Abstract
In recent years, there has been a dramatic increase in research papers about machine learning (ML) and artificial intelligence in radiology. With so many papers around, it is of paramount importance to make a proper scientific quality assessment as to their validity, reliability, effectiveness, and clinical applicability. Due to methodological complexity, the papers on ML in radiology are often hard to evaluate, requiring a good understanding of key methodological issues. In this review, we aimed to guide the radiology community about key methodological aspects of ML to improve their academic reading and peer-review experience. Key aspects of ML pipeline were presented within four broad categories: study design, data handling, modelling, and reporting. Sixteen key methodological items and related common pitfalls were reviewed with a fresh perspective: database size, robustness of reference standard, information leakage, feature scaling, reliability of features, high dimensionality, perturbations in feature selection, class balance, bias-variance trade-off, hyperparameter tuning, performance metrics, generalisability, clinical utility, comparison with traditional tools, data sharing, and transparent reporting.
Key Points
• Machine learning is new and rather complex for the radiology community.
• Validity, reliability, effectiveness, and clinical applicability of studies on machine learning can be evaluated with a proper understanding of key methodological concepts about study design, data handling, modelling, and reporting.
• Understanding key methodological concepts will provide a better academic reading and peer-review experience for the radiology community.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
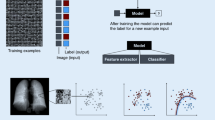
As a subfield of artificial intelligence, machine learning (ML) is the study of computer algorithms that learn from the input data and make predictions on unseen instances [1, 2]. ML algorithms are designed to operate without specific rule-based instructions, improving themselves by learning and correcting through experience [1,2,3,4]. ML is broadly grouped into two categories with a key difference in the use of labels for model development. While supervised learning needs labels in training, unsupervised learning requires no labels in discovering patterns in data sets. Several ML algorithms exist with a wide range of complexity levels. Of which, deep learning is a particular field of ML and capable of handling very large amount of data, even without any need for traditional feature extraction.
Owing to the advances in deep learning, ML has been rapidly expanding in recent years, invading as much radiology environment as possible [5, 6]. Being mostly based on supervised learning, the application area of ML in radiology is vast, ranging from image acquisition to outcome prediction [1]. Common ML tasks in radiology are prioritising worklists [7], classification of reports [8], risk assessment [9], screening [10], detection [11], segmentation [12], histopathologic diagnosis [13], radiogenomics [14, 15], and image acquisition improvement [16]. Although ML offers various opportunities for radiology, it also brings along many methodological challenges and pitfalls [6, 17, 18]. Interestingly, many of these are not specific to radiology and have already been addressed in other fields such as genomics, biostatistics, and bioinformatics [19,20,21,22,23].
Despite the widespread interest in ML, the methodology of ML papers is mostly complex and elusive for the radiology community. Without a good understanding of key methodological concepts, it might be very hard for radiologists to make a proper assessment and critique of the works as to their validity, reliability, effectiveness, and clinical applicability.
In this paper, we aimed to provide radiologists with a fresh perspective on how to evaluate the published literature and manuscript drafts that use ML in radiology. To achieve this, we concentrated on sixteen key methodological quality concepts of ML.
Key methodological concepts
ML-based pipelines vary to a large extend [24]. On the other hand, core concepts usually remain the same. Important steps of an ML pipeline can be grouped as follows: design, data handling, modelling, and reporting. A simplified illustration of these steps is given in Fig. 1. Key concepts that need attention in the evaluation of ML papers are given in Fig. 2. Before going into detail, a glossary of basic ML terminology is given in Table 1.
Key concepts of study design
Common pitfalls and recommendations for study design are summarised in Table 2.
Database size
ML projects need large and heterogeneous data sets to ensure generalisability. However, this is usually hardly achieved in radiology research due to a variety of reasons. A common pitfall that can be avoided is to train a model with an extremely small data set. Such a premature strategy poses many challenges to deal with, for instance, overfitting, noise, and outliers.
To our best knowledge, there is currently no well-adopted method for determining the optimal database size for ML and all proposed strategies are empirical. Statistical power calculations might result in thousands of instances, even for establishing the testing set, which seems hardly achievable for all radiology tasks. To minimise the effects of overfitting and improve the quality of predictive performance metrics, the inclusion of at least 50 instances might be sufficient for initial research [5, 25,26,27]. On the other hand, this number would be inappropriate for the development of highly generalisable and clinically useful real-world ML applications. Another common recommendation is to have a data size that is more than ten times the number of features [28, 29]. Furthermore, the complexity of algorithms (e.g., k-nearest neighbours versus deep learning) and tasks (e.g., substantially heterogeneous population or subtle discretionary features) should always be considered when deciding the appropriateness of database size. Aside from these recommendations, another well-known strategy is to plot a learning curve for error or accuracy values versus training data size [30].
Robustness of reference standard
The reference standard is usually an accepted test or a gold standard or expert diagnosis. Source and rationale of the reference standard must be clearly mentioned in ML papers.
Robustness of reference standard corresponds to the stability of labels in varying conditions such as different readers, scanners, or technical protocols, which is critical not only for high-quality model development but also for the overall success of the project [31]. Strategies for reducing such variabilities would be consensus evaluations by experts, majority voting, or selecting a reference standard that is less sensitive to variabilities. It is noteworthy to mention that these concerns on the robustness of reference standards are much more important in medicine compared with other fields because even a small difference in predictive performance might have a significant influence on a large patient population.
Key concepts of data handling
Common pitfalls and recommendations for data handling are summarised in Table 2.
Information leakage
Information leakage is one of the most significant pitfalls in ML modelling. It can be simply defined as the transmission of information among training, validation, and testing datasets, due to incomplete separation of the data.
The information leakage might occur in any stage of the ML pipeline. One should be very careful in detecting this pitfall because its occurrence might not be so obvious and can be easily missed [32], even if separate validation and test partitions are reported. Information leakage can be frequently encountered in following data handling steps: feature scaling, dimension reduction or feature selection, and hyperparameter tuning. It can be minimised or completely avoided through careful data separation [33, 34].
It should also be kept in mind that data leakage usually occurs at the first steps of the pipeline. Therefore, the data split must be done at the beginning of the pipeline. In other words, the separation of the data set must be done just after designating the raw data inputs because even preprocessing of the images (e.g., grey-level discretisation according to bin-width) before feature extraction might lead to leakage, causing optimistic results.
Feature scaling
Feature values are usually presented in different scales, which need to be considered in many ML classification tasks because the parameters of some algorithms are influenced by the scale of features. Particularly, distance-based algorithms like support vector machine, k-nearest neighbours, and artificial neural networks significantly benefit from feature scaling. On the other hand, some other algorithms like tree-based random forest do not need such requirements. Feature scaling can be done in a few ways. The most common approaches are standardisation, normalisation, and logarithmic transformation [35]. It is also important to note that scaling is an integral part of the neural network and deep learning architectures [36, 37].
It is worth mentioning that feature scaling is a completely different task from the scaling of images or image intensities [24]. The latter is commonly used to avoid some challenges posed by the scanner or site-specific variabilities in the field of newly emerging radiomics [24]. Neglecting feature scaling may lead to overrepresentation or underrepresentation of some features and cause bias in the analysis.
Reliability of features
The reliability of features defines the reproducibility of the features during extraction. Reliable features are highly resistant to changing conditions of the feature extraction process, for instance, segmentation margin differences [38], use of different image slice (i.e., slice selection bias in 2D analysis) [39, 40], and scanning protocol differences [41,42,43]. When analysing medical images, some preprocessing steps (e.g., pixel/voxel resampling, intensity normalisation) are necessary to obtain reliable features [44]. However, despite these measures, reliability remains a challenge [42, 45, 46]. The reliability can be assessed with several approaches such as intra-reader and inter-reader agreement analysis for detection of manual and semi-automatic segmentation differences [39, 47, 48], test-retest analysis for automatic methods [49], reproducibility analysis with different scanners or scanning protocols [46], and phantom or simulation measurements [43].
Development of models without a reliability assessment may lead to significant generalisability problems [50]. Nevertheless, there are some very interesting works with no attempt in selecting reliable features for their models [51,52,53]. Disregarding the reliability of features should be acceptable on the condition that the work has a true external independent validation cohort that is large enough to avoid bias and misleading conclusions.
High dimensionality
Advances in radiomics approaches have led to the extraction of a very high number of features, that is, high dimensionality. High dimensionality is considered a challenge to be dealt with in ML because it may induce multicollinearity, overfitting, and false discovery. Hence, redundant features should be eliminated through certain dimension reduction strategies.
Several methods can be used for dimension reduction [54]. Intra-reader and inter-reader feature reliability analysis, multicollinearity analysis, clustering, principal component analysis, and independent component analysis are the most common unsupervised methods. On the other hand, algorithm-based feature selection (e.g., wrapper, embedded, or filtering methods) is the most widely used supervised method [55, 56].
Perturbations in feature selection
The algorithm-based feature selection process has some inherent susceptibilities about the data structure such as data size, the order of input data, and initiations. Particularly when using small data, such susceptibilities might be much more apparent. Therefore, a common pitfall is to select features without considering possible perturbations in feature selection, which might lead to inappropriate feature selection and in turn generalisability problems [57, 58]. To minimise such susceptibilities of feature selection, the easiest way is to select features with multiple sampling, folding, or random initiations.
Key concepts of modelling
Common pitfalls and recommendations for modelling are summarised in Table 3.
Class balance
Class imbalance is an important issue in ML [45]. In case of severe imbalance, some algorithms are tended to vote for the majority class, inducing unrealistic outcomes and in turn very poor generalisability [59]. Ignoring the class imbalance is a major pitfall in ML modelling. For this reason, ML-based analysis with severe imbalance should include certain measures such as oversampling (synthetic or original), undersampling, or training with a trade-off between sensitivity and specificity.
It is also noteworthy to mention that the sampling strategies are usually recommended only for the training set. The rule of thumb is that no sampling should be done on the testing set. This is particularly important in the medical context because balancing the classes in test data might distort the actual disease prevalence, yielding poor clinical risk predictions. Also, undersampling should be used cautiously in the medical context because it might increase the risk of overfitting [60].
Bias-variance trade-off
Finding the trade-off between bias and variance is a vital task in ML to obtain models that generalise well (Fig. 3). Bias-variance trade-off can be achieved by a couple of methods. First, different algorithms with a wide range of complexity levels along with different penalisation and regularisation strategies might be evaluated with systematic validation methods. Then, the one that minimises the total prediction error is chosen. Second, some strong resampling techniques can be incorporated into modelling. For instance, bootstrap aggregating or bagging can be used to reduce variance. Third, optimisation or tuning of the models with adjustments of hyperparameters can be used. Besides, the number of hyperparameters can also be changed in this context. Fourth, data size can be altered to establish optimal trade-off.
Bias-variance trade-off and related concepts. (a) In simple terms, bias and variance are simply prediction errors. Bias is the difference between predictions (black dots) and actual values (light blue areas) that occurs when prediction models are prejudiced. Variance, on the other hand, is the level of variability and spread between predictions (black dots) and actual values (light blue areas). (b) If an algorithm is too complex, it may learn noise in training data, leading to good training and poor test performance, which is called overfitting. If an algorithm is too simple, it may not learn important aspects of data, leading to poor performance in training and testing, which is called underfitting. There is an opposite relationship between bias and variance. If one increases, the other one decreases, or vice versa. Suboptimal bias-variance trade-off leads overfitting or underfitting. High variance leads to overfitting, whereas high bias leads to underfitting. Finding trade-off between bias and variance is an important task in machine learning modelling to obtain models that generalise well. Usually, what matters is the total error, not particular items like bias and variance. In practice, there is no single analytical method to find optimal trade-off zone. In finding trade-off, it is critical to experiment with different model complexity levels to find the one that minimises overall error most
Hyperparameter tuning
ML models include parameters and hyperparameters with which modelling behaviour is configured for a given task. Parameters (e.g., support vectors in support vector machine, and weights in artificial neural networks) are internally calculated from the input data, whereas hyperparameters (e.g., C of support vector machine and learning rate of artificial neural network) are configured externally. Practitioners of ML cannot directly interfere with model parameters while the model operates. However, the selection of some parameters (e.g., type of loss function) before training is totally depended on the practitioner and peculiarities of the data being studied (e.g., organ, disease).
Determining the best hyperparameters, which is called hyperparameter tuning or optimisation, is also a crucial task in ML-based modelling [61]. In hyperparameter tuning, the aim is to find the optimal set of hyperparameters that reduces a predefined loss function and increases the predictive performance of the model on independent test data. In this context, one must always question the methodology of the papers as to the use of default hyperparameter configuration and copying from previous related works. The most common hyperparameter tuning strategies are manual configuration, automated random search, and grid search.
Performance metrics
The discriminative performance of ML models is generally evaluated using accuracy or area under the receiver operating characteristic curve. Furthermore, sensitivity, specificity, positive predictive value, negative predictive value, and the confusion matrix should be the minimum requirements in reporting the predictive performance in the medical context. It is also noteworthy that the confusion matrix itself is not only important for the calculation of other various metrics but also for eligibility in future meta-analyses. Concordance index and dice coefficient are other common relevant discriminative performance metrics used for survival analysis and segmentation performance, respectively. For the regression models with continuous results, the following metrics should be included: R squared, mean squared error, root mean squared error, root mean squared logarithmic error, and mean absolute error.
Both for classification and regression tasks, all performance metrics should be separately reported both for the training and testing sets because these are informative in the assessment of the fitting status of models. Furthermore, in comparative studies such as ML versus human expert reading, care should be taken to report the same metrics while comparing the performance of methods being used.
In the case of class imbalance, the Matthews correlation coefficient, F1 measure, area under the receiver operating characteristic curve, and area under the precision-recall curve are important metrics to be included in the results. Furthermore, a detailed evaluation of the confusion matrix and “no-information rate” is also helpful in the assessment of any work that suffers from class imbalance.
Metrics at a single point might be misleading in performance evaluation. This is particularly important when dealing with small data. Thus, the variability of performance metrics should be reported as well. The confidence interval, standard deviation, and standard error are common indicators of performance variability.
Generalisability
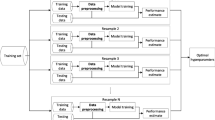
Generalisability in ML can be defined as the adaptability of models to previously unseen examples. It is assessed with two strategies: internal validation and independent validation. However, internal validation might lead to an overestimation of performance. Thus, the assessment of generalisability using an independent data set is important. For a true generalisability assessment, independent validation set must correctly represent the actual population of interest, for instance, in terms of disease prevalence and demographics, etc. It is very common to encounter a lack of transparency as to whether the independent validation set was truly independent owing to inconsistent terminology. An independent validation can ideally be achieved by the participation of external institutions. On the other hand, it is noteworthy that scanner-based independent validation in the same institution could be as valuable as the institution-based external independent validation. Validation terminology and simplified strategies are summarised in Fig. 4.
Simplified validation strategies in machine learning. In general, machine learning projects include three data partitions: training, validation, and testing. The training set is iteratively used to establish optimal parameter values that are special to each machine learning algorithm. Internal performance of the model is evaluated through a validation set (i.e., tuning set). Following many iterations of training and validation, the model is fed to unseen test data for its final performance evaluation. (a) Splitting data into training and testing sets. If the testing set includes instances from the same institution or same scanner, the method is called hold-out. If it comes from another institution or another scanner, the method is called independent. The training set includes other validation or sometimes testing parts. The training part should be used in dimension reduction, model development, and hyperparameter tuning. The testing part must be locked at the beginning of the study, to prevent bias in performance evaluation. (b) Cross-validation. This method has no overlap among the validation parts. The validation part can be a proportion of data (e.g., ten-fold cross-validation) or a single instance (i.e., leave-one-out cross-validation) in each sampling. (c) Random sampling. This method has overlaps among validation parts. On the other hand, its major strength is the number of iterations that is much more than that of simple cross-validation. Most common techniques with random sampling are bootstrap validation and random subsampling, with a key difference in replacement technique. (d) Nested cross-validation. Being rather a complex method, it includes separate testing parts, without overlap. Thus, it simulates previously described hold-out method. V, validation
Clinical utility
ML papers usually focus on performance metrics in the assessment of the diagnostic value of the method proposed. Assessment for clinical utility is often disregarded in ML-based classification tasks. Therefore, the claims about the improved predictive performance of ML tools in classifications remain uncertain and weak. The most common tools for this purpose are calibration statistics [62] and decision curve analysis [63].
Calibration statistics is the process in determining whether the predicted probability scores match with the actual probability scores. Rather than categorical outputs of ML models such as benign versus malignant, the use of probability scores for each target class might be much more useful in radiological decision-making, providing confidence in the diagnosis. A clinically useful model should be well-calibrated, having a balance between real and predicted probability scores. A calibration plot can be used to better present the calibration of the models (Fig. 5).
Decision curve analysis provides complementary information about the net benefits of the model proposed [63, 64]. This is a powerful clinical tool because it takes into account both discriminatory predictive performance and calibration of the models. A simple decision curve example and its basic interpretation are presented in Fig. 6.
Decision curve analysis for classification tasks. Higher the curve, the higher the sensitivity. Flatter the curve, higher the specificity. Each model curve is interpreted according to a reasonable probability range. The standard line of no medical action (e.g., intervention, surgery, drug therapy, additional diagnostic test) (a) for all instances. The standard line of full medical action (b) for all instances, regardless of diagnosis. Line of a model with high sensitivity and specificity (c), which is better than the other two models (d, e). Line of a model with high sensitivity and low specificity (d). Line of a model with low sensitivity and high specificity (e). TP, true positive
Comparison with traditional tools
As for all newly emerging techniques, the usefulness of ML in radiology should be assessed through comparisons with the traditional methods. Unless a new ML technique offers improvements over traditional methods, it is not intuitive to propose that technique for clinical usage. Therefore, ML papers should include relevant comparisons with traditional statistical modelling or clinical tools. Otherwise, reporting the ML results in isolation would not reflect and influence the clinical practice, limiting our ability to deploy in real-world health-care practice. Potential targets for comparisons would be traditional modelling techniques such as logistic regression and other clinical tools (e.g., qualitative expert readings) that have already been used in daily radiology practice. Such comparisons should be made on the same data sets. While making comparisons, potential negative results are also as valuable as the positive results and should be reported in the publications.
Key concepts of reporting
Common pitfalls and recommendations for reporting are summarised in Table 3.
Sharing data
Sharing data is important for replicability, proper quality assessment, and improvement of the proposed methodology. However, most research papers do not share their relevant data. This could be because of a few reasons. The authors might not be aware of the importance of data transparency. They might want to protect their data from potential misuse. Furthermore, they might even have a fear of falsification or negative comments from other researchers.
Authors of ML papers in radiology should consider sharing their image data, feature data, scripts used for modelling, and resultant model file. Sharing image data might be difficult due to the high volume and technical issues along with ethical and privacy-related concerns [65]. However, feature data, code scripts, and model files can be easily shared using online repositories.
Transparent reporting
Considering the abundance of easy-to-use and open-source toolboxes, it has never been so easy to develop an ML model for a given medical task. In such an environment, transparent reporting in every part of the study is the key to maintain the quality and replicability of the studies. Besides, the factors that limit the generalisability of an ML model to a certain case should not be ignored and must be transparently reported.
Adhering the checklists or guidelines would be the best practice in transparent reporting. Recent seminal work produced a significant checklist called CLAIM (Checklist for Artificial Intelligence in Medical Imaging) that is particularly designed for reporting the artificial intelligence–based research in the field of medical imaging [66]. Also, one can benefit from the following references for the same purpose [67,68,69].
Conclusions
In this paper, we systematically provided the key methodological concepts of ML to improve the academic reading and peer-review experience of radiology community. Although the recommendations given in this paper are not exclusive and do not guarantee an error-free evaluation, we hope it will serve as a guide for high-quality assessment.
Abbreviations
- ML:
-
Machine learning
References
Choy G, Khalilzadeh O, Michalski M et al (2018) Current applications and future impact of machine learning in radiology. Radiology 288:318–328. https://doi.org/10.1148/radiol.2018171820
Wang S, Summers RM (2012) Machine learning and radiology. Med Image Anal 16:933–951. https://doi.org/10.1016/j.media.2012.02.005
Jordan MI, Mitchell TM (2015) Machine learning: trends, perspectives, and prospects. Science 349:255–260. https://doi.org/10.1126/science.aaa8415
Kohli M, Prevedello LM, Filice RW, Geis JR (2017) Implementing machine learning in radiology practice and research. AJR Am J Roentgenol 208:754–760. https://doi.org/10.2214/AJR.16.17224
Sollini M, Antunovic L, Chiti A, Kirienko M (2019) Towards clinical application of image mining: a systematic review on artificial intelligence and radiomics. Eur J Nucl Med Mol Imaging 46:2656–2672. https://doi.org/10.1007/s00259-019-04372-x
Hosny A, Parmar C, Quackenbush J, Schwartz LH, Aerts HJWL (2018) Artificial intelligence in radiology. Nat Rev Cancer 18:500–510. https://doi.org/10.1038/s41568-018-0016-5
Do HM, Spear LG, Nikpanah M et al (2020) Augmented radiologist workflow improves report value and saves time: a potential model for implementation of artificial intelligence. Acad Radiol 27:96–105. https://doi.org/10.1016/j.acra.2019.09.014
Lou R, Lalevic D, Chambers C, Zafar HM, Cook TS (2020) Automated detection of radiology reports that require follow-up imaging using natural language processing feature engineering and machine learning classification. J Digit Imaging 33:131–136. https://doi.org/10.1007/s10278-019-00271-7
Mokrane F-Z, Lu L, Vavasseur A et al (2020) Radiomics machine-learning signature for diagnosis of hepatocellular carcinoma in cirrhotic patients with indeterminate liver nodules. Eur Radiol 30:558–570. https://doi.org/10.1007/s00330-019-06347-w
Schaffter T, Buist DSM, Lee CI et al (2020) Evaluation of combined artificial intelligence and radiologist assessment to interpret screening mammograms. JAMA Netw Open 3:e200265. https://doi.org/10.1001/jamanetworkopen.2020.0265
Chauvie S, De Maggi A, Baralis I et al (2020) Artificial intelligence and radiomics enhance the positive predictive value of digital chest tomosynthesis for lung cancer detection within SOS clinical trial. Eur Radiol. https://doi.org/10.1007/s00330-020-06783-z
Fischer AM, Varga-Szemes A, Martin SS et al (2020) Artificial intelligence-based fully automated per lobe segmentation and emphysema-quantification based on chest computed tomography compared with global initiative for chronic obstructive lung disease severity of smokers. J Thorac Imaging. https://doi.org/10.1097/RTI.0000000000000500
Kocak B, Durmaz ES, Ates E, Kaya OK, Kilickesmez O (2019) Unenhanced CT texture analysis of clear cell renal cell carcinomas: a machine learning-based study for predicting histopathologic nuclear grade. AJR Am J Roentgenol:W1–W8. https://doi.org/10.2214/AJR.18.20742
Kocak B, Durmaz ES, Ates E, Ulusan MB (2019) Radiogenomics in clear cell renal cell carcinoma: machine learning-based high-dimensional quantitative CT texture analysis in predicting PBRM1 mutation status. AJR Am J Roentgenol 212:W55–W63. https://doi.org/10.2214/AJR.18.20443
Kocak B, Durmaz ES, Ates E et al (2020) Radiogenomics of lower-grade gliomas: machine learning-based MRI texture analysis for predicting 1p/19q codeletion status. Eur Radiol 30:877–886. https://doi.org/10.1007/s00330-019-06492-2
Greffier J, Hamard A, Pereira F et al (2020) Image quality and dose reduction opportunity of deep learning image reconstruction algorithm for CT: a phantom study. Eur Radiol. https://doi.org/10.1007/s00330-020-06724-w
Parmar C, Barry JD, Hosny A, Quackenbush J, Aerts HJWL (2018) Data analysis strategies in medical imaging. Clin Cancer Res 24:3492–3499. https://doi.org/10.1158/1078-0432.CCR-18-0385
Thrall JH, Li X, Li Q et al (2018) Artificial intelligence and machine learning in radiology: opportunities, challenges, pitfalls, and criteria for success. J Am Coll Radiol 15:504–508. https://doi.org/10.1016/j.jacr.2017.12.026
Leek JT, Scharpf RB, Bravo HC et al (2010) Tackling the widespread and critical impact of batch effects in high-throughput data. Nat Rev Genet 11:733–739. https://doi.org/10.1038/nrg2825
Johnson WE, Li C, Rabinovic A (2007) Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8:118–127. https://doi.org/10.1093/biostatistics/kxj037
Quackenbush J (2002) Microarray data normalization and transformation. Nat Genet 32(Suppl):496–501. https://doi.org/10.1038/ng1032
Lee ML, Kuo FC, Whitmore GA, Sklar J (2000) Importance of replication in microarray gene expression studies: statistical methods and evidence from repetitive cDNA hybridizations. Proc Natl Acad Sci U S A 97:9834–9839. https://doi.org/10.1073/pnas.97.18.9834
Yu K-H, Beam AL, Kohane IS (2018) Artificial intelligence in healthcare. Nat Biomed Eng 2:719–731. https://doi.org/10.1038/s41551-018-0305-z
Koçak B, Durmaz EŞ, Ateş E, Kılıçkesmez Ö (2019) Radiomics with artificial intelligence: a practical guide for beginners. Diagn Interv Radiol 25:485–495. https://doi.org/10.5152/dir.2019.19321
Hernández B, Parnell A, Pennington SR (2014) Why have so few proteomic biomarkers “survived” validation? (sample size and independent validation considerations). Proteomics 14:1587–1592. https://doi.org/10.1002/pmic.201300377
Way TW, Sahiner B, Hadjiiski LM, Chan H-P (2010) Effect of finite sample size on feature selection and classification: a simulation study. Med Phys 37:907–920. https://doi.org/10.1118/1.3284974
Chan HP, Sahiner B, Wagner RF, Petrick N (1999) Classifier design for computer-aided diagnosis: effects of finite sample size on the mean performance of classical and neural network classifiers. Med Phys 26:2654–2668. https://doi.org/10.1118/1.598805
Sollini M, Cozzi L, Antunovic L, Chiti A, Kirienko M (2017) PET Radiomics in NSCLC: state of the art and a proposal for harmonization of methodology. Sci Rep 7:358. https://doi.org/10.1038/s41598-017-00426-y
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278:563–577. https://doi.org/10.1148/radiol.2015151169
Perlich C (2010) Learning curves in machine learning. In: Sammut C, Webb GI (eds) Encyclopedia of machine learning. Springer US, Boston, MA, pp 577–580
Krause J, Gulshan V, Rahimy E et al (2018) Grader variability and the importance of reference standards for evaluating machine learning models for diabetic retinopathy. Ophthalmology 125:1264–1272. https://doi.org/10.1016/j.ophtha.2018.01.034
Zwanenburg A (2019) Radiomics in nuclear medicine: robustness, reproducibility, standardization, and how to avoid data analysis traps and replication crisis. Eur J Nucl Med Mol Imaging 46:2638–2655. https://doi.org/10.1007/s00259-019-04391-8
Mwangi B, Tian TS, Soares JC (2014) A review of feature reduction techniques in neuroimaging. Neuroinformatics 12:229–244. https://doi.org/10.1007/s12021-013-9204-3
Zwanenburg A, Löck S (2018) Why validation of prognostic models matters? Radiother Oncol 127:370–373. https://doi.org/10.1016/j.radonc.2018.03.004
Huber W, von Heydebreck A, Sültmann H, Poustka A, Vingron M (2002) Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinformatics 18(Suppl 1):S96–S104. https://doi.org/10.1093/bioinformatics/18.suppl_1.s96
Ioffe S, Szegedy C (2015) Batch normalization: accelerating deep network training by reducing internal covariate shift. ArXiv150203167 Cs
Ba JL, Kiros JR, Hinton GE (2016) Layer normalization. ArXiv160706450 Cs stat
Kocak B, Ates E, Durmaz ES, Ulusan MB, Kilickesmez O (2019) Influence of segmentation margin on machine learning-based high-dimensional quantitative CT texture analysis: a reproducibility study on renal clear cell carcinomas. Eur Radiol 29:4765–4775. https://doi.org/10.1007/s00330-019-6003-8
Kocak B, Durmaz ES, Kaya OK, Ates E, Kilickesmez O (2019) Reliability of single-slice-based 2D CT texture analysis of renal masses: influence of intra- and interobserver manual segmentation variability on radiomic feature reproducibility. AJR Am J Roentgenol 213:377–383. https://doi.org/10.2214/AJR.19.21212
Koçak B (2019) Reliability of 2D magnetic resonance imaging texture analysis in cerebral gliomas: influence of slice selection bias on reproducibility of radiomic features. Istanb Med J 20:413–417
Um H, Tixier F, Bermudez D, Deasy JO, Young RJ, Veeraraghavan H (2019) Impact of image preprocessing on the scanner dependence of multi-parametric MRI radiomic features and covariate shift in multi-institutional glioblastoma datasets. Phys Med Biol 64:165011. https://doi.org/10.1088/1361-6560/ab2f44
Berenguer R, Pastor-Juan MDR, Canales-Vázquez J et al (2018) Radiomics of CT features may be nonreproducible and redundant: influence of CT acquisition parameters. Radiology 288:407–415. https://doi.org/10.1148/radiol.2018172361
Zhovannik I, Bussink J, Traverso A et al (2019) Learning from scanners: bias reduction and feature correction in radiomics. Clin Transl Radiat Oncol 19:33–38. https://doi.org/10.1016/j.ctro.2019.07.003
Bologna M, Corino V, Mainardi L (2019) Technical note: virtual phantom analyses for preprocessing evaluation and detection of a robust feature set for MRI-radiomics of the brain. Med Phys 46:5116–5123. https://doi.org/10.1002/mp.13834
He H, Garcia EA (2009) Learning from imbalanced data. IEEE Trans Knowl Data Eng 21:1263–1284
Meyer M, Ronald J, Vernuccio F et al (2019) Reproducibility of CT radiomic features within the same patient: influence of radiation dose and CT reconstruction settings. Radiology 293:583–591. https://doi.org/10.1148/radiol.2019190928
Qiu Q, Duan J, Duan Z et al (2019) Reproducibility and non-redundancy of radiomic features extracted from arterial phase CT scans in hepatocellular carcinoma patients: impact of tumor segmentation variability. Quant Imaging Med Surg 9:453–464. https://doi.org/10.21037/qims.2019.03.02
Owens CA, Peterson CB, Tang C et al (2018) Lung tumor segmentation methods: impact on the uncertainty of radiomics features for non-small cell lung cancer. PLoS One 13:e0205003. https://doi.org/10.1371/journal.pone.0205003
Estrada S, Lu R, Conjeti S et al (2020) FatSegNet: a fully automated deep learning pipeline for adipose tissue segmentation on abdominal Dixon MRI. Magn Reson Med 83:1471–1483. https://doi.org/10.1002/mrm.28022
Lambin P, Leijenaar RTH, Deist TM et al (2017) Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol 14:749–762. https://doi.org/10.1038/nrclinonc.2017.141
Leger S, Zwanenburg A, Pilz K et al (2017) A comparative study of machine learning methods for time-to-event survival data for radiomics risk modelling. Sci Rep 7:13206. https://doi.org/10.1038/s41598-017-13448-3
Vallières M, Kay-Rivest E, Perrin LJ et al (2017) Radiomics strategies for risk assessment of tumour failure in head-and-neck cancer. Sci Rep 7:10117. https://doi.org/10.1038/s41598-017-10371-5
Sun R, Limkin EJ, Vakalopoulou M et al (2018) A radiomics approach to assess tumour-infiltrating CD8 cells and response to anti-PD-1 or anti-PD-L1 immunotherapy: an imaging biomarker, retrospective multicohort study. Lancet Oncol 19:1180–1191. https://doi.org/10.1016/S1470-2045(18)30413-3
Parmar C, Grossmann P, Bussink J, Lambin P, Aerts HJWL (2015) Machine learning methods for quantitative radiomic biomarkers. Sci Rep 5:13087. https://doi.org/10.1038/srep13087
Guyon I, Elisseeff A (2003) An introduction to variable and feature selection. J Mach Learn Res 3:1157–1182
Brown G, Pocock A, Zhao M-J, Luján M (2012) Conditional likelihood maximisation: a unifying framework for information theoretic feature selection. J Mach Learn Res 13:27–66
Kalousis A, Prados J, Hilario M (2006) Stability of feature selection algorithms: a study on high-dimensional spaces. Knowl Inf Syst 12:95–116. https://doi.org/10.1007/s10115-006-0040-8
Haury A-C, Gestraud P, Vert J-P (2011) The influence of feature selection methods on accuracy, stability and interpretability of molecular signatures. PLoS One 6:e28210. https://doi.org/10.1371/journal.pone.0028210
Mazurowski MA, Habas PA, Zurada JM, Lo JY, Baker JA, Tourassi GD (2008) Training neural network classifiers for medical decision making: the effects of imbalanced datasets on classification performance. Neural Netw Off J Int Neural Netw Soc 21:427–436. https://doi.org/10.1016/j.neunet.2007.12.031
van Smeden M, Moons KG, de Groot JA et al (2019) Sample size for binary logistic prediction models: beyond events per variable criteria. Stat Methods Med Res 28:2455–2474. https://doi.org/10.1177/0962280218784726
Olson RS, La Cava W, Mustahsan Z, Varik A, Moore JH (2018) Data-driven advice for applying machine learning to bioinformatics problems. Pac Symp Biocomput 23:192–203
Dankers FJWM, Traverso A, Wee L, van Kuijk SMJ (2019) Prediction modeling methodology. In: Kubben P, Dumontier M, Dekker A (eds) Fundamentals of clinical data science. Springer, Cham
Vickers AJ, Elkin EB (2006) Decision curve analysis: a novel method for evaluating prediction models. Med Decis Making 26:565–574. https://doi.org/10.1177/0272989X06295361
Vickers AJ, van Calster B, Steyerberg EW (2019) A simple, step-by-step guide to interpreting decision curve analysis. Diagn Progn Res 3:18. https://doi.org/10.1186/s41512-019-0064-7
de Sitter A, Visser M, Brouwer I et al (2020) Facing privacy in neuroimaging: removing facial features degrades performance of image analysis methods. Eur Radiol 30:1062–1074. https://doi.org/10.1007/s00330-019-06459-3
Mongan J, Moy L, Kahn CE (2020) Checklist for Artificial Intelligence in Medical Imaging (CLAIM): a guide for authors and reviewers. Radiology Artificial Intelligence 2:e200029. https://doi.org/10.1148/ryai.2020200029
Luo W, Phung D, Tran T et al (2016) Guidelines for developing and reporting machine learning predictive models in biomedical research: a multidisciplinary view. J Med Internet Res 18:e323. https://doi.org/10.2196/jmir.5870
Collins GS, Reitsma JB, Altman DG, Moons KGM (2015) Transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD): the TRIPOD statement. Ann Intern Med 162:55–63. https://doi.org/10.7326/M14-0697
Collins GS, Moons KGM (2019) Reporting of artificial intelligence prediction models. Lancet 393:1577–1579. https://doi.org/10.1016/S0140-6736(19)30037-6
Funding
The authors state that this work has not received any funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Burak Kocak, MD.
Conflict of interest
The authors of this manuscript declare no relationships with any companies whose products or services may be related to the subject matter of the article.
Statistics and biometry
No statistical methods were necessary for this paper.
Informed consent
Not required.
Ethical approval
Not required.
Methodology
• Review Article
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kocak, B., Kus, E.A. & Kilickesmez, O. How to read and review papers on machine learning and artificial intelligence in radiology: a survival guide to key methodological concepts. Eur Radiol 31, 1819–1830 (2021). https://doi.org/10.1007/s00330-020-07324-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-020-07324-4