Abstract
Key message
TaPGLP1, a chloroplast stromal 2-phosphoglycolate phosphatase of wheat, is an ATG8-interacting protein and undergoes autophagic degradation in starvation-treated wheat mesophyll protoplasts.
Abstract
Selective autophagy in plants has been shown to target diverse cellular cargoes including whole chloroplasts (Chlorophagy) and several chloroplast components (Piecemeal chlorophagy). Most cargoes of selective autophagy are captured by the autophagic machinery through their direct or indirect interactions with the autophagy-essential factor ATG8. Here, we reported a new ATG8-interacting cargo of piecemeal chlorophagy, the wheat photorespiratory 2-phosphoglycolate phosphatase TaPGLP1. The TaPGLP1-mCherry fusions expressed in wheat protoplasts located in the chloroplast stroma. Strikingly, these fusions are translocated into newly formed chloroplast surface protrusions after a long time incubation of protoplasts in a nutrition-free solution. Visualization of co-expressed TaPGLP1-mCherry and the autophagy marker GFP-TaATG8a revealed physical associations of TaPGLP1-mCherry-accumulating chloroplast protrusions with autophagic structures, implying the delivery of TaPGLP1-mCherry fusions from chloroplasts to the autophagic machinery. TaPGLP1-mCherry fusions were also detected in the GFP-TaATG8a-labelled autophagic bodies undergoing degradation in the vacuoles, which suggested the autophagic degradation of TaPGLP1. This autophagic degradation of TaPGLP1 was further demonstrated by the enhanced stability of TaPGLP1-mCherry in protoplasts with impaired autophagy. Expression of TaPGLP1-mCherry in protoplasts stimulated an enhanced autophagy level probably adopted by cells to degrade the over-produced TaPGLP1-mCherry fusions. Results from gene silencing assays showed the requirement of ATG2s and ATG7s in the autophagic degradation of TaPGLP1. Additionally, TaPGLP1 was shown to interact with ATG8 family members. Collectively, our data suggest that autophagy mediates the degradation of the chloroplast stromal protein TaPGLP1 in starvation-treated mesophyll protoplasts.
Similar content being viewed by others

Avoid common mistakes on your manuscript.
Introduction
Chloroplasts are plant-specific organelles providing carbohydrates and energy for cells through photosynthesis and CO2 fixation. An intrinsic degradation or turnover of sunlight-damaged chloroplast components is essential for the health of the photosynthesis apparatus. Chloroplast degradation also occurs in naturally senescent leaves. Chloroplast proteins account for 75–80% of total leaf nitrogen, and the chloroplastic ribulose-1,5-bisphosphate (RuBP) carboxylase/oxygenase (RubisCO) proteins account for 50% of total leaf soluble proteins (Mae and Ohira 1981; Masclaux et al. 2001). Chloroplast recycling in senescent leaves mobilizes nutrition and energy into developing tissues and storage organs (Mae and Ohira 1981; Masclaux-Daubresse et al. 2010). In addition, chloroplast degradation occurs under unfavorable conditions, enabling plants to survive stresses through maintaining nutrient supply and cellular homeostasis (Izumi and Nakamura 2018). Both entire chloroplasts and their partial components are targets of degradation, and the degradation of chloroplast components can be mediated by intra- and extra-chloroplast mechanisms. The reported intra-chloroplast mechanisms involve chloroplast-intrinsic proteases and chlorophyll-degrading enzymes (Kuai et al. 2018; Nishimura et al. 2017). The reported extra-chloroplast mechanisms involve the ubiquitin–proteasome pathway (Ling et al. 2019), the small lytic senescence-associated vacuoles (SAVs) (Martinez et al. 2008), the chloroplast vesiculation-containing vesicles (CVVs) (Wang and Blumwald 2014), or the process of autophagy (Izumi and Nakamura 2018).
Autophagy is a conserved pathway for material degradation and nutrient cycling in eukaryotic cells. In autophagy, cellular cargoes are engulfed into double-membraned vesicles termed autophagosomes and delivered as inner membrane-enclosed autophagic bodies into the vacuoles/lysosomes for breakdown (Li and Vierstra 2012). Many AuTophaGy-related (ATG) factors participate in the biogenesis of autophagosomes and the autophagosome-vacuole fusion process (Li and Vierstra 2012). According to selectivity, autophagy can be divided into two types, bulk autophagy and selective autophagy (Michaeli et al. 2016). Plant selective autophagy have been shown to target diverse substances including entire or portions of organelles such as peroxisome, chloroplast, mitochondrion, endoplasmic reticulum, ribosome, and proteasome (Marshall and Vierstra 2018; Bu et al. 2020).
The autophagic degradation of entire chloroplasts (Chlorophagy) or a portion of chloroplast components (Piecemeal chlorophagy) can be induced by senescence, starvation conditions of carbon deprival and darkness, and stresses of salt, pathogen infection and irradiation with UVB or strong visible light (Dong and Chen 2013; Izumi and Nakamura 2018). The precise mechanisms underlying the recognition and capturing of chloroplast cargoes by the autophagic machinery are not well understood. Previous studies revealed the ab initio formation of chloroplast-derived vesicle-like structures, the rubisco-containing bodies (RCBs), the ATI1-positive (ATI1-PS) bodies or the small starch granule-like (SSGL) bodies, for wrapping chloroplast cargoes and delivering them to the autophagic machinery (Izumi and Nakamura 2018). The ESCRT (Endosomal Sorting Complex Required for Transport) subunit CHMP1 is essential for the RCB-involving piecemeal chlorophagy. The ATG8-interacting protein ATI1 was proposed to function as a cargo receptor in the ATI1-PS body-involving piecemeal chlorophagy (Michaeli et al. 2014; Spitzer et al. 2015). So far several chloroplastic components have been identified as cargoes of piecemeal chlorophagy including the stromal protein RubisCO and the envelope protein MEX1 carried by the RCBs, the thylakoid protein NPQ4 and the thylakoid/envelope protein APE1 carried by the ATI1-PS bodies, and chloroplast starch granules and granule-bound starch synthase I carried by the SSGL bodies (Ishida et al. 2008; Wang et al. 2013; Michaeli et al. 2014; Spitzer et al. 2015). Piecemeal chlorophagy may not be limited to these identified cargoes. In fact, a number of other chloroplast components accumulate in chmp1 mutant plants or interact with ATI1 (Michaeli et al. 2014; Spitzer et al. 2015). Therefore, it remains to be elucidated whether there are any more chloroplast components which can be targeted by known or unknown pathways of piecemeal chlorophagy.
ATG8 is one of the core ATG factors decorating on both layers of autophagic membranes (Nakatogawa et al. 2007). In selective autophagy, the capturing of specific cargoes by the autophagic machinery is generally related to pairwise interactions between ATG8 and different ATG8-interacting proteins (Marshall and Vierstra 2018; Bu et al. 2020). So far, a number of ATG8-interacting proteins have been identified in plants (Bu et al. 2020; Marshall and Vierstra 2018). Some ATG8-interacting proteins serve as direct cargoes of selective autophagy such as the autophagy regulators ATG1 and SH3P2, the virulence factor βC1 of CLCuMuV (cotton leaf curl Multan virus), the replication initiator C1 of TLCYnV (tomato leaf curl Yunnan virus), the NO signaling pathway regulator GSNOR1, and the drought negative regulator COST1 (Suttangkakul et al. 2011; Zhuang et al. 2013; Haxim et al. 2017; Zhan et al. 2018; Bao et al. 2020; Li et al. 2020). There are also some ATG8-interacting proteins serving as autophagy receptors/adaptors tethering their associated cargoes to the autophagic machinery via interacting with ATG8 (Marshall and Vierstra 2018; Bu et al. 2020). Such autophagy receptors/adaptors identified in plants include ATI1 engaged in the autophagic capturing of chloroplast cargoes, RPN10 engaged in the autophagic capturing of proteasome cargoes, PEX1/6/10 engaged in the autophagic capturing of peroxisome cargoes, and ATI3, Rtn1/2 and Sec62 engaged in the autophagic capturing of endoplasmic reticulum cargoes (Michaeli et al. 2014; Marshall et al. 2015; Xie et al. 2016; Zhou et al. 2018; Hu et al. 2020; Zhang et al. 2020). For their binding with ATG8, a majority of known ATG8-interacting proteins rely on a conserved ATG8-interacting motif (AIM, also known as LC3 interacting region [LIR]) and a few rely on a ubiquitin-interacting motif (UIM) (Marshall and Vierstra 2018; Marshall et al. 2019; Bu et al. 2020). Large-scale screening for protein–protein interactions and the in silico identification of AIM/UIM in plant proteins have generated a large number of potential ATG8-interacting proteins functioning in diverse biological processes (Xie et al. 2016; Wang et al. 2018; Marshall et al. 2019; Zess et al. 2019).
In this study, we report the identification of a new ATG8-interacting protein, the chloroplastic 2-phosphoglycolate phosphatase TaPGLP1 of wheat (Triticum aestivum L.), and provide evidence to show that TaPGLP1 undergoes autophagic degradation in starvation-treated wheat mesophyll protoplasts.
Materials and methods
Wheat materials and growth conditions
Seedlings of the wheat cultivar Yangmai158 were planted in peat soil at 22 ℃ under a 16-h light/8-h darkness photoperiod. The leaves of two-leaf-stage seedlings were used for RNA extraction. For protoplast preparation, one-leaf stage seedlings were moved to a 24 h dark photoperiod for further growth. The second etiolated leaves of dark-treated seedlings at the two-leaf stage were used for protoplast preparation.
Sequence characterization of the wheat 2-phosphoglycolate phosphatases (PGLPs)
The cDNA and protein sequences of the three wheat PGLP-encoding genes, TraesCS2A02G3485 00, TraesCS2B02G366900, and TraesCS2D02G346900 were downloaded from the wheat genome database (plants.ensembl.org/Triticum_aestivum/). The protein sequences of Arabidopsis AtPGLP1 (At5g36700) and AtPGLP2 (At5g47760) were downloaded from the UniProt database (www.uniprot.org/). Multiple sequence alignment of protein sequences was performed in Bioedit 7.2.5. Information on protein subcellular localization was predicted with TargetP 2.0 (www.cbs.dtu.dk/services/TargerP/).
Plasmid constructs
The GFP, GFP-TaATG8a and GFP-TaATG8h-expressing vectors have been described previously (Pei et al. 2014). All primers used in this study were listed in Supplementary Table S1. Total RNA was extracted with Trizol (Invitrogen) and used for the synthesis of first-strand cDNA with the TIANScript II RT Kit (Tiangen). The coding sequence fragment of TaPGLP1 lacking the stop codon was PCR amplified from first-strand cDNA, and recombinated into the mCherry-expressing vector pAN583 by using the pEASY-Basic Seamless Cloning and Assembly Kit (TransGen), resulting in the vector for expressing TaPGLP1-mCherry.
Yeast two-hybrid (Y2H) vectors were constructed based on the vectors pGBKT7 and pGADT7, respectively, expressing yeast GAL4’s DNA binding domain (BD) and activation domain (AD). To construct the AD-TaPGLP1-expressing vector, the coding sequence fragment of TaPGLP1 was PCR amplified, digested with EcoRI and XhoI, and inserted into the pGADT7 vector. Previously, we have reported the identification of nine wheat ATG8 family genes (TaATG8a-8g) (Pei et al. 2014). To construct vectors expressing BD-TaATG8s, the ORF sequence of TaATG8h lacking the codon of the C-terminal exposed glycine (Gly), and those of TaATG8a and 8g lacking the codons of a series of C-terminal Gly-beginning residues were each amplified, digested by NcoI/PstI, and inserted into the vector pGBKT7. During autophagy, the C-terminal conserved Gly of ATG8s, either innately exposed or exposed after processing by the protease ATG4, is covalently linked to the lipid molecule phosphatidyl ethanolamine (PE) and thereby mediate the positioning of ATG8s to autophagic membranes (Nakatogawa et al. 2007; Pei et al. 2014). Since a nuclear localization of BD-TaATG8s was required in Y2H assays, the deletion of the C-terminal Gly of TaATG8s here was intended to avoid the possible positioning of BD-TaATG8s on autophagic membranes in the cytoplasm of yeast cells.
Bimolecular fluorescence complementation (BiFC) vectors were constructed based on the vectors pUC-SPYNE and pUC-SPYCE, respectively, expressing YFP N-terminal fragment (YN) and C-terminal fragment (YC) (Walter et al. 2004). To construct the vector expressing TaPGLP1-YN, the ORF sequence of TaPGLP1 lacking the stop codon was PCR amplified, digested by XbaI/KpnI, and inserted into the vector pUC-SPYNE. To construct vectors expressing TaATG8a/8g/8h-YC, the ORF sequences of TaATG8a, 8g, and 8h, with the same codon deletions to the sequences cloned in the Y2H vectors, were each amplified, digested by XbaI/SalI, and inserted into the vector pUC-SPYCE. Here, the deletion of the C-terminal Gly of TaATG8s was intended to avoid removal of the C-terminally fused YC fragment by the protease TaATG4s in protoplasts.
RNA interference (RNAi) vectors for silencing wheat ATG2s or ATG7s were constructed based on the hairpin RNA-expressing vector pWMB006 (Wang et al. 2017). Blast searching against the reference genome sequence of wheat (http://plants.ensembl.org/Triticum_aestivum) by using Arabidopsis AtATG2 (At3g19190) or AtATG7 (At5g45900) protein sequences as queries was performed to identify wheat ATG2- and ATG7-encoding genes. Then two fragments of sequence to be used as RNAi triggers, one 170 bp fragment and one 232 bp fragment, were, respectively, chosen from conserved regions of ATG2s and ATG7s with the standard that the trigger sequence contained more than one nucleotide stretches longer than 23 bp and 100% identical among homeologous genes. Each RNAi trigger was PCR amplified as two fragments, a BamHI-KpnI and a SacI-SpeI fragment, from the first-strand cDNA and inserted into the same pWMB006 vector in the sense orientation (BamHI-KpnI fragments) or antisense orientation (SacI-SpeI fragments) to form an arrangement of inverted repeats.
Preparation and transfection of wheat mesophyll protoplasts
About 30 mg hand-cut leaf strips (0.5–1 mm) were submerged in 1 ml of enzyme solution containing 20 mM MES at pH5.8, 20 mM KCl, 10 mM CaCl2, 0.1% BSA, 5 mM β-mercaptoethanol, 0.5 M mannitol, 1.5% cellulase R-10 (Yakult, Japan), and 0.5% macerozyme R-10 (Yakult, Japan), vacuum-infiltrated for 30 min and then digested at 28 °C in the dark with gentle shaking for 5 h. The enzyme solution with released mesophyll protoplasts were then mixed with 1 ml W5 solution (154 mM NaCl, 125 mM CaCl2, 5 mM KCl, 5 mM glucose, and 0.03% MES at pH 5.8), and protoplasts were filtered through 40 μm nylon meshes into 1.5 ml tubes. Protoplasts were collected after spinning at 100 g for 3 min, washed once with 1 ml W5 solution and resuspended in the MMG solution (15 mM MgCl2, 0.5 M mannitol, and 0.1% MES at pH 5.6) at a concentration of 2 × 105 cells mL−1.
Endotoxin-free plasmids were extracted with a NucleoBond Xtra Midi EF kit (Macherey-Nagel) and adjusted to a concentration of 1 μg/μl. Transfection of plasmids into protoplasts was conducted with the PEG-Ca2+-mediated transfection method (Yoo et al. 2007). Briefly, 100 μl protoplasts were mixed with 10 μl plasmids for single-vector transfection, or mixed with 20 μl plasmids containing 10 μl of each of the two vectors for two-vector transfection. Transfection was started by adding an equal volume of freshly prepared PEG-Ca2+ solution (40% PEG4000, 0.4 M mannitol, and 100 mM CaCl2) into the above mixture. After 15 min transfection at room temperature in the dark, 440 μl W5 solution was added and mixed thoroughly with the transfection mixture to stop transfection. Transfected protoplasts were pelleted by spinning at 100 g for 3 min, washed once with the W5 solution and suspended in 1 ml nutrient-free WI solution (0.5 M mannitol, 20 mM KCl, and 4 mM MES at pH 5.6). For treatment with chemicals, 100 μM E-64d or 5 mM 3-Methyladenine (3-MA) was added into the WI solution immediately after transfected protoplasts were suspended in it. An equivalent volume of the solvent, water, was added to the control of the 3-MA treatments.
Fluorescence microscopy
The fluorescence of GFP and mCherry, and the autofluorescence of chloroplasts in transfected protoplasts were visualized under a laser scanning confocal microscope (Nikon ECLIPSE Ti2).
RNAi assays
Protoplasts were co-transfected with the RNAi vector for silencing ATG2s (or that for silencing ATG7s) and the GFP-expressing vector. Control protoplasts were co-transfected with the empty RNAi vector and the GFP-expressing vector. Total RNA of protoplasts was extracted with Trizol (Invitrogen) and treated with RNase-free DNase. The first-strand cDNA was synthesized with the TIANScriptII RT Kit (Tiangen). The expression levels of RNAi target genes were assayed by quantitative real-time PCR (qRT-PCR) with gene-specific primers (Supplementary Table S1). Results were normalized to the reference genes β-tubulin or GFP using the 2−ΔΔCt method (Livak and Schmittgen 2001).
Stability analysis of TaPGLP1
The stability of TaPGLP1 was quantified by measuring the size of chloroplast protrusions with accumulated TaPGLP1-mCherry fluorescence. Protoplasts expressing TaPGLP1-mCherry were photographed under a laser scanning confocal microscope. The size of chloroplast protrusion was measured from at least 20 chloroplast protrusions for each treatment with Image J 1.46r and shown as mean ± SD from three independent experiments.
Evaluation of autophagy activity
The activity of autophagy was evaluated by quantifying the autophagic structures. The number of GFP-TaATG8a- or GFP-TaATG8h-labelled autophagic structures per protoplast was counted from 30 protoplasts for each treatment and expressed as mean ± SD from three independent experiments.
Y2H assays
Y2H assays were carried out in cells of yeast (Saccharomyces cerevisiae) strain YH109. Pairwise gene combinations in pGADT7 and pGBKT7 (or empty vector controls) were co-transformed into yeast cells with the Yeastmaker Yeast Transformation System 2 (Takara). Serial dilutions of yeast transformant cultures were plated on synthetically defined (SD) medium lacking leucine, tryptophan, and histidine (SD-Trp-Leu-His) and on SD medium lacking leucine, tryptophan, histidine, and adenine (SD-Trp-Leu-His-Ade) and incubated at 29 ℃ in the dark. Protein–protein interactions were identified by yeast growth on SD-Trp-Leu-His and strong interactions were identified by yeast growth on SD-Trp-Leu-His-Ade.
BiFC assays
Protoplasts were co-transfected with pairwise combinations of YN (or YN fusions)-expressing vectors and YC (or YC fusions)-expressing vectors. Transfected protoplasts suspended in the WI solution were incubated in the dark at room temperature. Protein–protein interactions were identified by visualization of the fluorescence emitted by reconstructed YFP in transfected protoplasts under a laser scanning confocal microscope.
Statistical analysis
All experiments were repeated three times with similar results. Quantitative data were statistically analyzed using Student’s t test (P < 0.05) using the IBM SPSS19.0 software package.
Results
Identification of the 2-phosphoglycolate phosphatase gene TaPGLP1 in wheat
To discover previously unknown ATG8-interacting proteins, we used TaATG8a, a member of the wheat ATG8 family (Pei et al. 2014), as the bait to screen a prey cDNA library of the wheat cultivar Yangmai158 through the Y2H approach. Among the positive cDNA clones detected, one encoded 2-phosphoglycolate phosphatase PGLP, the first enzyme of the plant photorespiration pathway. Arabidopsis has two PGLP (AtPGLP1 and AtPGLP2) (Schwarte et al. 2007) and this wheat PGLP was more similar to AtPGLP1 (Fig. 1a), so it was named TaPGLP1. According to the reference genome sequence of the wheat cultivar Chinese Spring, TaPGLP1 in hexaploid wheat includes three homeologous members of TaPGLP1-2A, -2B, and -2D, respectively, from chromosomes 2A, 2B, and 2D. The sequences of the three TaPGLP1s, 359–362 amino acids in length, have 98% similarity, and they share a very high similarity of 84% with Arabidopsis AtPGLP1 (Fig. 1a). Varied residues between TaPGLP1s and AtPGLP1 occur predominantly in the N-terminal 72 residues (positions relative to TaPGLP1-2A) (Fig. 1a). The full-length coding sequence of TaPGLP1-2D was then amplified from Yangmai158 cDNA, which encoded an identical protein sequence to the TaPGLP1-2D from Chinese Spring (Fig. 1 a). This TaPGLP1-2D of Yangmai158 was used for further analysis as a representative of wheat TaPGLP1s, and hereafter was referred to as TaPGLP1 for short.
Characterization of wheat TaPGLP1. a Sequence alignment of plant PGLPs. The aligned sequences included two proteins (AtPGLP1 and AtPGLP2) from Arabidopsis, three (TaPGLP1-2A(CS), -2B(CS), and -2D(CS)) from the wheat cultivar Chinese Spring, and one (TaPGLP1-2D(Y158)) from the wheat cultivar Yangmai 158. b Subcellular localization of TaPGLP1. Protoplasts were transfected with mCherry or TaPGLP1-mCherry. The fluorescence of mCherry and the autofluorescence of chloroplasts were visualized at 24 h after transfection. Scale bar 10 μm
Arabidopsis AtPGLP1 localizes in chloroplasts (Schwarte et al. 2007). Here, TaPGLP1-mCherry was expressed in wheat protoplasts to determine if TaPGLP1 has a similar subcellular localization. At 24 h after transfection (HAT) of protoplasts, TaPGLP1-mCherry fluorescence was observed spreading in chloroplasts and completely overlapping with the chloroplast autofluorescence (Fig. 1b), indicating the localization of TaPGLP1 in the chloroplast stroma. The N-terminal fragment (55 residues) of TaPGLP1, which has been shown to cover most of the varied sites between TaPGLP1s and AtPGLP1, was predicted to be a chloroplast transit peptide by TargetP 2.0.
TaPGLP1 gradually translocates into chloroplast protrusions
In addition to spreading in the chloroplast stroma, TaPGLP1-mCherry fluorescence was also observed defining the boundary of the chloroplast autofluorescence in some protoplasts at 24 HAT (Fig. 2), implying a translocation behavior of TaPGLP1-mCherry fusions. We thus monitored TaPGLP1-mCherry fluorescence on a long time scale. At 48 HAT, the TaPGLP1-mCherry fluorescence either gathered more densely at the chloroplast boundaries or even concentrated in the chloroplast surface protrusions (Fig. 2). These results suggested that, when protoplasts were subjected to a long time of incubation, TaPGLP1-mCherry fusions could gradually translocate to the peripheries of chloroplasts and finally gather in the chloroplast protrusions.
TaPGLP1 undergoes autophagic degradation
In the reported pathways of piecemeal chlorophagy, a portion of the chloroplast stroma containing autophagic cargoes protrudes outward into the cytoplasm before these cargoes are delivered to the autophagic machinery (Yamane et al. 2012; Izumi et al. 2019). Since similar chloroplast protrusions containing accumulated TaPGLP1-mCherry fusions were detected here, we then examined whether TaPGLP1 underwent autophagic degradation. To achieve this, TaPGLP1-mCherry and GFP-TaATG8a were co-expressed in protoplasts and their fluorescence signals were examined in detail. The GFP/RFP-tagged forms of ATG8 have been widely used to indicate the occurrence of autophagic structures (Li et al. 2018). E-64d, an inhibitor of vacuolar proteases, was used to inhibit the degradation of autophagic bodies and thus to accumulate autophagic structures (Wang et al. 2013). As shown in Fig. 3a, GFP-TaATG8a-labelled autophagic structures, as well as chloroplasts protrusions with accumulated TaPGLP1-mCherry fusions, was clearly observed in the transfected protoplasts. More importantly, some chloroplast protrusions with accumulated TaPGLP1-mCherry were detected to co-localize with spot-shaped autophagic structures (Fig. 3a, arrowheads) or to be surrounded by cup- or ring-shaped autophagic structures (Fig. 3a, arrows). Such physical associations implied a delivery of TaPGLP1-mCherry fusions from chloroplast protrusions to the autophagic machinery. The previously described wrapping of whole chloroplasts by the autophagic structures was also detected (Fig. 3a, double arrowheads) (Izumi et al. 2017).
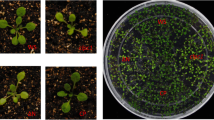
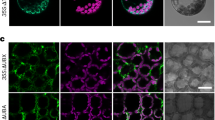
TaPGLP1 is subjected to autophagic degradation. a Chloroplast protrusions with accumulated TaPGLP1-mCherry fusions physically associate with autophagic structures. The autophagic structures were labeled by GFP-TaATG8a. The chloroplast protrusions were indicated by the TaPGLP1-mCherry fluorescence accumulated in them. Arrows indicate the chloroplast protrusions surrounded by cup- or ring-shaped autophagic structures. Arrowheads indicate the chloroplast protrusions overlapping with spot-shaped autophagic structures. Double arrowheads indicate the whole chloroplasts surrounded by cup-shaped autophagic structures. b TaPGLP1-mCherry fusions are delivered to the vacuole through the autophagy pathway. Arrowheads indicate the co-localized GFP-TaATG8a fluorescence and TaPGLP1-mCherry fluorescence on small round bodies in the vacuole. Protoplasts were co-transfected with the GFP-TaATG8a-expressing vector and the TaPGLP1-mCherry-expressing vector. Transfected protoplasts were suspended in the nutrient-free WI solution with 100 μM E-64d and incubated at 25 ℃ in darkness. The fluorescence of GFP and mCherry and the autofluorescence of chloroplasts were visualized at the indicated time points after transfection. Scale bar 10 μm
In the process of autophagy, the autophagosomes with enclosed cargoes were destined to the vacuoles for degradation. Here, co-localized GFP-TaATG8a fluorescence and TaPGLP1-mCherry fluorescence were seen on small round bodies undergoing the random motion characteristic of Brownian movement in the vacuoles, resembling autophagic bodies undergoing degradation in the vacuoles (Fig. 3b). TaATG8h is another member of the multiprotein ATG8 family of wheat (Pei et al. 2014). We also co-expressed GFP-TaATG8h and TaPGLP1-mCherry in protoplasts and observed similar vacuolar bodies with co-localized GFP-TaATG8h fluorescence and TaPGLP1-mCherry fluorescence (Supplementary Fig. S1). Most of such bodies showed weak fluorescence signals probably due to partial degradation of GFP/mCherry fusions by vacuolar proteases. These results indicated that TaPGLP1-mCherry fusions, after accumulating in chloroplast protrusions, were delivered into the vacuoles for degradation by the autophagy machinery.
3-MA treatment results in impaired autophagy and enhanced stability of TaPGLP1
The treatment with E-64d seemed to produce enlarged chloroplast protrusions with over-accumulated TaPGLP1-mCherry fluorescence at 48 HAT (Fig. 3a). This implied that disturbing autophagy with E-64d interfered with the autophagic degradation of TaPGLP1-mCherry. E-64d does not inhibit the upstream steps of the autophagy process and thus may not be optimal for addressing the process of cargo capturing into autophagic structures. For this reason, we further used 3-MA, a potent autophagy inhibitor, to treat protoplasts and whereby to investigate the stability of TaPGLP1 (Li et al. 2018). 3-MA inhibits an early stage of autophagy by inhibiting the class III phosphoinositide 3-kinase, an essential regulator of autophagy (Wang et al. 2013). When 5 mM 3-MA was applied to protoplasts in which GFP-TaATG8a or GFP-TaATG8h was expressed to label autophagic structures, impaired autophagy was detected at 24 and 48 HAT as evidenced by the significantly reduced number of autophagic structures (Fig. 4a, b). Moreover, the application of 5 mM 3-MA to protoplasts expressing TaPGLP1-mCherry resulted in enlarged chloroplast protrusions with over-accumulated red fluorescence (Fig. 4c, d), suggesting an enhanced stability of TaPGLP1-mCherry probably caused by the impairing of autophagy.
3-MA treatment results in impaired autophagy and enhanced stability of TaPGLP1. a Autophagy activity in protoplasts revealed by the quantity of autophagic structures. Protoplasts were transfected with the GFP-TaATG8a-expressing vector or the GFP-TaATG8h-expressing vector. Transfected protoplasts were suspended in the nutrient-free WI solution with or without 5 mM 3-MA (3-MA or Mock) and incubated at 25 ℃ in darkness. The green spots representing autophagic structures labelled by GFP-TaATG8a or GFP-TaATG8h were visualized at the indicated time points after transfection. Scale bar 10 μm. b Statistics of the autophagic structures shown in (a). The number of autophagic structures per protoplast was counted from 30 protoplasts for each treatment and shown as mean ± SD (standard deviation) from three independent experiments. Asterisks indicate a significant difference (P < 0.05, Student’s t test). c Stability of TaPGLP1 revealed by the size of chloroplast protrusions with accumulated TaPGLP1-mCherry fusions. Protoplasts were transfected with the TaPGLP1-mCherry-expressing vector. Transfected protoplasts were suspended in the WI solution with or without 5 mM 3-MA (3-MA or Mock) and incubated at 25 ℃ in darkness. The chloroplast protrusions with TaPGLP1-mCherry fluorescence and the autofluorescence of chloroplasts were visualized at 48 h after transfection. Scale bar 10 μm. d Measurement and statistics of the chloroplast protrusion size shown in (c). Protrusion size was measured from at least 20 protrusions for each treatment with Image J 1.46r and shown as mean ± SD from three independent experiments. Asterisks indicate a significant difference (P < 0.05, Student’s t test)
ATG2 and ATG7 are required in the autophagic degradation of TaPGLP1
A number of ATG proteins are involved in the biogenesis of autophagy (Li and Vierstra 2012). ATG2, together with ATG18 and ATG9, functions in the delivery of lipids to the expanding autophagic membranes (Li and Vierstra 2012). ATG7 is a ubiquitin-activating enzyme (E1)-like protein, which is essential for two ubiquitination-like reactions whereby ATG8 is connected to the lipid phosphatidylethanolamine (PE) and thus targeted to the surfaces of autophagic membranes (Li and Vierstra 2012). Arabidopsis AtATG2 and AtATG7 are both single-copy genes, and mutations in them have been reported rending efficient blocking of autophapy (Inoue et al. 2006; Chung et al. 2010; Wang et al. 2011). Here, we indentified the ATG2- and ATG7-encoding genes in the wheat genome and investigated their requirement in the autophagic degradation of TaPGLP1 through gene silencing. The hexaploid wheat genome possesses a family of six ATG2s (TaATG2s) and a family of three ATG7s (TaATG7s). The six TaATG2s can be divided into two groups with group 1 containing three homeologous genes of TaATG2-6A (TraesCS6A02G169400), TaATG2-6B (TraesCS6B02G197400), and TaATG2-6D (TraesCS6D02G159300) and group 2 containing another set of three homeologous genes of TaATG2-7A (TraesCS7A02G208300), TaATG2-7B (TraesCS7B02G115500), and TaATG2-7D (TraesCS7D02G210700) (Supplementary Fig. S2). The group 1 TaATG2s encode peptide sequences of 1917–1948 amino acid (a.a.) sharing 94–97% similarity with each other and 52–53% similarity with the sequence of Arabidopsis AtATG2 (Supplementary Fig. S2). The group 2 TaATG2s encode peptide sequences of 1932–1933 a.a. sharing 99% similarity with each other, 81–83% similarity with the members of group 1, and 55% similarity with the sequence of Arabidopsis AtATG2 (Supplementary Fig. S2). The three TaATG7s are homeologous genes of TaATG7-3A (TraesCS3A02G220600), TaATG7-3B (TraesCS3B02G250900), and TaATG7-3D (TraesCS3D02G231900), and they encode peptide sequences of 1020–1021 a.a. sharing 98% similarity with each other and 65% similarity with the sequence of Arabidopsis AtATG7 (Supplementary Fig. S3). The reason for the longer sequence size of TaATG7s than AtATG7 is that TaATG7s have two repeated E1-representing ThiF domains (ThiF1 and ThiF2) while AtATG7 has only one (Supplementary Fig. S3). The expression of All TaATG2s and TaATG7s was verified by the existence of corresponding mRNA sequences in the dbEST database of Genbank.
RNAi-based gene silencing in protoplasts was conducted by expressing one hairpin RNA targeting a conserved region of all three homeologous TaATG7s or another one targeting a conserved region of all three homeologous group 1 TaATG2s. The reason for only group 1 TaATG2s were selected as target genes of RNAi is that silencing of all six TaATG2s by expressing one hairpin RNA might be unfeasible due to the nucleotide sequence divergence between the two groups of TaATG2s. qRT-PCR assays with common primers for target genes of RNAi showed that the total transcript levels of TaATG7s or group 1 TaATG2s at 24 HAT, relative to whether the endogeneous tubulin gene or the exogeneous GFP gene, were significantly lower in the protoplasts expressing a cognate hairpin RNA than in the protoplasts expressing the empty vector (Supplementary Fig. S4). This suggested the feasibility of silencing TaATG7s or group 1 TaATG2s in protoplasts by RNAi. To determine the autophagy activity in protoplasts subjected to gene silencing, GFP-TaATG8a was co-expressed with either of the two hairpin RNAs and the activity of autophagy was evaluated by quantifying the GFP-TaATG8a-labelled autophagic structures per protoplast. Control protoplasts were those co-expressing GFP-TaATG8a and the empty RNAi vector. The results showed that silencing of either of these two sets of genes resulted in significantly reduced activity of autophagy at 24 and 48 HAT just like the effect of 3-MA (Fig. 5a, b). We then co-expressed either of the two hairpin RNAs with TaPGLP1-mCherry in protoplasts to determine if silencing of TaATG7s or group 1 TaATG2s can affect the autophagic degradation of TaPGLP1. As shown in Fig. 5c, d, silencing of either of these two sets of genes resulted in enhanced stability of TaPGLP1-mCherry manifested by the enlarged chloroplast protrusions with over-accumulated TaPGLP1-mCherry fluorescence. These results, together with the findings on the 3-MA-treated protoplasts, confirm the occurrence of autophagic degradation of TaPGLP1 in protoplasts and demonstrated the requirement of TaATG7s and group 1 TaATG2s in this autophagic degradation process.
ATG2 and ATG7 are required in the autophagic degradation of TaPGLP1. a Autophagy activity in protoplasts revealed by the quantity of autophagic structures. Protoplasts were co-transfected with the GFP-TaATG8a-expressing vector and either of the two RNAi vectors for silencing group1 TaATG2s (ATG2s-RNAi) or TaATG7s (ATG7s-RNAi). Control protoplasts were co-transfected with the GFP-TaATG8a-expressing vector and the empty RNAi vector (EV). Transfected protoplasts were suspended in the WI solution and incubated at 25 ℃ in darkness. The green spots representing autophagic structures labelled by GFP-TaATG8a were visualized at the indicated time points after transfection. Scale bar 10 μm. b Statistics of the autophagic structures shown in (a). The number of autophagic structures per protoplast was counted from 30 protoplasts for each treatment and shown as mean ± SD from three independent experiments. Asterisks indicate a significant difference (P < 0.05, Student’s t test). c Stability of TaPGLP1 revealed by the size of chloroplast protrusions with accumulated TaPGLP1-mCherry fusions. Protoplasts were co-transfected with the TaPGLP1-mCherry-expressing vector and either of the two RNAi vectors for silencing group1 TaATG2s (ATG2s-RNAi) or TaATG7s (ATG7s-RNAi). Control protoplasts were co-transfected with the TaPGLP1-mCherry-expressing vector and the empty RNAi vector (EV). Transfected protoplasts were suspended in the WI solution and incubated at 25 ℃ in darkness. The chloroplast protrusions with TaPGLP1-mCherry fluorescence and the autofluorescence of chloroplasts were visualized at 48 h after transfection. Scale bar 10 μm. d Measurement and statistics of the chloroplast protrusion size shown in (c). Protrusion size was measured from at least 20 protrusions for each treatment with Image J 1.46r and shown as mean ± SD from three independent experiments. Asterisks indicate a significant difference (P < 0.05, Student’s t test)
Over-expression of TaPGLP1 enhanced the autophagy activity
In view of the autophagic degradation of TaPGLP1, we hypothesized that enhanced autophagy activity may be required to degrade over-produced TaPGLP1-mCherry fusions in protoplasts. To prove this, we compared the autophagy activity in protoplasts expressing TaPGLP1-mCherry with that in protoplasts expressing mCherry or expressing neither TaPGLP1-mCherry nor mCherry. For evaluation of autophagy activity, GFP-TaATG8a was co-expressed in these protoplasts to label autophagic structures. Compared with the expression of GFP-TaATG8a alone and the co-expression of mCherry with GFP-TaATG8a, the co-expression of TaPGLP1-mCherry with GFP-TaATG8a resulted in an increased number of GFP-TaATG8a-labelled autophagic structures per protoplast at 24 and 48 HAT representing a response of enhanced autophagy activity to over-produced TaPGLP1-mCherry fusions (Fig. 6a, b).
Over-expression of TaPGLP1 enhances the autophagy activity in protoplasts. a Autophagy activity in protoplasts revealed by the quantity of autophagic structures. Protoplasts were transfected with mCherry (mCherry), TaPGLP1-mCherry (TaPGLP1-mCherry), or none of them (-). GFP-TaATG8a was co-expressed in these protoplasts to label autophagic structures. Transfected protoplasts were maintained in the nutrient-free WI solution and incubated at 25 ℃ in darkness. The fluorescence of mCherry and the fluorescence of autophagic structures labelled by GFP-TaATG8a were visualized at the indicated time points after transfection. Scale bar 10 μm. b Quantification of autophagic structures in protoplasts shown in (a). The number of autophagic structures per protoplast was counted from 30 protoplasts. Data are presented as mean ± SD of three biological replicates. Asterisks indicate a significant difference (P < 0.05, Student’s t test)
TaPGLP1 interacts with wheat ATG8 family members
Since TaPGLP1 was initially identified as a TaATG8a-interacting protein through Y2H screening of a wheat cDNA library, we conducted pairwise Y2H and BiFC assays to confirm this interaction. The wheat ATG8 family comprises of three groups of members, and TaATG8a, 8g, and 8h are members of group 1, group 2, and group 3, respectively (Pei et al. 2014). Here, TaATG8g and 8h were also included in these protein–protein interaction assays. In the Y2H assays, serial dilutions of yeast cultures expressing pairwise combinations of AD (or AD fusions) and BD (or BD fusions) were plated on different selective media to detect activation of the reporter genes, HIS3 and ADE2. Yeast transformants co-expressing AD-TaPGLP1 and BD-TaATG8a/8g/8h were able to grow on SD-Trp-Leu-His selection plates, while control yeast transformants co-expressing AD and BD, AD and BD-TaATG8a/8g/8h, or AD-TaPGLP1 and BD were unable to proliferate (Fig. 7a). In the BiFC assays, no signals of reconstructed YFP fluorescence were detected in one group of control protoplasts expressing YN and YC, and diffused signals, possibly due to non-specific interactions, were detected in the other group of control protoplasts expressing YN and TaATG8a/8g/8h-YC (Fig. 7b, Controls). Although such non-specific interactions may occur, specific interactions between TaPGLP1 and TaATG8a/8g/8h could be interpreted from the distinct punctate fluorescence of reconstructed YFP in the cytoplasm of protoplasts co-expressing TaPGLP1-YN and TaATG8a/8g/8h-YC (Fig. 7b). These results suggested that TaPGLP1 interacts with wheat ATG8 family members TaATG8a, 8g, and 8h.
TaPGLP1 interacts with wheat ATG8 family members. a Pairwise Y2H assays on the interaction of TaPGLP1 with TaATG8a/8g/8h. Serial dilutions of yeast transformant cultures expressing pairwise combinations of AD (or AD fusions) and BD (or BD fusions) were plated on the medium lacking tryptophan and leucine (SD-Trp-Leu) and on the selective media lacking leucine, tryptophan, and histidine (SD-Trp-Leu-His) or lacking leucine, tryptophan, histidine, and adenine (SD-Trp-Leu-His-Ade). b Pairwise BiFC assays on the interaction of TaPGLP1 with TaATG8a/8g/8h. Protoplasts were co-transfected with pairwise combinations of YN (or YN fusions)-expressing and YC (or YC fusions)-expressing vectors. Reconstructed YFP fluorescence was visualized at 24 h after transfection. Arrowheads indicate the fluorescence spots of reconstructed YFP in protoplasts co-expressing TaPGLP1-YN and TaATG8a/8g/8h-YC. Scale bar 10 μm. c Quantification of the fluorescence spots shown in (b) in protoplast cells co-expressing TaPGLP1-YN and TaATG8a/8g/8h-YC. The number of separated fluorescence spots per protoplast was counted from 30 protoplasts. Data are presented as mean ± SD of three biological replicates. Values not sharing a common letter are significantly different (P < 0.05, Student’s t test)
Differential interaction strength was also revealed among the pairwise TaPGLP1-TaATG8 interactions. In the Y2H assays, the TaPGLP1-TaATG8h interaction enabled the most vigorous growth, and the TaPGLP1-TaATG8g interaction enabled the less vigorous growth of yeast on the selection medium SD-Trp-Leu-His (Fig. 7a). Only the TaPGLP1-TaATG8h interaction supported the growth of yeast on the more stringent selection medium SD-Trp-Leu-His-Ade (Fig. 7a). Consistently, the TaPGLP1-TaATG8h interaction produced the largest number of, and the TaPGLP1-TaATG8g interaction produced the smallest number of reconstructed YFP fluorescence spots per protoplast in the BiFC assays (Fig. 7b, c). The YFP fluorescence spots produced by the TaPGLP1-TaATG8h interaction were very large and bright compared with those produced by the TaPGLP1-TaATG8a and TaPGLP1-TaATG8g interactions (Fig. 7b). These results indicated that TaATG8h is the strongest interacting partner of TaPGLP1, followed by TaATG8a and TaATG8g.
Discussion
Plant photorespiration is a light-dependent and O2-consuming catabolic pathway occurring simultaneously with photosynthesis. The photosynthetic carbon fixation starts with the RubisCO-catalyzed carboxylation of the primary CO2 acceptor RuBP and the decomposition of the intermediate product. This reaction produces two molecules of 3-phosphoglycerate (3PGA) which enter the Calvin–Benson cycle (CBC) for the synthesis of complex carbohydrates. The enzyme RubisCO also catalyzes the oxidation of RuBP to produce one molecule of 3PGA and one molecule of 2-phosphoglycolate (2PG) particularly under conditions of high O2 and low CO2 concentrations. Since 2PG is not only a non-CBC metabolite representing withdrawal of metabolites from CBC but also an inhibitor of many CBC enzymes, the formation of 2PG severely counteracts CBC of carbon fixation (Eisenhut et al. 2019). To deal with such situations, plants adopt the multistep photorespiration pathway to metabolize 2PG through recycling it into the CBC intermediate 3PGA (Eisenhut et al. 2019). The chloroplastic 2-phosphoglycolate phosphatase PGLP catalyzes the first-step reaction of photorespiration, the conversion of 2PG into glycolate (Eisenhut et al. 2019). In this study, we identified a wheat PGLP-encoding gene, TaPGLP1. TaPGLP1 located in the chloroplast stroma, and its sequence shares a high similarity with Arabidopsis AtPGLP1 which has a known function in photorespiration (Schwarte et al. 2007). Such evidence points to a role of TaPGLP1 in wheat photorespiration.
Previously, several chloroplast components including RubisCO, MEX1, NPQ4, APE1, and starch granules have been shown to undergo autophagic degradation via piecemeal chlorophagy (Izumi et al. 2019). They are exported from chloroplasts by distinct chloroplast-derived vesicles of RCBs, ATI1-PS bodies or SSGL bodies and subsequently captured by the autophagic machinery (Ishida et al. 2008; Wada et al. 2009; Wang et al. 2013; Michaeli et al. 2014; Izumi et al. 2015). Here, we reported that wheat TaPGLP1 is also a cargo of piecemeal chlorophagy in starvation-treated mesophyll protoplasts. The sorting of the TaPGLP1-mCherry fusions into the chloroplast protrusions resembled the first step of those known processes of piecemeal chlorophagy (Yamane et al. 2012; Izumi et al. 2019). The previously described physical associations of autophagic structures with chloroplast extensions were detected at the chloroplast protrusions with accumulated TaPGLP1-mCherry fusions (Spitzer et al. 2015). This reflected a delivery of TaPGLP1-mCherry fusions from chloroplasts to the autophagic machinery. The autophagic degradation of TaPGLP1 was confirmed by the observation of autophagic bodies containing TaPGLP1-mCherry fusions in the vacuoles and by the detection of enhanced stability of TaPGLP1-mCherry in protoplasts with impaired autophagy. Expression of TaPGLP1-mCherry in protoplasts stimulated an enhanced autophagy level probably adopted by cells to degrade the over-produced TaPGLP1-mCherry fusions. In addition, results from gene silencing assays suggest the requirement of ATG2s and ATG7s in the autophagic degradation of TaPGLP1.
The temporarily-formed chloroplast extensions such as protrusions or protrusion-derived stromules seem to be common collecting and distributing centers for the cargoes of piecemeal chlorophagy (Yamane et al. 2012; Izumi et al. 2019). The RCBs and SSGL bodies were thought to be formed as a result of the segmentation or tip shedding of protrusion-derived stromules containing autophagic cargoes (Ishida et al. 2008; Wang et al. 2013). The ATI1-PS bodies form in the stroma and bud from chloroplasts with enclosed autophagic cargoes (Michaeli et al. 2014). Since TaPGLP1-mCherry fusions showed similar accumulation in the chloroplast protrusions before they were delivered to the autophagic machinery, it cannot be excluded that TaPGLP1 is degraded via the known piecemeal chlorophagy processes. Both of the RCBs and ATI1-PS bodies have been proposed to contain a mix of chloroplast components with unidentified autophagic cargoes (Michaeli et al. 2014; Spitzer et al. 2015). To clearly clarify the relationship between the autophagic degradation process of TaPGLP1 and those known processes of piecemeal chlorophagy, further studies are needed to identify whether any chloroplast-derived vesicles are responsible for the delivery of TaPGLP1 proteins and whether these vesicles are per se RCBs or ATI1-PS bodies.
In selective autophagy, most reported cargoes are recognized by the autophagic machinery through their direct interactions with the autophagic membrane-targeting ATG8 or through receptors/adaptors bridging their interactions with ATG8 (Marshall and Vierstra 2018; Bu et al. 2020). The ATG8-interacting protein ATI1 is an organizer of ATI1-PS bodies, picking out its interacting chloroplast proteins and mediating their targeting by the autophagic machinery (Michaeli et al. 2014). Whether any specific ATG8-interacting proteins/cargo receptors function in RCB- and SSGL body-mediated piecemeal chlorophagy is unclear (Ishida et al. 2008; Wang et al. 2013). In our study, TaPGLP1 was shown to directly interact with wheat ATG8 family members, TaATG8a, 8g, and 8h. These interactions may contribute to the capturing of TaPGLP1 proteins by the autophagy machinery.
References
Bao Y, Song WM, Wang P, Yu X, Li B, Jiang C, Shiu SH, Zhang H, Bassham DC (2020) COST1 regulates autophagy to control plant drought tolerance. Proc Natl Acad Sci U S A 117(13):7482–7493
Bu F, Yang M, Guo X, Huang W, Chen L (2020) Multiple functions of ATG8 family proteins in plant autophagy. Front Cell Dev Biol 8:466
Chung T, Phillips AR, Vierstra RD (2010) ATG8 lipidation and ATG8-mediated autophagy in Arabidopsis require ATG12 expressed from the differentially controlled ATG12A and ATG12B loci. Plant J 62(3):483–893
Dong J, Chen W (2013) The role of autophagy in chloroplast degradation and chlorophagy in immune defenses during Pst DC3000 (AvrRps4) infection. PLoS One 8(8):e73091.
Eisenhut M, Roell MS, Weber APM (2019) Mechanistic understanding of photorespiration paves the way to a new green revolution. New Phytol 223(4):1762–1769
Haxim Y, Ismayil A, Jia Q, Wang Y, Zheng X, Chen T, Qian L, Liu N, Wang Y, Han S, Cheng J, Qi Y, Hong Y, Liu Y (2017) Autophagy functions as an antiviral mechanism against geminiviruses in plants. Elife 6:e23897.
Hu S, Ye H, Cui Y, Jiang L (2020) AtSec62 is critical for plant development and is involved in ER-phagy in Arabidopsis thaliana. J Integr Plant Biol 62(2):181–200
Inoue Y, Suzuki T, Hattori M, Yoshimoto K, Ohsumi Y, Moriyasu Y (2006) AtATG genes, homologs of yeast autophagy genes, are involved in constitutive autophagy in Arabidopsis root tip cells. Plant Cell Physiol 47(12):1641–1652
Ishida H, Yoshimoto K, Izumi M, Reisen D, Yano Y, Makino A, Ohsumi Y, Hanson MR, Mae T (2008) Mobilization of rubisco and stroma-localized fluorescent proteins of chloroplasts to the vacuole by an ATG gene-dependent autophagic process. Plant Physiol 148(1):142–155
Izumi M, Nakamura S (2018) Chloroplast protein turnover: the influence of extraplastidic processes Including Autophagy. Int J Mol Sci 19(3):828
Izumi M, Hidema J, Wada S, Kondo E, Kurusu T, Kuchitsu K, Makino A, Ishida H (2015) Establishment of monitoring methods for autophagy in rice reveals autophagic recycling of chloroplasts and root plastids during energy limitation. Plant Physiol 167(4):1307–1320
Izumi M, Ishida H, Nakamura S, Hidema J (2017) Entire photodamaged chloroplasts are transported to the central vacuole by autophagy. Plant Cell 29:377–394
Izumi M, Nakamura S, Li N (2019) Autophagic turnover of chloroplasts: its roles and regulatory mechanisms in response to sugar starvation. Front Plant Sci 10:280
Kuai B, Chen J, Hörtensteiner S (2018) The biochemistry and molecular biology of chlorophyll breakdown. J Exp Bot 69(4):751–767
Li F, Vierstra RD (2012) Autophagy: a multifaceted intracellular system for bulk and selective recycling. Trends Plant Sci 17(9):526–537
Li KX, Liu YN, Yu BJ, Yang WW, Yue JY, Wang HZ (2018) Monitoring autophagy in wheat living cells by visualization of fluorescence protein-tagged ATG8. Plant Cell Tiss Organ Cult 134:481–489
Li F, Zhang M, Zhang C, Zhou X (2020) Nuclear autophagy degrades a geminivirus nuclear protein to restrict viral infection in solanaceous plants. New Phytol 225(4):1746–1761
Ling Q, Broad W, Trösch R, Töpel M, Demiral Sert T, Lymperopoulos P, Baldwin A, Jarvis RP (2019) Ubiquitin-dependent chloroplast-associated protein degradation in plants. Science 363(6429):eaav4467.
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2 –ΔΔCT method. Methods 25(4):402–408
Mae T, Ohira K (1981) The remobilization of nitrogen related to leaf growth and senescence in rice plants (Oryza sativa L.). Plant Cell Physiol 22:1067–1074
Marshall RS, Vierstra RD (2018) Autophagy: The master of bulk and selective recycling. Annu Rev Plant Biol 69:173–208
Marshall RS, Li F, Gemperline DC, Book AJ, Vierstra RD (2015) Autophagic degradation of the 26S proteasome is mediated by the dual ATG8/ubiquitin receptor RPN10 in Arabidopsis. Mol Cell 58(6):1053–1066
Marshall RS, Hua Z, Mali S, McLoughlin F, Vierstra RD (2019) ATG8-binding UIM proteins define a new class of autophagy adaptors and receptors. Cell 177(3):766-781.e24
Martínez DE, Costa ML, Gomez FM, Otegui MS, Guiamet JJ (2008) “Senescence-associated vacuoles” are involved in the degradation of chloroplast proteins in tobacco leaves. Plant J 56(2):196–206
Masclaux C, Quillere I, Gallais A, Hirel B (2001) The challenge of remobilisation in plant nitrogen economy. A survey of physio-agronomic and molecular approaches. Ann Appl Biol 138:69–81
Masclaux-Daubresse C, Daniel-Vedele F, Dechorgnat J, Chardon F, Gaufichon L, Suzuki A (2010) Nitrogen uptake, assimilation and remobilization in plants: challenges for sustainable and productive agriculture. Ann Bot 105(7):1141–1157
Michaeli S, Honig A, Levanony H, Peled-Zehavi H, Galili G (2014) Arabidopsis ATG8-INTERACTING PROTEIN1 is involved in autophagy-dependent vesicular trafficking of plastid proteins to the vacuole. Plant Cell 26(10):4084–4101
Michaeli S, Galili G, Genschik P, Fernie AR, Avin-Wittenberg T (2016) Autophagy in plants–What’s new on the menu? Trends Plant Sci 21(2):134–144
Nakatogawa H, Ichimura Y, Ohsumi Y (2007) Atg8, a ubiquitin-like protein required for autophagosome formation, mediates membrane tethering and hemifusion. Cell 130(1):165–178
Nishimura K, Kato Y, Sakamoto W (2017) Essentials of proteolytic machineries in chloroplasts. Mol Plant 10(1):4–19
Pei D, Zhang W, Sun H, Wei XJ, Yue JY, Wang HZ (2014) Identification of autophagy-related genes ATG4 and ATG8 from wheat (Triticum aestivum L.) and profiling of their expression patterns responding to biotic and abiotic stresses. Plant Cell Rep 33(10):1697–1710.
Schwarte S, Bauwe H (2007) Identification of the photorespiratory 2-phosphoglycolate phosphatase, PGLP1 in Arabidopsis. Plant Physiol 144(3):1580–1586
Spitzer C, Li F, Buono R, Roschzttardtz H, Chung T, Zhang M, Osteryoung KW, Vierstra RD, Otegui MS (2015) The endosomal protein CHARGED MULTIVESICULAR BODY PROTEIN1 regulates the autophagic turnover of plastids in Arabidopsis. Plant Cell 27(2):391–402
Suttangkakul A, Li F, Chung T, Vierstra RD (2011) The ATG1/ATG13 protein kinase complex is both a regulator and a target of autophagic recycling in Arabidopsis. Plant Cell 23(10):3761–3779
Wada S, Ishida H, Izumi M, Yoshimoto K, Ohsumi Y, Mae T, Makino A (2009) Autophagy plays a role in chloroplast degradation during senescence in individually darkened leaves. Plant Physiol 149(2):885–893
Walter M, Chaban C, Schütze K, Batistic O, Weckermann K, Näke C, Blazevic D, Grefen C, Schumacher K, Oecking C, Harter K, Kudla J (2004) Visualization of protein interactions in living plant cells using bimolecular fluorescence complementation. Plant J 40(3):428–438
Wang S, Blumwald E (2014) Stress-induced chloroplast degradation in Arabidopsis is regulated via a process independent of autophagy and senescence-associated vacuoles. Plant Cell 26(12):4875–4888
Wang Y, Nishimura MT, Zhao T, Tang D (2011) ATG2, an autophagy-related protein, negatively affects powdery mildew resistance and mildew-induced cell death in Arabidopsis. Plant J 68(1):74–87
Wang Y, Yu B, Zhao J, Guo J, Li Y, Han S, Huang L, Du Y, Hong Y, Tang D, Liu Y (2013) Autophagy contributes to leaf starch degradation. Plant Cell 25(4):1383–1399
Wang K, Liu H, Du L, Ye X (2017) Generation of marker-free transgenic hexaploid wheat via an Agrobacterium-mediated co-transformation strategy in commercial Chinese wheat varieties. Plant Biotechnol J 15(5):614–623
Wang FX, Luo YM, Ye ZQ, Cao X, Liang JN, Wang Q, Wu Y, Wu JH, Wang HY, Zhang M, Cheng HQ, Xia GX (2018) iTRAQ-based proteomics analysis of autophagy-mediated immune responses against the vascular fungal pathogen Verticillium dahliae in Arabidopsis. Autophagy 14(4):598–618
Xie Q, Tzfadia O, Levy M, Weithorn E, Peled-Zehavi H, Van Parys T, Van de Peer Y, Galili G (2016) hfAIM: A reliable bioinformatics approach for in silico genome-wide identification of autophagy-associated Atg8-interacting motifs in various organisms. Autophagy 12(5):876–887
Yamane K, Mitsuya S, Taniguchi M, Miyake H (2012) Salt-induced chloroplast protrusion is the process of exclusion of ribulose-1,5-bisphosphate carboxylase/oxygenase from chloroplasts into cytoplasm in leaves of rice. Plant Cell Environ 35(9):1663–1671
Yoo SD, Cho YH, Sheen J (2007) Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat Protoc 2(7):1565-1572 https://doi.org/10.1038/nprot.2007.199
Zess EK, Jensen C, Cruz-Mireles N, De la Concepcion JC, Sklenar J, Stephani M, Imre R, Roitinger E, Hughes R, Belhaj K, Mechtler K, Menke FLH, Bozkurt T, Banfield MJ, Kamoun S, Maqbool A, Dagdas YF (2019) N-terminal β-strand underpins biochemical specialization of an ATG8 isoform. PLoS Biol 17(7):e3000373.
Zhan N, Wang C, Chen L, Yang H, Feng J, Gong X, Ren B, Wu R, Mu J, Li Y, Liu Z, Zhou Y, Peng J, Wang K, Huang X, Xiao S, Zuo J (2018) S-nitrosylation targets GSNO reductase for selective autophagy during hypoxia responses in plants. Mol Cell 71(1):142-154.e6
Zhang X, Ding X, Marshall RS, Paez-Valencia J, Lacey P, Vierstra RD, Otegui MS (2020) Reticulon proteins modulate autophagy of the endoplasmic reticulum in maize endosperm. Elife 9:e51918.
Zhou J, Wang Z, Wang X, Li X, Zhang Z, Fan B, Zhu C, Chen Z (2018) Dicot-specific ATG8-interacting ATI3 proteins interact with conserved UBAC2 proteins and play critical roles in plant stress responses. Autophagy 14(3):487–504
Zhuang X, Wang H, Lam SK, Gao C, Wang X, Cai Y, Jiang L (2013) A BAR-domain protein SH3P2, which binds to phosphatidylinositol 3-phosphate and ATG8, regulates autophagosome formation in Arabidopsis. Plant Cell 25(11):4596–4615
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 31971829), the Knowledge Innovation and Training Program of Tianjin (No. 135305JF78), and the Knowledge Innovation Program of Tianjin Normal University (No. 1353P2XC1604).
Funding
National Natural Science Foundation of China, 31971829, Huazhong Wang, Knowledge Innovation and Training Program of Tianjin, 135305JF78, Huazhong Wang, Knowledge Innovation Program of Tianjin Normal University, 1353P2XC1604, Huazhong Wang
Author information
Authors and Affiliations
Contributions
HW: conceived and designed the study. JN, YL, YX, XY, LJ, and JY: conducted the experiments. HW, JN and JY: analyzed and interpreted the results. HW and JN: wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Zheng-Yi Xu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ni, J., Li, Y., Xiang, Y. et al. Autophagic degradation of the chloroplastic 2-phosphoglycolate phosphatase TaPGLP1 in wheat. Plant Cell Rep 41, 473–487 (2022). https://doi.org/10.1007/s00299-021-02820-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-021-02820-3