Abstract
The C57BL/6 mouse, the most frequently utilized animal model in biomedical research, is in use as several substrains, all of which differ by a small array of genomic differences. Two of these substrains, C57BL/6J (B6J) and C57BL/6N (B6N), are commonly used but it is unclear how phenotypically similar or different they are. Here, we tested whether adolescent B6N mice have a bone phenotype and respond to the loss of weightbearing differently than B6J. At 9 weeks of age, normally ambulating B6N had lower trabecular bone volume fraction but greater bone formation rates and osteoblast surfaces than corresponding B6J. At 11 weeks of age, differences in trabecular indices persisted between the substrains but differences in cellular activity had ceased. Cortical bone indices were largely similar between the two substrains. Hindlimb unloading (HLU) induced similar degeneration of trabecular architecture and cellular activity in both substrains when comparing 11-week-old HLU mice to 11-week-old controls. However, unloaded B6N mice had smaller cortices than B6J. When comparing HLU to 9 weeks baseline control mice, deterioration in trabecular separation, osteoblast indices, and endocortical variables was significantly greater in B6N than B6J. These data indicate specific developmental differences in bone formation and morphology between B6N and B6J mice, giving rise to a differential response to mechanical unloading that may be modulated, in part, by the genes Herc2, Myo18b, and Acan. Our results emphasize that these substrains cannot be used interchangeably at least for investigations in which the phenotypic makeup and its response to extraneous stimuli are of interest.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genetic architecture of an individual regulates not only normal physiological development but also plays a role in the predisposition to diseases and their pathogenesis. For polygenic traits, many genes with predominantly small effect sizes influence the phenotype, but cumulatively, these genes can produce large inter-individual phenotypic variation (Plomin et al. 2009). For instance, genetic makeup accounts for up to 80% of the differences in body height (Visscher et al. 2008), a level similar to the genetic contribution to the variability in bone mineral density (Eisman 1999). Analogous to development, genetic predisposition or mediation has been implicated in many diseases including cancer, migraine, non-alcoholic fatty liver disease, and osteoporosis (Eisman 1999; Haan et al. 1997; Hansen and Cavenee 1987; Romeo et al. 2008). Linkage analysis and genome wide association studies (GWAS), correlating genomic architecture with phenotypic variation, have allowed the identification of genetic polymorphisms that either promote susceptibility or resistance to disorders (Frazer et al. 2009; Korte and Farlow 2013). For instance, specific mutations in BRCA1 and BRCA2 introduce a heightened risk of breast cancer (Mavaddat et al. 2010) while certain mutations in PTPN22 lower the risk of acquiring Crohn’s disease (Bayat et al. 2004; Frazer et al. 2009; Korte and Farlow 2013; Vang et al. 2005).
Discovery of putative genes often relies on associations and inherently, gene function has to be confirmed directly, such as through transgenic mouse models (van Amerongen and Berns 2006). In knockout (KO) mice, however, the genetic background from which the gene is deleted can modify the phenotypic outcome (Lloyd et al. 2015). For instance, IL10 null mice bred on a C3H/HeJBir (C3H) background were highly susceptible to colitis whereas deletion of IL10 from a C57B6/J (B6J) background produced a mouse largely impervious to colitis pathogenesis (Beckwith et al. 2005; Mähler and Leiter 2002). This ostensible dichotomy should not be entirely surprising when considering the large genetic diversity between the C3H and B6 strains (Wilson et al. 2001), including more than 350 coding site single nucleotide polymorphisms that can alter the amino acid sequence of the resultant protein (non-synonymous SNPs). This large genetic variability between these two inbred strains is also reflected at the tissue level. For bone, C3H mice have a 50% greater femoral BMD and a greatly reduced susceptibility to disuse compared to B6J mice (Beamer et al. 1996; Rosen et al. 2009).
Unlike the extensive genomic differences between inbred mouse strains, much less is known about phenotypic differences between inbred substrains of mice that comprise only subtle genetic variability. Knockout mice have predominantly been bred on the C57B6/J (B6J) substrain but the C57B6/N (B6N) substrain became popular with the implementation of the Knockout Mouse Project (KOMP) (Austin et al. 2004; Simon et al. 2013). B6N and B6J mice only differ by 34 single nucleotide polymorphisms (SNP) and two insertion–deletion (INDELS) in the coding regions of their genome, in addition to 15 structural variations (SV) within chromosomal regions. B6N mice have reduced vision, lower systolic pressure, and greater fat mass compared to B6J substrain (Simon et al. 2013). Not only do these substrains exhibit basal differences in certain physiological indices but they can also respond differently to a given stimulus. In a model of diet-induced obesity, B6J mice experienced greater weight gain than B6N accompanied by greater serum glucose and serum leptin levels (Nicholson et al. 2010). These differences were linked to a spontaneous loss-of-function mutation in the nicotinamide nucleotide transhydrogenase (Nnt) gene in the B6J mouse. In spite of clear evidence for dissimilarity between the B6J and B6N substrains, no bone phenotype or differential skeletal response was previously found in young adult 14-week-old B6N mice (Simon et al. 2013).
Phenotypic changes resulting from gene deletion can exhibit age dependency. For instance, lymphoid enhancement factor 1 (Lef1) null mice have low bone mass at 13 weeks of age but not at 34 weeks (Noh et al. 2009). Similarly, bone’s response to catabolic signals such as unloading can depend on age (Li et al. 2016). In an effort to test for potential skeletal differences between B6N and B6J, here, we targeted an age at which mice have reached sexual maturity but are still growing significantly, and exhibit a high susceptibility to disuse (Li et al. 2016). Specifically, we tested whether (1) the B6N and B6J substrains exhibit distinct trabecular and cortical bone morphology and cellular indices at 9 and 11 weeks of age and (2) whether they respond differently to 2 weeks of disuse (hindlimb unloading).
Methods
Experimental design
All procedures were reviewed and approved by Stony Brook University’s Institutional Animal Care and Use Committee (IACUC). Female mice of the C57BL/6J (Stock No. 000664) and C57BL/6N (Stock No. 005304) substrains were received from The Jackson Laboratory (Bar Harbor, ME). Mice were either euthanized at 9 weeks of age (baseline control, 9wk, n = 10), were ambulatory between 9 and 11 weeks and euthanized at 11 weeks (ambulatory age-matched control, 11wk, n = 12), or exposed to hindlimb unloading by tail suspension/traction (Globus and Morey-Holton 2016) between 9 and 11 weeks and euthanized at 11 weeks (hindlimb unloaded, HLU, n = 12). The experiments were performed on adolescent mice as at this age, overall bone mass of the C57BL/6J appendicular skeleton has not plateaued yet but the age-related decline of trabecular quantity and architecture has commenced (Glatt et al. 2007).
All mice were housed individually, had access to rodent chow and water ad libitum, and were maintained on a 12 h light/dark cycle. Body mass was recorded at the beginning of the experiment and monitored daily (HLU mice) or twice a week (ambulatory controls). Mice were injected intraperitoneally (i. p) with 10 mg/kg calcein (Sigma Aldrich) 2d and 9d prior to euthanasia for in vivo fluorescent labeling. After euthanasia, femora were harvested, fixed in 10% neutral buffered formalin for 36–48 h, and stored in 70% ethanol for further processing. Left femurs were used for μCT scanning and histology while right femurs were processed for histomorphometry.
Microcomputed tomography
Trabecular and cortical bone quantity and architecture of the distal femoral metaphysis was evaluated by high-resolution (10 μm) μCT scanning (μCT 40, Scanco Medical, SUI). Scans were performed at energy levels of 55 kV and 145 μA, with a 300 ms integration time and 1000 projections. A region beginning 650 μm proximal to the epiphyseal growth plate and extending 1500 μm towards the diaphysis was chosen for the analysis of the distal femur.
Trabecular and cortical bone were analyzed after separating trabecular struts from the cortical compartment using a semi-automated algorithm (Lublinsky et al. 2007). A thresholding procedure that was identical for both substrains of mice reconstructed trabecular and cortical morphology (Judex et al. 2004b). Trabecular indices including tissue volume (TV), bone volume (BV), bone volume fraction (BV/TV), trabecular thickness (Tb.th), trabecular number (Tb.N), trabecular separation (Tb.Sp), connectivity density (Conn.D), and bone surface to bone volume (BS/BV) were computed. Cortical indices included cortical area (Ct.Ar), cortical thickness (Ct.Th), total area (Tt.Ar), and marrow area (Ma.Ar).
Histology
After µCT scanning, left femurs were decalcified in 14% EDTA (ethylene diamine tetra acetic acid). Decalcified bones were embedded in paraffin, and two 5 µm sections per bone were stained with either toluidine blue to measure osteoblast surface per bone surface (Ob.S/BS, %) and osteoblast number per bone surface (Ob.N/BS), or with tartrate resistant acid phosphatase (TRAP) to quantify osteoclast surface to bone surface (Oc.S/BS, %). For cortical bone, these indices were calculated with either endocortical perimeter (Ec.Pm) or periosteal perimeter (Pe.Pm) as referent; Ob.S/Ec.Pm, Ob.N/Ec.Pm and Oc.S/ Ec.Pm at the endocortical surface and Ob.S/Pe.Pm, Ob.N/ Pe.Pm, Oc. S/ Pe.Pm at the periosteal surface. All analysis was performed in Osteomeasure, (Osteometrics, Inc., Decatur, GA, USA).
Histomorphometry
Undecalcified right femora underwent sequential dehydration in increasing concentrations of ethanol, cleared in petroleum ether, and infiltrated with and embedded in methyl methacrylate, n-butyl phthalate and benzoyl peroxide (Squire et al. 2008). Three 8 µm frontal longitudinal sections per femur were cut using a microtome (Leica Biosystems Inc., IL, USA) and analyzed for dynamic indices of bone formation using Osteomeasure (Green et al. 2012; Li et al. 2016). Mineralizing surface (MS/BS), mineral apposition rate (MAR), and bone formation rate (BFR/BS) were quantified for trabecular bone in the distal femur. One femur of the 11wk group (B6J) could not be processed due to technical issues and was excluded from the histomorphometric analysis. No double labels were detected in one 11wk sample of both B6N and B6J substrains as well as in one B6J HLU sample. MAR and BFR/BS were assumed to be zero in these samples whereas MS/BS, derived from single labels, was included in the quantification. No histomorphometric analysis was performed on cortical bone because of technical issues.
Allelic differences in coding genomic regions
SNPs in the coding regions of the B6N and B6J genome were identified using the Mouse Phenome Database (MPD, The Jackson Laboratory). The SNP variation query function within MPD determined non-synonymous SNPs (SNPs that alter the amino acid sequence of the encoded protein) between B6N and B6J mice using Sanger SNP data (Alfredson et al. 1996; Grubb et al. 2014). SNPs within the same gene were consolidated. The list of genes harboring SNPs between B6N and B6J were compared to the genetic targets identified by our previous genetic linkage study in which we identified bone quantitative trait loci (QTLs) in a F2 mouse population both at baseline (4 mo old) and after 3 weeks of unloading (Judex et al. 2013; Sankaran et al. 2015). To this end, a pool of genes within the unloading QTL intervals that modulated trabecular BV/TV, Tb.N, Tb.Th, Conn.D, and BS/BV was generated (Judex et al. 2013). Olfactory receptor genes were removed and SNPs within the same gene were combined. The genetic targets within different bone indices were merged to generate a distinct set of unloading QTL genes which was then compared to the list of SNP differences between B6N and B6J.
Statistics
Data were reported as mean ± SD. Differences in body mass between the two substrains at 9 weeks and at 11 weeks were determined with two-tailed unpaired t-tests. Comparisons of initial and final body mass within each substrain were performed via two-tailed paired t-tests. Differences between 9wk, 11wk, and HLU mice within each substrain were detected with ANOVA followed by a Tukey post-hoc test. Between-strain differences (B6N vs. B6J) were tested with two-tailed unpaired t-tests at 9 and 11 weeks. Differences in the magnitude of bone’s response to hindlimb unloading between the two substrains were quantified using two-way ANOVA. Body mass at 9 weeks was used as a covariate in all µCT comparisons between B6N and B6J mice. Statistical tests were performed in SPSS for Windows. Significance was set at α < 5%.
Results
Body mass
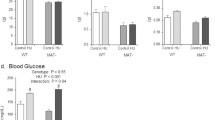
At 9 weeks of age and for both substrains, body mass of 9wk and HLU groups were not different from each other. However, the body mass of 11wk B6N mice was 7.5% lower compared to the corresponding group on the B6J substrain (p < 0.05) (Fig. 1). Despite lower starting body mass, 11wk B6N mice gained about 13% in body mass between 9 and 11 weeks, while 11wk B6J mice gained only about 5% in body mass during the same period (all p < 0.05) and consequently, the differences between these groups at 9 weeks did not persist at 11 weeks. Between 9 and 11 weeks of age, HLU B6J mice lost about 6% in body mass (p < 0.05) while body mass of HLU B6N mice was not different between these two time points (Fig. 1 ). There were no differences in body mass for HLU mice between substrains at 11 weeks. There were no statistically significant differences in body mass between groups within a substrain.
a Body mass of 9wk, 11wk and HLU mice at 9 weeks of age. b Relative mean change in body mass (and standard error of the difference) over the experimental period in control and HLU of B6N and B6J mice. Relative mean differences (and standard error of the difference) in (c) BV/TV, Tb.N, and Tb.Th and (d) Ct.Ar, Ct.Th, Ma.Ar, and Tt.Ar between 9wk B6N and B6J mice (n = 10 each). Microcomputed tomography images of the (e) trabecular and (f) cortical distal femoral metaphysis of 9wk, B6N and B6J mice. *p < 0.05 for difference between B6N 9wk vs. B6J 9wk; **p < 0.05 for difference in body mass between B6N and B6J
B6N versus B6J at 9 and 11 weeks (no HLU)
Trabecular bone
At 9 weeks of age, trabecular bone quantity and architectural indices were consistently lower in B6N than in B6J mice. 9wk B6N mice had lower BV/TV (−21%), Tb.Th (−6%), and BS/BV (−8%) when compared to 9wk B6J (all p < 0.05) (Fig. 1; Table 1 ). However, bone formation rates and osteoblastic surface were greater in 9wk B6N than B6J. B6N mice had greater MS/BS (63%), BFR/BS (46%), Ob.S/BS (35%), and Ob.N/BS (23%) (Table 3 ) when compared to their B6J counterparts (all p < 0.05). At 11 weeks, differences in trabecular microarchitectural indices persisted. 11wk B6N mice had lower BV/TV (−29%), Tb.N (−10%), Tb.Th (−7%). Tb.Sp (11%), and BS/BV (9%) were greater in B6N than B6J (all p < 0.05) (Table 1 ). No differences were observed in bone formation rates or osteoblastic surface between 11wk B6N and B6J mice. Osteoclastic activity was similar between B6N and B6J mice at both 9 and 11 weeks. When body mass was not used as covariate (µCT), the differences between B6N and B6J mice persisted at both time points.
Developmental growth between 9 and 11 weeks was compared between the two substrains by comparing trabecular indices between 11wk mice and 9wk mice. Trabecular microarchitectural indices were lower in 11wk mice when compared to 9wk mice in both substrains. When compared to 9wk mice, 11wk B6N mice had lower BV/TV (−26%), Tb.N (−11%), Tb.Sp (13%), and Conn.D (−49%) (all p < 0.05). Similarly, in the B6J substrain, 11wk B6J mice had lower BV/TV (−19%), Tb.N (−7%), Tb.Sp (8%) and Conn.D (−31%) when compared to 9wk B6J mice (all p < 0.05). Bone formation rates and osteoblastic activity were either lower or were not different in 11wk B6N mice when compared to corresponding 9wk B6N mice. 11wk B6N mice had 29% lower MS/BS and 18% lower Ob.S/BS when compared to 9wk B6N mice (all p < 0.05). Other indices of bone formation and osteoblastic activity were similar between 11wk and 9wk B6N mice (Fig. 1; Tables 1, 3 ). In B6J mice, bone formation and osteoblastic surface were greater at 11 weeks than at 9 weeks with 92% greater BFR/BS, and 23% greater Ob.S/BS (all p < 0.05). Osteoclastic activity was not different between 11wk and 9wk mice in both substrains (Table 3 ). In B6N mice, trabecular growth had plateaued or slowed at 11 weeks compared to 9 weeks (MS/BS: −29%, BFR/BS: nsd, Ob.S/BS: −18%). In contrast, trabecular growth indices in B6J had accelerated at 11 weeks when compared to mice at 9 weeks (MS/BS, nsd; BFR/BS, 92%; Ob.S/Bs, 23%) (gene*age, p < 0.05).
Cortical bone
Cortical geometry was similar between B6N and B6J mice both at 9 weeks and 11 weeks of age independent of whether body mass was used as covariate (Table 2 ). Osteoblastic activity and osteoblast number were also similar between B6N and B6J mice on both the periosteal and endocortical surface at both time points, with the exception that at 11 weeks, B6N mice had 29% lower Ob.N/Ec.Pm than B6J mice (p < 0.05). Osteoclast activity was not different between B6N and B6J both on the endocortical and periosteal surface at both time points (Table 4 ).
Unlike the trabecular compartment in which bone diminished between 9 and 11 weeks, cortical indices were greater at 11 weeks than at 9-week mice in both substrains. Compared to 9wk mice, in the B6N substrain, 11wk mice had 15 and 17% greater Ct.Ar and Ct.Th, while 11wk mice in the B6J substrain had 13 and 17% greater Ct.Ar and Ct.Th (all p < 0.05). Ma.Ar, Tt.Ar and osteoblast number and surface on both the endocortical and periosteal surface were not different between 11wk and 9wk mice in both substrains (Tables 2, 4 ). Cortical growth between 9 and 11 weeks followed similar patterns and were not different between the two substrains (Tables 2, 4 ).
Hindlimb unloading
Trabecular bone
The effects of hindlimb unloading were quantified by comparing HLU mice to both 11wk and 9wk normal-control mice. In both substrains, HLU mice had a severely depleted trabecular compartment when compared to either age-matched 11wk controls or 9wk baseline controls. The compromised trabecular structure in unloaded mice was reflected by architectural degeneration, suppressed bone formation, and lower osteoblastic surface.
In the B6N substrain and compared to 11wk controls, HLU mice had lower BV/TV (−52%), Tb.N (−11%), Tb.Th (−27%), Tb.Sp (12%), BS/BV (40%), and Conn.D (−74%) (all p < 0.05). Further, HLU mice had 51 and 48% lower Ob.S/BS and Ob.N/BS and 73% greater Oc.S/BS when compared to their corresponding 11wk controls (p < 0.05), with no statistically significant differences in dynamic indices of bone formation between these two groups (Fig. 2; Table 1 ). In the B6J substrain, HLU mice had lower BV/TV (−31%), Tb.N (−5%), Tb.Th (−19%), and BS/BV (24%) when compared to 11wk controls (all p < 0.05). This architectural decline was accompanied by lower MS/BS (−39%), BFR/BS (−56%), Ob.S/BS (−40%), and Ob.N/B.Pm (−31%) (all p < 0.05). HLU induced differences in BS/BV were greater in B6N than in B6J mice (40 vs. 24%; Gene*HLU, p < 0.05). No other differences in trabecular variables were observed between B6N and B6J when comparing HLU to 11wk (Fig. 2; Table 1 ).
Relative mean differences (and standard error of the difference) in Tb.Sp, BS/BV, Tb.N, and Tb.Th between HLU (n = 12/substrain) and 9wk (n = 10/substrain) mice of B6N and B6J substrains. *p < 0.05 for differences between HLU and 9wk within each substrain; Gene*HLU = p value for the interaction between the factors gene (substrain) and HLU (unloading)
Even when compared to 9wk baseline control mice, rather than 11wk-old age-matched controls, HLU mice in both substrains had lower trabecular indices. In the B6N substrain, HLU mice had lower BV/TV (−65%), Tb.N (−20%), Tb.Th (−27%), Tb.Sp (26%), BS/BV (44%), and Conn.D (−86%). B6J HLU mice also had lower BV/TV (−44%), Tb.N (−12%), Tb.Th (−18%), Tb.Sp (14%), BS/BV (27%), and Conn.D (−50%) (all p < 0.05; Fig. 2; Table 1 ). For cellular indices, HLU B6N mice had lower MS/BS (−29%), Ob.S/BS (−60%), and Ob.N/B.Pm (−52%) while HLU B6J mice had 26% lower Ob.S/BS (all p < 0.05; Fig. 3; Table 3). When comparing the magnitude of unloading-induced trabecular degeneration between the two substrains, B6N mice were more susceptible than B6J. Compared to their respective 9-week control groups, unloaded B6N mice had a nearly twice as large increase in Tb.Sp (26 vs. 14%) and BS/BV (44 vs. 27%) than unloaded B6J mice (Gene*HLU, p < 0.05). Further, statistical trends of greater differences in Tb.Th and Tb.N were observed in B6N mice when compared to the B6J substrain (Gene*HLU, p = 0.08 and 0.06; Fig. 2; Table 1 ). When body mass was not used as covariate, differences persisted. Further, differences in the HLU response between B6N and B6J were also observed in dynamic indices of bone formation and bone cell indices. Briefly, HLU B6N mice exhibited greater suppression of Ob.S/BS (−60 vs. −26%) and Ob.N/BS (−52% vs. nsd) with respect to its corresponding 9-week mice when compared to its B6J counterparts (Gene*HLU, all p < 0.05; Fig. 3; Table 4 ). No difference in osteoclastic activity was observed between HLU and 9-week controls in both substrains.
Relative mean differences (and standard error of the difference) in osteoblast surface/bone surface (Ob.S/BS), and osteoblast number (N.Ob/B.Pm) between HLU (n = 12/substrain) and 9wk mice (n = 10/substrain). *p < 0.05 for differences between HLU and 9wk within each substrain; Gene*HLU = p value for the interaction between the factors gene (substrain) and HLU (unloading)
Cortical bone
HLU also compromised the cortical compartment of both B6N and B6J substrains of mice. When compared to 11wk controls, HLU B6N mice had lower Ct.Ar (−18%), Ct.Th (−9%), and Tt.Ar (−5%), while Ma.Ar was not different between the two groups (all p < 0.05). When compared to 11wk controls, HLU B6J mice had 15% lower Ct.Ar and 7% lower Ct.Th (all p < 0.05; Table 2; Fig. 4 ), while Ma.Ar and Tt.Ar was not different between the two groups. Compared to the B6J substrain, B6N mice exhibited greater HLU induced suppression of Ct.Ar (−18 vs. −15%) and Tt.Ar (−5% vs. nsd) when 9-week body mass was used as covariate (Gene*HLU, p < 0.05). No differences were observed between the two substrains if no covariates were used. Further, B6N HLU mice had 47 and 52% lower Ob.N/Ec.Pm and Ob.S/Ec.Pm when compared to 11wk mice while HLU B6J mice had 30 and 41% lower Ob.N/Ec.Pm and Ob.S/Ec.Pm (all p < 0.05). However, no difference in the magnitude of the cellular response was detected between the two substrains. On the periosteal surface, HLU B6N mice exhibited a greater suppression of Ob.N/Pe.Pm (−34% vs. nsd) and Ob.S/Pe.Pm (−37% vs. nsd) when compared to the B6J substrain (Gene*HLU, p < 0.05). On the endocortical surface, osteoclastic surface was not different between HLU and 11wk mice of the B6J substrain. In the B6N substrain, 2% of the endocortical surface of HLU group exhibited osteoclastic surface compared to 1% of the endocortical surface in 11wk B6N mice (p < 0.05 for the difference; Table 4 ). On the periosteal surface of B6N mice, HLU did not significantly affect osteoclast surface. In B6J mice, however, osteoclast surface increased fourfold with HLU (p < 0.05), an increase that was significantly greater in B6J than B6N (Gene*HLU, p < 0.05; Table 4 ).
Relative mean difference (and standard error of the difference) in Ct.Ar and Tt.Ar between HLU (n = 12/substrain) and 11wk (n = 12/substrain) mice of B6N and B6J substrains. *p < 0.05 for differences between HLU and 11wk within each substrain; Gene*HLU = p value for the interaction between the factors gene (substrain) and HLU (unloading)
In both substrains, cortical morphology was largely similar between HLU and 9wk controls, with the exception of greater Ct.Th (8%) in HLU B6J mice than 9wk controls (p < 0.05; Table 2). However, HLU resulted in suppressed osteoblastic surface and number in both substrains. HLU B6N mice had 45 and 50% lower Ob.N/Ec.Pm and Ob.S/Ec.Pm when compared to 9wk mice, while HLU B6J mice had 30 and 48% lower Ob.N/Ec.Pm and Ob.S/Ec.Pm when compared to 9wk mice (all p < 0.05). On the periosteal surface, HLU B6N mice exhibited greater suppression of Ob.N/Pe.Pm (−45% vs. nsd) and Ob.S/Pe.Pm (−46% vs. nsd) than HLU B6J mice when using 9-week mice as referent (Gene*HLU, p < 0.05; Table 4). In the B6N substrain, unloading significantly increased (fivefold) osteoclastic resorptive surfaces at the endocortical surface (p < 0.05; Table 4 ), contrasting with B6J mice in which no increase osteoclast surfaces was observed (Gene*HLU, p < 0.05). At the periosteal surface, HLU did not increase osteoclastic surface in B6N while in B6J a three-fold difference between HLU and 9wk controls was observed (all p < 0.05; Table 4), a differential response between the two substrains which did not reach statistical significance in two-way ANOVA.
Allelic differences in coding genomic regions
To identify genomic variations between B6N and B6J mice that may have contributed to the differences in baseline bone architecture and the response to unloading, we compared the non-synonymous SNP variations between B6N and B6J mice to an earlier genetic linkage study (Judex et al. 2013). This genetic linkage study in F2 BALBxC3H mice was aimed at identifying QTLs that modulate basal bone morphology, as well as its response to mechanical unloading. From the baseline QTLs in this genetic linkage study, we identified 326 SNPs that potentially modulate BV/TV, which amounted to 165 distinct targets after consolidating SNPs within the same gene and removing olfactory genes. Following identical data retrieval procedures for other µCT indices, we identified 95 SNPs and 56 genetic targets for BS/BV; 216 SNPs and 102 genetic targets for Tb.Th; 592 SNPs and 282 genetic targets for Tb.N; and 312 SNPs and 160 genetic targets for Conn.D. Similarly, from the SNPs within the unloading QTLs (Judex et al. 2013; Sankaran et al. 2015), we identified 681 SNPs and 335 distinct genes for BV/TV, 574 SNPs, and 263 genetic targets for BS/BV, 410 SNPs, and 162 genetic targets for Tb.Th, 125 SNPs, and 65 genetic targets for Tb.N, and 739 SNPs and 353 genetic targets for Conn.D. The SNPs at baseline and during unloading were separately compared to the SNPs between B6N and B6J mice. The mouse phenome database generated 60 non-synonymous SNPs between B6N and B6J mice. After consolidating SNPs within the same gene and removing olfactory receptor genes, we identified 52 distinct genetic targets that could contribute to the differences between B6N and B6J mice. From this list, the Herc2 and Acan were the only two genes that overlapped between the baseline QTL gene list and the SNP differences between B6N and B6J generated gene list, potentially contributing to basal differences between B6N and B6J substrains. Comparison of unloading QTL genes and the allelic differences between B6N and B6J yielded Herc2, Myo18b, and Acan as potential genes that may have influenced the differential response of B6N and B6J to unloading.
Discussion
The extent of genetic differences between commonly used substrains of the C57BL/6 mouse (Austin et al. 2004; Simon et al. 2013) is small. Nevertheless, C57BL/6N versus C57BL/6J substrains displayed distinct bone architecture and indices of bone formation at 9 and 11 weeks of age. At 9 weeks, B6N mice had lower trabecular indices than B6J whereas cortical indices were similar between the two substrains. In B6N, cellular indices of trabecular metabolism decreased between 9 and 11 weeks while in the B6J substrain, bone formation and bone cell indices increased. In contrast, cortical osteoblastic surface and number were not different between 9 and 11 weeks in both substrains. When compared to age-matched ambulatory controls, hindlimb unloading, a strongly catabolic (anti-anabolic) signal to bone, produced a similarly devastated trabecular compartment deterioration in the two strains. However, as trabecular bone development was different between 6N and 6J in ambulatory controls over the 2 weeks experimental duration, the response to HLU differed between the two substrains. B6N mice exposed to HLU exhibited a significantly greater deterioration of trabecular architecture and cellular indices from their 9 weeks baseline levels than B6J mice. Unloading also resulted in greater suppression of the cortical apposition in B6N. The differential sensitivity of bone to subtle genomic differences associated with individual mouse substrains may not only be important to the skeletal field but, by extrapolation, may also be applicable to other tissues and systems (Kiselycznyk and Holmes 2011).
Our data demonstrated that B6N mice had lower trabecular microarchitectural indices (BV/TV and Tb.Th) but greater osteoblastic indices (BFR/BS and Ob.N/BS) than B6J mice at 9 weeks (no differences at 11 weeks) while cortical architecture and osteoblastic indices were largely similar both at 9 and 11 weeks of age. This contrasts with a previous study in 14-week-old male and female B6N and B6J mice investigating differences in tibial bone properties as well as serum levels of osteocalcin (bone formation marker) and urinary creatinine (resorption marker) (Simon et al. 2013). No differences between the two substrains were found in either male or female mice. Given that trabecular differences in our study were observed only at 9 weeks but not 11 weeks without concomitant cortical differences, it is likely that the discrepancy between the two studies is related to the age of the mice (9 vs. 14 weeks) (Glatt et al. 2007). Moreover, the earlier study (Simon et al. 2013) used systemic markers versus site-specific histology in our study, and differences in sensitivity, specificity, and circadian influence on systemic markers, may have also contributed to differences in results (Eastell et al. 1992; Megraud 1996; Otter et al. 1997).
Unloading-induced trabecular degeneration was measured comparing HLU mice to either 9- or 11-week normal-control mice. The differential trabecular response between B6N and B6J substrains was evident only when comparing HLU to 9wk mice, with losses significantly greater in B6N mice when compared to B6J. In B6N mice, trabecular bone formation indices declined between 9 and 11 weeks of age and HLU further suppressed these indices. In B6J mice, however, trabecular bone formation indices increased between 9 and 11 weeks and HLU largely prevented this increase. Previous studies have shown that even in the absence of genetic differences (i.e., within an inbred strain), the response to mechanical signals (or lack thereof) can be modulated by basal bone formation indices (Squire et al. 2008). For instance, young adult (16 weeks) C3H mice with moderately high basal osteoblastic function were only mildly responsive to mechanical loading, but at 32 weeks when basal osteoblastic activity tapered, these mice responded to cyclic mechanical loading significantly with anabolic bone formation (Poliachik et al. 2008). Conversely, 16 weeks old C3H mice were unaffected by 2 or 3 weeks of hindlimb unloading (Amblard et al. 2003; Judex et al. 2004a) but growing C3H mice (9 weeks) with very high basal bone formation indices were highly susceptible to unloading (Li et al. 2016). Thus, skeletal regulation of mechanical loading and unloading are distinct (Judex et al. 2013; Kesavan et al. 2006) but basal bone formation rates can play a critical role in bones’ response to these signals. Extrapolated to our study, it is possible that the differences in HLU induced bone indices between B6N and B6J were mediated by the distinctly different basal bone formation rate and differential bone development pattern inherent to these two substrains.
Genetic control of skeletal structure and adaptation has been investigated both in human subjects and mouse models. For instance, Afro-Caribbean and African American men have significantly greater bone mineral density across femoral neck, hip, and lumbar spine when compared to Caucasian men (Nam et al. 2010). Preclinical studies consistently show large differences in bone’s susceptibility to disuse between inbred strains of mice (Amblard et al. 2003; Judex et al. 2004a; Squire et al. 2004). The difference between these previous and our current study is that the extent of the polymorphisms across the genome is much smaller. Identification of specific genes underlying the differences in the magnitude of the skeletal response to disuse is challenging. Inherently, the pool of candidate genes grows with the number of polymorphisms between mouse strains and may comprise several hundred genetic targets (Judex et al. 2013). By comparison to genes suggested in a previous genetic linkage study, here, we identified Herc2 and Acan as potential genes accounting for the basal skeletal differences between B6N and B6J and Herc2, Myo18b, and Acan as potential genetic modulators of the differential skeletal adaptation to unloading. These genes add to a previously identified structural variation in the intronic portion of CSMD1 between B6N and B6J mice (Simon et al. 2013), which also lies within the interval of our earlier unloading QTL analysis (Judex et al. 2013).
Herc2 plays a role in regulatory ubiquitination in damaged DNA (Bekker-Jensen et al. 2009). Mice with mutations in the Herc2 gene also exhibit neuromuscular disorders (Lehman et al. 1998; Walkowicz et al. 1999) and suffer from impaired growth and movement coordination (Scheffner and Kumar 2014). CSMD1 (Cub and Sushi multiple domain protein 1) is postulated to be a membrane protein (Kamal et al. 2010) and as a tumor suppressor, plays a role in signal transduction. Loss or mutations of this gene have been reported in cancers (Ding et al. 2010; Kamal et al. 2010; Yokota et al. 1999). The regulatory function of Herc2 and the signal transduction capability of CSMD1 could be activated differently or to a varying extent in B6N and B6J mice upon initiating the catabolic signal of hindlimb unloading. Our data, however, somewhat argue against this possibility as the differential response to unloading between B6N and B6J was likely due to differential growth rather than to differences in signal transduction.
Myo18b is commonly expressed in cardiac and skeletal muscle, regulating intracellular transport or controlling gene transcription (Salamon et al. 2003; Simon and Wilson 2011). Polymorphism in Myo18b may have differentially modulated muscle adaptation between B6N and B6J mice (not measured here) and, via altered muscle-bone interactions, directly affected basal bone morphology as well as bone’s susceptibility to unloading (Burr 1997). Aggrecan, the product of the Acan gene, is found predominantly in cartilage and regulates tissue hydration and mechanical properties. A 50% reduction in aggrecan gene expression was observed with unloading in rats, and while the protein is mainly expressed in cartilage, it is also found in bone (Sibonga et al. 2000; Wong et al. 1992). Thus, a polymorphism in Acan between B6N and B6J has the potential to trigger a differential unloading response. Other polymorphisms known between these two substrains of mice, for instance the ns-SNP in BMPr2 (bone morphogenetic protein receptor 2), ADAMTS3 or ADAMTS12 (a disintegrin-like and metalloprotease, thrombospondin) also play a role in bone metabolism and growth (Le Goff et al. 2006; Shi and Massagué 2003) and may have influenced differences observed between B6N and B6J mice.
The distinct bone microarchitecture, developmental paradigm, and unloading response between B6N and B6J mice identified here, when taken in the context of the heterogeneous human population, suggest that patient specific medical care is not only beneficial for genetically diverse ethnic/racial demographics but may even be applicable to a relatively homogenous population (Schwartz 2001; Wilson et al. 2001; Witherspoon et al. 2007). For preclinical models, this data highlight that subtle allelic differences within a mouse strain can induce a bone phenotype, suggesting that for many investigations the B6N and B6J substrains cannot be used interchangeably and that transgenic mouse models built on these two closely-related substrains may be afflicted with differential physiologic responses.
References
Alfredson H, Nordstrom P, Lorentzon R (1996) Total and regional bone mass in female soccer players. Calcif Tissue Int 59:438–442
Amblard D, Lafage-Proust MH, Laib A, Thomas T, Rüegsegger P, Alexandre C, Vico L (2003) Tail suspension induces bone loss in skeletally mature mice in the C57BL/6J strain but not in the C3H/HeJ strain. J Bone Miner Res 18:561–569
Austin CP, Battey JF, Bradley A, Bucan M, Capecchi M, Collins FS, Dove WF, Duyk G, Dymecki S, Eppig JT (2004) The knockout mouse project. Nat Genet 36:921–924
Bayat A, Barton A, Ollier WE (2004) Dissection of complex genetic disease: implications for orthopaedics. Clin Orthop Relat Res 419:297–305
Beamer W, Donahue L, Rosen C, Baylink D (1996) Genetic variability in adult bone density among inbred strains of mice. Bone 18:397–403
Beckwith J, Cong Y, Sundberg JP, Elson CO, Leiter EH (2005) Cdcs1, a major colitogenic locus in mice, regulates innate and adaptive immune response to enteric bacterial antigens. Gastroenterol 129:1473–1484
Bekker-Jensen S, Danielsen JR, Fugger K, Gromova I, Nerstedt A, Lukas C, Bartek J, Lukas J, Mailand N (2009) HERC2 coordinates ubiquitin-dependent assembly of DNA repair factors on damaged chromosomes. Nat Cell Biol 12:80–86
Burr DB (1997) Muscle strength, bone mass, and age-related bone loss. J Bone Miner Res 12:1547–1551
Ding L, Ellis MJ, Li S, Larson DE, Chen K, Wallis JW, Harris CC, McLellan MD, Fulton RS, Fulton LL (2010) Genome remodelling in a basal-like breast cancer metastasis and xenograft. Nature 464:999–1005
Eastell R, Calvo MS, Burritt MF, Offord KP, Russell R, Riggs BL (1992) Abnormalities in circadian patterns of bone resorption and renal calcium conservation in type I osteoporosis. J Clin Endocrinol Metab 74:487–494
Eisman JA (1999) Genetics of osteoporosis. Endocr Rev 20:788–804
Frazer KA, Murray SS, Schork NJ, Topol EJ (2009) Human genetic variation and its contribution to complex traits. Nat Rev Genet 10:241–251
Glatt V, Canalis E, Stadmeyer L, Bouxsein ML (2007) Age-related changes in trabecular architecture differ in female and male C57BL/6J mice. J Bone Miner Res 22:1197–1207
Globus RK, Morey-Holton E (2016) Hindlimb unloading: rodent analog for microgravity. J Appl Physiol 120:1196–1206
Green DE, Adler BJ, Chan ME, Rubin CT (2012) Devastation of adult stem cell pools by irradiation precedes collapse of trabecular bone quality and quantity. J Bone Miner Res 27:749–759
Grubb SC, Bult CJ, Bogue MA (2014) Mouse phenome database. Nucleic Acids Res 42:D825-D834
Haan J, Terwindt GM, Ferrari MD (1997) Genetics of migraine. Neurol Clin 15:43–60
Hansen MF, Cavenee WEEK (1987) Genetics of cancer predisposition. Cancer Res 47:5518–5527
Judex S, Garman R, Squire M, Busa B, Donahue LR, Rubin C (2004a) Genetically linked site-specificity of disuse osteoporosis. J Bone Miner Res 19:607–613
Judex S, Garman R, Squire M, Donahue LR, Rubin C (2004b) Genetically based influences on the site-specific regulation of trabecular and cortical bone morphology. J Bone Miner Res 19:600–606
Judex S, Zhang W, Donahue LR, Ozcivici E (2013) Genetic loci that control the loss and regain of trabecular bone during unloading and reambulation. J Bone Miner Res 28:1537–1549
Kamal M, Shaaban AM, Zhang L, Walker C, Gray S, Thakker N, Toomes C, Speirs V, Bell SM (2010) Loss of CSMD1 expression is associated with high tumour grade and poor survival in invasive ductal breast carcinoma. Breast Cancer Res Treat 121:555–563
Kesavan C, Mohan S, Srivastava AK, Kapoor S, Wergedal JE, Yu H, Baylink DJ (2006) Identification of genetic loci that regulate bone adaptive response to mechanical loading in C57BL/6J and C3H/HeJ mice intercross. Bone 39:634–643
Kiselycznyk C, Holmes A (2011) All (C57BL/6) mice are not created equal. Front Neurosci 5:10
Korte A, Farlow A (2013) The advantages and limitations of trait analysis with GWAS: a review. Plant Methods 9:1
Le Goff C, Somerville RP, Kesteloot F, Powell K, Birk DE, Colige AC, Apte SS (2006) Regulation of procollagen amino-propeptide processing during mouse embryogenesis by specialization of homologous ADAMTS proteases: insights on collagen biosynthesis and dermatosparaxis. Development 133:1587–1596
Lehman AL, Nakatsu Y, Ching A, Bronson RT, Oakey RJ, Keiper-Hrynko N, Finger JN, Durham-Pierre D, Horton DB, Newton JM (1998) A very large protein with diverse functional motifs is deficient in rjs (runty, jerky, sterile) mice. Proc Natl Acad Sci 95:9436–9441
Li B, Sankaran JS, Judex S (2016) Trabecular and cortical bone of growing C3H mice is highly responsive to the removal of weightbearing. PLoS ONE 11:e0156222
Lloyd K, Franklin C, Lutz C, Magnuson T (2015) Reproducibility: use mouse biobanks or lose them. Nature 522:151–153
Lublinsky S, Ozcivici E, Judex S (2007) An automated algorithm to detect the trabecular-cortical bone interface in micro-computed tomographic images. Calcif Tissue Int 81:285–293
Mähler M, Leiter EH (2002) Genetic and environmental context determines the course of colitis developing in IL–10-deficient mice. Inflamm Bowel Dis 8:347–355
Mavaddat N, Antoniou AC, Easton DF, Garcia-Closas M (2010) Genetic susceptibility to breast cancer. Mol Oncol 4:174–191
Megraud F (1996) Advantages and disadvantages of current diagnostic tests for the detection of Helicobacter pylori. Scand J Gastroenterol 31:57–62
Nam H-S, Shin M-H, Zmuda J, Leung P, Barrett-Connor E, Orwoll E, Cauley J, Group OFiMR (2010) Race/ethnic differences in bone mineral densities in older men. Osteoporos Int 21, 2115–2123
Nicholson A, Reifsnyder PC, Malcolm RD, Lucas CA, MacGregor GR, Zhang W, Leiter EH (2010) Diet-induced obesity in two C57BL/6 substrains with intact or mutant nicotinamide nucleotide transhydrogenase (Nnt) gene. Obesity 18:1902–1905
Noh T, Gabet Y, Cogan J, Shi Y, Tank A, Sasaki T, Criswell B, Dixon A, Lee C, Tam J (2009) Lef1 haploinsufficient mice display a low turnover and low bone mass phenotype in a gender-and age-specific manner. PLoS ONE 4:e5438
Otter A, Jeffrey M, Scholes S, Helmick B, Wilesmith J, Trees A (1997) Comparison of histology with maternal and fetal serology for the diagnosis of abortion due to bovine neosporosis. Vet Rec 141:487–489
Plomin R, Haworth CM, Davis OS (2009) Common disorders are quantitative traits. Nat Rev Genet 10:872–878
Poliachik SL, Threet D, Srinivasan S, Gross TS (2008) 32 wk old C3H/HeJ mice actively respond to mechanical loading. Bone 42:653–659
Romeo S, Kozlitina J, Xing C, Pertsemlidis A, Cox D, Pennacchio LA, Boerwinkle E, Cohen JC, Hobbs HH (2008) Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nat Genet 40:1461–1465
Rosen CJ, Ackert-Bicknell C, Rodriguez JP, Pino AM (2009) Marrow fat and the bone microenvironment: developmental, functional, and pathological implications. Critical Rev™ Eukaryot Gene Expr 19:109–124
Salamon M, Millino C, Raffaello A, Mongillo M, Sandri C, Bean C, Negrisolo E, Pallavicini A, Valle G, Zaccolo M (2003) Human MYO18B, a novel unconventional myosin heavy chain expressed in striated muscles moves into the myonuclei upon differentiation. J Mol Biol 326:137–149
Sankaran JS, Li B, Donahue LR, Judex S (2015) Modulation of unloading-induced bone loss in mice with altered ERK signaling. Mamm Genome 27:1–15
Scheffner M, Kumar S (2014) Mammalian HECT ubiquitin-protein ligases: biological and pathophysiological aspects. Biochim Biophys Acta Mol Cell Res 1843:61–74
Schwartz RS (2001) Racial profiling in medical research. N Engl J Med 344:1392–1393
Shi Y, Massagué J (2003) Mechanisms of TGF-β signaling from cell membrane to the nucleus. Cell 113:685–700
Sibonga J, Zhang M, Evans G, Westerlind K, Cavolina J, Morey-Holton E, Turner R (2000) Effects of spaceflight and simulated weightlessness on longitudinal bone growth. Bone 27:535–540
Simon DN, Wilson KL (2011) The nucleoskeleton as a genome-associated dynamic’network of networks’. Nat Rev Mol Cell Biol 12:695–708
Simon MM, Greenaway S, White JK, Fuchs H, Gailus-Durner V, Wells S, Sorg T, Wong K, Bedu E, Cartwright EJ (2013) A comparative phenotypic and genomic analysis of C57BL/6J and C57BL/6N mouse strains. Genome Biol 14:R82
Squire M, Donahue LR, Rubin C, Judex S (2004) Genetic variations that regulate bone morphology in the male mouse skeleton do not define its susceptibility to mechanical unloading. Bone 35:1353–1360
Squire M, Brazin A, Keng Y, Judex S (2008) Baseline bone morphometry and cellular activity modulate the degree of bone loss in the appendicular skeleton during disuse. Bone 42:341–349
van Amerongen R, Berns A (2006) Knockout mouse models to study Wnt signal transduction. Trends Genet 22:678–689
Vang T, Congia M, Macis MD, Musumeci L, Orrú V, Zavattari P, Nika K, Tautz L, Taskén K, Cucca F (2005) Autoimmune-associated lymphoid tyrosine phosphatase is a gain-of-function variant. Nat Genet 37:1317–1319
Visscher PM, Hill WG, Wray NR (2008) Heritability in the genomics era—concepts and misconceptions. Nat Rev Genet 9:255–266
Walkowicz M, Ji Y, Ren X, Horsthemke B, Russell LB, Johnson D, Rinchik EM, Nicholls RD, Stubbs L (1999) Molecular characterization of radiation-and chemically induced mutations associated with neuromuscular tremors, runting, juvenile lethality, and sperm defects in jdf2 mice. Mamm Genome 10:870–878
Wilson JF, Weale ME, Smith AC, Gratrix F, Fletcher B, Thomas MG, Bradman N, Goldstein DB (2001) Population genetic structure of variable drug response. Nat Genet 29:265–269
Witherspoon DJ, Wooding S, Rogers AR, Marchani EE, Watkins WS, Batzer MA, Jorde LB (2007) Genetic similarities within and between human populations. Genetics 176:351–359
Wong M, Lawton T, Goetinck P, Kuhn J, Goldstein S, Bonadio J (1992) Aggrecan core protein is expressed in membranous bone of the chick embryo. Molecular and biomechanical studies of normal and nanomelia embryos. J Biol Chem 267:5592–5598
Yokota T, Yoshimoto M, Akiyama F, Sakamoto G, Kasumi F, Nakamura Y, Emi M (1999) Localization of a tumor suppressor gene associated with the progression of human breast carcinoma within a 1-cm interval of 8p22–p23. 1. Cancer 85:447–452
Acknowledgements
We are grateful for financial support from NASA (NNX12AL25G). The authors thank Leah Rae Donahue for providing the mice and Alyssa Tuthill, Tee Pamon, and Sherin Kuriakose for technical assistance. This work was in part presented in the doctoral dissertation of Jeyantt Sankaran, the first author of this manuscript (Sankaran JS, 2016, Identification of Genes that Modulate Bone Loss during Mechanical Unloading. State University of New York at Stony Brook).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Statement on the Welfare of Animals
All procedures were reviewed and approved by Stony Brook University’s Institutional Animal Care and Use Committee (IACUC).
Rights and permissions
About this article
Cite this article
Sankaran, J.S., Varshney, M. & Judex, S. Differences in bone structure and unloading-induced bone loss between C57BL/6N and C57BL/6J mice. Mamm Genome 28, 476–486 (2017). https://doi.org/10.1007/s00335-017-9717-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00335-017-9717-4