Abstract
In all living organisms, genome replication and cell division must be coordinated to produce viable offspring. In the event of DNA damage, bacterial cells employ the SOS response to simultaneously express damage repair systems and halt cell division. Extensive characterization of SOS-controlled cell division inhibition in Escherichia coli has laid the ground for a long-standing paradigm where the cytosolic SulA protein inhibits polymerization of the central division protein, FtsZ, and thereby prevents recruitment of the division machinery at the future division site. Within the last decade, it has become clear that another, likely more general, paradigm exists, at least within the broad group of Gram-positive bacterial species, namely membrane-localized, SOS-induced cell division inhibition. We recently identified such an inhibitor in Staphylococci, SosA, and established a model for SosA-mediated cell division inhibition in Staphylococcus aureus in response to DNA damage. SosA arrests cell division subsequent to the septal localization of FtsZ and later membrane-bound division proteins, while preventing progression to septum closure, leading to synchronization of cells at this particular stage. A membrane-associated protease, CtpA negatively regulates SosA activity and likely allows growth to resume once conditions are favorable. Here, we provide a brief summary of our findings in the context of what already is known for other membrane cell division inhibitors and we emphasize how poorly characterized these intriguing processes are mechanistically. Furthermore, we put some perspective on the relevance of our findings and future developments within the field.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacterial cell division is a tightly controlled process where the polymerization of FtsZ into a ring structure at the future division site plays a central role by acting as an anchor for assembly of the divisome. Although not fully conserved between Gram-negative and Gram-positive species, divisome assembly is considered a sequential process. A membrane subcomplex, FtsBLQ, is localized subsequent to FtsZ and presumably serves a regulatory function via multiple protein–protein interactions. This is followed by recruitment of FtsW and FtsI (a core divisome penicillin-binding protein named differently in different species) providing the enzymatic activity for septal peptidoglycan synthesis (Aarsman et al. 2005; Gamba et al. 2009; den Blaauwen and Luirink 2019; Reichmann et al. 2019; Taguchi et al. 2019; Wang et al. 2019). Multiple essential or stabilizing protein–protein interactions exist in the divisome, and the divisome is generally referred to as a single, complex entity, although recent super-resolution microscopy suggests that individual components may be arranged in a spatially distinct manner (Söderström and Daley 2017; Söderström et al. 2019). A plethora of additional protein components assists directly in the synthesis of septal peptidoglycan, while others function as regulatory spatiotemporal control elements. In rod-shaped model organisms such as Escherichia coli and Bacillus subtilis, for instance, the Min and nucleoid occlusion (SlmA/Noc) systems ensure that division takes place at mid-cell and as an event succeeding chromosomal segregation only (Adams and Errington 2009). It is also important that cell division is discontinued upon defective replication or if the DNA has sustained damage. Accordingly, delayed cell division should be an integral part of the bacterial SOS response.
The SOS response and cell division inhibition in E. coli
The existence of an inducible SOS repair system, first described and coined in E. coli in the early 1970s (Radman 1975), is now considered an almost universal stress response for bacteria experiencing compromised DNA integrity. SOS induction is mediated by RecA which upon binding to single-stranded DNA stimulates auto-proteolysis and inactivation of the conserved transcriptional repressor LexA. The LexA-controlled genes, the SOS regulon that following DNA damage becomes de-repressed, encode activities involved in DNA repair and cell division control [see reviews by Kelley (2006), Kreuzer (2013), Baharoglu and Mazel (2014), Henrikus et al. (2018), and Culyba (2019)]. Cell division inhibition ensuing SOS induction in E. coli is mediated by SulA. This cytosolic protein halts cell division by binding to FtsZ, inhibiting its polymerization and thereby abolishing formation of the septal ring that marks the future division site. A hallmark of SulA-mediated arrest of cell division is the characteristic filamentous growth phenotype resulting from continued cell elongation in the absence of division (Huisman and D'Ari 1981; Huisman et al. 1984; Higashitani et al. 1995; Mukherjee et al. 1998; Trusca et al. 1998; Cordell et al. 2003). Cell division inhibition needs to be controlled in a way that allows cells to resume division once relieved of SOS-inducing conditions. In line with the filamentous phenotype of E. coli lon mutants, the Lon protease has been identified as a direct negative regulator of SulA (Mizusawa and Gottesman 1983; Schoemaker et al. 1984; Sonezaki et al. 1995), likely assisted by another cytoplasmic protease ClpYQ (Seong et al. 1999; Wu et al. 1999). SulA-mediated division control in E. coli is by far the most well-understood SOS checkpoint amongst bacteria. It appears, though, that the processes by which more distantly related bacteria ensure delayed cell division are notably different. Most, if not all, subsequently described cell division inhibitors have in common that they are membrane bound and do not target FtsZ directly.
SOS-mediated cell division inhibition via membrane-localized effectors
The first Gram-positive, SOS-regulated cell division inhibitor to be described was YneA in B. subtilis. Initial characterization showed that yneA causes cell filamentation in a LexA (DinR)-dependent manner and that suppression of cell division coincides with decreased FtsZ localization at the division site (Kawai et al. 2003). However, following YneA overexpression, FtsZ rings are formed at mid-cell, suggesting that the observed halt in cell division is mediated via binding of YneA to an alternative, potentially membrane-bound, target in the divisome (Mo and Burkholder 2010). This was supported by mutational studies showing that specific mutations in the single transmembrane segment of the YneA protein reduce its activity without affecting translocation or stability. A C-terminally truncated version of YneA also lacking a LysM domain, a motif suggesting peptidoglycan binding, is also non-functional but capable of being exported (Mo and Burkholder 2010). As such, both the membrane domain and an extracellular portion of the protein could be functionally important. The target mechanism of YneA is, however, presently unknown. Importantly, it was proposed and established experimentally that cell division inhibition by YneA is a reversible process and that resumption of cell division is likely achieved via extracellular proteolysis of YneA (Mo and Burkholder 2010). YneA homologs are also present in other Bacillales such as Listeria monocytogenes (van der Veen et al. 2007) and Bacillus megaterium (Buchholz et al. 2013).
In the serious human pathogen, Staphylococcus aureus, sosA is expressed as part of the SOS response and encoded in a divergent orientation next to the lexA locus (Anderson et al. 2006; Cirz et al. 2007; Cohn et al. 2011). By inference, this genetic synteny suggested that it encodes a suppressor of cell division (i.e., analogous to yneA in B. subtilis (Kawai et al. 2003)), although no significant sequence similarity is evident in the encoded proteins. Indeed, we recently disclosed that SosA in Staphylococcus is a member of the SOS-induced cell division inhibitor family (Bojer et al. 2019).
Importantly, we found that the 77 amino acid membrane protein is critical for survival of cells subjected to DNA damage by the mutagen Mitomycin C and a major determinant of cell size. Under DNA-damaging conditions, sosA mutant cells continue cell division at the expense of viability. Ectopic expression of SosA alone arrests cell division and consequently leads to cell enlargement and impairment of the ability to form colonies. Based on the primary sequence of SosA, it is predicted to consist of a small N-terminal part, a single membrane-spanning segment, and a larger extracellular C-terminal part of approximately 47 amino acids, but in contrast to YneA it does not harbor a LysM domain. When investigating different C-terminally truncated variants of SosA and mutants with substitutions of conserved amino acids, we could show that the extracellular portion of the protein is of critical importance for activity. The divisomal target of SosA remains undisclosed, but expression of SosA stalls the division at a point where the divisome is capable of initiating but not completing septal peptidoglycan synthesis. Consistent with this finding, central divisome proteins such as FtsZ, EzrA and GpsB are correctly localized in SosA-arrested cells (Bojer et al. 2019).
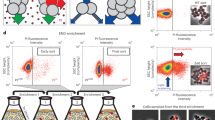
Serendipitously, we found that a small (10 amino acid) deletion of the C-terminal of SosA leads to apparent accumulation of the protein and augmentation of its division inhibitory activity. Inspired by the inactivation-by-proteolysis mechanism indicated for YneA/B. subtilis by Mo and Burkholder (2010), we sought to identify a membrane protease that could serve as a negative regulator of SosA. We revealed that the carboxy-terminal processing protease, CtpA, is likely to control the activity/abundance of SosA and that an S. aureus ctpA mutant is hypersensitive to DNA damage as well as to SosA expression (Bojer et al. 2019). Importantly, this finding was corroborated by independent experiments conducted by Burby et al. (2018) showing that in B. subtilis YneA is a substrate for the proteolytic activity of CtpA. This represents a conceptual advance in the field in that membrane-bound cell division inhibitors among Gram-positive bacteria may be controlled in situ by membrane proteases like CtpA. Figure 1 shows a simplified model of how cell division is regulated upon SOS induction in S. aureus, a model we hope to be expanded and elaborated upon in future studies.
Conceptual model of how cell division in S. aureus is regulated upon and after induction of the SOS response. Cell division inhibitor SosA prevents progression but not initiation of septum synthesis. Membrane protease CtpA controls SosA activity and allows continued division once DNA damage has been repaired
Apart from YneA in Bacillus and SosA in Staphylococcus, a few other SOS-induced, membrane-bound cell division inhibitors have been described. ChiZ (Rv2719c) is a membrane protein that inhibits cell division in Mycobacterium tuberculosis. It is likewise under the control of LexA and of importance for survival of M. tuberculosis under genotoxic stress. No direct interaction with FtsZ and no effect on polymerization of the protein have been reported, while interactions with FtsI and FtsQ could provide an important clue to its division inhibitory activity. As for YneA in B. subtilis, ChiZ has an extracytoplasmic LysM peptidoglycan-binding domain, and localization studies have shown that ChiZ is targeted to zones of nascent peptidoglycan synthesis. Moreover, the protein was reported to have cell wall hydrolase activity (Chauhan et al. 2006; Vadrevu et al. 2011). The LysM domain is, however, dispensable for the cell division inhibitory activity of the protein (Vadrevu et al. 2011). This, together with the reporting that the cell wall hydrolase activity, which is confined to the remaining N-terminal part of the protein, might be an experimental artifact (Escobar and Cross 2018), leaves ChiZ poorly understood mechanistically.
Corynebacterium glutamicum encodes a protein, DivS, which is expressed during and involved in survival under DNA-damaging conditions, and it is in itself sufficient to cause cell elongation in the absence of SOS induction. Like S. aureus SosA, DivS does not contain a LysM domain. However, in contrast to SosA, DivS is characterized by a large N-terminal segment but a small C-terminal segment relative to the predicted transmembrane domain. Microscopic analysis suggests that the protein interferes with FtsZ ring formation and septal peptidoglycan synthesis (Ogino et al. 2008). It is unclear to what extent these morphological effects are indirect consequences of cell division suppression at a later stage, as no direct interaction with FtsZ have been reported. Noteworthy, large deletions of the predicted intracellular N-terminus of the protein are tolerated with respect to activity, whereas a minor truncation to the predicted extracellular C-terminus abolishes activity of the protein (Ogino et al. 2008). It is unknown if this finding informs directly on an extracellular target/activity or if it is related to compromised localization and/or instability of the protein product.
Abovementioned membrane-bound cell division inhibitors were all identified in Gram-positive species. However, in the Gram-negative bacterium Caulobacter crescentus, cell division arrest and cell elongation upon DNA damage is mediated by a small 29 amino acid membrane protein, SidA. Detailed experimental analyses have shown that, while expression of SidA alone is sufficient to suppress division, expression of the inhibitor does not prevent polymerization of FtsZ at mid-cell, nor does it impede the full assembly of the late-acting membrane-located divisome constituents (e.g., FtsW, FtsN, and FtsI) (Modell et al. 2011). SidA does, however, interact with this divisomal subcomplex, most likely FtsW, as established by bacterial two-hybrid system analysis in combination with mapping of suppressor mutants being non-responsive to SidA overproduction. Interestingly, SidA does not hinder septal peptidoglycan synthesis, suggesting a critical checkpoint activity in preventing final septum constriction (Modell et al. 2011).
The collected evidence suggests that diverse bacteria cope with DNA damage by SOS response-mediated expression of membrane-bound cell division inhibitors. Despite this shared theme, these endogenous cell division inhibitors vary considerably in sequence and size and as they are not well characterized functionally, it is unknown how much they overlap with respect to target mechanism.
Perspectives
Bacterial cell division is a fascinating event and is orchestrated by a sequential, interdependent assembly of proteins; some of which are core constituents of the divisome with high conservation between different bacteria, whereas others appear restricted to certain genera. Both groups may represent valuable targets for antibiotic development (Lock and Harry 2008; den Blaauwen et al. 2014). Bacteria have evolved different means of regulating cell division as a correlative to DNA replication and associated counterproductive DNA damage. For an overview on bacterial cell division inhibition via both SOS-independent and -dependent processes, we refer the reader to a recent concise review by Burby and Simmons (2020). Across bacterial genera, the SOS response leads to expression of specific proteins that suppress cell division. Conceivably, these cell division inhibitors target essential processes and we believe that these proteins, in particular, could lead to future therapeutic target discovery and that they deserve further study in this respect. Especially for the expanding group of membrane-bound cell division inhibitors, although at present their molecular mechanism of action is too vaguely understood to allow prediction of druggable targets.
Future work should aim to describe how SosA arrests cell division in Staphylococci. Our preliminary data, e.g., obtained by two-hybrid analysis and suppressor mutant screens (unpublished data), support the idea that SosA acts on (or via) core membrane divisome proteins. We are currently undertaking alternative experimental approaches to better delineate endogenous cell division inhibition in this species.
Molecular studies of bacterial cell division are complicated due to the essentiality of many components and the interdependence of those. Current experimental approaches rely to a large extent on microscopy in combination with fluorescent protein fusions or alternative staining methods. Such phenotypic studies are not trivial since bacteria often grow asynchronously with individual cells being in different phases of cell division. We observed that SosA expression efficiently arrests S. aureus cells at a pre-septational stage (Bojer et al. 2019), and we foresee that this activity could be used for investigational purposes to generate synchronized cell populations.
References
Aarsman ME, Piette A, Fraipont C, Vinkenvleugel TM, Nguyen-Distèche M, den Blaauwen T (2005) Maturation of the Escherichia coli divisome occurs in two steps. Mol Microbiol 55:1631–1645
Adams DW, Errington J (2009) Bacterial cell division: assembly, maintenance and disassembly of the Z ring. Nat Rev Microbiol 7:642–653
Anderson KL, Roberts C, Disz T, Vonstein V, Hwang K, Overbeek R, Olson PD, Projan SJ, Dunman PM (2006) Characterization of the Staphylococcus aureus heat shock, cold shock, stringent, and SOS responses and their effects on log-phase mRNA turnover. J Bacteriol 188:6739–6756
Baharoglu Z, Mazel D (2014) SOS, the formidable strategy of bacteria against aggressions. FEMS Microbiol Rev 38:1126–1145
Bojer MS, Wacnik K, Kjelgaard P, Gallay C, Bottomley AL, Cohn MT, Lindahl G, Frees D, Veening JW, Foster SJ, Ingmer H (2019) SosA inhibits cell division in Staphylococcus aureus in response to DNA damage. Mol Microbiol 112:1116–1130
Buchholz M, Nahrstedt H, Pillukat MH, Deppe V, Meinhardt F (2013) yneA mRNA instability is involved in temporary inhibition of cell division during the SOS response of Bacillus megaterium. Microbiology 159:1564–1574
Burby PE, Simmons LA (2020) Regulation of cell division in bacteria by monitoring genome integrity and DNA replication status. J Bacteriol 202:e00408-19
Burby PE, Simmons ZW, Schroeder JW, Simmons LA (2018) Discovery of a dual protease mechanism that promotes DNA damage checkpoint recovery. PLoS Genet 14:e1007512
Chauhan A, Lofton H, Maloney E, Moore J, Fol M, Madiraju MV, Rajagopalan M (2006) Interference of Mycobacterium tuberculosis cell division by Rv2719c, a cell wall hydrolase. Mol Microbiol 62:132–147
Cirz RT, Jones MB, Gingles NA, Minogue TD, Jarrahi B, Peterson SN, Romesberg FE (2007) Complete and SOS-mediated response of Staphylococcus aureus to the antibiotic ciprofloxacin. J Bacteriol 189:531–539
Cohn MT, Kjelgaard P, Frees D, Penadés JR, Ingmer H (2011) Clp-dependent proteolysis of the LexA N-terminal domain in Staphylococcus aureus. Microbiology 157:677–684
Cordell SC, Robinson EJ, Lowe J (2003) Crystal structure of the SOS cell division inhibitor SulA and in complex with FtsZ. Proc Natl Acad Sci USA 100:7889–7894
Culyba MJ (2019) Ordering up gene expression by slowing down transcription factor binding kinetics. Curr Genet 65:401–406
den Blaauwen T, Luirink J (2019) Checks and balances in bacterial cell division. MBio 10:e00149–e219
den Blaauwen T, Andreu JM, Monasterio O (2014) Bacterial cell division proteins as antibiotic targets. Bioorg Chem 55:27–38
Escobar CA, Cross TA (2018) False positives in using the zymogram assay for identification of peptidoglycan hydrolases. Anal Biochem 543:162–166
Gamba P, Veening JW, Saunders NJ, Hamoen LW, Daniel RA (2009) Two-step assembly dynamics of the Bacillus subtilis divisome. J Bacteriol 191:4186–4194
Henrikus SS, van Oijen AM, Robinson A (2018) Specialised DNA polymerases in Escherichia coli: roles within multiple pathways. Curr Genet 64:1189–1196
Higashitani A, Higashitani N, Horiuchi K (1995) A cell division inhibitor SulA of Escherichia coli directly interacts with FtsZ through GTP hydrolysis. Biochem Biophys Res Commun 209:198–204
Huisman O, D'Ari R (1981) An inducible DNA replication-cell division coupling mechanism in E. coli. Nature 290:797–799
Huisman O, D'Ari R, Gottesman S (1984) Cell-division control in Escherichia coli: specific induction of the SOS function SfiA protein is sufficient to block septation. Proc Natl Acad Sci USA 81:4490–4494
Kawai Y, Moriya S, Ogasawara N (2003) Identification of a protein, YneA, responsible for cell division suppression during the SOS response in Bacillus subtilis. Mol Microbiol 47:1113–1122
Kelley WL (2006) Lex marks the spot: the virulent side of SOS and a closer look at the LexA regulon. Mol Microbiol 62:1228–1238
Kreuzer KN (2013) DNA damage responses in prokaryotes: regulating gene expression, modulating growth patterns, and manipulating replication forks. Cold Spring Harb Perspect Biol 5:a012674
Lock RL, Harry EJ (2008) Cell-division inhibitors: new insights for future antibiotics. Nat Rev Drug Discov 7:324–338
Mizusawa S, Gottesman S (1983) Protein degradation in Escherichia coli: the lon gene controls the stability of sulA protein. Proc Natl Acad Sci USA 80:358–362
Mo AH, Burkholder WF (2010) YneA, an SOS-induced inhibitor of cell division in Bacillus subtilis, is regulated posttranslationally and requires the transmembrane region for activity. J Bacteriol 192:3159–3173
Modell JW, Hopkins AC, Laub MT (2011) A DNA damage checkpoint in Caulobacter crescentus inhibits cell division through a direct interaction with FtsW. Genes Dev 25:1328–1343
Mukherjee A, Cao C, Lutkenhaus J (1998) Inhibition of FtsZ polymerization by SulA, an inhibitor of septation in Escherichia coli. Proc Natl Acad Sci USA 95:2885–2890
Ogino H, Teramoto H, Inui M, Yukawa H (2008) DivS, a novel SOS-inducible cell-division suppressor in Corynebacterium glutamicum. Mol Microbiol 67:597–608
Radman M (1975) SOS repair hypothesis: phenomenology of an inducible DNA repair which is accompanied by mutagenesis. Basic Life Sci 5A:355–367
Reichmann NT, Tavares AC, Saraiva BM, Jousselin A, Reed P, Pereira AR, Monteiro JM, Sobral RG, VanNieuwenhze MS, Fernandes F, Pinho MG (2019) SEDS-bPBP pairs direct lateral and septal peptidoglycan synthesis in Staphylococcus aureus. Nat Microbiol 4:1368–1377
Schoemaker JM, Gayda RC, Markovitz A (1984) Regulation of cell division in Escherichia coli: SOS induction and cellular location of the SulA protein, a key to lon-associated filamentation and death. J Bacteriol 158:551–561
Seong IS, Oh JY, Yoo SJ, Seol JH, Chung CH (1999) ATP-dependent degradation of SulA, a cell division inhibitor, by the HslVU protease in Escherichia coli. FEBS Lett 456:211–214
Söderström B, Daley DO (2017) The bacterial divisome: more than a ring? Curr Genet 63:161–164
Söderström B, Chan H, Daley DO (2019) Super-resolution images of peptidoglycan remodelling enzymes at the division site of Escherichia coli. Curr Genet 65:99–101
Sonezaki S, Ishii Y, Okita K, Sugino T, Kondo A, Kato Y (1995) Overproduction and purification of SulA fusion protein in Escherichia coli and its degradation by Lon protease in vitro. Appl Microbiol Biotechnol 43:304–309
Taguchi A, Welsh MA, Marmont LS, Lee W, Sjodt M, Kruse AC, Kahne D, Bernhardt TG, Walker S (2019) FtsW is a peptidoglycan polymerase that is functional only in complex with its cognate penicillin-binding protein. Nat Microbiol 4:587–594
Trusca D, Scott S, Thompson C, Bramhill D (1998) Bacterial SOS checkpoint protein SulA inhibits polymerization of purified FtsZ cell division protein. J Bacteriol 180:3946–3953
Vadrevu IS, Lofton H, Sarva K, Blasczyk E, Plocinska R, Chinnaswamy J, Madiraju M, Rajagopalan M (2011) ChiZ levels modulate cell division process in mycobacteria. Tuberculosis (Edinb) 91:S128–135
van der Veen S, Hain T, Wouters JA, Hossain H, de Vos WM, Abee T, Chakraborty T, Wells-Bennik MH (2007) The heat-shock response of Listeria monocytogenes comprises genes involved in heat shock, cell division, cell wall synthesis, and the SOS response. Microbiology 153:3593–3607
Wang M, Fang C, Ma B, Luo X, Hou Z (2019) Regulation of cytokinesis: FtsZ and its accessory proteins. Curr Genet. https://doi.org/10.1007/s00294-019-01005-6
Wu WF, Zhou Y, Gottesman S (1999) Redundant in vivo proteolytic activities of Escherichia coli Lon and the ClpYQ (HslUV) protease. J Bacteriol 181:3681–3687
Acknowledgements
The work in the authors’ lab was supported by grants from the Danish Council for Independent Research (1337-00129 and 1335-00772 to MSB) and the Danish National Research Foundation (DNRF120 to HI). We thank members of our own lab as well as members from the laboratories of Prof. Simon Foster (University of Sheffield) and Prof. Jan-Willem Veening (University of Lausanne) for encouraging discussions and collaboration.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Bojer, M.S., Frees, D. & Ingmer, H. SosA in Staphylococci: an addition to the paradigm of membrane-localized, SOS-induced cell division inhibition in bacteria. Curr Genet 66, 495–499 (2020). https://doi.org/10.1007/s00294-019-01052-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-019-01052-z