Abstract
Chalcone synthase (CHS, EC 2.3.1.74) is a member of the plant polyketide synthase superfamily; it catalyzes the first committed step in the flavonoid biosynthetic pathway. In this study, we identified and characterized five CHS superfamily genes from Nicotiana tabacum. Our phylogenetic analysis suggested that the five tobacco CHS genes have diverged into two subgroups, including the classical CHS genes for NtCHS1–4 and CHS-Like genes for NtCHS5. Solanaceae CHSs diverged into different subfamilies prior to the divergence of the genera. We used RNA-seq data to explore the spatial and temporal expression patterns of the NtCHS1–5 genes. NtCHS1–4 were widely expressed in pigmented or/and non-pigmented vegetative tissues and floral organs. NtCHS5 was expressed strongly in the calyx and corolla organs at the full-bloom stage. Quantitative RT-PCR analysis was performed; it verified our RNA-seq results. Further, the expression of NtCHS1 and NtCHS3 increased markedly under drought and high salt stress. NtCHS2 expression and NtCHS4 expression were obviously induced by ABA and MeJA treatment, and NtCHS4 expression was specifically induced by 2,4-D. NtCHS5 expression was not induced by any of these treatments. Our results represent new data about the differentiation of gene expression and environmental responsiveness in the expression of CHS superfamily genes in Nicotiana tabacum.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
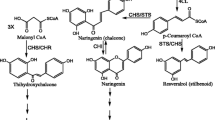
Chalcone synthase (CHS, EC 2.3.1.74) is the first key enzyme of the flavonoid pathway in higher plants. It catalyzes the condensation of one molecule of 4-coumaroyl coenzyme A (CoA) with three molecules of malonyl CoA, yielding chalcone as the product. Chalcone is the entry compound of the first step of flavonoid and anthocyanin biosynthesis. Combined with the same gene superfamily members plant CHS-Like (CHSL) genes, CHS and CHSL genes belong to the plant type III polyketide synthases (PKSs). Type III PKSs play major roles in the biosynthesis of important plant compounds including secondary metabolites. CHSs have been found in all plant species, and these enzymes are known to play diverse and important roles in flowering plants, such as producing floral pigments, antimicrobial compounds, UV protectants, and insect repellents. More than four thousand plant CHSs have been cloned since the first plant CHS was cloned in 1983 (Reimold and others 1983); they have been cloned from species of the Fabaceae, Solanaceae, Brassicaceae, and Gramineae families, among others.
Chalcone synthases are encoded by multigene families in many plants. At least three gerbera CHS genes were described in Gerbera hybrid (Deng and others 2014), five CHS genes were found in Phalaenopsis Orchid (Han and others 2006), six CHS genes were found in Brassica rapa (turnip) (Zhou and others 2013), and eight CHS genes were found in Petunia hybrida (Koes and others 1989). Gene duplication and subsequent positive selection results in the formation of gene families (Flagel and Wendel 2009). Multiple gene family members typically exhibit varied expression patterns in different tissues and different development stages, thus suggesting diverse functionality (Han and others 2006; Zhou and others 2013).
Plants are constantly challenged by a myriad of environmental factors and undergo complex developmental programs. The various functional roles and activities of CHSs are known to be regulated by numerous endogenous and external stimuli. For endogenous regulation, such as flower development (Sun and others 2015), nodulin development (Krause and others 1997), and male sterility (Napoli and others 1999; Budahn and others 2014) could induce the accumulation of CHSs in plants. External stimuli, including UV radiation (Dao and others 2011; Sun and others 2014), wounding (Richard and others 2000), low temperature (Zhou and others 2007), high salt (Chen and others 2015), microbial pathogen attack (Nagy and others 2004; Zabala and others 2006), and treatment with salicylic acid and methyl jasmonate (Schenk and others 2000), can alter the expression of CHSs. CHS superfamily members have evolved to function in varying roles in these complicated developmental and environmental contexts.
In this study, we successfully cloned five CHS (NtCHS1–5) genes from the Honghuadajinyuan (HD) cultivar of Nicotiana tabacum. We then investigated their sequence information and phylogenetic relationships. The spatial and temporal expression patterns of these tobacco CHS gene family members were then evaluated. We also measured the expression levels of NtCHS1–5 under abiotic stress and phytohormone treatments to assess if the expression of these genes was responsive to such signals.
Materials and Methods
Cloning of Chalcone Synthase Genes by PCR
Total RNA was extracted from Honghuadajinyuan (Nicotiana tabacum) leaves using Tripure isolation reagent (Roche, Germany) according to the manufacturer’s instructions. A Transcriptor First Strand cDNA synthesis kit (Roche, Germany) was used for transcribing first-strand cDNA. Genomic DNA was extracted according to the CTAB method. Based on the chalcone synthase (CHS) of tobacco (Nicotiana tabacum) in China Tobacco Genome Database (www.tobaccodb.org), PCR amplification primers (Supplemental Table 1) were designed to obtain the full-length cDNA and the genomic DNA sequence of chalcone synthases. The final volume of the PCR reaction mixture was 50 µl, and included 25 µl PrimSTAR mix, 1 µl forward primer, 1 µl reverse primer, 1 µl template cDNA, and 22 µl sterile water. The following thermal cycling program was used: 2 min at 98 °C, 35 cycles (10 s at 98 °C, 15 s at Tm (listed in Supplemental Table 1), 30 s at 72 °C) and 7 min at 72 °C. PCR products were purified and cloned with the pEASY-Blunt Cloning Kit (TransGen Biotech, China) for sequencing.
Sequence Analysis
Members of the CHS superfamily from solanaceous species and Arabidopsis thaliana were identified and downloaded from NCBI. All sequences were aligned with ClustalW2 (http://www.ebi.ac.uk/Tools/msa/clustalw2/) and the redundant sequences were excluded. The intron–exon structures of the CHS genes were analyzed with GSDS2.0 (http://gsds.cbi.pku.edu.cn). Phylogenetic analysis with a poisson model method was performed using MEGA6.0 with a neighbor-joining method. The robustness of the tree topology was assessed using 1000 bootstrap replicates. The 2K upstream sequences of NtCHS1–5 from the transcriptional start site were selected as the promoter sequences and were downloaded from the China Tobacco Genome Database. Promoter sequences of the CHS genes were analyzed by PLACE (Database of Plant Cis-acting Regulatory DNA Elements, http://www.dna.affrc.go.jp/PLACE/signalscan.html) to identify cis-element motifs.
Plant Materials and Treatments
Seeds of the common tobacco cultivar Honghuadajinyuan (Nicotiana tabacum) were planted on 1/2 strength MS medium in sterile plates and cultured in growth chambers (16 h of light/8 h of dark cycle, 25 °C). For phytohormone treatments, 2-week-old seedlings were transferred to 1/2 strength MS medium containing 10 μM 2,4-D, 10 μM MeJ, or 10 μM ABA, respectively (Liu and others 2014; Concha and others 2013). The seedlings were sampled after 0, 1, 2, and 5 days of treatment, the strict 24-h interval from the time that we started the phytohormone treatments was implemented. For the abiotic stress treatments, tobacco plants were grown in a greenhouse(16 h of light/8 h of dark cycle, 25 °C). Plants that were about one-month-old, with four or five fully expanded leaves, were used in this experiment. Plants were transferred to abluent sand pots prior to treatment with the experimental solutions. The test solutions were PEG 6000 (20%) and NaCl (200 mM). Leaves and roots were harvested after 0, 1, and 2 days of treatment, the strict 24-h interval from the time that we started the abiotic treatments was implemented, according to the method of Chen (2011). All collected samples were frozen immediately with liquid nitrogen and stored at −80 °C.
Expression Profiles of NtCHS1–5 in RNA-seq
Expression levels were measured at five growth stages of common tobacco, including the seedling, rosette, bud, full-bloom, and seed stages. The tissues sampled included: leaf, root, lateral root, fibril, stem, flower, calyx, corolla, stamen, pistil, anther, and seed, and leaves at different positions (counted from the bottom up) including the fifth leaf (L5), the tenth leaf (L10), and the fifteenth leaf (L15). Samples at the seedling stage with seven fully expanded leaves, at the rosette stage with fifteen fully expanded leaves, at the bud stage when the first flower started to bud in the central inflorescence, at the full-bloom stage 19 days after the first flower bloom and the fresh capsules at seed stage, were collected in this experiment. All collected samples were frozen immediately with liquid nitrogen and stored at −80 °C and sequenced using an Illumina HiSeq™ 2000 platform, performed at the Beijing Genomics Institute (BGI, http://www.genomics.cn/index; Shenzhen, China). RPKM values (reads per kilobase of exon model per million mapped reads) were calculated to represent the expression level for each gene. The expression profiles of NtCHS1–5 in various tissues and stages of development in common tobacco were examined with the RNA-seq dataset We then constructed heatmaps—using HemI software (Heatmap Illustrator, version 1.0) (http://hemi.biocuckoo.org/index.php).
Quantitative RT-PCR Analysis (qRT-PCR)
qRT-PCR analysis was used to verify the expression profiles in the RNA-seq dataset and to profile the expression patterns of NtCHS1–5 under treatment with various abiotic stresses and plant hormones. First-strand cDNA was synthesized using a Transcriptor First Strand cDNA Synthesis Kit (Roche, Germany) according to the manufacturer’s instructions. The qRT-PCR was performed using FastStart Universal SYBR Green Master mix (Rox) (Roche, Germany). The qRT-PCR primers are listed in Supplemental Table 1. Tobacco actin gene (NTU60495) served as an internal control. The reaction mixtures were 20 and 10 µl SYBR Green mix, 0.6 µl forward qRT-PCR primer, 0.4 µM reverse primer, 0.8 µM template cDNA, and 6.8 µl sterile water. The thermal cycling parameters were 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 58 °C for 1 min. The comparative cycle threshold (ΔΔC T) method was used to calculate the relative expression levels of the target genes.
Measurement of Flavonoid Content
Flavonoid content was quantified by the NaNO2-Al (NO3)3-NaOH spectrophotometry method (Fu and others 2006; Luo and others 2015), with some modifications. Briefly, approximately 0.6 g of dry tissue was ground to a fine powder in liquid nitrogen and dissolved in 70% ethanol solution and then transferred to a 10-mL volumetric flask to reach the constant volume. The phenolics were extracted for 12 h at room temperature. Each extraction was repeated three times, and the materials were centrifuged at 13,000 rpm for 10 min. The supernatants were then transferred to a 50-mL flask and 70% ethanol solution was added up to the constant volume. Approximately 5 mL of extract were then transferred to a new 10-mL flask, and 30% alcohol was added up to 5.4 mL. To this was added 0.3 mL 5% NaNO3; samples were then held stable for 6 min. Ten percent Al(NO3)3 (0.3 mL) was then added and the samples were held stable for 6 min, after which 4.0 mL of 4% NaOH was added up to a total volume of 10 mL. The samples were then mixed and subsequently held stable for 15 min. The absorbance at 510 nm was measured with a Beckman Coulter DU 640 ultraviolet spectrophotometer. Three measurements for each biological replicate sample were performed. A standard curve was calculated according to the following formula: A = 0.0124C + 0.0068. Rutin was used as the standard to construct the standard curve. The flavonoid content was measured according to the following formula: y = 2440.5x + 2.3804.
Results
Identification of Nicotiana tabacum Chalcone Synthase Family Genes
We successfully isolated five chalcone synthase genes from common tobacco, including the full-length cDNA and the genomic DNA sequence. Basic information about the five CHS family genes, including gene size, chromosomal location, and the predicted sizes and isoelectric points of the proteins they encode, is listed in Table 1. High amino acid level homology was found with other Solanaceae CHS and among the five family members. NtCHS1 was exactly the same in its amino acid sequence as a previously reported chalcone synthase from Nicotiana tabacum (AAK49457). NtCHS2 shares 99% sequence similarity with chalcone synthase 2 (AHK22740), with one amino acid change, Ile(50) to Thr(50). NtCHS3 and NtCHS4 share 96% similarity with the predicted chalcone synthase A (XP_009764938) of Nicotiana sylvestris and the predicted chalcone synthase 2 (XP_009616637.) of Nicotiana tomentosiformis, respectively. NtCHS5 shares almost 81% similarity with a Solanum tuberosum chalcone synthase J-like gene (XP_006367318). There was a 98.46% similarity between NtCHS1 and NtCHS3. NtCHS2 and NtCHS4 share about 97.94% identity. NtCHS5 shares about 72% similar with each of the other four CHSs.
As with the other reported plant CHS genes, all five of the tobacco CHS genes have one intron and diverse intron length was observed (Supplemental Fig. 1). Most of the plant CHSs have Cys in the consensus sequence of (K/Q)R(M/I) C(D/E)KS as the intron splice site (Harashima and others 2004). NtCHS1, 2, 3, and 5 each have one conserved intron at this Cys (T/GT). NtCHS4 contains one intron at Gly (G/GT), either Gly (G/GT) or Cys (T/GT) is a phase-1 intron, as is the case in all plant CHS superfamily genes whose gene structures are known. Conforming to the GT-AG rule, all of the introns begin with the nucleotides ‘GT’ and end with the nucleotides, ‘AG.’
High identity was observed between NtCHS1 and NtCHS3, and also between NtCHS2 and NtCHS4. To confirm if they are from different genomic regions rather than two different transcriptional variants, we compared the intron sequences of NtCHS1 and NtCHS3, NtCHS2 and NtCHS4, respectively. There was 66.13% identity detected between NtCHS1 and NtCHS3, 54.07% identity was detected between NtCHS2 and NtCHS4. Moreover, numbers of insertions or deletions exist between each two introns (Supplemental Fig. 2a, b). The results demonstrated that NtCHS1 and NtCHS3 are from a different genomic region, the same situation was observed with NtCHS2 and NtCHS4.
Variation in Residues at the Active Sites in Tobacco Chalcone Synthases
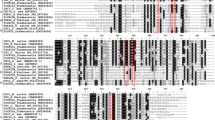
Alignment of the predicted protein sequences of the tobacco CHSs revealed that the five tobacco CHSs contained previously reported active site amino acids (1999; Han and others 2006), including T132, S133, M137, C164, T194, T197, F215, I254, G256, F265, H303, P304, P307, N336, and S338 (Fig. 1). Changes in NtCHS5 at conserved amino acid sites S133, M137, T194, T197, F265, P304, P307, and S338 were detected, whereas NtCHS1–4 did not vary at these sites.
Molecular Evolution of Tobacco Chalcone Synthase Family Genes
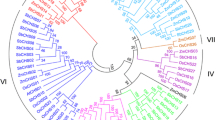
Based on amino acid sequences, a phylogenetic tree of the CHSs was constructed using the neighbor-joining method (Fig. 2). The tree revealed two important features. First, Nicotiana tabacum CHSs were distributed into two subgroups: NtCHS1, 2, 3, and 4 clustered together with the typical CHSs of Solanaceae species and Arabidopsis thaliana and belonged to subgroup I. NtCHS5 clustered with CHSL proteins from Solanum tuberosum and Solanum lycopersicum as subgroup II, indicating that NtCHS5 may be a CHS-like protein. Secondly, the solanaceous CHSs (including those of Solanum tuberosum, Solanum lycopersicum, Capsicum annuum, Iochroma cyaneum, Nicotiana tabacum, and Petunia hybrid) did not form genus-specific clusters. Instead, all the examined solanaceous CHSs clustered into two subgroups. Our phylogenetic analysis suggests that the solanaceous CHSs diverged into two subgroups prior to the divergence of these genera.
Phylogenetic tree based on the amino acid sequences of eudicot CHSs. The CHSs isolated in this study are highlighted as black dots. The tree was constructed using the MEGA 6.0 and neighbor-joining method with 1000 bootstrap replicates. The GenBank accession numbers of the sequences used are as follows: SlyCHS1 (NP_001234033), SlyCHS2 (NP_001234036), SlyCHSB (XP_004239898), SlyCHSJ (XP_010322085), SlyCHSJ-L (XP_010314857), from Solanum lycopersicum, StCHS2 (NP_001275296), StCHS1A (NP_001275352), StCHSJ-L1 (XP_006367318), StCHSJ-L2 (XP_006349940), StCHSG-L (XP_006367320), StCHSJ-L (XP_006367319) from Solanum tuberosum, CaCHS (AFL47798) from Capsicum annuum, IcCHS1 (AIY22756), IcCHS2 (AIY22757), IcCHS3 (AIV99854) from Iochroma calycinum, PahCHS1 (AAF60297) and PahCHS2 (CAA32731) from Petunia × hybrid, NtCHS (AHK22740) (AHK22740), NtCHS (AAK49457) (AAK49457), NtCHS1 (Ntab 0575960), NtCHS2 (Ntab 0434910), NtCHS3 (Ntab 0206900), NtCHS4 (Ntab 0320210), NtCHS5 (Ntab 0357400) from Nicotiana tabacum, and AtCHS (AED91961) from Arabidopsis thaliana
Expression Profiles of NtCHS1–5 in Various Tissues and Stages
The spatial and temporal expression patterns of the NtCHS1–5 genes were analyzed. Figure 3a showed the expression of the CHS genes in various tissues and stages of development in tobacco, as determined in the RNA-seq data.
CHS gene expression patterns in various tissues and stages of development in tobacco. a CHS gene expression patterns data was determined in RNA-seq. Expression data were measured in five growth stages of common tobacco, including seedling, rosette, bud, full-bloom, and seed stage. Tissues including leaf (L), root (R), lateral root (LR), fibril (FR), stem (S), flower (F), calyx (Ca), corolla (Co), stamen (St), pistil (P), anther (An), and seed (Sd); leaves at different positions which counted from the bottom up, the fifth leaf (L5), the tenth leaf (L10), and the fifteenth leaf (L15). b The relative expression data from the qRT-PCR analysis of the NtCHS1–5 gene. Expression data were measured in two growth stages of common tobacco, including seedling (S) and full-bloom (M) stage. Tissues sampled included leaf (L), stem (S), root (R), bud (B), flower (F), and anther (AN)
It is clear that NtCHS1–4 were widely expressed; their transcripts were detected in almost all of the tissues and all development stages, including in seedling leaves (L_seedling), the floral and calyx of the bud stage (F_bud and Ca_bud), calyx, corolla, and stamen of the full-bloom stage (Ca_F, Co_F, and Stamen_F), and in seed. The expression of NtCHS1–4 in corolla and calyx increased sharply along with the developmental progression of these floral organs; a similar phenomenon was observed previously in orchid and in a freesia hybrid (Han and others 2006; Sun and others 2015). The expression of NtCHS1–4 in leaves varied in the course of the development stages, and gradually decreased as the tobacco plants matured. The expression patterns showed moderate increases from the fifth leaf (L5) to the fifteenth leaf (L15) (counted from down to up), which demonstrated that NtCHS1–4 were expressed at a higher level in young leaves than in older leaves. NtCHS5 was moderately expressed in leaves of almost all the growth stages, but was especially highly expressed in the calyx and the corolla at the full-bloom stage.
To validate the RNA-seq data, real-time PCR was performed. Total RNA from different tissues collected at two developmental stages (Fig. 3b), including the seedling stage and the mature stage, was used to measure the transcription levels of NtCHS1–5. Due to sequence similarity, NtCHS1 and NtCHS3 could not be analyzed individually. The real-time PCR data were in agreement with RNA-seq data. The close correlation of these results indicated that the RNA-seq results were accurate in representing the gene expression levels.
Promoter Region Analysis of NtCHS1–5
Cis-elements (including, for example, TF binding sites) are functional DNA elements that influence temporal and spatial transcriptional activity. The regions in 2K upstream of the transcriptional start sites of the NtCHS1–5 loci were evaluated as ‘promoter regions.’ At the nucleotide sequence level, the five promoters share 50.9–73.8% similarity. A relatively high similarity was found between the promoter regions of NtCHS1 and 3 and between those of NtCHS2 and 4. MEME (Multiple Em for Motif Elicitation) software, with default settings, was used to identify motifs in the promoter regions of NtCHS1–5. Five motifs were found in NtCHS1–5 (Supplemental Fig. 3). We used PLACE (Database of Plant Cis-acting Regulatory DNA Elements) to identify the cis-elements of the promoter sequences.
The promoter regions of NtCHS1–5 were found to contain conserved putative drought- and light-regulated elements, ABRE, and GT. These cis-elements in the promoters of NtCHS1–5 suggest that they may function in plant responses to drought or light stress (Kaplan-Levy and others 2012; Ma and others 2014; Park and others 2004; Vijayan and others 2015). A G-box, an ASF-1 element, and a nodulin consensus motif were detected in NtCHS1–4. The G-box has a binding site for bHLH transcription factors and has been reported to confer salt tolerance in plants (Boter and others 2004; Ahmad and others 2015). The ASF1 element is a bZIP transcription factor binding site and is known to be involved in salicylic acid and auxin-induced regulation of gene expression (Niggeweg and others 2000; Hwang and Hwang 2010). CTCTT is one of two nodulin consensus motifs present in the promoter regions of nodulin genes and is known to be involved in nodulin development (Macknight and others 1995).
NtCHS1 and 3 both contain a W-box and a RAV1 element. W-boxes are found in genes that are recognized by WRKY DNA binding proteins, and are known to be involved in the regulation of plant tolerance to pathogens and salt stress (Choi and others 2015; Li and others 2015). The AP2 transcription factor RAV1 is unique in higher plants; it recognizes the RAV1 binding site CAACA motif and is known to modulate plant sensitivity to drought and salt stress (Fu and others 2014; Li and others 2015). NtCHS2 and 4 both have pollen-specific regulator elements (AGAAA) and the phytohormone-response elements GARE, ABRE, and auxin responsive element. AGAAA is one of two co-dependent regulators required for pollen specific expression (Filichkin and others 2004).
Drought and High Salt Stress Responsiveness of NtCHS1–5
Changes in NtCHS1–5 expression in response to drought and high salt stress were assessed by real-time PCR analysis. Leaves and roots of the samples treated with PEG 6000 (20%) or NaCl (200 mM) were collected after 0, 1, and 2 days. Due to sequence similarity, NtCHS1 and NtCHS3 could not be analyzed individually. NtCHS1 and 3 were responsive to both drought and high salt stress (Fig. 4a, b). The expression level of these two genes increased significantly under treatment with PEG 6000 (20%) and NaCl (200 mM). The expression level of NtCHS2 was increased significantly in root, but not in leaf, under PEG 6000 (20%); no significant change in expression was detected for NtCHS2 under NaCl (200 mM). These results suggest that NtCHS2 is responsive to drought stress but is not responsive to high salt treatment. The only significant change in expression observed for NtCHS4 was a significant decrease in leaf tissue under PEG 6000 (20%). NtCHS4 thus has a different expression pattern responsiveness to PEG 6000 treatment than does NtCHS1,3, and NtCHS4 is not responsive to high salt stress. NtCHS5 was not induced by drought or high salt stress.
The expression patterns of the NtCHS1–5 genes under different treatments. a Drought stress, the plants were treated with PEG 6000 (20%) and leaf and root samples were collected after 0, 1, and 2 days of treatment. CK-L and CK-R represents control leaf and root samples lacking PEG6000 treatment, respectively. PL and PR represent leaf and root samples treated with PEG6000. 1D and 2D represents treatment times of 1 and 2 days, respectively. b High salt stress, the plants were treated with 200 mM NaCl, and leaf and root samples were collected after 0, 1, and 2 days of treatment. NL and NR represents leaf and root samples treated with NaCl. c 2,4-D treatment; d ABA treatment; e MeJA treatment. Intact seedlings treated with a 10 μM concentration of the given phytohormones sampled at 0, 1, 2, and 5 days for RNA extraction and gene expression analysis
Phytohormone Induction Responsiveness of NtCHS1–5
For phytohormone treatment, 2-week-old seedlings were transplanted to 1/2 strength MS medium with 10 μM 2,4-D, 10 μM MeJ, or 10 μM ABA(Liu and others 2014; Concha and others 2013). The seedlings were collected after 0, 1, 2, and 5 days of treatment. In response to ABA treatment (Fig. 4c), the expression patterns of NtCHS1 and 3 did not change after 2 days of treatment, but expression of both genes decreased sharply by day 5 of treatment. The expression levels of NtCHS2 and NtCHS4 were significantly increased after ABA treatment, peaking at day 2 and then sharply declining by day 5. NtCHS5 expression was decreased in a trend consistent with ABA treatment time. NtCHS4 expression was strongly induced by the 2,4-D treatment (Fig. 4d), and its expression was up-regulated to a peak of more than 250-fold greater than the control at day 2 of treatment. For the MeJA treatment (Fig. 4e), it was notable that the expression level of NtCHS2 and 4 peaked at day 1 of treatment and then sharply decreased with increasing treatment time. NtCHS1 and 3 were down-regulated significantly under MeJA treatment. NtCHS5 was moderately regulated by 2,4-D or MeJA.
Flavonoid Accumulation in Response to Abiotic Stress and Phytohormone Treatment
To measure any changes in flavonoid content and to examine any correlated relationships between flavonoid content and the expression patterns of NtCHS1–5 genes during abiotic stress and phytohormone treatment, flavonoid accumulation was measured using a spectrophotometry method. The total flavonoid content of tobacco leaves did not change at day 1 of drought and high salt stress treatment, but decreased dramatically after 2 days of these treatments (Fig. 5a, b). In the phytohormone treatment experiments, the flavonoid content increased after ABA and MeJA treatment at 1, 2, and 5 days (Fig. 5c, d and e). This was the same trend as the expression levels of NtCHS2 and 4 in tobacco leaves. In the 2,4-D treatment, the flavonoid content increased at the day 1 and day 5 time points, but decreased at the day 2 time point. The result may suggest a moderate increase in the expression of NtCHS4 was induced by 2,4-D treatment and likely caused increased flavonoid accumulation, but the extreme expressed NtCHS4 may suppress accumulation of flavonoid at a certain extent.
Flavonoid accumulation under different treatments. a, b represent the drought and high salt stress experiments. The plants were treated with PEG 6000 (20%) or 200 mM NaCl, and leaves were sampled after 0, 1, and 2 days of treatment. c 2,4-D treatment; d ABA treatment; e MeJA treatment. Whole seedlings were sampled after 0, 1, 2, and 5 days of treatment with a 10 μM concentration of the aforementioned phytohormones
Discussion
In this study, we successfully isolated five CHS genes from Honghuadajinyuan (HD) cultivar of Nicotiana tabacum, named NtCHS1–5 genes. Among them, NtCHS1 and NtCHS2 share high amino acid similarity with previously reported CHS1 (AAK49457) and CHS2 (AHK22740). Considering the possibility of sequence difference between the different tobacco cultivars, the result suggested that NtCHS1 and CHS1 (AAK49457), NtCHS2 and CHS2 (AHK22740) may be the same genes, respectively.
Evidence of gene duplication and subsequent nucleotide substitutions were observed in NtCHS1 and 3 and NtCHS2 and 4. Common tobacco is known to be an allotetraploid formed by the duplication of the hybrid progeny of N. tomentosiformis and N. sylvestris. In duplication events, genes can be duplicated and the newly duplicated genes must be positively selected if they are going to persist over long periods of evolutionary time. NtCHS1 and 3 showed high sequence similarity and expression patterns, although they have distinct promoter sequences and intron sequences. The same situation was observed with NtCHS2 and 4. These results indicated that NtCHS3 and NtCHS4 may be the donor CHS genes of the genome.
Chalcone synthase includes many representative active site amino acids, which play important roles in regulating the catalytic process (Jez and others 2000; Dao and others 2011). In this study, amounts of active site amino acids were detected (Fig. 1), and several residues were believed to be indispensable for the enzyme activity of CHS (Ferrer and others 1999). Moreover, changes in those active site amino acids could cause the function variation of CHS (Suh and others 2000; Sun and others 2015). Residues involved in determination of substrate specificity (T132, S133 and T194) and CoA-binding (P307), in cyclization (M137 and F265), in controlling polyketide size (T197 and S338) and cavity size (P304), were observed changed in NtCHS5. The results indicated the different structure and may be the cause of the diverse function of NtCHS5.
Based on our phylogenetic results, we found that NtCHS1–4 and NtCHS5 were two independent subgroups. NtCHS1–4 shared high sequence homology with the classical CHS genes of eudicots, whereas NtCHS5 shared high homology with some CHS-Like genes (CHSL) of other Solanaceae species. The CHS genes of Solanaceae plants split into two subgroups prior to the divergence of the genera. Similar results were observed in orchid CHSs (Han and others 2006, 2014).
NtCHS1–5 showed different spatial and temporal expression patterns. NtCHS1–4 were widely expressed in pigmented or/and non-pigmented vegetative tissues and floral organs (Fig. 3a, b), suggesting that NtCHS1–4 may be involved both in the biosynthesis of anthocyanins and of other flavonoids. Specifically, the expression of NtCHS5 (Fig. 3a, b) in calyx and corolla indicated that NtCHS5 may be involved in anthocyanin biosynthesis. The expression patterns of NtCHS1 and 3 in seedlings may suggest that their enzymatic product(s) may be involved in more flavonoid pathways besides anthocyanin biosynthesis, for example, flavonol biosynthesis, and so on. The high NtCHS2 and 4 expression levels in stamens and anthers may indicate their potential specific functions in reproduction. Similar results were reported in Phalaenopsis orchid, gerbera (Gerbera hybrida), Freesia hybrid, and tomato (Han and others 2006; Deng and others 2014; Sun and others 2015; Laura and others 2014).
Abiotic stress, including UV radiation, high salt, and drought stress can accelerate the generation and accumulation of ROS in planta (Apel and Hirt 2004; Suzuki and others 2012). Flavonoids serve as antioxidants that function to diminish intracellular generation of ROS and increase ROS scavenging activity (Liu and others 2014; Agati and others 2012). More recently, Chen (2015) revealed that overexpression of EaCHS1 (Eupatorium adenophorum) in tobacco enhances plant tolerance to high salt stress and suggested that the accumulation of flavonoids contributed to plant tolerance to salinity stress by maintaining ROS homeostasis. In this study, the flavonoid content of tobacco leaves was found to have decreased dramatically under high salt stress and drought treatment (Fig. 5a, b), suggesting that flavonoids were employed to scavenge ROS. Meanwhile, the depletion and shortage of flavonoids induced the dramatically increased expression of NtCHS1 and 3 (Fig. 4a, b). The results indicated that NtCHS1 and 3 were induced by both high salt and drought stress, and contribute to the ROS scavenging activity.
Abscisic acid, auxin, salicylic acid, and methyl jasmonate have been reported to influence flavonoid biosynthesis (Tombesi and others 2015; Trivellini and others 2014; Concha and others 2013; Li and others 2014). In this study, the expression of NtCHS2 and 4 was strongly induced by ABA treatment (Fig. 4d) and caused corresponding flavonoid accumulation. ABA has been reported to affect the metabolism of fruit pigments and to regulate the expression of genes in the anthocyanin biosynthesis pathway, including 4CL, CHS, DFR, F3H, among others (Li and others 2015). It is notable that the expression of NtCHS2 and 4 peaked after 1 day of MeJA treatment and then sharply decreased with subsequent treatment time (Fig. 4e). A similar expression pattern of a CHS gene was observed in white spruce and in Fragaria chiloensis fruit (Richard and others 2010; Concha and others 2013). Exogenous MeJA suppressed the expression of NtCHS1 and 3 (Fig. 4e). The opposite expression pattern of NtCHS1,3 and NtCHS2,4 as regulated by the MeJA may highlight the different functions among them. It was notable that after two days of 2,4-D treatment, the accumulation of flavonoid was inhibited (Fig. 5c). Shi and Xie (2011) reported a similar conclusion with 2,4-D treatment strongly inhibiting anthocyanin accumulation. Shortage of flavonoid may induce the strong expression of NtCHS4 (Fig. 4c). The result indicated that NtCHS2 and 4 may be involved in anthocyanin biosynthesis and strongly induced by ABA, MeJA, and 2,4-D.
In conclusion, we identified and explored the functions of five chalcone synthase genes (NtCHS1–5) from Nicotiana tabacum. NtCHS1–5 had different gene structures, different evolution patterns, different expression profiles, and different stress responsiveness. Our results suggest the diverse functional roles of NtCHS1–5 in plant growth and development. This study extends the knowledge about plant CHS genes and will contribute to research about the molecular evolution, expression, and regulation of these genes. More functional analysis will need to be conducted to further characterize the exact functions of the NtCHS1–5 genes of Nicotiana tabacum.
References
Agati G, Azzarello E, Pollastri S, Tattini M (2012) Flavonoids as antioxidants in plants: location and functional significance. Plant Sci 196:67–76. doi:10.1016/j.plantsci.2012.07.014
Ahmad A, Niwa Y, Goto S, Ogawa T, Shimizu M, Suzuki A, Kobayashi K, Kobayashi H (2015) bHLH106 integrates functions of multiple genes through their G-Box to confer salt tolerance on Arabidopsis. PLoS ONE 10(5):e0126872. doi:10.1371/journal.pone.0126872.eCollection
Apel K, Hirt H (2004) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399
Boter M, Ruiz-Rivero O, Abdeen A, Prat S (2004) Conserved MYC transcription factors play a key role in jasmonate signaling both in tomato and Arabidopsis. Genes Dev 18:1577–1591
Budahn H, Barański R, Grzebelus D, Kiełkowska A, Straka P, Metge K, Linke B, Nothnagel T (2014) Mapping genes governing flower architecture and pollen development in a double mutant population of carrot. Front Plant Sci 8(5):504. doi:10.3389/fpls.2014.00504.eCollection2014
Chen S, Liu GS, Wang YY, Sun YH, Chen J (2011) Cloning of a calcium-dependent protein kinase gene NtCDPK12, and its induced expression by high-salt and drought in Nicotiana tabacum. Agric Sci China 10(12):1851–1860
Chen LJ, Guo HM, Lin Y, Cheng HM (2015) Chalcone synthase EaCHS1 from Eupatorium adenophorum functions in salt stress tolerance in tobacco. Plant Cell Rep 34:885–894. doi:10.1007/s00299-015-1751-7
Choi C, Hwang SH, Fang IR, Kwon SI, Park SR, Ahn I, Kim JB, Hwang DJ (2015) Molecular characterization of Oryza sativa WRKY6, which binds to W-box-like element 1 of the Oryza sativa pathogenesis-related (PR) 10a promoter and confers reduced susceptibility to pathogens. New Phytol 208(3):846–859. doi:10.1111/nph.13516
Concha CM, Figueroa NE, Poblete LA, Oñate FA, Schwab W, Figueroa CR (2013) Methyl jasmonate treatment induces changes in fruit ripening by modifying the expression of several ripening genes in Fragaria chiloensis fruit. Plant Physiol Biochem 70:433–444. doi:10.1016/j.plaphy.2013.06.008
Dao TT, Linthorst HJ, Verpoorte R (2011) Chalcone synthase and its functions in plant resistance. Phytochem Rev 10:397–412
Deng X, Bashandy H, Ainasoja M, Kontturi J, Pietiäinen M, Laitinen RA, Albert VA, Valkonen JP, Elomaa P, Teeri TH (2014) Functional diversification of duplicated chalcone synthase genes in anthocyanin biosynthesis of Gerbera hybrida. New Phytol 201(4):1469–1483. doi:10.1111/nph.12610
Ferrer JL, Jez JM, Bowman ME, Dixon RA, Noel JP (1999) Structure of chalcone synthase and the molecular basis of plant polyketide biosynthesis. Nat Struct Biol 6:775–784
Filichkin SA, Leonard JM, Monteros A, Liu PP, Nonogaki H (2004) A novel endo-beta-mannanase gene in tomato LeMAN5 is associated with anther and pollen development. Plant Physiol 134:1080–1087
Flagel LE, Wendel JF (2009) Gene duplication and evolutionary novelty in plants. New Phytol 183:557–564. doi:10.1111/j.1469-8137.2009.02923.x
Fu GM, Wan Y, Zhou JB (2006) Comparison of methods of determining the content of flavonoids in extracts from Evcommia Vlmoides Oliv. Leaves. J Chin Inst Food Sci Technol 6(1):224–229
Fu M, Kang HK, Son SH, Kim SK, Nam KH (2014) A subset of arabidopsis RAV transcription factors modulates drought and salt stress responses independent of ABA. Plant Cell Physiol 55(11):1892–1904. doi:10.1093/pcp/pcu118
Han YY, Ming F, Wang W, Wang JW, Ye MM, Shen DL (2006) Molecular evolution and functional specialization of chalcone synthase superfamily from Phalaenopsis orchid. Genetica 128:429–438
Han Y, Zhao W, Wang Z, Zhu J, Liu Q (2014) Molecular evolution and sequence divergence of plant chalcone synthase and chalcone synthase-like genes. Genetica 142(3):215–225. doi:10.1007/s10709-014-9768-3
Harashima S, Takano H, Ono K, Takio S (2004) Chalcone synthase-like gene in the liverwort, Marchantia paleacea var. diptera. Plant Cell Rep 23:167–173
Hwang SH, Hwang DJ (2010) Isolation and characterization of the rice NPR1 promoter. Plant Biotechnol Rep 4:29–35
Jez JM, Austin MB, Ferrer J, Bowman ME, Schröder J, Noel JP (2000) Structural control of polyketide formation in plant-specific polyketide synthases. Chem Biol 7(12):919–930
Kaplan-Levy RN, Brewer PB, Quon T, Smyth DR (2012) The trihelix family of transcription factors—light, stress and development. Trends Plant Sci 17(3):163–171. doi:10.1016/j.tplants.2011.12.002
Koes RE, Spelt CE, van den Elzen PJ, Mol JN (1989) Cloning and molecular characterization of the chalcone synthase multigene family of Petunia hybrida. Gene 81:245–257
Krause A, Lan VT, Broughton WJ (1997) Induction of chalcone synthase expression by rhizobia and nod factors in root hairs and roots. Mol Plant Microbe Interact 10(3):388–393
Li T, Jia KP, Lian HL, Yang X, Li L, Yang HQ (2014) Jasmonic acid enhancement of anthocyanin accumulation is dependent on phytochrome A signaling pathway under far-red light in Arabidopsis. Biochem Biophys Res Commun 454(1):78–83. doi:10.1016/j.bbrc.2014.10.059
Li D, Li L, Luo Z, Mou W, Mao L, Ying T (2015) Comparative transcriptome analysis reveals the influence of abscisic acid on the metabolism of pigments, ascorbic acid and folic acid during strawberry fruit ripening. PLoS ONE 10(6):e0130037. doi:10.1371/journal.pone.0130037.eCollection
Liu Z, Shi MZ, Xie DY (2014) Regulation of anthocyanin biosynthesis in Arabidopsis thaliana red pap1-D cells metabolically programmed by auxins. Planta 239:765–781. doi:10.1007/s00425-013-2011-0
Luo XY, Fang HB, Huang RY, Zhao DF, Zhu T, Guo YY, Yi-ping Zeng, Liu RY, Zhu D (2015) Research on the spectrophotometry for determination of flavonoids in waste residue of Stevia rebaudiana. Guang Pu Xue Yu Guang Pu Fen Xi 35(10):2820–2824
Ma Z, Hu X, Cai W, Huang W, Zhou X, Luo Q, Yang H, Wang J, Huang J (2014) Arabidopsis miR171-targeted scarecrow-like proteins bind to GT cis-elements and mediate gibberellin-regulated chlorophyll biosynthesis under light conditions. PLoS Genet 10(8):e1004519. doi:10.1371/journal.pgen.1004519.eCollection
Macknight RC, Reynolds PH, Farnden KJ (1995) Analysis of the lupin Nodulin-45 promoter: conserved regulatory sequences are important for promoter activity. Plant Mol Biol 27(3):457–466
Nagy NE, Fossdal CG, Krokene P, Krekling T, Lönneborg A, Solheim H (2004) Induced responses to pathogen infection in Norway spruce phloem: changes in polyphenolic parenchyma cells, chalcone synthase transcript levels and peroxidase activity. Tree Physiol 24(5):505–515
Napoli CA, Fahy D, Wang HY, Taylor LP (1999) White anther: a petunia mutant that abolishes pollen flavonol accumulation, induces male sterility, and is complemented by a chalcone synthase transgene. Plant Physiol 120(2):615–622
Niggeweg R, Thurow C, Kegler C, Gatz C (2000) Tobacco transcription factor TGA2.2 is the main component of as-1-binding factor ASF-1 and is involved in salicylic acid- and auxin-inducible expression of as-1-containing target promoters. J Biol Chem 275(26):19897–19905
Park HC, Kim ML, Kang YH et al (2004) Pathogen- and NaCl-induced expression of the SCaM-4 promoter is mediated in part by a GT-1 box that interacts with a GT-1-like transcription factor. Plant Physiol 135:2150–2161
Reimold U, Kröger M, Kreuzaler F, Hahlbrock K (1983) Coding and 3′ non-coding nucleotide sequence of chalcone synthase messenger RNA and assignment of amino acid sequence of the enzyme. EMBO J 2(10):1801–1805
Richard S, Lapointe G, Rutledge RG, Séguin A (2000) Induction of chalcone synthase expression in white spruce by wounding and jasmonate. Plant Cell Physiol 41(8):982–987
Schenk PM, Kazan K, Wilson I et al (2000) Coordinated plant defense responses in Arabidopsis revealed by microarray analysis. Proc Natl Acad Sci USA 97:11655–11660
Shi MZ, Xie DY (2011) Engineering of red cells of Arabidopsis thaliana and comparative genome-wide gene expression analysis of red cells versus wild-type cells. Planta 233(4):787–805. doi:10.1007/s00425-010-1335-2
Suh DY, Kagami J, Fukuma K, Sankawa U (2000) Evidence for catalytic cysteine–histidine dyad in chalcone synthase. Biochem Biophys Res Commun 275(3):725–730
Sun LL, Li Y, Li SS, Wu XJ, Hu BZ, Chang Y (2014) Identification and characterisation of DfCHS, a chalcone synthase gene regulated by temperature and ultraviolet in Dryopteris fragrans. Cell Mol Biol (Noisy-le-grand) 60(6):1–7
Sun W, Meng X, Liang L, Jiang W, Huang Y, He J, Hu H, Almqvist J, Gao X, Wang L (2015) Molecular and biochemical analysis of chalcone synthase from Freesia hybrid in flavonoid biosynthetic pathway. PLoS ONE 10(3):e0119054. doi:10.1371/journal.pone.0119054.eCollection
Suzuki N, Koussevitzky S, Mittler R, Miller G (2012) ROS and redox signalling in the response of plants to abiotic stress. Plant Cell Environ 35:259–270. doi:10.1111/j.1365-3040.2011.02336.x
Tombesi S, Nardini A, Frioni T, Soccolini M, Zadra C, Farinelli D, Poni S, Palliotti A (2015) Stomatal closure is induced by hydraulic signals and maintained by ABA in drought-stressed grapevine. Sci Rep 5:12449. doi:10.1038/srep12449
Trivellini A, Gordillo B, Rodríguez-Pulido FJ, Borghesi E, Ferrante A, Vernieri P, Quijada-Morín N, González-Miret ML, Heredia FJ (2014) Effect of salt stress in the regulation of anthocyanins and color of hibiscus flowers by digital image analysis. J Agric Food Chem 62(29):6966–6974. doi:10.1021/jf502444u
Vijayan J, Devanna BN, Singh NK, Sharma TR (2015) Cloning and functional validation of early inducible Magnaporthe oryzae responsive CYP76M7 promoter from rice. Front Plant Sci 6:371. doi:10.3389/fpls.2015.00371.eCollection2015
Zabala G, Zou J, Tuteja J, Gonzalez DO, Clough SJ, Vodkin LO (2006) Transcriptome changes in the phenylpropanoid pathway of Glycine max in response to Pseudomonas syringae infection. BMC Plant Biol 3:6–26
Zhou B, Wang Y, Zhan Y, Li Y, Kawabata S (2013) Chalcone synthase family genes have redundant roles in anthocyanin biosynthesis and in response to blue/UV-A light in turnip (Brassica rapa; Brassicaceae). Am J Bot 100(12):2458–2467. doi:10.3732/ajb.1300305
Acknowledgements
This work was supported by The Agricultural Science and Technology Innovation Program (ASTIP-TRIC01).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, S., Pan, X., Li, Y. et al. Identification and Characterization of Chalcone Synthase Gene Family Members in Nicotiana tabacum . J Plant Growth Regul 36, 374–384 (2017). https://doi.org/10.1007/s00344-016-9646-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00344-016-9646-6