Akpa-Vincesla M, Sauvage H, Desaire S, Houot S, Laval K (2011) Fungal, bacterial and plant dsDNA contributions to soil total DNA extracted from silty soils under different farming practices: relationships with chloroform-labile carbon. Soil Biol Biochem 43:431–437. https://doi.org/10.1016/j.soilbio.2010.11.012
Gattinger A, Ruser R, Schloter M, Munch JC (2002) Microbial community structure varies in different soil zones of a potato field. J Plant Nutr Soil Sci 165:421–428. https://doi.org/10.1002/1522-2624(200208)165:4%3C421::AID-JPLN421%3E3.0.CO;2-N
Geisen S, Lara E, Mitchell E (2023) Contemporary issues, current best practice and ways forward in soil protist ecology. Mol Ecol Resour:1–11. https://doi.org/10.1111/1755-0998.13819
Gérard F (2016) Clay minerals, iron/aluminum oxides, and their contribution to phosphate sorption in soils - a myth revisited. Geoderma 262:213–226. https://doi.org/10.1016/j.geoderma.2015.08.036
Gómez-Brandón M, Ascher-Jenull J, Bardelli T, Fornasier F, Sartori G, Pietramellara G, Arfaioli P, Markus Egli M, Beylich A, Insam H, Graefe U (2017) Ground cover and slope exposure effects on micro- and mesobiota in forest soils. Ecol Indic 80:174–185. https://doi.org/10.1016/j.ecolind.2017.05.032
Gong P, Guan X, Witter E (2001) A rapid method to extract ergosterol from soil by physical disruption. Appl Soil Ecol 17:285–289. https://doi.org/10.1016/S0929-1393(01)00141-X
Gong H, Du Q, Xie S, Hu W, Akram MA, Hou Q, Dong L, Sun Y, Manan A, Deng Y, Ran J, Deng J (2021) Soil microbial DNA concentration is a powerful indicator for estimating soil microbial biomass C and N across arid and semi-arid regions in northern China. Appl Soil Ecol 160:103869. https://doi.org/10.1016/j.apsoil.2020.103869
Guerra V, Beule L, Lehtsaar E, Liao HL, Karlovsky P (2020) Improved protocol for DNA extraction from subsoils using phosphate lysis buffer. Microorganisms 8:532. https://doi.org/10.3390/microorganisms8040532
Harden T, Joergensen RG, Meyer B, Wolters V (1993) Mineralization of straw and formation of soil microbial biomass in a soil treated with simazine and dinoterb. Soil Biol Biochem 25:1273–1276. https://doi.org/10.1016/0038-0717(93)90224-Y
Hartmann M, Frey B, Mayer J, Mäder P, Widmer F (2015) Distinct soil microbial diversity under long-term organic and conventional farming. ISME J 9:1177–1194. https://doi.org/10.1038/ismej.2014.210
Heidrich V, Beule L (2022) Are short-read amplicons suitable for the prediction of microbiome functional potential? A critical perspective. iMeta 1:e38. https://doi.org/10.1002/imt2.38
Hemkemeyer M, Pronk GJ, Heister K, Kögel-Knabner I, Martens R, Tebbe CC (2014) Artificial soil studies reveal domain-specific preferences of microorganisms for the colonisation of different soil minerals and particle size fractions. FEMS Microbiol Ecol 90:770–782. https://doi.org/10.1111/1574-6941.12436
Hemkemeyer M, Schwalb SA, Heinze S, Joergensen RG, Wichern F (2021) Functions of elements in soil microorganisms. Microbiol Res 252:126832. https://doi.org/10.1016/j.micres.2021.126832
Hemkemeyer M, Schwalb SA, Berendonk C, Geisen S, Heinze S, Joergensen RG, Li R, Lövenich P, Xiong W, Wichern F (2024) Potato yield and quality are linked to cover crop and soil microbiome, respectively – evidence from a long-term trial. Biol Fertil Soils. (re-submitted)
Herrera ML, Vallor AC, Gelfond JA, Patterson TF, Wickes BL (2009) Strain-dependent variation in 18S ribosomal DNA copy numbers in Aspergillus fumigatus. J Clin Microbiol 47:1325–1332. https://doi.org/10.1128/JCM.02073-08
Hessenberger A, Leppard GG, Herndl GJ (1996) Relationship between the intracellular integrity and the morphology of the capsular envelope in attached and free-living marine bacteria. Appl Environ Microbiol 62:4521–4528. https://doi.org/10.1128/aem.62.12.4521-4528.1996
Höper H, Kleefisch B (2001) Untersuchung bodenbiologischer Parameter im Rahmen der Boden-Dauerbeobachtung in Niedersachsen– Bodenbiologische Referenzwerte und Zeitreihen. Arbeitshefte Boden 2001/4, NLfB, Hannover
Huang YT, Lowe DJ, Zhang H, Cursons R, Young JM, Churchman GJ, Schipper LS, Rawlence NJ, Wood JR, Cooper A (2016) A new method to extract and purify DNA from allophanic soils and paleosols, and potential for paleoenvironmental reconstruction and other applications. Geoderma 274:114–125. https://doi.org/10.1016/j.geoderma.2016.04.003
IUSS Working Group WRB (2022) World reference base for soil resources. International soil classification system for naming soils and creating legends for soil maps, 4th edn. International Union of Soil Sciences (IUSS), Vienna
Jenkinson DS (1988) The determination of microbial biomass carbon and nitrogen in soil. In: Wilson JR (ed) Advances in nitrogen cycling in agricultural ecosystems. CABI, Wallingford, pp 368–386
Joergensen RG (1996) The fumigation-extraction method to estimate soil microbial biomass: calibration of the kEC value. Soil Biol Biochem 28:25–31. https://doi.org/10.1016/0038-0717(95)00102-6
Joergensen RG (2010) Organic matter and micro-organisms in tropical soils. In: Dion P (ed) Soil biology and agriculture in the tropics. Springer, Berlin, pp 17–44. https://doi.org/10.1007/978-3-642-05076-3_2
Joergensen RG (2022) Phospholipid fatty acids in soil – drawbacks and future prospects. Biol Fertil Soil 58:1–6. https://doi.org/10.1007/s00374-021-01613-w
Joergensen RG, Emmerling C (2006) Methods for evaluating human impact on soil microorganisms based on their activity, biomass, and diversity in agricultural soils. J Plant Nutr Soil Sci 169:295–309. https://doi.org/10.1002/jpln.200521941
Joergensen RG, Mueller T (1996) The fumigation-extraction method to estimate soil microbial biomass: calibration of the kEN value. Soil Biol Biochem 28:33–37. https://doi.org/10.1016/0038-0717(95)00101-8
Joergensen RG, Wichern F (2008) Quantitative assessment of the fungal contribution to microbial tissue in soil. Soil Biol Biochem 40:2977–2991. https://doi.org/10.1016/j.soilbio.2008.08.017
Joergensen RG, Wichern F (2018) Alive and kicking: why dormant soil microorganisms matter. Soil Biol Biochem 116:419–430. https://doi.org/10.1016/j.soilbio.2017.10.022
Joergensen RG, Brookes PC, Jenkinson DS (1990) Survival of the soil microbial biomass at elevated temperatures. Soil Biol Biochem 22:1129–1136. https://doi.org/10.1016/0038-0717(90)90039-3
Jörgensen RG, Raubuch M, Brandt M (2002) Soil microbial properties down the profile of a black earth buried by colluvium. J Plant Nutr Soil Sci 165:274–280. https://doi.org/10.1002/1522-2624(200206)165:3%3C274::AID-JPLN274%3E3.0.CO;2-2
Kaiser EA, Mueller T, Joergensen RG, Insam H, Heinemeyer O (1992) Evaluation of methods to estimate the soil microbial biomass and the relationship with soil texture and organic matter. Soil Biol Biochem 24:675–683. https://doi.org/10.1016/0038-0717(92)90046-Z
Kätterer T, Reichstein M, Andren O, Lomander A (1998) Temperature dependence of organic matter decomposition: a critical review using literature data analyzed with different models. Biol Fertil Soils 27:258–262. https://doi.org/10.1007/s003740050430
Khan KS, Heinze S, Joergensen RG (2009) Simultaneous measurement of S, macronutrients, and heavy metals in the soil microbial biomass with CHCl3 fumigation and NH4NO3 extraction. Soil Biol Biochem 41:309–314. https://doi.org/10.1016/j.soilbio.2008.11.001
Kirschbaum MUF (2006) The temperature dependence of organic matter decomposition - still a topic of debate. Soil Biol Biochem 38:2510–2518. https://doi.org/10.1016/j.soilbio.2006.01.030
Kuzyakov Y, Blagodatskaya E (2015) Microbial hotspots and hot moments in soil: concept & review. Soil Biol Biochem 83:184–199. https://doi.org/10.1016/j.soilbio.2015.01.025
Lakshmi V, Jackson TJ, Zehrfuhs D (2003) Soil moisture-temperature relationships: results from two field experiments. Hydrol Proced 17:3041–3057. https://doi.org/10.1002/hyp.1275
Lauber CL, Strickland MS, Bradford MA, Fierer N (2008) The influence of soil properties on the structure of bacterial and fungal communities across land-use types. Soil Biol Biochem 40:2407–2415. https://doi.org/10.1016/j.soilbio.2008.05.021
Leckie SE, Prescott CE, Grayston SJ, Neufeld JD, Mohn WW (2004) Comparison of chloroform fumigation-extraction, phospholipid fatty acid, and DNA methods to determine microbial biomass in forest humus. Soil Biol Biochem 36:529–532. https://doi.org/10.1016/j.soilbio.2003.10.014
Lekberg Y, Vasar M, Bullington LS, Sepp SK, Antunes PM, Bunn R, Larkin BG, Öpik M (2018) More bang for the buck? Can arbuscular mycorrhizal fungal communities be characterized adequately alongside other fungi using general fungal primers? New Phytol 220:971–976. https://doi.org/10.1111/nph.15035
Levy-Booth DJ, Campbell RG, Gulden RH, Hart MM, Powell JR, Klironomos JN, Pauls KP, Swanton CJ, Trevors JT, Dunfield KE (2007) Cycling of extracellular DNA in the soil environment. Soil Biol Biochem 39:2977–2991. https://doi.org/10.1016/j.soilbio.2007.06.020
Lloyd-Jones G, Hunter DWF (2001) Comparison of rapid DNA extraction methods applied to contrasting New Zealand soils. Soil Biol Biochem 33:2053–2059. https://doi.org/10.1016/S0038-0717(01)00133-X
Loeppmann S, Semenov M, Kuzyakov Y, Evgeniya Blagodatskaya E (2018) Shift from dormancy to microbial growth revealed by RNA:DNA ratio. Ecol Indic 85:603–612. https://doi.org/10.1016/j.ecolind.2017.11.020
Lofgren LA, Uehling JK, Branco S, Bruns TD, Martin F, Kennedy PG (2019) Genome-based estimates of fungal rDNA copy number variation across phylogenetic scales and ecological lifestyles. Molec Ecol 28:721–730. https://doi.org/10.1111/mec.14995
Mäder P, Fließbach A, Dubois D, Gunst L, Fried P, Niggli U (2002) Soil fertility and biodiversity in organic farming. Science 296:1694–1697. https://doi.org/10.1126/science.1071148
Marstorp H, Witter E (1999) Extractable dsDNA and product formation as measures of microbial growth in soil upon substrate addition. Soil Biol Biochem 31:1443–1453. https://doi.org/10.1016/S0038-0717(99)00065-6
Marstorp H, Guan X, Gong P (2000) Relationship between dsDNA, chloroform labile C and ergosterol in soils of different organic matter contents and pH. Soil Biol Biochem 32:879–882. https://doi.org/10.1016/S0038-0717(99)00210-2
Martens R (1995) Current methods for measuring microbial biomass C in soil: potentials and limitations. Biol Fertil Soils 19:87–99. https://doi.org/10.1007/BF00336142
Martin KJ, Rygiewicz PT (2005) Fungal-specific PCR primers developed for analysis of the ITS region of environmental DNA extracts. BMC Microbiol 5:28. https://doi.org/10.1186/1471-2180-5-28
Meyer S, Grüning MM, Beule L, Karlovsky P, Joergensen RG, Sundrum A (2021) Soil N2O flux and nitrification and denitrification gene responses to feed-induced differences in the composition of dairy cow faeces. Biol Fertil Soils 57:767–779. https://doi.org/10.1007/s00374-021-01566-0
Moyano FE, Manzoni S, Chenu C (2013) Responses of soil heterotrophic respiration to moisture availability: an exploration of processes and models. Soil Biol Biochem 59:72–85. https://doi.org/10.1016/j.soilbio.2013.01.002
Moyer CL, Morita RY (1989) Effect of growth rate and starvation survival on cellular DNA, RNA, and protein of a psychrophilic marine bacterium. Appl Environ Microbiol 55:2710–2716. https://doi.org/10.1128/aem.55.10.2710-2716.1989
Müller T, Höper H (2004) Soil organic matter turnover as a function of the soil clay content: consequences for model applications. Soil Biol Biochem 36:877–888. https://doi.org/10.1016/j.soilbio.2003.12.015
Nieder R, Harden T, Martens R, Benbi DK (2008) Microbial biomass in arable soils of Germany during the growth period of annual crops. J Plant Nutr Soil Sci 171:878–885. https://doi.org/10.1002/jpln.200700024
Oskonbaeva Z, Maitykov T, Schwalb SA, Joergensen RG, Wichern F (2023) No evidence of an elevation effect on soil microbial properties in a walnut-fruit forest in Kyrgyzstan. J Soil Sci Plant Nutr 23:2662–2672. https://doi.org/10.1007/s42729-023-01222-6
Řezáčová V, Gryndler M, Bukovská P, Šmilauer P, Jansa J (2016) Molecular community analysis of arbuscular mycorrhizal fungi - Contributions of PCR primer and host plant selectivity to the detected community profiles. Pedobiol 59:179–187. https://doi.org/10.1016/j.pedobi.2016.04.002
Rousk J, Brookes PC, Bååth E (2009) Contrasting soil pH effects on fungal and bacterial growth suggest functional redundancy in carbon mineralization. Appl Environ Microbiol 75:1589–1596. https://doi.org/10.1128/AEM.02775-08
Rummel PS, Beule L, Hemkemeyer M, Schwalb SA, Wichern F (2021) Black soldier fly diet impacts soil greenhouse gas emissions from frass applied as fertilizer. Front Sustain Food Syst 5:709993. https://doi.org/10.3389/fsufs.2021.709993
Sardinha M, Müller T, Schmeisky H, Joergensen RG (2003) Microbial performance in a temperate floodplain soil along a salinity gradient. Appl Soil Ecol 23:237–244. https://doi.org/10.1016/S0929-1393(03)00027-1
Schwalb SA, Li S, Hemkemeyer M, Heinze S, Joergensen RG, Mayer J, Mäder P, Wichern F (2023a) Long-term differences in fertilisation type change the bacteria:archaea:fungi ratios and reveal a heterogeneous response of the soil microbial ionome. Soil Biol Biochem 177:108892. https://doi.org/10.1016/j.soilbio.2022.108892
Schwalb SA, Khan KS, Hemkemeyer M, Heinze S, Oskonbaeva Z, Joergensen RG, Wichern F (2023b) Chloroform-labile trace elements in soil via fumigation-extraction: steps towards the soil microbial ionome beyond C:N:P. Eur J Soil Sci 74:e13356. https://doi.org/10.1111/ejss.13356
Schwalb SA, Hemkemeyer M, Christensen BT, Heinze S, Oliva RL, Joergensen RG, Wichern F (2024) Response of the soil microbial ionome to mineral fertilisation in the long-term trial at Askov, Denmark. Soil Biol Biochem. (re-submitted)
Schweizer SA, Mueller CW, Höschen C, Ivanov P, Kögel-Knabner I (2021) The role of clay content and mineral surface area for soil organic carbon storage in an arable toposequence. Biogeochemistry 156:401–420. https://doi.org/10.1007/s10533-021-00850-3
Semenov M, Blagodatskaya E, Stepanov A, Kuzyakov Y (2018) DNA-based determination of soil microbial biomass in alkaline and carbonaceous soils of semi-arid climate. J Arid Environ 150:54–61. https://doi.org/10.1016/j.jaridenv.2017.11.013
Starke R, Jehmlich N, Alfaro T, Dohnalkova A, Capek P, Bell SL, Hofmockel KS (2019) Incomplete cell disruption of resistant microbes. Sci Rep 9:5618. https://doi.org/10.1038/s41598-019-42188-9
Stoddard SF, Smith BJ, Hein R, Roller BRK, Schmidt TM (2015) rrnDB: improved tools for interpreting rRNA gene abundance in bacteria and archaea and a new foundation for future development. Nucleic Acids Res 43:D593–D598. https://doi.org/10.1093/nar/gku1201
Strickland MS, Rousk J (2010) Considering fungal:bacterial dominance in soils - methods, controls, and ecosystem implications. Soil Biol Biochem 42:1385–1395. https://doi.org/10.1016/j.soilbio.2010.05.007
Song Z, Vail A, Sadowsky MJ, Schilling JS (2014) Quantitative PCR for measuring biomass of decomposer fungi in planta. Fungal Ecol 7:39–46. https://doi.org/10.1016/j.funeco.2013.12.004
Tamez-Hidalgo P, Christensen BT, Lever MA, Elsgaard L, Lomstein BA (2016) Endospores, prokaryotes, and microbial indicators in arable soils from three long-term experiments. Biol Fertil Soils 52:101–112. https://doi.org/10.1007/s00374-015-1057-5
Tellenbach C, Grünig CR, Sieber TN (2010) Suitability of quantitative real-time PCR to estimate the biomass of fungal root endophytes. Appl Environ Microbiol 76:5764–5772. https://doi.org/10.1128/AEM.00907-10
Tomlinson PJ, Savin MC, Moore PA Jr (2008) Phosphatase activities in soil after repeated untreated and alum-treated poultry litter applications. Biol Fertil Soils 44:613–622. https://doi.org/10.1007/s00374-007-0245-3
van Veen JA, Ladd JN, Frissel MJ (1984) Modelling C and N turnover, through the microbial biomass in soil. Plant Soil 76:257–274. https://doi.org/10.1007/BF02205585
Vance ED, Brookes PC, Jenkinson DS (1987) An extraction method for measuring soil microbial biomass C. Soil Biol Biochem 19:703–707. https://doi.org/10.1016/0038-0717(87)90052-6
Victorino ÍMM, Berruti A, Orgiazzi A, Voyron S, Bianciotto V, Lumini E (2020). High-throughput DNA sequence-based analysis of AMF communities. In: Ferrol N, Lanfranco L (eds) Arbuscular mycorrhizal fungi. Methods in molecular biology, vol 2146. Humana, New York. https://doi.org/10.1007/978-1-0716-0603-2_9
Wang Y, Yan Y, Thompson KN, Bae S, Accorsi EK, Zhang Y, Shen J, Vlamakis H, Hartmann EM, Huttenhower C (2021) Whole microbial community viability is not quantitatively reflected by propidium monoazide sequencing approach. Microbiome 9:17. https://doi.org/10.1186/s40168-020-00961-3
Wardle DA (1998) Controls of temporal variability of the soil microbial biomass: a global scale synthesis. Soil Biol Biochem 30:1627–1637. https://doi.org/10.1016/S0038-0717(97)00201-0
Watson C, Schlösser C, Vögerl J, Wichern F (2021) Excellent excrement? Frass impacts on a soil’s microbial community, processes and metal bioavailability. Appl Soil Ecol 168:104110. https://doi.org/10.1016/j.apsoil.2021.104110
Weete JD, Abril M, Blackwell M (2010) Phylogenetic distribution of fungal sterols. PLoS ONE 5:e10899. https://doi.org/10.1371/journal.pone.0010899
Wentzel S, Schmidt R, Piepho HP, Semmler-Busch U, Joergensen RG (2015) Response of soil fertility indices to long-term application of biogas and raw slurry under organic farming. Appl Soil Ecol 96:99–107. https://doi.org/10.1016/j.apsoil.2015.06.015
Wichern F, Joergensen RG (2009) Soil microbial properties along a precipitation transect in southern Africa. Arid Land Res Manag 23:115–126. https://doi.org/10.1080/15324980902817071
Wichern J, Wichern F, Joergensen RG (2006) Impact of increased salinity on soil microbial communities and decomposition of maize in acidic soils. Geoderma 137:100–108. https://doi.org/10.1016/j.geoderma.2006.08.001
Wichern F, Islam R, Hemkemeyer M, Watson C, Joergensen RG (2020) Organic amendments alleviate salinity effects on soil microorganisms and mineralisation processes in aerobic and anaerobic paddy rice soils. Front Sust Food Syst 4:30. https://doi.org/10.3389/fsufs.2020.00030
Widmer F, Fließbach A, Laczkó E, Schulze-Aurich J, Zeyer J (2001) Assessing soil biological characteristics: a comparison of bulk soil community DNA-, PLFA-, and Biolog™-analyses. Soil Biol Biochem 33:1029–1036. https://doi.org/10.1016/S0038-0717(01)00006-2
Wu J, Joergensen RG, Pommerening B, Chaussod R, Brookes PC (1990) Measurement of soil microbial biomass C by fumigation-extraction - an automated procedure. Soil Biol Biochem 22:1167–1169. https://doi.org/10.1016/0038-0717(90)90046-3
Yokoyama S, Yuri K, Nomi T, Komine M, Nakamura S, Hattori H, Rai H (2017) The high correlation between DNA and chloroform-labile N in various types of soil. Appl Soil Ecol 117–118:1–9. https://doi.org/10.1016/j.apsoil.2017.04.002
Yu Y, Lee C, Kim J, Hwang S (2005) Group-specific primer and probe sets to detect methanogenic communities using quantitative real-time polymerase chain reaction. Biotechnol Bioengin 89:670–679. https://doi.org/10.1002/bit.20347
Zhao S, Gibbons JG (2018) A population genomic characterization of copy number variation in the opportunistic fungal pathogen Aspergillus fumigatus. PLoS ONE 13:e0201611. https://doi.org/10.1371/journal.pone.0201611
Zielińska S, Radkowski P, Blendowska A, Ludwig-Gałęzowska A, Łoś JM, Łoś M (2017) The choice of the DNA extraction method may influence the outcome of the soil microbial community structure analysis. Microbial Open 6:e453. https://doi.org/10.1002/mbo3.453
Acknowledgements
We highly appreciate the support of Lisa Blöcker, Linda Claaßen, Shiwei Li, Sabrina Meisen, Laura Schages, Rita Theelen, and several student assistants at Rhine-Waal University, Germany for soil sampling and laboratory work. We also like to thank the technicians of the Agricultural Chamber of North Rhine-Westphalia (Germany) for their field work and Diana Werner (Rostock University, Germany) for C/N analyses. To obtain the datasets we received further support by Bent T. Christensen (Aarhus University, Denmark), Clara Berendonk, Ingo Dünnebacke, and Peter Lövenich (Agricultural Chamber of North Rhine-Westphalia), Paul Mäder and Jochen Mayer (Agroscope, Switzerland), Md. Rafiqul Islam (Bangladesh Agricultural University, Bangladesh), and Stefanie Heinze (Ruhr University Bochum, Germany). A further warm thank you goes to Conor Watson (Rhine-Waal University) for his contribution in DOC measurements and critically commenting on the manuscript draft.
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was supported by the German Research Foundation through the research unit DFG-FOR 2337 (project DASIM, grant number DI 546/4–2), by the German Federal Ministry of Education and Research in the framework of BONARES (project SIGNAL, funding codes: 031A562A and 031B0510A), by the Ministry of [Climate Action,] Environment, Agriculture, Nature Conservation and Consumer Protection of the State North Rhine-Westphalia (project EffiZwisch), and by the Ministry of Culture and Science of the State North Rhine-Westphalia in association with Project Management Jülich (project Soil ionoMICS, grant number 005–1703-0025). Zhyldyz Oskonbaeva and Janyl Iskakova received grants from the German Academic Exchange Service (DAAD).
Author information
Authors and Affiliations
Contributions
RGJ performed the statistical analysis and wrote the first draft. MH, ZO, SAS, and CW sampled the soil and conducted the laboratory work. FW contributed to the conception and design of the study. All authors contributed to the manuscript revision, read, and approved the submitted version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Joergensen, R.G., Hemkemeyer, M., Beule, L. et al. A hitchhiker’s guide: estimates of microbial biomass and microbial gene abundance in soil. Biol Fertil Soils 60, 457–470 (2024). https://doi.org/10.1007/s00374-024-01810-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00374-024-01810-3