Abstract
Anthocyanins are secondary metabolites derived from the general phenylpropanoid pathway and are widespread throughout the plant kingdom. Anthocyanin accumulation is regulated by transcription factors. However, little is known about the role of CaMYC from the bHLH family in pepper anthocyanin synthesis. In this study, a purple-leaved pepper (Capsicum annuum L.) was selected and subjected to light and cold stress at the seedling stage. The result showed that anthocyanin content in purple leaves was higher than in green leaves, which was related to anthocyanins synthesis. CaMYC is an important factor and may be responsible for anthocyanin synthesis. Light and cold stress can result in strong anthocyanin accumulation. CaMYC plays a key role in increasing structural gene expression, which regulates anthocyanin synthesis by the combination of CaMYB and CaWD40. Further analysis showed that CaMYC-silencing resulted in low expression of structural genes in the anthocyanin synthesis pathway, suggesting that CaMYC has a positive role in pepper anthocyanin metabolism. Our study will provide an insight for anthocyanin synthesis in pepper.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Anthocyanins are colored water-soluble pigments belonging to the phenolic group, and primarily responsible for the colors, such as red, purple, and blue, in fruits and vegetables (Raghvendra and Shakya 2011; Khoo et al. 2017). Owing to its importance in plant metabolism and breeding programs, researchers have paid more and more attention to anthocyanin synthesis in recent years (Cheng et al. 2018). Anthocyanins possess various bioactivities, and give a distinctive and attractive color to plant organs including leaves (Zhao et al. 2017), flowers (Tatsuzawa et al. 2012) and fruits (Shiomi et al. 1996). Thus, anthocyanin-rich plants have attracted increasing attention in planta for their special biological value (Gao 2010).
Recently, the anthocyanin biosynthesis pathway has been widely described. All structure genes in the pathway, including chalcone synthase (CHS), chalcone isomerase (CHI), flavanone 3-hydroxylase (F3H), flavonoid 3′5′-hydroxylase (F3′5′H), dihydrolavonols 4-reductase (DFR), anthocyanin synthase (ANS), UDP-glucose: flavonoid 3-glucosyltransferase (UFGT), anthocyanin permease (ANP) and glutathione S-transferase (GST), have been identified in some crops (Lalusin et al. 2006; Liu et al. 2013a). Anthocyanin content is closely related to control of expression patterns of structural genes, which in turn is reflected in the color of plant organs/tissues (Tiffin et al. 1998; Boulton 2001). These structure genes usually are regulated by multiple transcription factors, especially MYB, bHLH, and WD40 proteins (Liu et al. 2013b). The three transcription factors can form a regulatory complex MYB-bHLH-WD40 (MBW) by interacting with each other to directly regulate the transcription of the structural genes (Spelt et al. 2000; Petroni and Tonelli 2011; see EMS_1, Liu et al. 2013a, b). Anthocyanin production is transcriptionally regulated by the MBW complex which controls anthocyanin levels by activating the late biosynthetic genes (Li 2014). The interaction between these transcription factors and the structure genes can protect plants from environmental factors including light and temperature (Ambawat et al. 2013). In general, the synthesis of anthocyanins can be induced or upregulated intensively by the exposure of plants to environmental stressors as diverse as high irradiance, extremes of temperature and high light (Shao et al. 2008; Peng et al. 2015). Light has been thought to be an important factor affecting anthocyanins synthesis. It can regulate anthocyanin levels by controlling the stability of transcriptional factors in the process of anthocyanin synthesis (Maier et al. 2013). Cominelli et al. (2008) reported that, in Arabidopsis, expression of CHS, DFR, and F3H are significantly up-regulated under white light induction. In contrast, mutants lacking the capacity for anthocyanin synthesis exhibit sensitivity to light (Yang et al. 2017). These data suggest the positive role of light in anthocyanin metabolism. Similarly, temperature as another abiotic stress also plays a key role in anthocyanin synthesis. Low temperature can induce up-regulatory expression of anthocyanins synthesis genes and transcriptional factors, such as DFR, CHS, PAL, MYB and so on (Islam et al. 2005; Zhang et al. 2012). Overexpressing the MYB gene leads to high accumulation of anthocyanin and enhanced resistance to chilling and oxidative stress. This was also confirmed by Christie et al. (1994), who reported that anthocyanin pigmentation was higher under cold stress than under normal conditions. Furthermore, there was a difference in anthocyanins levels at different growth stages. Along with plant growth, the expression of key genes involved in anthocyanin biosynthesis, including CHS, CHI, and F3H, vary (Honda et al. 2002).
Pepper (Capsicum annuum L.) is an important vegetable crop grown and consumed worldwide (Visser 2014). Pepper from anthocyanin-rich cultivars can become specialty peppers with high anti-oxidant activity (Sharma et al. 2016) and have the enormous potential to develop some anthocyanin-rich new pepper cultivars (Kim et al. 2017a). Taylor (2014) has extracted, separated and characterized anthocyanins in purple leaves of colored pepper cultivars. Therefore, color-leaved peppers have been popular with consumers recently due to much higher economic, viewing and desirable values. A previous study showed that anthocyanin content in pepper is closely related to the regulation of transcription factors (Borovsky et al. 2004), and the characteristics of some transcription factors including MYB and WD40 were identified in pepper. Silencing of these transcription factors intensively inhibited anthocyanin synthesis (Aguilar-Barragán and Ochoa-Alejo 2014). A recent study on pepper has reported that anthocyanin structural gene transcription requires the expression of at least one member of each of three transcription factor families, MYC, MYB, and WD40. These transcription factors form a complex that binds to structural gene promoters, thereby modulating gene expression (Stommel et al. 2009). However, these studies were limited to the MBW complex and the role of these transcriptional factors in pepper metabolism remains ambiguous. In a previous study, we found a new transcription factor, MYC, from the bHLH family in pepper plants, but little is known about its central role in anthocyanin synthesis and its response to stress. In this article, we investigate the role of MYC in anthocyanin synthesis, and analyze its response to light and cold stresses in pepper. Our study may give insight on the regulatory mechanism of CaMYC by which we may understand anthocyanin synthesis in the future.
Materials and Methods
Plant Materials and Growth Conditions
Z1 (Capsicum annuum), a purple-leaved cultivar, was provided by the College of Horticulture, Northwest A&F University, Yang Ling (34°16′N, 108°4′W). Flowers and fruits of the pepper cultivar were purple throughout the growth period (EMS_2). However, during seedling growth, color at the first to sixth leaf positions is green, whereas leaf color is purple at other positions. Pre-germinated seeds were sown in pots under dark conditions at 28 °C for 2 days. The seedlings were grown in a growth cabinet with a 12-h day/night (d/n) cycle at 60/80% (d/n) relative humidity, 23/18 °C (d/n) temperatures and 300 µmol m−2 s−1 light intensity. The experiments were conducted in the growth cabinet and commenced when the plants reached the 6-leaf to 8-leaf stage (seedling stage). The youngest fully expanded leaves of the seedlings, which developed during stress treatment, were randomly selected for measurement of contributed parameters.
Stress Treatments
Z1 seedlings at the 6-leaf stage were exposed to cold stress (4 °C) for 2 days, and young leaves were sampled at 0, 3, 6, 12, 24 and 48 h. For the treatment of light stress, pepper seedlings were grown under low light (< 300 µmol m−2 s−1), moderate light (300 µmol m−2 s−1) and high light (500 µmol m−2 s−1) for 48 h. The samples were collected 48 h later. The design was completely randomized within the growth chamber with three replicates per treatment.
RNA Extraction, and Quantitative Real-Time PCR
Total RNA was extracted using a Plant RNA Kit (OMEGA) according to the manufacturer’s instructions, and then reverse transcription was performed using the Primescript™ first strand cDNA Synthesis Kit (TaKaRa, Dalian, China). Quantitative real-time PCR (qRT-PCR) was implemented according to the method of Ali et al. (2018) and Guo et al. (2014). The ubiquitin-conjugating gene CaUbi3 (AY486137) was used as the reference gene for pepper (Wan et al. 2011). The gene-specific primers used for expression analysis are listed in EMS_2. The relative expression levels were calculated following the 2−ΔΔCT method (Livak and Schmittgen 2001).
Extraction and Content Determination of Anthocyanin
The samples were collected at the first to sixth leaf position. Purple- and green-colored leaves were separated into two groups; the green-leaf group and the purple-leaf group, respectively. Anthocyanin content was measured according to the method described by Xiang et al. (2015) with a little modification. For measurement of anthocyanin in pepper leaves, frozen leaves were ground to a powder, extracted with the extracting solution (3 M HCl-H2O-methanol solution with the volume ratio of 1:3:16) overnight at 4 °C in the dark. After centrifugation for 10 min, the supernatant was measured at the absorbance of 530 and 657 nm. The relative anthocyanin content in pepper leaves was calculated on a per g fresh weight basis by the formula [(A530—A657)/mg FW tissue] × 1000.
Virus-Induced Gene Silencing (VIGS) of CaMYC in Pepper Plants
The optimal tobacco rattle virus (TRV)-based VIGS system was employed to silence CaMYC expression in pepper purple-leaved line Z1 according to the method of Wang et al. (2013). CaPDS has been used as a visual marker for the effectiveness of VIGS in pepper plants. A fragment of the coding region of CaPDS was amplified using the gene-specific primers VCaPDSF with an EcoRI restriction site and Reverse VCaPDSR with a BamHI restriction site. The resulting product was inserted into a TRV2 vector to generate TRV2:CaPDS.
A fragment of the coding region of CaMYC was amplified using the gene-specific primers with restriction sites. The obtained product was inserted into a TRV2 vector to generate TRV2:CaMYC. The TRV1, TRV2, TRV2:CaPDS, and TRV2:CaMYC vectors were individually transformed into the Agrobacterium tumefaciens strain GV3101. The Agrobacterium strain GV3101 carrying TRV1 was separately mixed with TRV2, TRV2:CaPDS, the empty vector TRV2:00, and TRV2:CaMYC at a 1:1 ratio. The suspensions of the agrobacterium inoculation containing TRV1, TRV2, TRV2:CaPDS, and TRV2:CaMYC (OD600 = 1.0) were infiltrated into the fully expanded cotyledons of pepper plants (Z1 cultivar) using a 1.0-mL sterilized syringe without a needle. The agrobacterium-inoculated pepper plants were grown in a growth chamber at 18 °C in the dark for 2 days with 45% relative humidity, and then transferred into a growth chamber at 22 °C under a 16-h light/8-h dark photoperiod cycle with 60% relative humidity. The primers for VIGS and real-time quantitative PCR are listed in EMS_3.
Expression Analysis for Anthocyanins-Related Genes in Pepper
When the photobleaching symptom appeared on the leaves of the pepper seedlings inoculated with TRV2:CaPDS after 1 month, the young leaves of TRV2:CaMYC and TRV2:00 were sampled for the testing of silencing efficiency and gene expression. Total RNA was isolated using the method of Guo et al. (2012), and cDNA was synthesized using the PrimeScript Kit (Takara) according to the manufacturer’s instructions. Primers were designed to generate 150- to 250-bp fragments using PRIMER5 software. Primers for real-time quantitative PCR (qRT-PCR) are listed in EMS_2. The ubiquitin-conjugating gene CaUbi3 (AY486137) was used as the reference gene for pepper (Wan et al. 2011). PCR and detection were performed as described previously (Feng et al. 2012). The 2−△△CT method was used to analyze the relative transcript levels in gene expression with the means from three replications. The qRT-PCR process was performed in a qRT-PCR instrument (iQ5; BIO-RAD, USA). A completely random design was used in this experiment, and the treatment for each organ was conducted with three biological replicates with each replicate containing five pepper seedlings.
Data Analysis
The treatments were arranged in a randomized complete block design with three biological replicates. The data were analyzed using a one-way analysis of variance (ANOVA) as implemented in SPSS 11.0 software. Statistical significance was inferred at p < 0.05. All figures were edited by Sigma plot 11.0. The analyzed data are presented as means ± SE of three replicates for all measured parameters.
Results
Anthocyanin Content in Pepper Leaves under Normal Condition
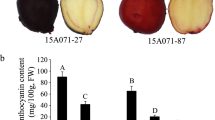
Figure 1A show the performance of Z1 pepper cultivars under normal conditions. Leaf color at the first to sixth positions of Z1 are purple-green under normal light. At normal light intensities, the anthocyanin per unit leaf area of purple-colored leaves was 0.20 mg/g, which was about twofold higher than that of green-colored leaves (Fig. 1B). This result suggested that the presentation of purple color in the leaves of pepper is related to anthocyanin content.
Anthocyanin content in the leaves of pepper at the seedling stage. A Plants at the seedling stage. B Anthocyanin content. Vertical bars represent the mean ± SD of three independent biological replicates. The leaves were sampled at the 6-leaf to 8-leaf stage and each treatment contains three pepper seedlings
Transcript levels of genes involved in anthocyanin synthesis in pepper leaves were investigated by real-time quantitative PCR. The results showed that all the investigated structural genes, especially CaCHS, CaCHI, CaF3H, CaF3′5′H, CaDFR, CaANS, CaUFGT, CaANP, and CaGST, were expressed in leaves, and more strongly in the purple-leaved group as compared to in the green-leaved group (Fig. 2). As for transcription factor, CaMYC expression in the purple-leaved group was significantly higher than that in the green-leaved group, whereas no differences in levels of CaMYB and CaWD40 expression were observed between the purple-leaved group and green-leaved groups. This result indicated that the bHLH transcription factor CaMYC may participate in regulation of these structural genes.
Expression of genes involved in anthocyanin synthesis. The leaves of pepper were collected at the 6-leaf to 8-leaf stage. The expression of all genes was normalized by that of the ubiquitin-con-jugating protein gene CaUbi3. The experiment was conducted with three biological replicates and each replicate contained three pepper seedlings. Error bars represent the mean ± SD of three independent biological replicates
Transcript Levels of Regulatory Genes under Light Treatment
Under different light treatments, the differences in anthocyanin content might be the result of differences in levels of regulatory gene expression. Along with the increasing of light intensity, leaf color turned purple gradually, and the purple color in the high-light treatment was much darker than in the moderate- and low-light treatments (Fig. 3A). Correspondingly, under the treatment of high light intensity, an obvious increase in anthocyanin content was observed as compared to in low-light and moderate-light treatments (Fig. 3B). This result indicated that anthocyanin synthesis was regulated by light intensity and that high light promoted accumulation of anthocyanins.
Effect of light intensity on anthocyanin content of pepper leaves. A Performance of the leaves of peppers at the 6-leaf to 8-leaf stage under different light intensities. B Anthocyanins content. For the treatment of light intensity, the whole seedlings at the 6-leaf to 8-leaf stage were treated with low (< 300 µmol·m−2 s−1), middle (300 µmol m−2 s−1) and high light intensities (500 µmol m−2 s−1) for 48 h. Each treatment contains three pepper seedlings. Error bars represent SD for three biological replicates
To understand the involvement of regulatory genes in purple-leaved pepper sensitivity to light, the expression of all regulatory and structural genes in anthocyanin synthesis was investigated by qRT-PCR under light stress. The results showed that all genes showed expression when treated with light. High light enhanced the levels in the expression of CaMYB and CaMYC, but more strongly in the latter than in the former. In contrast, under low light treatment, levels in the expression of all regulatory genes including CaMYB, CaMYC and CaWD40 decreased compared to in the moderate light treatment, but the level in the expression of CaMYC is lower. Further analysis suggested that high transcriptional levels of CaMYB and CaMYC under high light intensity lead to improved expression of structural genes (Fig. 4). Furthermore, no obvious change in expression of all genes was observed under moderate light treatment. These data suggested a central role of the regulatory gene CaMYC in anthocyanin synthesis in leaves of pepper exposed to light.
Response of genes involved in anthoycanin synthesis in the leaves of pepper in different light intensity treatments. Seedlings at 6-leaf to 8-leaf stage were subjected to light intensities of low light (< 300 µmol m−2 s−1), moderate light (300 µmol m−2 s−1) or high light (500 µmol m−2 s−1) for 48 h. Each treatment contains three pepper seedlings. Vertical bars represent standard deviations (SD, n = 3). Different letters denote significant differences between low light, moderate light and high light intensities in purple-leaved pepper seedlings (p < 0.05)
Accumulation of Anthocyanin in Pepper Leaves under Cold Stress
To monitor anthocyanin biosynthesis under cold stress, anthocyanin content and genes related to anthocyanin synthesis were measured. Anthocyanin content significantly increased progressively during the hours of treatment. Anthocyanin content of plants exposed to cold stress at 6 h was significantly higher than the control, and the level reached to the highest at 48 h post-cold stress, which is about 1.5-fold times that of the control. This result suggested the positive role of cold stress in anthocyanin synthesis. (Fig. 5).
Effect of cold stress on anthocyanin content in the leaves of pepper. The samples at the 6-leaf to 8-leaf stage were collected at different time points (0, 3, 6, 12, 24, 48 h). Mean values and SDs for three replicates are shown. Error bars represent SD for three biological replicates, and the letters show the significance level at α = 0.05
The transcriptional level of the three members (CaMYB, CaMYC and CaWD40) from the MBW transcription complex was monitored by real-time quantitative PCR. Leaves with cold stress treatment showed an obvious increase in the level of all regulatory gene expression at the early stage (6 h post-cold stress), whereas the transcriptional level of CaMYC improved continuously at the following late stage (12 h post-cold stress). Further analysis showed that the relative expression of the CaMYB gene was reduced 24 h after cold stress treatment (Fig. 6A). As for CaMYC expression, a continuous increase was observed throughout the entire process of cold stress (Fig. 6B). Similarly, leaves also possessed a high CaWD40 expression level 6 h after cold stress, but a low expression then increased expression at the late stage (Fig. 6C). To understand the involvement of these transcriptional factors in anthocyanin synthesis, we also investigated the expression of structural genes at the early stage (6 h post-cold stress). Levels in the expression of most of all investigated structural genes were increased at 6 h post-treatment as compared to the control (0 h) (EMS_4). The result suggests that the MBW complex might respond to cold stress at the early stage of cold stress treatment, whereas the CaMYC gene can respond to a low-temperature signal chronically.
Expression profiles of regulatory genes related to anthocyanin synthesis in response to cold stress. A–C Seedlings at the 6-leaf to 8-leaf stage were exposed to 4 °C, and levels in the expression of ACaMYB, BCaMYC and CCaWD40 were investigated by qRT-PCR. The samples were collected at different time points (3, 6, 12, 24, 48 h) with three replicates. Error bars represent SDs for three biological replicates
Anthocyanins-Related Gene Expression in CaMYC-Silencing Peppers
To understand its involvement in pepper anthocyanin synthesis, CaMYC expression was silenced through the virus-induced gene silencing (VIGS) technique, in which the Agrobacterium stain harboring TRV2:00, TRV2:CaMYC and TRV2:CaPDS were injected into cotyledons of pepper purple-leaved line Z1, respectively (Fig. 7A). After 1 month of injection, photobleaching was presented on the leaves of the positive control with TRV2:CaPDS, and obvious green leaves were observed in the silenced peppers with the TRV2:CaMYC vector. However, no obvious symptoms were observed on pepper seedlings with TRV2:00 (Fig. 7B). These data suggest the reliability of the VIGS technique in silencing pepper gene expression, which is further confirmed by a high silencing efficiency of CaMYC gene expression after 1 month of injection (Fig. 7C).
TRV-mediated silencing of CaMYC in purple-leaved pepper plants. A Phenotypes of pepper plants infiltrated with TRV2: CaPDS construct (the positive control). B Leaves in CaMYC-silencing plants exhibiting varying phenotypes in comparison to the negative control with TRV2:00 vector. C Silencing efficiency in the seedlings with TRV2:CaMYC vector. The silencing efficiency was analyzed by qRT-PCR. The experiment was conducted after photobleaching was presented on the leaves of positive control with TRV2:CaPDS. Values are the means ± SD from three separate experiments
Real-time quantitative PCR was conducted to measure transcription levels of genes involved in anthocyanin synthesis. The result showed that CaMYC silencing did not impact CaWD40 expression. Compared to the control of TRV2:00, the CaPAL, CaC4H, and Ca4CL showed higher expression. In contrast, CaCHS, CaCHI, CaF3H, CaF3′5′H, CaDFR, CaANS, CaANP, CaGST, and CaUFGT were repressed after CaMYC-silencing was conducted due to low expression observed in silenced peppers compared with the negative control (Fig. 8).
Expression of genes involved in anthocyanin synthesis in CaMYC-silencing peppers. The samples were collected when photobleaching was observed on the leaves of the positive control with TRV2:CaPDS. TRV2:00: negative control plants; TRV2:CaMYC: CaMYC-silencing plants. The expression of all genes was normalized by that of ubiquitin-con-jugating protein gene CaUbi3. The experiment was conducted with three biological replicates and each replicate contained three pepper seedlings. Error bars represent SD for three biological replicates
Discussion
Optimal environment stress can improve plant adaption to the environment, whereas excess stress results in poor plant growth. However, the importance of anthocyanins for plant development is supported by their utility in the plant kingdom, primarily reflected in resistance to environmental stress and attraction of pollinators (Gould et al. 2000; Shang et al. 2011). As an important pigment, the participation of anthocyanins in the plant response to light and low temperature has been intensively studied in recent years (Lima et al. 2005; Dong et al. 2018). A similar pattern is observed in our study: anthocyanin content is significantly enhanced under high light intensity and cold stress. This may be explained by the result of Xu et al. (2017), who thought that optimal reactive oxygen caused by stress can activate anthocyanin biosynthesis by regulating related genes.
In previous research work, all genes involved in the anthocyanin pathway were identified, and the important role of transcription factors in regulation of anthocyanin biosynthesis in pepper plants was determined (Lightbourn et al. 2007; Stommel et al. 2009). In the pathway of anthocyanin metabolism, the expression profiles of these structural genes were intensively dependent on transcriptional factors. Thus, the interaction between structural genes and transcriptional factors jointly determines anthocyanin synthesis (Ambawat et al. 2013). By further analysis of the expression of these genes in this study, we found that reduction in transcriptional factor gene expression, especially CaMYC, in green leaves may be the main factors to result in down-regulated expression of structural genes. Interestingly, no difference was observed in the levels of the other two transcription factor expressions (CaWD40 and CaMYB) between purple leaves and green leaves. There is a positive relationship between MYC and structural genes in the expression profiles. Therefore, we speculated that regulation of the investigated structural genes may be related to the CaMYC gene, which is similar to a study in wheat (Zong et al. 2017). As an important family, some members from bHLH can bind to the promoters of the anthocyanin biosynthesis genes, such as DFR and UFGT, and the regulatory gene MYB1 to activate their expression (Xie et al. 2017). In Arabidopsis, the ratio of MYC and MYB transcription factors also regulates anthocyanin production at some degree (Ma et al. 2008). These transcriptional factors control anthocyanin synthesis by the regulation of structural genes. These data suggest the positive regulatory function of CaMYC in the regulation of anthocyanin synthesis.
Similar to the results of research in Arabidopsis, light treatment induced high expression of genes related to anthocyanin synthesis (Cominelli et al. 2008). This is also confirmed by Albert et al. (2009), who reported that CHS, F3H, DFR, and UFGT showed down-regulation under low light intensity; even dark treatment resulted in no expression of DFR and ANS genes. In the current study, high light improved CaMYC and CaMYB gene expression, and the highest expression was observed in the CaMYC gene. Correspondingly, structural gene expression was strong in the high-light treatment. A previous study has reported that MYC overexpression reliably lead to high anthocyanin concentrations (Majnik et al. 2000). Light can interact with R2R3-MYBs or bHLHs to organize or disrupt the formation of the MBW complex, leading to enhanced or compromised flavonoid production (Li 2014). Enhanced expression of bHLHs members can result in the accumulation of anthocyanins by interaction with MYBs (Feng et al. 2015). Based on these data, considering the higher expression of CaMYC than CaMYB in peppers, it can be speculated that CaMYC may be responsible for anthocyanin synthesis under light treatment.
To confirm anthocyanin response to low temperature, anthocyanin content and related gene expression were measured in pepper leaves under cold stress. As expected, in our study, anthocyanin content improved under cold stress. When suffering cold stress, anthocyanin genes were induced by cold stress and resulted in the accumulation of anthocyanins (Li et al. 2015). This may be the result of anthocyanins attenuating reactive oxygen caused by cold stress (Kim et al. 2017a, b; Sun et al. 2016). Interestingly, these transcription factors may play a key role in pepper response to cold stress in its early stage (Fig. 6). It is well known that the MBW complex can control anthocyanin products by activating the early biosynthetic steps in plants (Härtl et al. 2017). A previous study also reported that overexpression of the MBW complex induced a high accumulation of anthocyanins both earlier and stronger (Liu et al. 2013b). Therefore, it is possible that these transcription factors are expressed at 6 h after cold stress treatment to improve anthocyanin synthesis.
VIGS is an effective tool for gene function analysis in plants. Over the last decade, VIGS has been used as both a forward and a reverse genetics technique for gene function analysis in various model plants (Ramegowda et al. 2014). To further confirm the contribution of transcription factors on pepper growth, CaMYC silencing was conducted in pepper seedlings. After CaMYC silencing was conducted, the expression of structural genes was decreased in silenced pepper, suggesting a positive role of CaMYC in the anthocyanin synthesis pathway. Although MYC is a protein encoding a bHLH motif and affects anthocyanin accumulation, it is not essential for anthocyanin synthesis (Pattanaik et al. 2008; Zong et al. 2017). A silenced plant can express structural genes related to anthocyanin at a low level. Furthermore, CaMYC controlling anthocyanin synthesis may be correlated with the MBW complex, because CaMYC silencing also resulted in a reduction in levels of CaMYB and CaWD40 expression (EMS_3). Similarly, CaMYB silencing in pepper leaves also showed a similar change (Zhang et al. 2015). Therefore, we speculate that the CaMYC gene may be with a MBW complex, especially CaMYB, to jointly regulate anthocyanins. Although CaMYC is considered to have a key role controlling anthocyanin synthesis, the combined effects and mechanisms of multiple factors on anthocyanin biosynthesis is not clear. Further studies should be focused on the impact of multiple factors on anthocyanin regulatory mechanisms.
Conclusions
Briefly, in this study, we found the importance of anthocyanin-related gene expression in anthocyanin synthesis, thereby demonstrating the close relationship between leaf color and anthocyanins. Regulatory gene expression is positively related to expression of structural genes. Furthermore, anthocyanin content can be induced strongly under high light and cold stress, which is reflected primarily in high levels of genes related to anthocyanin synthesis. CaMYC is a primary regulatory factor in pepper anthocyanin synthesis, and its silencing in pepper resulted in poor anthocyanin accumulation by reducing other structural gene expression. This study provides a better understanding for the role of CaMYC in anthocyanin metabolism. Further study will be focused on the mechanism of interaction between CaMYC and CaMYB/WD40.
References
Aguilar-Barragán A, Ochoa-Alejo N (2014) Virus-induced silencing of MYB and WD40 transcription factor genes affects the accumulation of anthocyanins in chilli pepper fruit. Biol Plant 58:567–574. https://doi.org/10.1007/s10535-014-0427-4
Albert NW, Lewis DH, Zhang H et al (2009) Light-induced vegetative anthocyanin pigmentation in Petunia. J Exp Bot 60:2191–2202. https://doi.org/10.1093/jxb/erp097
Ali M, Luo D, Khan A et al (2018) Classification and genome-wide analysis of chitin-binding proteins gene family in pepper (Capsicum annuum L.) and transcriptional regulation to Phytophthora capsici, Abiotic stresses and hormonal applications. Int J Mol Sci 19:2216. https://doi.org/10.3390/ijms19082216
Ambawat S, Sharma P, Yadav NR, Yadav RC (2013) MYB transcription factor genes as regulators for plant responses: an overview. Physiol Mol Biol Plant 19:307–321. https://doi.org/10.1007/s12298-013-0179-1
Borovsky Y, Oren-Shamir M, Ovadia R et al (2004) The A locus that controls anthocyanin accumulation in pepper encodes a MYB transcription factor homologous to Anthocyanin2 of Petunia. Theor Appl Genet 109:23–29. https://doi.org/10.1007/s00122-004-1625-9
Boulton R (2001) The copigmentation of anthocyanins and its role in the color of red wine: a critical review. Am J Enol Viticul 52:67–87
Cheng GX, Li RJ, Wang M et al (2018) Variation in leaf color and combine effect of pigments on physiology and resistance to whitefly of pepper (Capsicum annuum L.). Sci Hortic 229:215–225. https://doi.org/10.1016/j.scienta.2017.11.014
Christie PJ, Alfenito MR, Walbot V (1994) Impact of low-temperature stress on general phenylpropanoid and anthocyanin pathways: enhancement of transcript abundance and anthocyanin pigmentation in maize seedlings. Planta 194:541–549. https://doi.org/10.1007/BF00714468
Cominelli E, Gusmaroli G, Allegra D et al (2008) Expression analysis of anthocyanin regulatory genes in response to different light qualities in Arabidopsis thaliana. J Plant Physiol 165:886–894. https://doi.org/10.1016/j.jplph.2007.06.010
Dong Y, Qu Y, Qi R et al (2018) Transcriptome analysis of the biosynthesis of anthocyanins in Begonia semperflorens under low-temperature and high-light conditions. Forests 9:1–17. https://doi.org/10.3390/f9020087
Feng H, Li Y, Liu Z, Liu J (2012) Mapping of or, a gene conferring orange color on the inner leaf of the Chinese cabbage (Brassica rapa L. ssp. pekinensis). Mol Breed 29:235–244. https://doi.org/10.1007/s11032-010-9542-x
Feng S, Sun S, Chen X et al (2015) PyMYB10 and PyMYB10.1 interact with bHLH to enhance anthocyanin accumulation in pears. PLoS ONE 10:e0142112. https://doi.org/10.1371/journal.pone.0142112
Gao C (2010) Research advances on the effects of culture conditions on the production of anthocyanin by plant cell culture. J Anhui Agric Sci 38:10–12. (In Chinese)
Gould KS, Markham KR, Smith RH, Goris JJ (2000) Functional role of anthocyanins in the leaves of Quintinia serrata A. Cunn. J Exp Bot 51:1107–1115. https://doi.org/10.1093/jexbot/51.347.1107
Guo WL, Chen RG, Gong ZH et al (2012) Exogenous abscisic acid increases antioxidant enzymes and related gene expression in pepper (Capsicum annuum) leaves subjected to chilling stress. Genet Mol Res 11:4063–4080. https://doi.org/10.4238/2012.September.10.5
Guo M, Zhai YF, Lu J-P et al (2014) Characterization of CaHsp70-1, a pepper heat-shock protein gene in response to heat stress and some regulation exogenous substances in Capsicum annuum L. Int J Mol Sci 15:19741–19759. https://doi.org/10.3390/ijms151119741
Härtl K, Denton A, Franz-Oberdorf K et al (2017) Early metabolic and transcriptional variations in fruit of natural white-fruited Fragaria vesca genotypes. Sci Rep 7:45113. https://doi.org/10.1038/srep45113
Honda C, Kotoda N, Wada M et al (2002) Anthocyanin biosynthetic genes are coordinately expressed during red coloration in apple skin. Plant Physiol Biochem 40:955–962. https://doi.org/10.1016/S0981-9428(02)01454-7
Islam MS, Jalaluddin M, Garner JO et al (2005) Artificial shading and temperature influence on anthocyanin compositions in sweetpotato leaves. HortSci 40:176–180
Khoo HE, Azlan A, Tang ST, Lim SM (2017) Anthocyanidins and anthocyanins: colored pigments as food, pharmaceutical ingredients, and the potential health benefits. Food Nutr Res 61:1361779. https://doi.org/10.1080/16546628.2017.1361779
Kim J, Park M, Jeong ES et al (2017a) Harnessing anthocyanin-rich fruit: a visible reporter for tracing virus-induced gene silencing in pepper fruit. Plant Methods 13:3. https://doi.org/10.1186/s13007-016-0151-5
Kim JN, Han SN, Ha TJ, Kim H-K (2017b) Black soybean anthocyanins attenuate inflammatory responses by suppressing reactive oxygen species production and mitogen activated protein kinases signaling in lipopolysaccharide-stimulated macrophages. Nutr Res Prac 11:357–364. https://doi.org/10.4162/nrp.2017.11.5.357
Lalusin AG, Nishita K, Kim S-H et al (2006) A new MADS-box gene (IbMADS10) from sweet potato (Ipomoea batatas (L.) Lam) is involved in the accumulation of anthocyanin. Mol Genet Genom 275:44–54. https://doi.org/10.1007/s00438-005-0080-x
Li S (2014) Transcriptional control of flavonoid biosynthesis: fine-tuning of the MYB-bHLH-WD40 (MBW) complex. Plant Signal Behav 9:e27522. https://doi.org/10.4161/psb.27522
Li SJ, Bai YC, Li CL et al (2015) Anthocyanins accumulate in tartary buckwheat (Fagopyrum tataricum) sprout in response to cold stress. Acta Physiol Plant 37:1–8
Lightbourn GJ, Stommel JR, Griesbach RJ (2007) Epistatic interactions influencing anthocyanin gene expression in Capsicum annuum. J Am Soc Hortic Sci 132:824–829
Lima VLAG, De Mélo EDA, Lima DEDS (2005) Efeito da luz e da temperatura de congelamento sobre a estabilidade das antocianinas da pitanga roxa. Cienc Tecnol Aliment 25:92–94. https://doi.org/10.1590/S0101-20612005000100015
Liu X, Feng C, Zhang M et al (2013a) The MrWD40-1 gene of Chinese bayberry (Myrica rubra) interacts with MYB and bHLH to enhance anthocyanin accumulation. Plant Mol Biol Rep 31:1474–1484. https://doi.org/10.1007/s11105-013-0621-0
Liu X, Li F, Yin X, Kunsong C (2013b) Research progress on transcriptional regulation of anthocyanin biosynthesis. Acta Hortic Sin 40:2295–2306
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods (San Diego Calif) 25:402–408. https://doi.org/10.1006/meth.2001.1262
Ma H, Pooler M, Griesbach R (2008) Ratio of Myc and Myb transcription factors regulates anthocyanin production in orchid flowers. J Am Soc Hortic Sci 133:133–138.
Maier A, Schrader A, Kokkelink L et al (2013) Light and the E3 ubiquitin ligase COP1/SPA control the protein stability of the MYB transcription factors PAP1 and PAP2 involved in anthocyanin accumulation in Arabidopsis. Plant J 74:638–651. https://doi.org/10.1111/tpj.12153
Majnik J, Djordjevi MA, Rolf BG et al (2000) Anthocyanin regulatory gene expression in transgenic white clover can result in an altered pattern of pigmentation. Aust J Plant Physiol 7:659–667
Pattanaik S, Xie CH, Yuan L (2008) The interaction domains of the plant Myc-like bHLH transcription factors can regulate the transactivation strength. Planta 227:707–715. https://doi.org/10.1007/s00425-007-0676-y
Peng L, Zou L, Su Y et al (2015) Effects of light on growth, levels of anthocyanin, concentration of metabolites in Fagopyrum tataricum sprout cultures. Int J Food Sci Technol 50:1382–1389. https://doi.org/10.1111/ijfs.12780
Petroni K, Tonelli C (2011) Recent advances on the regulation of anthocyanin synthesis in reproductive organs. Plant Sci 181:219–229. https://doi.org/10.1016/j.plantsci.2011.05.009
Raghvendra SV, Shakya A et al (2011) Chemical and potential aspects of anthocyanins—a water-soluble vacuolar flavonoid pigments: a review. Int J Pharm Sci Rev Res 6:28–33
Ramegowda V, Mysore KS, Senthil-Kumar M (2014) Virus-induced gene silencing is a versatile tool for unraveling the functional relevance of multiple abiotic-stress-responsive genes in crop plants. Front Plant Sci 5:323. https://doi.org/10.3389/fpls.2014.00323
Shang Y, Venail J, Mackay S et al (2011) The molecular basis for venation patterning of pigmentation and its effect on pollinator attraction in flowers of Antirrhinum. New Phytol 189:602–615. https://doi.org/10.1111/j.1469-8137.2010.03498.x
Shao L, Shu Z, Peng CL et al (2008) Enhanced sensitivity of Arabidopsis anthocyanin mutants to photooxidation: a study with fluorescence imaging. Funct Plant Biol 35:714–724. https://doi.org/10.1071/fp08069
Sharma V, Chandel C, Kumar R et al (2016) Influence of planting time and fruit maturity stage on antioxidant activity, phenols and anthocyanin contents in sweet pepper. Res Crops 17:360. https://doi.org/10.5958/2348-7542.2016.00061.9
Shiomi S, Wamocho LS, Agong SG (1996) Ripening characteristics of purple passion fruit on and off the vine. Postharvest Biol Technol 7:161–170. https://doi.org/10.1016/0925-5214(95)00023-2
Spelt C, Quattrocchio F, Mol JN, Koes R (2000) anthocyanin1 of petunia encodes a basic helix-loop-helix protein that directly activates transcription of structural anthocyanin genes. Plant Cell 12:1619–1632. https://doi.org/10.1105/tpc.12.9.1619
Stommel JR, Lightbourn GJ, Winkel BS, Griesbach RJ (2009) Transcription factor families regulate the anthocyanin biosynthetic pathway in Capsicum annuum. J Am Soc Hortic Sci 134:244–251
Sun W, Wang Z, Cao J et al (2016) Cold stress increases reactive oxygen species formation via TRPA1 activation in A549 cells. Cell Stress Chaperon 21:367–372. https://doi.org/10.1007/s12192-015-0663-3
Tatsuzawa F, Aiba Y, Morino T et al (2012) Copigmentation with acylated anthocyanin and kaempferol glycosides in violet and purple flower cultivars of Aubrieta × cultorum (Brassicaceae). J Jap Soc Hortic Sci 81:275–284. https://doi.org/10.2503/jjshs1.81.275
Taylor CL (2014) Purple pepper plants, An anthocyanin powerhouse: extraction, Separation and Characterization. Dissertations & Theses, Gradworks
Tiffin P, Miller RE, Rausher MD (1998) Control of expression patterns of anthocyanin structural genes by two loci in the common morning glory. Genes Genet Syst 73:105–110. https://doi.org/10.1266/Ggs.73.105
Visser L (2014) Development of a pepper (Capsicum annuum L.) hybrid variety with resistance to potato virus Y (PVY) using molecular breeding. University of KwaZulu-Natal
Wan H, Yuan W, Ruan M et al (2011) Identification of reference genes for reverse transcription quantitative real-time PCR normalization in pepper (Capsicum annuum L.). Biochem Biophys Res Commun 416:24–30. https://doi.org/10.1016/j.bbrc.2011.10.105
Wang J-E, Li D-W, Gong Z-H, Zhang Y-L (2013) Optimization of virus-induced gene silencing in pepper (Capsicum annuum L.). Genet Mol Res 12:2492–2506. https://doi.org/10.4238/2013.july.24.4
Xiang L, Liu X, Li X et al (2015) A novel bHLH transcription factor involved in regulating anthocyanin biosynthesis in Chrysanthemums (Chrysanthemum morifolium Ramat.). PLoS ONE 10:e0143892. https://doi.org/10.1371/journal.pone.0143892
Xie X, Zhao J, Hao Y-J et al (2017) The ectopic expression of apple MYB1 and bHLH3 differentially activates anthocyanin biosynthesis in tobacco. Plant Cell Tissue Org 131:183–194. https://doi.org/10.1007/s11240-017-1275-7
Xu Z, Mahmood K, Rothstein SJ (2017) ROS induces anthocyanin production via late biosynthetic genes and anthocyanin deficiency confers the hypersensitivity to ROS-generating stresses in arabidopsis. Plant Cell Physiol 58:1364–1377. https://doi.org/10.1093/pcp/pcx073
Yang JF, Chen YZ, Kawabata S et al (2017) Identification of light-independent anthocyanin biosynthesis mutants induced by ethyl methane sulfonate in Turnip “Tsuda” (Brassica rapa). Int J Mol Sci 18:1288. https://doi.org/10.3390/ijms18071288
Zhang B, Hu Z, Zhang Y et al (2012) A putative functional MYB transcription factor induced by low temperature regulates anthocyanin biosynthesis in purple kale (Brassica Oleracea var. acephala f. tricolor). Plant Cell Rep 31:281–289. https://doi.org/10.1007/s00299-011-1162-3
Zhang Z, Li D-W, Jin J-H et al (2015) VIGS approach reveals the modulation of anthocyanin biosynthetic genes by CaMYB in chili pepper leaves. Front Plant Sci 6:500. https://doi.org/10.3389/fpls.2015.00500
Zhao Z, Xiao L, Xu L et al (2017) Fine mapping the BjPl1 gene for purple leaf color in B2 of Brassica juncea L. through comparative mapping and whole-genome re-sequencing. Euphytica 213:80. https://doi.org/10.1007/s10681-017-1868-6
Zong Y, Xi X, Li S et al (2017) Allelic variation and transcriptional isoforms of wheat TaMYC1 gene regulating anthocyanin synthesis in pericarp. Front Plant Sci 8:1645. https://doi.org/10.3389/fpls.2017.01645
Acknowledgements
This work was supported through funding from the National Natural Science Foundation of China (No. U1603102), the National Key R&D Program of China (No. 2016YFD0101900), and the Independent Innovation Fund Project of Agricultural Science and Technology in Jiangsu [No.CX (17) 3040].
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Bo-Ya Lu and Guo-Xin Cheng: co-first authors in this paper.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lu, BY., Cheng, GX., Zhang, Z. et al. CaMYC, A Novel Transcription Factor, Regulates Anthocyanin Biosynthesis in Color-leaved Pepper (Capsicum annuum L.). J Plant Growth Regul 38, 574–585 (2019). https://doi.org/10.1007/s00344-018-9871-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00344-018-9871-2