Abstract
Since its discovery in the early 70s, the 2 micron plasmid of Saccharomyces cerevisiae continues to intrigue researchers with its high protein-coding capacity and a selfish nature yet high stability, earning it the title of a ‘miniaturized selfish genetic element’. It codes for four proteins (Rep1, Rep2, Raf1, and Flp) vital for its own survival and recruits several host factors (RSC2, Cohesin, Cse4, Kip1, Bik1, Bim1, and microtubules) for its faithful segregation during cell division. The plasmid maintains a high-copy number with the help of Flp-mediated recombination. The plasmids organize in the form of clusters that hitch-hike the host chromosomes presumably with the help of the plasmid-encoded Rep proteins and host factors such as microtubules, Kip1 motor, and microtubule-associated proteins Bik1 and Bim1. Although there is no known yeast cell phenotype associated with the 2 micron plasmid, excessive copies of the plasmid are lethal for the cells, warranting a tight control over the plasmid copy number. This control is achieved through a combination of feedback loops involving the 2 micron encoded proteins. Thus, faithful segregation and a concomitant tightly controlled plasmid copy number ensure an optimized benign parasitism of the 2 micron plasmid within budding yeast.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The 2 micron plasmid
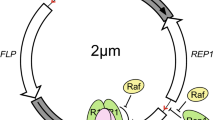
The yeast 2 micron circular DNA is an endogenous plasmid of 6318 bp (Hartley and Donelson 1980) resident in the nuclei of most of the Saccharomyces strains (Stevens and Moustacchi 1971; Clark-Walker and Miklos 1974; Petes and Williamson 1975; Boach 1983) with an average copy number close to 60 (Clark-Walker and Miklos 1974; Gerbaud et al. 1979; Murray and Cesareni 1986). This is a classic example of selfish DNA that codes for the elements required for its own maintenance (Broach et al. 1979), but persists in the host without jeopardizing the fitness of the host. The 2 micron plasmid is schematically represented by notion as a dumb-bell-shaped structure (Fig. 1a), where the two unique halves of the plasmid are separated by the two Flp recognition targets (FRTs); the Flp-mediated recombination at the FRTs causes the plasmid to switch from A form to B form depending on the relative orientation of the two halves with respect to each other (Beggs 1978; Broach and Hickst 1980; Hartley and Donelson 1980; Broach et al. 1982; Broach 1982). The plasmid confers a negligible growth disadvantage to the host at its normal copy number which is justified for a parasite to impose minimum burden on the host. Intriguingly, although the plasmid does not cause any known selective advantage to the host (Futcher and Cox 1983; Mead et al. 1986), it is among the most stable plasmids with a loss rate of about 10−5 per cell per generation (Futcher and Cox 1983). Owing to this high stability and copy number, the components of the 2 micron plasmid are being used as multicopy vectors successfully for many years for high-copy gene expression in yeast (Gerbaud et al. 1979; Boach 1983). Importantly, this high chromosome-like stability is due to two plasmid-borne systems, namely, partitioning and amplification systems, encoded by four ORFs and two cis-acting sequences (Broach et al. 1979; Hartley and Donelson 1980; Jayaram et al. 1983) (Fig. 1b). Besides the ORFs coding for the four known proteins, several short and long transcripts are also transcribed with unknown products and functions (Broach et al. 1979) (Fig. 1b).
Organization of the 2 µm plasmid genome. a Usual dumb-bell representation of the 2 micron plasmid showing the two unique halves and the intervening inverted repeat (FRTs). Site-specific recombination mediated by the Flp–FRT recombination system changes one form of plasmid to the other form and vice versa. b Map of a 2 micron plasmid showing the four ORFs, the partitioning locus STB, the FRTs, and the origin of replication Ori. STB is designated by the SnaBI and PstI sites, and STB-distal (dist.) and proximal (prox.) with respect to the Ori are demarcated by the HpaI restriction site. The 2 micron map also shows the four ORFs (REP1, REP2, FLP, and RAF1) encoding the proteins that are responsible for the stable maintenance of the plasmid. The promoters are designated by their ability to fire the expression of a LacZ reporter. Apart from the four ORFs, there are several other, yet uncharacterized, transcripts (shown in red) found to be transcribed by the 2 micron genome. c STB proximal showing the five 69 bp repeats as 1–5 and the percent similarity with the consensus sequence (TTTTTPuPyAGAACAAAAAYGCAACGCGAGAGCGCTAATTTT(A/T)CAAACAAAGAATCTGAGCTTCA) in parentheses (Hartley and Donelson 1980)
The plasmid-encoded proteins Rep1 and Rep2 along with the cis-acting locus STB form the partitioning system which helps in the equal segregation of the plasmids between the mother and the daughter cells during cell division (Cashmore et al. 1986). Apart from these plasmid-borne elements, an increasing number of host-encoded factors are found to be involved in the faithful segregation of the 2 micron plasmid (Kikuchi 1983; Velmurugan et al. 1998; Mehta et al. 2002, 2005; Hajra et al. 2006; Ghosh et al. 2007, 2009; Cui et al. 2009; Ma et al. 2013; Prajapati et al. 2017). However, missegregation events occur despite the presence of an efficient partitioning system. Under such conditions, a drop in the plasmid copy number (PCN) has to be recovered during the replication in the S phase of the cell cycle. It has been shown that plasmid replication is restricted to S phase and the 2 micron origin of replication fires once per S phase (Zakian et al. 1979). Therefore, multiple initiations of replication within a span of one cell cycle cannot be an option to increase the PCN. It is believed that when PCN drops in the daughter cell due to a rare missegregation event, the steady-state PCN is restored through the activation of the amplification system which is comprised of Flp site-specific recombinase and its target site FRTs (Volkert and Broach 1986). Once the steady-state PCN is reached, the amplification system is switched off (Futcher 1986). A high PCN causes extra metabolic burden resulting in reduced growth rate and petite colonies (Volkert and Broach 1986; Chen et al. 2005). However, it is not known how any increase in the PCN due to any runaway amplification is brought back to the steady-state level.
Despite several studies on the elucidation of functions of Rep proteins and Flp recombinase in partitioning and copy number amplification of the 2 micron plasmid, the characterization of the plasmid-encoded protein, Raf1, remains scanty. An earlier study revealed a role of Raf1 in the plasmid stability and the protein had been shown to interact with the partitioning locus STB in vitro (Hadfield et al. 1995). Whereas the dependence of the interaction of Raf1 with STB on the Rep proteins is debatable from the in vitro experiments, very recently, using in vivo experiments, we found that this interaction is independent of the Rep proteins (Rizvi et al. 2017). Apart from its interaction with STB and influencing plasmid partitioning, Raf1 is believed to have a regulatory role on the plasmid copy number control as well as it has been shown to activate FLP (Murray et al. 1987), although the exact mechanism of this FLP activation is still speculative. From the earlier and our recent data, it appears that there are roles of Raf1 in both the partitioning and the copy number controls. This dual role may stem from this protein being an anti-repressor to the Rep1–Rep2 repressor complex, which is at the center of both the partitioning and the amplification systems as discussed in the subsequent sections (Murray et al. 1987; Rizvi et al. 2017).
Besides REP1, REP2, RAF1, and FLP ORFs, there are several, yet uncharacterized, transcripts transcribed by the 2 micron genome. These transcripts include both short transcripts spanning a single ORF (such as a 700 bp transcript spanning the RAF1 ORF), and very long transcripts (such as 2600 and 1950 bp transcripts) spanning REP2–FLP and REP1–RAF1 ORFs, respectively (Fig. 1b). Apart from these transcripts, there are other transcripts transcribed from the opposite strand of the plasmid. The biological functions of all these transcripts are unknown, but it is speculated that either these transcripts undergo differential splicing to produce the mature transcripts of the four ORFs, or they might act as attenuators controlling the translation from the four ORFs (Broach et al. 1979).
The partitioning system
The partitioning system of the 2 micron plasmid is responsible for its faithful segregation during cell division (Cashmore et al. 1986). Effectively, the partitioning system comprising of the proteins Rep1, Rep2, and the DNA locus STB (Jayaram et al. 1983; Kikuchi 1983) provides a mechanism to overcome the mother bias, a diffusion barrier that keeps the plasmid trapped in the mother cell during the cell division (Murray and Szostak 1983; Shcheprova et al. 2008; Gehlen et al. 2011; Khmelinskii et al. 2011). An important question that arises here is why does a multicopy plasmid (which reduces the chances of no segregation) needs an active partitioning system despite having a robust amplification system that can restore the copy number if any inadvertent missegregation occurs? The answer may lie in the organization of the plasmid inside the nucleus. When the FlORS (fluorescence operator–reporter system) was used to visualize the plasmids and study their localization and dynamics in real time, it was found that the plasmids are localized in the nucleus as a cluster made up of dynamic foci (3–5). Such a clustered organization reduces the effective copy number of the plasmids to one for partitioning (Ahn et al. 1997; Scott-Drew and Murray 1998; Velmurugan et al. 2000; Scott-Drew et al. 2002). Thus, a mechanism involving simple diffusion followed by amplification may not be effective. The partitioning system might have evolved to facilitate cluster segregation, thereby overcoming this diffusion barrier. This plasmid cluster (similar to a chromosome) utilizes host factors as part of its partitioning system for equal segregation.
Partitioning locus STB
The cis-acting locus STB of the 2 micron plasmid is a ~600 bp DNA sequence between the AvaI and PstI restriction sites (Murray and Cesareni 1986). STB is essential for the faithful partitioning of the 2 micron plasmid along with the trans-acting ORFs REP1 and REP2 (Jayaram et al. 1983; Kikuchi 1983). In the absence of STB (or the trans-acting ORFs REP1 or REP2), the 2 micron-based plasmids are rapidly lost due to extreme mother bias during mitosis; a behavior similar to an ARS plasmid (Murray and Szostak 1983). Structurally, STB is divided into two segments—STB distal (PstI–HpaI fragment, farther from the Ori) and the STB proximal (HpaI–AvaI fragment, proximal to the Ori) depending on their proximity to the 2 micron origin of replication (Ori). The STB proximal has five tandem repeats of a 62 bp (Fig. 1c) sequence and is sufficient for the stability of the plasmid. The STB distal overlaps the 3′ end of the RAF1 ORF and is supposed to be responsible for halting the transcription from RAF1 and preventing the runaway transcription through the STB proximal and the Ori and thus might help in plasmid stability indirectly (Fig. 1) (Murray and Cesareni 1986; Papacs et al. 2004). Another consensus sequence [TC(T rich)13,15 ATCTTG] within STB is shared by other different loci within the plasmid. The number of repeats harboring this consensus sequence varies among the loci such as one repeat each within −10 to +20 of the REP2 ORF and +714 to +732 of the REP1 ORF and three repeats each within −90 to +1 of the FLP ORF and within the PstI–SnaBI fragment of the STB (Murray and Cesareni 1986). It is speculated that these loci are the recognition target sites for the Rep1–Rep2 complex (Murray and Cesareni 1986; Rizvi et al. 2017). STB shows a substantial polymorphism among different isolates both in the consensus sequence of the core 62 bp repeat unit and the number of such units present in the tandem repeats. Sequence comparison among the isolates suggests a combination of both recombination and drift-driven variation in STB (Wei et al. 1991).
The Rep proteins
Apart from STB, the Rep proteins (Rep1 and Rep2) are the primary factors responsible for the 2 micron plasmid stability (Jayaram et al. 1983; Kikuchi 1983; Cashmore et al. 1986). Both Rep1 and Rep2 have an N-terminal oligomerization domain and a C-terminal DNA binding domain and both form homo- and hetero-complexes (Ahn et al. 1997; Velmurugan et al. 1998; Sengupta et al. 2001). Rep1 and Rep2 colocalize to the nucleus in the form of discrete foci, but in the absence of their C-terminal regions, they fail to localize to the nucleus. The nuclear localization can be restored by adding an NLS (SV40 T antigen) to the truncated protein, thus suggesting that the Rep protein’s C-terminal ends function as an NLS (Velmurugan et al. 1998). The characterization of the functional domains of the Rep proteins revealed that N-terminal regions help in the interaction among the Rep proteins, whereas the C-terminal regions harbor the DNA binding property and help the proteins to bind STB (Sengupta et al. 2001). Although no structural information is available for the Rep proteins, it was speculated that Rep1 may form a fibrous structure and may attach to the nuclear pore complex, nuclear matrix, or to lamina through its C-terminal domain based. This is based on the sequence similarity of its C-terminal domain with that of coiled-coil proteins such as myosin heavy chain, vimentin and nuclear lamins A and C, and co-purification of Rep1 with a nuclear karyoskeletal fraction (Wu et al. 1987).
The Rep–STB system
The Rep–STB system is composed of two plasmid-encoded proteins Rep1 and Rep2 and a cis-acting locus STB (Fig. 1). The Rep proteins show association to the STB and are localized in the form of discrete clusters inside the nucleus. The average number of these foci ranges from 4 to 5 in the G1 cells to ~8, and by the time, the cell enters into mitosis (Scott-Drew and Murray 1998). Nuclear localization of Rep1 and Rep2 is not independent of each other. The presence of Rep2 is necessary for the post-translational stability (Pinder et al. 2013) and complete nuclear localization of Rep1 (Ahn et al. 1997). However, nuclear localization of Rep2 is independent of Rep1. Notably, the number of plasmid clusters and, hence, the number of foci are independent of the PCN. However, an increased number of plasmid and, hence, higher levels of STB enhance the nuclear localization of the Rep proteins with an increase in the compactness of the foci (Scott-Drew and Murray 1998). The increased compactness of the foci also suggests a possible role of the Rep proteins in the tight packaging of the 2 micron plasmid into clusters.
Although Rep1 and Rep2 form a complex, fluorescence intensities of the stained Rep1 and Rep2 do not follow an identical pattern. Fluorescence intensity of Rep1 peaks up in the S/G2 phase, while for Rep2, it is at the G2/M phase (Scott-Drew and Murray 1998). This temporal difference may suggest distinct functions of Rep1 and Rep2 at individual levels. It may also suggest the requirement of the Rep1–Rep2 complex in different stoichiometric proportions of the subunits at different stages of the cell cycle.
Interestingly, when both the Rep proteins are expressed together, they were found to co-localize with the chromosome in a [Cir0] cell ([Cir0] cells are devoid of the 2 micron plasmid as opposed to [Cir+] cells) irrespective of the presence of a resident STB locus (Mehta et al. 2002). This observation suggests that Rep proteins can attach to the chromatin independent of a resident 2 micron plasmid and, therefore, can act as a landing platform for the plasmids to be tethered to the chromatin. The point to be noted here is that the interaction of the Rep proteins is not only limited to the endogenous yeast chromatin but also was found to bind to the chromatin of mammalian cells symmetrically in the form of discrete foci in a chromatin spread (Liu et al. 2016). This suggests that the Rep proteins by themselves might have an intrinsic property, as discussed above, to bind to the chromatin in a sequence-independent manner. This reinforces the fact that the Rep proteins can promote a chromosome hitch-hike mechanism of plasmid segregation by tethering plasmids to the chromosomes. As a matter of similitude, episomal form of several mammalian viruses also co-segregate with their host chromosomes (discussed later).
Host factors involved in plasmid partitioning
The remarkable high mitotic stability of the 2 micron plasmid close to the level of chromosomes warrants that a robust mechanism must be engaged in the maintenance of the plasmid. Hadfield et al. revealed for the first time that some host factors may bind to STB in addition to the 2 micron encoded proteins (Hadfield et al. 1995). Later, Shf1 was demonstrated to bind to the STB and was identified as the first host factor involved in the plasmid stability (Velmurugan et al. 1998). In the subsequent studies using fluorescence labeled plasmids, it was observed that the plasmid and the chromosome segregations are coupled, and in chromosome segregation mutants (such as in ipl1–2 with nonfunctional Aurora B kinase), both plasmid and chromosomes were found to be equally erroneous (Velmurugan et al. 2000). These observations led to a hypothesis that the chromosome-segregation machineries may be involved as host factors in the faithful partitioning of the plasmids. The hypothesis turned out to be true as a series of studies have revealed the involvement of several host factors in the plasmid segregation as discussed below.
-
1.
Cohesin Timely loading and cleavage of cohesin are the key events for equal chromosome segregation during cell cycle (Nasmyth and Haering 2009; Mehta et al. 2012). It was found that cohesin loading occurs at ‘STB’ of plasmid the same as ‘CEN’ of the chromosome and is a key factor for plasmid segregation from mother to daughter (Mehta et al. 2002) (Fig. 2). The active and independent recruitment of cohesin to the plasmid is supported by two observations—(1) the association of Mcd1 (kleisin subunit of cohesin) to the STB was independent of CEN when Mcd1 was expressed ectopically during G1 (Mehta et al. 2002) and (2) loading of cohesin at STB requires intact nuclear microtubules in contrast to the CEN (Mehta et al. 2005). The binding of whole cohesin complex was confirmed by showing the association of Smc1 and Smc3 to the STB (Mehta et al. 2002). Use of non-cleavable cohesin revealed that the cleavage of cohesin is essential for plasmid disjunction just like the chromosomes (Mehta et al. 2002). To demonstrate that the acquisition of cohesin is really required for the act of cohesion of the replicated plasmids, a single copy derivative of the plasmid was developed (Ghosh et al. 2007). Using fluorescent-tagged single-copy plasmid, it was shown that the plasmids, following replication, remain cohesed in a Ctf7 (cohesion establishment factor)-dependent way. Almost every host factor that was identified to be involved in plasmid partitioning affects the cohesion between sister plasmids supporting the fact that it is a rate limiting step in the partitioning pathway. Later, entrapment of sister plasmids by monomeric cohesin ring was demonstrated (Ghosh et al. 2009) similar to what has been shown for the sister chromatids (Ivanov and Nasmyth 2005, 2007; Haering et al. 2008), suggesting an evolutionary functional similarity between the plasmid and the chromosome segregation.
Fig. 2 Working model depicting the roles of different host factors in 2 micron plasmid segregation (Prajapati et al. 2017). As the cell enters into S phase, Rsc8 and Rsc58 associate with the plasmid and ‘Rep1–Rep2–Kip1′ sub-complex dissociates from the plasmid for a short-time window to reassociate (Ma et al. 2013). A ‘pre-partitioning complex’ (Hajra et al. 2006) is formed at STB through temporal association of Bik1, Bim1, and Kip1. This pre-partitioning complex (Hajra et al. 2006) with the aid of the microtubules may facilitate the movement of the plasmids to a location probably near to the centromere/SPB which helps in loading of the other host factors such as Cse4, Rsc2, and Mcd1 (cohesin) lead to the formation of a mature partitioning complex. Finally, cohesin cleavage after metaphase allows the plasmid clusters to segregate along with the chromosomes. During anaphase, the plasmids may move towards the minus end (towards SPB) with the help of minus-end-directed activity of Kip1 motor and segregate faithfully between the mother and the daughter cells (figure not drawn to scale)
-
2.
RSC2 (Restructure of chromatin remodeler) Remodeling of chromatin helps in the rearrangement of the nucleosomes along the chromatin that can influence certain DNA–protein interactions which can culminate into changes in the organization of the DNA locus with respect to protein binding and/or in gene expression (Cairns et al. 1996; Tsuchiya et al. 1998). The Rsc2 (a key subunit of the RSC2 complex) chromatin remodeler has been found to have a unique role in the subsistence of the 2 micron plasmid by maintaining a proper chromatin architecture at STB (Wong et al. 2002) which turned out to be crucial for the loading of Rep1 and cohesin (Yang et al. 2004). Recently, in search of novel host factors, Rep1 and Rep2 complexes were purified using Tandem Affinity Purification method and identification of the interactors showed other subunits of the RSC2 complex such as Rsc8, Rsc58, and Sth1 which further signifies the role of the RSC2 complex in plasmid segregation (Ma et al. 2013) (Fig. 2). These results reconfirmed the previously shown data that Sth1 associates to the STB and plays a significant role in the maintenance of the 2 micron plasmid by promoting an association between the plasmid and cohesin (Huang et al. 2004).
-
3.
Cse4 (The Histone 3 variant) Almost in every eukaryote, centromeric chromatin is marked by a variant form of histone H3, namely, CENP-A which is essential for kinetochore assembly and equal segregation of the chromosomes (Black et al. 2010; Mehta et al. 2010). In Saccharomyces cerevisiae, this variant form is called Cse4 (yeast homolog of CENP-A). There is a functional kinship between STB and centromere as partitioning loci for the plasmids and the chromosomes, respectively, where both appear to recruit supra-molecular complex to facilitate genome segregation. This led us to believe that STB might possess Cse4 just like centromere which is turned out to be true (Hajra et al. 2006). As a consequence of Cse4 incorporation, the STB chromatin also shows positive supercoiling similar to CEN chromatin which distinguishes these loci from bulk of the H3 containing chromatin (Furuyama and Henikoff 2009; Huang et al. 2011). However, unlike centromere, assembly of Cse4 at STB is believed to depend on the Rep proteins and once assembled, it further allows the RSC2 complex to functionally activate the STB chromatin for the acquisition of cohesin that eventually ensures faithful partitioning of the plasmid between mother and daughter (Hajra et al. 2006) (Fig. 2). This is in concordance with an earlier study, where RSC function has been implicated in the generation of cohesion (Huang et al. 2004)
-
4.
Microtubules It was noticed that in the absence of nuclear microtubules, the plasmids missegregate along with the chromosomes (Mehta et al. 2005). It was found that the acquisition of most of the above host factors by the plasmids, through their binding at STB, depends on the presence of intact microtubule (Mehta et al. 2005; Hajra et al. 2006; Prajapati et al. 2017). However, the binding of the plasmid-encoded Rep proteins at STB was not found hindered in the absence of the microtubules (Mehta et al. 2005). These data along with the observations that plasmids reside very close to the spindle pole body (SPB) or centromere (Velmurugan et al. 2000; Mehta et al. 2005) led to a belief that plasmids need to reach to a subnuclear locale through microtubule (perhaps close to the centromere) for acquiring the proteins involved in chromosome segregation (Mehta et al. 2005). Although the later results suggest that the subnuclear locale could be the telomeric regions at least in meiosis (Sau et al. 2014). Nevertheless, the requirement of the microtubule for the plasmid segregation led to a hypothesis that some adapter molecules might associate both with the plasmids and the microtubules to connect these two. The hypothesis turns out to be true as described below.
-
5.
Kip1 The fact that the plasmid utilizes microtubule, perhaps as a track, to reach to a subnuclear locale presumed that a molecular motor may be involved in the process. Since the plasmids were observed close to the SPB and centromere (Velmurugan et al. 2000; Mehta et al. 2005), it was believed that the subnuclear locale could be somewhere near the + end of the microtubules and a + end-directed motor function may be crucial to achieve this. It was observed that among the budding yeast motors, the + end-directed Kip1 of kinesin 5 family proteins, contributes significantly more to the maintenance of the 2 micron plasmid as compared to another motor of the same directionality and family, Cin8 (Cui et al. 2009). However, Cin8 has a more significant role to play than Kip1 in chromosome segregation, suggesting a distribution of the duties between these two proteins for plasmid and chromosome segregation, possibly a clever strategy optimized by the plasmids by not putting the burden on its host. The absence of the Kip1 motor protein disturbs the nuclear locale of the plasmid which further abates the loading of many host factors (Cui et al. 2009). A recent study showed a bi-directional movement (both towards minus and plus ends) of Kip1 motor (Fridman et al. 2013). It is possible that in the late anaphase, the plasmid clusters move towards minus end with the help of Kip1 and segregate along with chromosomes and SPBs (Fig. 2).
-
6.
Bik1 and Bim1 Since the functions of microtubules are largely regulated by both the motor and non-motor microtubule-associated proteins (MAPs), we wished to address if the observed interaction between the plasmid and the motor/microtubules, and hence, the efficiency of the plasmid partitioning is influenced by any MAP that has function at the + end of the microtubules. Bik1 and Bim1 (mammalian homolog of CLIP170 and EB1, respectively) are + end tracking proteins (+TIP) required for overall microtubule stabilization and proper nuclear division (Miller et al. 2006). From these functions at the + ends, we speculated that to attach to the microtubules, 2 micron plasmid might utilize Bik1 and Bim1 along with the Kip1 as an adapter. The speculation turned out to be true as we found that the missegregation of the 2 micron plasmid is substantially higher in the absence of both the proteins, compared to a centromeric plasmid (Prajapati et al. 2017). Interestingly, both Bik1 and Bim1 could bind to STB in the absence of the Rep proteins suggesting an early binding of both the proteins which was further supported by the observation that Bik1 precedes the Kip1 motor in its association to STB. The requirement of the cargo-binding domain (C-terminus) of Bik1 for binding to the plasmid and its stability suggests that the plasmids are utilized as cargos. Overall, our study identifies Bik1 and Bim1 as the missing link between the plasmids and the motor/microtubules. The model predicts that the plasmid acquires Bik1 and Bim1 to connect to the motor and the microtubules which further helps in relocating the plasmid to a defined address in the nucleus, where other factors associated with the plasmid (Fig. 2) present a cumulative model for equal segregation of plasmids (Prajapati et al. 2017), depicting the cell cycle-dependent association and dissociation of all the known major host factors identified until date in the process (Ma et al. 2013).
‘Kinetochore’ at CEN vs ‘partitioning complex’ at STB
Chromosome segregation relies on a very important nucleoprotein structure called ‘kinetochore’ which is made up of >100 proteins and centromeric DNA in budding yeast (Westermann et al. 2007; Biggins 2013; Yamagishi et al. 2014). The structure is required for the successful attachment of the chromosomes to the microtubules (McAinsh et al. 2003). The high stability of 2 micron plasmid (nearly similar to chromosome) and a requirement of the microtubules for this process suggest that a structure made of some kinetochore proteins may form at STB that may capture the microtubule. However, none of the representative kinetochore proteins from different subcomplexes (inner, middle, and outer) (Fig. 3) was detected at the STB in a previous study [unpublished but cited in (Mehta et al. 2005)], except Cse4 (Hajra et al. 2006) and (Fig. 3b). The process of recruitment of Cse4 to the centromere and to STB appears to be different as the centromere CBF3 complex is required, whereas the latter needs the Rep1 and Rep2 proteins (Malik 2006; Hajra et al. 2006). However, interdependent ChIP assays have revealed that, like kinetochore, the assembly of proteins at STB, called ‘partitioning complex’ (Hajra et al. 2006), occurs in an ordered fashion sensing a cell cycle cue (Ma et al. 2013). The point to be noted is that although there is an organizational kinship between the kinetochore and the partitioning complex, where a multiprotein complex attaches the DNA locus (CEN or STB, respectively) with the microtubules (Fig. 3), the strength of attachment appears to be much stronger for CEN over STB. This is because attempts to segregate the plasmid with two CENs can cause the DNA breakage but not with two or more STBs [(Mann and Davis 1983) and unpublished observation, discussed in Liu et al. (2014)]. Recently, it was observed in fission yeast that epigenetic alterations at the centromeric chromatin lead to an overall change in microtubule dynamics and kinetochore microtubule attachment (George and Walworth 2016) which further requires a proper chromosome segregation. Although such a study in budding yeast was focused only on kinetochore function (Choy et al. 2011), it would be interesting to investigate whether such epigenetic markings are also present at the STB chromatin. Interestingly, it has been proposed that a centromere might have evolved from STB when a free living 2 micron plasmid happened to accidentally integrate into the yeast genome which was followed by the acquisition of crucial new strategies required for a much more efficient segregation of the chromosomes (Malik 2006; Liu et al. 2013). How the host factors with a sub-stoichiometric concentration are distributed among multiple copies is an open question. However, it is presumed that either plasmids recycle these factors among the copies (Chan et al. 2013) or utilize these minimally by forming a cluster.
Organizational similarity between the protein complexes linking DNA loci, CEN or STB to the microtubules. a Different subcomplexes are organized in a hierarchical pattern to form a kinetochore structure to connect centromere to the microtubules. b Different proteins assemble in a temporal fashion to form a ‘partitioning complex’ to connect STB to the microtubules (figure not drawn to scale)
Models of plasmid segregation: chromosome hitch-hike or independent?
Out of the two models proposed (Mehta et al. 2002) for the plasmid segregation, the experimental evidence is increasingly accumulating to support the chromosome hitch-hike model, where the plasmid clusters hitch-hike/tether with the host chromosomes and segregate in a chromosome-dependent manner (Fig. 4a). However, the second model in which the plasmid-cluster segregation is independent of the chromosomes (Fig. 4b) could not be ruled out completely (discussed below). In support of the hitch-hike model, it has been demonstrated that the 2 micron plasmid cosegregated with the sister chromatids when they were forced to segregate either to the mother or to the daughter cell (Liu et al. 2013). The strategy of partitioning of an extra-chromosomal genome through tethering to the host chromosome has been well adapted for the segregation of the viral episomes (Kanda et al. 2007; You et al. 2004). In these systems, EBNA1 from Epstein–Bar virus (EBV) and E2 from Bovine papilloma virus (BPV) are viral-encoded partitioning proteins (functionally similar to Rep proteins) which bind to the iterons (virus-borne partitioning locus). hEBP2 (for EBV) and Brd4 (for BPV) are host-encoded proteins which help in a symmetric association of viral episomes to the host chromatin (Kapoor et al. 2005; McPhillips et al. 2005). Such a tethering strategy assumes that the plasmids may associate to certain specific site(s) on the chromosomes. In concordance with this, it has been shown that the plasmid may associate with telomeres, at least during meiosis, as the plasmids were found to follow the dynamics similar to the telomeres, perhaps utilizing the nuclear envelope-associated motors Rap1 or Mps3 (Sau et al. 2014). However, our observation with different fluorescently marked centromere, telomere, and plasmid in the same cell suggests that in mitosis, the plasmids may not always associate with the telomeres (Prajapati and Ghosh, unpublished observations).
Models of plasmid segregation. a 1 Plasmid clusters are tethered to the chromosome through Rep–STB system. 2 The plasmid clusters are cohesed together by cohesin in a fashion similar to the sister chromatids during the replication. 3 The plasmids, therefore, hitch-hike the chromosomes and following cohesin cleavage move to the daughter cells accompanied by the chromosomes. b 1 Plasmid clusters do not attach to the chromosome. The cohesin assembly occurs in the S phase separately on both plasmids and chromosomes. 3 Cohesin cleavage separates both sister plasmids and sister chromosomes, but plasmid clusters do not hitch-hike the chromosomes and move to the daughter cells independently perhaps guided by the spindle
Besides hitchhiking the chromosome, it has been proposed that the plasmids may also tether to other cellular structure such as the nuclear matrix (Wu et al. 1987) or the nuclear membrane for its partitioning (Khmelinskii et al. 2011). However, the latter possibility has been ruled out using 2 micron-derived plasmid in contrast to an ARS-derived plasmid (Liu et al. 2016).
The question of chromosome-independent segregation (Fig. 4b) of the plasmids remains elusive, but a successful uncoupling of plasmid and chromosome segregation in few instances could indeed be achieved. Following are some of the cases as described previously (Ma et al. 2013): (1) microtubule restoration in G2/M after depolymerization in the G1 phase of the cell cycle does not restore the equal plasmid segregation, but chromosomes segregate faithfully following the restoration. (2) Loading of Cse4 at STB is independent to CBF3 complex (Ndc10), whereas the loading is disrupted at CEN in ndc10-1 mutant (Hajra et al. 2006). (3) The absence of Kip1 (Cui et al. 2009) or Rsc2 (Wong et al. 2002) or Bim1 (Prajapati et al. 2017) differentially affects plasmid segregation compared to the chromosome. This is probably due to the presence of redundant proteins Cin8, Rsc1, and Bik1 for Kip1, Rsc2, and Bim1, respectively. However, the absence of Bik1 affects the stability of the 2 micron plasmid along with the chromosome indicating bi-functionality of this protein. These evidences in favor of a ‘chromosome independent’ mode of segregation suggest that both the mechanisms (Mehta et al. 2002) may be utilized by 2 micron plasmid segregation and the two mechanisms may not be mutually exclusive to each other.
The amplification system
Despite all the elegance, the partitioning system fails occasionally. Under such rare situations, the plasmids missegregate and daughter cells receive a reduced number of plasmids due to mother bias. The 2 micron plasmid has an efficient PCN amplification system to restore the steady-state copy number under such rare events (Futcher 1986; Volkert and Broach 1986; Murray et al. 1987). This amplification system consists of a site-specific recombinase Flp encoded by the plasmid-borne FLP gene and two FRT sites that divide a plasmid into two unique regions (Andrews et al. 1985; Senecoff et al. 1985; McLeod et al. 1986) (Fig. 1).
Flp recombinase
The 2 micron plasmid-encoded protein Flp is a conservative site-specific recombinase of the integrase family (Sadowski 1995) with a turnover number of ~0.12 min−1 per Flp monomer (Gates and Cox 1988). The frequent inversion of one unique region of the 2 micron plasmid with respect to the other (Beggs 1978) that generates A and B forms of the plasmid (Fig. 1) is predominantly Flp-dependent (Gerbaud et al. 1979; Broach and Hickst 1980). However, site specific but Flp independent mitotic and meiotic recombination has also been observed at a very low frequency (Broach et al. 1982). Apart from flipping one half of the plasmid with respect to the other, the Flp-mediated recombination is also believed to be responsible for the inter-conversion of the plasmid replication between the theta and the rolling circle modes of replication (Futcher 1986). Flp cleaves the FRT sites (discussed in the next section) at the ends of an 8 bp core DNA sequence and remains covalently attached to the 3′-phosphate via Tyr343 present within the active site (Gronostajskil and Sadowski 1985; Andrews et al. 1985; Senecoff et al. 1985; Amin and Sadowski 1989; Pan et al. 1993) in a fashion analogous to the Int and Cre recombinases. Mutations Y343S and Y343F of Flp can abolish the recombinase function without affecting the DNA-binding function of the Flp suggesting that the Tyr343 is the key residue for Flp’s catalytic function and has minimal or perhaps no role in the DNA binding function (Prasad et al. 1987). The slow catalytic activity presumably enables a fine-tuned regulation of the recombination process under a narrow window of time (the time between the replication of the two FRTs) to change the orientation of the replication forks during plasmid amplification (discussed below). In vivo, the Flp activity appears to be regulated by post-translational modification of Flp (discussed later, Xiong et al. 2009).
Flp–FRT recombination system
The Flp–FRT recombination is central to the amplification system. The structure of minimal FRT region which is relevant to the Flp-mediated recombination is a 34 bp core sequence consisting of two 13 bp Flp-binding elements 1a and 1′a flanking an 8 bp spacer region at which the actual recombination occurs through a single-strand cleavage (Senecoff et al. 1985). In addition to the flanking repeats 1a and 1′a, is the sequence 1′b identical to 1′a, which is in the same orientation (Fig. 5a). However, the recombination occurs within the ‘minimal FRT’ without 1′b sequence. The asymmetry within the core region specifies the directionality for the recognition and cleavage at the FRTs (McLeod et al. 1986). Efficient recombination at the FRTs requires a homology between the core regions of the partner strands (Andrews et al. 1985), although non-homologous partners can also undergo recombination with reduced efficiency (Andrews et al. 1986). Studies on the topology of the plasmids bearing two FRTs have shown that the reduced-recombination efficiency observed between the FRTs with non-homologous cores is due to the unstable heteroduplex which readily recombines to form a non-recombinant homo-duplex (Azam et al. 1997; Ma et al. 2014). The Flp–FRT recombination system is, in general, in agreement with other tyrosine family recombinases such as Int, Cre, and XerC/XerD, but differs in the assembly of its active site. In contrast to other recombinases, where a monomer is assembled at the active site in cis, the Flp recombinase active site involves the assembly of a dimer in trans (Fig. 5b).

Adopted from (Futcher 1986)
Flp–FRT recombination. a Organization of FRT site. There are three regions of inverted repeats, namely, 1a, 1′a, and 1′b. The arrows show their relative orientation at the top. The 34 bp sequence consists of two inverted repeats 1a and 1′a with an 8 bp spacer sequence in between, where recombination occurs. b Assembly of Flp dimers at the active site and ensuing geometrical changes during Flp-mediated recombination. Two FRTs in opposite orientation each bound by two Flp monomers are brought together. Flp monomers are assembled at the 1a and 1′a regions of the FRT sites and strand cleavage occurs at the junction of spacer region and the flanking inverted repeats. One active site is assembled at the identical ends of each DNA fragment. Strands are cleaved and exchanged to form a Holliday intermediate, H1. Second round of cleavage and strand exchange occurs at the other identical ends of the fragment pair in the H2 Holliday structure to resolve the strands. c Plasmid before the replication starts with black blocks showing the origin and FRT sites, I and II. Replication starts at the origin replicating one FRT site before the other producing three FRT sites, III and IV. Intramolecular recombination between two FRT sites converts the θ structure to the rolling circle structure, where the replication forks chase each other, V. Replication proceeds forming multimeric plasmid molecule, VI and VII. A second recombination at the FRT site reconverts the rolling circle again to θ mode, VIII. The two replication forks converge to give one monomeric plasmid and another multimeric plasmid molecule, IX which undergoes further recombination to resolve into monomeric plasmid molecules, X
Futcher’s model of Flp-mediated amplification
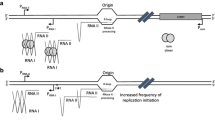
The amplification system becomes activated to restore the steady-state PCN in the event of missegregation and consequent perturbations in the PCN. It has been demonstrated that this activation depends on Flp-mediated recombination at the FRT sites (Futcher 1986; Volkert and Broach 1986). Futcher (1986) proposed a model for the Flp-mediated plasmid amplification which depends on two structural features of the 2 micron plasmid—(1) Flp–FRT recombination system and (2) Asymmetric location of 2 micron origin of replication with respect to the two FRT sites.
The model accommodating these two features is described as follows (Fig. 5c). Under a steady-state condition, the plasmids replicate by theta mode of replication, thus duplicating the plasmid number during the S phase. In theta mode, the two replication forks move away from each other. While the plasmid replicates, the FRT closer to the Ori replicates early, and the one farther replicates late. Consequently, at a certain point of time, there are three FRTs present within the same molecule, and a recombination between one of the two replicated FRTs, and the yet non-replicated FRT can reverse the orientation of the replication forks. This leads to a rolling circle mode of replication, where one replication fork chases the other, thus spooling out a long concatemer made of several plasmids connected at their ends. The concatemer may later be resolved into single plasmid molecules by the Flp mediated or by homologous recombination or can be trapped in the form of replication intermediates (RIs) of different possible configurations. This way a single monomer can generate multiple monomers within one S phase causing PCN amplification.
However, electron micrograph analysis of the RIs of the replicating plasmids, which are topologically interlinked, suggests a mechanism which differs from that expected from the above-described Futcher’s model (Petes and Williamson 1994). Instead of interlinked θ structures (as expected if the amplification follows Futcher’s model), the micrograph shows a structure in which two circular forms are connected by a linear stretch of DNA appearing like a Pince-nez, hence the name PN. Although this structure is different from what can be expected from the Futcher’s model, it does not contradict the Flp–FRT recombination paradigm (Petes and Williamson 1994).
The Futcher’s model assumes that the recombination event, upon activation of Flp, has to be timed precisely to convert the θ mode of replication to the rolling circle mode which is the key to turn ON the PCN amplification system. It also believes that Flp activation occurs only when there is a drop in PCN. Moreover, the intramolecular recombination might not be limited to the re-orientation of the replication forks alone, since different possible structural variants of the plasmid (Petes and Williamson 1975, 1994) might coexist leading to a complex set of outcomes. Once the S phase is over, the partially replicated portions will remain as such and may get pinched off during successive recombination events leading to the resolution of the plasmids into monomers and the removal of partially replicated plasmids.
Crosstalk between amplification and partitioning systems: a mechanism for plasmid maintenance
From the above discussions, it is apparent that the amplification and the partitioning systems together ensure stable maintenance of the 2 micron plasmid in the cell. Presumably, when the partitioning system fails and PCN drops to a sub steady-state level, the amplification system is turned on. The system is turned off when the average PCN is restored. This indicates that there should be a mechanism of direct readout of the PCN inside the cell which allows the amplification system to be turned on or off and that mechanism must be intricately linked to the partitioning system.
It is intriguing to address how a steady-state PCN is maintained through a probable crosstalk between the partitioning and the amplification systems. From earlier and our studies, we believe that Rep proteins and the Raf1 protein might be involved in the crosstalk as discussed below. From the transcriptional assays, it has been proposed that Rep1–Rep2 complex forms a bipartite repressor and attenuates Flp activity (Som et al. 1988). The in vitro and in vivo interactions of the Rep1 and Rep2 proteins have been demonstrated by co-IP (Scott-Drew and Murray 1998) and by yeast two-hybrid assay (Sengupta et al. 2001), respectively. Raf1, on the other hand, acts as a positive regulator of Flp which has been demonstrated by an increased level of FLP transcripts when Raf1 is over-expressed from an inducible GAL1 promoter. Furthermore, Raf1 is believed to antagonize the Rep1–Rep2 repressor complex and enhances Flp activity (Reynolds et al. 1987; Som et al. 1988). The functions of the Rep1–Rep2 complex along with Raf1 have been summarized in the context of their probable roles in the crosstalk between the partitioning and the amplification systems in the following sections.
The Rep1–Rep2 complex
The salient features of the complex are: it primarily binds to STB proximal as a part of the partitioning system and facilitates equal partition of the plasmids presumably by recruiting several host factors (Jayaram et al. 1983; Hadfield et al. 1995). It is believed that with a relatively decreased affinity, it also binds to a sequence upstream of FLP (probably at the promoter region) to reduce the level of FLP transcription as PCN increase (Som et al. 1988). It also binds to a region terminal to RAF1 ORF. The region is a part of STB distal and is believed to act as a transcriptional ‘silencer’ to block any runaway transcript from RAF1 ORF to arrive into STB proximal (Papacs et al. 2004). The Rep1–Rep2 complex also has an auto-regulatory function by controlling the transcription of REP1, perhaps through its interaction with the REP1 promoter region.
Among the four plasmid-encoded proteins, the functions of Rep1, Rep2, and Flp have been extensively studied in the earlier studies (Futcher and Cox 1983; Jayaram et al. 1983; Reynolds et al. 1987; Gates and Cox 1988). However, the function of Raf1 was not clearly understood from the earlier studies (Cashmore et al. 1988; Xiao et al. 1991; Hadfield et al. 1995). Very recently, we have taken up a study to reveal the function of Raf1; the outcome of which is discussed below.
Raf1
The belief that Raf1 enhances FLP expression came from the experiment, where Raf1 was overproduced in a cell where, consequently, an increased transcription of FLP, hence a higher PCN, was observed (Murray et al. 1987). However, the involved mechanism was not clear. On the other hand, Raf1 was shown to bind to STB, but whether this binding depends on the Rep proteins was not clear as contradicting results were obtained from the in vivo and in vitro studies (Cashmore et al. 1988; Hadfield et al. 1995; Velmurugan et al. 1998). In our study, we wished to reinvestigate Raf1 function by generating raf1 null cells and found a decrease in the plasmid stability but an increase in PCN (Rizvi et al. 2017). To address how Raf1 can affect both the partitioning and the amplification systems, we found that Raf1 interacts with both Rep1 and Rep2 independently and blocks their interaction thus reducing the cellular concentration of the Rep1–Rep2 complex that acts a repressor of REP1, FLP, and RAF1 genes. This blockage resulted in reduced plasmid stability and increased PCN. Interestingly, both the deletion and overexpression of Raf1 had a similar effect on the plasmid stability and copy number. This could be due to the two feedback loops connected together through the Rep1–Rep2 repressor complex (Fig. 6a). Rep1 (X) and Rep1–Rep2 repressor complex (Y) are connected to each other in a negative feedback loop. This loop is further connected to Raf1 (Z), as anti-repressor through a positive feedback loop (Fig. 6c). This arrangement is analogous to a repressor amplified negative feedback loop discussed elsewhere (Novák and Tyson 2008). This control is different from a simple negative feedback (Fig. 6b) due to the presence of a positive feedback by an anti-repressor that pushes the concentration of the repressor away from the equilibrium, thus increasing the concentration of the repressor. This also causes the repressor concentration to oscillate (Ananthasubramaniam and Herzel 2014). When Raf1 is deleted, the positive feedback loop is removed and the repressor concentration decreases, resulting in an increased PCN and decreased plasmid stability (Fig. 6d). Overexpression of Raf1, on the other hand, directly blocks the interaction of Rep1 and Rep2 independent of the control circuit, thus producing the same but more severe effect as compared to the deletion of RAF1.
The 2 micron plasmid transcription control network. a Rep1, Rep1–Rep2 repressor complex, and Raf1 are connected through two feedback loops. Loop 1 is a negative feedback loop and loop 2 is a positive feedback loop. b In the absence of Raf1, the control network reduces to a single negative feedback loop with the repressor Y represses its own expression. c Wild-type control network is analogous to a repressor amplified negative feedback loop, where the repressor Y, represses the expression of X and is antagonized by an anti-repressor Z. In a repressor amplified negative feedback loop, the concentration of the repressor Y is pushed away from the equilibrium, thus increasing the average concentration of the repressor. d In the absence of Raf1, the positive feedback is removed resulting in a reduced average cellular concentration of the Rep1–Rep2 repressor complex that results in a decreased plasmid stability but an increased PCN
The host–plasmid relationship
Until date, the plasmid is not known to cause any advantage to its host, the yeast cell. On the other hand, the plasmid selfishly utilizes the host resources for its own faithful segregation. It is, therefore, surprising from an evolutionary perspective that how this host–plasmid relationship is maintained, where the selfish plasmid is highly ubiquitous and stable. Efforts have been made to assess this host–parasite relationship using competition experiments, where isogenic [Cir+] and [Cir0] cells were allowed to compete in the same flask (Futcher and Cox 1983; Mead et al. 1986). The results showed that at a steady-state PCN, a [Cir0] cell has 1–1.4% advantage over a [Cir+] cell. Competition experiments in a fed-batch culture condition with diploid and haploid strains demonstrated a greater PCN and stability of the 2 micron plasmids in diploids suggesting that higher ploidy of the host helps in the maintenance of the 2 micron plasmid (Mead et al. 1986). Although these early experiments were successful in establishing a parasitic nature of the plasmid, the growth advantage of the [Cir0] strain over the [Cir+] strain was marginal. However, to assess whether there is indeed any difference between the [Cir+] and [Cir0] cells, the experimental conditions demand a more sophisticated treatment using modern techniques of monitoring growth parameters at single cell level, such as spatial light interference microscopy (Mir et al. 2011). Such single cell techniques may be helpful in resolving parametric differences, if any, which get masked during population-level studies done in culture flasks.
Other factors in plasmid maintenance
Recently, another layer of control on the plasmid maintenance has been revealed which occurs through post-translational modification of the plasmid-encoded proteins. It has been shown that SUMOylation of the plasmid-encoded proteins is also involved both in the stability (Dobson et al. 2005; Pinder et al. 2013) and PCN control (Chen et al. 2005). SMT3 (Saccharomyces genome database: YDR510 W) is the only SUMO present in budding yeast. Like all the other members of the SUMO family, it covalently attaches to the lysine residue of its target. SUMO modification is involved in diverse cellular processes such as regulation of chromatid cohesion, chromosome segregation, DNA replication, and septin dynamics (Johnson and Blobel 1999). It has been demonstrated that SUMO modification of Flp has a significant role in maintaining a steady-state PCN within the host (Chen et al. 2005). Study of the cells showing typical nib phenotype [irregularly shaped colonies, cold sensitivity, and cell cycle delay (Holm 1982)], has demonstrated that the SUMOylation of Flp is required to restrict the over-replication of the 2 micron plasmid and hence to keep the PCN under check at a steady state (Chen et al. 2005; Xiong et al. 2009). The nib phenotype is now revealed and occurs due to a very high-copy number of the plasmid (more than 100) that imposes an extra burden on the cells. This high-copy number is attained, because the cells are unable to SUMOylate Flp and thereby restrict its activity. Deletion of ULP1 gene that codes for the Ulp1 protease responsible for the removal of Smt3 from its substrate is lethal for the cells irrespective of the presence of the 2 micron plasmid. However, strains bearing partially active Ulp1 are sensitive to the presence of the 2 micron plasmid. Deletion of ULP2, another SUMO protease (Li and Hochstrasser 2000), or simultaneous deletion of MLP1 and MLP2 [coding myosin-like proteins required for the stabilization and localization of Ulp1 to the nuclear pore (Zhao et al. 2004)] also results in a 2 micron-associated nib phenotype (Dobson et al. 2005). Rep1 and Rep2 have also been demonstrated to be SUMOylated (Pinder et al. 2013). Several lysine-to-arginine mutations in Rep1 and Rep2 completely abolish SUMOylation which leads to plasmid loss and missegregation (Pinder et al. 2013). Together, these results show that defect in the SUMOylation of the Rep1, Rep2, and, Flp proteins directly affect the plasmid stability and copy number.
Conclusion
Since its probable invasion within the yeast cell, the 2 micron plasmid has evolved into a highly optimized molecular parasite so much so that its host, S. cerevisiae is appeared to have naturalized the plasmid as a part of its own genome and is not perturbed by its presence unless the PCN increases beyond a sustainable limit. The amplification system encoded by the plasmid thus ensures not only just a high PCN but also a feedback control preventing any runaway increase in PCN beyond this sustainable limit. This requirement to maintain a steady-state level of PCN is intricately linked to its mechanism of partitioning that utilizes several key host factors that become limiting if the PCN becomes too high. These factors seem to be primarily dedicated to hitch-hike the plasmid to the host chromosomes, which can later be equally distributed between the mother and the daughter cells. Besides the host factors, how only four proteins using a network of transcriptional control amongst them can form a minimal, but a highly robust system of stable genome segregation is itself a classic example of ‘simplicity is the ultimate sophistication’. In the future, analysis of the large transcripts encoded by the plasmid may reveal another layer of control on the expression of the four proteins. In summary, the 2 micron plasmid has not only been successful in tailoring its genetic makeup to code a highly sophisticated control mechanism for its maintenance but has also developed means to recruit host-segregation machinery to overcome the problem of missegregation imposed by the mother bias. Future work using more advanced tools for growth analysis along with challenging cells with [Cir+] or without [Cir0] 2 micron plasmids under various stress conditions may reveal the significance of adoption of these plasmids by yeast as a host.
Importantly, it has been found that the chromosome-tethering mechanism is also acquired by several mammalian viruses (during their episomal form) such as EBV, BPV, and HPV (human papilloma virus) (Bastien and McBride 2000; Kapoor and Frappier 2003), presumably to survive exclusion during the open mitotic segregation. However, this reasoning fails in case of closed mitosis observed in S. cerevisiae. Furthermore, the plasmid segregation is not coupled to the nuclear pore complex either (Liu et al. 2016), which was earlier thought to be responsible for the segregation of the plasmid (Wu et al. 1987). The chromosome-tethering mechanism may, therefore, be reminiscent of its viral origin which got selected to overcome the mother bias in the absence of its ability to attach to the nuclear membrane or membrane-associated structures. Interestingly, as a matter of similitude to the 2 micron plasmids, the segregation of the viral episomes is found to be non-random and cohesin dependent (Holdorf et al. 2011; Mehta et al. 2015). Future work on 2 micron plasmid may reveal a hitherto unidentified mechanism of survival of viral episomes within the mammalian host. Finally, based on the recent evidence for the association of Rep proteins to the mammalian chromosomes (Liu et al. 2016), the highly stable genetically engineered 2 micron-derived plasmid for the mammalian system may be generated, which could be utilized for developing a new generation of mammalian vectors for high expression of proteins.
Abbreviations
- PCN:
-
Plasmid copy number
- SPB:
-
Spindle pole body
- TAP:
-
Tandem affinity purification
References
Ahn YT, Wu XL, Biswal S et al (1997) The 2 micron-plasmid-encoded Rep1 and Rep2 proteins interact with each other and colocalize to the nucleus. J Bacteriol 179:7479–7506
Amin AA, Sadowski PD (1989) Synthesis of an enzymatically active FLP recombinase in vitro: search for a DNA-binding domain. Mol Cell Biol 9:1987–1995. doi:10.1128/MCB.9.5.1987.Updated
Ananthasubramaniam B, Herzel H (2014) Positive feedback promotes oscillations in negative feedback loops. PLoS One. doi:10.1371/journal.pone.0104761
Andrews BJ, Proteau GA, Beatty LG, Sadowski PD (1985) The FLP recombinase of the 2µ circle DNA of yeast: interaction with its target sequences. Cell 40:795–803
Andrews BJ, McLeod M, Broach J, Sadowski PD (1986) Interaction of the FLP recombinase of the Saccharomyces cerevisiae 2 µm plasmid with mutated target sequences. Mol Cell Biol 6:2482–2489
Azam N, Dixon JE, Paul D (1997) Topological analysis of the role of homology in Flp-mediated recombination. J Biol Chem 272:8731–8738
Bastien N, McBride AA (2000) Interaction of the papillomavirus E2 protein with mitotic chromosomes. Virology 270:124–134. doi:10.1006/viro.2000.0265
Beggs JD (1978) Transformation of yeast by a replicating hybrid plasmid. Nature 275:104–109. doi:10.1038/275104a0
Biggins S (2013) The composition, functions, and regulation of the budding yeast kinetochore. Genetics 194:817–846. doi:10.1534/genetics.112.145276
Black BE, Jansen LET, Foltz DR, Cleveland DW (2010) Centromere identity, function, and epigenetic propagation across cell divisions. Cold Spring Harb Symp Quant Biol 75:403–418. doi:10.1101/sqb.2010.75.038
Boach JR (1983) Construction of high copy yeast vectors using 2-µm circle sequences. Methods Enzymol 101:307–325
Broach JR (1982) The yeast plasmid 2µ circle. Cell 28:203–204
Broach JR, Hickst JB (1980) Replication and recombination functions associated with the yeast plasmid, 2µ circle. Cell 21:501–508
Broach JR, Atkins JF, Mcgill C, Chow L (1979) Identification and mapping of the transcriptional and translational products of the yeast plasmid 2µ circle. Cell 16:827–839
Broach JR, Guarascio VR, Jayaram M (1982) Recombination within the yeast plasmid 2µ circle is site-specific. Cell 29:227–234. doi:10.1016/0092-8674(82)90107-6
Cairns BR, Lorch Y, Li Y et al (1996) RSC, an essential, abundant chromatin-remodeling complex. Cell 87:1249–1260. doi:10.1016/S0092-8674(00)81820-6
Cashmore AM, Hadfield C, Albury MS, Meacock PA (1986) Genetic analysis of partitioning functions encoded by the 2 µm circle of Saccharomyces cerevisiae. Mol Gen Genet 203:154–162
Cashmore AM, Albury MS, Hadfield C, Meacock PA (1988) The 2 µm D region plays a role in yeast plasmid maintenance. Mol Gen Genet 212:426–431
Chan K-M, Liu Y-T, Ma C-H et al (2013) The 2 micron plasmid of Saccharomyces cerevisiae: a miniaturized selfish genome with optimized functional competence. Plasmid 70:2–17. doi:10.1016/j.plasmid.2013.03.001
Chen XL, Reindle A, Johnson ES (2005) Misregulation of 2 µm circle copy number in a SUMO pathway mutant. Mol Cell Biol 25:4311–4320. doi:10.1128/MCB.25.10.4311
Choy JS, Acuña R, Au WC, Basrai MA (2011) A role for histone H4K16 hypoacetylation in Saccharomyces cerevisiae kinetochore function. Genetics 189:11–21. doi:10.1534/genetics.111.130781
Clark-Walker GD, Miklos GL (1974) Localization and quantification of circular DNA in yeast. Eur J Biochem 41:359–365
Cui H, Ghosh SK, Jayaram M (2009) The selfish yeast plasmid uses the nuclear motor Kip1p but not Cin8p for its localization and equal segregation. J Cell Biol 185:251–264. doi:10.1083/jcb.200810130
Dobson MJ, Pickett AJ, Velmurugan S et al (2005) The 2 µm plasmid causes cell death in Saccharomyces cerevisiae with a mutation in Ulp1 protease. Mol Cell Biol 25:4299–4310. doi:10.1128/MCB.25.10.4299
Fridman V, Gerson-Gurwitz A, Shapira O et al (2013) Kinesin-5 Kip1 is a bi-directional motor that stabilizes microtubules and tracks their plus-ends in vivo. J Cell Sci 126:4147–4159. doi:10.1242/jcs.125153
Furuyama T, Henikoff S (2009) Centromeric nucleosomes induce positive DNA supercoils. Cell 138:104–113. doi:10.1016/j.cell.2009.04.049
Futcher AB (1986) Copy number amplification of the 2 micron circle plasmid of Saccharomyces cerevisiae. J Theor Biol 119:197–204
Futcher AB, Cox BS (1983) Maintenance of the 2 microns circle plasmid in populations of Saccharomyces cerevisiae. J Bacteriol 154:612–622
Gates CA, Cox MM (1988) FLP recombinase is an enzyme. Proc Natl Acad Sci USA 85:4628–4632
Gehlen LR, Nagai S, Shimada K et al (2011) Nuclear geometry and rapid mitosis ensure asymmetric episome segregation in yeast. Curr Biol 21:25–33. doi:10.1016/j.cub.2010.12.016
George AA, Walworth NC (2016) Microtubule dynamics decoded by the epigenetic state of centromeric chromatin. Curr Genet 62:691–695. doi:10.1007/s00294-016-0588-0
Gerbaud C, Fournier P, Blanc H et al (1979) High frequency of yeast transformation by plasmids carrying part or entire 2-micron yeast plasmid. Gene 5:233–253
Ghosh SK, Hajra S, Jayaram M (2007) Faithful segregation of the multicopy yeast plasmid through cohesin-mediated recognition of sisters. Proc Natl Acad Sci USA 104:13034–13039. doi:10.1073/pnas.0702996104
Ghosh SK, Huang CC, Hajra S, Jayaram M (2009) Yeast cohesin complex embraces 2 micron plasmid sisters in a tri-linked catenane complex. Nucleic Acids Res 38:570–584. doi:10.1093/nar/gkp993
Gronostajskil RM, Sadowski PD (1985) The FLP recombinase of the Saccharomyces cerevisiae 2 µm plasmid attaches covalently to DNA via phosphotyrosyl linkage. Microbiology 5:3274–3279
Hadfield C, Mount RC, Cashmore AM (1995) Protein binding interactions at the STB locus of the yeast 2 µm plasmid. Nucleic Acids Res 23:995–1002
Haering CH, Farcas A-M, Arumugam P et al (2008) The cohesin ring concatenates sister DNA molecules. Nature 454:297–301. doi:10.1038/nature07098
Hajra S, Ghosh SK, Jayaram M (2006) The centromere-specific histone variant Cse4p (CENP-A) is essential for functional chromatin architecture at the yeast 2-micron circle partitioning locus and promotes equal plasmid segregation. J Cell Biol 174:779–790. doi:10.1083/jcb.200603042
Hartley JL, Donelson JE (1980) Nucleotide sequence of the yeast plasmid. Nature 286:860–865
Holdorf MM, Cooper SB, Yamamoto KR, Miranda JJL (2011) Occupancy of chromatin organizers in the Epstein-Barr virus genome. Virology 415:1–5. doi:10.1016/j.virol.2011.04.004
Holm C (1982) Clonal lethality caused by the yeast plasmid 2μ DNA. Cell 29:585–594. doi:10.1016/0092-8674(82)90174-X
Huang J, Hsu JM, Laurent BC (2004) The RSC nucleosome-remodeling complex is required for cohesin’s association with chromosome arms. Mol Cell 13:739–750. doi:10.1016/S1097-2765(04)00103-0
Huang C-C, Hajra S, Ghosh SK, Jayaram M (2011) Cse4 (CenH3) association with the Saccharomyces cerevisiae plasmid partitioning locus in its native and chromosomally integrated states: implications in centromere evolution. Mol Cell Biol 31:1030–1040. doi:10.1128/MCB.01191-10
Ivanov D, Nasmyth K (2005) A topological interaction between cohesin rings and a circular minichromosome. Cell 122:849–860. doi:10.1016/j.cell.2005.07.018
Ivanov D, Nasmyth K (2007) A physical assay for sister chromatid cohesion in vitro. Mol Cell 27:300–310. doi:10.1016/j.molcel.2007.07.002
Jayaram M, Li YY, Broach JR (1983) The yeast plasmid 2µ circle encodes components required for its high copy propagation. Cell 34:95–104
Johnson ES, Blobel G (1999) Cell cycle-regulated attachment of the ubiquitin-related protein SUMO to the yeast septins. J Cell Biol 147:981–994
Kanda T, Kamiya M, Maruo S, Iwakiri D, Takada K (2007) Symmetrical localization of extrachromosomally replicating viral genomes on sister chromatids. J cell sci 120:1529–1539
Kapoor P, Frappier L (2003) EBNA1 partitions Epstein-Barr virus plasmids in yeast cells by attaching to human EBNA1-binding protein 2 on mitotic chromosomes. J Virol 77:6946–6956. doi:10.1128/JVI.77.12.6946-6956.2003
Kapoor P, Lavoie BD, Frappier L (2005) EBP2 plays a key role in Epstein-Barr virus mitotic segregation and is regulated by aurora family kinases. Mol Cell Biol 25:4934–4945
Khmelinskii A, Meurer M, Knop M, Schiebel E (2011) Artificial tethering to nuclear pores promotes partitioning of extrachromosomal DNA during yeast asymmetric cell division. Curr Biol 21:R17–R18
Kikuchi Y (1983) Yeast plasmid requires a cis-acting locus and two plasmid proteins for its stable maintenance. Cell 35:487–493
Li SJ, Hochstrasser M (2000) The yeast ULP2 (SMT4) gene encodes a novel protease specific for the ubiquitin-like Smt3 protein. Mol Cell Biol 20:2367–2377. doi:10.1128/MCB.20.7.2367-2377.2000
Liu YT, Ma CH, Jayaram M (2013) Co-segregation of yeast plasmid sisters under monopolin-directed mitosis suggests association of plasmid sisters with sister chromatids. Nucleic Acids Res 41:4144–4158. doi:10.1093/nar/gkt096
Liu Y-T, Fan H-F, Kachroo AH, Jayaram M, Rowley PA, Sau S, Chang K-M, Ma C-H (2014) The partitioning and copy number control systems of the selfish yeast plasmid: an optimized molecular design for stable persistence in host cells. Microbiol Spectr 2(5). doi:10.1128/microbiolspec.PLAS-0003-2013
Liu Y-T, Chang K-M, Ma C-H, Jayaram M (2016) Replication-dependent and independent mechanisms for the chromosome-coupled persistence of a selfish genome. Nucleic Acids Res. doi:10.1093/nar/gkw694
Ma C-H, Cui H, Hajra S et al (2013) Temporal sequence and cell cycle cues in the assembly of host factors at the yeast 2 micron plasmid partitioning locus. Nucleic Acids Res 41:2340–2353. doi:10.1093/nar/gks1338
Ma C-H, Liu Y-T, Savva CG et al (2014) Organization of DNA partners and strand exchange mechanisms during Flp Site-specific recombination analyzed by difference topology, single molecule FRET and single molecule TPM. J Mol Biol 426:793–815. doi:10.1016/j.jmb.2013.11.017
Malik HS (2006) A hitchhiker’s guide to survival finally makes CENs. J Cell Biol 174:747–749. doi:10.1083/jcb.200608107
Mann C, Davis RW (1983) Instability of dicentric plasmids in yeast. Proc Natl Acad Sci USA 80:228–232
McAinsh AD, Tytell JD, Sorger PK (2003) Structure, function, and regulation of budding yeast kinetochores. Annu Rev Cell Dev Biol 19:519–539. doi:10.1146/annurev.cellbio.19.111301.155607
McLeod M, Craft S, Broach JR (1986) Identification of the crossover site during FLP-mediated recombination in the Saccharomyces cerevisiae plasmid 2 µm circle. Mol Cell Biol 6:3357–3367
McPhillips MG, Ozato K, McBride AA (2005) Interaction of bovine papillomavirus E2 protein with Brd4 stabilizes its association with chromatin. J virol 79:8920–8932
Mead DJ, Gardner DC, Oliver SG (1986) The yeast 2 µ plasmid: strategies for the survival of a selfish DNA. Mol Gen Genet 205:417–421
Mehta S, Yang XM, Chan CS et al (2002) The 2 micron plasmid purloins the yeast cohesin complex: a mechanism for coupling plasmid partitioning and chromosome segregation? J Cell Biol 158:625–637. doi:10.1083/jcb.200204136
Mehta S, Yang X, Jayaram M, Velmurugan S (2005) A novel role for the mitotic spindle during DNA segregation in yeast: promoting 2 µm plasmid-cohesin association. Mol Cell Biol 25:4283–4298. doi:10.1128/MCB.25.10.4283
Mehta GD, Agarwal MP, Ghosh SK (2010) Centromere identity: a challenge to be faced. Mol Genet Genom 284:75–94. doi:10.1007/s00438-010-0553-4
Mehta GD, Rizvi SMA, Ghosh SK (2012) Cohesin: a guardian of genome integrity. Biochim Biophys Acta 1823:1324–1342. doi:10.1016/j.bbamcr.2012.05.027
Mehta K, Gunasekharan V, Satsuka A, Laimins LA (2015) Human papilloma viruses activate and recruit SMC1 cohesin proteins for the differentiation-dependent life cycle through association with CTCF insulators. PLoS Pathog 11:1–25. doi:10.1371/journal.ppat.1004763
Miller RK, D’Silva S, Moore JK, Goodson HV (2006) The CLIP-170 orthologue Bik1p and positioning the mitotic spindle in yeast. Curr Top Dev Biol 76:49–87. doi:10.1016/S0070-2153(06)76002-1
Mir M, Wang Z, Shen Z et al (2011) Optical measurement of cycle-dependent cell growth. Proc Natl Acad Sci 108:13124–13129. doi:10.1073/pnas.1100506108
Murray JAH, Cesareni G (1986) Functional analysis of the yeast plasmid partition locus STB. EMBO J 5:3391–3399
Murray AW, Szostak JW (1983) Pedigree analysis of plasmid segregation in yeast. Cell 34:961–970
Murray JAH, Scarpal M, Rossi N, Cesareni G (1987) Antagonistic controls regulate copy number of the yeast 2µ plasmid. EMBO J 6:4205–4212
Nasmyth K, Haering CH (2009) Cohesin: its roles and mechanisms. Annu Rev Genet 43:525–558. doi:10.1146/annurev-genet-102108-134233
Novák B, Tyson JJ (2008) Design principles of biochemical oscillators. Nat Rev Mol Cell Biol 9:981–991. doi:10.1038/nrm2530
Pan G, Luetke K, Sadowski PD (1993) Mechanism of cleavage and ligation by FLP recombinase: classification of mutations in FLP protein by in vitro complementation analysis. Mol Cell Biol 13:3167–3175. doi:10.1128/MCB.13.6.3167.Updated
Papacs LA, Sun Y, Anderson EL et al (2004) REP3-mediated silencing in Saccharomyces cerevisiae. Genetics 166:79–87
Petes TD, Williamson DH (1975) Replicating circular DNA molecules in yeast. Cell 4:249–253. doi:10.1016/0092-8674(75)90172-5
Petes TD, Williamson DH (1994) A novel structural form of the 2 micron plasmid of the yeast Saccharomyces cerevisiae. Yeast 10:1341–1345. doi:10.1002/yea.320101011
Pinder JB, McQuaid ME, Dobson MJ (2013) Deficient sumoylation of yeast 2-micron plasmid proteins Rep1 and Rep2 associated with their loss from the plasmid-partitioning locus and impaired plasmid inheritance. PLoS One 8:e60384. doi:10.1371/journal.pone.0060384
Prajapati HK, Rizvi SMA, Rathore I, Ghosh SK (2017) Microtubule-associated proteins, Bik1 and Bim1, are required for faithful partitioning of the endogenous 2 micron plasmids in budding yeast. Mol Microbiol 103:1046–1064. doi:10.1111/mmi.13608
Prasad PV, Young LJ, Jayaram M (1987) Mutations in the 2-microns circle site-specific recombinase that abolish recombination without affecting substrate recognition. Proc Natl Acad Sci USA 84:2189–2193
Reynolds AE, Murray AW, Szostak JW (1987) Roles of the 2 µm gene products in stable maintenance of the 2 µm plasmid of Saccharomyces cerevisiae. Mol Cell Biol 7:3566–3573
Rizvi SMA, Prajapati HK, Nag P, Ghosh SK (2017) The 2-μm plasmid encoded protein Raf1 regulates both stability and copy number of the plasmid by blocking the formation of the Rep1–Rep2 repressor complex. Nucleic Acids Res. doi:10.1093/nar/gkx316
Sadowski PD (1995) The Flp recombinase of th 2-μm plasmid of Saccharomyces cerevisiae. Prog Nucleic Acid Res Mol Biol 51:53–91
Sau S, Conrad MN, Lee C-Y et al (2014) A selfish DNA element engages a meiosis-specific motor and telomeres for germ-line propagation. J Cell Biol 205:643–661. doi:10.1083/jcb.201312002
Scott-Drew S, Murray JAH (1998) Localisation and interaction of the protein components of the yeast 2µ circle plasmid partitioning system suggest a mechanism for plasmid inheritance. J Cell Sci 111:1779–1789
Scott-Drew S, Wong CMVL, Murray JAH (2002) DNA plasmid transmission in yeast is associated with specific sub-nuclear localisation during cell division. Cell Biol Int 26:393–405
Senecoff JF, Bruckner RC, Cox MM (1985) The FLP recombinase of the yeast 2-µm plasmid: characterization of its recombination site. Proc Natl Acad Sci USA 82:7270–7274
Sengupta A, Blomqvist K, Pickett AJ et al (2001) Functional domains of yeast plasmid-encoded Rep proteins. J Bacteriol 183:2306–2314. doi:10.1128/JB.183.7.2306
Shcheprova Z, Baldi S, Frei SB et al (2008) A mechanism for asymmetric segregation of age during yeast budding. Nature 454:728–734. doi:10.1038/nature07212
Som T, Armstrong KA, Broach JF, Volkert C (1988) Autoregulation of 2 µm circle gene expression provides a model for maintenance of stable plasmid copy levels. Cell 52:27–37
Stevens BJ, Moustacchi E (1971) γ Satellite DNA and small twisted circular molecules in yeast. Exp Cell Res 64:259–266
Tsuchiya E, Hosotani T, Miyakawa T (1998) A mutation in NPS1/STH1, an essential gene encoding a component of a novel chromatin-remodeling complex RSC, alters the chromatin structure of Saccharomyces cerevisiae centromeres. Nucleic Acids Res 26:3286–3292. doi:10.1093/nar/26.13.3286
Velmurugan S, Ahn YT, Yang XM et al (1998) The 2 µm plasmid stability system: analyses of the interactions among plasmid- and host-encoded components. Mol Cell Biol 18:7466–7477
Velmurugan S, Yang XM, Chan CS et al (2000) Partitioning of the 2-µm circle plasmid of Saccharomyces cerevisiae: functional coordination with chromosome segregation and plasmid-encoded rep protein distribution. J Cell Biol 149:553–566
Volkert FC, Broach JR (1986) Site-specific recombination promotes plasmid amplification in yeast. Cell 46:541–550
Wei XA, Pelcher LE, Rank GH (1991) DNA sequence divergence and functional conservation at the STB locus of yeast 2 µm circle variants. J Bacteriol 173:1181–1186
Westermann S, Drubin DG, Barnes G (2007) Structures and functions of yeast kinetochore complexes. Ann Rev Biochem 76:563–591. doi:10.1146/annurev.biochem.76.052705.160607
Wong MCVL, Scott-Drew SRS, Hayes MJ et al (2002) RSC2, encoding a component of the RSC nucleosome remodeling complex, is essential for 2 microm plasmid maintenance in Saccharomyces cerevisiae. Mol Cell Biol 22:4218–4229. doi:10.1128/MCB.22.12.4218-4229.2002
Wu LCC, Fisher PA, Broach JR (1987) A yeast plasmid partitioning protein is a karyoskeletal component. J Biol Chem 262:883–891
Xiao W, Pelcher LE, Rank GH (1991) Sequence diversity of yeast 2 µm RAF gene and its co-evolution with STB and REP1. Gene 101:75–80
Xiong L, Chen XL, Silver HR et al (2009) Deficient SUMO attachment to Flp recombinase leads to homologous recombination-dependent hyperamplification of the yeast 2 µm circle plasmid. Mol Biol Cell 20:1241–1251. doi:10.1091/mbc.E08
Yamagishi Y, Sakuno T, Goto Y, Watanabe Y (2014) Kinetochore composition and its function: lessons from yeasts. FEMS Microbiol Rev 38:185–200
Yang X-M, Mehta S, Uzri D et al (2004) Mutations in a partitioning protein and altered chromatin structure at the partitioning locus prevent cohesin recruitment by the Saccharomyces cerevisiae plasmid and cause plasmid missegregation. Mol Cell Biol 24:5290–5303. doi:10.1128/MCB.24.12.5290-5303.2004
You J, Croyle JL, Nishimura A, Ozato K, Howley PM (2004) Interaction of the bovine papillomavirus E2 protein with Brd4 tethers the viral DNA to host mitotic chromosomes. Cell 117:349–360
Zakian VA, Brewer BJ, Fangman WL (1979) Replication of each copy of the yeast 2 micron DNA plasmid occurs during the S phase. Cell 17:923–934
Zhao X, Wu CY, Blobel G (2004) Mlp-dependent anchorage and stabilization of a desumoylating enzyme is required to prevent clonal lethality. J Cell Biol 167:605–611. doi:10.1083/jcb.200405168
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work is funded by DST (SB/SO/BB-0125/2013) Govt. of India, Council of Scientific and Industrial Research [38/1267/10/EMR-II] Govt. of India. HKP and SMAR are supported by MHRD and UGC (Ref. no: 20-06/2010(i)EU-IV) fellowship, respectively.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Rizvi, S.M.A., Prajapati, H.K. & Ghosh, S.K. The 2 micron plasmid: a selfish genetic element with an optimized survival strategy within Saccharomyces cerevisiae . Curr Genet 64, 25–42 (2018). https://doi.org/10.1007/s00294-017-0719-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-017-0719-2