Abstract
Key message
PDE1 acts as a mediator of primary root growth in response to Pi deficiency.
Abstract
Phosphorus is commonly considered as a limiting nutrient for plant growth, which is mainly due to the immobility and uneven distribution of phosphate (Pi) in soils so that available Pi is not adequate in the rhizosphere. Although various mediators have been identified in Pi sensing and response, more details need to be uncovered in plant Pi-deficiency tolerance. Here, we isolated a mutant, termed pde1 (phosphate-deficiency sensitive 1), showing the hypersensitive Pi-deficiency-induced growth inhibition of primary roots. PDE1 encodes a hydroxyphenylpyruvate reductase with rare activity in vitro and repressed by Pi deficiency. Histochemical analysis displayed that Pi-deprived pde1 accumulated more Fe and reactive oxygen species (ROS) in primary roots than the wild type (WT). Addition of ferrozine, a Fe2+ chelator, or a ROS scavenger (e.g., thiourea and potassium iodide), alleviated the sensitivity of Pi-deficiency in pde1 primary roots. By contrast, pde1 showed reduced cotyledon expansion rate with treatment of H2O2 compared to WT. Taken together, these results suggested that PDE1 is responsible for regulating primary root growth in response to Pi deficiency, which is associated with ROS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As a basic constituent of molecules such as ATP, nucleic acid, and phospholipid, phosphorus is essential for plant growth and development (Péret et al. 2011). Root system has been recognized as the major structure in response to phosphate (Pi) deficiency, in which the inhibition of primary root growth was observed under Pi-deficient condition (Williamson et al. 2001; Lynch et al. 2001). Growing evidence has been emerged to uncover the regulatory mechanism of plant Pi-deficiency tolerance, and numerous genes were identified to participate in the process over the past 20 years (Liu 2021). LPR1 (low phosphate root 1) was a key mediator in the regulation of primary root growth under Pi-deprived condition (Svistoonoff et al. 2007). LPR1 encodes a multicopper oxidase and possesses a ferroxidase activity for conversion of Fe2+ to Fe3+. The LPR1-dependent iron redox cycle could promote ROS accumulation in root tips (Müller et al. 2015), and ALS3 (aluminum sensitive 3) cooperated with LPR1 to mediate the root iron homeostasis in response to Pi deficiency (Dong et al. 2017). STOP1 (sensitive to proton toxicity 1), a transcription factor, and ALMT1 (Al-activated malate transporter 1), a malate transporter protein, were involved in the Pi-deficiency tolerance (Hoekenga et al. 2006; Balzergue et al. 2017). Low Pi activated the STOP1-ALMT1 module to facilitate malate transport and malate-dependent iron accumulation in the apoplast (Mora-Macías et al. 2017). In addition, ALS3 interacted with STAR1 (sensitive to aluminum rhizotoxicity 1) to form an ABC transporter complex for the mediation of Pi-deficiency root architecture remodeling (Dong et al. 2017), and ALS3/STAR1 repressed STOP1 accumulation in the nucleus to increase the tolerance against Pi deficiency (Wang et al. 2019). Recently, blue light was found to trigger the malate-mediated photo-Fenton reaction for growth inhibition of primary roots under Pi-deficient condition (Zheng et al. 2019). Blue light, thereby, in accompany with Fe2+, Fe3+, malate, H2O2, and low pH was indispensable to inhibit primary root growth in response to Pi deficiency (Liu 2021). Aside from this, Pi deficiency also slightly inhibited primary root growth in darkness (Silva-Navas et al. 2019).
ROS play a crucial role in root growth and development (Liszkay et al. 2004; Tsukagoshi et al. 2010; Manzano et al. 2014). Accumulation of O2·− and H2O2 inhibited the primary root growth and root hair formation in Arabidopsis thaliana (Foreman et al. 2003; Dunand et al. 2007), and the respiratory burst oxidase homolog (RBOH)-mediated ROS production could promote the lateral root development (Orman-Ligeza et al. 2016). In addition to being a signaling molecule in root system architecture morphogenesis (Prakash et al. 2020; Bian et al. 2021), ROS were capable of regulating root growth in response to nutrient deficiencies (Shin and Schachtman 2004; Wang et al. 2021). The H2O2 content and expression levels of genes encoding ROS-producing enzymes were up-regulated, after roots were exposed to nitrogen, phosphorus or potassium-deficiency environment (Shin et al. 2005). Reyt et al. (2015) reported that iron homeostasis was associated with the generation of ROS to orchestrate the root growth (Reyt et al. 2015).
In this study, a mutant pde1 (phosphate-deficiency sensitive 1), which displayed the Pi-deficiency sensitivity in primary root growth, was identified from a forward genetic screen. The length of primary roots in pde1 was significantly shorter than that in wild type (WT) under Pi-deprived condition. The pde1 mutant had a T-DNA insertion site in the second exon of At2g45630 which encodes a hydroxyphenylpyruvate reductase (HPPR) with rare enzyme activity for 4-hydroxyphenylpyruvic acid (pHPP), phenylpyruvic acid and hydroxypyruvic acid substrates (Xu et al. 2018). Expression of At2g45630 in pde1 could complement the inhibited growth of primary roots in Pi-deprived pde1, and PDE1 was suppressed by Pi deficiency. Under Pi-deprived condition, more Fe was accumulated in pde1 primary roots, and addition of ferrozine, a Fe2+ chelating agent, rescued the inhibited primary root growth. Pi-deprived pde1 also showed enhanced accumulation of H2O2 in primary roots by DAB staining. Supplement of Thiourea or potassium iodide in Pi-deprived medium alleviated the sensitivity of Pi-deficiency in pde1 primary roots. Additionally, the cotyledon expansion rate in pde1 was much lower than that in WT with treatment of 3 mM H2O2. Overall, these results suggested the role of PDE1 in the Pi-deficiency tolerance that was associated with ROS.
Materials and methods
Plant materials and growth conditions
All Arabidopsis plants used in this study were Columbia-0 (Col-0) background, and the T-DNA insertional mutants of pde1 (SALK_201658) and hppr3-2 (SALK_019014) were obtained from the Arabidopsis Biological Resource Center (ABRC, Ohio State University, Columbus, OH, USA). hppr2cr was generated via lustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR‐associated nuclease 9 (Cas9) genome editing system (Wang et al. 2015). Plants were grown aseptically or on soil in a growth chamber under a 16-h light/8-h dark cycle at 22 °C and 60% relative humidity. For aseptic growth, seeds were surface-sterilized in 25% bleach and 0.1% Triton X-100 for 4 min, washed thoroughly with sterile water, and placed for 3 d at 4 °C in darkness. Then, the seeds were sown onto Petri dishes containing different types of MS medium. The +Pi medium (pH 5.7) consisted of MS medium (MS524, PhytoTech) containing 0.17 g/L KH2PO4, 1% Suc, and 1% agar. The −Pi medium (pH 5.7) consisted of MS medium without nitrogen, phosphorous, and potassium (MS407, PhytoTech) supplemented with 1.65 g/L NH4NO3, 1.9 g/L KNO3, 1% Suc, and 1% agar. After grown on medium for 1–2 weeks, seedlings were transferred into a 1:1:8 mixture of vermiculite, perlite, and peat moss and watered every other day for soil growth.
Mutant screening
Approximately 1000 surface-sterilized mutant individuals from a T-DNA insertion mutant library were grown on −Pi medium for 9 days at 22 °C under a 16-h light/8-h dark cycle. Then, the primary root length was considered as a phenotypic criterion for the mutant screening. The library was used for the glucose-sensitive mutant screen in our previous study (Zheng et al. 2020).
Mutant identification and plant transformation
For genotyping of pde1, genomic DNA was extracted from plant leaves and subjected to PCR using specific primers of T-DNA and genome. The homozygous seeds obtained from F2 progenies were used for analysis. For the complementation assay, the open reading frame of PDE1 was cloned into the pCAMBIA1305-EGFP plasmid at the Mlu I and Kpn I restriction sites to generate the 35S::PDE1-EGFP vector. Then the plasmid was introduced into Agrobacterium tumefaciens (GV3101) and transformed into pde1 via a floral-dip method (Clough and Bent 1998). The transgenic plants were screened on 1/2 MS medium containing 50 mg/L hygromycin. For the phenotypic analysis, T3 or T4 homozygous lines were used.
Light-avoidance experiment of root
The light-avoidance experiment of root was performed as described previously (Zheng et al. 2019) with minor modification. Briefly, WT or pde1 seeds were grown on the Pi-replete or -deprived medium for 4 days, then aluminum foil was used to cover the root part for another 4-day growth in avoidance of light.
Perls staining
Perls staining was used to detect iron accumulation in root tips. In brief, the whole seedlings were collected and vacuum infiltrated with Perls staining solution containing equal volumes of 4% (v/v) HCl and 4% (w/v) potassium ferrocyanide for 15 min. After infiltration, the samples were washed with deionized water three times to stop the reaction and immersed in the clearing solution (1 g/ml chloral hydrate in 15% glycerol). Then, the root tips were examined and imaged under bright-field illumination on an optical microscope (Uop, UB103i, China).
DAB staining
DAB staining was performed as previously described (Zheng et al. 2020). Nine-day-old seedlings were incubated in the staining solution (1 mg/ml DAB, 10 mM Na2HPO4, 0.05% Tween 20, pH7.0) at room temperature for 6 h, and then fixed in a solution of ethanol:lactic acid:glycerol (3:1:1). After being boiled for 5 min, root tips were cut-off and imaged under bright-field illumination on an optical microscope (Uop, UB103i, China).
RNA isolation and PCR
Total RNA was extracted using RNAiso plus (TAKARA) according to the manufacturer’s instructions. RNA was further purified by chloroform/isoamyl alcohol extraction and isopropanol precipitation. 1 μg total RNA was used to synthesize cDNA using the Reverse Transcriptase M-MLV kit (TAKARA) after removing the residual DNA by AccuRT Genomic DNA Removal Kit (abm). Quantitative real-time PCR (qPCR) was performed with the LightCycler® 96 system (Roche) using EvaGreen 2× qPCR MasterMix (abm). The primers are listed in Table S1.
Statistical analysis
ImageJ software was used for determining the primary root length and quantifying the DAB staining intensity. All experiments were performed at least three technical replicates and two to four biological replicates. GraphPad Prism 5 and Microsoft Excel 2019 were used for calculating mean and standard deviation, except that qPCR data were analyzed via LightCycle® 96 SW1.1 software. Statistical significance was determined according to the student’s t test and one-/two-way ANOVA.
Results
Inhibition of primary root growth in pde1 under Pi-deprived condition
To identify the essential regulators in response to Pi deficiency, a T-DNA insertion mutant pde1 (phosphate-deficiency sensitive 1) in Arabidopsis was identified by a forward genetic screen. When seedlings were grown on Pi-replete (+Pi) medium under white light (16-h light/8-h dark cycle) for 9 days, there was not significant difference between pde1 and the wild type (WT) in the primary root length. In contrast, the primary root length of pde1 was approximately two thirds of that of WT under Pi-deprived (−Pi) condition (Fig. 1). Previous study reported that blue light triggered the photo-Fenton reaction, and a canonical Fenton reaction produced ·OH that can inhibit primary root growth, no matter whether seedlings were grown on +Pi or −Pi medium (Zheng et al. 2019). Therefore, we investigated whether the sensitive phenotype of pde1 to Pi deprivation was affected by light. Under −Pi condition, seedlings were exposed to white light for 4 days followed by growth with aluminum foil-covering roots for another 4 days. As shown in Fig. S1, pde1 showed shorter primary roots than WT with or without the root cover (Fig. S1), implying that the inhibition of primary root growth in pde1 was not dependent on light under −Pi condition. Taken together, these results suggested the involvement of PDE1 in Pi-deficiency tolerance.
pde1 showed the growth-inhibited phenotype of primary roots under Pi-deprived condition. a Representative images of 9-day-old WT, pde1, and pde1 complementary lines (COM#1 and COM#2) grown on MS medium with 1.25 mM phosphate (+Pi) or no phosphate (−Pi) in the presence of light (16-h light/8-h dark cycle). b The statistical analysis of primary root length in WT, pde1, COM#1 and COM#2 grown as described in a. The values are means ± the standard deviation. One-way analysis of variance was followed by Tukey’s test, and different letters represent significant difference (n = 60, p < 0.01). Scale bar = 1 cm
Identification of PDE1
To investigate the function of PDE1 in Pi-deficiency tolerance, we first confirmed the T-DNA insertion site in pde1 via genotyping and found that the insertion site was located in the second exon of At2g45630 open reading frame (ORF) (Fig. 2a, b). RT-PCR results showed that the expression levels of PDE1 in pde1 were not detected, suggesting that the T-DNA insertion disrupted PDE1 transcripts (Fig. 2b). To verify that the Pi-deprived phenotype of pde1 in primary root growth was due to lack of PDE1, 35S::PDE1-GFP was constructed and transformed in pde1. The primary root length of pde1 harboring 35S::PDE1-GFP was similar to that of WT under −Pi condition (Figs. 1a, b, 2b), videlicet, overexpression of At2g45630 in pde1 rescued the inhibited primary root growth of pde1. Overall, the mutation in At2g45630 was responsible for the sensitivity of Pi deficiency in pde1.
Identification and characterization of PDE1. a Schematic structure of the pde1 T-DNA insertion site. Black box, black line, and grey line indicate exon, intron, and untranslated regions, respectively. Inverted triangle indicates the T-DNA insertion site. LP/RP, left/right genomic primer; BP, T-DNA border primer. b Genotyping and PDE1 transcript analysis (RT-PCR) in WT, pde1, and pde1 complementary lines (COM#1 and COM#2). 35S-F, CaMV35S promoter forward primer; PDE1-R, PDE1 reverse primer. Nine-day-old seedlings were used to extract genomic DNA and total RNA. c Tissue-specific expression of PDE1. Se, seedlings; R, roots; St, stems; RL, rosette leaves; CL, cauline leaves; F, flowers; Si, siliques. d PDE1 was repressed by Pi deficiency. Roots of 8-day-old seedlings on +Pi or −Pi medium were used as samples. The values are means ± the standard deviation (n ≥ 3, t test: ** p < 0.01). UBQ5 was used as the control (c, d)
At2g45630 was previously reported to encode a hydroxyphenylpyruvate reductase (HPPR4) that possessed little catalytic activity and was localized to the cytoplasm (Xu et al. 2018). There are three HPPRs (HPPR2/3/4) in Arabidopsis (Fig. S2), thus it was ascertained whether lack of HPPR2 or HPPR3 displayed the similar phenotype to pde1 in primary root growth under −Pi condition. A HPPR2 mutant (hppr2cr) was generated by the CRISPR/Cas9 system, which had a T insertion (a missense mutation) in the first exon, resulting in a premature translation-termination, and a HPPR3 mutant (hppr3-2) with a T-DNA insertion in the second exon was obtained from the TAIR stock center (Fig. S3a–c). Under −Pi condition, hppr2cr and hppr3-2 did not show the significant difference in primary root length compared to WT (Fig. S3d–e). The expression profile of PDE1 was examined by qRT-PCR, and PDE1 transcripts were detected in all tested tissues. As shown in Fig. 2c, leaves had the higher expression levels of PDE1 than other tissues such as stems and flowers (Fig. 2c). To test PDE1 expression in response to Pi deficiency, total RNA was extracted from 8-day-old roots, and PDE1 was repressed under −Pi condition (Fig. 2d).
Effects of Fe on pde1 primary root growth under Pi-deprived condition
Fe accumulation was found to enhance in Arabidopsis was in response to Pi deficiency, and Pi and Fe exhibited the antagonistic effect in the primary root growth (Misson et al. 2005; Hirsch et al. 2006; López-Bucio et al. 2019). Thus, we first examined Fe in the primary roots with Perls staining that mainly recognizes Fe3+ by redox reaction (Meguro et al. 2007). There was no obvious difference in the staining intensity and distribution between pde1 and WT grown on +Pi medium for 9 days, whereas pde1 showed enhanced staining intensity and altered distribution compared with WT under −Pi condition (Fig. 3a). Accumulation of Fe caused by Pi deficiency in pde1 was detected along the root axis containing root cap, root apical meristem, elongation zone, and maturation zone, but that in WT was only observed in root maturation zone (Fig. 3a right panel). Additionally, ferrozine [3-(2-pyridyl)-5,6-bis (4-phenylsulfonic acid)-1,2,4-triazine], a Fe2+ chelating agent (Simonzadeh and Jaselskis 1984), was used to evaluate association between the inhibition of primary root growth in Pi-deprived pde1 and Fe accumulation. As shown in Fig. 3b and c, the addition of ferrozine rescued the inhibited primary root growth of not only pde1 but also WT (Fig. 3b, c). In contrast, ferrozine had no obvious effect on the primary root growth of Pi-replete pde1 and WT (Fig. S4), suggesting the role of Fe in primary root growth under Pi-deprived condition. Together, lack of PDE1 accumulated more Fe and altered Fe deposition pattern in the Pi-deprived primary roots, which could account for the Pi-deprived phenotype of pde1. We also examined the expression levels of several genes related to Pi deficiency (RNS1, LPR1, ALS3, STAR1, STOP1, and ALM1). All the six genes were significantly up-regulated in both primary roots of pde1 and WT under Pi-deprived condition, in which the Pi deficiency-induced expression levels of LPR1, ALS3, and ALMT1 were higher in pde1 than that in WT (Fig. 4).
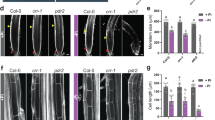
pde1 over-accumulated Fe in primary roots under Pi-deprived condition. a Nine-day-old WT and pde1 on +Pi or −Pi medium were stained by Perls staining for Fe detection. Representative images (b) and the statistical analysis of primary root length (c) in WT and pde1 on −Pi medium with or without 500 μM ferrozine (Fer). The values are means ± the standard deviation (n = 60, t test: **p < 0.01). Scale bar = 0.2 mm (a) and 0.5 cm (b)
The expression levels of Pi-related genes in pde1 under Pi-deprived condition. The primary root apexes (2–3 mm) of 5-day-old seedlings grown on +Pi or −Pi medium were collected for total RNA extraction. ACTIN1 was used as the control. Error bars represent the standard deviation. Two-way analysis of variance was followed by Tukey’s test, and different letters represent significant difference (n ≥ 3, p < 0.01)
The involvement of ROS in PR inhibition growth of Pi-deprived pde1
LPR1 was known to act as a ferroxidase for oxidation of Fe2+ to Fe3+ and H2O2 production, and redox signaling triggered by LPR1-dependent Fe2+ oxidation could regulate callose deposition in root meristems under low Pi condition (Svistoonoff et al. 2007; Müller et al. 2015). H2O2 levels were thereby examined in the primary roots by DAB staining. Under +Pi condition, the brown staining intensity in primary roots of pde1 was similar to that of WT (Fig. 5a right panel). In contrast, pde1 showed darker brown in primary roots compared to WT under −Pi condition (Fig. 5a left panel, b), albeit only the background brown was observed in cotyledons of pde1 and WT (Fig. S5). These results suggested that H2O2 accumulation in primary roots of pde1 was significantly enhanced in response to Pi deficiency.
Additionally, thiourea (Thi) and potassium iodide (KI), two antioxidants, were added into −Pi medium, respectively. Thi is able to scavenge superoxide radicals, hydroxyl radicals and H2O2 (Kelner et al. 1990), and KI could detoxify H2O2 (Dunand et al. 2007). As expected, both Thi and KI could attenuate the inhibition in primary root growth of pde1 under −Pi condition, and KI showed more favorable effect against ROS on the Pi-deprived primary root growth (Fig. 6a, b). Next, the sensitivity of pde1 to H2O2 was examined during early seedling growth. There was no notable difference in the radicle germination rate between pde1 and WT with or without 3 mM H2O2 (Fig. 6c, d); however, the cotyledon expansion rate of pde1 was lower than that of WT in the presence of H2O2 (Fig. 6e), implying the role of PDE1 in oxidative stress. Taken together, these results suggested that ROS was involved in the Pi-deficiency tolerance of pde1.
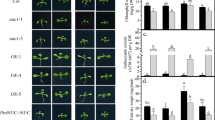
Association of ROS with growth inhibition of primary roots in Pi-deprived pde1. Representative images (a) and the statistical analysis of primary root length (b) in WT and pde1 on −Pi medium with or without 300 μM thiourea (Thi)/500 μM potassium iodide (KI). c Representative images of 7-day-old WT and pde1 on MS medium with or without 3 mM H2O2. The statistical analysis of radicle germination rate (d) and cotyledon expansion rate (e) in 7-day-old WT and pde1 in response to H2O2. CK, control check. The values are means ± the standard deviation (n = 60 (b) and ≥ 100 (d–e), t test: * p < 0.05, ** p < 0.01). Scale bar = 0.5 cm (a, c)
Discussion
In previous studies, PDE1 was recognized as a hydroxyphenylpyruvate reductase with weak catalytic activity, which may be involved in the biosynthesis of tyrosine-derived metabolites (Timm et al. 2008; Xu et al. 2018). Here, we found that PDE1 contributed to the mediation of Arabidopsis primary root growth in response to Pi deficiency. Under −Pi condition, accumulation of Fe in 9-day-old primary roots of WT was observed in the maturation zone, rarely in the elongation zone and root apical meristem; however, pde1 accumulated more Fe in the elongation zone and root apical meristem of primary roots compared to WT (Fig. 3a). It has been reported that Fe was accumulated mostly in the maturation zone of Pi-deprived wild-type primary roots in the early growth phase, and progressively extended to the tip of the primary roots along with the increased growth till root apical meristem was completely exhausted which inhibited growing (Dong et al. 2017; Hoehenwarter et al. 2016; Wang et al. 2019). Liu (2021) indicated that in response to Pi deficiency, plants with the same ecotype showed differences in primary root growth depending on compounds in the medium such as the salt composition, sucrose levels and gelling agents, as well as light intensity (Liu 2021). Different Arabidopsis ecotypes also displayed the variation of primary root growth under limited Pi availability (Chevalier et al. 2003). Therefore, the wild type, as a critical control, is indispensable for the detection of Fe distribution in primary roots without Pi. For instance, we used agar instead of agarose, MS medium instead of 1/2 MS medium, and Pi-free medium instead of 10 µM Pi-containing medium as the −Pi medium in this study (Nacry et al. 2005; Dong et al. 2017; Xu et al. 2020), which could cause the difference of Fe distribution during the similar growth phase of primary roots. Fe and Pi were known to have the antagonistic effect on primary root growth (Müller et al. 2015). Pi-deficiency promotes Fe content in roots and redistribution towards root apical meristem where Fe acts as a source for ROS in the apoplast (Gutierrez-Alanis et al. 2018). ROS could trigger root apical meristem exhaustion and callose production to cause the growth inhibition of primary roots (Crombez et al. 2019). As expected, addition of Fe chelator into −Pi medium rescued the inhibited growth of primary roots in both pde1 and WT (Fig. 3b and c).
It has been demonstrated that Pi-deficiency-induced ALMT1 acting upstream of LPR1 was responsible for malate secretion into the apoplast of primary roots, which not only facilitated LPR1-derived Fe3+ accumulation by formation of Fe3+-malate complex but also promoted the expression of LPR1 by malate (Mora-Macías et al. 2017). The LPR1-derived Fe3+ production could contribute to ROS formation that activated callose deposition in the apoplast to block plasmodesmata, resulting in disrupted primary root growth (Müller et al. 2015). Lack of PDE1 enhanced the expression of Pi-deficiency-induced ALMT1 and LPR1 in primary roots (Fig. 4). Pi-deprived pde1 also showed more accumulation of H2O2 in primary roots (Fig. 5) and the sensitivity of H2O2 (Fig. 6c). It could be speculated that PDE1 in response to Pi deficiency may be involved in the ALMT1-LPR1-ROS regulatory cascade.
Arabidopsis genome encodes three HPPRs (HPPR2/3/4) with different subcellular localization, expression pattern and substrate preference (Xu et al. 2018). Both HPPR2 and HPPR3 were capable of reducing pHPP to pHPL (hydroxyphenylacetic acid) in the presence of NAD(P)H, and HPPR3 showed higher affinity for pHPP. pHPL is known as a critical intermediate in tyrosine metabolism for rosmarinic acid biosynthesis in plants, especially for Lamiaceae plant species (Petersen and Simmonds 2003). Silencing of CbHPPR in Coleus blumei hairy roots repressed rosmarinic acid production, and overexpression of CbHPPR had the opposite effect (Hücherig and Petersen 2013). The hairy roots of Salvia miltiorrhiza overexpressing SmHPPR showed enhanced rosmarinic acid content (Wang et al. 2017). Rosmarinic acid was firstly found in rosemary and recently reported to have several biological activities like antioxidant and antimicrobial properties that could contribute to plant defense response (Trócsányi et al. 2020). For instance, exogenous application of rosmarinic acid improved tomato heat tolerance with enhanced activity of some antioxidant enzymes (Zhou et al. 2022). Rosmarinic acid extracted from Zostera marina exhibited the nematicidal activity against pine wood nematode and the antibacterial activity the nematode carries, which may be beneficial for controlling pine wilt disease (Wang et al. 2012). Although HPPR4 (PDE1) showed 39.35% and 47.94% amino acid identity with HPPR2 and HPPR3, respectively (Fig. S2), HPPR4 did not possess the similar enzyme activity to HPPR2 and HPPR3 in vitro (Xu et al. 2018). Lack of PDE1 might affect rosmarinic acid production, resulting in the reduction of antioxidant capability. However, the growth inhibition phenotype of primary roots was not observed in the Pi-deprived hrrp2cr and hrrp3-2 (Fig. S3). Different from Lamiaceae plant species that have the high content of rosmarinic acid, Arabidopsis appears to mainly elicit phytohormone, not rosmarinic acid, as an accumulated compound in response to stress. PDE1 was thereby supposed to have the specific role in primary root growth under −Pi condition aside from acting as the potential HPPR in rosmarinic acid production. Future studies should elucidate PDE1 interactive proteins and its position in the regulatory network of plant Pi-deficiency tolerance.
In summary, PDE1 was identified to confer plant Pi-deficiency tolerance. Lack of PDE1 caused the growth inhibition phenotype in Pi-deprived primary roots, and enhanced accumulation of Fe and ROS was correlated with the phenotype.
Data availability
All relevant data are available from the corresponding author on request.
References
Balzergue C, Dartevelle T, Godon C, Laugier E, Meisrimler C, Teulon JM, Creff A, Bissler M, Brouchoud C, Hagège A, Müller J, Chiarenza S, Javot H, Becuwe-Linka N, David P, Péret B, Delannoy E, Thibaud MC, Armengaud J, Abel S, Pellequer JL, Nussaume L, Desnos T (2017) Low phosphate activates STOP1-ALMT1 to rapidly inhibit root cell elongation. Nat Commun 8:15300
Bian L, Wang Y, Bai H, Li H, Zhang C, Chen J, Xu W (2021) Melatonin-ROS signal module regulates plant lateral root development. Plant Signal Behav 16:1901447
Chevalier F, Pata M, Nacry P, Doumas P, Rossignol M (2003) Effects of phosphate availability on the root system architecture: large-scale analysis of the natural variation between Arabidopsis accessions. Plant, Cell Environ 26:1839–1850
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Crombez H, Motte H, Beeckman T (2019) Tackling plant phosphate starvation by the roots. Dev Cell 48:599–615
Dong J, Piñeros MA, Li X, Yang H, Liu Y, Murphy AS, Kochian LV, Liu D (2017) An Arabidopsis ABC transporter mediates phosphate deficiency-induced remodeling of root architecture by modulating iron homeostasis in roots. Mol Plant 10:244–259
Dunand C, Crèvecoeur M, Penel C (2007) Distribution of superoxide and hydrogen peroxide in Arabidopsis root and their influence on root development: possible interaction with peroxidases. New Phytol 174:332–341
Foreman J, Demidchik V, Bothwell JH, Mylona P, Miedema H, Torres MA, Linstead P, Costa S, Brownlee C, Jones JD, Davies JM, Dolan L (2003) Reactive oxygen species produced by NADPH oxidase regulate plant cell growth. Nature 422:442–446
Gutierrez-Alanis D, Ojeda-Rivera JO, Yong-Villalobos L, Cardenas-Torres L, Herrera-Estrella L (2018) Adaptation to phosphate scarcity: tips from Arabidopsis roots. Trends Plant Sci 23:721–730
Hirsch J, Marin E, Floriani M, Chiarenza S, Richaud P, Nussaume L, Thibaud MC (2006) Phosphate deficiency promotes modification of iron distribution in Arabidopsis plants. Biochimie 88:1767–1771
Hoehenwarter W, Monchgesang S, Neumann S, Majovsky P, Abel S, Muller J (2016) Comparative expression profiling reveals a role of the root apoplast in local phosphate response. BMC Plant Biol 16:106
Hoekenga OA, Maron LG, Piñeros MA, Cançado GM, Shaff J, Kobayashi Y, Ryan PR, Dong B, Delhaize E, Sasaki T, Matsumoto H, Yamamoto Y, Koyama H, Kochian LV (2006) AtALMT1, which encodes a malate transporter, is identified as one of several genes critical for aluminum tolerance in Arabidopsis. Proc Natl Acad Sci USA 103:9738–9743
Hücherig S, Petersen M (2013) RNAi suppression and overexpression studies of hydroxyphenylpyruvate reductase (HPPR) and rosmarinic acid synthase (RAS) genes related to rosmarinic acid biosynthesis in hairy root cultures of Coleus blumei. Plant Cell Tiss Organ Cult (PCTOC) 113:375–385
Kelner MJ, Bagnell R, Welch KJ (1990) Thioureas react with superoxide radicals to yield a sulfhydryl compound. Explanation for protective effect against paraquat. J Biol Chem 265:1306–1311
Liszkay A, van der Zalm E, Schopfer P (2004) Production of reactive oxygen intermediates (O2⋅−, H2O2, and ⋅OH) by maize roots and their role in wall loosening and elongation growth. Plant Physiol 136:3114–3123; discussion 3001
Liu D (2021) Root developmental responses to phosphorus nutrition. J Integr Plant Biol 63:1065–1090
López-Bucio JS, Salmerón-Barrera GJ, Ravelo-Ortega G, Raya-González J, León P, de la Cruz HR, Campos-García J, López-Bucio J, Guevara-García ÁA (2019) Mitogen-activated protein kinase 6 integrates phosphate and iron responses for indeterminate root growth in Arabidopsis thaliana. Planta 250:1177–1189
Lynch JP, Brown KM (2001) Topsoil Foraging an—Architectural Adaptation of Plants to Low Phosphorus Availability. 237:225–237
Manzano C, Pallero-Baena M, Casimiro I, De Rybel B, Orman-Ligeza B, Van Isterdael G, Beeckman T, Draye X, Casero P, Del Pozo JC (2014) The emerging role of reactive oxygen species signaling during lateral root development. Plant Physiol 165:1105–1119
Meguro R, Asano Y, Odagiri S, Li C, Iwatsuki H, Shoumura K (2007) Nonheme-iron histochemistry for light and electron microscopy: a historical, theoretical and technical review. Arch Histol Cytol 70:1–19
Misson J, Raghothama KG, Jain A, Jouhet J, Block MA, Bligny R, Ortet P, Creff A, Somerville S, Rolland N, Doumas P, Nacry P, Herrerra-Estrella L, Nussaume L, Thibaud MC (2005) A genome-wide transcriptional analysis using Arabidopsis thaliana Affymetrix gene chips determined plant responses to phosphate deprivation. Proc Natl Acad Sci USA 102:11934–11939
Mora-Macías J, Ojeda-Rivera JO, Gutiérrez-Alanís D, Yong-Villalobos L, Oropeza-Aburto A, Raya-González J, Jiménez-Domínguez G, Chávez-Calvillo G, Rellán-Álvarez R, Herrera-Estrella L (2017) Malate-dependent Fe accumulation is a critical checkpoint in the root developmental response to low phosphate. Proc Natl Acad Sci USA 114:E3563–E3572
Müller J, Toev T, Heisters M, Teller J, Moore KL, Hause G, Dinesh DC, Bürstenbinder K, Abel S (2015) Iron-dependent callose deposition adjusts root meristem maintenance to phosphate availability. Dev Cell 33:216–230
Nacry P, Canivenc G, Muller B, Azmi A, Van Onckelen H, Rossignol M, Doumas P (2005) A role for auxin redistribution in the responses of the root system architecture to phosphate starvation in Arabidopsis. Plant Physiol 138:2061–2074
Orman-Ligeza B, Parizot B, de Rycke R, Fernandez A, Himschoot E, Van Breusegem F, Bennett MJ, Périlleux C, Beeckman T, Draye X (2016) RBOH-mediated ROS production facilitates lateral root emergence in Arabidopsis. Development 143:3328–3339
Péret B, Clément M, Nussaume L, Desnos T (2011) Root developmental adaptation to phosphate starvation: better safe than sorry. Trends Plant Sci 16:442–450
Petersen M, Simmonds MS (2003) Rosmarinic acid. Phytochemistry 62:121–125
Prakash V, Vishwakarma K, Singh VP, Rai P, Ramawat N, Tripathi DK, Sharma S (2020) NO and ROS implications in the organization of root system architecture. Physiol Plant 168:473–489
Reyt G, Boudouf S, Boucherez J, Gaymard F, Briat JF (2015) Iron- and ferritin-dependent reactive oxygen species distribution: impact on Arabidopsis root system architecture. Mol Plant 8:439–453
Shin R, Schachtman DP (2004) Hydrogen peroxide mediates plant root cell response to nutrient deprivation. Proc Natl Acad Sci USA 101:8827–8832
Shin R, Berg RH, Schachtman DP (2005) Reactive oxygen species and root hairs in Arabidopsis root response to nitrogen, phosphorus and potassium deficiency. Plant Cell Physiol 46:1350–1357
Silva-Navas J, Conesa CM, Saez A, Navarro-Neila S, Garcia-Mina JM, Zamarreño AM, Baigorri R, Swarup R, Del Pozo JC (2019) Role of cis-zeatin in root responses to phosphate starvation. New Phytol 224:242–257
Simonzadeh N, Jaselskis B (1984) Reaction of iron(III) with tiron in the presence of ferrozine, and determination of tiron. Talanta 31:715–716
Svistoonoff S, Creff A, Reymond M, Sigoillot-Claude C, Ricaud L, Blanchet A, Nussaume L, Desnos T (2007) Root tip contact with low-phosphate media reprograms plant root architecture. Nat Genet 39:792–796
Timm S, Nunes-Nesi A, Pärnik T, Morgenthal K, Wienkoop S, Keerberg O, Weckwerth W, Kleczkowski LA, Fernie AR, Bauwe H (2008) A cytosolic pathway for the conversion of hydroxypyruvate to glycerate during photorespiration in Arabidopsis. Plant Cell 20:2848–2859
Trócsányi E, György Z, Zámboriné-Németh É (2020) New insights into rosmarinic acid biosynthesis based on molecular studies. Curr Plant Biol 23:100162
Tsukagoshi H, Busch W, Benfey PN (2010) Transcriptional regulation of ROS controls transition from proliferation to differentiation in the root. Cell 143:606–616
Wang J, Pan X, Han Y, Guo D, Guo Q, Li R (2012) Rosmarinic acid from eelgrass shows nematicidal and antibacterial activities against pine wood nematode and its carrying bacteria. Mar Drugs 10:2729–2740
Wang ZP, Xing HL, Dong L, Zhang HY, Han CY, Wang XC, Chen QJ (2015) Egg cell-specific promoter-controlled CRISPR/Cas9 efficiently generates homozygous mutants for multiple target genes in Arabidopsis in a single generation. Genome Biol 16:144
Wang GQ, Chen JF, Yi B, Tan HX, Zhang L, Chen WS (2017) HPPR encodes the hydroxyphenylpyruvate reductase required for the biosynthesis of hydrophilic phenolic acids in Salvia miltiorrhiza. Chin J Nat Med 15:917–927
Wang X, Wang Z, Zheng Z, Dong J, Song L, Sui L, Nussaume L, Desnos T, Liu D (2019) Genetic dissection of Fe-dependent signaling in root developmental responses to phosphate deficiency. Plant Physiol 179:300–316
Wang Y, Dai X, Xu G, Dai Z, Chen P, Zhang T, Zhang H (2021) The Ca2+-CaM signaling pathway mediates potassium uptake by regulating reactive oxygen species homeostasis in tobacco roots under low-K+ Stress. Front Plant Sci 12:658609
Williamson LC, Ribrioux SP, Fitter AH, Leyser HM (2001) Phosphate availability regulates root system architecture in Arabidopsis. Plant Physiol 126:875–882
Xu JJ, Fang X, Li CY, Zhao Q, Martin C, Chen XY, Yang L (2018) Characterization of Arabidopsis thaliana hydroxyphenylpyruvate reductases in the tyrosine conversion pathway. Front Plant Sci 9:1305
Xu JM, Wang ZQ, Wang JY, Li PF, Jin JF, Chen WW, Fan W, Kochian LV, Zheng SJ, Yang JL (2020) Low phosphate represses histone deacetylase complex1 to regulate root system architecture remodeling in Arabidopsis. New Phytol 225:1732–1745
Zheng Z, Wang Z, Wang X, Liu D (2019) Blue light-triggered chemical reactions underlie phosphate deficiency-induced inhibition of root elongation of Arabidopsis seedlings grown in petri dishes. Mol Plant 12:1515–1523
Zheng M, Zhu C, Yang T, Qian J, Hsu YF (2020) GSM2, a transaldolase, contributes to reactive oxygen species homeostasis in Arabidopsis. Plant Mol Biol 104:39–53
Zhou Z, Li J, Zhu C, Jing B, Shi K, Yu J, Hu Z (2022) Exogenous rosmarinic acid application enhances thermotolerance in tomatoes. Plants 11:1172
Funding
This work was supported by the Chongqing science and technology forestry project (YB 2023–4) and the Chongqing Municipal Education Commission for postgraduates innovation program (CYB21101).
Author information
Authors and Affiliations
Contributions
MZ and YFH designed the research. LYW, ML, HZ and XY performed the experiments. JQ and LYW analyzed the data. LYW, JQ, MZ and YFH wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by Li Tian.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, L., Qian, J., Li, M. et al. Arabidopsis PDE1 confers phosphate-deficiency tolerance in primary root growth. Plant Cell Rep 43, 8 (2024). https://doi.org/10.1007/s00299-023-03120-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00299-023-03120-8