Abstract
Most organisms on Earth use glucose, a photosynthetic product, as energy source. The chloroplast, the home of photosynthesis, is the most representative and characteristic organelle in plants and is enclosed by the outer envelope and inner envelope membranes. The chloroplast biogenesis and unique functions are very closely associated with proteins in the two envelope membranes of the chloroplast. Especially, the chloroplast outer envelope membrane proteins have important roles in signal transduction, protein import, lipid biosynthesis and remodeling, exchange of ions and numerous metabolites, plastid division, movement, and host defense. Therefore, biogenesis of these membrane proteins of chloroplast outer envelope membrane is very important for biogenesis of the entire chloroplast proteome as well as plant development. Most proteins among the outer envelope membrane proteins are encoded by the nuclear genome and are post-translationally targeted to the chloroplast outer envelope membrane. In this process, cytoplasmic receptor and import machineries are required for efficient and correct targeting of these membrane proteins. In this review, we have summarized recent advances on the sorting, targeting, and insertion mechanisms of the outer envelope membrane proteins of chloroplasts and also provide future direction of the study on these topics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plastids, found ubiquitously in algae and plants, are very diverse, complex, and versatile organelles (Li and Chiu 2010; Whatley 1978), and chloroplasts are the most important and well-known members of the plastid family. Chloroplasts evolved monophyletically from a photosynthetic bacterial endosymbiont similar to cyanobacteria (Margulis 1981; Mereschkowsky 1905), and participate in various essential metabolic and cellular processes in algae and plants, including photosynthesis, biosynthesis of primary and secondary metabolites, communication, and host defense (Kessler and Schnell 2009; Reyes-Prieto et al. 2007). Chloroplasts are surrounded by two envelope membranes, the outer envelope membrane (OEM) and the inner envelope membrane (IEM), which function as biochemical and physical barriers to enable unique metabolic reactions and functions, and to efficiently communicate with other organelles and the cytosol. These communications are related to physical interactions with other cellular compartments, protein import, and exchanges of metabolites and ions (Inoue 2011). Such events on the chloroplast envelope membrane are closely related to the intrinsic functions of chloroplast OEM proteins. All chloroplast OEM proteins are nuclear-encoded proteins that are synthesized on 80S ribosomes in the cytosol and targeted to the chloroplast OEM in a post-translational targeting process (Keegstra and Cline 1999; Leister 2003). Generally, chloroplast OEM proteins are classified into three groups, signal-anchored (SA), tail-anchored (TA), and β-barrel proteins. The targeting of these proteins to the chloroplast is different from the transit peptide-mediated protein import into chloroplasts (Baldwin and Inoue 2006; Inoue and Keegstra 2003; Lee and Hwang 2018; Tranel and Keegstra 1996). During post-translational targeting, chloroplast OEM proteins face tremendous challenges related to the maintenance of proper proteostasis in an aqueous environment. While traversing that environment, the proteins are prone to misfolding or aggregation. As an essential part of proteostasis, membrane protein-targeting machineries must provide specific receptors or molecular chaperones to protect the cargo proteins from aggregation and to keep them in a translocation-competent state (Jaru-Ampornpan et al. 2010; Kim and Hwang 2013). To elucidate the targeting mechanisms related to these proteins, it is essential to identify their targeting signals as well as the molecular machineries involved in the targeting. Recently, significant progresses have been made in the identification of the targeting signals and cytosolic targeting factors of chloroplast OEM proteins (Bae et al. 2008; Dhanoa et al. 2010; Kim et al. 2014, 2015; Lee et al. 2011; Walther et al. 2009). In this review, we have mainly focused on recent advances in understanding the cytosolic events associated with the targeting of proteins to the OEM of chloroplasts in plants.
Biogenesis of OEM signal-anchored proteins: targeting signals and cytosolic targeting factors
SA proteins mainly perform very important functions in various biological processes including as a receptor for the import of chloroplast precursor proteins and as a lipid-biosynthetic enzyme (Cline and Keegstra 1983; Moellering et al. 2010; Qbadou et al. 2007; Sohrt and Soll 2000). SA proteins of the chloroplast OEM lack a cleavable targeting sequence, the transit peptide that is responsible for proteins import into chloroplasts (Lee et al. 2001, 2004, 2006, 2008; Li and Chiu 2010). The targeting signals for SA proteins of chloroplast OEM have three chemical or structural features: a TMD, a C-terminal positively charged region (CPR), and hydrophobicity in the TMD. Characteristically, SA proteins contain a single TMD at their N-terminal region. The TMD is a stretch of about 20 amino acid residues consisting primarily of hydrophobic amino acids and functions in providing correct localization in the cell and for insertion into the hydrophobic lipid membrane (Lee et al. 2001; Schleiff and Becker 2011; Schleiff et al. 2001). Additionally, the TMDs of the SA proteins of the chloroplast OEM have moderate hydrophobicity. Intriguingly, increasing the hydrophobicity in the TMD of AtToc64 altered its localization from the chloroplast to the plasma membrane (Lee et al. 2004), indicating that moderate hydrophobicity in the TMD of SA proteins is important for targeting SA proteins to the chloroplast OEM. Generally, a TMD should have a hydrophobicity value below 0.4 on the Wimley and White hydrophobicity (WWH) scale with more than 89% of chloroplast SA proteins having hydrophobicity values below 0.4 on the WWH scale (Lee et al. 2011). The other structural feature involved in the targeting of chloroplast OEM SA proteins is the CPR of the TMD. Generally, the CPR in the TMD of a chloroplast OEM-targeted SA protein has an important role in signals related to signal recognition particles evading and determining the topology in membrane (Lee et al. 2001, 2004, 2011; von Heijne 1992). The CPR is mainly located at the short C-terminal flanking region of the TMD and usually contains three or more positively charged amino acid arginine and/or lysine residues (Lee et al. 2011). These three basic amino acid residues in the CPR are a minimum requirement. Substitution of these basic residues with neutral amino acids in the CPR alters the localization of SA proteins OEP7 and AtToc64 to the plasma membrane in Arabidopsis (Lee et al. 2001, 2004, 2011). Although the characteristics of the CPR have not yet been clearly established, additional features, such as amino acid composition and density of basic residues in the CPR and distance of the CPR from the TMD, are also important for CPR function. However, both the basic requirements of CPR and the moderate hydrophobicity of TMD are needed to determine the targeting specificity of an SA protein. Because endoplasmic reticulum (ER) and mitochondrial SA proteins have similar targeting signals, a challenging issue in plant research is to clearly define the CPR and determine how targeting signals confer target specificity.
When nascent proteins of organelles emerge from the exit tunnel of ribosomes during translation, they may interact with specific targeting factors and/or molecular chaperones that assist in specific targeting of the protein to the proper intracellular compartment (Lee et al. 2014; Schlünzen et al. 2005; Spreter et al. 2005; Ullers et al. 2003). Hydrophobic SA proteins of the chloroplast OEM are targeted post-translationally. One challenging point is how hydrophobic SA proteins of the chloroplast OEM can maintain their proper proteostasis in the aqueous environment during post-translational targeting. Therefore, protein-targeting machineries need to provide chaperones or cytosolic receptors to protect the hydrophobic SA proteins from the aqueous environment and to keep them under more favorable conditions for membrane anchoring. Recently, important progress has been made in studying the sorting and targeting mechanism of the chloroplast OEM-targeted SA proteins. Ankyrin repeat-containing protein 2 (AKR2), AKR2A and AKR2B, have been identified as a cytosolic targeting factor for targeting of SA proteins of chloroplast OEM (Bae et al. 2008). AKR2 interacts with the targeting signals of SA client proteins including the TMD and CPR through its N-terminal region that contains PEST, C1, and C2 domains (Bae et al. 2008; Kim et al. 2014). The first step in cargo recognition by AKR2 is in the translation process. The targeting signal of a nascent cargo protein of AKR2 residing in the ribosomal exit tunnel induces AKR2 docking to ribosomal RPL23A located near the exit site (Fig. 1). Subsequently, the RPL23A-bound AKR2 binds to the targeting signal when it becomes exposed from ribosomal exit tunnel (Kim et al. 2015). Therefore, AKR2 displays chaperone activity attributes and prevents non-specific aggregation of its client proteins by binding to the hydrophobic TMD (Bae et al. 2008). However, failure of AKR2 binding to RPL23A severely disrupts protein targeting to the chloroplast OEM and biogenesis of the entire chloroplast proteome as well as plant development (Kim et al. 2015). Therefore, the timing and positioning of AKR2 on the translating ribosome are very critical for correct recognition and protection of the hydrophobic targeting signals of nascent SA proteins during translation. Another important question is how dose AKR2 recognize chloroplasts and insert their SA proteins into the chloroplast OEM. AKR2 binds to two chloroplast OEM lipids, monogalactosyldiacylglycerol (MGDG) and phosphatidylglycerol (PG), through its C-terminal ankyrin repeat domain (ARD) and facilitates membrane insertion of its SA cargo proteins, in which protein import channel Toc75 assists with their insertion (Bae et al. 2008; Kim et al. 2014; Tu et al. 2004). Molecular modeling, structural analysis, and mutational analysis of the ARD have identified two adjacent sites, L1 and L2, for coincidental and synergistic binding of MGDG and PG, respectively. Interestingly, AKR2 of modern-day land plants evolved from the ARD of the host cell by the stepwise addition of three domains (PEST, C1, and C2) in its N-terminal, and two lipids (MGDG and PG) of the endosymbiont were selected to function as the AKR2 receptor for SA protein biogenesis (Fig. 1; Kim et al. 2014). These findings strongly suggest that the targeting mechanism of chloroplast OEM proteins was established by using components from both the host cell and the endosymbiont through gradual modification of the protein–protein interacting module ARD into a lipid-binding domain. Furthermore, the C-terminal ARD of AKR2 also interacts with the dimer form of sHsp17.8, a member of the cytosolic class I small heat shock protein (sHsp) family, as a cofactor, and these interactions facilitate AKR2-mediated targeting of SA proteins to the chloroplast OEM (Kim et al. 2011). In the future, more detailed analysis of membrane insertions of the SA proteins into the chloroplast OEM is needed. Additionally, since the targeting signals of chloroplast and mitochondrial SA proteins in plants are very similar, a challenging study is needed on how the target specificities between these two organelles have been established.
Targeting mechanism of signal-anchored (SA) proteins to the chloroplast outer envelope membranes. Targeting signals of chloroplast OEM SA proteins have three chemical or structural features: a TMD, a CPR, and hydrophobicity in the TMD. The N-terminal moderate hydrophobic TMD of the SA protein has a function as a targeting signal and a membrane anchor. In addition, the CPR following the TMD plays an important role in signal for SRP-evading and determining the topology in membrane. The targeting signal of nascent cargo protein residing in the ribosomal exit tunnel induce AKR2 docking to the ribosomal RPL23A located near the exit site. Subsequently, RPL23A-bound AKR2 binds to the targeting signal when it becomes exposed from ribosomal exit tunnel. The activity of AKR2 for targeting of the SA proteins is enhanced by dimer form of the class I small heat shock protein sHsp17.8. Two adjacent grooves of the ARD specifically binds to two chloroplast lipids MGDG and PG and that this lipid binding is a driving force for the chloroplast recognition of AKR2. Interestingly, AKR2 of modern-day land plants evolved from the ARD of the host cell by stepwise addition of its N-terminal three domains (left box). AKR2 ankyrin repeat-containing protein 2, ARD ankyrin repeat domain, C1 conserved domain 1, C2 conserved domain 2, CPR C-terminal positively charged flanking region, MGDG monogalactosyldiacylglycerol, PEST proline, glutamic acid, serine and threonine rich sequence, PG phosphatidylglycerol, TMD transmembrane domain, +++ positively charged amino acid residues, ? postulated factor, dashed arrow, postulated pathway
Biogenesis of OEM tail-anchored proteins: targeting signals and cytosolic targeting factors
Although a number of TA proteins of the chloroplast OEM have been identified (Inoue 2007; Simm et al. 2013), the targeting signals and biogenesis mechanisms of these proteins have not yet been studied in detail. The targeting signals for TA proteins of chloroplast OEM have four chemical or structural features: a TMD, a C-terminal sequence (CTS, similar to CPR) following the TMD, hydrophobicity in the TMD, and a GTPase domain (in some cases). However, these features of TA proteins of the chloroplast OEM differ in their importance depending on the type of TA protein. TA proteins are another class of chloroplast OEM proteins that do not contain a cleavable signal sequence for chloroplast OEM targeting (Borgese et al. 2007), but TA proteins do have a single TMD at their C-terminal region that functions as a targeting signal and in membrane anchoring (Bauer et al. 2002; Smith et al. 2002; Dhanoa et al. 2010). A noticeable feature of TA proteins of the chloroplast OEM is that the hydrophobicity value in TMD appears to vary significantly compared to that of chloroplast SA proteins and mitochondrial TA proteins containing a moderately hydrophobic TMD (Lee et al. 2014). The hydrophobicity of TMD in TA proteins of the chloroplast OEM is considered to be less important in determining targeting specificity to chloroplast OEM. Moreover, in the case of TA proteins of mitochondrial OEM, the TMD and CTS are necessary and sufficient for mitochondrial OEM targeting (Dukanovic and Rapaport 2011; Hwang et al. 2004; Kaufmann et al. 2003; Rapaport 2003), whereas the C-terminal part of Toc33 or Toc34, including the TMD and CTS, is necessary but not sufficient for targeting to the chloroplast OEM in vivo (Dhanoa et al. 2010). Interestingly, plants are thought to have unique targeting mechanism as has been demonstrated experimentally for the GTPase domain acting as a target signal in Toc33 and Toc34 (Dhanoa et al. 2010). Toc33 and Toc34, which are involved in precursor protein import into the chloroplast interior, have two important domains including the N-terminal GTPase domain and the C-terminal TMD for targeting of and anchoring to the chloroplast OEM. The GTPase domain of Toc33 interacts with the GTPase domain of Toc159, which allows specific targeting of Toc33 to the chloroplast OEM (Bauer et al. 2002; Smith et al. 2002). On the other hand, the GTPase domain of Toc159 binds to the chloroplast OEM in vitro, but a truncated form of Toc159 that lacks the GTPase domain is still capable of binding to the chloroplast OEM (Smith et al. 2002). Therefore, the clear role of the GTPase domain, such as determining targeting specificity in the targeting mechanism of TA proteins of the chloroplast OEM, seems to be limited to certain proteins such as Toc33 and Toc34. In addition, the other structural feature associated with the targeting of chloroplast TA proteins is the CTS. Basic amino acid residues located at a C-terminal flanking region or on both sides of the TMD produce a positive net charge. Although the positive net charge in the CTS is an important structural feature for the proper location of mitochondrial TA proteins, the net charge in the CTS of chloroplast TA proteins appears to have no significant function in some TA proteins such as Toc33 (Dhanoa et al. 2010). The CTS of Toc33 even inhibits the chloroplast OEM targeting of TA protein OEP9. On the other hand, the TMD and CTS in OEP9 are necessary and sufficient for targeting to the chloroplast OEM (Dhanoa et al. 2010). Interestingly, when the CTS of tung tree (Aleurites fordii) mitochondrial cytochrome b5 reductase (Cb5) is replaced with the CTS of OEP9, the chimeric protein was targeted to the chloroplast OEM. Moreover, the CTS of OEP9 mediates chloroplast OEM targeting of the GTPase domain deletion mutant of Toc33. In contrast, substitution mutants of positively charged or negatively charged amino acid residues in the CTS of OEP9 lead to mitochondrial OEM targeting instead of chloroplast OEM targeting, which suggests that the positive net charge or the distribution of positive amino acid residues in the CTS of OEP9 is crucial for determining the targeting specificity of chloroplast TA proteins (Dhanoa et al. 2010). Nevertheless, there are still many unanswered questions regarding the targeting of TA proteins of the chloroplast OEM. More detailed studies are required to clearly define the relationship between the positive net charge or the distribution of positive amino acid residues in the CTS and the targeting specificity of TA proteins of the chloroplast OEM. As in the case of SA proteins, TA proteins of the ER, mitochondria, and chloroplast also show high similarity in the structural and physical–chemical properties of their targeting signals. Therefore, it is necessary to study how the target specificity of TA proteins is determined among the chloroplast OEM and other organelles.
The ribosomal exit tunnel retains the C-terminal, approximately 40 amino acids of a nascent chain, during translation (Blobel and Sabatini 1970). Generally, since the targeting signal TMD and CTS of TA proteins are located within ~ 40 amino acids from the end of the C-terminus, the nascent TA proteins are released from the ribosome before the C-terminal TMD and CTS emerge from the ribosomal exit tunnel; as a consequence, the TMD will be recognized after translation (Stefanovic and Hegde 2007). Therefore, the C-terminal location of the hydrophobic TMD and its exposure timing from the ribosomal exit tunnel cause additional complexity during the targeting of TA proteins (Chartron et al. 2012). Although the targeting mechanisms underlying selective targeting of TA proteins are well established in mammalian cells and yeast, little is known about the molecular mechanism for the biogenesis of TA proteins of the chloroplast OEM. Chloroplast OEM TA proteins such as OEP9 and Toc33/Toc34 interact with the AKR2 which, as a cytosolic targeting factor, is also involved in targeting SA proteins to the chloroplast OEM (Fig. 2; Bae et al. 2008; Dhanoa et al. 2010). However, the targeting signal of OEP9 is partially different from that of Toc33/Toc34. OEP9 has a single hydrophobic TMD and a long hydrophilic CTS comprising 32 amino acids acting as a targeting signal (Dhanoa et al. 2010). Although Toc33 and Toc34 also have TMD and CTS at their C-termini, their targeting to the chloroplast OEM is largely dependent on almost the entire protein sequence including the N-terminal GTPase domain rather than the C-terminal TMD and CTS. This difference implies that the targeting of TA proteins to the chloroplast OEM may not solely depend on AKR2, and there is a possibility of involvement of other factors in this process. Recent studies on green algae (Chlamydomonas reinhardtii) showed that arsenite transporter (ArsA1) appears to be required for the accumulation of TA protein Toc34, an essential component of the chloroplast OEM translocon (TOC) complex (Formighieri et al. 2013; Maestre-Reyna et al. 2017). Interestingly, there are two ARSA homolog genes, ARSA1 and ARSA2, in the C. reinhardtii genome that are homologs of cytosolic targeting factors TRC40 and GET3 in mammalian cells and yeast, respectively (Formighieri et al. 2013). CrArsA1 and CrArsA2 can form a complex with co-expressed TA proteins CrSec61β and CrToc34, respectively. Based on structural modeling, molecular dynamics simulation, and protein-binding assay-based studies, the ArsAs in C. reinhardtii are not arsenite transporters, and the targeting specificity of CrArsAs is achieved at the ligand level, with ArsA1 mainly carrying TA proteins to the chloroplast OEM, while CrArsA2 carries them to the ER (Maestre-Reyna et al. 2017). However, the three GET3 paralogs of Arabidopsis were localize to the cytosol (AtGET3a), chloroplast (AtGET3b), and mitochondria (AtGET3c), respectively (Xing et al. 2017). AtGET3b, a homolog of CrArsA1, is predicted to have a transit peptide, and is clearly targeted to the chloroplast inside. The intracellular localization of AtGET3b and CrArsA1 is completely different. Therefore, further studies are needed on how CrArsA1 and AtGET3 functions in the cytosol and chloroplast, respectively. Additionally, more detailed analyses are required of when and how AKR2 participates in the targeting process of chloroplast TA proteins and of whether AKR2 communicates with ArsA1 or its homologs for specific targeting of TA proteins to the chloroplast OEM in green algae and plants. Another factor involved in the targeting and membrane insertion of chloroplast TA proteins is lipid molecules. In cell-free competitive targeting assays, targeting of chloroplast TA protein can occur efficiently and with high fidelity in the absence of cytosolic factors, but the addition of cytosolic fractions, Hsp70, and Hsp90, respectively, increased the targeting efficiency of chloroplast TA proteins but not the fidelity (Kriechbaumer and Abell 2012). Additionally, compared with OEP9, Toc33 and Toc34 differ in their membrane insertions and those insertions are dependent on themselves and the unique lipid composition of the chloroplast OEM. In addition, membrane insertion of OEP9 into the chloroplast OEM is dependent on a proteinaceous factor (Dhanoa et al. 2010). These results suggest that targeting of the TA proteins of chloroplast OEM is primarily dependent on events at the chloroplast OEM, and the main role of cytosolic targeting factors in this process is to increase targeting efficiency by maintaining an insertion competent status at the outer envelope. Nevertheless, cytosolic targeting factors are probably needed to protect the hydrophobic TMD, which acts as a target signal, and to keep nascent membrane proteins in membrane insertion competent status by preventing their aggregation or by minimizing abnormal interactions with other proteins or metabolites in the cellular environment (Ellis and Minton 2006; Flores-Pérez and Jarvis 2013; Lee et al. 2014).
Targeting mechanism of tail-anchored (TA) proteins to the chloroplast outer envelope membrane. Little information is available about the mechanisms and molecular machineries of the targeting of TA proteins to the chloroplast OEM. Basically, the targeting signals for TA proteins of chloroplast OEM have four chemical or structural features: a TMD, a C-terminal sequence (CTS, similar to CPR) following the TMD, hydrophobicity in the TMD, and the GTPase domain (in the case of Toc33/Toc34). However, these features differ in their importance depending on the type of TA proteins. AKR2 also binds to the chloroplast OEM-targeted TA proteins such as OEP9, Toc33, and Toc34. In addition, the cytosolic targeting factor ArsA1, which is homologs of TRC40 and Get3 function as cytosolic targeting factor for the TA proteins in yeast and mammalian cells, mediates the biogenesis and targeting of Toc34 in green algae (Chlamydomonas reinhardtii). In the case of Toc33, the GTPase domain of Toc33 interacts with the GTPase domain of Toc159 which allows specific targeting of Toc33 to the chloroplast OEM. Moreover, the unique lipid composition of the chloroplast OEM is also important for the efficient insertion of the TA proteins. AKR2 ankyrin repeat-containing protein 2, ArsA1 arsenite transport 1, CTS C-terminal sequence, MGDG monogalactosyldiacylglycerol, PG phosphatidylglycerol, TA tail-anchored, TMD transmembrane domain, +++ positively charged amino acid residues, ? postulated factor, dashed arrow, postulated pathway
Biogenesis of OEM β-barrel proteins: targeting signals and cytosolic targeting factors
The third group of chloroplast OEM proteins includes the β-barrel proteins that mainly function as transporters for ions, metabolites, and chloroplast interior and IEM proteins. Although little is known about the targeting mechanism of the chloroplast OEM β-barrel proteins, the targeting mechanism of Toc75-III has been studied extensively. Toc75-III is a common element of TOC complexes and it has a central role in general protein import as well as in both the assembly and function of TOC GTPases such as Toc34 and Toc159. In bacteria, yeast, and mammalian cell studies, the targeting signal of β-barrel proteins is not restricted to specific motifs or domains in the primary sequence. Instead, it has been proposed that the targeting signal is dispersed throughout the protein and is displayed in the form of a secondary and/or tertiary structure (Court et al. 1996; Rapaport and Neupert 1999; Walther et al. 2009). Unlike β-barrel proteins of other organelles, the chloroplast OEM β-barrel protein Toc75-III has a cleavable N-terminal bipartite targeting signal that is essential for protein targeting from the cytosol to the chloroplasts after translation (Tranel and Keegstra 1996). Intriguingly, the N-terminal target signal of Toc75-III is composed of two parts; the first part is a canonical transit peptide, and the second part is a glycine-rich region comprised a polyglycine stretch and hydrophobic residues following the transit peptide (Inoue and Keegstra 2003; Tranel and Keegstra 1996). Chimeric fusion proteins of the N-terminal region of precursor Toc75-III can be targeted to the stroma, which means that the N-terminal region has a function as a typical transit peptide (Tranel and Keegstra 1996). In addition, the function of the glycine-rich region is to insert Toc75 into the chloroplast OEM (Baldwin and Inoue 2006; Tranel and Keegstra 1996). On the other hand, other chloroplast OEM β-barrel proteins, OEP24, OEP37, and Toc75-V/OEP80, are also predicted to have a transit peptide at their N-terminal end, whereas OEP23 and TGD4 do not (Lee et al. 2014; Patel et al. 2008). In the case of Toc75-V/OEP80 in Arabidopsis, the approximately 52 amino acid residues of the N-terminal are dispensable in the targeting, insertion, or functionality of this protein (Hsu et al. 2012; Patel et al. 2008). However, whether the predicted transit peptides in the other proteins are involved in chloroplast targeting has not yet been experimentally confirmed. Collectively, these results suggest that the location and form of targeting signals vary depending on each protein and indicate the possibility that various targeting mechanisms exist for chloroplast OEM-targeted β-barrel proteins. In the future, more detailed analyses are required to identify the targeting signals in the various β-barrel proteins of the chloroplast OEM and to determine how the targeting signals of chloroplast OEM-targeted β-barrel proteins is different from the targeting signals of the β-barrel proteins targeted to other organelles.
Consequently, specific cytosolic targeting factors for the sorting, targeting, and membrane insertion of β-barrel proteins of the chloroplast OEM have not yet been identified in plants. Although OEP24, OEP37, and Toc75-V/OEP80 are predicted to have a cleavable targeting signal, all currently studied chloroplast OEM proteins except Toc75-III are synthesized in the cytosol in mature form without a cleavable targeting signal and are targeted to the chloroplast OEM (Hofmann and Theg 2005; Lee et al. 2014; Li and Chiu 2010; Schleiff and Becker 2011). Toc75-III has a cleavable bipartite targeting signal consisting of a canonical transit peptide and a polyglycine stretch, which can be cleaved off by the stromal processing peptidase (SPP) and a membrane-bound plastidic type I signal peptidase (Plsp1), respectively (Fig. 3; Inoue et al. 2005; Tranel and Keegstra 1996). The intermediate form of Toc75-III, the N-terminal transit peptide-cleaved form, does not accumulate in the stroma but is arrested in the intermembrane space between the outer and inner envelope membranes, and the glycine-rich region in Toc75-III is cleaved by a type I signal peptidase, resulting in mature and functional Toc75-III (Inoue and Keegstra 2003; Inoue et al. 2005; Shipman and Inoue 2009). Interestingly, the glycine-rich region in the intermediate form of Toc75-III appears to be critical for arresting the import of the protein in the intermembrane space and for stimulating the integration of the Toc75-III mature form into the chloroplast OEM (Inoue and Keegstra 2003). In addition, the reduction of OEP80 in Arabidopsis by a dexamethasone-inducible RNA interference strategy results in the reduced accumulation of Toc75-III, raising the possibility that OEP80 might constitute the core of the machinery needed for β-barrel protein insertion into the chloroplast OEM (Hsu and Inoue 2009; Huang et al. 2011; Schleiff and Soll 2005). However, direct evidence for the involvement of OEP80 in membrane insertion of β-barrel proteins has not been reported. Overall, the exact mechanism of and cytosolic components for sorting, targeting and insertion of chloroplast OEM-targeted β-barrel proteins, including Toc75-III, are still undescribed. Moreover, it is unclear whether the N-terminal transit peptide of β-barrel proteins is functionally similar to that of the traditional transit peptide in chloroplast interior proteins and whether molecular chaperones such as Hsp70, Hsp90, and 14-3-3 are also involved in the targeting of these proteins.
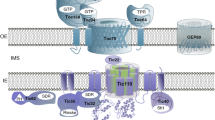
Targeting mechanism of β-barrel proteins to the chloroplast outer envelope membrane. Although little is known about the targeting mechanism of the chloroplast OEM-targeted β-barrel proteins, the targeting mechanism of Toc75-III has been studied extensively. Unlike β-barrel proteins with other organelles, the β-barrel protein Toc75 in the chloroplast OEM has a cleavable N-terminal bipartite targeting signal that is essential for targeting to the chloroplasts. The N-terminal target signal of Toc75-III is composed of two parts; the first part is a canonical transit peptide cleaved in the stroma by the SPP, whereas the second part is a glycine-rich region removed by the Plsp1 in the intermembrane space between the outer and inner envelope membrane. On the other hand, other chloroplast OEM β-barrel protein OEP24, OEP37, and Toc75-V/OEP80 are also predicted to have a transit peptide at its N-terminal end, whereas OEP23 and TGD4 do not. However, the exact mechanism and cytosolic components for sorting, targeting, and insertion of the chloroplast OEM-targeted β-barrel proteins including Toc75-III is still unknown. IMS intermembrane space, Plsp1 plastidic type I signal peptidase, SPP stromal processing peptidase, TIC translocon at the inner envelope membrane of chloroplasts, TOC translocon at the outer envelope membrane of chloroplasts, ? postulated factor, dashed arrow, postulated pathway
Conclusions and prospects
The mechanisms for protein targeting to various subcellular locations have been studied extensively in bacteria, yeast, and mammalian cells. Now detailed molecular mechanisms have been elucidated for ER and mitochondrial protein distribution systems. Understanding the biogenesis mechanisms of proteins targeted to each organelle can provide important clues to elucidating the physiological function of each organelle and to understanding the principles of related life phenomena. Recently, significant progresses have been made in the identification of the sorting mechanism of SA client proteins and the recognition mechanism of the chloroplast OEM by the cytosolic receptor AKR2 in plants. The chloroplast OEM proteins can be classified into three groups: SA proteins, TA proteins, and β-barrel proteins. Especially, the SA and TA proteins have a hydrophobic TMD acting as a targeting signal and a membrane insertion region. Therefore, the biogenesis of membrane proteins needs to be understood in terms of four key stages: coordination and sorting of ribosomes during translation, recognition of nascent client proteins by targeting factors, receptors on the target organelle for targeting factors, and transporters or translocases for insertion of membrane proteins. For the elucidation of the biogenesis process of chloroplast OEM proteins, further studies are needed to find many puzzle pieces in order to fully understand these four key stages. In addition, since membrane proteins of ER and mitochondria also have hydrophobic TMDs as targeting signals, it is also necessary to study how the targeting signals of chloroplasts, ER, and mitochondria can be differentiated to ensure proper targeting. Furthermore, AKR2, a cytosolic receptor for chloroplast OEM-targeted SA proteins, has evolved through the stepwise addition of N-terminal PEST, C1, and C2 domains from the ARD of the host cell during the evolution from green algae to land plants. Therefore, future studies should be directed to find clues as to how the targeting signal of the chloroplast OEM proteins, its cytosolic factors, and the components involved in the membrane insertion have evolved in parallel through communication with each other during endosymbiosis of cyanobacteria. Such studies will contribute greatly to understanding the organellogenesis process of endosymbionts.
Author contribution statement
DHK devised and supervised the project. JK, YJN, SJP, and SB designed the figures. JK, YJN, and DHK wrote the manuscript. All authors reviewed, revised and approved the manuscript.
References
Bae W, Lee YJ, Kim DH, Lee J, Kim S, Sohn EJ, Hwang I (2008) AKR2A mediated import of chloroplast outer membrane proteins is essential for chloroplast biogenesis. Nat Cell Biol 10:220–227
Baldwin AJ, Inoue K (2006) The most C-terminal tri-glycine segment within the polyglycine stretch of the pea Toc75 transit peptide plays a critical role for targeting the protein to the chloroplast outer envelope membrane. FEBS J 273:1547–1555
Bauer J, Hiltbrunner A, Weibel P, Vidi PA, Alvarez-Huerta M, Smith MD, Schnell DJ, Kessler F (2002) Essential role of the G-domain in targeting of the protein import receptor atToc159 to the chloroplast outer membrane. J Cell Biol 159:845–854
Blobel G, Sabatini DD (1970) Controlled proteolysis of nascent polypeptides in rat liver cell fractions. I. Location of the polypeptides within ribosomes. J Cell Biol 45:130–145
Borgese N, Brambillasca S, Colombo S (2007) How tails guide tail-anchored proteins to their destinations. Curr Opin Cell Biol 19:368–375
Chartron JW, Clemons WM, Suloway CJM (2012) The complex process of GETting tail-anchored membrane proteins to the ER. Curr Opin Struc Biol 22:217–224
Cline K, Keegstra K (1983) Galactosyltransferases involved in galactolipid biosynthesis are located in the outer membrane of pea chloroplast envelopes. Plant Physiol 71(2):366–372
Court DA, Kleene R, Neupert W, Lill R (1996) Role of the N- and C-termini of porin in import into the outer membrane of Neurospora mitochondria. FEBS Lett 390:73–77
Dhanoa PK, Richardson LG, Smith MD, Gidda SK, Henderson MP, Adrews DW, Mullen RT (2010) Distinct pathways mediate the sorting of tail-anchored proteins to the plastid outer envelope. PLoS One 5(4):e10098
Dukanovic J, Rapaport D (2011) Multiple pathways in the integration of proteins into the mitochondrial outer membrane. Biochim Biophys Acta 1808:971–980
Ellis RJ, Minton AP (2006) Protein aggregation in crowded environments. Biol Chem 387:485–497
Flores-Pérez U, Jarvis P (2013) Molecular chaperone involvement in chloroplast protein import. Biochim Biophys Acta 1833:332–340
Formighieri C, Cazzaniga S, Kuras R, Bassi R (2013) Biogenesis of photosynthetic complexes in the chloroplast of Chlamydomonas reinhardtii requires ARSA1, a homolog of prokaryotic arsenite transporter and eukaryotic TRC40 for guided entry of tail-anchored proteins. Plant J 73:850–861
Hofmann NR, Theg SM (2005) Chloroplast outer membrane protein targeting and insertion. Trends Plant Sci 10:450–457
Hsu SC, Inoue K (2009) Two evolutionarily conserved essential β-barrel proteins in the chloroplast outer envelope membrane. Biosci Trends 3:168–178
Hsu SC, Nafati M, Inoue K (2012) OEP80, an essential protein paralogous to the chloroplast protein translocation channel Toc75, exists as a 70-kD protein in the Arabidopsis thaliana chloroplast outer envelope. Plant Mol Biol 78:147–158
Huang W, Ling Q, Bedard J, Lilley K, Jarvis P (2011) In vivo analyses of the roles of essential Omp85-related proteins in the chloroplast outer envelope membrane. Plant Physiol 157:147–159
Hwang YT, Pelitire SM, Henderson MP, Andrews DW, Dyer JM, Mullen RT (2004) Novel targeting signals mediate the sorting of different isoforms of the tail-anchored membrane protein cytochrome b5 to either endoplasmic reticulum or mitochondria. Plant Cell 16:3002–3019
Inoue K (2007) The chloroplast outer envelope membrane: The edge of light and excitement. J Integr Plant Biol 49(8):1100–1111
Inoue K (2011) Emerging roles of the chloroplast outer envelope membrane. Trends Plant Sci 16:550–557
Inoue K, Keegstra K (2003) A polyglycine stretch is necessary for proper targeting of the protein translocation channel precursor to the outer envelope membrane of chloroplasts. Plant J 34:661–669
Inoue K, Baldwin AJ, Shipman RL, Matsui K, Theg SM, Ohme-Takagi M (2005) Complete maturation of the plastid protein translocation channel requires a type I signal peptidase. J Cell Biol 171:425–430
Jaru-Ampornpan P, Shen K, Lam VQ, Ali M, Doniach S, Jia TZ, Shan S (2010) ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP. Nat Struct Mol Biol 17(6):696–702
Kaufmann T, Schlipf S, Sanz J, Neubert K, Stein R, Borner C (2003) Characterization of the signal that directs Bcl-xL, but not Bcl-2, to the mitochondrial outer membrane. J Cell Biol 160:53–64
Keegstra K, Cline K (1999) Protein import and routing systems of chloroplasts. Plant Cell 11(4):557–570
Kessler F, Schnell D (2009) Chloroplast biogenesis: diversity and regulation of the protein import apparatus. Curr Opin Cell Biol 21:494–500
Kim DH, Hwang I (2013) Direct targeting of proteins from the cytosol to organelles: the ER versus endosymbiotic organelles. Traffic 14(6):613–621
Kim DH, Xu Z-H, Na YJ, Yoo YJ, Lee J, Sohn EJ, Hwang I (2011) Small heat shock protein Hsp17.8 functions as an AKR2A cofactor in the targeting of chloroplast outer membrane proteins in Arabidopsis. Plant Physiol 157:132–146
Kim DH, Park MJ, Gwon GH, Silkov A, Xu ZY, Yang EC, Song S, Song K, Kim Y, Yoon HS, Honig B, Cho W, Cho Y, Hwang I (2014) An ankyrin repeat domain of AKR2 drives chloroplast targeting through coincident binding of two chloroplast lipids. Dev Cell 30(5):598–609
Kim DH, Lee JE, Xu ZY, Geem KR, Kwon Y, Park JW, Hwang I (2015) Cytosolic targeting factor AKR2A captures chloroplast outer membrane-localized client proteins at the ribosome during translation. Nat Commun 6:6843
Kriechbaumer V, Abell BM (2012) Chloroplast envelope protein targeting fidelity is independent of cytosolic components in dual organelle assays. Front Plant Sci 3:148
Lee DW, Hwang I (2018) Evolution and design principles of the diverse chloroplast transit peptides. Mol Cells 41:161–167
Lee YJ, Kim DH, Kim YW, Hwang I (2001) Identification of a signal that distinguishes between the chloroplast outer envelope membrane and the endomembrane system in vivo. Plant Cell 13:2175–2190
Lee YJ, Sohn EJ, Lee KH, Lee DW, Hwang I (2004) The transmembrane domain of AtToc64 and its C-terminal lysine-rich flanking region are targeting signals to the chloroplast outer envelope membrane [correction]. Mol Cells 17:281–291
Lee DW, Lee S, Lee GJ, Lee KH, Kim S, Cheong GW, Hwang I (2006) Functional characterization of sequence motifs in the transit peptide of Arabidopsis small subunit of rubisco. Plant Physiol 140:466–483
Lee DW, Kim JK, Lee S, Choi S, Kim S, Hwang I (2008) Arabidopsis nuclear-encoded plastid transit peptides contain multiple sequence subgroups with distinctive chloroplast-targeting sequence motifs. Plant Cell 20:1603–1622
Lee J, Lee H, Kim J, Lee S, Kim DH, Kim S, Hwang I (2011) Both the hydrophobicity and a positively charged region flanking the C-terminal region of the transmembrane domain of signal-anchored proteins play critical roles in determining their targeting specificity to the endoplasmic reticulum or endosymbiotic organelles in Arabidopsis cells. Plant Cell 23:1588–1607
Lee J, Kim DH, Hwang I (2014) Specific targeting of proteins to outer envelope membranes of endosymbiotic organelles, chloroplasts, and mitochondria. Front Plant Sci 5:1–11
Leister D (2003) Chloroplast research in the genomic age. Trends Genet 19:47–56
Li HM, Chiu CC (2010) Protein transport into chloroplasts. Ann Rev Plant Biol 61:157–180
Maestre-Reyna M, Wu SM, Chang YC, Chen CC, Maestre-Reyna A, Wang AH, Chang HY (2017) In search of tail-anchored protein machinery in plants: reevaluating the role of arsenite transporters. Sci Rep 7:46022
Margulis L (1981) Symbiosis in cell evolution. WH Freeman, New York
Mereschkowsky C (1905) Über Natur und Ursprung der Chromatophoren im Pflanzenreiche. Biol Centralbl 25:593–604
Moellering ER, Muthan B, Benning C (2010) Freezing tolerance in plants requires lipid remodeling at the outer chloroplast membrane. Science 330:226–228
Patel R, Hsu SC, Bedard J, Inoue K, Jarvis P (2008) The Omp85-related chloroplast outer envelope protein OEP80 is essential for viability in Arabidopsis. Plant Physiol 148:235–245
Qbadou S, Becker T, Bionda T, Reger K, Ruprecht M, Soll J, Schleiff E (2007) Toc64—a preprotein-receptor at the outer membrane with bipartide function. J Mol Biol 367(5):1330–1346
Rapaport D (2003) Finding the right organelle. Targeting signals in mitochondrial outer-membrane proteins. EMBO Rep 4:948–952
Rapaport D, Neupert W (1999) Biogenesis of Tom40, core component of the TOM complex of mitochondria. J Cell Biol 146:321–331
Reyes-Prieto A, Weber AP, Bhattacharya D (2007) The origin and establishment of the plastid in algae and plants. Annu Rev Genet 41:147–168
Schleiff E, Becker T (2011) Common ground for protein translocation: access control for mitochondria and chloroplasts. Nat Rev Mol Cell Biol 12:48–59
Schleiff E, Soll J (2005) Membrane protein insertion: mixing eukaryotic and prokaryotic concepts. EMBO Rep 6:1023–1027
Schleiff E, Tien R, Salomon M, Soll J (2001) Lipid composition of outer leaflet of chloroplast outer envelope determines topology of OEP7. Mol Biol Cell 12(12):4090–4102
Schlünzen F, Wilson DN, Tian P, Harms JM, Mclnnes SJ, Hansen HA, Albrecht R, Buerger J, Wilbanks SM, Fucini P (2005) The binding mode of the trigger factor on the ribosome: implications for protein folding and SRP interaction. Structure 13(11):1685–1694
Shipman RL, Inoue K (2009) Suborganellar localization of plastidic type I signal peptidase 1 depends on chloroplast development. FEBS Lett 583:938–942
Simm S, Papasotiriou DG, Ibrahim M, Leisegang MS, Müller B, Schorge T, Karas M, Mirus O, Sommer MS, Schleiff E (2013) Defining the core proteome of the chloroplast envelope membranes. Front Plant Sci 4:11
Smith MD, Hiltbrunner A, Kessler F, Schnell DJ (2002) The targeting of the atToc159 preprotein receptor to the chloroplast outer membrane is mediated by its GTPase domain and is regulated by GTP. J Cell Biol 159:833–843
Sohrt K, Soll J (2000) Toc64, a new component of the protein translocon of chloroplasts. J Cell Biol 148(6):1213–1221
Spreter T, Pech M, Beatrix B (2005) The crystal structure of archaeal nascent polypeptide-associated complex (NAC) reveals a unique fold and the presence of a ubiquitin-associated domain. J Biol Chem 280(16):15849–15854
Stefanovic S, Hegde RS (2007) Identification of a targeting factor for posttranslational membrane protein insertion into the ER. Cell 128(6):1147–1159
Tranel PJ, Keegstra K (1996) A novel, bipartite transit peptide targets OEP75 to the outer membrane of the chloroplastic envelope. Plant Cell 8:2093–2104
Tu SL, Chen LJ, Smith MD, Su YS, Schnell DJ, Li HM (2004) Import pathways of chloroplast interior proteins and the outer-membrane protein OEP14 converge atToc75. Plant Cell 16:2078–2088
Ullers RS, Houben EN, Raine A, ten Hagen-Jongman CM, Ehrenberg M, Brunner J, Oudega B, Harms N, Luirink J (2003) Interplay of signal recognition particle and trigger factor at L23 near the nascent chain exit site on the Escherichia coli ribosome. J Cell Biol 161(4):679–684
von Heijne G (1992) Membrane protein structure prediction. Hydrophobicity analysis and the positive-inside rule. J Mol Biol 225:487–494
Walther DM, Rapaport D, Tommassen J (2009) Biogenesis of beta-barrel membrane proteins in bacteria and eukaryotes: evolutionary conservation and divergence. Cell Mol Life Sci 66:2789–2804
Whatley JM (1978) A suggested cycle of plastid developmental interrelationships. New Phytol 80:489–502
Xing S, Mehlhorn DG, Wallmeroth N, Asseck LY, Kar R, Voss A, Denninger P, Schmidt VA, Schwarzländer M, Stierhof YD, Grossmann G, Grefen C (2017) Loss of GET pathway orthologs in Arabidopsis thaliana causes root hair growth defects and affects SNARE abundance. Proc Natl Acad Sci USA 114(8):E1544–E1553
Acknowledgements
This research was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT & Future Planning (2017R1A4A1015594, 2017R1C1B2009362).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial interests.
Additional information
Communicated by Inhwan Hwang.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kim, J., Na, Y.J., Park, S.J. et al. Biogenesis of chloroplast outer envelope membrane proteins. Plant Cell Rep 38, 783–792 (2019). https://doi.org/10.1007/s00299-019-02381-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-019-02381-6