Abstract
Key message
The loss-of-function mutants of the Arabidopsis orthologue of the wheat LRK10 gene shows ABA-insensitive and drought stress-sensitive phenotypes, suggesting that LRK10L1.2 is positively involved in ABA signaling.
Abstract
A subset of receptor-like kinases (RLKs) superfamily proteins play a key role in sensing internal and external signals. A gene encoding Arabidopsis thaliana Leaf rust 10 disease-resistance locus receptor-like protein kinase 1 (AtLRK10L1), most closely related to wheat LRK10, expresses two different transcripts, LRK10L1.1 and LRK10L1.2, using alternative promoters. The T-DNA insertion mutant, lrk10l1-2, that specifically shuts down LRK10L1.2 transcription displayed an abscisic acid (ABA)-insensitive phenotype in seed germination and seedling growth. However, the lrk10l1.2 mutant exhibited reduced tolerance to drought stress, compared with wild type, which is accompanied by alteration of stomatal apertures. The transgenic plants overexpressing full-length LRK10L1.2, which localizes to the plasma membrane (PM) complemented the phenotypes of lrk10l1-2 mutant background, while those expressing LRK10L1.2 Nu1, which switched its localization to the endoplasmic reticulum (ER) by skipping of a mini-exon, showed even higher ABA insensitivity and drought sensitivity than its mutant background. Our results suggest that ABA signaling involves the PM-localized LRK10L1.2.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Belonging to a monophyletic group of RLK/Pelle serine/threonine protein kinase superfamily, receptor-like protein kinases (RLKs) represent 615 members out of 1,041 total protein kinases in Arabidopsis (Shiu and Bleecker 2001). RLKs are defined by consisting of an N-terminal extracellular and the C-terminal protein kinase domains with a transmembrane (TM) patch between them. These proteins respond to a variety of internal and external stimuli through the N-terminal extracellular domain, and thus RLKs are subdivided into 23 subfamilies according to the presence and the arrangement of motifs or subdomains present in this domain involved in ligand-binding (Shiu and Bleecker 2003). A prominent characteristic of RLKs is that they resemble receptor tyrosine kinases in mammalian cells, so that most plant RLKs are anchored on the plasma membrane (PM) through the TM patch, and perceive internal and external ligands by the N-terminal extracellular domain and activate downstream signaling through phosphorylation by cytosolic serine/threonine protein kinase domain (Gish and Clark 2011).
The most well-known subclass of RLK superfamily proteins is leucine-rich repeat (LRR) containing RLKs encompassing 200 members in Arabidopsis. Because of the number and arrangements being diversified, LRR-RLK proteins are linked to a variety of molecular functions involving stem cell differentiation, patterning of the embryo, brassinosteroid signaling and pathogen recognition (Clark et al. 1997; Nodine et al. 2007; Kwak et al. 2005; Searle et al. 2003; Gomez–Gomez and Boller 2000). Other RLKs of known functions are those containing epidermal growth factor-like repeats, C-type lectin-like and extension-like motifs involved in biogenesis of cell wall (He et al. 1999; Ringli 2010; Nakhamchik et al. 2004), tumor necrosis factor-like motifs in epidermal development (Becraft et al. 1996), lysine motifs (LysM) in Nod factor perception (Limpens et al. 2003), self-incompatibility (S) domains in pollen recognition (Stein et al. 1991), and thaumatin-like motifs in pathogen recognition (Wang et al. 1996).
Abiotic stress responses also involve RLKs. For example, ABA signaling involves a LRR-RLK designated receptor-like kinase 1 (RPK1; Osakabe et al. 2005), proline-rich extension-like receptor kinase 4 (PERK4; Bai et al. 2009), guard cell hydrogen peroxide-resistant 1 (GHR1; Hua et al. 2012) and cysteine-rich receptor-like kinase 45 (CRK45; Zhang et al. 2013), while ABA signaling is negatively regulated by CRK36 (Tanaka et al. 2012). However, physiological and molecular functions of the LRK10-like N-terminal domains are as yet largely unknown. The gene encoding LRK10 was first identified from wheat (TaLRK10) by homology-based cloning from the basis of TaLr10 disease-resistance locus (Feuillet et al. 1997). LRK10L1, the most closely related Arabidopsis homolog of TaLRK10, produces two transcripts, AtLRK10L1.1 and AtLRK10L1.2, using alternative promoters (Accompanying paper). We have shown that all wheat and Arabidopsis LRK10L genes contain mini-exons of varying sizes and sequences. By expressing alternatively spliced variants of AtLRK10L1.2, we showed that the 45 nt mini-exon 2 is required for targetting of the protein to the plasma membrane (Accompanying paper). Here we report that the LRK10L1.2 protein produced from the latter transcript is involved in ABA signaling and drought resistance in Arabidopsis and requires to be targetted to the plasma membrane.
Materials and methods
Plant materials and Arabidopsis transformation
Arabidopsis thaliana T-DNA insertion lines GK_270F11 (lrk10l1-1), SALK_144810 (lrk10l1-2) and SALK_150401 (lrk10l1-3) were obtained from The European Arabidopsis Stock Centre (uNASC). The homozygous T3 lines were screened by PCR using the primers shown in Table S1, and used for further analyses. Col and Col-0 WT backgrounds, and the above T-DNA mutants were grown in soil, stratified at 4 °C for 3 days and grown under long-day condition in a conditioned chamber [16 h light (120 μmol photons m−2 s−1): 8 h dark]. A. tumefaciens strain GV3101 containing binary plant expression vector containing LRK10L1.2 No4 or LRK10L1.2 Nu1 fused to YFP under CaMV 35S promoter (Accompanying paper) was used for transformation of lrk10l1-2 by the flower-dipping method (Clough and Bent 1998). Seeds were screened on MS media containing 50 μg/ml basta (Duchefa, Haarlem, The Netherlands).
ABA treatments
Arabidopsis seeds were stratified at 4 °C for 3 days and sown on plates containing 0.5× MS agar plates supplemented with either 0.5 or 2.0 μM ABA. After 3 days, the germinated seeds were quantified. For measurement of root growth, Arabidopsis seeds were germinated on 0.5× MS agar media containing 0.5 μM ABA, and plates were placed vertically on shelves. The seedlings were grown with a day/night cycle of 16/8 h at 22 °C and in a light intensity of 130 μmol photons m−2 s−1.
Assays of drought tolerance
One-week-old seedlings were randomly planted in a tray containing soil mix (peat moss, perlite, and vermiculite, 9:1:1). Dehydration stress was imposed to plants by withholding watering. To determine the drought tolerance in a quantitative manner, leaves were detached from each plant and placed in petri dishes. The dishes were kept in a growth chamber with 40 % relative humidity, and the loss of fresh weight was determined at the indicated times.
For stomatal aperture bioassays, four rosette leaves from 4-week-old plants were detached and floated in stomatal opening solution (SOS: 50 mM KCl and 10 mM MES-KOH, pH 6.15, 10 mM CaCl2) in the light as described by Lee et al. (2009). After 2.5 h, SOS was replaced with SOS containing ABA of various concentrations. After 2.5 h further incubation, 80 stomata from each individual leaf sample were measured, and each experiment was performed in triplicate.
RT-PCR analysis
Twenty-one-day-old plants were used for RNA extraction. For drought treatments, whole rosette leaves were removed and incubated in a 22 °C chamber described above for the indicated time. For ABA treatments, plants were sprayed with 50 μM ABA and incubated for the indicated time. Reverse-transcriptase polymerase chain reactions were done by the method described previously (Kim et al. 2009). For real-time PCR, the resulting cDNA was amplified in a CFX96 Touch™ Real-Time PCR detection system (Bio-Rad, Hercules, CA, USA) with iQ™SYBR Green Supermix. Reactions were performed in triplicate and with the following cycling conditions: 95 °C for 5 min, 45 cycles at 95 °C for 20 s and 60 °C for 20 s, and then 72 °C for 20 s. The relative expression value for each gene was calculated using the ∆∆Ct method (Livak and Schmittgen 2001). Relative levels of the transcripts were normalized to the expression levels of genes used as internal controls, which included Actin2, Actin8, and EF1α. Semi-quantitative and quantitative RT-PCR analysis was performed with at least two biological and three technical replicates. Primers used for PCR are listed in Table S1.
Results
Loss-of-function mutants of LRK10L1.2 do not show phenotypic changes under normal growth conditions
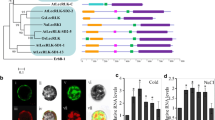
To investigate the molecular function of LRK10L1, we obtained the insertional mutants lrk10l1-1, lrk10l1-2 and lrk10l1-3 which contain T-DNA insertions in the different exon 1 sequences of LRK10L1.1 and LRK10L1.2, and 202 bp upstream of LRK10L1.2 transcription start site, respectively (Fig. 1a). The T3 homozygote lines were analyzed for LRK10L1.2 expression by RT-PCR using primer sets shown in Fig. 1a. No RT-PCR products were observed in lrk10l-2 from reactions using primer sets across the T-DNA insertion or downstream of the insertion (P1 + P2 and P2 + P3 in Fig. 1b, lane 3), suggesting that lrk10l-2 is a knockout mutant of LRK10L1.2. However, weak signal was detected in lrk10l-3 compared with WT Col-0 background (Fig. 1b, lane 5 and 4, respectively). As we previously showed that intron 1 of LRK10L1.1 in LRK10L1 gene contains enough upstream sequence for the LRK10L1.2 transcription (Accompanying paper), this finding indicates that at least some putative upstream elements required for full activation of LRK10L1.2 transcription may be essential and present between the T-DNA insertion site in lrk10l-3 and 5′ splice site (ss) of LRK10L1.1 IVS1. Notably, the T-DNA insertion in exon 1 of LRK10L1.1 (lrk10l-1) in turn produced stronger signals compared with WT (arrowheads in Fig. 1b, lanes 1 and 2, respectively), suggesting reciprocal negative regulation of LRK10L1 transcription by which the amount of LRK10L1.2 mRNA may be decreased when the LRK101.1 transcription is in progress, and vice versa.
a The T-DNA integration sites of mutant Arabidopsis plants used in this study are indicated. Exons are boxed; in which 5′ and 3′ UTRs and coding regions are filled in gray and black, respectively, while introns are indicated with solid lines. The arrows indicate relative positions of primers used for RT-PCR in b. b RT-PCR analysis (23 cycles) of the T-DNA mutants and the complementation lines using primer pairs described in a and listed in Fig. S1. The arrowheads indicate up-regulation of the fully spliced LRK10L1.2 in lrk10l-1 (lane 1) compared to wild type background (lane 2). The single, double and triple asterisks in lanes 9 and 10 indicate the splicing events, intron-unspliced, IVS2-spliced and cryptic 3′ ss in IVS2, respectively (Accompanying paper)
Three different homozygous transgenic lines containing WT coding sequence of Lrk10L1.2 (a splicing variant designated No4; Accompanying paper) under the 35S promoter overexpress fully spliced WT mRNA that localizes to PM as expected (Fig. 1b, lanes 6–8). Transgenic lines containing a splicing variant designated Nu1, which had been spliced at a 5′ cryptic ss in IVS1, generated a variety of splicing variants including transcripts with the 45 nt long mini-exon 2 deletion (bottom bands in Fig. 1b lanes 9 and 10 indicated by an arrow). All other transcripts shown as higher molecular weight bands (asterisks in Fig. 1b lanes 9 and 10) contain unspliced intron sequences and premature termination codons immediately after their abnormal splicing events (Accompanying paper), so that the transcripts are unable to be translated into full-length proteins. Therefore, the in-frame exon 2 skip variant of LRK10L1.2 Nu1 produces a protein with 15 amino acids deletion by which it is targetted to the ER instead of the PM.
Molecular functions of Arabidopsis LRK10Ls have not been documented except for a few whole genome microarray data reporting that the transcript of LRK10L1 and its homolog At5g38210 (AtLRK10L3) are down-regulated in null mutants of FLOWERING LOCUS D (FLD) and LUMINIDEPENDENS (LD) (Veley and Michaels 2008), and a DELLA, key regulators of flowering time, and gibberellin and other stress responses (Cao et al. 2006), respectively. In addition, the At5g38210 transcription level is increased in sucrose starvation condition (Contento et al. 2004). These findings suggest that Arabidopsis LRK10Ls may also be involved in plant development and stress responses, apart from disease resistance. To investigate the function of AtLRK10L1.2, we analyzed above transgenic plants expressing 35S-Lrk10L1.2 No4 and 35S-Lrk10L1.2 Nu1 cDNAs in lrk10l1-2 as well as their mutant background lrk10l1-2, and the knock-down mutant lrk10l1-3 shown in Fig. 1b. In normal long-day growth condition, they did not exhibit any apparent phenotypic changes compared with the WT, except that lrk10l1-2 flowers approximately 3 days later than WT (Fig. S1a), and the transgenic plants overexpressing full-length LRK10L1.2 under the 35S promoter (35S-LRK10L1.2 No4) in turn flowers much earlier than WT (Fig. S1a). In addition, the LRK10L1.2 transcript level decreases in 3-week-old T-DNA insertion mutants of LD (Fig. S1c) showing a late flowering phenotype (data not shown). These results support previous whole genome microarray data (Veley and Michaels 2008), and suggest that AtLRK10L1.2 may be at least partly involved in flowering time regulation.
LRK10L1.2 is involved in ABA signaling
To explore whether LRK10L1.2 is involved in stress response, the above plants were treated with a variety of plant growth regulators including glucose, maltose and salts, such as NaCl and KCl, involved in ABA signaling (Fig. S2; Zhou et al. 1998; Rook et al. 2001; Leon and Sheen 2003). The mutants, lrk10l1-2 and lrk10l1-3, were less sensitive to more than 300 mM manitol and 200 mM NaCl than WT (Fig. S2). In these conditions, two independent lines of 35S-LRK10L1.2 No4 recovered WT phenotypes. However, glucose, which is involved in ABA signaling by delaying seed germination and cotyledon greening (Zhou et al. 1998), had an effect at the higher concentration of 6 %, in which 35S-LRK10L1.2 No4′s showed a more sensitive phenotype compared with WT. These findings suggest that LRK10L1.2 may be involved in ABA signaling.
Since ABA inhibits seed germination and controls the responses of environmental stresses in plants, we measured the germination rate in the seeds of WT, mutant and overexpressing plants to see whether LRK10L1.2 is involved in ABA signaling (Fig. 2a). Germination was tested under high stringency of 2.0 μM ABA. While germination rates of untreated seeds did not significantly differ between mock and ABA-treated conditions, the mutant seeds, lrk10l1-2 and lrk10l1-3, were less sensitive to ABA compared to the Col-0 WT. The transgenic lines 35S-LRK10L1.2 No4 overexpressing PM-targeting WT protein showed similar germination rates to WT seeds. However, 35S-Lrk10L1.2 Nu1, which produces ER-localized exon 2 skip mutant protein, did not show the WT phenotype and was less sensitive to ABA similar to the two mutants. Also, the growth and cotyledon greening of the WT was strongly inhibited by 0.5 μM ABA, while lrk10l1-2 and lrk10l1-3 mutant plants showed more cotyledon greening and expansion at this condition (Fig. 2b), suggesting these mutants are insensitive to ABA. Moreover, 35S-LRK10L1.2 Nu1 again was less sensitive to ABA. The cotyledon greening rates of the mutants and 35S-LRK10L1.2 Nu1 plants during the post-germinative growth phase were significantly higher than that in WT on ABA containing medium, but similar on ABA-free medium (Fig. 2c). By contrast, 35S-LRK10L1.2 No4 lines showed less cotyledon greening rate than the mutant plants, similar to WT plants. These findings suggest that LRK10L1.2 is involved in ABA signaling, and that the PM localization of LRK10L1.2 is essential for this process.
Enhanced ABA insensitivity of the mutants and the transgenic plant 35S-LRK10L1.2 Nu1 at the germination (a) and seedling establishment (b, c) stages. The seeds were germinated on MS agar plates containing 2.0 (a) and 0.5 μM ABA (b, c) and measured for rates of germination (a) and for green cotyledons of each line, 3 (a) and 7 days (b, c) after planting. Data in (a) and (b) are the mean ± standard deviation of three independent experiments, where 100 seeds were evaluated. c The representative images in (b) were taken
We further examined the ABA responses of WT, mutants, and over-expression lines at the seedling stage (Fig. 3). While treatment of seedlings with 0.5 μM ABA did not significantly affect seed germination rates, it inhibited root growth of WT and 35S-LRK10L1.2 No4 plants—the root lengths of WT seedlings were 40–60 % shorter than those of mutant and 35S-LRK10L1.2 Nu1, while those between WT and 35S-LRK10L1.2 No4 plants were similar (Fig. 3a, b). Together with cotyledon greening, these results suggest that response to ABA during seedling stage requires AtLRK10L1.2, and that AtLRK10L1.2 Nu1, which has a defect in the PM localization, cannot support either ABA sensing or signaling.
ABA-insensitive root growth of WT, mutant and transgenic plants in early seedling growth in 0.5 μM ABA. The seedlings were grown vertically in 0.5× MS containing 0.5 μM ABA for 7 days, and the primary root lengths of 50 seedlings of each line were measured (a), and the representative images were taken (b)
LRK10L1.2 is involved in resistance to drought stress
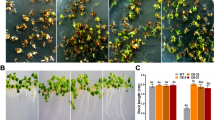
Since ABA response is critical in directing stomatal closure in response to shortage of water, we examined whether adult stages of the above plants respond differentially to drought conditions (Fig. 4). When grown in a well-watered conditions, both WT and mutant plants did not show any significant differences in phenotype. However, upon drought treatment by withholding watering, 35S-LRK10L1.2 Nu1 as well as the mutants, lrk10l1-2 and lrk10l1-3, exhibited hypersensitivity to drought compared with WT plants (Fig. 4ai, ii). However, there were no significant differences between WT and 35S-LRK10L1.2 No4 transgenic plants upon drought treatment (Fig. 4aii). In detail, approximately 85 % of WT and 35S-LRK10L1.2 No4 plants survived, while mutant plants lrk10l1-2 and lrk10l1-3, and 35S-LRK10L1.2 Nu1 had much lower survival rates ranging from 0 to approximately 18 %, where 0 % is observed from 35S-LRK10L-1.2 Nu1 (Fig. 4aii), indicating that resistance to drought stress is correlated with ABA sensitivity observed in Figs. 2 and 3, and that 35S-LRK10L1.2 Nu1 may work in a dominant negative way.
Response of lrk10l1.2 mutants and the transgenic plants to drought stress. a Representative images for “Watered” seedlings were taken before dehydration. Dehydration stress was given to WT and mutants, lrk10l1-2 and lrk10l1-3 (i), and WT and transgenic lines (ii) by withholding water for 7 days. The plants were then dehydrated for 2 days before they were photographed for “Drought”. b Transpiration rates of the WT, mutants and transgenic plants. Leaves were weighed at various time points after detaching. Data are the means ± standard errors (n = 10). c Stomatal aperture in leaves of WT, mutants and transgenic plants upon ABA treatment. Stomatal apertures were measured under the microscope after treatment with 10 μM ABA for 2 h 30 min as described in “Materials and methods”. Data are the means ± standard errors (n = 80)
To see whether hypersensitivity to drought stress in lrk10l1-2 and lrk10l1-3 mutants and 35S-LRK10L1.2 Nu1 transgenic plants are directly correlated with malfunctioning in stomatal closure, we determined the transpiration rates by measuring water loss from detached rosette leaves (Fig. 4b). Loss of fresh weight was greater in lrk10L1-2 and lrk10L1-2 mutants and 35S-LRK10L-1.2 Nu1 transgenic plants compared with those in WT and the 35S-LRK10L1.2 No4 transgenic plants, suggesting that the increased tolerance to drought is due, at least partly, to altered leaf transpiration (Fig. 4b).
Because the lrk10l1-2 and lrk10l1-3 exhibit altered transpiration rates, we presumed that the stomatal movement of these mutants may be altered. To determine whether LRK10L1.2 affects the responsiveness of guard cells to ABA, we measured stomatal apertures of leaves with or without ABA treatment (Fig. 4c). In the absence of ABA, there were no obvious differences in stomatal aperture among WT, mutants and transgenic plants. After treatment with 10 μM ABA, we observed the sizes of stomatal apertures were more increased in lrk10l1-2 and lrk10l1-3 and 35S-LRK10L1.2 Nu1 transgenic plants than those in WT and 35S-LRK10L1.2 No4 transgenic plants (Fig. 4c). These results indicate that defects in the LRK10L1.2 expression is responsible for decrease in ABA sensitivity of guard cells, leading to increased water loss under drought conditions.
LRK10L1.2 does not affect ABA marker gene expression
Multiple dehydration-inducible genes are induced by exogenous ABA treatment (Yamaguchi-Shinozaki and Shinozaki 2006), although the global transcriptional network activated in response to osmotic stress is cooperatively but not exclusively regulated by ABA-dependent and ABA-independent pathways.
We investigated the functional relationship of adaptation to drought stress with ABA by focusing on the LRK10L1.1 and the LRK10L1.2 transcripts (Fig. 5). As controls, levels of KIN1, RD22 and RAB18 transcripts were increasingly responsive to ABA and dehydration (Fig. 5) indicating both treatments are reliable. After dehydration and ABA treatment, the expression of LRK10L1.1 was clearly increased at 0.67 and 0.5 h upon drought and ABA treatments, respectively (Fig. 5). Interestingly, the level of the LRK10L1.2 transcript was decreased at the above time-points (Fig. 5). Drought treatment, which triggers endogenous ABA synthesis, showed clear distinction between LRK10L1.1 and LRK10L1.2. Down-regulation of LRK10L1.2 by drought and ABA treatment was accompanied by up-regulation of Lrk10L1.1 (Fig. 5). These results suggest that the LRK10L1.1 and the LRK10L1.2 transcripts may antagonistically regulate each other in drought stress as well as in ABA treatment.
Semi-quantitative RT-PCR analysis. Level of the LRK10L1.2 transcript decreases at early stages upon drought and ABA treatments. WT Col-0 plants were treated with drought and 50 μM ABA, before leaf samples were taken at indicated time (top) for RNA extraction. Gene-specific primers for the genes indicated on the left are listed in Table S1
Next, we performed qRT-PCR and semi-qRT-PCR analyses of WT, mutant and transgenic plants to determine involvement of the LRK10L1.2 gene in regulating the expression of ABA-signaling and drought-responsive genes under drought conditions. Unexpectedly overall expression level of these ABA-signaling and drought-responsive marker genes was unaffected across the mutants, WT and transgenic plants (Figs. S3, S4). We do not know why transgenic expression of LRK10L1.2 did not up-regulate expression of these genes. However, this finding suggests that ABA signaling through LRK10L1.2 may be unrelated in up-regulation of genes containing ABA-responsive elements (ABRE).
Discussion
We report here that the Arabidopsis homolog of wheat LRK10 produces two independent proteins by use of alternative promoters that share a common serine/threonine protein kinase domain, and that the second variant, LRK10L1.2, produces a PM protein involved in ABA signaling. The biological function of the first variant LRK10L1.1 is still unknown, since the mutant lrk10l-1 (Fig. 1) containing a T-DNA in exon 1 of At1g18390.1 showed a phenotype indistinguishable from WT upon ABA or drought treatment (data not shown). It is noticeable, however, that transcription of LRK10L1.1 antagonistically regulates that of LRK10L1.2 in the T-DNA insertion in exon 1 of LRK10L1.1 (Fig. 1b, lanes 1 and 2). This observation suggests that expression of LRK10L1.2 is in part regulated by LRK10L1.1 promoter.
Drought, high salinity, and low temperature are common water-deficit conditions that adversely affect plant growth. Many physiological changes in response to such stresses result from the plant hormone ABA that regulates expression of stress-related genes and that in turn leads to various adaptive responses at the cellular and whole plant levels (Shinozaki and Yamaguchi-Shinozaki 2000; Ramanjulu and Bartels 2002). ABA is mainly involved in adaptation to water-deprived states through regulatory circuits that control gene expression and stomatal closure (Luan 2002; Zhu 2002; Wasilewska et al. 2008). The cellular and molecular mechanisms underlying ABA-induced stomatal closure have been extensively studied (Cutler et al. 2010; Hubbard et al. 2010). For example, when ABA level increases in drought conditions, anion efflux via the anion channels induces depolarization and activation of outward K+-channels (Ward et al. 2008; Lee et al. 2009; Kim et al. 2010). Reduced ionic concentration in the cell causes water efflux and reduces guard cell volume thereby leading to stomatal closure (Ward et al. 1995; Wasilewska et al. 2008). The physiological and molecular mechanisms explaining such stress conditions are well established (Zhu 2002; Yamaguchi-Shinozaki and Shinozaki 2006), but detailed functional modifications caused by these stresses still less understood.
Nevertheless, the ABA signaling pathways via the ABA sensor PYRABACTIN RESISTANCE (TYR)/REGULATORY COMPONENT OF ABA RECEPTOR (TYR/RCAR) have been well documented, in which PYR/RCAR forms complexes with type A-protein phosphatase 2Cs (PP2CAs) such as ABA-INSENSITIVE 1 (ABI1) in the presence of ABA (Park et al. 2009; Ma et al. 2009). In this condition, PP2CA is unable to inhibit a subset of SNF1-related protein kinase 2’s (SnRK2’s; Fujii et al. 2009), which then phosphorylate ion channels and transcription factors that stimulate transcription of ABA-reponsive genes (Choi et al. 2000). We observed that known ABA-and drought-induced genes are unchanged across WT, lrk10l1-2 and complemented lines (Figs. S3, S4), suggesting that ABA signaling through LRK10L1.2 may work independently of PYR/RCAR-PP2C-SnRK2 pathway.
The gene encoding the wheat LRK10 has been cloned by homology-based cloning based on disease resistance against the leaf rust-causing fungal pathogen Puccinia recondita (Feuillet et al. 1997). Hence, the Arabidopsis ortholog LRK10L1.2 may be potentially involved in defense response to pathogens. A number of plant hormones, including ABA, jasmonic acid (JA) and salicylic acid (SA), play a role in plant defense response. In addition, several lines of evidence showed that ABA signaling is connected to the jasmonate and SA signalings, which is associated with necrotrophic and biotrophic pathogens, respectively. (Anderson et al. 2004; Lorenzo et al. 2004; Pieterse et al. 2009). In this study, however, we only elucidated the function of LRK10L1.2 in response to abiotic stress. Therefore, further studies are needed to understand the other function of LRK10L1.2 in defense response to biotic stress.
In conclusion, this study showed that LRK10L1.2 acts as a positive regulator in response to drought tolerance via the induction of stomatal closing, which may occur directly or indirectly via ABA-mediated signaling. Although we revealed the functional involvement of LRK10L1.2 in plant response to drought stress, it is still unclear how LRK10L1.2 serves as a positive regulator of abiotic stress response. Subsequent molecular analysis of the downstream target of LRK10L1.2 will improve our understanding of the function of LRK10L1.2 under biotic and abiotic stress conditions.
Author contribution statement
S. C. L. and S. H. K. designed research; C. W. L., S. H. Y. and K. H. S. performed research; S. C. L. and S. H. K. analyzed data and; S. C. L. and S. H. K wrote the paper.
Abbreviations
- ABA:
-
Abscisic acid
- AS:
-
Alternative splicing
- ER:
-
Endoplasmic reticulum
- IVS:
-
Intervening sequence
- LRK10L:
-
Leaf rust 10 disease-resistance locus receptor-like protein kinase-like
- PM:
-
Plasma membrane
- WT:
-
Wild type
- YFP:
-
Yellow fluorescent protein
References
Anderson JP, Badruzsaufari E, Schenk PM, Manners JM, Desmond OJ, Ehlert C, Maclean DJ, Ebert PR, Kazan K (2004) Antagonistic interaction between abscisic acid and jasmonate–ethylene signaling pathways modulates defense gene expression and disease resistance in Arabidopsis. Plant Cell 16:3460–3479
Bai L, Zhang G, Zhou Y, Zhang Z, Wang W, Du Y, Wu Z, Song CP (2009) Plasma membrane-associated proline-rich extensin-like receptor kinase 4, a novel regulator of Ca signalling, is required for abscisic acid responses in Arabidopsis thaliana. Plant J 60:314–327
Becraft PW, Stinard PS, McCarty DR (1996) CRINKLY4 – a TNFR-like receptor kinase involved in maize epidermal differentiation. Science 273:1406–1409
Cao D, Cheng H, Wu W, Soo HM, Peng J (2006) Gibberellin mobilizes distinct DELLA-dependent transcriptomes to regulate seed germination and floral development in Arabidopsis. Plant Physiol 142:509–525
Choi H, Hong J, Ha J, Kang J, Kim SY (2000) ABFs, a family of ABA-responsive element binding factors. J Biol Chem 275:1723–1730
Clark SE, Williams RW, Meyerowitz EM (1997) The CLAVATA1 gene encodes a putative receptor kinase that controls shoot and floral meristem size in Arabidopsis. Cell 89:575–585
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Contento AL, Kim SJ, Bassham DC (2004) Transcriptome profiling of the response of Arabidopsis suspension culture cells to Suc starvation. Plant Physiol 135:2330–2347
Cutler SR, Rodriguez PL, Finkelstein RR, Abrams SR (2010) Abscisic acid: emergence of a core signaling network. Annu Rev Plant Biol 61:651–679
Feuillet C, Schachermayr G, Keller B (1997) Molecular cloning of a new receptor-like kinase gene encoded at the Lr10 disease resistance locus of wheat. Plant J 11:45–52
Fujii H, Chinnusamy V, Rodrigues A, Rubio S, Antoni R, Park S-Y, Cutler SR, Sheen J, Rodriguez PL, Zhu J-K (2009) In vitro reconstitution of an abscisic acid signalling pathway. Nature 462:660–664
Gish LA, Clark SE (2011) The RLK/Pelle family of kinases. Plant J 66:117–127
Gomez-Gomez L, Boller T (2000) FLS2: an LRR receptor-like kinase involved in the perception of bacterial elicitor flagellin in Arabidopsis. Mol Cell 5:1003–1011
He ZH, Cheeseman I, He D, Kohorn BD (1999) A cluster of five cell wall-associated receptor kinase genes, Wak1-5, are expressed in specific organs of Arabidopsis. Plant Mol Biol 39:1189–1196
Hua D, Wang C, He J, Liao H, Duan Y, Zhu Z, Guo Y, Chen Z, Gong Z (2012) A plasma membrane receptor kinase, GHR1, mediates abscisic acid- and hydrogen peroxide-regulated stomatal movement in Arabidopsis. Plant Cell 24:2546–2561
Hubbard KE, Nishimura N, Hitomi K, Getzoff ED, Schroeder JI (2010) Early abscisic acid signal transduction mechanisms: newly discovered components and newly emerging questions. Genes Dev 24:1695–1708
Kim SH, Koroleva OA, Lewandowska D, Pendle AF, Clark GP, Simpson CG, Shaw PJ, Brown JWS (2009) Aberrant mRNA Transcripts and the Nonsense-Mediated Decay Proteins UPF2 and UPF3 Are Enriched in the Arabidopsis Nucleolus. Plant Cell 21:2045–2057
Kim TH, Bohmer M, Hu H, Nishimura N, Schroeder JI (2010) Guard cell signal transduction network: advances in understanding abscisic acid, CO2, and Ca2+ signaling. Annu Rev Plant Biol 61:561–591
Kwak SH, Shen R, Schiefelbein J (2005) Positional signaling mediated by a receptor-like kinase in Arabidopsis. Science 307:1111–1113
Lee SC, Lan W, Buchanan BB, Luan S (2009) A protein kinase-phosphatase pair interacts with an ion channel to regulate ABA signaling in plant guard cells. Proc Natl Acad Sci USA 106:21419–21424
Leon P, Sheen J (2003) Sugar and hormone connections. Trends Plant Sci 8:110–116
Limpens E, Franken C, Smit P, Willemse J, Bisseling T, Geurts R (2003) LysM domain receptor kinases regulating rhizobial Nod factor-induced infection. Science 302:630–633
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta C(T)) method. Methods 25:402–408
Lorenzo O, Chico JM, Sánchez-Serrano JJ, Solano R (2004) JASMONATE-INSENSITIVE1 encodes a MYC transcription factor essential to discriminate between different jasmonate-regulated defense responses in Arabidopsis. Plant Cell 16:1938–1950
Luan S (2002) Signalling drought in guard cells. Plant Cell Environ 25:229–237
Ma Y, Szostkiewicz I, Korte A, Moes D, Yang Y, Christmann A, Grill E (2009) Regulators of PP2C phosphatase activity function as abscisic acid sensors. Science 324:1064–1068
Nakhamchik A, Zhao Z, Provart NJ, Shiu SH, Keatley SK, Cameron RK, Goring DR (2004) A comprehensive expression analysis of the Arabidopsis proline-rich extensin-like receptor kinase gene family using bioinformatic and experimental approaches. Plant Cell Physiol 45:1875–1881
Nodine MD, Yadegari R, Tax FE (2007) RPK1 and TOAD2 are two receptor-like kinases redundantly required for Arabidopsis embryonic pattern formation. Dev Cell 12:943–956
Osakabe Y, Maruyama K, Seki M, Satou M, Shinozaki K, Yamaguchi-Shinozaki K (2005) Leucine-rich repeat receptor-like kinase1 is a key membrane-bound regulator of abscisic acid early signaling in Arabidopsis. Plant Cell 17:1105–1119
Park SY, Fung P, Nishimura N, Jensen DR, Fujii H, Zhao Y, Lumba S, Santiago J, Rodrigues A, Chow TF, Alfred SE, Bonetta D, Finkelstein R, Provart NJ, Desveaux D, Rodriguez PL, McCourt P, Zhu JK, Schroeder JI, Volkman BF, Cutler SR (2009) Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 324:1068–1071
Pieterse CM, Leon-Reyes A, Van der Ent S, Van Wees SC (2009) Networking by small-molecule hormones in plant immunity. Nat Chem Biol 5:308–316
Ramanjulu S, Bartels D (2002) Drought- and desiccation-induced modulation of gene expression in plants. Plant Cell Environ 25:141–151
Ringli C (2010) The hydroxyproline-rich glycoprotein domain of the Arabidopsis LRX1 requires Tyr for function but not for insolubilization in the cell wall. Plant J 63:662–669
Rook F, Corke F, Card R, Munz G, Smith C, Bevan MW (2001) Impaired sucrose induction mutants reveal the modulation of sugar-induced starch biosynthetic gene expression by abscisic acid signalling. Plant J 26:421–433
Searle IR, Men AE, Laniya TS, Buzas DM, Iturbe-Ormaetxe I, Carroll BJ, Gresshoff PM (2003) Long-distance signaling in nodulation directed by a CLAVATA1-like receptor kinase. Science 299:109–112
Shinozaki K, Yamaguchi-Shinozaki K (2000) Molecular responses to dehydration and low temperature: differences and cross-talk between two stress signaling pathways. Curr Opin Plant Biol 3:217–223
Shiu SH, Bleecker AB (2001) Receptor-like kinases from Arabidopsis form a monophyletic gene family related to animal receptor kinases. Proc Natl Acad Sci USA 98:10763–10768
Shiu SH, Bleecker AB (2003) Expansion of the receptor-like kinase/Pelle gene family and receptor-like proteins in Arabidopsis. Plant Physiol 132:530–543
Stein JC, Howlett B, Boyes DC, Nasrallah ME, Nasrallah JB (1991) Molecular cloning of a putative receptor protein kinase gene encoded at the self-incompatibility locus of Brassica oleracea. Proc Natl Acad Sci USA 88:8816–8820
Tanaka H, Osakabe Y, Katsura S, Mizuno S, Maruyama K, Kusakabe K, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) Abiotic stress-inducible receptor-like kinases negatively control ABA signaling in Arabidopsis. Plant J 70:599–613
Veley KM, Michaels SD (2008) Functional redundancy and new roles for genes of the autonomous floral-promotion pathway. Plant Physiol 147:682–695
Wang XQ, Zafian P, Choudhuary M, Lawton M (1996) The PRK5 receptor protein kinase from Arabidopsis thaliana is structurally related to a family of plant defense proteins. Proc Natl Acad Sci USA 93:2598–2602
Ward JM, Pei ZM, Schroeder JI (1995) Roles of Ion Channels in Initiation of Signal Transduction in Higher Plants. Plant Cell 7:833–844
Ward JM, Maser P, Schroeder JI (2008) Plant Ion Channels: gene Families, Physiology, and Functional Genomics Analysis. Annu Rev Physiol 71:59–82
Wasilewska A, Vlad F, Sirichandra C, Redko Y, Jammes F, Valon C, Frey NFd, Leung J (2008) An update on abscisic acid signaling in plants and more. Mol Plant 1:198–217
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803
Zhang X, Yang G, Shi R, Han X, Qi L, Wang R, Xiong L, Li G (2013) Arabidopsis cysteine-rich receptor-like kinase 45 functions in the responses to abscisic acid and abiotic stresses. Plant Physiol Biochem 67:189–198
Zhou L, Jang JC, Jones TL, Sheen J (1998) Glucose and ethylene signal transduction crosstalk revealed by an Arabidopsis glucose-insensitive mutant. Proc Natl Acad Sci USA 95:10294–10299
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Acknowledgments
This research was supported by a grant from the BioGreen 21 Program, Rural Development Administration of Republic of Korea (PJ00948402) to S. H. K. and (PJ00822201) to S. C. L., and the National Research Foundation (NRF) of Korea Grant funded by the Korean Government (MOE) (NRF-2011-0029568) to S. H. K. and (No. 2010-0024596) to S. H. Y.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by Youn-Il Park.
C. W. Lim and S. H. Yang contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
299_2014_1724_MOESM1_ESM.tif
Fig. S1 a Phenotypes of WT, lrk10l-2, and 35S-Lrk10L1.2 transgenic plants in long day growth condition. Thirty-day-old plants were photographed when the primary inflorescence stems of lrk10l1-2 are 0.5 cm long. b T-DNA insertion sites of ld insertional mutants (Salk_145773 and Salk_003047). The 5´ and 3´ UTR’s and CDS are drawn by open and closed boxes, respectively. Introns shown by solid lines. c RT-PCR analysis for the LRK10L1.2 transcript from WT (Col-0) and two T-DNA insertion mutants using the primers shown in Fig. 1a (P2 and P3). (TIFF 14020 kb)
299_2014_1724_MOESM2_ESM.tif
Fig. S2 Seedling growth of WT, mutants and transgenic plants in response to mannitol, glucose, and NaCl. The seedlings were grown in 0.5× MS containing different concentrations of mannitol and glucose, and 200 mM NaCl. The representative images were taken 7 days (for 300 mM manitol, 4 and 5 % glucose) or after 10 days (for 400 mM Mannitol, 6 % glucose, and 200 mM NaCl) after stratification. (TIFF 11031 kb)
299_2014_1724_MOESM4_ESM.tif
Fig. S4 LRK10L1.2 expression does not affect transcription of ABA marker genes, KIN1, RD22 and RAB18. Different plant samples indicated on the left were treated and collected as in Fig. 5 for RNA extraction. Gene-specific primers for the genes indicated on the right are listed in Table. S1. LRK10L1.2, the WT background of lrk10l1-2; No4, 35S-Lrk10L1.2 No4/lrk10l1-2 (Line 7) and; Nu1, 35S-Lrk10L1.2 Nu1/lrk10l1-2 (Line14). (TIFF 9488 kb)
Rights and permissions
About this article
Cite this article
Lim, C.W., Yang, S.H., Shin, K.H. et al. The AtLRK10L1.2, Arabidopsis ortholog of wheat LRK10, is involved in ABA-mediated signaling and drought resistance. Plant Cell Rep 34, 447–455 (2015). https://doi.org/10.1007/s00299-014-1724-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-014-1724-2