Abstract
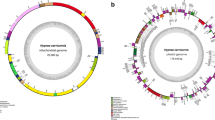
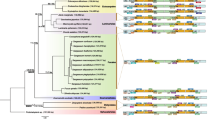
Pyropia species grow in the intertidal zone and are cold-water adapted. To date, most of the information about the whole plastid and mitochondrial genomes (ptDNA and mtDNA) of this genus is limited to Northern Hemisphere species. Here, we report the sequencing of the ptDNA and mtDNA of the Antarctic red alga Pyropia endiviifolia using the Illumina platform. The plastid genome (195 784 bp, 33.28% GC content) contains 210 protein-coding genes, 37 tRNA genes and 6 rRNA genes. The mitochondrial genome (34 603 bp, 30.5% GC content) contains 26 protein-coding genes, 25 tRNA genes and 2 rRNA genes. Our results suggest that the organellar genomes of Py. endiviifolia have a compact organization. Although the collinearity of these genomes is conserved compared with other Pyropia species, the genome sizes show significant differences, mainly because of the different copy numbers of rDNA operons in the ptDNA and group II introns in the mtDNA. The other Pyropia species have 2–3 distinct intronic ORFs in their cox 1 genes, but Py. endiviifolia has no introns in its cox 1 gene. This has led to a smaller mtDNA than in other Pyropia species. The phylogenetic relationships within Pyropia were examined using concatenated gene sets from most of the available organellar genomes with both the maximum likelihood and Bayesian methods. The analysis revealed a sister taxa affiliation between the Antarctic species Py. endiviifolia and the North American species Py. kanakaensis.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Abascal F, Zardoya R, Posada D. 2005. ProtTest: selection of best-fit models of protein evolution. Bioinformatics, 21 (9): 2 104–2 105.
Brodie J A, Irvine L M. 2003. Seaweeds of the British Isles. Volume 1 Rhodophyta. Part 3B Bangiophycidae. Natural History Museum, London.
Buschiazzo E, Ritland C, Bohlmann J, Ritland K. 2012. Slow but not low: genomic comparisons reveal slower evolutionary rate and higher dN/dS in conifers compared to angiosperms. BMC Evolutionary Biology, 12: 8.
Capella-Gutiérrez S, Silla-Martínez J M, Gabaldón T. 2009. trimAl: a tool for automated alignment trimming in largescale phylogenetic analyses. Bioinformatics, 25 (15): 1 972–1 973.
Chamberlain Y M. 1963. The identity of Monostroma endiviifolium A. and E.S. Gepp. Nova Hedwigia, 5: 151–155.
Darling A C E, Mau B, Blattner F R, Perna N T. 2004. Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome R esearch, 14 (7): 1 394–1 403.
Douglas S E. 1998. Plastid evolution: origins, diversity, trends. Current O pinion in G enetics & Development, 8 (6): 655–661.
Dutcher J A, Kapraun D F. 1994. Random amplified polymorphic DNA (RAPD) identification of genetic variation in three species of Porphyra (Bangiales, Rhodophyta). Journal of Applied Phycology, 6 (3): 267–273.
Gray M W, Burger G, Lang B F. 2001. The origin and early evolution of mitochondria. Genome Biology, 2 (6): reviews1018.1.
Guiry M D, Guiry G M. 2017. AlgaeBase. World-Wide Electronic Publication, National University of Ireland, Galway, https://doi.org/www.marinespecies.org/aphia.php?p=sourcedetails&id=37.
Hagopian J C, Reis M, Kitajima J P, Bhattacharya D, De Oliveira M C. 2004. Comparative analysis of the complete plastid genome sequence of the red alga Gracilaria tenuistipitata var. liui provides insights into the evolution of rhodoplasts and their relationship to other plastids. Journal of Molecular Evolution, 59 (4): 464–477.
Henry R J. 2005. Plant Diversity and Evolution: Genotypic and Phenotypic Variation in Higher Plants. CABI Publishing, Wallingford, Oxfordshire, UK.
Hernandez D, François P, Farinelli L, Østerås M, Schrenzel J. 2008. De novo bacterial genome sequencing: millions of very short reads assembled on a desktop computer. Genome R esearch, 18 (5): 802–809.
Huelsenbeck J P, Ronquist F. 2001. MRBAYES: bayesian inference of phylogenetic trees. Bioinformatics, 17 (8): 754–755.
Janouškovec J, Liu S L, Martone P T, Carré W, Leblanc C, Collén J, Keeling P J. 2013. Evolution of red algal plastid genomes: ancient architectures, introns, horizontal gene transfer, and taxonomic utility of plastid markers. PLoS One, 8 (3): e59001.
Kamikawa R, Masuda I, Demura M, Oyama K, Yoshimatsu S, Kawachi M, Sako Y. 2009. Mitochondrial group II introns in the raphidophycean flagellate Chattonella spp. suggest a diatom-to-Chattonella lateral group II intron transfer. Protist, 160 (3): 364–375.
Katoh K, Kuma K I, Toh H, Miyata T. 2005. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic A cids R esearch, 33 (2): 511–518.
Lee J, Cho C H, Park S I, Cho J W, Song H S, West J A, Bhattacharya D, Yoon H S. 2016. Parallel evolution of highly conserved plastid genome architecture in red seaweeds and seed plants. BMC Biology, 14: 75.
Lohse M, Drechsel O, Bock R. 2007. OrganellarGenomeDRAW (OGDRAW): a tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Current G enetics, 52 (5–6): 267–274.
Mayor C, Brudno M, Schwartz J R, Poliakov A, Rubin E M, Frazer K A, Pachter L S, Dubchak I. 2000. VISTA: visualizing global DNA sequence alignments of arbitrary length. Bioinformatics, 16 (11): 1 046–1 047.
Michel F, Jacquier A, Dujon B, 1982. Comparison of fungal mitochondrial introns reveals extensive homologies in RNA secondary structure. Biochimie, 64 (10): 867–881.
Mumford Jr T F, Miura A. 1988. Porphyra as food: cultivation and economics. In: Lembi C A, Waaland J R eds. Algae and Human Affairs. Cambridge University Press, Cambridge.
Niwa K, Kikuchi N, Iwabuchi M, Aruga Y. 2004. Morphological and AFLP variation of Porphyra yezoensis Ueda form, narawaensis Miura (Bangiales, Rhodophyta). Phycological Research, 52 (2): 180–190.
Ogihara Y, Yamazaki Y, Murai K, Kanno A, Terachi T, Shiina T, Miyashita N, Nasuda S, Nakamura C, Mori N, Takumi S, Murata N, Futo S, Tsunewaki K. 2005. Structural dynamics of cereal mitochondrial genomes as revealed by complete nucleotide sequencing of the wheat mitochondrial genome. Nucleic A cids Research, 33 (19): 6 235–6 250.
Patel R K, Jain M. 2012. NGS QC Toolkit: a toolkit for quality control of next generation sequencing data. PLoS One, 7 (2): e30619.
Porebski S, Bailey L G, Baum B R. 1997. Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant M olecular Biology Reporter, 15 (1): 8–15.
Reyes-Prieto A, Weber A P, Bhattacharya D. 2007. The origin and establishment of the plastid in algae and plants. Annu al Review Genetics, 41: 147–168.
Rodríguez-Ezpeleta N, Brinkmann H, Burey S C, Roure B, Burger G, Löffelhardt W, Bohnert H J, Philippe H, Lang B F. 2005. Monophyly of primary photosynthetic eukaryotes: green plants, red algae, and glaucophytes. Current Biology, 15 (4): 1 325–1 330.
Ruby J G, Bellare P, DeRisi J L. 2013. PRICE: software for the targeted assembly of components of (Meta) genomic sequence data. G3: Genes, Genomes, Genetics, 3 (5): 865–880.
Stamatakis A. 2014. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics, 30 (9): 1 312–1 313.
Sutherland J E, Lindstrom S C, Nelson W A, Brodie J, Lynch M D J, Hwang M S, Choi H G, Miyata M, Kikuchi N, Oliveira M C, Farr T, Neefus C, Mols-Mortensen A, Milstein D, Müller K M. 2011. A new look at an ancient order: generic revision of the Bangiales (Rhodophyta). Journal of Phycology, 47 (5): 1 131–1 151.
Taanman J W. 1999. The mitochondrial genome: structure, transcription, translation and replication. Biochimica et Biophysica Acta (BBA)-Bioenergetics, 1410 (2): 103–123.
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. 2013. MEGA6: molecular evolutionary genetics analysis version 6.0. Molecular Biology and E volution, 30 (12): 2 725–2 729.
Verbruggen H, Maggs C A, Saunders G W, Le Gall L, Yoon H S, De Clerck O. 2010. Data mining approach identifies research priorities and data requirements for resolving the red algal tree of life. BMC Evolutionary Biology, 10: 16.
Wang L, Mao Y X, Kong F N, Li G Y, Ma F, Zhang B L, Sun P P, Bi G Q, Zhang F F, Xue H F, Cao M. 2013. Complete sequence and analysis of plastid genomes of two economically important red algae: Pyropia haitanensis and Pyropia yezoensis. PLoS One, 8 (5): e65902.
Wiencke C, Clayton M N. 1998. The life history of Porphyra endiviifolium from the South Shetland Islands, Antarctica. Polar Biology, 19 (4): 257–263.
Xie C T, Chen C S, Xu Y, Ji D H. 2010. Construction of a genetic linkage map for Porphyra h aitanensis (Bangiales, Rhodophyta) based on sequence-related amplified polymorphism and simple sequence repeat markers. Journal of Phycology, 46 (4): 780–787.
Yang E C, Kim K M, Kim S Y, Lee J, Boo G H, Lee J H, Nelson W A, Yi G M, Schmidt W E, Fredericq S, Boo S M, Bhattacharya D, Yoon H S. 2015. Highly conserved mitochondrial genomes among multicellular red algae of the Florideophyceae. Genome Biology and E volution, 7 (8): 2 394–2 406.
Yoon H S, Müller K M, Sheath R G, Ott F D, Bhattacharya D. 2006. Defining the major lineages of red algae (Rhodophyta). Journal of Phycology, 42 (2): 482–492.
Zhao L, Li X, Zhang N, Zhang S D, Yi T S, Ma H, Guo Z H, Li D Z. 2016. Phylogenomic analyses of large-scale nuclear genes provide new insights into the evolutionary relationships within the rosids. Molecular Phylogenetics and Evolution, 105: 166–176.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supported by the National Natural Science Foundation of China (No. 31372517), the Scientific and Technological Innovation Project Financially Supported by Qingdao National Laboratory for Marine Science and Technology (No. 2015ASKJ02), and the National Infrastructure of Fishery Germplasm Resources (No. 2016DKA30470)
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Xu, K., Tang, X., Bi, G. et al. The first complete organellar genomes of an Antarctic red alga, Pyropia endiviifolia: insights into its genome architecture and phylogenetic position within genus Pyropia (Bangiales, Rhodophyta). J. Ocean. Limnol. 36, 1315–1328 (2018). https://doi.org/10.1007/s00343-018-7088-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00343-018-7088-7