Abstract
The assembly of eukaryotic ribosomes follows an assembly line-like pathway in which numerous trans-acting biogenesis factors act on discrete pre-ribosomal intermediates to progressively shape the nascent subunits into their final functional architecture. Recent advances in cryo-electron microscopy have led to high-resolution structures of many pre-ribosomal intermediates; however, these static snapshots do not capture the dynamic transitions between these intermediates. To this end, molecular genetics can be leveraged to reveal how the biogenesis factors drive these dynamic transitions. Here, we briefly review how we recently used the deletion of BUD23 (bud23∆) to understand its role in the assembly of the ribosomal small subunit. The strong growth defect of bud23∆ mutants places a selective pressure on yeast cells for the occurrence of extragenic suppressors that define a network of functional interactions among biogenesis factors. Mapping these suppressing mutations to recently published structures of pre-ribosomal complexes allowed us to contextualize these suppressing mutations and derive a detailed model in which Bud23 promotes a critical transition event to facilitate folding of the central pseudoknot of the small subunit. This mini-review highlights how genetics can be used to understand the dynamics of complex structures, such as the maturing ribosome.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ribosomes are the ribonucleoprotein complexes that translate the genetic code into proteins. Each ribosome contains two subunits: a small subunit (SSU or 40S) that decodes mRNA and a large subunit (LSU or 60S) that catalyzes protein synthesis. In eukaryotes, ribosomes are assembled along a hierarchical pathway that involves the synthesis of four ribosomal RNAs (rRNA), the incorporation of roughly 80 ribosomal proteins (RPs), and the function of more than 200 trans-acting biogenesis factors (Woolford and Baserga 2013). Many factors act as chaperones that ensure correct ribosomal structure while others drive necessary rRNA processing and catalyze structural rearrangements. Decades of biochemical and genetic studies in the model eukaryote Saccharomyces cerevisiae have informed our understanding of ribosome assembly. The early work pertaining to SSU assembly was primarily devoted to rRNA modification and processing (Venema et al. 1995; Lafontaine et al. 1995; Allmang et al. 1996; Granneman et al. 2006; Bleichert et al. 2006), while other studies identified additional biogenesis factors and roughly ordered the timing of their association with pre-ribosomal particles (Dragon et al. 2002; Grandi et al. 2002; Wehner et al. 2002; Schäfer et al. 2003; Gallagher et al. 2004; Bernstein et al. 2004; Pérez-Fernández et al. 2007). However, this gave us only an amorphous picture of the assembly pathway. It has only been with the advent of high-resolution cryo-electron microscopy (cryo-EM) that we have been able to visualize discrete intermediates of assembly (Barandun et al. 2017; Kater et al. 2017; Ameismeier et al. 2018; Zhou et al. 2019; Cheng et al. 2019, 2020; Du et al. 2020; Rai et al. 2021).
The structures obtained by cryo-EM provide static snapshots of metastable intermediate particles along the assembly pathway. However, these structures do not capture the dynamic events of ribosome assembly to tell us how the transitions between these intermediates are driven by biogenesis factors. To understand these dynamic transitions, we can turn to genetics to animate structures and reveal the functional roles of biogenesis factors. A genetic interaction occurs when an allele of one gene influences the phenotype caused by an allele of another gene. Such an interaction implies a functional relationship but not necessarily a physical relationship; the interpretation of genetic interactions has been extensively reviewed (Dixon et al. 2009; Costanzo et al. 2019). One class of genetic interaction is extragenic suppression, where the defects due to mutation of one gene are alleviated by the deletion or mutation of another. We have found that extragenic suppression is particularly well-suited to study the release of factors from multimeric complexes. On average, a mutation is more likely to disrupt a molecular interaction than promote one. Consequently, if factor A is needed to disrupt a complex of factors B and C, then a mutation in factor B or C that facilitates their separation will tend to bypass the need for factor A. Such a relationship can be observed in numerous transition events in ribosome biogenesis that often rely on the disassociation of biogenesis factors. For example, the loss of the essential kinase Hrr25 blocks the release of the 40S biogenesis factor Ltv1 and can be bypassed by phosphomimetic Ltv1 mutants that promote its release (Ghalei et al. 2015). We and others have previously made extensive use genetic interactions to delineate the cytoplasmic 60S maturation pathway (Hedges et al. 2005; Kemmler et al. 2009; Lo et al. 2009, 2010). When contextualized with structural information, genetic interactions can be used to develop verifiable mechanistic models that explain how a set of factors promote the progression of an intermediate (Patchett et al. 2017; Zhou et al. 2019; Mitterer et al. 2019). Recently, we used the deletion of BUD23 (bud23∆) to understand its role in 40S biogenesis (Black et al. 2020). In this mini-review, we highlight how screening for extragenic suppressors of bud23∆ revealed a network of biogenesis factors that, when considered in the light of recent structures of 40S precursors, reveal that Bud23 promotes a critical transition event in the assembly of the small ribosomal subunit.
The assembly and disassembly of the small subunit processome
Ribosome biogenesis begins with transcription of a primary transcript containing three of the four precursor rRNAs (pre-rRNA) flanked by spacer regions that are removed during processing. The SSU Processome, henceforth “Processome”, is an assemblage of approximately 70 biogenesis factors and RPs that encompass the 5′ portion of the primary transcript, containing the 5′ external transcribed spacer (5′ ETS), the 18S rRNA, and the internal transcribed spacer 1 (ITS1) (Dragon et al. 2002; Grandi et al. 2002; Pérez-Fernández et al. 2007). The Processome forms co-transcriptionally with the modular incorporation biogenesis factors and RPs (Pérez-Fernández et al. 2007; Chaker-Margot et al. 2015; Zhang et al. 2016). The 5′ ETS and its associated factors along with the U3 snoRNA establish the scaffolding upon which the Processome is built (Beltrame et al. 1994; Venema et al. 1995; Hunziker et al. 2016, 2019). The initial folding of the rRNA is driven by its intrinsic ability to form secondary structure resulting in four domains: the 5′, central, 3′ major, and 3′ minor. The Processome factors chaperone the tertiary structure of the rRNA domains by holding them in a splayed-open conformation that allows their independent maturation but prevents them from adopting their more compact conformations seen in downstream pre-40S intermediates (Kornprobst et al. 2016; Chaker-Margot et al. 2017; Sun et al. 2017; Barandun et al. 2017; Cheng et al. 2017). The Processome is present even under conditions in which ribosome assembly is largely halted (Talkish et al. 2016; Kos-Braun et al. 2017; Chaker-Margot et al. 2017), suggesting that it serves as a metastable intermediate that controls further progression of maturation.
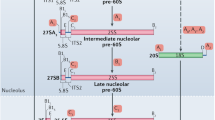
The transition from the Processome to the pre-40S particles requires the disassembly of the Processome which entails pre-rRNA cleavage and removal of the 5′ ETS by the RNA Exosome, the release of most Processome factors, and RNA cleavage within ITS1 to separate the SSU and LSU precursors (Schäfer et al. 2003; Thoms et al. 2015; Lau et al. 2021) Recent cryo-EM studies reveal that the Processome disassembles in a stepwise fashion (Cheng et al. 2020; Du et al. 2020; Lau et al. 2021). The shedding of the Processome elements is accompanied by dramatic structural rearrangements that result in compaction of the 18S rRNA as it approaches its mature structure (Fig. 1). A recently described late intermediate of the disassembly pathway, the Dis-C complex (Cheng et al. 2020), retains a handful of Processome factors, some of which prevent the 3′ major and minor domains from adopting their near-final positions. Among these factors is the U3 snoRNA, which scaffolds the Processome through hybridization to the 5′ ETS and pre-18S rRNAs (Beltrame and Tollervey 1995; Méreau et al. 1997; Sharma and Tollervey 1999; Kudla et al. 2011; Dutca et al. 2011; Marmier-Gourrier et al. 2011). Its interaction with pre-18S rRNA prevents the premature formation of the central pseudoknot (CPK), a universally conserved structural hallmark that organizes all four rRNA domains of 18S rRNA as well as the decoding center. The CPK is formed by the long-range base pairing of the loop of helix 1 at the 5′-end of 18S rRNA with nucleotides A1137-U1144 to form helix 2. Displacement of U3 snoRNA by the DEAH-box helicase Dhr1 and its activator Utp14 is required for the CPK to fold (Sardana et al. 2015; Zhu et al. 2016; Black et al. 2018). Strikingly, Dhr1 and Utp14 are primed to unwind U3 to drive its removal in the Dis-C complex (Cheng et al. 2020), but what promotes this final disassembly step was not clear.
Structural overview of small ribosomal subunit biogenesis in eukaryotes. Small subunit biogenesis is a multi-step process; several SSU assembly intermediates are shown to highlight the structural rearrangements that occur during SSU biogenesis. The SSU Processome (composite of PDBs 5WLC and 5WYJ) is an early precursor which contains assembly factors (AFs, light gray) and ribosomal proteins (RPs, dark gray) that independently chaperone the four domains: the 5′ major, central, 3′ major, and 3′ minor (respectively, colored blue, magenta, yellow, and green). A yet unknown signal triggers progression of the Processome which involves the shedding of the 5′-ETS as well as many AFs and RNA rearrangements (colored arrows) that transform it into a more compact final disassembly complex, termed “Dis-C” (PDB 6ZQG) (Cheng et al. 2020). However, a set of factors retained on Dis-C prevent its final transformation into the earliest pre-40S intermediate (PDB 6G4W). We have proposed that the coordinated efforts of Bud23 and the RNA helicase Dhr1 promote the final transition into the pre-40S, allowing the 3′ domains to adopt their near-mature positions (Black et al. 2020). A series of subsequent, less dramatic maturation events fine-tune the rRNA (colored lines; lower panel) to produce the mature 40S (PDB 4V88). The figure is adapted from (Black et al. 2020). Molecular visualizations were generated in UCSF ChimeraX v0.93 (Goddard et al. 2018)
Bud23 promotes the final disassembly of the Processome
BUD23 is a nonessential gene encoding a Trm112-associated methyltransferase that modifies nitrogen 7 of guanosine 1575 (G1575) of the 18S rRNA (White et al. 2008; Figaro et al. 2012; Sardana and Johnson 2012; Létoquart et al. 2014). Its deletion (bud23∆) reduces SSU levels by 70% and causes a strong growth defect (White et al. 2008); however, the methyltransferase activity of Bud23 is fully dispensable for growth and SSU assembly (White et al. 2008; Lin et al. 2012; Létoquart et al. 2014). Thus, the physical presence of Bud23 confers its primary function, but how exactly it promotes SSU biogenesis was unknown. Bud23 is often considered a pre-40S factor, and consistent with this notion human Bud23 was resolved on the earliest pre-40S intermediate (Ameismeier et al. 2018). However, a decade of studies in yeast suggested that Bud23 first binds to an earlier precursor, that retains some Processome factors, and remains associated with the particle during its transition to a pre-40S (Figaro et al. 2012; Sardana et al. 2013, 2014; Létoquart et al. 2014).
To understand the function of Bud23, we took advantage of the remarkable propensity of bud23∆ mutants to generate spontaneous suppressors of its slow growth phenotype. The high rate of generating suppressors appears to be due to the large number of mutations that can suppress bud23∆ cells rather than an elevated mutation rate (unpublished). Through a combination of screening for spontaneous extragenic suppressors and random mutagenesis of several target factors, we identified a total of 67 unique extragenic suppressor mutations of bud23∆ in genes encoding the Processome factors Dhr1, Utp14, Utp2, Imp4, and the GTPase Bms1 as well as the ribosomal protein Rps28 (Fig. 2a) (Sardana 2013; Sardana et al. 2013, 2014; Zhu et al. 2016; Black et al. 2020). The genetic connection of Bud23 to Imp4 and Dhr1 (Sardana 2013; Sardana et al. 2014) was particularly informative since Imp4 was known to be a U3-associated factor (Lee and Baserga 1999; Gallagher and Baserga 2004), and led us to identify Dhr1 as the helicase that unwinds U3 snoRNA from pre-rRNA (Sardana et al. 2015) and Utp14 as its co-factor (Zhu et al. 2016). The genetic interactions of BUD23 suggested a role for Bud23 in Processome disassembly. Indeed, we recently showed that intermediates of Processome disassembly accumulate in Bud23-depleted cells (Black et al. 2020).
Summary of the physical interaction network underlying bud23∆ suppression. a The extragenic suppressors of bud23∆ form a tight network of physical and functional interactions that includes U3 snoRNA. b Mapping this network to the structure of the SSU Processome reveals physical interactions connecting to U3 snoRNA; Bms1 (green), Imp4 (blue), Rps28 (cyan), Utp2 (orange), and Utp14 (brown). Imp4, Utp2, and Bms1 physically connect the U3 snoRNA (magenta) to the 3′ basal subdomain (dark gray), the future binding site of Bud23. The target base of Bud23 methyltransferase, G1575 (red), is shown for reference. c The majority of the Imp4 mutants (magenta sticks) mapped to its interaction interface with the 3′ basal subdomain of 18S RNA. d The Utp2 mutants (green sticks) mapped to its interface with Imp4 near the 3′ basal subdomain. e F58 of Utp2 fits into a hydrophobic pocket in Imp4 (upper) while V170 and P252 of Imp4 help establish this pocket (lower). Amino acid substitutions of these residues disrupt the interaction between Imp4 and Utp2 (Black et al. 2020). Molecular visualizations were generated in MacPyMOL: PyMOL v1.8.2.1 Enhanced for Mac OS X (Schrödinger LLC) using PDB 5WLC
With the recent publication of several structures of the Processome and disassembly intermediates (Kornprobst et al. 2016; Chaker-Margot et al. 2017; Sun et al. 2017; Barandun et al. 2017; Cheng et al. 2017, 2020), we can postulate how the suppressors of bud23∆ impact the disassembly pathway. These structures reveal that, like Bud23, the factors Bms1, Imp4, Utp2, and Rps28 all contact the 3′ basal subdomain, a compact subdomain of the 3′ major domain (Fig. 2b). Bms1, Imp4, and Utp2 also have long helical extensions that pierce deep into the Processome where they embrace, and appear to stabilize, the U3-18S rRNA heteroduplexes that will be unwound by Dhr1 and Utp14. Furthermore, Bms1, Imp4, and Utp2 all make multiple interactions amongst one another. Thus, the bud23∆ suppressing mutations describe a network of factors connecting the 3′ major rRNA domain to the U3 heteroduplexes within the Processome or Processome disassembly intermediates (Fig. 2a, b). Close inspection of the Imp4 mutants revealed that the majority of the amino acid changes cluster in its interface with the rRNA, suggesting that these mutations weaken the affinity of Imp4 for the rRNA (Fig. 2c). Similarly, many of the Utp2 mutants map to its interface with Imp4 (Fig. 2d). Notably, F58 of Utp2 inserts into a hydrophobic pocket of Imp4 (Fig. 2e, upper) and mutation of F58 of Utp2 or V170 or P252 of Imp4, which help to create this pocket (Fig. 2e, lower), weaken the interaction between these two proteins (Black et al. 2020). Meanwhile, Rps28 binds directly adjacent to the binding site of Bud23. The single suppressor mutation that we identified in RPS28A, G24D, lies in a residue that directly interacts with the rRNA of the 3′ basal subdomain (Black et al. 2020). Because Rps28 is a constituent of the mature 40S, we speculate that this amino acid change affects rRNA structure in a manner that facilitates the release of factors that bind the 3′ basal subdomain rather than the mutation promoting Rps28 release. These results led us to conclude that the Imp4, Utp2, and Rps28 mutants destabilize the scaffold that holds the 3′ major domain in place in the Processome. Because these destabilizing mutations bypass the absence of Bud23, it can be inferred that the role of Bud23 is to promote rearrangement of the 3′ major domain.
Bms1 is an essential GTPase of the Processome whose exact molecular function remains unknown (Gelperin et al. 2001; Wegierski et al. 2001; Karbstein et al. 2005; Karbstein and Doudna 2006). The cryo-EM structures revealed that Bms1 provides a rigid strut between the 3′ major and 5′-domains, thereby constraining the movement of the 3′ major domain. In our genetic screen for bud23∆ suppressors, we did not find mutations in Bms1 at its interface with the 3′ basal subdomain. However, we did identify five mutations in BMS1 that change amino acids located within intramolecular interfaces of Bms1 (Black et al. 2020). Comparison of Bms1 to structures of the related GTPase EF-Tu suggests that these amino acid changes likely mimic a conformational change that Bms1 undergoes during GTP hydrolysis (Black et al. 2020), raising the intriguing possibility that Bud23 influences Bms1 GTPase activity. Consistent with this notion, Bms1 is in the GTP-bound state in the Dis-C complex indicating that its GTPase has not been activated (Cheng et al. 2020). We speculate that Bms1 serves as a molecular switch that senses Bud23 binding and initiates disassembly of the scaffold holding the 3′ major domain in place.
A kinetic proofreading model for central pseudoknot formation
The functional connection of Bud23 to Dhr1 is particularly interesting as Dhr1 is a DEAH box helicase that removes the U3 snoRNA (Sardana et al. 2015; Choudhury et al. 2018). Dhr1 requires Utp14 for activation (Zhu et al. 2016; Choudhury et al. 2018; Boneberg et al. 2019) which depends on an interaction between an unresolved region of Utp14 with the catalytic domains of Dhr1 (Zhu et al. 2016). Amino acid changes within this region of Utp14 suppress bud23∆, diminish the Dhr1-Utp14 interaction, and consequentially reduce Dhr1 unwinding activity (Zhu et al. 2016). In our suppressor screen, we also identified numerous mutations in Dhr1, many of which affect residues on the surface of its catalytic domains, and like the Utp14 mutants, these Dhr1 mutants weaken the Dhr1–Utp14 interaction (Black et al. 2020). Thus, diminishing the Dhr1–Utp14 interaction, and consequently Dhr1 activity, leads to suppression of bud23∆.
The result that reduced helicase activity bypasses a defect in disassembly seems counterintuitive; why would reducing Dhr1 unwinding activity, which is needed for disassembly of the Processome, suppress the disassembly defect of bud23∆? Here, we can glean clues from the structures of the disassembly intermediates (Cheng et al. 2020). During Processome disassembly, RNA rearrangements allow the stem-loop of helix 1 of the central pseudoknot (CPK) to fold, while the loop remains paired with U3. In the mature CPK, nucleotides A1137-U1144 of 18S will replace U3 to form helix 2 of the CPK. Residues A1137-U1144 are unresolved in the Dis-C complex, but their position can be inferred because they originate from the 3′ basal subdomain (Fig. 3a). We surmise that productive unwinding of U3 by Dhr1, leading to the formation of helix 2, requires the correct positioning of residues A1137-U1144. Dhr1 binds to U3 on the 3′-side of the U3-rRNA duplexes (Sardana et al. 2015; Cheng et al. 2020). Dhr1 is expected to translocate 3′ to 5′ (Boneberg et al. 2019), essentially pulling U3 towards itself to unwind it from the rRNA. We propose that the concerted actions of Bud23 and Dhr1 promote helix 2 formation to form the CPK (Fig. 3b). In this model, Bud23 binding to the 3′ basal subdomain triggers Bms1-driven rearrangements that bring residues A1137-U1144 into close proximity of helix 1. Here, these residues are poised to base-pair with the stem-loop of helix 1 to form the CPK. Subsequently, the Utp14-dependent activity of Dhr1 unwinds U3 to allow productive CPK folding.
Model for how Bud23 promotes the final disassembly step of the SSU Processome. a The structures of Bms1, Imp4, Rps28, Utp2, Utp14, and Dhr1 as observed in the final disassembly complex (Dis-C, PDB 6ZQG). Factors are colored as in Fig. 2a. Helix 1 of the 18S rRNA has formed. Nucleotides A1137-U1144 of the 18S rRNA are unresolved but originate from the 3′ basal subdomain at G1146. Molecular visualizations were generated, MacPyMOL: PyMOL v1.8.2.1 Enhanced for Mac OS X (Schrödinger LLC). b The model for Bud23 function with proteins and RNAs colored as in panel A. Bud23 binding to the 3′ basal subdomain promotes Bms1 activation (black lines), allowing Imp4 and Utp2 eviction and the rRNA rearrangements (gray arrows) that bring residues A1137-U1144 into proximity with helix 1. Subsequent activation of Dhr1 by Utp14 (green arrow) promotes productive unwinding of U3, which allows the rRNA strand containing residues A1137-U1144 to base-pair with the stem-loop of Helix 1 to form Helix 2, completing the CPK
Our model proposes that Bud23 drives the correct positioning of A1137-U1144 (Fig. 3b). We suggest that this repositioning is critical to promote productive unwinding of U3 by Dhr1 to fold the CPK (Fig. 4a). Therefore, the absence of Bud23 (bud23∆) should hinder the repositioning of these residues, leading to unproductive unwinding of U3, as the CPK cannot fold (Fig. 4b); however, CPK folding must still occur at some low rate in the absence of Bud23 since bud23∆ is not lethal. With this framework for CPK formation, we propose a kinetic proofreading model to explain how Dhr1 and Utp14 mutants bypass bud23∆ to restore 40S assembly. For the mutants that we have tested, their impact on Dhr1 is a reduction in its unwinding activity (Zhu et al. 2016; Black et al. 2020). We suggest that such mutants allow a greater window of opportunity for A1137-U1144 to move into place before Dhr1 unwinds U3, increasing the likelihood of a productive unwinding event that allows the CPK to fold (Fig. 4c). Similar kinetic proofreading models have been used to understand spliceosome dynamics where DEAD/H-box helicase-dependent RNA rearrangements are needed for transitioning through intermediate states (Semlow and Staley 2012). This framework for CPK folding also accommodates how the Utp2, Imp4, and Bms1 mutants bypass bud23∆. We proposed that Bud23 triggers disassembly of the scaffolding that constrains the 3′ basal subdomain to allow its rearrangement in the maturing particle (Fig. 3b). Weakening the scaffolding that holds the subdomain in place, through mutations in Utp2, Imp4, or Bms1 would reduce the energy barrier for the rearrangements necessary that bring A1137-U1144 into proximity of helix 1 for productive unwinding by Dhr1 (Fig. 4c).
Kinetic proofreading model for how extragenic suppressors bypass bud23∆. a In wild-type cells, Bud23 binding to the Dis-C complex leads to release of Imp4 and Utp2 and Bms1-dependent repositioning of residues A1137-U1144 of the 18S rRNA into close proximity of U3. The subsequent unwinding of U3 by Dhr1 allows A1137-U1144 to base pair with the stem-loop of Helix 1 to form Helix 2, thereby completing the CPK. b In cells lacking Bud23, the repositioning of residues A1137-U1144 is hindered and the unwinding of U3 by Dhr1 leads to unproductive folding of the rRNA and failure in CPK formation. However, a low rate of productive folding must occur in bud23∆ cells because they are viable, though very slow growing. c In bud23∆ cells harboring a suppressor mutation, the rate of productive CPK folding is partially restored. This can be due to either (1) amino acid substitutions in Dhr1 or Utp14 that slow the Dhr1 activity to allow more time for residues A1137-U1144 to come into position before U3 unwinding occurs or (2) amino acid substitutions in Bms1, Imp4, or Utp2 that promote the repositioning of residues A1137-U1144
In the disassembly complex, Dhr1 and Bms1 are positioned on opposite sides of the structure (Fig. 3a). Interestingly, the N-terminus of Dhr1 extends across the complex and interacts with Bms1. Thus, Bms1 and Dhr1 can be mechanistically and physically connected. It is tempting to compare the relationship of Bms1 and Dhr1 in Processome disassembly to that of Snu114 and Brr2 that control spliceosome disassembly where the GTPase-dependent conformational changes in Snu114 promote Brr2 helicase activity (Small et al. 2006). It is tempting to speculate that there is a similar coordination between Dhr1 and Bms1 during Processome disassembly.
Leveraging genetic fulcrums to animate structure
In this mini-review, we have highlighted our recent in-depth screen for extragenic suppressors of bud23∆ that we carried out to understand the role of Bud23 in 40S assembly (Black et al. 2020). We identified an unexpectedly large number of suppressors in multiple factors involved in the function of the Processome making bud23∆ a particularly powerful entry point for genetic analysis. We liken the bud23∆ mutant to a kind of “genetic fulcrum”, a tool that allows us to leverage a genetic defect to understand the workings of the Processome. We believe this work also nicely illustrates how genetics can be used to animate structures of macromolecular complexes that undergo dynamic conformational and compositional changes. Assembly of such complexes requires the formation of myriad macromolecular contacts to produce a functional architecture, while its transition into a subsequent state entails the disruption of those contacts. The particular genetic relationship that we uncover, mutations in factors that destabilize the Processome bypass the function of Bud23, is particularly apt for studying disassembly of the Processome, as well as other macromolecular complexes. Such mutations in critical factors must strike a fine balance between sufficient loss of function to destabilize an interface and complete disruption which would result in lethality. Indeed, the suppressors of bud23∆ showed no obvious growth defects in Bud23-replete cells individually but did negatively impact fitness when combined (Zhu et al. 2016; Black et al. 2020). Although Cryo-EM has allowed the unprecedented visualization of many pre-ribosomal intermediates (Klinge and Woolford 2019; Frazier et al. 2020), genetic approaches, such as what we have reported (Black et al. 2020), allow us to understand the dynamics of these complexes. A similar genetic analysis has recently been used to explore splicesosomal dynamics (Brow 2019). Cryo-EM will continue to provide snapshots of numerous complexes, and the use of molecular genetics should be broadly applicable to animate those snapshots and understand their dynamic natures.
References
Allmang C, Henry Y, Wood H et al (1996) Recognition of cleavage site A(2) in the yeast pre-rRNA. RNA 2:51–62
Ameismeier M, Cheng J, Berninghausen O, Beckmann R (2018) Visualizing late states of human 40S ribosomal subunit maturation. Nature 558:249–253. https://doi.org/10.1038/s41586-018-0193-0
Barandun J, Chaker-Margot M, Hunziker M et al (2017) The complete structure of the small-subunit processome. Nat Struct Mol Biol. https://doi.org/10.1038/nsmb.3472
Beltrame M, Tollervey D (1995) Base pairing between U3 and the pre-ribosomal RNA is required for 18S rRNA synthesis. EMBO J 14:4350–4356. https://doi.org/10.1002/j.1460-2075.1995.tb00109.x
Beltrame M, Henry Y, Tollervey D (1994) Mutational analysis of an essential binding site for the U3 snoRNA in the 5’ external transcribed spacer of yeast pre-rRNA. Nucleic Acids Res 22:4057–4065. https://doi.org/10.1093/nar/22.20.4057
Bernstein KA, Gallagher JEG, Mitchell BM et al (2004) The small-subunit processome is a ribosome assembly intermediate. Eukaryot Cell 3:1619–1626. https://doi.org/10.1128/EC.3.6.1619-1626.2004
Black JJ, Wang Z, Goering LM, Johnson AW (2018) Utp14 interaction with the small subunit processome. RNA 24:1214–1228. https://doi.org/10.1261/rna.066373.118
Black JJ, Sardana R, Elmir EW, Johnson AW (2020) Bud23 promotes the final disassembly of the small subunit Processome in Saccharomyces cerevisiae. PLoS Genet 16:e1009215. https://doi.org/10.1371/journal.pgen.1009215
Bleichert F, Granneman S, Osheim YN et al (2006) The PINc domain protein Utp24, a putative nuclease, is required for the early cleavage steps in 18S rRNA maturation. Proc Natl Acad Sci USA 103:9464–9469. https://doi.org/10.1073/pnas.0603673103
Boneberg FM, Brandmann T, Kobel L et al (2019) Molecular mechanism of the RNA helicase DHX37 and its activation by UTP14A in ribosome biogenesis. RNA 25:685–701. https://doi.org/10.1261/rna.069609.118
Brow DA (2019) An allosteric network for spliceosome activation revealed by high-throughput suppressor analysis in Saccharomyces cerevisiae. Genetics 212:111–124. https://doi.org/10.1534/genetics.119.301922
Chaker-Margot M, Hunziker M, Barandun J et al (2015) Stage-specific assembly events of the 6-MDa small-subunit processome initiate eukaryotic ribosome biogenesis. Nat Struct Mol Biol 22:920–923. https://doi.org/10.1038/nsmb.3111
Chaker-Margot M, Barandun J, Hunziker M, Klinge S (2017) Architecture of the yeast small subunit processome. Science. https://doi.org/10.1126/science.aal1880
Cheng J, Kellner N, Berninghausen O et al (2017) 3.2-Å-resolution structure of the 90S preribosome before A1 pre-rRNA cleavage. Nat Struct Mol Biol 24:954–964. https://doi.org/10.1038/nsmb.3476
Cheng J, Baßler J, Fischer P et al (2019) Thermophile 90S pre-ribosome structures reveal the reverse order of co-transcriptional 18S rRNA subdomain integration. Mol Cell. https://doi.org/10.1016/j.molcel.2019.06.032
Cheng J, Lau B, La Venuta G et al (2020) 90S pre-ribosome transformation into the primordial 40S subunit. Science 369:1470–1476. https://doi.org/10.1126/science.abb4119
Choudhury P, Hackert P, Memet I et al (2018) The human RNA helicase DHX37 is required for release of the U3 snoRNP from pre-ribosomal particles. RNA Biol 00:1–15. https://doi.org/10.1080/15476286.2018.1556149
Costanzo M, Kuzmin E, van Leeuwen J et al (2019) Global Genetic Networks and the Genotype-to-Phenotype Relationship. Cell 177:85–100. https://doi.org/10.1016/j.cell.2019.01.033
Dixon SJ, Costanzo M, Baryshnikova A et al (2009) Systematic mapping of genetic interaction networks. Annu Rev Genet 43:601–625. https://doi.org/10.1146/annurev.genet.39.073003.114751
Dragon F, Gallagher JEG, Compagnone-Post PA et al (2002) A large nucleolar U3 ribonucleoprotein required for 18S ribosomal RNA biogenesis. Nature 417:967–970. https://doi.org/10.1038/nature00769
Du Y, An W, Zhu X et al (2020) Cryo-EM structure of 90S small ribosomal subunit precursors in transition states. Science 369:1477–1481. https://doi.org/10.1126/science.aba9690
Dutca LM, Gallagher JEG, Baserga SJ (2011) The initial U3 snoRNA:pre-rRNA base pairing interaction required for pre-18S rRNA folding revealed by in vivo chemical probing. Nucleic Acids Res 39:5164–5180. https://doi.org/10.1093/nar/gkr044
Figaro S, Wacheul L, Schillewaert S et al (2012) Trm112 is required for Bud23-mediated methylation of the 18S rRNA at position G1575. Mol Cell Biol 32:2254–2267. https://doi.org/10.1128/MCB.06623-11
Frazier MN, Pillon MC, Kocaman S et al (2020) Structural overview of macromolecular machines involved in ribosome biogenesis. Curr Opin Struct Biol 67:51–60. https://doi.org/10.1016/j.sbi.2020.09.003
Gallagher JEG, Baserga SJ (2004) Two-hybrid Mpp10p interaction-defective Imp4 proteins are not interaction defective in vivo but do confer specific pre-rRNA processing defects in Saccharomyces cerevisiae. Nucleic Acids Res 32:1404–1413. https://doi.org/10.1093/nar/gkh318
Gallagher JEG, Dunbar DA, Granneman S et al (2004) RNA polymerase I transcription and pre-rRNA processing are linked by specific SSU processome components. Genes Dev 18:2506–2517. https://doi.org/10.1101/gad.1226604
Gelperin D, Horton L, Beckman J et al (2001) Bms1p, a novel GTP-binding protein, and the related Tsr1p are required for distinct steps of 40S ribosome biogenesis in yeast. RNA 7:1268–1283. https://doi.org/10.1017/S1355838201013073
Ghalei H, Schaub FX, Doherty JR et al (2015) Hrr25/CK1delta-directed release of Ltv1 from pre-40S ribosomes is necessary for ribosome assembly and cell growth. J Cell Biol 208:745–759. https://doi.org/10.1083/jcb.201409056
Goddard TD, Huang CC, Meng EC et al (2018) UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci 27:14–25. https://doi.org/10.1002/pro.3235
Grandi P, Rybin V, Bassler J et al (2002) 90S pre-ribosomes include the 35S pre-rRNA, the U3 snoRNP, and 40S subunit processing factors but predominantly lack 60S synthesis factors. Mol Cell 10:105–115. https://doi.org/10.1016/S1097-2765(02)00579-8
Granneman S, Bernstein KA, Bleichert F, Baserga SJ (2006) Comprehensive mutational analysis of yeast DEXD/H box RNA helicases required for small ribosomal subunit synthesis. Mol Cell Biol 26:1183–1194. https://doi.org/10.1128/MCB.26.4.1183-1194.2006
Hedges J, West M, Johnson AW (2005) Release of the export adapter, Nmd3p, from the 60S ribosomal subunit requires Rpl10p and the cytoplasmic GTPase Lsg1p. EMBO J 24:567–579. https://doi.org/10.1038/sj.emboj.7600547
Hunziker M, Barandun J, Petfalski E et al (2016) UtpA and UtpB chaperone nascent pre-ribosomal RNA and U3 snoRNA to initiate eukaryotic ribosome assembly. Nat Commun 7:12090. https://doi.org/10.1038/ncomms12090
Hunziker M, Barandun J, Buzovetsky O et al (2019) Conformational switches control early maturation of the eukaryotic small ribosomal subunit. Elife 8:1–16. https://doi.org/10.7554/eLife.45185
Karbstein K, Doudna JA (2006) GTP-dependent formation of a ribonucleoprotein subcomplex required for ribosome biogenesis. J Mol Biol 356:432–443. https://doi.org/10.1016/j.jmb.2005.11.052
Karbstein K, Jonas S, Doudna JA (2005) An essential GTPase promotes assembly of preribosomal RNA processing complexes. Mol Cell 20:633–643. https://doi.org/10.1016/j.molcel.2005.09.017
Kater L, Thoms M, Barrio-Garcia C et al (2017) Visualizing the assembly pathway of nucleolar pre-60S ribosomes. Cell 171:1599-1610.e14. https://doi.org/10.1016/j.cell.2017.11.039
Kemmler S, Occhipinti L, Veisu M, Panse VG (2009) Yvh1 is required for a late maturation step in the 60S biogenesis pathway. J Cell Biol 186:863–880. https://doi.org/10.1083/jcb.200904111
Klinge S, Woolford JL (2019) Ribosome assembly coming into focus. Nat Rev Mol Cell Biol 20:116–131. https://doi.org/10.1038/s41580-018-0078-y
Kornprobst M, Turk M, Kellner N et al (2016) Architecture of the 90S pre-ribosome: a structural view on the birth of the eukaryotic ribosome. Cell 166:380–393. https://doi.org/10.1016/j.cell.2016.06.014
Kos-Braun IC, Jung I, Koš M (2017) Tor1 and CK2 kinases control a switch between alternative ribosome biogenesis pathways in a growth-dependent manner. PLoS Biol 15:e2000245. https://doi.org/10.1371/journal.pbio.2000245
Kudla G, Granneman S, Hahn D et al (2011) Cross-linking, ligation, and sequencing of hybrids reveals RNA-RNA interactions in yeast. Proc Natl Acad Sci USA 108:10010–10015. https://doi.org/10.1073/pnas.1017386108
Lafontaine D, Vandenhaute J, Tollervey D (1995) The 18S rRNA dimethylase Dim1p is required for pre-ribosomal RNA processing in yeast. Genes Dev 9:2470–2481
Lau B, Cheng J, Flemming D et al (2021) Structure of the maturing 90S pre-ribosome in association with the RNA exosome. Mol Cell 81:293-303.e4. https://doi.org/10.1016/j.molcel.2020.11.009
Lee SJ, Baserga SJ (1999) Imp3p and Imp4p, two specific components of the U3 small nucleolar ribonucleoprotein that are essential for pre-18S rRNA processing. Mol Cell Biol 19:5441–5452. https://doi.org/10.1128/MCB.19.8.5441
Létoquart J, Huvelle E, Wacheul L et al (2014) Structural and functional studies of Bud23-Trm112 reveal 18S rRNA N7–G1575 methylation occurs on late 40S precursor ribosomes. Proc Natl Acad Sci USA 111:E5518–E5526. https://doi.org/10.1073/pnas.1413089111
Lin J-L, Yu H-C, Chao J-L et al (2012) New phenotypes generated by the G57R mutation of BUD23 in Saccharomyces cerevisiae. Yeast 29:537–546. https://doi.org/10.1002/yea.2934
Lo K-Y, Li Z, Wang F et al (2009) Ribosome stalk assembly requires the dual-specificity phosphatase Yvh1 for the exchange of Mrt4 with P0. J Cell Biol 186:849–862. https://doi.org/10.1083/jcb.200904110
Lo K-Y, Li Z, Bussiere C et al (2010) Defining the pathway of cytoplasmic maturation of the 60S ribosomal subunit. Mol Cell 39:196–208. https://doi.org/10.1016/j.molcel.2010.06.018
Marmier-Gourrier N, Cléry A, Schlotter F et al (2011) A second base pair interaction between U3 small nucleolar RNA and the 5’-ETS region is required for early cleavage of the yeast pre-ribosomal RNA. Nucleic Acids Res 39:9731–9745. https://doi.org/10.1093/nar/gkr675
Méreau A, Fournier R, Grégoire A et al (1997) An in vivo and in vitro structure-function analysis of the Saccharomyces cerevisiae U3A snoRNP: protein-RNA contacts and base-pair interaction with the pre-ribosomal RNA. J Mol Biol 273:552–571. https://doi.org/10.1006/jmbi.1997.1320
Mitterer V, Shayan R, Ferreira-Cerca S et al (2019) Conformational proofreading of distant 40S ribosomal subunit maturation events by a long-range communication mechanism. Nat Commun 10:2754. https://doi.org/10.1038/s41467-019-10678-z
Patchett S, Musalgaonkar S, Malyutin AG, Johnson AW (2017) The T-cell leukemia related rpl10-R98S mutant traps the 60S export adapter Nmd3 in the ribosomal P site in yeast. PLoS Genet 13:e1006894. https://doi.org/10.1371/journal.pgen.1006894
Pérez-Fernández J, Román A, De Las RJ et al (2007) The 90S preribosome is a multimodular structure that is assembled through a hierarchical mechanism. Mol Cell Biol 27:5414–5429. https://doi.org/10.1128/MCB.00380-07
Rai J, Parker MD, Huang H et al (2021) An open interface in the pre-80S ribosome coordinated by ribosome assembly factors Tsr1 and Dim1 enables temporal regulation of Fap7. RNA 27:221–233. https://doi.org/10.1261/rna.077610.120
Sardana R (2013) From knobs to a central pseudoknot: understanding 40S ribosomal subunit biogenesis through Bud23. The University of Texas, Austin
Sardana R, Johnson AW (2012) The methyltransferase adaptor protein Trm112 is involved in biogenesis of both ribosomal subunits. Mol Biol Cell 23:4313–4322. https://doi.org/10.1091/mbc.E12-05-0370
Sardana R, White JP, Johnson AW (2013) The rRNA methyltransferase Bud23 shows functional interaction with components of the SSU processome and RNase MRP. RNA 19:828–840. https://doi.org/10.1261/rna.037671.112
Sardana R, Zhu J, Gill M, Johnson AW (2014) Physical and functional interaction between the methyltransferase Bud23 and the essential DEAH-box RNA helicase Ecm16. Mol Cell Biol 34:2208–2220. https://doi.org/10.1128/MCB.01656-13
Sardana R, Liu X, Granneman S et al (2015) The DEAH-box helicase Dhr1 dissociates U3 from the pre-rRNA to promote formation of the central pseudoknot. PLoS Biol 13:e1002083. https://doi.org/10.1371/journal.pbio.1002083
Schäfer T, Strauss D, Petfalski E et al (2003) The path from nucleolar 90S to cytoplasmic 40S pre-ribosomes. EMBO J 22:1370–1380. https://doi.org/10.1093/emboj/cdg121
Semlow DR, Staley JP (2012) Staying on message: ensuring fidelity in pre-mRNA splicing. Trends Biochem Sci 37:263–273. https://doi.org/10.1016/j.tibs.2012.04.001
Sharma K, Tollervey D (1999) Base pairing between U3 small nucleolar RNA and the 5’ end of 18S rRNA is required for pre-rRNA processing. Mol Cell Biol 19:6012–6019. https://doi.org/10.1128/mcb.19.9.6012
Small EC, Leggett SR, Winans AA, Staley JP (2006) The EF-G-like GTPase Snu114p regulates spliceosome dynamics mediated by Brr2p, a DExD/H box ATPase. Mol Cell 23:389–399. https://doi.org/10.1016/j.molcel.2006.05.043
Sun Q, Zhu X, Qi J et al (2017) Molecular architecture of the 90S small subunit pre-ribosome. Elife 6:e22086. https://doi.org/10.7554/eLife.22086
Talkish J, Biedka S, Jakovljevic J et al (2016) Disruption of ribosome assembly in yeast blocks cotranscriptional pre-rRNA processing and affects the global hierarchy of ribosome biogenesis. RNA 22:852–866. https://doi.org/10.1261/rna.055780.115
Thoms M, Thomson E, Baßler J et al (2015) The exosome is recruited to RNA substrates through specific adaptor proteins. Cell 162:1029–1038. https://doi.org/10.1016/j.cell.2015.07.060
Venema J, Henry Y, Tollervey D (1995) Two distinct recognition signals define the site of endonucleolytic cleavage at the 5’-end of yeast 18S rRNA. EMBO J 14:4883–4892. https://doi.org/10.1002/j.1460-2075.1995.tb00169.x
Wegierski T, Billy E, Nasr F, Filipowicz W (2001) Bms1p, a G-domain-containing protein, associates with Rcl1p and is required for 18S rRNA biogenesis in yeast. RNA 7:1254–1267. https://doi.org/10.1017/S1355838201012079
Wehner KA, Gallagher JEG, Baserga SJ (2002) Components of an interdependent unit within the SSU processome regulate and mediate its activity. Mol Cell Biol 22:7258–7267. https://doi.org/10.1128/MCB.22.20.7258-7267.2002
White J, Li Z, Sardana R et al (2008) Bud23 methylates G1575 of 18S rRNA and is required for efficient nuclear export of pre-40S subunits. Mol Cell Biol 28:3151–3161. https://doi.org/10.1128/MCB.01674-07
Woolford JL, Baserga SJ (2013) Ribosome biogenesis in the yeast Saccharomyces cerevisiae. Genetics 195:643–681. https://doi.org/10.1534/genetics.113.153197
Zhang L, Wu C, Cai G et al (2016) Stepwise and dynamic assembly of the earliest precursors of small ribosomal subunits in yeast. Genes Dev 30:718–732. https://doi.org/10.1101/gad.274688.115
Zhou Y, Musalgaonkar S, Johnson AW, Taylor DW (2019) Tightly-orchestrated rearrangements govern catalytic center assembly of the ribosome. Nat Commun 10:958. https://doi.org/10.1038/s41467-019-08880-0
Zhu J, Liu X, Anjos M et al (2016) Utp14 recruits and activates the RNA helicase Dhr1 to undock U3 snoRNA from the Preribosome. Mol Cell Biol 36:965–978. https://doi.org/10.1128/MCB.00773-15
Acknowledgements
We thank Dr. E. Sarinay-Cenik and Dr. R. Lin for their careful input and proofreading of this manuscript.
Funding
This work was supported by the National Institutes of Health (https://www.nih.gov/) Grant GM127127 awarded to AWJ and a fellowship from the University of Texas at Austin (https://www.utexas.edu/) Graduate School awarded to JJB.
Author information
Authors and Affiliations
Contributions
JJB and AWJ wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Communicated by Michael Polymenis.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Black, J.J., Johnson, A.W. Genetics animates structure: leveraging genetic interactions to study the dynamics of ribosome biogenesis. Curr Genet 67, 729–738 (2021). https://doi.org/10.1007/s00294-021-01187-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-021-01187-y