Abstract
Resistance to the antibiotic Cycloheximide has been reported for a number of fungal taxa. In particular, some yeasts are known to be highly resistant to this antibiotic. Early research showed that this resulted from a transition mutation in one of the 60S ribosomal protein genes. In addition to the yeasts, most genera and species in the Ophiostomatales are highly resistant to this antibiotic, which is widely used to selectively isolate these fungi. Whole-genome sequences are now available for numerous members of the Ophiostomatales providing an opportunity to determine whether the mechanism of resistance in these fungi is the same as that reported for yeast genera such as Kluyveromyces. We examined all the available genomes for the Ophiostomatales and discovered that a transition mutation in the gene coding for ribosomal protein eL42, which results in the substitution of the amino acid Proline to Glutamine, likely confers resistance to this antibiotic. This change across all genera in the Ophiostomatales suggests that the mutation arose early in the evolution of these fungi.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Ophiostomatales (Ascomycetes) are best known as arthropod associated fungi that include important pathogens of trees such as the Dutch elm disease fungi Ophiostoma ulmi and Ophiostoma novo-ulmi (Brasier1991; Gibbs 1978), human and animal pathogens in the genus Sporothrix (de Beer et al. 2003; Rodrigues et al. 2016) and agents of sap-stain in lumber (Seifert 1993). An unusual characteristic of species in the Ophiostomatales is that they are consistently highly tolerant to the antibiotic cycloheximide. This biochemical characteristic was initially recognized by Fergus (1956) who showed that some wood staining species of Ophiostoma shared this feature.
Fungi in the Ophiostomatales have had a long and complex taxonomic history. This has more specifically concerned to the separation of the genera Ophiostoma and Ceratocystis and their relatives (de Hoog and Scheffer 1984; Wingfield et al. 1993; Seifert et al. 2015). Confusion regarding the generic boundaries of these fungi dates back to a time when their taxonomy relied almost exclusively on morphology (Uphadhyay 1991; Wingfield et al. 1993). Specifically, their various shared morphological characteristics, arising from convergent evolution that facilitates associations with arthropod vectors resulted in confusion regarding the appropriate taxonomic boundaries between the genera Ophiostoma and Ceratocystis, which were collectively referred to as the Ophiostomatoid fungi (Wingfield et al. 1993; Seifert et al. 2015).
For many years, cycloheximide tolerance provided a useful non-morphological characteristic that clearly separated species related to Ceratocystis from those related to Ophiostoma (Harrington 1981). The more recent emergence of DNA sequence-based phylogenies has strongly supported the fact that these two groups of fungi are unrelated and reside, respectively, in unrelated Orders (Hausner et al. 1993a, b; Spatafora and Blackwell 1994). These are the Ophiostomatales defined by Ophiostoma sensu lato (de Beer et al. 2013) and the Microscales including genera in the Ceratocystidaceae (de Beer et al. 2014) and the Gondwanamycetaceae including species of Knoxdaviesia (Réblová et al. 2011). A recent revision of the Ophiostomatales based on multiple gene genealogies as well as whole genome data (de Beer et al. 2022) has defined 16 genera including all those species that have, in various studies, been shown to tolerate high levels of cycloheximide in culture.
Cycloheximide is a powerful antibiotic that is not generally applied for medical purposes. It is, however, commonly used in research experiments to inhibit translation of messenger RNA and thus protein synthesis in eukaryotic cells. For example, Rao and Grollman (1967) showed that its mechanism of action was associated with the 60S ribosomal subunit in Saccharomyces cerevisiae. Studies using S. cerevisiae and Tetrahymena thermophila mutants with low levels of resistance to cycloheximide showed that this was the result of an amino acid substitution in the ribosomal protein L29 (Käufer et al. 1983; Yao and Yao 1991).
Most Eukaryotes are sensitive cycloheximide. There are, however, various exceptions, other than in the Ophiostomatales mentioned above, such as in some ascomycetous yeasts (Saccharomycetaceae). For example, resistance to the antibiotic in species of Kluyveromyces, Candida and Schwanniomyces has been shown to result from the substitution of a Glutamine (Gln) in the place of a Proline (Pro) at position 56 in the ribosomal protein L41 (Dehoux et al. 1993; Sasnauskas et al. 1992). Using genetic transformants of Saccharomyces cerevisiae, Kawai et al. (1992) showed that a Pro to Gln change in the ribosomal protein L41 results in resistance to cycloheximide at concentrations of 100 μg/ml. More recently, Shen et al. (2021) have shown the importance of ribosomal protein eL42 in resistance to cycloheximide by Neurospora crassa. Likewise in the green alga Chlamydomonas reinhardtii, mutants with point mutations in the ribosomal protein gene L41 (RPL41) where a Proline at position 56 has been replaced with either Leucine or Serine are also resistant to cycloheximide (Stevens et al. 2001). The Leucine mutation in this case results in higher levels of resistance.
There is a reasonably robust literature showing that cycloheximide resistance arises from amino acid substitutions in specific ribosomal proteins. A complication in understanding this trait arises from the fact that the ribosomal proteins have been named variously for the prokaryotes and eukaryotes in the past (Wittmann et al. 1971; Kruiswijk and Planta 1974; Wool et al. 1995). Thus, to compare the names of these proteins in different publications, it is necessary to be aware of their variable nomenclature. Specifically, and pertinent to this study, ribosomal protein L41 was renamed L42 (Planta and Mager 1998) and is now referred to as eL42 (Ban et al. 2014). Thus references to substitutions in ribosomal protein L41 are most correctly referred to as being in ribosomal protein eL42.
In the recent revision of the Ophiostomatales, de Beer et al. (2022) included Genome sequences for 31 species representing 11 of 14 currently recognized genera (excluding Afroraffaelea, Aureovirgo and Paleoambrosia). The availability of these genome sequences has provided an opportunity to determine the basis of their resistance to cycloheximide and whether this might be similar to that described in many yeasts. The aim of this study was thus to use the available Ophiostomatales genome sequences to identify the amino acid sequence of the ribosomal protein eL42. Consequently, to determine whether the predicted amino acid Proline at position 56 has been substituted by Glutamine or some other amino acid.
Materials and methods
Taxon sampling and genome collection
To provide a phylogenomic framework for this study, a genome data set was assembled and analysed including 69 genomes for species in the Sordariomycetes and the Saccharomycetes. This dataset included all currently available genome sequences for genera in the Ophiostomatales (Ceratocystiopsis, Chrysosphaeria, Esteya, Fragosphaeria, Graphilbum, Grosmannia, Hawksworthiomyces, Intubia, Leptographium, Ophiostoma, Sporothrix and Raffaelea) and thus fungi known or expected to be tolerant to cycloheximide. For comparative purposes, genomes for representative genera in the Microascales including the Ceratocystidaceae (Ambrosiella, Bretziella, Catunica, Ceratocystis, Davidsoniella, Endoconidiophora, Huntiella and Thielaviopsis), Gondwanamycetaceae (Knoxdaviesia) and Microascaceae (Microascus) were included. With the exception of Microascus, these are known to be sensitive to the antibiotic. In addition, genomes for a selection of other Sordariomycetes genera reported to be cycloheximide sensitive (Colleototrichum, Cryphonectria, Diaporthe, Fusarium, Geosmithia, Magnaporthe, Neurospora, Phaeoacremonium, Thielavia and Trichoderma) were also included. To accommodate yeasts (Saccharomycetes) 16 species in 12 genera (Ascoidea, Brettanomyces, Candida, Eremothecium, Komagataella, Kluyveromyces, Lachancea, Ogataea, Pachysolen, Pichia, Saccharomyces and Saccharomycopsis), some of which are known to be either sensitive or tolerant to cycloheximide, were included (Table1). All genome sequences were downloaded from JGI Genome Portal or NCBI genome databases with accession numbers and references provided in Table 1.
Phylogenomic analyses
All genome sequences were subjected to BUSCO v4.0.5 analysis using the ascomycota_odb10 dataset (Seppey et al. 2019). Single copy BUSCO genes that were shared across all 69 species were identified and these were used to construct a species tree utilizing a coalescence approach. The amino acid sequences for each BUSCO gene were aligned with PRANK v.170427 (Löytynoja 2014) using the default parameters and trimmed with Trimal v1.4 (Capella-Gutiérrez et al. 2009) with the “automated1” option. After trimming, an additional filtering step was carried out to remove datasets with less than 100 sites in alignment length or less than 50 parsimony-informative characters. Datasets that did not include all taxa after the aligning and trimming steps were also excluded from further analyses.
Maximum likelihood trees were constructed on the remaining datasets using IQTREE v1 with automatic model selection and 1000 ultrafast bootstrap replicates (Hoang et al. 2018; Minh et al. 2020). After collapsing the branches having less than 10% bootstrap support from individual gene trees using Newick Utilities (Junier and Zdobnov 2010), the species phylogeny was inferred from the resulting gene trees in ASTRAL v5.7.7 (Mirarab et al. 2014). Finally, RaxML v 8.2.11 (Stamatakis 2014) was applied to estimate branch length for the species phylogeny with the concatenated alignment of all BUSCO genes used for species tree constructions.
Ribosomal protein eL42 annotation and comparison
Protein coding genes present in all genomes were predicted with Augustus v3.2.3 (Stanke et al. 2006) using the species models for Neurospora crassa and Kluyveromyces lactis as the representatives for taxa in the Sordariomycetes and the Saccharomycetes, respectively. Genes encoding the ribosomal protein eL42 were identified by carrying out a BLASTP analysis with the Kluyveromyces lactis ribosomal protein eL42 (GenBank accession M94988.1) as query against a protein database consisting of all amino acid sequences obtained with Augustus prediction of all 69 genomes. The genome sequences and DNA sequences of eL42 gene were extracted from all species and these were aligned in MAFFT v7 with the E-INS-i option (Katoh and Standley 2013). The resulting alignment was then used to verify and manually curate (where necessary) the protein coding sequences of the eL42 genes from all species. Finally, the eL42 amino acid sequences all species were aligned in MAFFT v7 (Katoh and Standley 2013) and the alignment was visualized on the phylogenomic tree with iTOL v4 (Letunic and Bork 2019).
Results
Phylogenomic tree construction
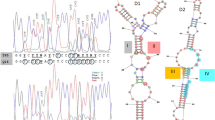
A total of 312 shared single copy BUSCO genes were identified across 69 species, 248 of which were retained for the construction of the species phylogeny. The phylogenomic tree inferred with ASTRAL showed two major lineages represented by species of the Saccharomycetes and Sordariomycetes, respectively (Fig. 1). The evolutionary relationships of species residing in the Saccharomycetes included in this study were consistent with those in the phylogeny produced by Krassowski et al. 2018. The Sordariomycete and Saccharomycete lineages grouped together, species in the Ophiostomatales formed a monophyletic clade and species in Microascales including the Ceratocystidaceae, Gondwanamycetaceae and Microascus grouped together.
Phylogenomic tree of all the species in this study. The red stars indicate species that are known to be resistant to cycloheximide. Presence of Glutamine (Q) or Proline (P) at amino acid position 56 in the ribosomal protein eL42 indicated. Glutamine (Q) is present only in species that are resistant to cycloheximide, whereas a Proline (P) is present in species that are known to be susceptible to this antibiotic (except for Microascus trigonosporus)
Ribosomal protein eL42 annotation and comparison
A single gene encoding for the eL42 protein was predicted from each of the 69 genomes included in this study. The total length of the predicted protein was 100 amino acids in all species investigated. The amino acid alignment of the protein sequence displayed a high level of conservation (supplementary Fig. 1) across species in the Saccharomycetes and those in the Sordariomycetes. There were, however, a range of introns present in the predicted gene sequences, from one (in species of the Saccharomycetes) to six in Knoxdaviesia protea (Sordariomycetes).
All species of the Ophiostomatales had a Glutamine (Q) at position 56 in the eL42 protein (Fig. 1). In contrast, all species in the Sordariomycetes known to be cycloheximide sensitive including those in the Ceratocystidaceae and Gondwanamycetaceae had a Proline (P) at position 56. In the case of Microascus trigonosporus, which is known to be cycloheximide resistant, there was a Proline (P) at position 56. All the other species in the Sordariomycetes included in this study have a Proline (P) at position 56 in the eL42 protein and are known to be cycloheximide sensitive. All the species in the Saccharomycetes with known resistance to cycloheximide had a Glutamine at position 56, in contrast to a Proline at this position for species that are susceptible to the antibiotic.
There were three additional amino acid substitutions in the predicted eL42 protein that are shared between the Ophiostomatales, but not present in the close relatives utilized as outgroups in this study. These were Threonine at positions 30 and 88 and Lysine at position 81. These amino acid differences are not shared with yeasts known to be highly resistant to cycloheximide and that have the Glutamine substitution in position 56 of eL42. These additional amino acid differences are thus unlikely to be linked to cycloheximide resistance in the Ophiostomatales.
Discussion
Cycloheximide resistance has been well known in species of the Ophiostomatales for many years. However, the molecular basis of this characteristic has never been considered. In this study, we were able to show that cycloheximide tolerance in these fungi is due to a substitution of the amino acid Proline in the ribosomal protein eL42 at position 56 with a Glutamine. This is the same as has been shown in various species of yeasts where the Proline at position 56 in eL42 is replaced with a Glutamine (deHoux et al. 1993; Sasnauskas et al. 1992).
We included in this study an analysis of the ribosomal protein eL42 in species of the Microscales, more specifically the Ceratocystidaceae and Gondwanamycetaceae. This was due to the long-standing confusion between members of these Families and the Ophiostomatales in the past. Unsurprisingly, none of these species had the Glutamine substitution in position 56 of eL42. This confirms the molecular basis of cycloheximide sensitivity in these fungi, which has been well known for those species and for which the trait has previously been tested (Harrington 1981).
Microascus trigonosporus was included in this study due to its placement in the Microscales and thus its relationship with the Ceratocystidaceae and Gondwanamycetaceae. This fungus is a dermatophyte and has been established as cycloheximide resistant in previous studies (Brasch et al. 2019). The fact that M. trigonosporus has a Proline at position 56 in the predicted protein eL42 suggests that the resistance of this fungus to cycloheximide is not as a consequence of a change in the protein eL42, but rather due to a different mechanism. Given that its close relatives in the Ceratocystidaceae and Gondwanamycetaceae are sensitive to cycloheximide, a different molecular basis for the trait is perhaps not surprising. This could for example be due to overexpression of the ATP-binding cassette (ABC) transporters, (Moran et al. 1998), the presence of the multi-drug resistance MDR 1 gene (Gupta et al. 1998) or the ability to convert cycloheximide to a less toxic derivative (Shearer and Sypherd 1988). Interestingly, this last form of resistance is not limited to fungi but has also been reported in carrot cell culture (Sung et al. 1981). Additionally, Shen et al. (2021) report a number of amino acid substitutions in ribosomal proteins that result in cycloheximide resistance in Neurospora crassa. In the case of eL42 these were P56L and F58L and for uL15 they reported two different mutations, Q38K and Q38L. None of these mutations are found in the genome of M. trigonosporus. Further research to determine the molecular basis of cycloheximide tolerance in M. trigonosporus is likely to yield interesting and useful findings.
Numerous yeasts, relatively widely distributed across the Saccharomycetaceae are known to be highly resistant to cycloheximide and the results of the present study are consistent with that fact. In the case of Candida and Kluyveromyces, cycloheximide resistance is the result of a single amino acid substitution in eL42 (Dehoux et al. 1993; Sasnauskas et al. 1992). This is the ribosomal protein to which cycloheximide binds and that underpins its mode of action. It is, therefore, not surprising that species in the Ophiostomatales, known to be highly resistant to this antibiotic have a substitution in the same ribosomal protein. What was perhaps unexpected is that the substitution is exactly the same as that found in various yeast taxa. In this regard, it suggests that the mutation allows for a functional protein but that also provides cycloheximide resistance. What is also interesting is that the amino acid (Glutamine), which is substituted in the Ophiostomatales, is also the same as that observed in yeasts. It seems likely that other cycloheximide resistant eukaryotes would have this same mutation and that this would have then arisen separately in different lineages.
It is particularly relevant that all species in the Ophiostomatales are tolerant to high levels of cycloheximide. This is a relatively large Order of the fungi and there are no known exceptions. The situation in the yeasts is different where this biological characteristic is present variously across the Saccharomycetales without any clear pattern of occurrence. This suggests that there has been a selection for cycloheximide tolerance early in the evolution of the Ophiostomatales and that this selective pressure has been maintained over a long evolutionary history. In contrast, the occurrence of this trait across the ascomycetous yeasts suggests that it has thus either arisen independently in different lineages or been lost across evolutionary time in lineages, where there is no selective pressure to maintain it.
The results of this and previous studies provide robust evidence that all species in the Ophiostomatales are highly tolerant to cycloheximide. This implies that there has been strong evolutionary pressure across a relatively large assemblage of fungi to maintain this unique characteristic. The Ophiostomatales are well-known associates of arthropods including various groups of insects and mites (Wingfield et al 2017b) and it is reasonable to speculate that cycloheximide tolerance has contributed to the establishment of this niche. Some evidence supporting this view emerges from the close association of between some wood boring beetles and Streptomyces (Actinomycetes) that produce cycloheximide (Grubbs et al. 2020). While this might only be a limited example, the fact that most if not all Ophiostomatales likely have some association with arthropods, including those such as mites that occur in soils, suggests that they have evolved in an environment rich in cycloheximide or together with organisms that produce this antibiotic. Further understanding this relationship is likely to be lucrative in new scientific discovery.
Data availability
The genome data used in this study are available in the NCBI repository, https://www.ncbi.nlm.nih.gov. The accession numbers for all genomes are indicated in Table 1. The Microascus genome data is available on the JGI mycocosm website (https://mycocosm.jgi.doe.gov/mycocosm/home). The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Aylward J, Steenkamp ET, Dreyer LL, Roets F, Wingfield BD et al (2016) Genome sequences of Knoxdaviesia capensis and K. proteae (Fungi: Ascomycota) from Protea trees in South Africa. Stand Genomic Sci 11:22. https://doi.org/10.1186/s40793-016-0139-9
Ban N, Beckmann R, Cate JH, Dinman JD, Dragon F (2014) A new system for naming ribosomal proteins. Curr Opin Struct Biol. https://doi.org/10.1016/j.sbi.2014.01.002
Berka RM, Grigoriev IV, Otillar R, Salamov A, Grimwood J et al (2011) Comparative genomic analysis of the thermophilic biomass-degrading fungi Myceliophthora thermophila and Thielavia terrestris. Nat Biotechnol. https://doi.org/10.1038/nbt.1976
Blanco-Ulate B, Rolshausen P, Cantu D (2013) Draft Genome Sequence of the Ascomycete Phaeoacremonium aleophilum Strain UCR-PA7, a causal agent of the Esca disease complex in grapevines. Genome Announc. https://doi.org/10.1128/genomeA.00390-13
Brasch J, Beck-Jendroschek V, Iturrieta-González I, Voss K, Gené J (2019) A human subcutaneous infection by Microascus ennothomasiorum sp. nov. Mycoses 62:157–164. https://doi.org/10.1111/myc.12861
Brasier CM (1991) Ophiostoma novo-ulmi sp. nov., causative agent of current Dutch elm disease pandemics. Mycopathologia 115:151–161. https://doi.org/10.1007/BF00462219
Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics. https://doi.org/10.1093/bioinformatics/btp348
Choo JH, Hong CP, Lim JY, Seo A, Kim YS et al (2016) Whole-genome de novo sequencing, combined with RNA-Seq analysis, reveals unique genome and physiological features of the amylolytic yeast Saccharomycopsis fibuligera and its interspecies hybrid. Biotechnol Biofuels. https://doi.org/10.1186/s13068-016-0653-4
Cliften P, Sudarsanam P, Desikan A, Fulton L et al (2003) Finding functional features in Saccharomyces genomes by phylogenetic footprinting. Science. https://doi.org/10.1126/science.1084337
Crouch JA, Dawe A, Aerts A, Barry K, Churchill ACL (2020) Genome sequence of the chestnut blight fungus Cryphonectria parasitica EP155: a fundamental resource for an archetypical invasive plant pathogen. Phytopathology. https://doi.org/10.1094/PHYTO-12-19-0478-A
Cuomo CA, Güldener U, Xu JR, Trail F, Turgeon BG et al (2007) The Fusarium graminearum genome reveals a link between localized polymorphism and pathogen specialization. Science. https://doi.org/10.1126/science.1143708
D’Alessandro E, Giosa D, Huang L, Zhang J, Gao W et al (2016) Draft genome sequence of the dimorphic fungus Sporothrix pallida, a nonpathogenic species belonging to Sporothrix, a genus containing agents of human and feline Sporotrichosis. Genome Announc. https://doi.org/10.1128/genomeA.00184-16
de Beer ZW, Harrington TC, Vismer HF, Wingfield BD, Wingfield MJ (2003) Phylogeny of the Ophiostoma stenoceras-Sporothrix schenckii complex. Mycologia 95:434–441
De Beer ZW, Seifert KA, Wingfield MJ (2013) The ophiostomatoid fungi: their dual position in the Sordariomycetes. In: Seifert KA, de Beer ZW, Wingfield MJ (eds).The Ophiostomatoid fungi: expanding frontiers. CBS-KNAW fungal biodiversity series, vol 12, pp 1–19
De Beer ZW, Erasmus M, Wingfield MJ, Marincowitz S, Duong TA (2022) Generic boundaries in the Ophiostomatales reconsidered and revised. Stud Mycol (in press)
De Beer ZW, Duong TA, Barnes I, Wingfield BD, Wingfield MJ (2014) Redefining Ceratocystis and allied genera. Stud Mycol. https://doi.org/10.1016/j.simyco.2014.10.001
De Hoog GS, Scheffer R (1984) Ceratocystis versus Ophiostoma: a reappraisal. Mycologia. https://doi.org/10.1080/00275514.1984.12023838
De Schutter K, LinYC TP, Hecke V, Glinka S et al (2009) Genome sequence of the recombinant protein production host Pichia pastoris. Nat Biotechnol. https://doi.org/10.1038/nbt.1544
Dean RA, Talbot NJ, Ebbole DJ, Farman ML, Mitchell TK et al (2005) The genome sequence of the rice blast fungus Magnaporthe grisea. Nature. https://doi.org/10.1038/nature03449
Dehoux P, Davies J, Cannon M (1993) Natural cycloheximide resistance in yeast. The role of ribosomal protein L41. Eur J Biochem. https://doi.org/10.1111/j.1432-1033.1993.tb17827.x
Dietrich FS, Voegeli S, Brachat S, Lerch A, Gates K, Steiner S et al (2004) The Ashbya gossypii genome as a tool for mapping the ancient Saccharomyces cerevisiae genome. Science 304:304–307
DiGuistini S, Wang Y, Liao NY, Taylor G, Tanguay P et al (2011) Genome and transcriptome analyses of the mountain pine beetle-fungal symbiont Grosmannia clavigera, a lodgepole pine pathogen. Proc Natl Acad Sci U S A. https://doi.org/10.1073/pnas.1011289108
Dujon B, Sherman D, Fischer G, Durrens P, Casaregola S et al (2004) Genome evolution in yeasts. Nature. https://doi.org/10.1038/nature02579
Fergus CL (1956) The influence of actidione on wood-staining fungi. Mycologia. https://doi.org/10.1080/00275514.1956.12024558
Forgetta V, Leveque G, Dias J, Grove D, Lyons R et al (2013) Sequencing of the Dutch elm disease fungus genome using the Roche/454 GS-FLX titanium system in a comparison of multiple genomics core facilities. J Biomol Tech. https://doi.org/10.7171/jbt.12-2401-005
Galagan J, Calvo S, Borkovich K et al (2003) The genome sequence of the filamentous fungus Neurospora crassa. Nature. https://doi.org/10.1038/nature01554
Gibbs JN (1978) Intercontinental epidemiology of Dutch Elm disease. Ann Rev Phytopathol. https://doi.org/10.1146/annurev.py.16.090178.001443
Griffin DN, Sullia SB, Salkin IF (1978) Resistance of selected saprobic and zoopathogenic fungi to cycloheximide. J Gen Microbiol. https://doi.org/10.1099/00221287-105-1-127
Grubbs KJ, Surup F, Biedermann PHW, McDonald BR, Klassen JL et al (2020) Cycloheximide-producing Streptomyces associated with Xyleborinus saxesenii and Xyleborus affinis fungus-farming ambrosia beetles. Front Microbiol. https://doi.org/10.3389/fmicb.2020.562140
Gupta V, Kohli A, Krishnamurthy S, Puri N, Aalamgeer SA et al (1998) Identification of polymorphic mutant alleles of CaMDR1, a major facilitator of Candida albicans which confers multidrug resistance and it in vitro transcriptional activation. Curr Genet. https://doi.org/10.1007/s002940050385
Haridas S, Wang Y, Lim L, Massoumi Alamouti S, Jackman S et al (2013) The genome and transcriptome of the pine saprophyte Ophiostoma piceae, and a comparison with the bark beetle-associated pine pathogen Grosmannia clavigera. BMC Genomics 2(14):373. https://doi.org/10.1186/1471-2164-14-373
Harrington TC (1981) Cycloheximide sensitivity as a taxonomic character in Ceratocystis. Mycologia. https://doi.org/10.2307/3759682
Hausner G, Reid J, Klassen GR (1993a) On the phylogeny of Ophiostoma, Ceratocystis s.s., and Microascus, and relationships within Ophiostoma based on partial ribosomal DNA sequences. Can J Bot. https://doi.org/10.1139/b93-148
Hausner G, Reid J, Klassen GR (1993b) On the subdivision of Ceratocystis s.l., based on partial ribosomal DNA sequences. Can J Bot. https://doi.org/10.1139/b93-007
Hoang DT, Chernomor O, von Haeseler A, Minh BQ, Vinh LS (2018) UFBoot2: improving the ultrafast bootstrap approximation. Mol Biol Evol. https://doi.org/10.1093/molbev/msx281
Howe R, Moore RH (1968) Acetylation of cycloheximide by Cunninghamella blakesleeana. Experientia. https://doi.org/10.1007/BF02138641
Huang L, Gao W, Giosa D, Criseo G, Zhang J et al (2016) Whole-genome sequencing and in silico analysis of two strains of Sporothrix globosa. Genome Biol Evol. https://doi.org/10.1093/gbe/evw230
Jeon J, Kim KT, Song H, Lee GW, Cheong K, Kim H, Choi G, Lee YH et al (2017) Draft genome sequence of the fungus associated with oak wilt mortality in South Korea, Raffaelea quercus-mongolicae KACC44405. Genome Announc 5:34. https://doi.org/10.1128/genomeA.00797-17
Junier T, Zdobnov EM (2010) The Newick utilities: high-throughput phylogenetic tree processing in the UNIX shell. Bioinformatics. https://doi.org/10.1093/bioinformatics/btq243
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. https://doi.org/10.1093/molbev/mst010
Käufer NF, Fried HM, Schwindinger WF, Jasin M, Warner JR (1983) Cycloheximide resistance in yeast: the gene and its protein. Nucleic Acids Res. https://doi.org/10.1093/nar/11.10.3123
Kawai S, Murao S, Mochizuki M, Shibuya I, Yano K, Takagi M (1992) Drastic alteration of cycloheximide sensitivity by substitution of one amino acid in the LA1 ribosomal protein of yeast. J Bacteriol. https://doi.org/10.1128/jb.174.1.254-262.1992
Khoshraftar S, Hung S, Khan S, Gong Y, Tyagi V et al (2013) Sequencing and annotation of the Ophiostoma ulmi genome. BMC Genomics. https://doi.org/10.1186/1471-2164-14-162
Krassowski T, Coughlan AY, Shen XX, Zhou X, Kominek J et al (2018) Evolutionary instability of CUG-Leu in the genetic code of budding yeasts. Commun Nat. https://doi.org/10.1038/s41467-018-04374-7
Kruiswijk T, Planta RJ (1974) Analysis of the protein composition of yeast ribosomal subunits by two-dimensional polyacrylamide gel electrophoresis. Mol Biol Rep. https://doi.org/10.1007/BF00385674
Lah L, Löber U, Hsiang T, Hartmann S (2017) A genomic comparison of putative pathogenicity-related gene families in five members of the Ophiostomatales with different lifestyles. Fungal Biol. https://doi.org/10.1016/j.funbio.2016.12.002
Letunic I, Bork P (2019) Interactive tree of life (iTOL) v4: recent updates and new developments. Nucleic Acids Res. https://doi.org/10.1093/nar/gkz239
Liu F, Chen S, Ferreira MA, Chang R, Sayari M et al (2019) Draft genome sequences of five Calonectria species from Eucalyptus plantations in China, Celoporthe dispersa, Sporothrix phasma and Alectoria sarmentosa. IMA Fungus. https://doi.org/10.1186/s43008-019-0023-5
Löytynoja A (2014) Phylogeny-aware alignment with PRANK. Methods Mol Biol. https://doi.org/10.1007/978-1-62703-646-7_10
Ma LJ, van der Does HC, Borkovich KA, Coleman JJ, Daboussi MJ et al (2010) Comparative genomics reveals mobile pathogenicity chromosomes in Fusarium. Nature. https://doi.org/10.1038/nature08850
Martinez D, Berka R, Henrissat B, Saloheimo M, Arvas M et al (2008) Genome sequencing and analysis of the biomass-degrading fungus Trichoderma reesei (syn. Hypocrea jecorina). Nat Biotechnol. https://doi.org/10.1038/nbt1403
Masuya H, Manabe R, Ohkuma M, Endoh R (2016) Draft genome sequence of Raffaelea quercivora JCM 11526, a Japanese oak wilt pathogen associated with the platypodid beetle, Platypus quercivorus. Genome Announc. https://doi.org/10.1128/genomeA.00755-16
Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD et al (2020) IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol. https://doi.org/10.1093/molbev/msaa015
Mirarab S, Reaz R, Bayzid MS, Zimmermann T, Swenson MS et al (2014) ASTRAL: genome-scale coalescent-based species tree estimation. Bioinformatics. https://doi.org/10.1093/bioinformatics/btu462
Morales-Cruz A, Amrine KCH, Blanco-Ulate B, Lawrence DP et al (2015) Distinctive expansion of gene families associated with plant cell wall degradation, secondary metabolism, and nutrient uptake in the genomes of grapevine trunk pathogens. BMC Genomics. https://doi.org/10.1186/s12864-015-1624-z
Moran GP, Sanglard D, Donnelly SM, Shanley DB, Sullivan DJ et al (1998) Identification and expression of multidrug transporters responsible for fluconazole resistance in Candida dubliniensis. Antimicrob Agents Chemother. https://doi.org/10.1128/AAC.42.7.1819
Nel WJ, de Beer ZW, Wingfield MJ, Poulsen M, Aanen DK et al (2021) Phylogenetic and phylogenomic analyses reveal two new genera and three new species of ophiostomatalean fungi from termite fungus combs. Mycologia. https://doi.org/10.1080/00275514.2021.1950455
O’Connell R, Thon M, Hacquard S, Amyotte SG, Kleemann J et al (2012) Lifestyle transitions in plant pathogenic Colletotrichum fungi deciphered by genome and transcriptome analyses. Nat Genet. https://doi.org/10.1038/ng.2372
Okagaki LH, Nunes CC, Sailsbery J, Clay B, Brown D et al (2015) Genome sequences of three phytopathogenic species of the magnaporthaceae family of fungi. G3 Genes Genomes Genet. https://doi.org/10.1534/g3.115.020057
Planta RJ, Mager WH (1998) The list of cytoplasmic ribisomal protein of Saccharomyces cerevisiae. Yeast. https://doi.org/10.1002/(SICI)1097-0061(19980330)14:5%3c471::AID-YEA241%3e3.0.CO;2-U
Rao SS, Grollman AP (1967) Cycloheximide resistance in yeast: a property of the 60s subunit. Biochem Biophys Res Commun. https://doi.org/10.1016/0006-291x(67)90273-2
Réblová M, Gams W, Seifert KA (2011) Monilochaetes and allied genera of the Glomerellales, and a reconsideration of families in the Microascales. Stud Mycol 2011(68):163–191. https://doi.org/10.3114/sim.2011.68.07
Riley R, Haridas S, Wolfe KH, Lopes MR, Hittinger CT et al (2016) Comparative genomics of biotechnologically important yeasts. Proc Natl Acad Sci U S A. https://doi.org/10.1073/pnas.1603941113
Roach MJ, Borneman AR (2020) New genome assemblies reveal patterns of domestication and adaptation across Brettanomyces (Dekkera) species. BMC Genomics. https://doi.org/10.1186/s12864-020-6595-z
Rodrigues AM, de Hoog GS, de Camargo ZP (2016) Sporothrix species causing outbreaks in animals and humans driven by animal-animal transmission. PLoS Pathog 12(7):e1005638. https://doi.org/10.1371/journal.ppat.1005638
Sasnauskas K, Jomantiene R, Lebediene E, Lebedys J, Januska A et al (1992) Cloning and sequence analysis of a Candida maltosa gene which confers resistance to cycloheximide. Gene. https://doi.org/10.1016/0378-1119(92)90636-4
Schuelke TA, Wu G, Westbrook A, Woeste K, Plachetzki DC et al (2017) Comparative genomics of pathogenic and nonpathogenic beetle-vectored fungi in the genus Geosmithia. Genome Biol Evol. https://doi.org/10.1093/gbe/evx242
Seifert K (1993) Sapstain of commercial lumber by species of Ophiostoma and Ceratocystis. In: Wingfield MJ, Seifert KA, Webber J (eds) Ceratocystis and Ophiostoma: taxonomy, ecology and pathogenicity. APS Press, St. Paul, pp 141–151
Seifert KA, de Beer ZW, Wingfield MJ (eds) (2015) The Ophiostomatoid fungi: expanding frontiers. In: CBS-KNAW fungal biodiversity series 12. CBS-KNAW Biodiversity Centre, Utrecht
Seppey M, Manni M, Zdobnov EM (2019) BUSCO: assessing genome assembly and annotation completeness. In: Kollmar M. (eds) Gene prediction. Methods in Molecular Biology. https://doi.org/10.1007/978-1-4939-9173-0_14
Shearer G Jr, Sypherd PS (1988) Cycloheximide efflux in antibiotic-adapted cells of the fungus Mucor racemosus. Antimicrob Agents Chemother. https://doi.org/10.1128/AAC.32.3.341
Shen L, Su Z, Yang K, Wu C, Becker T et al (2021) Structure of the translating Neurospora ribosome arrested by cycloheximide. Proc Natl Acad Sci U S A. https://doi.org/10.1073/pnas.2111862118
Shen XX, Opulente DA, Kominek J, Zhou X, Steenwyk JL et al (2018) Tempo and mode of genome evolution in the budding yeast subphylum. Cell. https://doi.org/10.1016/j.cell.2018.10.023
Souciet JL, Dujon B, Gaillardin C, Johnston M, Baret PV, Génolevures Consortium et al (2009) Comparative genomics of protoploid Saccharomycetaceae. Genome Res. https://doi.org/10.1101/gr.091546.109
Spatafora JW, Blackwell M (1994) The polyphyletic origins of ophiostomatoid fungi. Mycol Res. https://doi.org/10.1016/S0953-7562(09)80327-4
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. https://doi.org/10.1093/bioinformatics/btu033
Stanke M, Keller O, Gunduz I, Hayes A, Waack S, Morgenstern B (2006) AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. https://doi.org/10.1093/nar/gkl200
Stevens DR, Atteia A, Franzén LG, Purton S (2001) Cycloheximide resistance conferred by novel mutations in ribosomal protein L41 of Chlamydomonas reinhardtii. Mol Gen Genet. https://doi.org/10.1007/s004380000368
Sung ZR, Lazar GB, Dudits D (1981) Cycloheximide resistance in carrot culture: a differentiated function. Plant Physiol. https://doi.org/10.1104/pp.68.1.261
Teixeira MM, de Almeida LG, Kubitschek-Barreira P, Alves FL, Kioshima ES et al (2014) Comparative genomics of the major fungal agents of human and animal Sporotrichosis: Sporothrix schenckii and Sporothrix brasiliensis. BMC Genomics. https://doi.org/10.1186/1471-2164-15-943
Uphadyay HP (1981) A monograph of Ceratocystis and Ceratocystiopsis. University of Georgia Press, Athens
Vakirlis N, Sarilar V, Drillon G, Fleiss A, Agier N et al (2016) Reconstruction of ancestral chromosome architecture and gene repertoire reveals principles of genome evolution in a model yeast genus. Genome Res. https://doi.org/10.1101/gr.204420.11
van der Nest MA, Beirn LA, Crouch JA, Demers JE, de Beer ZW et al (2014a) Draft genomes of Amanita jacksonii, Ceratocystis albifundus, Fusarium circinatum, Huntiella omanensis, Leptographium procerum, Rutstroemia sydowiana, and Sclerotinia echinophila. IMA Fungus. https://doi.org/10.5598/imafungus.2014.05.02.11
van der Nest MA, Bihon W, De Vos L, Naidoo K, Roodt D et al (2014b) Draft genome sequences of Diplodia sapinea, Ceratocystis manginecans, and Ceratocystis moniliformis. IMA Fungus. https://doi.org/10.5598/imafungus.2014.05.01.13
Vanderpool D, Bracewell RR, McCutcheon JP (2018) Know your farmer: Ancient origins and multiple independent domestications of ambrosia beetle fungal cultivars. Mol Ecol. https://doi.org/10.1111/mec.14394
Wilken PM, Steenkamp ET, Wingfield MJ, de Beer ZW, Wingfield BD (2013) Draft nuclear genome sequence for the plant pathogen, Ceratocystis fimbriata. IMA Fungus. https://doi.org/10.5598/imafungus.2013.04.02.14
Wingfield MJ, Seifert KA, Webber JF (eds) (1993) Ceratocystis and Ophiostoma: taxonomy ecology and pathogenicity. American Phytopathological Society Press, St. Paul
Wingfield BD, Ades PK, Al-Naemi FA, Beirn LA, Bihon W et al (2015a) Draft genome sequences of Chrysoporthe austroafricana, Diplodia scrobiculata, Fusarium nygamai, Leptographium lundbergii, Limonomyces culmigenus, Stagonosporopsis tanaceti, and Thielaviopsis punctulata. IMA Fungus. https://doi.org/10.5598/imafungus.2015.06.01.15
Wingfield BD, Barnes I, de Beer ZW, De Vos L, Duong TA et al (2015b) Draft genome sequences of Ceratocystis eucalypticola, Chrysoporthe cubensis, C. deuterocubensis, Davidsoniella virescens, Fusarium temperatum, Graphilbum fragrans, Penicillium nordicum, and Thielaviopsis musarum. IMA Fungus. https://doi.org/10.5598/imafungus.2015b.06.02.13
Wingfield BD, Ambler JM, Coetzee MP, de Beer ZW, Duong TA et al (2016a) Draft genome sequences of Armillaria fuscipes, Ceratocystiopsis minuta, Ceratocystis adiposa, Endoconidiophora laricicola, E. polonica and Penicillium freii DAOMC 242723. IMA Fungus. https://doi.org/10.5598/imafungus.2016a.07.01.11
Wingfield BD, Duong TA, Hammerbacher A, van der Nest MA, Wilson A et al (2016b) Draft genome sequences for Ceratocystis fagacearum, C. harringtonii, Grosmannia penicillata, and Huntiella bhutanensis. IMA Fungus. https://doi.org/10.5598/imafungus.2016b.07.02.11
Wingfield BD, Berger DK, Steenkamp ET, Lim HJ, Duong TA et al (2017a) Draft genome of Cercospora zeina, Fusarium pininemorale, Hawksworthiomyces lignivorus, Huntiella decipiens and Ophiostoma ips. IMA Fungus. https://doi.org/10.5598/imafungus.2017a.08.02.10
Wingfield MJ, Barnes I, de Beer ZW, Roux J, Wingfield BD et al (2017b) Novel associations between ophiostomatoid fungi, insects and tree hosts: current status-future prospects. Biol Invasions. https://doi.org/10.1007/s10530-017-1468
Wingfield BD, M. Liu M, Nguyen HDT, Lane FA, Morgan SW et al. (2018) Nine draft genome sequences of Claviceps purpurea s. lat., including C. arundinis, C. humidiphila, and C. cf. spartinae, pseudomolecules for the pitch canker pathogen Fusarium circinatum, draft genome of Davidsoniella eucalypti, Grosmannia galeiformis, Quambalaria eucalypti, and Teratosphaeria destructans. IMA Fungus. https://doi.org/10.5598/imafungus.2018.09.02.10
Wittmann HG, Stoeffler G, Hindennach I, Kurland CG, Randall-Hazelbauer L et al (1971) Correlation of 30S ribosomal proteins of Escherichia coli isolated in different laboratories. Mol Gen Genet. https://doi.org/10.1007/BF00569784
Wool IG, Chan YL, Glück A (1995) Structure and evolution of mammalian ribosomal proteins. Biochem Cell Biol. https://doi.org/10.1139/o95-101
Yao M-C, Yao C-H (1991) Transformation of Tetrahymena to cycloheximide resistance with a ribosomal protein gene through sequence replacement. Proc Natl Acad Sci U S A. https://doi.org/10.1073/pnas.88.21.9493
Acknowledgements
We are grateful to the South African Department of Science and Innovation (DSI) and National Research Foundation (NRF) for financial support through the South African Research Chairs Initiative (SARChI), specifically BDW’s SARChI Chair in Fungal Genomics (UID98353). The Grant holders acknowledge that opinions, findings and conclusions or recommendations expressed in any publication generated by NRF-supported research are that of the author(s), and that the NRF accepts no liability whatsoever in this regard
Funding
This work was supported by grants from the South African National Research Foundation.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. The first draft of the manuscript was written by BW and all authors commented on subsequent versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no declare they have no competing financial interests.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Additional information
Communicated by Michael Polymenis.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
294_2022_1235_MOESM1_ESM.jpg
Figure S1. Phylogenomic tree (left) and alignment of the ribosomal protein eL42 sequences (right) in 69 fungal species residing in the Saccharomycetes and the Sordariomycetes. The red stars indicate species that are known to be resistant to cycloheximide
Rights and permissions
About this article
Cite this article
Wingfield, B.D., Wingfield, M.J. & Duong, T.A. Molecular basis of cycloheximide resistance in the Ophiostomatales revealed. Curr Genet 68, 505–514 (2022). https://doi.org/10.1007/s00294-022-01235-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-022-01235-1