Abstract
Arctic rainbow smelt Osmerus dentex is one of the most abundant fish species widely distributed in the NW Pacific and Arctic seas. Studying this species can contribute to understanding the microevolutionary processes of the Arctic fauna in unstable climatic conditions and new habitats. However, little is known about the genetic structure and the mechanisms underlying its population dynamics. Ten microsatellite loci were used to genotype 822 individuals in 20 locations throughout a large part of the species’ range in the Sea of Japan, Sea of Okhotsk, Bering, Kara, and White seas (across more than 11,000 km). O. dentex displayed substantial levels of genetic differentiation (overall FST = 0.067, RST = 0.091) and STRUCTURE identified eight distinct clusters. The significant regional variation was demonstrated in both genetic diversity and population structure, most notably between Arctic and Pacific basins. On a regional scale, contrasting patterns were revealed with limited gene flow between populations of the Arctic region (FST = 0.117) and high gene flow between the Bering Sea (FST = 0.017) or Sea of Okhotsk (FST = 0.031), which were probably associated with differences in larval distribution from the spawning grounds due to different oceanographic conditions of the studied areas. These results demonstrate the effect of local contemporary forces in shaping different patterns of genetic connectivity and diversity within O. dentex.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pacific or Arctic rainbow smelt Osmerus dentex Steindachner and Kner, 1870 (Nellbring 1989) is a typical anadromous species widely distributed in estuaries and coastal waters of European and Siberian shores of the Arctic Ocean from the White Sea to Chukotka in eastern Siberia and from the Bering Sea to Korea and Northern Japan in the Pacific Ocean (Klyukanov 1975; Chereshnev 1998; Kottelat and Freyhof 2007). Spawning of smelt usually takes place in early spring, the upstream run is generally short, a maximum of a few km, but on some large rivers it can rise for very considerable distances. Smelt mature at 2–4 years of age, and the eggs attach to gravel; after hatching, larvae are immediately carried downstream and develop in estuaries. Adults return to estuaries after spawning and may move into shallow waters; however, they do not make long-distance migrations (Berg 1948; Nellbring 1989; Vasilets 2000; Chereshnev et al. 2002; Gritsenko 2002).
Arctic rainbow smelt can potentially be structured on a very small spatial scale up to individual river systems. They are philopatric, and their spatial migrations are limited. In different areas of the Sea of Okhotsk, Sakhalin, Kamchatka, Chukotka, and the White and Barents seas, O. dentex is distinguished by a certain number of ecological groups associated mainly with the timing and duration of spawning, the length of spawning and wintering migrations, and some morphometric traits (Kirpichnikov 1935; Podushko 1970; Vasilets et al. 2000; Chereshnev et al. 2002; Gritsenko 2002; Bugaev 2007). The investigation of populations within the White and Barents seas using allozymes (Sendek et al. 2005) and microsatellites (Semenova et al. 2019a) was fragmentary; however, some regional differences at the level of large bays have been revealed. The genetic difference has been observed using microsatellites between populations inhabiting rivers on the East and West sides of the Kamchatka Peninsula (Semenova et al. 2019b). Simultaneously, mtDNA-based studies have shown weak population structure of smelt in most of its range despite the high molecular diversity (Skurikhina et al. 2015).
Examination of genetic differentiation within related species of family Osmeridae, the rainbow smelt (Osmerus mordax) and the eulachon (Thaleichthys pacificus), which possess a similar life history, suggested that divergence may be very low across a wide geographic scale (McLean and Taylor 2001; Beacham et al. 2005; Bradbury et al. 2006; Kovach et al. 2013). At the same time, in some parts of the species’ range, gene flow can be significantly limited (Bradbury et al. 2008a, b).
Arctic rainbow smelt is inferred to have expanded its range after the end of the latest Wisconsin glaciations from the main refuge in the NW Pacific (the Sea of Japan and the southern Sea of Okhotsk). In addition, it the existence of a small refuge in the White Sea (Skurikhina et al. 2018) has been proposed. Historical factors are thought to have played a significant role in the patterns of genetic variation in Arctic rainbow smelt. Meanwhile, contemporary mechanisms affecting the genetic structure of smelt remain largely unknown.
O. dentex is one of the most abundant fish species of the Osmeridae family, and it is important for commercial and sport fisheries in the Russian Far East waters (Andriashev and Chernova 1994; Bugaev et al. 2014). The economic importance of smelt clearly emphasises the need for a better understanding of the population structure of the species and the mechanisms underlying its population dynamics. Microsatellites can help identify cryptic population structure and distinguish historical and contemporary factors that influence the demographic histories of populations. Thus, understanding population structure based on neutral genetic variation has become a necessary step in the development of the conservation programs for many commercial exploited fish (Allendorf et al. 1987).
While earlier studies have identified the possibility of an additional small refuge in the White Sea, we reasoned that an investigation with nuclear markers might offer additional insights into this issue. It has been shown in several studies that the analyses of microsatellite data can be useful for resolving phylogenies at substantial divergence scales, including for estimating the number of post-glacial refugia that contributed to the re-colonisation of Northern Europe (Koskinen et al. 2002; Tonteri et al. 2005; Peterson and Ardren 2009).
The purpose of this study was to investigate the population genetic structure of Arctic rainbow smelt across its range in the Arctic waters, eastern coast of Bering Sea, Sea of Okhotsk, and Sea of Japan using ten microsatellite loci. Unfortunately, in this paper and the literature, there are no data on smelt from the Arctic coast of Siberia due to the inaccessibility of this region. This greatly limits our ability to discuss smelt relationships throughout the Arctic coastline. Our specific goals were to (i) characterise the overall patterns of genetic diversity as well as regional and range-wide genetic structure of O. dentex; (ii) assess historical and contemporary drivers of population structure; and (iii) discuss conservation and management measures for O. dentex populations.
Materials and methods
Sample collection, microsatellite genotyping and descriptive genetic data
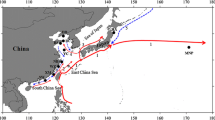
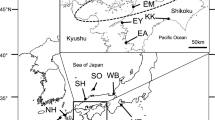
In this study, 20 locations of O. dentex from the White, Kara, Bering, Okhotsk, and Japan seas were analysed. These samples were collected in 2006–2019 at different periods of fish life cycles: in the rivers during spawning; in the mouth of the rivers and the estuaries during wintering, pre-spawning, or post-spawning migrations. We assume that all samples represent the populations of their specific rivers, since most of smelt in the range tend to adhere to local habitats for feeding or wintering (Table 1; Fig. 1).
DNA was extracted from the fin by the following standard methods using the Diatom DNA Prep kits (IsoGene, Russia). Ten microsatellite loci isolated from Pacific smelt or relative species were tested (Kaukinen et al. 2004; Coulson et al. 2006; Fisch et al. 2009). We designed new primers for six loci using the Primer-BLAST (Ye et al. 2012) (Online Resource 1). DNA amplification and separation of the PCR products were carried out according to the procedure described in Semenova et al. (2019b).
The genotype data were analysed with MICROCHECKER 2.2.3 to check for null alleles and scoring errors (Van Oosterhout et al. 2004). The populations were tested for deviations from HW equilibrium and linkage disequilibrium using GENEPOP 3.4 (Raymond and Rousset 1995).
The GDA 1.0 software (Lewis and Zaykin 2001) was used for calculating allele frequencies, the expected and observed heterozygosities (HE, HO), allelic diversity (A), and the number of private alleles (APR). The allelic richness (AR), corrected for the minimum sample size, was calculated using FSTAT 2.9.3 (Goudet 2001). Estimates of genetic variability were calculated for all localities and clusters revealed by STRUCTURE (see below). The differences in the average number of alleles (AR) and heterozygosity (HO) between sites were tested using a randomisation test (1000 permutations) in FSTAT.
Bottleneck events
To detect indications for a recent bottleneck or a reduction in population size, we used BOTTLENECK 1.2.02 (Piry et al. 1999) with the Wilcoxon test, based on 100,000 iterations using a stepwise mutation model (SMM), an infinite allele model (IAM), and a two-phase mutation model (TPM) (set at 70% SMM, 30% IAM). For each population, only polymorphic loci in HWE were used. We also used the M-ratio method with the critical value of M = 0.68 following Garza and Williamson (2001).
Population structure and connectivity
Population differentiation was estimated per population pair and overall using FST (θ) and RST (Weir and Cockerham 1984; Slatkin 1995) in GENEPOP and GDA. Fisher’s exact test in GENEPOP was used to test for significant differences in allele frequencies between populations. We compared the levels of variation among locations based on allele identity (FST) and allele size (RST). A permutation test (1,000 permutations) used to determine whether stepwise-like mutations contributed to genetic differentiation (RST = FST) was carried out using SPAGEDi 1.2 (Hardy and Vekemans 2002; Hardy et al. 2003). Sequential Bonferroni adjustments were applied to correct for the effect of multiple tests. We conducted a Mantel test to evaluate isolation by distance (IBD) using the standardised genetic distance (FST/1 − FST) and the logarithm of the geographic distance among locations, with 10,000 permutations, in the IBD 1.52 program (Jensen et al. 2005).
Bayesian analysis was implemented in STRUCTURE 2.3.4 (Pritchard et al. 2000), assuming the admixture model and correlated allele frequencies between clusters. STRUCTURE was run using MCMC simulations of 9,00,000 iterations, with a burn-in period of 1,00,000. Runs were conducted for the number of putative populations (i.e. K), set at 1–15, with each having 20 iterations. The optimum K value was assessed by analysing the Ln P(K) distribution, and the Evanno method ΔK (Evanno et al. 2005), as well as the bar plots showing individual assignments. All parameters were received on the STRUCTURE HARVESTER webpage (Earl and vonHoldt 2012). To study the intricate population structure within clusters, we used the ‘hierarchical STRUCTURE analysis’ (Vähä et al. 2007). Firstly, we estimated the optimal K‐value at the uppermost hierarchical level of structure, delineating large‐scale differences in our data set. Then we examined whether substructure existed within each of the populations identified at the uppermost level by estimating K within each of those populations. We used the CLUMPAK webserver (Kopelman et al. 2015) to align and visualise bar plots at multiple K values.
We performed an analysis of molecular variance (AMOVA; Excoffier et al. 1992) in ARLEQUIN 3.5 (Excoffier and Lischer 2010) to estimate the amount of genetic variation present at the following hierarchical levels: among major sea basins (Japan, Okhotsk, Bering, Kara, and White seas), among sites within major sea basins, and within sites (analysis #1). We conducted AMOVA analysis #2 at the hierarchical levels based on eight clusters previously identified by STRUCTURE: among major clusters, among sites within these clusters, and within sites.
BARRIER 2.2 (Manni et al. 2004) was used to highlight the geographical areas with pronounced genetic discontinuity between the sites, i.e. barriers. Geographical coordinates were supplied for each site and connected by Delauney triangulation, such that each connection had an association. Within the analysis, barriers were set from 1 to 5. To test the significance of the detected barriers, 1000 resampled bootstrapped (population pairwise) DA genetic distance matrices (Nei’s chord distance) were calculated in MICROSATELLITE ANALYSER (MSA) software (Dieringer and Schlötterer 2003).
Results
Data quality check, Hardy–Weinberg equilibrium, and linkage disequilibrium
A table of allele frequencies by populations is presented in Online Resource 2. MICROCHECKER suggested a possible null allele at the MGPL-Omo9 locus in six locations: AMU, KRU, BEL, BOL, NER14, and DVI. No other loci showed signs of possible null alleles. There was no evidence for scoring error due to shuttering or large allele drop-out for any microsatellite loci. Locus MGPL-Omo9 was discarded from further analyses of population structure.
Of the 200 single-locus tests for conformation to HWE, five had significant departure from the expectations following Bonferroni corrections (Online Resource 3). Four of them for the MGPL-Omo9 locus could be explained by the presence of null alleles. When the results of all loci were combined for each population, only BEL departed from HWE after correction (p = 0.000). Evidence for linkage disequilibrium was detected between MGPL-Omo3 and MGPL-Omo16 with all populations pooled (p < 0.0001). Therefore, MGPL-Omo16 was removed from the dataset.
Microsatellite diversity and tests for bottleneck
The mean number of alleles per location ranged from 3.1 to 6.7. The levels of observed heterozygosity (HO) and allelic richness (AR) per location ranged from 0.365 to 0.567 and from 2.6 to 4.9, respectively (Table 2; Online Resource 3). No private alleles (APR) with p > 0.05 were found in any of the localities. The lowest genetic diversity was shown in the smelt of the inner part of the White Sea (Kandalaksha and Dvina bays). HO and AR were declines in the direction from the Sea of Japan and Sea of Okhotsk to the Bering Sea, then to the Kara Sea and the White Sea. Measures of genetic diversity (AR, HO) significantly differed among sites (AR, p = 0.004; HO, p = 0.032).
There were no indications of bottlenecks in any of the populations for the IAM, TPM, or SMM models (0.461 < p < 1.000). Moreover, in all populations excluding BEL, we identified a deviation from mutational drift equilibrium and a significant heterozygosity deficit (Wilcoxon test, p < 0.0003–0.019). No evidence of bottleneck effects were detected by the M-ratio tests, as estimated M values ranging in locations from 0.709 to 0.888 were above the critical M = 0.68.
Population genetic structure
The global FST = 0.067 (95% CI: 0.038–0.099) revealed a significant level of genetic structuring among all locations. Individual locus FST varied considerably, from 0.021 at MGPL-Omo2 to 0.116 at MGPL-Omo4, all associated exact tests of global differentiation were significant (p < 0.05). The overall RST = 0.091 (95% CI 0.078–0.103) was also significant, as were all locus-specific tests (Online Resource 4).
A pairwise population analysis of O. dentex detected a significant difference in allele frequencies for 175 of the 190 possible tests (Exact tests, p < 0.05), with multilocus FST ranging from 0.0004 to 0.1781 (Table 3). Pairwise FST for collections in consecutive years (AMH16 and AMH18; NER14 and NER16) were not significant, so yearly collections from the same locations were pooled in further analyses. The genetic differentiation was also not significant between locations of the Tumnin and Amur rivers, between sites of Kandalaksha Bay from the White Sea, and between the majority of the sites from the western coast of Kamchatka. After correcting for multiple testing, nine more tests were found to be non-significant, including all sites of Western Kamchatka except BEL and smelt from Lake Nerpich’e vs the Uka River of Eastern Kamchatka. The highest levels of genetic differentiation were revealed in comparisons of smelt from the Arctic region (the White and Kara seas) vs the Pacific Basin (the Sea of Japan, Sea of Okhotsk, and Bering Sea), with pairwise FST that ranged from 0.056 to 0.199. However, among Arctic populations, pairwise FST were quite high (0.055–0.170), except the FST between KAN and KAB sites in the Kandalaksha Bay in the White Sea.
Pairwise RST followed a similar pattern and were significant in 135 out of 190 comparisons before Bonferroni adjustments and in 128 cases after it, ranging from 0.0020 to 0.4130 (Table 3). The greatest differences in FST and RST were shown between the smelt of the White Sea and the smelt of Pacific Basin.
The allele size permutation test found significant differences between all FST and RST estimates (RST > FST, p = 0.0013) and supported the existence of a population structure within O. dentex formed not only by drift and gene migration but also by the contribution of the SMM. Sixty-one out of 190 pairwise comparisons between locations resulted in a significant phylogeographic signal (RST significantly larger than FST; Table 3). These significant comparisons were generally population pairs from Kandalaksha Bay, the White Sea (KAR, KAB), and the remaining locations, except DVI vs KAB. Also, about 50% of the pairwise tests between populations from the Eastern Kamchatka (the Bering Sea) vs the Western Kamchatka (the Sea of Okhotsk) were considerable. In addition, some comparisons between the Sea of Japan and the southwestern part of the Sea of Okhotsk vs the populations of the Western Kamchatka were substantial. When testing the Arctic region or the Pacific Basin separately, there were no differences between the FST and RST indices.
Bayesian analysis in STRUCTURE of the first hierarchical level detected by ΔK test indicated two clusters corresponding with: (1) all location from the Pacific Basin and the Kara Sea, and (2) smelt of the White Sea. The likelihood of the data Ln P(K) was lowest at K = 13 and indicated the possibility of a deeper structuring of the smelt (Online Resource 5).At the second round of hierarchical analysis the White Sea initial cluster was separated into the three groups: the Mezen Bay, Dvina Bay, and Kandalaksha Bay. Also, there were two clusters detected within ‘the Pacific Basin’ cluster: (1) the Sea of Japan and Sea of Okhotsk, and (2) the Bering Sea and the Kara Sea. Most individual genotypes of AVA (the eastern coast of Kamchatka, the Pacific Ocean) indicated admixture between these clusters. However, > 50% genotypes referred to the cluster 1. A third round of analysis found further structuring within two groups, where the Sea of Japan-Sea of Okhotsk cluster was again divided into two clusters: (1) Tumnin and Amur rivers, and Sakhalin and Amakhton Bay; and (2) the western coast of Kamchatka and AVA. The Bering Sea and the Kara Sea were also separated into discrete clusters: (1) NER and UKA, and (2) KAR. Finally, the fourth round of STRUCTURE revealed TUM-AMU-SAH and AMH as a separated cluster, while no subdivision was indicated within Western Kamchatka-AVA cluster. The finest population structure detected by STRUCTURE therefore consisted of eight geographical groups (Fig. 2): (1–3) three groups among the White sea (the Kandalaksha, Dvina, and Mezen bays); (4) the Kara Sea; (5) the eastern coast of Kamchatka (the Bering Sea); (6) the western coast of Kamchatka (the Sea of Okhotsk) and a sample from Pacific Ocean (AVA); (7) Amakhton bay (the Sea of Okhotsk); and (8) Sakhalin, Amur River (the Sea of Okhotsk), and Tumnin River (the Sea of Japan).
The global AMOVA, analysis # 1, indicated that a significant majority of genetic variation was located within sites (FST: 92.2%; RST: 89.8%). Significant variation was also attributable to differences between sea basins and between sites within sea basins. Analysis #2 revealed a similar genetic variation within sites. Between eight STRUCTURE clusters were 6.3% genetic variation by FST and 9.1% by RST, whereas a small remainder of the variation was attributable to differences among sites within the clusters (Table 4).
BARRIER identified five discontinuities in gene flow across the species’ range supported with more 88% bootstrap level. The main barriers (1) delimited the Kandalaksha Bay populations (KAN, KAB) with the highest bootstrap support of 100%. Another strong barrier (2) (with up to 100% support) separated populations of smelt from the Kara and White seas from the remainder of the species’ range. Between the Kara Sea and the White Sea, the barrier (3) had 88% bootstrap support, and between Mezen Bay and Dvina Bay, the barrier (4) had 99% bootstrap support. TUM and AMU populations showed genetic isolation, with a 100% supported barrier (5) (Online Resource 6).
There was significant support for an overall isolation by distance both in FST and in RST. Significant IBD was also evidenced among the Arctic populations, as well as among the Pacific Basin populations. On a smaller spatial scale, within the western coast of Kamchatka, the eastern coast of Kamchatka or the Sea of Japan and Southwestern Sea of Okhotsk, IBD was observed only among the Eastern Kamchatka locations (Online Resource 7).
Discussion
In this study, we revealed considerable genetic diversity and population differentiation in Arctic smelt, following a hierarchical geographic pattern. Genetic differentiation was most pronounced between the Arctic region, and, especially, the population from Kandalaksha Bay (the White Sea), vs the smelt inhabiting basins of the Pacific Ocean. Lower levels of genetic variation within populations from the Kara and White seas were also revealed. Genetic differentiation among Arctic smelt was considerably stronger than among Pacific populations. The regional differences in gene flow and connectivity within smelt populations of the Arctic and Pacific basins were mainly associated with differences in larval distribution from the spawning grounds due to different oceanographic conditions of the studied areas. These results demonstrate the effect of local contemporary forces in shaping different patterns of genetic connectivity and diversity within O. dentex.
Genetic variation within sampling locations
The genetic variation in the Kandalaksha and Dvina bays (the White Sea) were among the lowest we observed, whereas estimates in the Sea of Okhotsk were almost two times greater. A decrease in the molecular diversity in the Arctic region has also been reported in mtDNA-based studies suggesting that the ancestral population had undergone an effective size reduction or a founder effect during re-colonisation (Skurikhina et al. 2015, 2018). These patterns are largely consistent with those of the Pacific herring Clupea pallasii and species of the family Osmeridae, Hypomesus olidus, which have similar geographic distributions in the Pacific Ocean and the Arctic waters (Semenova et al. 2015; Skurikhina et al. 2019).
The history of glaciations in the Northern hemisphere, environmental conditions, and the species life history appear to have an important influence on the level of genetic variation within O. dentex. It has typically shown increased genetic variation within populations located in or near areas of former glacial refuge compared to populations from glaciated areas presumably founded by small numbers of individuals (Bernatchez and Wilson 1998; Costello et al. 2003; Stamford and Taylor 2004). The movement O. dentex from the main refuge after the retreat of the glaciers was in the direction from the Sea of Okhotsk to the Bering Sea, and then, to the Kara Sea and occurred through the Bering Strait about < 15 Kya (Taylor and Dodson 1994; Gladenkov and Gladenkov 2004; Dodson et al. 2007; Skurikhina et al. 2018). Decreases in the effective number of smelt populations may have occurred and in modern time, due to the species’ habitation in the harsh climate of the Arctic, subject to periodic fluctuations.
Comparisons of F ST and R ST estimates
The results showed that some populations of O. dentex were isolated from each other long enough for mutations to cause genetic differences between them (Hardy et al. 2003). It is especially true for the sites from Kandalaksha Bay, and there is a possibility this is caused by a long time of divergence. One likely reason for this may be post-glacial colonisation by ancestral smelt of the White Sea from more than one glacial refuge. The existence of small refuges in the White Sea, in the Late Pleistocene, was previously assumed by Skurikhina et al. (2018). Colonisation from two refuges has also been detected for Atlantic salmon in the White Sea (Tonteri et al. 2005). Moreover, it has been recently reported that a pond smelt, H. olidus inhabiting the Barents and Kara seas, might have survived in glacial refuge of the Polar Ural region (Skurikhina et al. 2019).
However, other locations from the Arctic basin demonstrated from one (KAR) to three (MEZ) pairwise RST larger than FST compared to the Pacific sites. Also, some locations from the Pacific Basin have significant permutation tests compared to other Pacific populations (Table 3). Estoup and Angers (1998) argued that mutations are expected to have a substantial effect on population divergence only after 2000 generations. Generation lengths for smelt are usually about 2–4 years (Chereshnev et al. 2002; Bugaev 2007), which can be transformed in no less than 4000–8000 years of interspecies divergence. Thus, it can be assumed that the duration of the phylogenetic history of smelt already exceeds the time required to the mutation process to become significant in genetic differentiation and, therefore, the RST larger than FST does not necessarily indicate the origin of populations from different refuges.
Genetic population structure of O. dentex across the entire range
There were signals of IBD observed for the entire geographical range, as within the Arctic and Pacific basins separately. However, with these analyses, it is difficult to make an inference regarding migration-drift equilibrium, because these results may be due to the restricted gene flow (barrier effect), which contribute a strong IBD pattern (Hutchinson and Templeton 1999; Koizumi et al. 2006). At the small spatial scale, there was no correlation between genetic and geographical distances, excluding Eastern Kamchatka. However, these analyses involved too few populations (from three to seven), so the power of detecting IBD signal may be limited (Jenkins et al. 2010). Nevertheless, it can be assumed that geography (or distance) is one of the main factors determining the population structure of O. dentex throughout its large-scale range.
Genetic differentiation was particularly pronounced between populations of the Arctic region and other populations located in the Pacific (Table 3). At the same time, genetic differentiation among the Arctic populations was considerably stronger than among the populations from the Pacific Basin. So, the overall FST estimate of the Arctic region (FST = 0.117, RST = 0.144) was significantly (p < 0.001) greater than for the Sea of Japan/Okhotsk (FST = 0.031, RST = 0.031) or the Bering Sea (FST = 0.017, RST = 0.002).
The overall level of genetic differentiation we observed was similar to that found in related species with analogous life histories, namely O. mordax, across the broad-scale including Newfoundland and mainland Canada (Bradbury et al. 2011) and corresponded to the differentiation indices for groups with a moderate genetic structure (Zhivotovsky 1991). Moreover, we demonstrated the various patterns of genetic differentiation within the Arctic and the Pacific geographical areas. A similar spatial genetic structure was also found in O. mordax. So, the microsatellite divergence within the mainland Canada and the USA was an order of magnitude lower than in Newfoundland due to the re-colonisation of the island by two glacial races (Bradbury et al. 2006, 2011; Kovach et al. 2013). Within other anadromous osmerid species the level of geographic differentiation was usually low (McLean and Taylor 2001; Beacham et al. 2005).
The differentiation of smelt in the Pacific Basin
The microsatellites showed significant differences on a large geographic scale between groups from the western coast of Kamchatka (north-eastern Sea of Okhotsk), and the northern (AMH) and southwestern populations that were consistent with the division based on mtDNA (Skurikhina et al. 2015, 2018). The isolation of the southwestern Sea of Okhotsk cluster is also supported by the results of the BARRIER analysis (Online Resource 6). Interestingly, the most considerable mtDNA separation was observed for the Sea of Japan, whereas in our study, the microsatellite differentiation of smelt from the Sea of Japan versus the Sea of Okhotsk was low. According to mtDNA data, the boundary between the diverged populations of the Sea of Japan and the Sea of Okhotsk probably passes in the Nevelskoy Strait (the northern part of Tatar Strait) connecting these seas. The smelt collection areas from the Sea of Japan analysed by Skurikhina et al. (2018) are located almost 400 km to the south of the site (TUM) from our study, which belongs to the Tatar Strait. It can be assumed that the biogeographic border between the seas is located south of the Tatar Strait; however, a more extensive sampling program is needed to clarify the genetic structure and geographical boundaries of O. dentex distribution in the Sea of Japan.
The locations collected along the western coast of Kamchatka separated by more than 500 km demonstrated a low level of genetic differentiation and did not follow an IBD pattern. Only the Belogolovaya River (BEL) had significant pairwise FST and RST with other Western Kamchatka sites and was differentiated from even its nearest neighbours. According to fisheries statistics and certain morphometric traits of smelt in Western Kamchatka two geographically distinct population groups were defined: the ‘northern’ group has the main breeding centre in the rivers of the Northwestern Kamchatka [Kovran (KOV), Khairyuzova (HAI), Belogolovaya (BEL), and Moroshechnaya], and the ‘southern’ group has the main breeding centre in the Bolshaya (BOL) and Kol’ (KOL) rivers in the Southwestern Kamchatka (Vasilets et al. 2000; Bugaev et al. 2014). However, the assumption about the independence of smelt groups in the northern and southern areas of the Western Kamchatka was not supported by our data. Moreover, the genetic differentiation between sites in SW and NW Kamchatka was lower in certain cases than differentiation within the locations from the NW coast.
It has been shown for O. mordax that gene flow was limited beyond to approximately 200 km (Kovach et al. 2013; Bradbury et al. 2008a). Fish species of the genus Osmerus are characterised by short upstream migrations and small pelagic marine larvae that exit the river within hours of hatching (Berg 1948; Nellbring 1989); however, even among the osmerids, the level of gene flow observed in O. dentex along the western coast of Kamchatka is high. This indicates that extensive mixing occurs either in the feeding grounds or during larval drift from the spawning site. It is assumed that adult smelt, both O. mordax and O. dentex, do not make long feeding migrations and population structure is largely determined by the distribution of the early life studies (Baby et al. 1991; Bernatchez and Martin, 1996; Bradbury et al. 2006; Kovach et al. 2013). The number of genetically distinct populations results from the number of distinct larvae retention zones (Baby et al. 1991), which may be determined by geographical, physical, or oceanographic factors. So the genetic similarity of smelt from the Western Kamchatka may be explained by hydrographic factors influenced on larvae dispersal. There is a main West Kamchatka Current (WKC) along the western coast of Kamchatka. The scheme of water circulation in the Sea of Okhotsk is subject to inter-annual dynamics, and in addition to the WKC, in some years an intense shelf current in the South direction is formed (Compensation Current) (Chernyavsky 1981; Khen et al. 2019). The dynamics of currents determine the spatial distribution of fish larvae, so it has been shown that capelin Mallotus villosus and pollock Theragra halcogramma larvae can be carried far from the coast and widely distributed along the coast of Kamchatka in years of intense WKC action (Varkentin et al. 2001; Naumova and Kolomeitsev 2015). The high similarity of smelt between the sites along the Western Kamchatka is likely due to the intensive circulation patterns.
The differentiation of smelt in the Arctic Region
The Arctic rainbow smelt are characterised by high morpho-ecological variability: several local shoals or even geographic races of smelt are distinguished based on growth and maturation rates, meristic and morphometric indices, and habitats: Kandalaksha, Onega, Dvina, and Czech groups (Kirpichnikov 1935; Altukhov and Yerastova 1974). An early allozymes-based study described genetic isolation for the smelt from Onega Bay of the White Sea, on the one hand, and from Dvina Bay and the South-Eastern Barents Sea, on the other hand (Sendek et al. 2005).
The pairwise estimates among the five locations from the Arctic region using microsatellites were high, and significant allele frequency differences were observed among all sites excluding KAN vs KAB. These data suggest that each location from the different bays of the White and Kara seas represents a genetically distinct population and that the Kandalaksha Bay contains a single population. The main discontinuities in gene flow defined by BARRIER between each of the bays, well-defined STRUCTURE clustering and the high pairwise FST and RST estimates, indicate that little genetic exchange occurs among the population of Arctic basins.
The observed patterns of divergence may indicate that some groups of smelt have colonised Kandalaksha Bay from another ice-age refuge and may represent historically isolated populations preserved at glacial margins during the Pleistocene glaciations. Probably, during the colonisation, the ancient population could be distributed not across the entire White Sea, but along some rivers, such as the rivers in the Kandalaksha Bay. The hypothesis that relict smelt populations inhabit only the Kandalaksha Bay was partially confirmed by the microsatellite data. Thus, the other locations in the White Sea from the Dvina and Mezen bays revealed pairwise estimates of FST and RST with populations from the Bering Sea almost three times smaller than those between the Kandalaksha Bay population vs the Bering Sea.
However, historically isolated populations often contain private alleles not observed in other populations (Slatkin 1985; Allendorf and Luikart 2007). But no private alleles were observed in the Kandalaksha Bay population as we might expect if this was a relict population. Furthermore, the distribution of allele size of smelt in the Kandalaksha Bay was consistent with other sites and didn’t reveal evidence of a multimodal size distribution for any loci that could also indicate distinct historical origin (Lu et al. 2001; Saint-Laurent et al. 2003).
The combination of low genetic variability in populations of the White Sea and high levels of their interpopulation differentiation could also arise if the populations originated from the same ancestral refuge due to a founder effect and ancient bottleneck (McCauley 1993; Costello et al. 2003). In addition, the patterns of genetic differentiation may be influenced by contemporary factors, such as migration barriers. It has been shown, for example, that for European grayling (Thymallus thymallus) post-colonisation divergence of populations could also be effective in the absence of the homogenising effect of migration (Koskinen et al. 2002).
There is almost no possibility for modern mixing between the studied smelt populations in the White Sea. Thus, a very limited distribution of adults from spawning grounds has been shown (Kirpichnokov 1935; Berger 2007). It can also be assumed that there is a limited distribution of smelt larvae from estuarine areas of their native rivers. Unfortunately, large-scale studies of the distribution of smelt larvae in the White Sea have not been carried out; however, for White Sea herring, the restriction of larvae dispersion from the spawning ground was shown (Evseenko and Mishin 2011; Kobylyanskii et al. 2014). The retention of larvae in the shallow waters is due to the specific hydrological characteristics of the White Sea—the coastal structural frontal zones of increased gradients of the hydrophysical properties. These front zones are quasi-stationary, have different causes of occurrence and are usually expressed only in the spring and summer periods (Pantyulin 2003, 2012), the time of their existence just coincides with the time of stay of smelt larvae in sea waters (Parukhina 2009, 2016). The most obvious role of frontal zones is the barrier function, i.e. the retention of fish larvae within the water area bounded by the front (Sabates and Olivar 1996; Grioche and Koubbi 1997). The important role of frontal zones in forming aggregations of larvae of various species of pelagic fish and their development has been shown in several papers (e.g. Munk et al. 1999). That is largely consistent with the member-vagrant hypothesis (Iles and Sinclair 1982) that the number of genetically distinct populations is determined by the number of distinct larval retention areas. So, barriers to migration of smelt in the early stages of development appear to be an important factor influencing patterns of genetic variability among populations of O. dentex.
Stock management
Arctic rainbow smelt is a commercial fish species in the Pacific Ocean, so this genetic data will serve as a basis for making informed decisions on the management of this species in Russia. Our results determine that smelt shows high levels of population subdivision and genetic structure in the Arctic region. Such a high level of genetic subdivision suggests a high level of demographic independence. In addition, low rates of genetic variability of smelt populations in the White Sea may mean a small effective size of these populations and their particular vulnerability to external influences, including anthropogenic factors. Currently, the activity of the fishing industry in the White, Kara, and Laptev seas is reduced due to the socio-economic processes taking place in the country (Stasenkov et al. 2011; Ulchenko et al. 2016; Filatov et al. 2019). Nevertheless, the White Sea fishery should be carefully conducted to conserve the diversity of smelt in each particular bay, because genetically divergent populations are increasingly being recognised as appropriate units for conservation. The state of smelt stocks in different parts of the Pacific Basin is not currently a concern, but there is no doubt that the state of smelt populations needs to be assessed considering the geographic unit that may be genetically distinct (Bugaev et al. 2014; Burlak 2018).
References
Allendorf FW, Luikart G (2007) Conservation and the genetics of populations. Blackwell Publishing, Malden
Allendorf FW, Ryman N, Utter F (1987) Genetics and fishery management. In: Ryman N, Utter F (eds) Population genetics and fisheries management. University of Washington Press, Seattle and London, pp 1–19
Altukhov KA, Yerastova VM (1974) Biological characteristics of smelt, Osmerus eperlanus L. from Karelian White Sea coast. J Ichthyol 14:67–78
Andriashev AP, Chernova NV (1994) Annotated list of fishlike vertebrates and fish of the Arctic seas and adjacent waters. J Ichthyol 35:81–123
Baby MC, Bernatchez L, Dodson JJ (1991) Genetic structure and relationships among anadromous and landlocked populations of rainbow smelt, Osmerus mordax, Mitchill, as revealed by mtDNA restriction analysis. J Fish Biol 39:61–68
Beacham TD, Hay DE, Le KD (2005) Population structure and stock identification of eulachon (Thaleichthys pacificus), an anadromous smelt, in the Pacific Northwest. Mar Biotechnol 7:363–372
Berg LS (1948) Freshwater fishes of the USSR and adjacent countries. Akademiya Nauk SSSR, Moscow-Leningrad (in Russian)
Berger VY (2007) Production Potential of the White Sea. St. Petersburg Zoological Institute RAS, Saint-Petersburg (in Russian)
Bernatchez L, Martin S (1996) Mitochondrial DNA diversity in anadromous rainbow smelt, Osmerus mordax Mitchill: a genetic assessment of the member-vagrant hypothesis. Can J Fish Aquat Sci 53:424–433
Bernatchez L, Wilson CC (1998) Comparative phylogeography of Nearctic and Palearctic fishes. Mol Ecol 7:431–452
Bradbury IR, Coulson MW, Campana SE, Bentzen P (2006) Morphological and genetic differentiation in anadromous smelt Osmerus mordax (Mitchill): disentangling the effects of geography and morphology on gene flow. J Fish Biol 69:95–114
Bradbury IR, Campana SE, Bentzen P (2008a) Low genetic connectivity in an estuarine fish with pelagic larvae. Can J Fish Aquat Sci 65:147–158
Bradbury IR, Campana SE, Bentzen P (2008b) Estimating contemporary early life-history dispersal in an estuarine fish: integrating molecular and otolith elemental approaches. Mol Ecol 17:1438–1450
Bradbury IR, Coulson MW, Campana SE et al (2011) Contemporary nuclear and mitochondrial genetic clines in a north temperate estuarine fish reflect Pleistocene vicariance. Mar Ecol Prog Ser 438:207–218
Bugaev AV (2007) The fish of the Kamchatka River watershed. Kamchatpress, Petropavlovsk-Kamchatsky (in Russian)
Bugaev AV, Amelchenko YuN, Lipnyagov SV (2014) Rainbow smelt Osmerus mordax dentex in the shelf zone and inland water bodies of Kamchatka: state of stock, fishery, and biological structure. Izv TINRO 178:3–24 (in Russian)
Burlak OV (2018) Research of the Azian smelt Osmerus mordax in the Amur river basin in 2017. Prospects for fisheries and aquaculture in the modern world, p 66 (in Russian)
Chereshnev IA (1998) Biogeography of freshwater fishes of far east of Russia. Dal’nauka, Vladivostok (in Russian)
Chereshnev IA, Volobuev VV, Shestakov AV, Frolov SV (2002) Salmonoid fishes in Russian north-east. Dal’nauka, Vladivostok (in Russian)
Chernyavsky VI (1981) Circulation systems in the Sea of Okhotsk. Russian Proc TINRO 105:13–19
Costello AB, Down TE, Pollard SM et al (2003) The influence of history and contemporary stream hydrology on the evolution of genetic diversity within species: an examination of microsatellite DNA variation in bull trout, Salvelinus confluentus (Pisces: Salmonidae). Evolution 57:328–344
Coulson MW, Paterson IG, Green A et al (2006) Characterization of di- and tetranucleotide microsatellite markers in rainbow smelt (Osmerus mordax). Mol Ecol Res 6:942–944
Dieringer D, Schlötterer C (2003) Microsatellite analyser (MSA): a platform independent analysis tool for large microsatellite data sets. Mol Ecol Notes 3:167–169
Dodson JJ, Tremblay S, Colombani F et al (2007) Trans-Arctic dispersals and the evolution of a circumpolar marine fish species complex, the capelin (Mallotus villosus). Mol Ecol 16:5030–5043
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Estoup A, Angers B (1998) Microsatellites and minisatellites for molecular ecology: theoretical and experimental considerations. In: Carvalho GR (ed) Advances in molecular ecology. IOS Press, Amsterdam, pp 55–86
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Evseenko SA, Mishin AV (2011) On the distribution of larvae and localization of spawning stocks of White Sea herring Clupea pallasii marisalbi. J Ichthyol 51:776–787
Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Filatov NN, Nazarova LE, Druzhinin PV (2019) Influence of climatic and anthropogenic factors of the White Sea-catchment system. Proc KNT RAS 9:30–50. https://doi.org/10.17076/lim1117
Fisch KM, Petersen JL, Baerwald MR et al (2009) Characterization of 24 microsatellite loci in delta smelt, Hypomesus transpacificus, and their cross-species amplification in two other smelt species of the Osmeridae family. Mol Ecol Res 9:405–408
Garza JC, Williamson EG (2001) Detection of reduction in population size using data from microsatellite loci. Mol Ecol 10:305–318
Gladenkov AY, Gladenkov YB (2004) Onset of connections between the Pacific and Arctic Oceans through the Bering Strait in the Neogene. Stratigr Geol Correl 12:175–187
Goudet J (2001) FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3). http://www.unil.ch/izea/softwares/fstat.html
Grioche A, Koubbi P (1997) A preliminary study of the influence of a coastal frontal structure on ichthyoplankton assemblages in the English Channel. ICES J Mar Sci 54:93–104
Gritsenko OF (2002) Diadromous fishes of Sakhalin (systematics, ecology, fisheries). VNIRO, Moscow (in Russian)
Hardy OJ, Vekemans X (2002) SPAGEDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620
Hardy OJ, Charbonnel N, Freville H, Heuertz M (2003) Microsatellite allele sizes: a simple test to assess their significance on genetic differentiation. Genetics 163:1467–1482
Hutchinson DW, Templeton AR (1999) Correlation of pairwise genetic and geographic distance measures inferring the relative influences of gene flow and drift on the distribution of genetic variability. Evolution 53:1898–1914
Iles TD, Sinclair M (1982) Atlantic herring: stock discreteness and abundance. Science 215:627–633
Jenkins DG, Carey M, Czerniewska J et al (2010) A meta-analysis of isolation by distance: relic or reference standard for landscape genetics? Ecography 33:315–320
Jensen JL, Bohonak AJ, Kelley ST 2005. Isolation by distance, web service. BMC Genet 6:13. http://ibdws.sdsu.edu/
Kaukinen KH, Supernault KJ, Miller KM (2004) Development of microsatellite loci in eulachon (Thaleichthys pacificus). Mol Ecol Res 4:632–634
Khen GV, Ustinova EI, Sorokin Y (2019) Principal climate indices for the North Pacific: nature and history (a review). Russian Proc TINRO 197:166–181
Kirpichnikov VS (1935) Biological and systematic essay on smelt of the White Sea, Czech Bay, and Pechora River. Tr. VNIRO 2:103–191 (in Russian)
Klyukanov VA (1975) The status of Osmeridae in the system of order Salmoniformes. J Ichthyol 15:3–20
Kobylyanskii SG, Drits AV, Mishin AV et al (2014) Small scale distribution of the White Sea herring larvae (Clupea pallasii marisalbi) in relation to hydrophysical features. Oceanology 54:752–762
Koizumi I, Yamamoto S, Maekawa K (2006) Decomposed pairwise regression analysis of genetic and geographic distances reveals a metapopulation structure of stream-dwelling Dolly Varden charr. Mol Ecol 15:3175–3189
Kopelman NM, Mayzel J, Jakobsson M et al (2015) Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour 15:1179–1191
Koskinen MT, Nilsson J, Veselov AJ et al (2002) Microsatellite data resolve phylogeographic patterns in European grayling, Thymallus thymallus, Salmonidae. Heredity 88:391–401
Kottelat M, Freyhof J (2007) Handbook of European freshwater fishes. Kottelat Cornol and Freyhof, Berlin
Kovach AI, Breton TS, Enterline C, Berlinsky DL (2013) Identifying the spatial scale of population structure in anadromous rainbow smelt (Osmerus mordax). Fish Res 141:95–106
Lewis PO, Zaykin D (2001) Genetic data analysis: computer program for the analysis of allelic data: Version 1.0 (d 16c). http://lewis.eeb.unconn.edu/lewishome/software.html
Lu G, Basley DJ, Bernatchez L (2001) Contrasting patterns of mitochondrial DNA and microsatellite introgressive hybridization between lineages of lake whitefish (Coregonus clupeaformis); relevance for speciation. Mol Ecol 10:965–985
Manni F, Guérard E, Heyer E (2004) Geographic patterns of (genetic, morphologic, linguistic) variation: how barriers can be detected by “Monmonier’s algorithm.” Human Biol 76(2):173–190
McCauley DE (1993) Genetic consequences of extinction and recolonization in fragmented habitats. In: Kareiva PM, Kingsolver JG, Huey RB (eds) Biotic interactions and global change. Sinauer Associates, Sunderland, pp 217–233
McLean J, Taylor E (2001) Resolution of population structure in a species with high gene flow: microsatellite variation in the eulachon (Osmeridae: Thaleichthys pacificus). Mar Biol 139:411–420
Munk P, Larsson PO, Danielssen DS, Moksness E (1999) Variability in frontal zone formation and distribution of gadoid fish larvae at the shelf break in the northeastern North Sea. Mar Ecol Prog Ser 177:221–233
Naumova TN, Kolomeitsev VV (2015) Some data about biology and distribution of larval capelin (Mallotus villosus catervarius) on the shelf of west Kamchatka. Issled Vodn Biol Resur Kamchat Sev-Zap Chasti Tikhogo Okeana 38:84–92 (in Russian)
Nellbring S (1989) The ecology of smelts (genus Osmerus): a literature review. Nord J Freshw Res 65:116–145
Pantyulin AN (2003) Hydrological system of the White Sea. Oceanology 43:1–14
Pantyulin AN (2012) Dynamics, structure, and water masses. In: Lisitzin AP (ed) The White Sea system. Water column and interacting atmosphere, cryosphere, river flow and biosphere. Nauchnyi Mir, Moscow, pp 309–379 (in Russian)
Parukhina LV (2009) Summer ichthyoplankton of the Dvina Bay of the White Sea. In: Proceedings of the XXVIII international conference on biological resources of the White Sea and Inland Reservoirs of European North, KNT RAS, Petrozavodsk, pp 407–411 (in Russian)
Parukhina LV (2016) Abundance and some features of biology of smelt Osmerus mordax dentex larvae in the Onega Bay of the White Sea. In: Proceedings of the conference on marine biological research: achievements and perspectives, ECOSI-Hydrophisic, Sevastopol, pp 239–241 (in Russian)
Peterson DP, Ardren WR (2009) Ancestry, population structure, and conservation genetics of Arctic grayling (Thymallus arcticus) in the upper Missouri River, USA. Can J Fish Aquat Sci 66:1758–1774
Piry S, Luikart G, Conuet JM (1999) Bottleneck: a computer program for detecting recent reduction in the effective population size using allele frequency data. J Hered 90:502–503
Podushko YN (1970) Biological characteristic of the rainbow smelt Osmerus eperlanus dentex Steindachner in Lower Amur. Izv. TINRO 74:130–138 (in Russian)
Pritchard J, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Raymond M, Rousset F (1995) An exact test for population differentiation. Evolution 49:1280–1283
Sabates A, Olivar MP (1996) Variation of larval fish distributions associated with variability in the location of a shelf-slope front. Mar Ecol Prog Ser 135:11–20
Saint-Laurent R, Legault M, Bernatchez L (2003) Divergent selection maintains adaptive differentiation despite high gene flow between sympatric rainbow smelt ecotypes (Osmerus mordax Mitchill). Mol Ecol 12:315–330
Semenova AV, Stroganov AN, Afanas’ev KI, Rubtsova GA (2015) Population structure and variability of Pacific herring (Clupea pallasii) in the White Sea, Barents and Kara Seas revealed by microsatellite DNA analyses. Polar Biol 38:951–965
Semenova AV, Stroganov AN, Ponomareva EV, Afanas’ev KI (2019a) Microsatellite variability of the Arctic rainbow smelt Osmerus dentex from the White Sea. Russ J Genet 55:770–772
Semenova AV, Stroganov AN, Bugaev AV et al (2019b) An analysis of microsatellite polymorphism in the population of the Arctic rainbow smelt Osmerus dentex from Eastern and Western Kamchatka. Russ J Genet 55:77–88
Sendek DS, Studenov II, Sherstkov VS et al (2005) Genetic differentiation among smelt fish of the genus Osmerus (Osmeridae, Salmoniformes) in the European North of Russia. In: Salmonid fishes of eastern Fennoscandia. Karelsk Nauch Tsentr Ross Akad Nauk, Petrozavodsk, pp 148–157 (in Russian)
Skurikhina LA, Oleinik AG, Kukhlevsky AD et al (2015) Genetic differentiation of Pacific smelt Osmerus mordax dentex inferred from the data of mitochondrial DNA analysis. Rus J Genet 51:1221–1232
Skurikhina LA, Oleinik AG, Kukhlevsky AD et al (2018) Phylogeography and demographic history of the Pacific smelt Osmerus dentex inferred from mitochondrial DNA variation. Polar Biol 41:877–896
Skurikhina LA, Oleinik AG, Kukhlevsky AD et al (2019) Molecular genetic analysis of the relationships and origin of smelt (Hypomesus, Osmeridae), a new component of the fauna of the Barents Sea. Rus J Genet 55:1082–1092
Slatkin M (1985) Rare alleles as indicators of gene flow. Evolution 39:53–65
Slatkin M (1995) A measure of population subdivision based on microsatellite allele frequencies. Genetics 139:457–462
Stamford MD, Taylor EB (2004) Phylogeographical lineages of Arctic grayling (Thymallus arcticus) in North America: divergence, origins and affinities with Eurasian Thymallus. Mol Ecol 13:1533–1549
Stasenkov VA, Studenov II, Novoselov AP et al (2011) Pomor fisheries. Pravda Severa, Arkhangelsk ([in Russia])
Taylor EB, Dodson JJ (1994) A molecular analysis of relationships and biogeography within a species complex of Holarctic fish (genus Osmerus). Mol Ecol 3:235–248
Tonteri A, Titov S, Veselov A et al (2005) Phylogeography of anadromous and non-anadromous Atlantic salmon (Salmo salar) from northern Europe. Ann Zool Fennici 42:1–22
Ulchenko VA, Matkovskiy AK, Stepanov SI et al (2016) Fish resources and their development in the estuaries of the Kara Sea and the Laptev Sea. Proc VNIRO 160:116–132 (in Russian)
Vähä J-P, Erkinaro J, Niemelä E, Primmer CR (2007) Life-history and habitat features influence the within-river genetic structure of Atlantic salmon. Mol Ecol 16:2638–2654
Van Oosterhout C, Hutchinson WF, Wills DP, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Varkentin AI, Buslov AV, Tepnin OB (2001) Some features of spawning and distribution of pollock eggs in the waters of Western Kamchatka. Proc TINRO 128:177–187 (in Russian)
Vasilets PM (2000) On scale structure and growth rate of the Pacific smelt Osmerus mordax dentex (Osmeridae) from the Kamchatka waters during the first two years of life. J Ichthyol 40:397–401
Vasilets PM, Trofimov IK, Raevskii RV (2000) Morphological differentiation of Rainbow Smelt, Osmerus mordax dentex Kamchatka Waters. Tr. KamchatNIRO 5:101–105 (in Russian)
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Ye J, Coulouris G, Zaretskaya I et al (2012) PrimerBLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinform 13:134
Zhivotovsky LA (1991) Population biometrics. Nauka, Moscow (in Russian)
Acknowledgements
The reported study was funded by Russian Foundation for Basic Research, Project Number 19-04-00244-a. The use of DNA markers for population differentiation was supported by the Russian Science Foundation No. 19-16-00101. The work was also carried out within the framework of the state theme GZ 0112-2019-0002 (sub-topic ‘Ecological and genetic structure of the species’).We are grateful to Alla N. Gorbacheva and Edward V. Golzha for their help in samples collection. We thank the Editor in Chief, Dieter Piepenburg, and the three reviewers Niall McKeown, Cristian B. Canales-Aguirre, and Vidar Wennevik for their valuable comments on an earlier draft of this manuscript.
Author information
Authors and Affiliations
Contributions
AVS performed and designed research, analysed data, and wrote the manuscript. EP and OV conducted field work. ANS and KA provided research opportunity. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Semenova, A.V., Stroganov, A.N., Ponomareva, E.V. et al. Large-scale genetic structure and diversity of Arctic rainbow smelt Osmerus dentex Steindachner et Kner, 1870 throughout its distributional range based on microsatellites. Polar Biol 44, 927–940 (2021). https://doi.org/10.1007/s00300-021-02848-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00300-021-02848-x