Abstract
Salinity is a key abiotic constraint on crop production worldwide. To minimize the effect of salinity on rice production, crop improvement strategies based on molecular marker techniques and genetic engineering are being used to develop salt-tolerant rice varieties. The development of such varieties, however, is hindered by limited parental resources. In this study, Haidao 86 (HD86) was observed to exhibit strong salt tolerance during its entire growth period, suggesting that it is a good donor for breeding salt-tolerant rice varieties. To dissect the molecular characteristics underlying the salt tolerance of HD86, we performed quantitative trait locus (QTL) analysis at the seed germination stage of an F2 segregating population derived from crossing HD86 with Nipponbare under 2% NaCl treatment (EC of 32.90 dS/m). One salt-tolerant QTL, qST1, was identified on the long arm of chromosome 1 between the markers CHR1-30M and CHR1-36M, corresponding to a genetic distance of 17.9 cM, and explained about 9.27% of the phenotypic variance. Using 1500 plants of the HD86 BC4F2 introgression lines in the Nipponbare genetic background with heterozygous segments between markers CHR1-30M and CHR1-36M, the exact location of qST1 was narrowed down to a 900 kb region. Combined with a bulk segregant analysis using two extreme phenotype pools of F2, eleven candidates were predicted among the mapped qST1 region. Quantitative RT-PCR analyses revealed that eleven candidates were differentially expressed in response to salt stress at the germination stage in HD86 compared with Nipponbare, suggesting the possible role of these genes in regulating salt tolerance in HD86.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salinity represents a key abiotic constraint on global crop production. Out of the 230 million ha of irrigated land worldwide, 45 million ha (approximately 20%) are affected by salt (Sharma et al. 2016). To survive on salt-affected soils, crops must compete with soil salts for water and cope with ion toxicity, nutritional disorders, and poor soil physical conditions, with a consequent reduction in productivity (Semagn et al. 2006). Rice is the world’s main food crop, feeding more than half the world’s population. As the global population increases, the demand for rice increases accordingly. Rice is susceptible to salinity, especially during its early vegetative and later reproductive stages. The effect of salinity on seedling establishment, seedling growth, and grain yield components such as spikelet number, tiller number, and fertility has successively led to a reduction in grain yield (Khatun and Flowers 1995; Lutts et al. 1995). It is estimated that high salinity reduces agricultural productivity of more than 20% of the cultivated land worldwide (Ganie et al. 2019).
Tolerance to salinity is a complex quantitative trait controlled by polygenes and attributed to multiple mechanisms (Baby and Sujatha 2010; Bizimana et al. 2017; Jahan et al. 2020). In order to breed new high salt-tolerant varieties, we must first identify the genetic basis of the salt resistance mechanisms. DNA-based molecular markers have been extensively used to assess the genetic diversity across most crop species and to map quantitative trait loci (QTLs) across a large number of traits (Mondal et al. 2019). In recent decades, several hundred QTLs for salt tolerance in rice have been identified using the bi-parental linkage mapping of plants at different development stages (Lin et al. 2004; Wang et al. 2012; Ghomi et al. 2013; Tiwari et al. 2016; Shi et al. 2017; Rahman et al. 2017, 2019; Islam et al. 2019; Sun et al. 2019a, b). For example, Lin et al. (2004) identified eight QTLs that contribute to determining Na+ and K+ concentrations in the roots and shoots of seedlings using an F2 population derived from a cross between a high salt-tolerant Xian/Indica variety, Nona Bokra, and a susceptible elite Geng/Japonica variety, Koshihikari (Lin et al. 2004). Among them, two major QTLs with very large effects, qSNC-7 for shoot Na+ concentration and qSKC-1 for shoot K+ concentration, explained 48.5% and 40.1% of the total phenotypic variance, respectively. Wang et al. (2012) reported eleven salt tolerance-associated QTLs that determined seedling height, dry shoot weight, dry root weight, and Na/K ratios in the seedling stages using recombinant inbred lines (RILs, F2:9) derived from a cross between the salt-tolerant variety, Jiucaiqing, and the salt-sensitive variety, IR26 (Wang et al. 2012). As rice is also sensitive to salt at the reproductive stage, some studies also focused on the role of salt tolerance in determining yield traits including plant height, number of tillers, and number of spikelets (Mondal et al. 2019; Sun et al. 2019b; Tiwari et al. 2016). A few studies also reported salt-tolerant QTLs at the rice seed germination stage. For example, Wang et al. (2011) reported a total of 16 QTLs explaining 4.6–43.7% of the variation in the seed imbibition rate and germination percentage under a salt treatment using RILs derived from a cross between a Geng/Japonica rice salt-tolerant landrace and a salt-sensitive Xian/Indica rice variety (Wang et al. 2011). With the development of next-generation sequencing, increasingly more research is applying genome-wide association mapping studies (GWAS) based on linkage disequilibrium to identify numerous QTLs and genes related to salt tolerance (Al-Tamimi et al. 2016; Shi et al. 2017; Sun et al. 2019a; Tsai et al. 2019). Eleven loci containing 22 significant salt tolerance-associated single-nucleotide polymorphisms (SNPs) were identified based on the stress-susceptibility indicators of vigor index and mean germination time using the resequencing of 467 diverse rice accessions at the seed germination stage (Shi et al. 2017). Furthermore, based on the whole-genome resequencing of two extreme pools demonstrating a significantly different phenotype, bulk segregant analysis (BSA) is more effective in rapidly locating candidate genomic regions that underlie target genes (Sun et al. 2019a; Takagi et al. 2013). Sun et al. 2019a, b) identified six candidate regions for salt tolerance on chromosome 1 via BSA-seq using the extreme populations derived from the cross between Changmaogu (salt-tolerant rice landrace) and a salt-sensitive cultivar, Zhefu802 (Sun et al. 2019a). Though many salt-tolerant QTLs have been detected in different rice lines, only a few major salt-tolerant rice QTLs or genes, such as qSKC1 (Ren et al. 2005), qSNC7 (Lin et al. 2004), Saltol (Thomson et al. 2010), OsRR22 (Takagi et al. 2015), and qSE3 (He et al. 2019), have been identified by genomic methods (Sun et al. 2019a).
To minimize the effect of salinity on rice production, crop improvement strategies based on molecular marker techniques and genetic engineering are being used in conjunction with traditional breeding efforts to develop salt-tolerant rice varieties. Limited parental resources, however, hinder the development of such varieties. The rice variety Haidao86 was first discovered in mangrove tidal flats in 1986 (Chen et al. 2017). Haidao 86 was named sea rice 86 in Chen’s study (Chen et al. 2017). However, sea rice literally refers to a kind of rice variety that can grow in seawater or be irrigated using seawater. In reality, no rice varieties have hitherto survived in seawater. Thus, the word “sea rice” is easily misunderstood and exaggerates the salt tolerance of Haidao 86 (Ling 2018). Accordingly, in this study, we renamed this salt-tolerant variety Haidao86 (HD86), based on the Chinese phonetic alphabet. HD86 can grow in saline-alkaline and infertile soil, and has the ability of submergence and water logging tolerance in marginal lands while producing acceptable average yield of ~ 2250 kg/ha yields (Chen et al. 2017). Thus, it is considered as a strategic germplasm resource for new rice variety development. It was reported that HD86 has higher seed germination rates compared to the salt-tolerant cultivar, Yanfeng47, under salt stress (Chen et al. 2017). However, the molecular characterization of salt tolerance remains unknown in HD86.
In this study, we assessed the salt tolerance of HD86 during the entire growth period and confirmed that HD86 has strong salt tolerance during the seedling stage both in paddy fields and in the laboratory. In addition, HD86 and its hybrid progeny population exhibited higher salt tolerance in saline–alkaline soils. In order to characterize the mechanism of salt tolerance in HD86, we constructed a segregated population derived from a cross between HD86 and Nipponbare. Based on the whole-genome resequencing of HD86, we developed 138 polymorphism markers between two parents on agarose gel and constructed a genetic linkage map. Using a combined linkage map with a phenotype of seed germination rate for the F2 population under salt treatment, we identified one QTL on the long arm of chromosome 1 at about a 900 kb physical distance. According to the BSA analysis, eleven candidates were predicted. The transcript levels of the candidate genes in response to salt stresses were analyzed by quantitative RT-PCR (qRT-PCR) analyses.
Materials and Methods
Plant Materials, Growth Conditions, and Salt Stress Treatments
The F2 population used in this study, containing 209 lines, was derived from a cross between Nipponbare (O. sativa ssp. Geng/Japonica) and HD86 (O. sativa ssp. Xian/Indica), which were harvested under the same growth conditions in Sanya, Hanan province, China.
For the germination assay, seeds were firstly placed for 7 ds at 42 °C to break any possible dormancy. First, we conducted preliminary germination experiment of HD86 and Nipponbare under salt conditions for 10 days using NaCl content with 0, 85.6, 171.1, 256.7, and 342.2 mM, corresponding to an EC of 0, 9.20, 17.67, 25.40, and 32.90 dS/m, respectively, as measured by conductivity meter, Mettler Toledo FE30, Switzerland. The germination rate was 100, 100, 92, 76, 46%, respectively for HD86, and 100, 100, 90, 64, 18.7%, respectively for Nipponbare. Under the salt condition with an EC of 32.90 dS/m, the germination rate was significantly different between HD86 and Nipponbare. Thus, we used an EC of 32.90 dS/m as salt stress condition in the F2 population germination assay. Fifty seeds with the uniform maturity of each F2:3 line were surface-sterilized, placed on filter paper on sterile glass plates, and incubated in the dark (30 °C). Among 209 lines, 68 lines were not used because they lacked sufficient seeds. Owing to the sterility of the hybrid of Xian/Geng (HD86/Nipponbare) rice, more than half lines of the F2:3 segregation population obtained less than 300 seeds. Thus, it is not be possible for us to perform the seed germination assay in salt-free water as mock. In this population, there were 16 lines with seed germination rate less than 20% under salt conditions. There were only 3 lines with germination rates less than 100% under salt-free water, which were excluded in the next experiments. Ultimately, assessment of the seed germination rate was based on 3 replicates for each of the 138 lines and the two parent lines.
In agricultural practice until to now, few rice cultivars can survive under salt content with more than 0.7% (119.8 mM, EC of 12.80 dS/m). Thus, rice was suffered to salt stress with an EC of 12.80 dS/m in the lab and paddy field. For the seedling salt tolerance assays, seeds of Nipponbare and HD86 were de-husked, surface-sterilized, and grown in 96-well plates in a growth chamber (15 h light /28 °C, 9 h dark /26 °C). The growth medium, Yoshida’s culture solution (Patnaik et al. 1965), was renewed every 2 days. After 14 days of growth, the seedlings were treated with either Yoshida’s culture solution or Yoshida’s culture solution containing 200 mM NaCl (an EC of 20.30 dS/m). At the same time, the 2-week-old seedlings were transplanted into soil in a 1 L glass bottle and treated with diluted seawater of 12.80 dS/m for 1 month.
For the paddy field assays, seedlings of HD86 and its hybrids with the cultivars (Nipponbare (O. sativa ssp. Geng/Japonica), Yanfeng47 (O. sativa ssp. Geng/Japonica), and Yangeng218 (O. sativa ssp. Geng/Japonica) were transplanted into an irrigated field in Dongying, China (latitude 38°15′N; longitude 118°50′E) in early June. The planting density was 25 × 14 cm, with one plant per hill in each 3 m2 block. Saline water with an EC of 9.20 dS/m was prepared by mixing fresh water and underground water with an EC of 40.00 dS/m (pH 8.0). One week after transplantation, rice plants were irrigated by saline water with an EC of 9.20 dS/m (pH 8.0) from the seedling through to the mature stage. The salinity of the soil was monitored and adjusted by irrigation with saline water whenever necessary. Fertilization and other agronomic management measures were implemented according to local rice cultivation practices.
BSA Using F2 Segregation Population
An equal amount of DNA from twenty of the 138 F2 plants with the highest seed germination rates (0.70–0.92) of F2:3 families was mixed to construct the salt-tolerant pool (T-pool). Then, the DNA from another twenty plants with the lowest seed germination rates (0.02–0.26) was mixed to construct the salt-sensitive pool (S-pool). The sequence data of the two bulks and parents of HD86 were determined by an Illumina HiSeq4000 (Novogene limited company, Beijing, China). Briefly, a total of 1.5 μg DNA per sample was used as the input material for the DNA sample preparations. Sequencing libraries were generated using a Truseq Nano DNA HT Sample preparation Kit (Illumina USA) following the manufacturer’s recommendations, and the index codes were added to attribute the sequences to each sample. After that, the DNA sample was sonicated to produce fragments with a size of 350 bp; the DNA fragments were then end-polished, A-tailed, and ligated with the full-length adapter for Illumina sequencing with further PCR amplification. Lastly, the PCR products were purified (AMPure XP system), and libraries were analyzed for size distribution by Agilent2100 Bioanalyzer and quantified using real-time PCR.
These constructed libraries were sequenced using an Illumina HiSeq4000 platform, and 150 bp paired-end reads were generated with an insert size around 350 bp.
To ensure the reads are reliable and free of artificial bias (i.e., low-quality paired reads, which mainly result from base-calling duplicates and adapter contamination), in the following analyses, the raw data (raw reads) in fasta format were firstly processed through a series of quality control (QC) procedures involving removing reads with ≥ 10% unidentified nucleotides, reads with > 50% bases with a phred quality < 5, reads with > 10 nt aligned to the adapter, and putative PCR duplicates generated by PCR amplification in the library construction process.
BWA was used to align the clean reads of the two bulks against the Oryza sativa Nipponbare (IRGSP Build 4) reference genome. Alignment files were converted to BAM files using the SAM tools software. If multiple read pairs had identical external coordinates, we only retained the pair with the highest mapping quality. SNP/InDel variants calling were performed for all samples by using the Unified Genotyper function in the GATK software. ANNOVAR was used to annotate SNP or InDel based on the GFF3 files for the reference genome.
The homozygous SNPs/InDels between the two parents were extracted from the vcf files for SNP/InDel. The reads depth information for the homozygous SNPs/InDels above in the offspring pools was obtained to calculate the SNP/InDel index. We used the genotype of one parent as the reference and to count reads number of this parent’s genotype or the others in the offspring pool. We then calculated the ratio of the number of different reads in the total number, which is the SNP/InDel index of the base sites. We filtered out the points in the SNP/InDel indices in both pools that were less than 0.3. Sliding window methods were used to present the SNP/InDel index of the whole genome. The average of all SNP/InDel indices in each window was used as the SNP/InDel index for this window. Usually we use window size of 1 Mb and a step size of 10 Kb as default settings. The differences of the SNP/InDel indices of two pools were calculated as the delta SNP/InDel index, which means ΔSNP/InDel index = SNP/InDel index (T-pool) − SNP/InDel index (S-pool). A ten thousand replacement sampling of individuals in two bulks was performed to calculate the reads depth of the given number of alleles. A 95% confidence level was selected as the threshold for the candidate interval.
Genome Sequencing Analysis
Genomic DNA (gDNA) was extracted from HD86 leaf tissue by the CTAB method. Sequencing libraries were constructed using an Illumina TruSeq DNA sample preparation kit (Illumina, Santiago, California, USA). Briefly, high-quality gDNA was sheared into fragments using an ultrasonicator (Covaris, Woburn, MA, USA) to obtain a final library with an average insert size of 350 bp. The sheared DNA was then subjected to end repair and the adenylation of 3′ ends followed by the ligation of indexed paired-end adapters. Finally, the libraries were sequenced on the Illumina HiSeq 2000 NGS platform.
The original image data generated by the Illumina sequencer were converted into sequence data via base calling (Illumina pipeline CASAVA v1.8.2). These data were then subjected to quality control procedures to remove unusable reads, which included the reads containing library construction adapters and those containing more than 10% unknown bases or more than 50% low-quality bases (sequencing quality value ≤ 5).
The sequencing reads were aligned to the reference genome using the Burrows–Wheeler alignment tool (BWA) (Li and Durbin 2010) with default parameters. Subsequent processing, including duplicate removal, was performed using SAMtools and PICARD (http://picard.sourceforge.net).
The raw SNP/InDel sets were called using the parameters ‘− q 1 − C 50 − m2 − F0.002 − d1000’ and then filtered according to two criteria: mapping quality > 20 and depth of variant position > 4. BreakDancer and CNVnator were used for SV and CNV detection, respectively. ANNOVAR was used for functional annotation of variants. UCSC known UCSC genes were used for gene and regional annotations.
Construction of Genetic Linkage Maps and QTL Analysis
Structural variations were exploited to develop polymorphism markers based on comparing the HD86 genome resequencing data with Nipponbare. One hundred and thirty-eight molecular markers were identified to be polymorphic between HD86 and Nipponbare on an agarose gel (Table S1). These markers were equally distributed on 12 chromosomes and used to construct a genetic linkage map using Joinmap4.1 software (Van Ooijen 2006). The genetic map distance (centimorgans, cM) was calculated using the Kosambi function.
QTL mapping analysis for seed germination rate was performed via the composite interval mapping (CIM) method using WinQTLCart2.5 software (Wang et al. 2007). The LOD threshold at a significance level of 0.05 was determined based on 1000 permutations.
qRT-PCR Analyses
In the seed germination rate assay, fifty seeds were surface-sterilized, placed on filter paper on sterile glass plates, and incubated in the dark (30 °C). The seeds were de-hulked and sampled after being treated with 2% NaCl solution or water for 2 days. Three biological replicates were used for each treatment group.
Total RNA was isolated from the seeds using RNAiso (Takara, Dalian, China). qRT-PCR was performed using an ABI 7500 Fast Real-Time PCR system (Applied Biosystems, Foster City, CA, USA) with SYBR Premix Ex Taq (Takara). The rice eEF-1a gene (AK061464) was used as an internal control to quantify the relative level of each target gene (Jain et al. 2006). Three replicate biological experiments were performed.
Results
The Obvious Salt-Tolerant Phenotypes of Hd86 During the Entire Growth Period
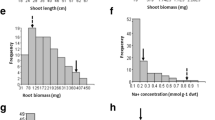
To evaluate HD86 salt tolerance, the seeds or seedlings of HD86 were exposed to either seawater or an NaCl-containing hydroponic solution. As shown in Fig. 1a, b, 46.0% of HD86 seeds, but only 18.7% of Nipponbare seeds, germinated by the 10th day after treatment with NaCl of EC 32.90 dS/m. We also investigated the salt tolerance of the seedlings grown in a salt solution. When the 2-week-old seedlings were treated with NaCl solution of EC 20.30 dS/m for 7 days, more leaves remained green on HD86 than on Nipponbare (Fig. 1c). Approximately 77.7% of the HD86 seedlings grew robustly; by contrast, only 4.6% of the Nipponbare plants grew new leaves after restoration to normal conditions for 7 days (Fig. 1d). The 2-week-old seedlings were transferred to soil with diluted seawater of EC 12.80 dS/m and were grown for 1 month, and HD86 grew well compared to Nipponbare (Fig. 1e). After transplantation into soil, HD86 and its hybrids with various cultivars thrived, and their seeds set in the rice paddy fields of the Yellow River Delta that were irrigated with water with 0.7% salinity, whereas the Nipponbare plants died, and no seeds were obtained (Fig. 1f). All of these results indicate that HD86 had stronger tolerance to salinity relative to Nipponbare during the entire growth period. HD86 is thus likely a good donor for breeding salt-tolerant rice varieties.
Phenotypes of HD86 and Nipponbare under salt stress conditions. a Seed germination of HD86 and Nipponbare in NaCl solution of EC 32.90 dS/m for 10 days. Scale bar = 1 cm. b Seed germination rates of HD86 and Nipponbare in NaCl solution of EC 32.90 dS/m for 10 days. Asterisks (**) denote significant difference (P < 0.01) between Nipponbare and HD86 according to a t test. c Phenotypes of 2-week-old seedlings of HD86 and Nipponbare exposed to NaCl solution of EC 20.30 dS/m for 7 days and restored in water for 7 days. Scale bar = 1 cm. d Survival rates of seedlings restored in water for 7 days after 7-day treatment with NaCl solution of EC 20.30 dS/m. Asterisks (**) denote significant difference (P < 0.01) between Nipponbare and HD86 according to a t test. e Phenotypes of 2-week-old seedlings of HD86 and Nipponbare exposed to the diluted seawater of EC 12.80 dS/m. Scale bar = 5 cm. f Phenotypes of HD86, Nipponbare, HD86-derived hybrids and some cultivars in paddy filed of saline-alkali soil in Dongying, Shandong Province, China. YG218 (Yangeng 218) and YF47 (Yanfeng 47) are two salt-tolerant cultivars. YG218/HD86 F2 means F2 segregation population of hybrid crossed by YG218 and HD86. YF47/HD86 F2 means F2 segregation population of hybrid crossed by YF47 and HD86
Genome-Wide DNA Polymorphisms in HD86 Based on Whole-Genome Resequencing
In order to dissect the molecular characteristics behind salt tolerance, whole-genome resequencing was performed on the HD86 genomic DNA using the Illumina Hiseq2000 platform. We obtained 12.4 billion clean sequence reads, among which approximately 94% were of high quality (a quality score above Q30) with 20× genome coverage (Table S2). These short reads were mapped onto the Nipponbare genome (IRGSP Build 5) using BWA software (Li and Durbin 2010), and 96.86% of the obtained reads were successfully mapped to chromosomes with an average depth of 29.19× (Table S3). We also mapped the sequence reads onto Xian/Iindica 9311 and R498 (Du et al. 2017) using the same analysis methods; 97.49 and 97.87% of the reads were located on the chromosomes of 9311 and R498, with 29.06× and 28.82× average depths, respectively (Table S3).
Using the HD86 whole-genome resequencing data, we identified 4,121,246 DNA polymorphisms with the Nipponbare reference on a genome-wide scale (Fig. S1, Table 1). Almost all SNPs and insertion and deletion (InDels) were evenly distributed across individual chromosomes, with densities of approximately 10 and 1 Kb−1, respectively. The distribution of structural variants (SVs), however, was uneven across chromosomes, with chromosomes 12 and 3 having the highest (98.8 Mb−1) and lowest (33.6 Mb−1) densities, respectively (Fig. S1, Table 1).
Identification of Salt-Tolerant QTL by BSA and Map-Based Genetic Linkage Analysis
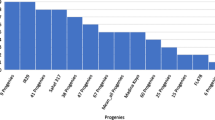
As HD86 had stronger tolerance to salinity relative to Nipponbare during the entire growth period, we used the seed germination rate to assess the F2 population phenotype obtained by crossing Nipponbare and HD86 under salt treatment. In order to verify whether QTLs referred to the salt tolerance of HD86, we conducted a BSA based on next-generation sequencing technologies. As shown in Fig. 2a, the seed germination rate ranged from 2 to 92%, demonstrating a normal frequency distribution among the F2 population. The seed germination rate of HD86 was 76.6%, which is significantly higher than that of Nipponbare, which was 37.3%. Interestingly, lines with seed germination rates that were both higher and lower than those of the parents were observed in the population, such as in the case of L45 and L327 (Fig. 2b).
The phenotype of seed germination of F2 population derived from a cross with HD86 and Nipponbare under NaCl solution of EC 32.90 dS/m for 10 days. a The frequency distribution of seed germination rate of the population and the two parents. b Seed germination of typical progeny lines under salt conditions
Twenty lines with the highest seed germination rate (0.70–0.92) and twenty lines with the lowest seed germination rate (0.02–0.26) were selected and resequenced as the salt-tolerant pool (T-pool) and the salt-sensitive pool (S-pool), respectively. A total of 27.2 Gb raw data were generated, and 27 Gb clean data were obtained after being filtered with 95% high quality (> Q30) (Table S4). These sequence data were mapped onto the Nipponbare genome (IRGSP Build 5) using BWA software (Li and Durbin 2010). In total, 94.03% of the obtained reads were successfully mapped to chromosomes in the T-pool with an average depth of 28.92 × 97.23% of the obtained reads successfully mapped to chromosomes in the S-pool with an average depth of 31.95 × (Table S5).
We identified 4098526 SNP/InDels in HD86 with respect to the Nipponbare reference on a genome-wide scale (Table 1). The average homozygous SNPs/InDels index in the two pools and ΔSNP/InDel index between the two pools were gained to calculate in a 1 Mb intervals using a 10 kb sliding window. One genome region on chromosome 1 was identified with a significance above the threshold 95% confidence level and had a physical distance of about 25.97 to 40.79 Mb, which is consistent with the results from the preliminary map-based QTL analysis (Fig. S2). Overall, 115 candidate genes were predicted to determine the variations between the two pools.
To validate the location of salt-tolerant QTL by BSA-Seq, we constructed QTL analysis using the F2 segregation population by crossing HD86 and Nipponbare. The continuous distribution of the phenotype indicates that the salt-tolerant trait was controlled by multiple genes and could be identified by QTL analysis. Thus, we developed polymorphism markers between the parental lines. Based on whole-genome resequencing of HD86, 138 polymorphism markers were identified and used to construct a genetic linkage map (Table S1). This map covered 1301.0 cM of 12 chromosomes with an average distance of 9.43 cM between markers (Fig. S3). One QTL was identified on chromosome 1, designated as qST1, located between marker CHR1-30M and CHR1-36M, corresponding to a genetic distance of 17.9 cM (a physical distance of about 30–36 Mb), and explained about 9.27% of the phenotypic variance (Fig. 3a, Table 2, Fig. S4). The location region of qST1 was consistent with that of one from BSA. In addition, the alleles of HD86 at the qST1 position increased the germination rate under salt treatment (Table 2).
In order to narrow down the exact region corresponding to qST1, two HD86 BC4F1 introgression lines in the Nipponbare genetic background with heterozygous regions between marker CHR1-30M and CHR1-36M were selected to generate a BC4F2 segregation population with 1500 F2 plants. Another eight markers in the QTL located region were identified (Table S1). In total, 26 F2 plants featuring recombination between CHR1-30M and CHR1-36M were identified from the segregating population (Fig. 3b, c). An analysis of the variance between the nine genotypes among the 26 recombinant F2 families revealed three distinct phenotypic groups. The genotype data and corresponding phenotypic results allowed us to map qST1 between the markers M324 and M334, with a physical distance of about 900 kb (Fig. 3b). Combined with the QTL region located obtained by fine-mapping using the BC4F2 segregating population and 115 candidate genes deduced from BSA, eleven candidate genes were identified, among which two genes (Os01g0781100 and Os01g0781200) were nonsynonymous in HD86 compared with the Nipponbare genome reference (Table S6).
qRT-PCR Analysis of the Candidate Genes Corresponding to qST1
Since qST1 likely regulates salt tolerance at the seed germination stage, we first validated the expression levels of these candidates in germinating seeds under salt treatment. As shown in Fig. 4, the candidates could be divided into four groups. The first group included Os01g0772800, Os01g0778800, and Os01g0779300 encoding the putative signal recognition particle 54 kDa protein, the putative insulin-degrading enzyme, and the putative expressed protein, respectively. The expression levels of these three genes were significantly upregulated after salt treatment of Nipponbare rice seeds, but were not obviously different in HD86 (Fig. 4a–c). Moreover, the expression levels of the two genes were significantly higher in Nipponbare relative to those in HD86 under treatment with either salt solution of EC 32.90 dS/m or salt-free water. The second group included Os01g0773200 and Os01g0783600, the former encoding the putative transcription factor jumonji and the latter encoding the putative frataxin (iron–sulfur cluster protein 22, OsISC22). The expression levels of these two genes were significantly higher in rice seeds of Nipponbare than those of HD86, though both were upregulated after salt treatment (Fig. 4d, e). The third group contained two disease-resistant genes, Os01g0781100 and Os01g0781200, which encoded disease resistance RPP13-like protein 1 and Magnaporthe grisea resistance-64 (Pi64), respectively. These transcripts were nearly indetectable in the HD86 seeds for both salt-free water and salt solution of EC 32.90 dS/m treatments. However, the expression levels of Os01g0781100 and Os01g0781200 were dramatically upregulated and downregulated, respectively, in the seeds of Nipponbare after salt treatment (Fig. 4f, g). Os01g0770800, Os01g0771100, Os01g0784700, and Os01g0786800 constituted the fourth group, encoding the copper transporter family protein, putative mitochondrial glycoprotein, serine/threonine protein kinase receptor precursor, and putative membrane protein, respectively. The expression levels of these four genes were significantly upregulated after salt treatment in the Nipponbare rice seeds but were obviously downregulated in HD86. Moreover, the expression levels were dramatically higher in Nipponbare seeds than in HD86, especially under NaCl solution of EC 32.90 dS/m treatment (Fig. 4h–k). In summary, the changes in the expression levels of these genes were smaller in the germinated seeds of HD86 than that in Nipponbare when treated with salt stress.
Discussion
In this study, we identified one salt-tolerant QTL (qST1) related to salt tolerance at the seed germination stage using the F2 population derived from crossing Nipponbare with HD86, a high salinity-tolerant Xian/Indica rice cultivar. Seed vigor is a complex and important agricultural trait which determines the potential for rapid, uniform emergence, and for the establishment of seedlings under a wide range of field conditions, especially in a direct-seeding planting system, which is the most popular type of rice planting system throughout the world. Thus, salt tolerance at the seed germination stage has become a major rice breeding goal in many Asian countries where paddy fields are salinized to some degree (Shi et al. 2017). The seed germination rate was chosen as an indicator to determine the phenotype of the segregation population. As we excluded lines with a low germination rate under natural conditions (see Materials and Methods), qST1 was identified as the QTL regulating seed germination under salt stress.
In our study, the distribution of the seed germination rate under salt conditions was continuous, suggesting that the salt-tolerant trait is controlled by multiple genes (Fig. 2). However, only one QTL was detected on chromosome 1, which explained 9.27% of the phenotypic variation in this F2 population (Table 2). The reason for this result might be as follows. The parents of the mapping populations must have sufficient variation for the traits of interest at level of both the DNA sequence and phenotype (Bizimana et al. 2017; Meksem and Kahl 2005; Semagn et al. 2006). However, Nipponbare was moderately tolerant to 40 mM NaCl but not to 80 mM, and showed an ‘ion balance’ mechanism (Hossain et al. 2015). In this sense, Nipponbare might not be a good choice as another parent to construct a segregation population with HD86 due to loss of minor effect QTLs. SKC1, encoding a Na+/K+ transporter protein, was first cloned in the rice cultivar Nona Bokra (Ren et al. 2005). The amino acid of SKC1 in Nona Bokra (SKC1Nona) differs at four base positions from that of SKC1 in Koshihikari, a salt-sensitive cultivar, resulting in a higher expression level of the gene in Nona Bokra than in Koshihikari. In our study, the amino acid sequence of SKC1HD86 was the same as that of SKC1Nona, and SKC1Nip was the same as that of SKC1Koshihikari (data not shown). However, we did not detect SKC1 in this population, indicating that the effect of SKC1 was less than that of qST1 or qST1 only had function in seed germination stage in this population.
As we only detected one salt-tolerant QTL on chromosome 1, we compared its location with those of other salt-tolerant QTLs on chromosome 1 reported in previous studies (Table S7). As shown in Fig. 5, two hotspots of QTLs were found around a physical distance of 10 and 30–40 Mb, respectively, and SKC1 was located at a position around 10 Mb. However, most of the QTLs were identified in the seedling stages for shoot/root length, shoot/root fresh/dry weight, Na+/K+ content, and biomass, while only few for plant height, grain yields, and 1000-grain weight in the reproductive stages (Bimpong et al. 2014; Hossain et al. 2014; Mondal et al. 2019; Sun et al. 2019b; Tiwari et al. 2016). qST1 overlapped with the QTL regulating the root dry weight reported in the seedling stages by Ghomi et al. (2013) and QTLs regulating the plant height, panicle length, pollen fertility, Na + concentration, and Na/K ratio in the flag leaves in the reproductive stages by Hossain et al. (2014). It is unclear if they were in the same loci. To date, few studies have focused on evaluating the salt tolerance of rice at the germination stage (Shi et al. 2017). Wang et al. (2011) reported 16 QTLs regulating the imbibition rate and germination percentage under salt stress, which were found on all 12 chromosomes with the exception of chromosome 1 and 5. Eleven loci containing 22 significant salt tolerance-associated SNPs were identified based on the stress-susceptibility indicators of the vigor index and mean germination time through a genome-wide association study based on the resequencing of 478 diverse rice accessions (Shi et al. 2017). However, no candidate genes were located in the qST1 region. Sun et al. identified six candidate regions for salt tolerance on chromosome 1 via BSA-seq for two extreme bulks derived from the cross between Changmaogu and Zhefu802 at the germination and young seedling stages (Sun et al. 2019a). However, qST1 did not overlap with these QTLs (Fig. 5). Thus, qST1 is a new QTL that regulates seed germination under salt conditions. Due to an insufficient number of seeds, we could not conduct a QTL analysis in the seedling stages and could not confirm whether qST1 was functional at this stage. Regardless of whether qST1 had a function in the seedling stage, pyramiding it with the positive effects of salt-tolerant QTLs at the other growth stages of rice via selection with tightly linked markers would allow for the generation of more salt-tolerant rice cultivars.
Based on the BSA-seq analysis and mapping, eleven candidates were selected for further analysis by qRT-PCR. As shown in Fig. 4, changes in the expression levels of these genes were less significant in the germinated seeds of HD86 than those in Nipponbare after the salt treatment, indicating that the seeds of HD86 had a higher resistance capability. Among these candidates, nine genes (Os01g0770800, Os01g0771100, Os01g0772800, Os01g0773200, Os01g0778800, Os01g0779300, Os01g0783600, Os01g0784700, and Os01g0786800) had variations in the regulation-region or intron in HD86 compared with the Nipponbare genome reference (Table S6). The expression levels of them were also reduced by salt stress, which suggested these candidates are probably involved in the salt tolerance of HD86 by regulating the expression levels owing to the variations in up- or down-regulation region on the genome. A previous study also reported the salt tolerance of HD86 based on transcriptome analysis (Chen et al. 2017). They found that the genes of families of PPR, peroxidase, dirigent, MATE, GST, NBC-ARC, NBS, and kinesin were likely associated with salt tolerance of HD86. However, no genes were located in the qST1 region, which indicated these genes might act in the downstream of qST1-regulated salt tolerance. Future work should perform fine-mapping using a larger advanced segregation population to narrow down the location of qST1 to confirm whether some or all of the candidates coordinately regulate the salt resistance of HD86 during seed germination.
Data Availability
All datasets generated for this study are included in the article/Supplementary Material.
References
Al-Tamimi N, Brien C, Oakey H, Berger B, Saade S, Ho YS, Schmockel SM, Tester M, Negrao S (2016) Salinity tolerance loci revealed in rice using high-throughput non-invasive phenotyping. Nat Commun 7:13342. https://doi.org/10.1038/ncomms13342
Baby JDJ, Sujatha S (2010) Biological and physiological perspectives of specificity in abiotic salt stress response from various rice plants. Asian J Agric Sci 2:90–105
Bimpong IK, Manneh B, Diop B, Ghislain K, Sow A, Amoah NKA, Gregorio G, Singh RK, Ortiz R, Wopereis M (2014) New quantitative trait loci for enhancing adaptation to salinity in rice from Hasawi, a Saudi landrace into three African cultivars at the reproductive stage. Euphytica 200:45–60. https://doi.org/10.1007/s10681-014-1134-0
Bizimana JB, Luzi-Kihupi A, Murori RW, Singh RK (2017) Identification of quantitative trait loci for salinity tolerance in rice (Oryza sativa L.) using IR29/Hasawi mapping population. J Genet 96:571–582. https://doi.org/10.1007/s12041-017-0803-x
Chen R, Cheng Y, Han S, Van Handel B, Dong L, Li X, Xie X (2017) Whole genome sequencing and comparative transcriptome analysis of a novel seawater adapted, salt-resistant rice cultivar—sea rice 86. BMC Genomics 18:655. https://doi.org/10.1186/s12864-017-4037-3
Du H, Yu Y, Ma Y, Gao Q, Cao Y, Chen Z, Ma B, Qi M, Li Y, Zhao X, Wang J, Liu K, Qin P, Yang X, Zhu L, Li S, Liang C (2017) Sequencing and de novo assembly of a near complete indica rice genome. Nat Commun 8:15324. https://doi.org/10.1038/ncomms15324
Ganie SA, Molla KA, Henry RJ, Bhat KV, Mondal TK (2019) Advances in understanding salt tolerance in rice. Theor Appl Genet 132:851–870. https://doi.org/10.1007/s00122-019-03301-8
Ghomi K, Rabiei B, Sabouri H, Sabouri A (2013) Mapping QTLs for traits related to salinity tolerance at seedling stage of rice (Oryza sativa L.): an agrigenomics study of an Iranian rice population. OMICS 17:242–251. https://doi.org/10.1089/omi.2012.0097
He Y, Yang B, He Y, Zhan C, Cheng Y, Zhang J, Zhang H, Cheng J, Wang Z (2019) A quantitative trait locus, qSE3, promotes seed germination and seedling establishment under salinity stress in rice. Plant J 97:1089–1104. https://doi.org/10.1111/tpj.14181
Hossain MR, Pritchard J, Ford-Lloyd BV (2015) Qualitative and quantitative variation in the mechanisms of salinity tolerance determined by multivariate assessment of diverse rice (Oryza sativa L.) genotypes. Plant Genetic Resour 14:91–100. https://doi.org/10.1017/s1479262115000118
Hossain H, Rahman MA, Alam MS, Singh RK (2014) Mapping of quantitative trait loci associated with reproductive-stage salt tolerance in rice. J Agron Crop Sci 201:17–31. https://doi.org/10.1111/jac.12086
Islam MS, Ontoy J, Subudhi PK (2019) Meta-analysis of quantitative trait loci associated with seedling-stage salt tolerance in rice (Oryza sativa L.). Plants 8:33
Jahan N, Zhang Y, Lv Y, Song M, Zhao C, Hu H, Cui Y, Wang Z, Yang S, Zhang A, Hu J, Ye G, Qian Q, Gao Z, Guo L (2020) QTL analysis for rice salinity tolerance and fine mapping of a candidate locus qSL7 for shoot length under salt stress. Plant Growth Regul 90:307–319. https://doi.org/10.1007/s10725-019-00566-3
Jain M, Nijhawan A, Tyagi AK, Khurana JP (2006) Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR. Biochem Biophys Res Commun 345:646–651. https://doi.org/10.1016/j.bbrc.2006.04.140
Khatun S, Flowers TJ (1995) Effects of salinity on seed set in rice. Plant Cell Environ 18:61–67
Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26:589–595. https://doi.org/10.1093/bioinformatics/btp698
Lin HX, Zhu MZ, Yano M, Gao JP, Liang ZW, Su WA, Hu XH, Ren ZH, Chao DY (2004) QTLs for Na+ and K+ uptake of the shoots and roots controlling rice salt tolerance. Theor Appl Genet 108:253–260. https://doi.org/10.1007/s00122-003-1421-y
Ling QH (2018) Discussion on the related problems of rice planting in saline-alkali soil. China Rice 24:1–2
Lutts S, Bouharmont J (1995) Changes in plant response to NaCl during development of rice (Oryza sativa L.) varieties differing in salinity resistance. J Exp Bot 46:1843–1852
Meksem K, Kahl G (2005) The handbook of plant genome mapping: genetic and physical mapping. Wiley-Vch Verlag GmbH, Weinheim, Germany
Mondal S, Borromeo TH, Diaz MGQ, Amas J, Rahman MA, Thomson MJ, Gregorio GB (2019) Dissecting QTLs for reproductive stage salinity tolerance in rice from BRRI dhan 47. Plant Breeding Biotechnol 7:302–312. https://doi.org/10.9787/pbb.2019.7.4.302
Patnaik S, Misra CS, Bhadrachalam A (1965) Studies on the nutrition of the rice plant (Oryza satvia L.). Proc Indian Acad Sci 61:309–315
Rahman MA, Bimpong IK, Bizimana JB, Pascual ED, Arceta M, Swamy BPM, Diaw F, Rahman MS, Singh RK (2017) Mapping QTLs using a novel source of salinity tolerance from Hasawi and their interaction with environments in rice. Rice 10:47. https://doi.org/10.1186/s12284-017-0186-x
Rahman MA, Thomson MJ, De Ocampo M, Egdane JA, Salam MA, Shah EAM, Ismail AM (2019) Assessing trait contribution and mapping novel QTL for salinity tolerance using the Bangladeshi rice landrace Capsule. Rice 12:63. https://doi.org/10.1186/s12284-019-0319-5
Ren ZH, Gao JP, Li LG, Cai XL, Huang W, Chao DY, Zhu MZ, Wang ZY, Luan S, Lin HX (2005) A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat Genet 37:1141–1146. https://doi.org/10.1038/ng1643
Semagn K, Bjornstad A, Ndjiondjop MN (2006) Principles, requirements and prospects of genetic mapping in plants. Afr J Biotech 5:2569–2587
Sharma A, Rana C, Singh C, Katoch V (2016) Soil salinity causes, effects, and management in cucurbits. In: Pessarakli M (ed) Handbook of cucurbits: growth, cultural practices, and physiology. CRC Press, Taylor and Francis Group, pp 419–434
Shi Y, Gao L, Wu Z, Zhang X, Wang M, Zhang C, Zhang F, Zhou Y, Li Z (2017) Genome-wide association study of salt tolerance at the seed germination stage in rice. BMC Plant Biol 17:92. https://doi.org/10.1186/s12870-017-1044-0
Sun BR, Fu CY, Fan ZL, Chen Y, Chen WF, Zhang J, Jiang LQ, Lv S, Pan DJ, Li C (2019a) Genomic and transcriptomic analysis reveal molecular basis of salinity tolerance in a novel strong salt-tolerant rice landrace Changmaogu. Rice 12:99. https://doi.org/10.1186/s12284-019-0360-4
Sun J, Xie D, Zhang E, Zheng H, Wang J, Liu H, Yang L, Zhang S, Wang L, Zou D (2019b) QTL mapping of photosynthetic-related traits in rice under salt and alkali stresses. Euphytica 215:147. https://doi.org/10.1007/s10681-019-2470-x
Takagi H, Abe A, Yoshida K, Kosugi S, Natsume S, Mitsuoka C, Uemura A, Utsushi H, Tamiru M, Takuno S, Innan H, Cano LM, Kamoun S, Terauchi R (2013) QTL-seq: rapid mapping of quantitative trait loci in rice by whole genome resequencing of DNA from two bulked populations. Plant J 74:174–183. https://doi.org/10.1111/tpj.12105
Takagi H, Tamiru M, Abe A, Yoshida K, Uemura A, Yaegashi H, Obara T, Oikawa K, Utsushi H, Kanzaki E, Mitsuoka C, Natsume S, Kosugi S, Kanzaki H, Matsumura H, Urasaki N, Kamoun S, Terauchi R (2015) MutMap accelerates breeding of a salt-tolerant rice cultivar. Nat Biotechnol 33:445–449. https://doi.org/10.1038/nbt.3188
Thomson MJ, Ocampo MD, Egdane J, Rahman MA, Sajise AG, Adorada DL, Tumimbang-Raiz E, Blumwald E, Seraj ZI, Singh RK (2010) Characterizing the saltol quantitative trait locus for salinity tolerance in rice. Rice 3:148–160
Tiwari S, Sl K, Kumar V, Singh B, Rao AR, Mithra SVA, Rai V, Singh AK, Singh NK (2016) Mapping QTLs for salt tolerance in rice (Oryza sativa L.) by bulked segregant analysis of recombinant inbred lines using 50K SNP Chip. PLoS ONE 11:e0153610. https://doi.org/10.1371/journal.pone.0153610
Tsai YC, Chen KC, Cheng TS, Lee C, Lin SH, Tung CW (2019) Chlorophyll fluorescence analysis in diverse rice varieties reveals the positive correlation between the seedlings salt tolerance and photosynthetic efficiency. BMC Plant Biol 19:403. https://doi.org/10.1186/s12870-019-1983-8
Van Ooijen JW (2006) Joinmap 4, software for the calculation of genetic linkage maps in experimental population. Kyazma B.V., Wageningen, Netherlands
Wang S, Basten CJ, Zeng ZB (2007) Windows QTL cartographer 2.5. Department of Statistics North Carolina State University, Raleigh
Wang Z, Wang J, Bao Y, Wu Y, Zhang H (2011) Quantitative trait loci controlling rice seed germination under salt stress. Euphytica 178:297–307. https://doi.org/10.1007/s10681-010-0287-8
Wang Z, Cheng J, Chen Z, Huang J, Bao Y, Wang J, Zhang H (2012) Identification of QTLs with main, epistatic and QTL × environment interaction effects for salt tolerance in rice seedlings under different salinity conditions. Theor Appl Genet 125:807–815. https://doi.org/10.1007/s00122-012-1873-z
Acknowledgements
We thank Mr. Risheng Chen for providing Haidao 86 seeds. We also thank MDPI author service (https://www.mdpi.com/authors/english#English_Editing_Services) for special editing the English text of a draft of this manuscript.
Funding
This work was supported by Grants from Agricultural Variety Improvement Project of Shandong Province (2019LZGC003), Strategic Priority Research Program of the Chinese Academy of Sciences (XDA24030101-6), The National Science Fund for Young Scholars (31700251), Young Talents Training Program of Shandong Academy of Agricultural Sciences, The Youth Scientific Research Foundation of Shandong Academy of Agricultural Sciences (2016YQN10), and Science and Technology Innovation Program of Shandong Academy of Agricultural Sciences (CXGC2018E16 and CXGC2019G02).
Author information
Authors and Affiliations
Contributions
XX and LX designed the experiments and analyzed the data. LX, CZ, WL, and MP performed most experiments, with assistance of GZ, WS, XW, and XZ. LX and WL performed the genome sequence analyses. XX and LX wrote the paper. All authors read and commented on the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Alexander Christmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xie, L., Zheng, C., Li, W. et al. Mapping and Identification a Salt-Tolerant QTL in a Salt-Resistant Rice Landrace, Haidao86. J Plant Growth Regul 41, 2347–2358 (2022). https://doi.org/10.1007/s00344-021-10448-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00344-021-10448-6