Abstract
Transposons are an important source of genetic variation. The phytopathogen Moniliophthora perniciosa shows high level of variability but little is known about the role of class I elements in shaping its genome. In this work, we aimed the characterization of a new gypsy/Ty3 retrotransposon species, named MpSaci, in the M. perniciosa genome. These elements are largely variable in size, ranging from 4 to 15 kb, and harbor direct long terminal repeats (LTRs) with varying degrees of similarity. Approximately, all of the copies are non-autonomous as shifts in the reading frame and stop codons were detected. Only two elements (MpSaci6 and MpSaci9) code for GAG and POL proteins that possess functional domains. Conserved domains that are typically not found in retrotransposons were detected and could potentially impact the expression of neighbor genes. Solo LTRs and several LARDs (large retrotransposon derivative) were detected. Unusual elements containing small sequences with or without interruptions that are similar to gag or different pol domains and presenting LTRs with different levels of similarities were identified. Methylation was observed in MpSaci reverse transcriptase sequences. Distribution analysis indicates that MpSaci elements are present in high copy number in the genomes of C-, S- and L-biotypes of M. perniciosa. In addition, C-biotype isolates originating from the state of Bahia have fragments in common with isolates from the Amazon region and two hybridization profiles related to two chromosomal groups. RT-PCR analysis reveals that the gag gene is constitutively expressed and that the expression is increased at least three-fold with nutrient depravation even though no new insertion were observed. These findings point out that MpSaci collaborated and, even though is primarily represented by non-autonomous elements, still might contribute to the generation of genetic variability in the most important cacao pathogen in Brazil.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In various organisms, transposable elements are an important source of genetic variability and significantly impact genome organization and evolution. Transposable elements can be directly involved in spontaneous mutations via base insertions and deletions, gene expression alterations, chromosomal rearrangements, and other changes (Daboussi and Capy 2003; Shnyreva 2003; Pereira et al. 2006; Wicker et al. 2007). The investigation of the diversity, distribution and expression of these elements can provide important information about molecular biology of an organism.
Transposons are divided into two major classes according to their mechanism of transposition: class I, whose elements are transposed by reverse transcription of an RNA intermediate, and class II, whose transposition acts by a cut-and-paste mechanism not requiring an RNA intermediate. The class I elements that are flanked by long terminal repeats (LTRs) and possess a pol region, encoding a protease, reverse transcriptase, RNase H and integrase in this particular order, in addition to the gag gene are termed gypsy/Ty3-like. In contrast, class I elements with an integrase located between the protease and reverse transcriptase are referred to as copia/Ty1-like (Daboussi and Capy 2003; Wicker et al. 2007). Copia/Ty1 elements are classified into Pseudoviridae family while gypsy/Ty3 elements belong to the Metaviridae family, which is separated into three genus, Metavirus, Errantivirus and Semotivirus (King et al. 2012). Retrotransposons from the gypsy/Ty3 group (Metaviridae) are the major transposable elements found in phytopathogenic fungi (Daboussi and Capy 2003; Pereira et al. 2006). Given the fact that transposons encode for proteins necessary for transposition, transposons can be classified as follows: autonomous elements encode for all the domains; defective elements contain small mutations but still possess intact coding regions; and non-autonomous elements lack the complete domain(s) necessary for transposition (Wicker et al. 2007). Non-autonomous elements are capable of propagating using the proteins coded by other elements (Tanskanen et al. 2007). In addition, if these elements display sufficient sequence similarity to other copies, these elements can be used by the cell machinery for various types of rearrangements through recombination. Changes in transposon sequences can be performed by defense mechanisms like RIP (repeated-induced point mutation) where C:G to A:T transversions are generated in repetitive DNA frequently leading to epigenetic silencing through DNA methylation (Galagan and Selker 2004). In fact, those transversions were detected in reverse transcriptase sequences from M. perniciosa evidencing the presence of a putative RIP-like process (Pereira et al. 2007).
Moniliophthora perniciosa (Agaricales, Tricholomataceae) is the most important pathogen of cocoa in Brazil. After that phytopathogen arrived in the state of Bahia, the main cocoa-producing region in the country, Brazil shifted from one of the largest world producers to a major importer (Evans et al. 2013). Other plants in addition to cocoa are infected by M. perniciosa, and pathological data is used to divide this species into biotypes as the C-biotype, which infects species from the Theobroma and Herrania (Sterculiaceae) genera; the S-biotype, which infects species from the Solanaceae family; and the L-biotype, which colonizes a variety of plants (Meinhardt et al. 2008). While the L-biotype shows an out-crossing reproductive strategy (bifactorial heterothallism) and, consequently, higher levels of genetic diversity, C- and S-biotypes exhibit primary homothallism being the basidiospores produced when the disease cycle is completed (Griffith and Hedger 1994). Curiously, the C-biotype possesses A- and B-mating-type-like genes orthologous to the ones of heterothallic basidiomycetes (Kües and Navarro-González 2010).
Genetic diversity in M. perniciosa has been evaluated using several methods, and various levels of variability have been reported. In general, S- and L-biotype isolates display considerable diversity. C-biotype isolates from the Amazon region are more heterogeneous than isolates from Bahia, where two genotype groups have been identified. For example, restriction fragment length polymorphism targeting the mitochondrial DNA and ribosomal DNA ITS regions display no differences among isolates from various biotypes (Arruda et al. 2003a). In contrast, AFLP (Amplified Fragment Length Polymorphism) displays substantial variation among C-biotypes isolates from Bolivia, Brazil, Ecuador and Trinidad, but it shows similarity among isolates from the same geographic region (Ploetz et al. 2005). However, differences among C- and S-biotype isolates were detected using RAPD (Andebrhan et al. 1999), rDNA IGS region sequencing (Arruda et al. 2003a) and ERIC-PCR (Arruda et al. 2003b). IRAP and REMAP analyses revealed intraspecific variability among C-biotype isolates from different regions (Santana et al. 2012). Using pulsed field gel electrophoresis, distinct chromosomal patterns among S- and L- biotypes isolates were identified compared with C-biotype isolates (Rincones et al. 2003, 2006). In the Bahia state, C-biotype isolates typically possess eight chromosomes with different electrophoretic patterns; some isolates have eight bands, and others isolate display six bands with two overlapping bands. Microsatellites have also been isolated from M. perniciosa (Gramacho et al. 2007; Silva et al. 2008), indicating various polymorphisms for each locus. Some of these microsatellites and the ITS and IGS regions of the rDNA were evaluated in Colombian M. perniciosa isolates, and clear genetic variability was demonstrated (Osorio-Solano et al. 2012). Diversity studies have also been performed using somatic compatibility and SDS- PAGE (Ferreira et al. 2012).
In a survey of M. perniciosa genome sequences, gypsy/Ty3 elements were reported as the most abundant transposon type identified in this species (Mondego et al. 2008). A gypsy/Ty3-like reverse transcriptase, which is distributed across the M. perniciosa genome (Rincones et al. 2003), was characterized by our group (Pereira et al. 2007). Other types of elements, such as a transposase from the TC1-mariner superfamily (Ignacchiti et al. 2011) and an interesting element from the PIF/harbinger superfamily called Boto (Pereira et al. 2013), demonstrate that class II transposons are also distributed throughout the genome of this pathogen. Transposons were identified in the differentially expressed transcripts between biotrophic and saprophytic M. perniciosa mycelia (Rincones et al. 2008); however, Boto element expression has been detected in mycelium grown normally in culture medium (Pereira et al. 2013). It has been postulated that stresses can activate the transposition of these elements, thereby increasing the genetic variability necessary for adaptation to various environmental conditions (McClintock 1984). To date, it has been demonstrated that elements from various phytopathogenic fungi species are activated by some type of biotic or abiotic stress, such as heat shock, nutrient stress, toxic substances, or others (Anaya and Roncero 1996; Mes et al. 2000; Eto et al. 2001; Ikeda et al. 2001; Rep et al. 2005; Bouvet et al. 2008; Amyotte et al. 2012; Chadha and Sharma 2014).
Given that transposable elements are important in the evolution of new variants of plant pathogenic fungi, it is important to analyze the presence and activity of transposable elements to understand the biology of a species and the mechanisms that may be involved in the generation of genetic variability. In this article, we characterize several gypsy/Ty3 transposable elements that belong to a new species, named MpSaci, in the M. perniciosa genome. The majority of the elements are non-autonomous as only two of the characterized copies produce GAG and POL proteins with intact domains. We also report the distribution of this species in various biotype isolates, phylogenetic analyses and element expression under nutrient deprivation. MpSaci elements are likely involved in the generation of genetic variability in M. perniciosa given the number of elements, their putative activity, the rearrangements observed among the unusual elements, the presence of solo LTR and conserved domains not typically found in transposable elements.
Materials and methods
MpSaci sequences
The Moniliophthora perniciosa genome was obtained from the Witches’ Broom Genome Project database (www.lge.ibi.unicamp.br/vassoura). LTR transposons were detected using LTR-Finder software (Xu and Wang 2007). The resulting sequences were aligned with the reverse transcriptase sequence characterized by Pereira et al. (2007), leading to the identification of the MpSaci1 element. This element was used to identify additional MpSaci elements in the sequences generated by the LTR-finder software, and similarities greater than 1e−10 were not considered.
M. perniciosa isolates
Moniliophthora perniciosa isolates examined in this study are listed in Table 1. Stock cultures were maintained at room temperature after growth on PDA (Potato Dextrose Agar) at 27 °C for 2 weeks.
Primers
To amplify a 394 bp DNA fragment corresponding to a region of the MpSaci reverse transcriptase coding region, we used the primers 5′ TGCCAAAGAAACAGGGACTT 3′ and 5′ GAATTTGCGAGACCGAAAAA 3′ described by Pereira et al. (2007). The primers SNF1-R (5′ GCTGTCCGTATGTCCAAGGT 3′) and SNF1-L (5′ GATGTACCACACGCCTGAGA 3′) that amplify part of the M. perniciosa snf1 gene were used as RT-PCR experimental controls (Medina 2006). The primers ITS1 and ITS4 were used to amplify the rDNA ITS region as described by White et al. (1990). The primers GAG1 (5′ AACTTGACGAGGCGAATCAT 3′) and GAG2 (5′ TTTGACCGACTTCAGGAAGC 3′) were constructed based on the MpSaci14 sequence using the Primer3 program (http://www-genome.wi.mit.edu/cgibin/primer/primer3.cgi) (Rozen and Skaletsky 2000). These primers were used to amplify a 400 bp DNA fragment corresponding to a portion of the gag coding region. For the IRAP and REMAP analyses (Kalendar et al. 1999), primers CPLTR1F (5′ CTCGCTTCTTCAGCTTGACC 3′), CPLTR1R (5′ GTGCAAGCGACACACAAACT 3′), CPLTR2F (5′ GCCGTACTTAGCGTACTTCTTT 3′), CPLTR2R (5′ GCAGTGTTTCTACAGCGAAG 3′), MS1 (5′ GGCGGCGGCGGCGGCGGCGGCT 3′), MS2 (5′ ATGATGATGATGATGATGATGG 3′), and MS4 (5′ CACACACACACACACACACAG 3′) developed by Santana et al. (2012) were used in different combinations (Fig. 7).
Sequence analysis
The MpSaci sequences were compared with sequences deposited in GenBank using the BLAST program (Basic Local Alignment Search Tool) (Altschul et al. 1997). Subsequent analyses were performed using ClustalW (Thompson et al. 1994) and CDD to search for conserved domains (Marchler-Bauer et al. 2011) with a maximum E value of 1e−5. For the alignment analysis, the following GAG and POL sequences (with the respective accession numbers) were used: grh (M77661, M77661), MAGGY (AAA33419, AAA33420), marY1 (BAA78624, BAD10925, BAA78625), REAL (BAA89271, BAA89272), and skippy (AAA88790, S60179). For the phylogenetic analysis of the MpSaci reverse transcriptase protein, representative sequences of the Metaviridae (genus Metavirus, Errantivirus and Semotivirus) and Pseudoviridae families were obtained using the following access numbers: Bel (U23420), CfT-1 (Z11866), copia (D10880), grh (M77661), gypsy (X03734), MAGGY (AAA33420), marY1 (BAA78625), ninja (D83207), Pao (L09635), REAL (BAA89272), skippy (S60179), suzu (AF537216), Tf1-107 (M38526), Tom (Z24451), Tv1 (AF056940), Ty1 (Z48149), and yoyo (U60529). Phylogenetic analyses were performed with the ClustalW program (Thompson et al. 1994) via the neighbor-joining method (Saitou and Nei 1987) with bootstrap values using 1,000 replicates.
DNA extraction and PCR
Total DNA was extracted as described by Pereira et al. (2014). PCR was performed in a thermocycler (PTC-100—MJ Research) programmed for 40 cycles of 1 min at 94 °C; 1 min at 55 °C for the reverse transcriptase, 55 °C for gag or 52 °C for the ITS region; 1 min at 72 °C and a final extension at 72 °C for 10 min. The reactions were prepared in a final volume of 25 μl containing 1× thermophilic DNA poly Buffer (Promega), 2.5 mM MgCl2 (Promega), 100 μM of each dNTP, 0.5 μM of each primer, 20 ng of DNA and one unit of Taq DNA polymerase (Promega). Negative controls (without DNA) were used for each set of experiments to assess non-specific amplification. The DNA fragments were analyzed by electrophoresis using a 1.5 % agarose gel.
Southern hybridization analysis
Total DNA (3.0 μg) was cleaved with BamHI, which does not cleave the reverse transcriptase sequence used as a probe, HpaII or MspI, which recognize the same sequence (5′ CCGG 3′) but the cut is dependent of the cytosine methylation status (Bender 2004). The cleaved DNA was electrophoresed overnight in 0.7 % agarose gel and transferred to Duralon membrane-UVTM (Stratagene) according to standard protocols (Sambrook et al. 1989). Hybridization was performed at 65 °C using the 394 bp DNA fragment corresponding to part of the reverse transcriptase as the probe as well as the “Gene ImagesTM Random Primer Labeling Module” and “CDP-StarTM Detection Module” (Amersham) according to the manufacturer’s instructions.
RNA extraction, RT-PCR, IRAP and REMAP
To obtain the M. perniciosa mycelial mass for the nutritional stress experiments, 7-mm mycelial disks were incubated on plates containing PDA medium at 27 °C for 10 days. After growth, 30 small mycelium fragments were transferred to 125-ml Erlenmeyer flasks containing 50 ml of liquid minimal medium (6.8 g KH2PO4, 8.6 g K2HPO4, 6.0 g KNO3, 0.52 g KCl, 0.01 g FeSO4, 0.01 g ZnSO4, 10 g glucose, and 1,000 ml distilled water) and incubated for 7 days at 27 °C/180 rpm. After 7 days, the mycelia were filtered, washed with distilled water, transferred to fresh minimal medium with and without a carbon or nitrogen source, and incubated for 10 h at 27 °C/180 rpm. The mycelia was used for total DNA extraction (as described above) and total RNA extraction as described by Watkinson et al. (2008). For the RT reactions, total RNA was treated with RNase-Free RQI DNase (Promega) and quantified at 260 nm. The first cDNA strand was generate using 5 μg of RNA, 1× RT buffer (Promega), 0.5 mM dNTPs, 500 ng oligo (dT)15 (Promega), 20 U of ribonuclease inhibitor RNasin (Promega) and 10 U Avian Myeloblastosis Virus Reverse Transcriptase (Promega). The volume of each reaction was adjusted to 20 μl, and the reactions were incubated at 25 °C for 5 min and then at 42 °C for 60 min. PCR reactions were performed to detect the cycles corresponding to the logarithmic phase of DNA amplification. For these analyses, we used 1 μl cDNA and SNF1-L/R primers that amplify a 230 or 300 bp fragment of the M. perniciosa snf1 gene depending on whether the template is cDNA or genomic DNA (Medina 2006). The amplification program was 1 min at 94 °C, 1 min at 57 °C and 1 min at 72 °C. Aliquots were removed at 20, 22, 24, 26 and 40 cycles. Based on the results from these analyses, all PCR reactions were performed using 25 cycles, and the entire content of each amplification reaction (25 μl) was separated on a 1.5 % agarose gel. The intensity of the amplified fragments was analyzed using ImageJ software (Abramoff et al. 2004) obtained from http://rsb.info.nih.gov/ij/. The expression of the gag gene was calculated in relation to snf1 gene expression. The total DNA was analyzed by IRAP (inter-retrotransposon amplified polymorphism) and REMAP (retrotransposon-microsatellite amplified polymorphism) techniques with PCR amplification and thermocycling conditions as described by Santana et al. (2012). Amplifications were performed twice.
Results
Harvesting the MpSaci elements
A reverse transcriptase (RT) sequence previously characterized by our group (Pereira et al. 2007) was used to identify a complete LTR retrotransposon from the sequences in the M. perniciosa genome harvested by the LTR-Finder software. The first element identified, named MpSaci1, is 8,499 bp with 529 bp direct long terminal repeats (LTRs) displaying 99 % similarity. MpSaci1 possesses two open read frames (ORFs) coding for the GAG and POL proteins; however, the first ORF is interrupted by two premature stop codons (Supplementary Fig. 1). The MpSaci1 sequence was used to search for other elements from this species in the sequences harvested by the LTR-Finder software. We were able to obtain several sequences demonstrating various levels of similarity with the MpSaci1 sequence. Fourteen of these sequences displayed an organization typically found in gypsy-Ty3 elements; however, the majority of the copies did not code for intact GAG and POL proteins due to premature stop codons, frameshift mutations, or the lack of start codons and coding capacity (Fig. 1). The additional sequences demonstrated several rearrangements leading to truncated elements with low similarity to GAG or to some POL protein domains and LTRs in the same or opposite direction. These regions either flanked or did not flank sequences with similarities to GAG and/or POL (Fig. 2). Given that only two elements coded for all the domains necessary for transposition, non-autonomous elements primarily represent the MpSaci species in the M. perniciosa genome.
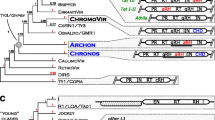
Schematic representation of the fourteen MpSaci elements in M. perniciosa genome that display organization similar to the gypsy/Ty3 group. For convenience, all elements are presented in the same orientation. The GAG protein contains a zinc finger domain (Zn) displayed in black. The POL protein contains protease (PR), reverse transcriptase (RT), RNase H (RH) and integrase (IN) domains. The asterisks correspond to premature stop codons, and the arrows indicate frameshift mutations. N indicates sequences with a stretch of undefined bases. ? indicates protein sizes that could not be detected. The bars are proportional to the element size of the elements and putative proteins (not considering the stop codons when possible). The sizes of the GAG and POL proteins in aa (amino acids) are indicated considering stop codons. The LTR sizes and similarity between the LTR 5′ and 3′ are indicate below each element. The numbers inside the circles represent the presence of the following conserved domains: 1 chromatin organization modifier, 2 GAL4-like Zn2Cys6 binuclear cluster DNA-binding domain, 3 Tymovirus 45/70Kd protein, 4 C-terminal domain of homeodomain 1 for mating typing, 5 Atrophin-1, 6 AF-4 proto-oncoprotein, 7 Herpes-TAF50 super family, and 8 DUF3295 super family
Schematic representation of the unusual MpSaci sequences displaying various levels of homology with the MpSaci1 element. For convenience, all elements are presented in the same orientation. N indicates sequences with a stretch of undefined bases. The bars are proportional to the sequence size. The LTRs inside the boxes (gray horizontal arrowheads) are similar to the LTRs from the MpSaci1 to Mpsaci14 elements. The LTRs indicated by white horizontal arrowheads are not similar to the MpSaci elements. The LTR sizes are indicated below each element. The dark boxes represent the domains of a Ty1/copia-like retrotransposon inside the MpSaci15 element. The numbers inside the circles represent the presence of the following conserved domains: 1 chromatin organization modifier, 2 GAL4-like Zn2Cys6 binuclear cluster DNA-binding domain, 9 DnaQ-like-exo super family, 10 SDH-beta super family, 11 hAT family dimerization domain, 12 Glyco-hydrolase-16 super family, 13 hAT transposase, 14 DDE superfamily endonuclease, and 15 large tegument protein UL36
Characterization of the MpSaci species
The direct LTRs identified in the 14 elements with gypsy/Ty3 organization display similarities ranging from 94 to 100 % and sizes ranging from 223 to 581 bp. Most of the LTRs are larger than the ones found in the majority of gypsy/Ty3 LTR retroelements (200–450 bp); however, the Rhizot element has LTRs greater than 1 kb (Abe et al. 2009). Various characteristics have been reported for LTR of fungal retrotransposons, including inverted repeats at the ends, tRNA primer binding sites, and transcriptional initiation and polyadenylation signals. Inverted terminal repeats are typically found in the majority of retroviral and retrotransposon LTRs (Bingham and Zachar 1989). Similar to the LTRs found in skippy, CfT-1, Boty, Cgret, REAL and ANiTa1 elements, gypsy/Ty3 group elements typically end in 5′-TG…CA-3′ (McHale et al. 1992; Dobinson et al. 1993; Anaya and Roncero 1995; Diolez et al. 1995; Kaneko et al. 2000; Zhu and Oudemans 2000; Braumann et al. 2007). The LTRs of the majority of MpSaci elements also end in inverted repeats containing 5′-TG…CA-3′ (Fig. 3a). However, additional characteristics of the LTRs from other filamentous fungi were not found in the MpSaci elements, such as the primer binding site that is typically required for transcription initiation and is located immediately upstream of the 5′ LTR containing a conserved TGG in Drosophila (Bingham and Zachar 1989). For the REAL and marY1 elements, the primer binding site was not found, but purine-rich sequences that correlate with the duplication process were identified (Kaneko et al. 2000; Murata and Yamada 2000). Three CT rich regions, one sequence similar to CCATT and two putative TATA boxes potentially linked to the transcription start site were identified in the marY1 element. The majority of eukaryotic LTR retroelements are also flanked by small repeats related to the target site duplication; however, these repeats could not be precisely identified for the MpSaci elements. Interestingly, the alignment of the LTRs from the 14 copies of MpSaci indicates minimal sequence similarity among some of the copies. In addition, these copies can be divided into different groups (Fig. 3b). The gag, pol and LTRs appear to be the target of several rearrangements because the MpSaci15 to MpSaci23 elements display an unusual pattern and solo LTRs were found (Fig. 2). Several sequences have LTRs but do not contain gag or pol coding regions, and these sequences can be classified as LARDs (large retrotransposon derivative) based on their sizes (MpSaci24 to MpSaci30) (Havecker et al. 2004). Two of the unusual elements (MpSaci15 and MpSaci26) display rearrangements leading to the presence of other transposable elements (Ty1/copia and hAT) between the LTRs of MpSaci. Transposable elements containing additional domains from other transposons inserted into the structure of a typical Ty3 element were also observed by Muszewska et al. (2011). MpSaci15 was considered part of the MpSaci species since most part of the sequence (around 80 %) that is flanked by LTRs is correlated to a gypsy/Ty3 retroelement.
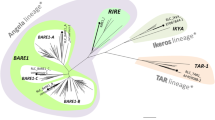
Analyses of the LTRs and reverse transcriptases from the fourteen MpSaci elements displaying organization similar to the gypsy/Ty3 group. a Sequence analysis of the first and last five bases in the 5′ and 3′ LTRs. Black circles before the name of the element represent elements possessing similar inverted repeats. b Relationship among the LTRs from the fourteen MpSaci elements. c Phylogenetic tree presenting the relationship among the reverse transcriptase domains from MpSaci1, MpSaci6, MpSaci9, MpSaci12 and MpSaci15 as well as other fungal retrotransposons and representatives of the Metaviridae (genus Metavirus, Errantivirus and Semotivirus) and Pseudoviridae families. The arrowhead indicates the second reverse transcriptase sequence found in MpSaci15 that belongs to the Ty1/copia group of retroelements. The numbers in each branch indicate the bootstrap based on 1,000 replicates. The trees were constructed using the neighbor-joining method (Saitou and Nei 1987)
Intact ORFs coding for GAG (MpSaci4, MpSaci6, MpSaci9, MpSaci11, MpSaci13 and MpSaci14) and POL (MpSaci1, MpSaci6, MpSaci9, MpSaci12 and MpSaci15) proteins were detected. MpSaci6 and MpSaci9 are the only elements that code for both intact proteins. High sequence similarity among the GAG and POL proteins encoded by MpSaci elements and transposons from other filamentous fungi were detected. Conserved domains were identified in all of the elements. However, the region surrounding the GAG zinc finger in MpSaci6 displayed reduced similarity. In addition, the glutamic acid from the integrase DDE motif in the POL protein from MpSaci1 was changed to a lysine, likely resulting in a non-functional protein (Supplementary Fig. 2). Thus, MpSaci6 and MpSaci9 are the only elements that encode all of the domains typically required for transposition.
The sizes of the functional GAG proteins range from 346 to 375 aa with predicted molecular masses from 39.6 to 40.6 kDa. The mass is less than the masses reported for other retrotransposons with the exception of the marY1 element, which has a 42 kDa GAG protein (Murata and Yamada 2000). However, a putative ribosomal frameshift site was identified in marY1, which could result in a 63.6 kDa GAG protein. This size is similar to the mass reported for other retrotransposons (60–80 kDa). Putative ribosomal frameshift sites were not detected in the MpSaci elements. A conserved zinc finger domain (Cys-X2-Cys-X4-His-X4-Cys), which is also present in other retrotransposon and retroviral GAGs, was detected in the GAGs from MpSaci4, MpSaci6, MpSaci9, MpSaci11, MpSaci13 and MpSaci14 (Supplementary Fig. 2A). The functional POL protein is formed by four different domains (protease, reverse transcriptase, RNase H and integrase) and ranges in size from 1,061 to 1,220 aa with a predicted molecular mass of 122.2–141.5 kDa. The POL protein displays high similarity to the marY1 element from Tricholoma matsutake (Murata and Yamada 2000), REAL (Kaneko et al. 2000), MAGGY from Magnaporthe grisea (Farman et al. 1996), and skippy from Fusarium oxysporum (Anaya and Roncero 1995). The protease domain possesses the conserved motif DSG (Asp, Ser, Gly), which is the proposed active site (Katoh et al. 1987). The reverse transcriptase displays the seven domains described by Xiong and Eickbush (1990) and the motif YXDD (Tyr, Met, Asp, Asp), which is the proposed enzyme active site (Varmus and Brown 1989). The integrase possesses the motif DD(35)E (Asp, Asp, Glu), which is highly conserved among retrotransposon integrases (Supplementary Fig. 2E). Phylogenetic analyses using reverse transcriptase domains from various elements reveal that MpSaci is related to the gypsy-Ty3 group. In addition, the other reverse transcriptase sequence found in MpSaci15 is clearly related to a Ty1/copia element (Fig. 3c).
Another characteristic of the MpSaci elements is the long distance between the end of the pol coding region and the 3′ LTR; this region is greater than 2 kb in MpSaci1, MpSaci2, MpSaci5, MpSaci7, MpSaci8, MpSaci11 and MpSaci13. In different elements, this region possesses seven conserved domains that are not related to the domains typically found in retrotransposon sequences (Figs. 1, 2). The most common domains are the chromatin organization modifier (chromo) domain; a DNA-binding domain that is similar to the domain found in transcription regulators, such as GAL4; and the C-terminal domain of homeodomain 1, which is involved in fungi mating types. Some of the conserved domains detected in the MpSaci elements were previously reported to negatively or positively impact transcription (Marmorstein and Harrison 1994; Koonin et al. 1995).
Distribution of MpSaci
The MpSaci distribution was evaluated by PCR and Southern hybridization. Portions of the gag and reverse transcriptase coding region were amplified in all isolates (Fig. 4), indicating that the primers annealing sites were conserved among all isolates. Hybridization with the RT sequence indicates that MpSaci copies are present in the genomes of C-, S- and L-biotypes (Fig. 5). Although the resolution of the Southern analysis is not clear for most of the genotypes, the lowest MpSaci copy number, which was estimated to be seven, was detected for the FA609 isolate (L-biotype). Since the bioinformatics analysis resulted in 18 sequences with homology to the probe, we estimated the MpSaci copy number from seven to 18. However, considering the unusual elements, that number could increase to 30 sequences as detected for the CP02 genome (Figs. 1, 2). The C-biotype isolates originated from the Amazon region and the Bahia state isolates have different hybridization profiles; these two distinct profiles correlate with the two chromosomal groups, CG1 and CG2, described by Rincones et al. (2006). Although DNA fragments greater than 3 kb are not separated and hence more difficult to analyze, some DNA fragments from the Amazon isolates correlate with Bahia state isolates (Fig. 5). One common 1,630 bp fragment is present in isolate 3 (originated from Rondônia) and the CG1 isolates (from Bahia). This fragment is absent in isolates 2 and 4 (from the Pará state) as well as the CG2 isolates (Bahia). This finding creates a link among the different chromosomal groups from the Bahia isolates and C-biotypes from different locations in the Amazon region. L- and S-biotype isolates also possess an increased MpSaci element copy number; however, none of the copies could be correlated with a specific genotype or location (Fig. 5).
PCR amplification of various M. perniciosa isolates with primers specific for the MpSaci elements. a Amplification of a 394 bp DNA fragment corresponding to part of the reverse transcriptase coding region; b amplification of a 400 bp DNA fragment corresponding to part of the gag coding region; and c amplification of the rDNA-ITS region as a control for DNA quality. M indicates DNA size marker (ϕX174 Hae III), and n indicates the negative control (reaction without total DNA). The isolate identification corresponds to the data presented in Table 1
Hybridization of the 394 bp DNA fragment containing part of the MpSaci reverse transcriptase coding region with total DNA from the M. perniciosa isolates. BamHI cutted DNA of a C-biotypes and b L- and S-biotypes isolates. c DNA of CP02 isolate cleaved with HpaII or MspI. The isolate identification is presented in Table 1. “CG1” and “CG2” refer to the two chromosomal groups of the C-biotype from Bahia. The position of some fragments is presented in the left; the size and position of the fragment discussed in the text are displayed
MpSaci reverse transcriptase methylation status
Hybridization of the MpSaci reverse transcriptase revealed a different profile depending if the total DNA was cleaved with HpaII or MspI (Fig. 5). The cleavage by HpaII was inhibited generating larger bands while MspI was able to cut the transcriptase sequences more often. The different hybridization profile is evidence that MpSaci reverse transcriptase sequence is methylated.
Expression analysis
RT-PCR was performed to evaluate the effect of nutritional stress (lack of nutrients) on gag gene expression (Fig. 6). After determining that the logarithmic phase of DNA amplification occurs between cycles 24 and 26 (Fig. 6c), the PCR reaction was ended at the 25th cycle. An intron was detected in the amplification of the control gene (snf1) (Fig. 6e). Amplification of the 400 bp DNA fragment that corresponds to a region of the MpSaci gag gene revealed that this element is constitutively expressed in M. perniciosa grown in minimal medium. In addition, nutritional stress activates element expression (Fig. 6f). A minimum of three-fold increase in DNA intensity was detected in stress conditions (Fig. 6i). No introns were detected in the 400 bp DNA fragment from the MpSaci gag gene. Using GenBank (http://www.ncbi.nlm.nih.gov/nucleotide/), we also detected several MpSaci gag and pol sequences generated from expression studies, indicating that MpSaci expression was also detected in other experiments (Supplementary Table 1).
RT-PCR analysis of gag expression from MpSaci. a Representation of the experiment. b Quality of the RNA extracted. c, d Evaluation of the amplification curve using the snf1 gene to detect the logarithm phase of DNA amplification. Amplifications of the snf1 control (e) and part of the MpSaci gag region (f) were performed until the 25th cycle. g, h Densitometric analysis of the amplified products. i Relative gag gene expression from the MpSaci elements in relation to the snf1 gene. ϕX indicates the DNA size marker (ϕX174 Hae III); “MM” indicates minimum medium; w/glu indicates minimum medium without glucose; w/nit indicates minimum medium without nitrogen source; DNA indicates the total DNA from the CP02 isolate; and n indicates the negative control (amplification without adding cDNA or DNA). The DNA fragment sizes amplified are presented in c, e and f
IRAP and REMAP
To search for new insertions, DNA extracted from nutritionally stressed mycelia was used to amplify regions between LTRs (IRAP) or LTR insertions near to microsatellite markers (REMAP). Different primers combinations were used being the amplification profile highly similar among the stressed and non-stressed mycelia (Fig. 7). Differences in the amplification profile were observed in the stressed mycelia when the microsatellite primer MS2 was used in different combinations. New insertions were not detected.
IRAP and REMAP analyses (2 % agarose gel) in DNA extracted from the nutritionally stressed mycelia. Some fragments of the molecular size marker are presented in the left. MM indicates minimum medium; w/glu indicates minimum medium without glucose; w/nit indicates minimum medium without nitrogen source. White rectangle indicates differences in the amplification profile
Discussion
This report is the first one to characterize all members of a species, belonging to the class I of retrotransposons, in the most important pathogen of cocoa in Brazil. Since transposable elements are associated with increased adaptability and the phenotypic variation demonstrated by phytopathogenic fungi (Daboussi and Capy 2003; Shnyreva 2003; Pereira et al. 2006), the characterization of the retrotransposon MpSaci is an important step forward in understanding the genomic organization and adaptability of M. perniciosa. Almost entirely represented by non-autonomous elements, the MpSaci species contains elements whose gene organization are similar to gypsy/Ty3 elements but also unusual elements and solo LTRs. This species is distributed across different biotypes and it is activated under nutritional stress although the lack of some structural characteristics might prevent an efficient transposition.
High copy numbers of the MpSaci elements exist in C-, S- and L-biotypes of M. perniciosa that originate from various geographic regions (Fig. 5). Based on a transposition mechanism wherein a new copy is inserted in a new target site, high copy numbers of retrotransposons are found in the genomes of phytopathogenic fungi. In general, transposable elements constitute less than 20 % of fungal genomes (Daboussi and Capy 2003; Wicker et al. 2007) although they can represent around 85 % of Blumeria graminis f.sp. tritici genome (Parlange et al. 2011). Class I elements are more abundant than class II elements in M. perniciosa genome (Mondego et al. 2008); however, the percentage of the genome that is comprised of these elements remains unknown. Phylogenetic analysis of the reverse transcriptase protein, the most conserved protein among retrotransposons (Havecker et al. 2004), and the organization of the pol region indicate that MpSaci is another species of the Metaviridae family (gypsy/Ty3) belonging to the genus Metavirus (Fig. 3c). The gypsy/Ty3 group is the most abundant retrotransposon in the M. perniciosa genome (Mondego et al. 2008), and this group is typically the most abundant retrotransposon in other filamentous fungi (Muszewska et al. 2011). According to Daboussi (1996), gypsy/Ty3-like retrotransposons are ubiquitous because one common ancestor contained a copy of these elements and the elements were transferred vertically along with the other components of the genome during speciation events. The vertical transmission is also based on phylogenetic analyses of the reverse transcriptase. The majority of the LTR-retrotransposons are clustered, thereby creating a monophyletic group of fungal transposons; this finding is consistent with the vertical heritage (Daboussi 1996).
The high copy number of transposable elements in fungi can potentially lead to chromosome-length polymorphism (Zolan 1995). Not only the transposition can cause chromosomal breaks but also the homologous recombination between copies spread throughout the genome can promote chromosomal rearrangements (Mieczkowski et al. 2006). With regard to phytopathogenic fungi, a relationship between karyotype instability and transposable elements has been reported in F. oxysporum and M. grisea (Shull and Hamer 1996a, b; Nitta et al. 1997; Davière et al. 2001; Thon et al. 2006; Starnes et al. 2012). In M. perniciosa, two different types of chromosomal patterns were reported for C-biotypes originating from the Bahia state (Rincones et al. 2003, 2006), and these chromosomal patterns display two unique hybridization patterns when analyzed using a MpSaci reverse transcriptase sequence (Fig. 5). Similarly, the Boto element from the PIF/Harbinger superfamily (Pereira et al. 2013) and a portion of an element belonging to the TC1-Mariner superfamily (Ignacchiti et al. 2011) are also distributed in M. perniciosa genome based on the chromosomal groups. MpSaci is the third species of transposable elements analyzed by our group in the M. perniciosa genome. Given that all of these elements are distributed according to the Bahia state C-biotype chromosomal groups, we hypothesize that the chromosomal polymorphism causes the different transposons profiles or, alternatively, the transposable elements are potentially linked to the origin of the two C-biotype chromosomal groups. Clearly, more concrete evidence should be provided to support these hypotheses. In addition to the link between MpSaci distribution and the chromosomal groups, two distinct MpSaci hybridization profiles were detected in the C-biotype isolates from the Bahia state. This finding indicates the presence of two genotypic groups, and each group displays similarities with different genotypes from the Amazon region. The presence of two distinct C-biotype groups in the Bahia state has been reported previously by Andebrhan et al. (1999) through RAPD analysis; this study concluded that witches’ broom was introduced in Bahia via two independent points. In addition, a relationship between the Amazon and Bahia isolates was also reported by other authors (Andebrhan et al. 1999; Arruda et al. 2003a, b; Ploetz et al. 2005; Ignacchiti et al. 2011).
Interestingly, transposons can contribute to the evolution of phytopathogenic fungi by promoting phenotype alterations. Various alterations originate through inactivation via insertional mutagenesis; this mechanism is directly linked to transposition promoted by autonomous elements. For example, the insertion of Pot3 in AVR-Pita promotes virulence of M. grisea in a cultivar containing the Pi-Ta resistance gene (Kang et al. 2001). This element was also identified in the AVR-Pita gene of other isolates (Zhou et al. 2007; Singh et al. 2014). Moreover, transposon insertions promoted gain of virulence in C. fulvum and increased drug resistance in B. cinerea (Luderer et al. 2002; Kretschmer et al. 2009). In addition, changes generated by recombination are also reported, thereby providing evidence that non-autonomous transposable elements can also contribute to the evolution of plant pathogens. For example, retrotransposons near the avirulence gene AvrLm1 in L. maculans could have increased the frequency of recombination events in that region, creating deletions that are linked to virulence in some isolates (Fudal et al. 2007). Transposons in another avirulence gene, AVR-Pita, potentially contribute to the increased proportion of translocation in the genomes of various M. oryzae isolates (Chuma et al. 2011). In addition, the super-expression of CYP51, which confers resistance to fungicide, is positively correlated with a truncated LINE-like element in the CYP51 coding region in Blumeriella jaapii (Ma et al. 2006). In Neurospora spp., retrotransposons play an important role in the transition between homothallic and heterothallic systems (Gioti et al. 2012). In P. tritici-repentis, alterations induced by transposons are likely to be involved in the emergence of pathogenicity (Manning et al. 2013). Thus, although the MpSaci species is primarily represented by non-autonomous elements, these elements still can contribute to genetic variability in M. perniciosa. It is also important to consider that some of the unusual domains in the MpSaci sequences could contribute to alterations in the expression of neighboring genes because these domains could potentially impact transcription (Marmorstein and Harrison 1994; Koonin et al. 1995).
The following characteristics demonstrate that the majority of MpSaci elements sequenced in the wild type isolate are likely non-autonomous: the absence of structures typically found in LTRs, the presence of premature stop codons and frameshift mutations, the complete absence of sequences resembling the coding region and the reduced similarity/lack of conserved domains (Figs. 1, 2). The reduced similarity among LTRs from various elements of the same species potentially suggests that the MpSaci is an old resident of the M. perniciosa genome. This finding might indicate that the host genome suppressed the majority of the deleterious effects of the transposable elements. Non-autonomous retrotransposons are typically found in various fungal species and can represent the majority of element copies in the genome (Muszewska et al. 2011). For instance, no autonomous elements were identified in the N. crassa genome (Galagan et al. 2003; Schulman 2012). These elements can be inactivated through genetic or epigenetic mechanisms by the host genome (Kazazian 2004). Two evidences indicate that a silencing mechanism related to RIP (repeat-induced point mutation) could be acting on M. perniciosa genome: methylated reverse transcriptase sequences as observed in Southern analysis (Fig. 5) and numerous G:C to A:T transitions at the MpSaci reverse transcriptase previously reported by Pereira et al. (2007). RIP is the mechanism of action implicated as a primary reason for the origin of degenerate transposable elements reported in other phytopathogenic fungi (Nakayashiki et al. 1999; Attard et al. 2005). Although RIP is reported in numerous fungal species (Clutterbuck 2011), this defense mechanism has not been shown to exist in M. perniciosa. In addition to RIP, other mechanisms act on MpSaci elements, leading to truncated copies, LTRs with reduced similarity to elements (Fig. 3b), solo LTR and several LARDs (Fig. 2). Solo LTRs have been detected in the Phanerochaete chrysosporium and B. graminis f.sp. tritici genomes (Martinez et al. 2004; Parlange et al. 2011) as well as in the Ty1, ANiTa1 and yeti retrotransposons of Saccharomyces paradoxus, Aspergillus niger and Podospora anserina. LTR–LTR recombination, which is based on homologous recombination between LTRs, is thought to be involved in the generation of solo LTRs (Kim et al. 1998; Hamann et al. 2000; Moore et al. 2004; Braumann et al. 2008). Primers based on the MpSaci LTRs were used for inter-retrotransposon amplified polymorphism (IRAP) to generate polymorphic loci in the different isolates, and the solo LTR and LARDs sequences may have also contributed to the polymorphism (Santana et al. 2012). It is also important to note that we analyzed the genome of one M. perniciosa isolate, and different isolates can potentially retain autonomous copies of MpSaci.
Gene expression analysis of transposable elements (Okuda et al. 1998; Kaneko et al. 2000; Kito et al. 2003; Rep et al. 2005; Bouvet et al. 2008; Amyotte et al. 2012; Pereira et al. 2013) is used to demonstrate transposon activity as well as gene inactivation (Daboussi et al. 1992; Langin et al. 1995; Maurer et al. 1997; Gómez-Gómez et al. 1999) and the appearance of new copies in the genome (Anaya and Roncero 1996; Mes et al. 2000; Pereira et al. 2013; Chadha and Sharma 2014). Stress-induced transposition is described in other fungi. For example, nutritional stress activates the transposition of the hAT-like Drifter element in F. oxysporum and various elements in Verticillium dahliae (Rep et al. 2005; Amyotte et al. 2012). Nutritional stress results in at least 3-fold increase in gag gene expression from MpSaci (Fig. 6). Given that we detected several gag and pol sequences from cDNA libraries (Supplementary Table 1), these mRNAs could produce active GAG and POL proteins. Although only two MpSaci elements produce intact GAG and POL proteins, in trans activation of the transposition cannot be ruled out. However, the LTRs, the primer-binding site and a polypyrine tract are minimally required for transposon replication (Havecker et al. 2004). Given that some of these characteristics were not detected in the MpSaci elements, in trans mobilization appears to be unlikely in the analyzed M. perniciosa genome. In fact, new fragments were not detected by IRAP and REMAP techniques in the mycelia obtained after nutritional stress (Fig. 7).
In conclusion, our results highlight that MpSaci species is highly represented by non-autonomous elements. Even the elements that code for intact proteins lack structures necessary for efficient transposition. This could explain why no new insertions were detected although nutritional stress can activate the expression of gag and reverse transcriptase genes. Nevertheless, these elements could be involved in the generation of polymorphisms through homologous recombination between two copies and also could impact the transcription of neighboring genes since conserved domains were detected inside all copies. The copy number, expression and distribution of the MpSaci elements throughout the genomes of various isolates added valuable information about the molecular biology of M. perniciosa. In addition, further studies can address the questions raised here about the possible link between transposable elements and the appearance of two chromosomal groups in C-biotype isolates and the presence of defense mechanisms that promote the degeneration of most of the copies in a transposon species.
References
Abe A, Asano K, Sone T (2009) Identification and characterization of Rhizot, a novel LTR retrotransposon of Rhizopus oryzae and R. delemar. Biosci Biotechnol Biochem 73:1860–1862
Abramoff MD, Magelhaes PJ, Ram SJ (2004) Image processing with ImageJ. Biophotonics Int 11:36–42
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Amyotte SG, Tan X, Pennerman K, Jimenez-Gasco MM, Klosterman SJ, Ma LJ, Dobinson KF, Veronese P (2012) Transposable elements in phytopathogenic Verticillium spp.: insights into genome evolution and inter- and intra-specific diversification. BMC Genomics 13:314
Anaya N, Roncero MIG (1995) Skippy, a retrotransposon from the fungal plant pathogen Fusarium oxysporum. Mol Gen Genet 249:637–647
Anaya N, Roncero MIG (1996) Stress-induced rearrangement of Fusarium retrotransposon sequences. Mol Gen Genet 253:89–94
Andebrhan T, Figueira A, Yamada MM, Cascardo J, Furtek DB (1999) Molecular fingerprinting suggests two primary outbreaks of witches’ broom disease (Crinipellis perniciosa) of Theobroma cacao in Bahia, Brazil. Eur J Plant Pathol 105:167–175
Arruda MCC, Ferreira MASV, Miller RNG, Resende MLV, Felipe MSS (2003a) Nuclear and mitochondrial rDNA variability in Crinipellis perniciosa from different geographic origins and hosts. Mycol Res 107:25–37
Arruda MCC, Miller RNG, Ferreira MASV, Felipe MSS (2003b) Comparison of Crinipellis perniciosa isolates from Brazil by ERIC repetitive element sequence-based PCR genomic fingerprint. Plant Pathol 52:236–244
Attard A, Gout L, Ross S, Parlange F, Cattolico L, Balesdent MH, Rouxel T (2005) Truncated and RIP-degenerated copies of the LTR retrotransposon Pholy are clustered in a pericentromeric region of the Leptosphaeria maculans genome. Fungal Genet Biol 42:30–41
Bender J (2004) DNA methylation and epigenetics. Annu Rev Plant Biol 55:41–68
Bingham PM, Zachar Z (1989) Retrotransposons and the FB transposons from Drosophila melanogaster. In: Berg DE, Howe MM (eds) Mobile DNA. American Society for Microbiology, Washington, pp 485–502
Bouvet GF, Jacobi V, Plourde KV, Bernier L (2008) Stress-induced mobility of OPHIO1 and OPHIO2, DNA transposons of the Dutch elm disease fungi. Fungal Genet Biol 45:565–578
Braumann I, van den Berg M, Kempken F (2007) Transposons in biotechnologically relevant strains of Aspergillus niger and Penicillium chrysogenum. Fungal Genet Biol 44:1399–1414
Braumann I, van den Berg MA, Kempken F (2008) Strain-specific retrotransposon-mediated recombination in commercially used Aspergillus niger strain. Mol Genet Genomics 280:319–325
Chadha S, Sharma M (2014) Transposable elements as stress adaptive capacitors induce genomic instability in fungal pathogen Magnaporthe oryzae. PLoS One 9(4):e94415. doi:10.1371/journal.pone.0094415
Chuma I, Isobe C, Hotta Y, Ibaragi K, Futamata N, Kusaba M, Yoshida K, Terauchi R, Fujita Y, Nakayashiki H, Valent B, Tosa Y (2011) Multiple translocation of the AVR-Pita effector gene among chromosomes of the rice blast fungus Magnaporthe oryzae and related species. PLoS Pathog 7:e1002147
Clutterbuck AJ (2011) Genomic evidence of repeat-induced point mutation (RIP) in filamentous ascomycetes. Fungal Genet Biol 48:306–326
Daboussi MJ (1996) Fungal transposable elements: generators of diversity and genetic tools. J Genet 75:325–339
Daboussi MJ, Capy P (2003) Transposable elements in filamentous fungi. Annu Rev Microbiol 57:275–299
Daboussi MJ, Langin T, Brygoo Y (1992) Fot1, a new family of fungal transposable elements. Mol Gen Genet 232:12–16
Davière JM, Langin T, Daboussi MJ (2001) Potential role of transposable elements in the rapid reorganization of the Fusarium oxysporum genome. Fungal Genet Biol 34:177–192
Diolez A, Marches F, Fortini D, Brygoo Y (1995) Boty, a long-terminal-repeat retroelement in the phytopatogenic fungus Botrytis cinerea. Appl Environ Microbiol 61:103–108
Dobinson KF, Harris RE, Hamer JE (1993) Grasshoper, a long terminal repeat (LTR) retroelement in the phytopathogenic fungus Magnaporthe grisea. Mol Plant Microbe Interact 6:114–126
Eto Y, Ikeda K, Chuma I, Kataoka T, Kuroda S, Kituchi N, Don LD, Kusaba M, Nakayashiki H, Tosa Y, Mayama S (2001) Comparative analyses of the distribution of various transposable elements in Pyricularia and their activity during and after the sexual cycle. Mol Gen Genet 264:565–577
Evans HC, Bezerra JL, Barreto RW (2013) Of mushrooms and chocolate trees: aetiology and phylogeny of witches’ broom and frosty pod diseases of cacao. Plant Pathol 62:728–740
Farman ML, Tosa Y, Nitta N, Leong S (1996) MAGGY, a retrotransposon in the genome of the rice blast fungus Magnaporthe grisea. Mol Gen Genet 251:665–674
Ferreira LFR, Duarte KMR, Gomes LH, Carvalho RS, Leal Junior GA, Aguiar MM, Armas RD, Tavares FCA (2012) Genetic diversity of polysporic isolates of Moniliophthora perniciosa (Tricholomataceae). Genet Mol Res 11:2559–2568
Fudal I, Ross S, Gout L, Blaise F, Kuhn ML, Eckert MR, Cattolico L, Bernard-Samain S, Balesdent MH, Rouxel T (2007) Heterochromatin-like regions as ecological niches for avirulence genes in the Leptosphaeria maculans genome: map-based cloning of AvrLm6. Mol Plant Microbe Interact 20:459–470
Galagan JE, Selker EU (2004) RIP: the evolutionary cost of genome defense. Trends Genet 20:417–423
Galagan JE, Calvo SE, Borkovich KA et al (2003) The genome sequence of the filamentous fungus Neurospora crassa. Nature 422:859–868
Gioti A, Mushegian AA, Strandberg R, Stajich JE (2012) Unidirectional evolutionary transitions in fungal mating systems and the role of transposable elements. Mol Biol Evol 29:3215–3226
Gómez-Gómez E, Anaya N, Roncero MIG, Hera C (1999) Folyt1, a new member of the hAT family, is active in the genome of the plant pathogen Fusarium oxysporum. Fungal Genet Biol 27:67–76
Gramacho KP, Risterucci AM, Lanaud C, Lima LS, Lopes UV (2007) Characterization of microsatellites in the fungal plant pathogen Crinipellis perniciosa. Mol Ecol Notes 7:153–155
Griffith GW, Hedger JN (1994) The breeding biology of biotypes of the witches’ broom pathogen of cocoa, Crinipellis perniciosa. Heredity 72:278–289
Hamann A, Feller F, Osiewacz HD (2000) Yeti, a degenerate gypsy-like LTR retrotransposon in the filamentous ascomycete Podospora anserina. Curr Genet 38:132–140
Havecker ER, Gao X, Voytas DF (2004) The diversity of LTR retrotransposons. Genome Biol 5:225
Ignacchiti MDC, Santana MF, Araújo EF, Queiroz MV (2011) The distribution of a transposase sequence in Moniliophthora perniciosa confirms the occurrence of two genotypes in Bahia, Brazil. Trop Plant Pathol 36:276–286
Ikeda K, Nakayashiki H, Takagi M, Tosa Y, Mayama S (2001) Heat shock, copper sulfate and oxidative stress active the retrotransposon MAGGY resident in the plant pathogenic fungus Magnaporthe grisea. Mol Genet Genomics 266:318–325
Kalendar R, Grob T, Regina M, Suoniemi A, Schulman A (1999) IRAP and REMAP: two new retrotransposon-based DNA fingerprinting techniques. Theor Appl Genet 98:704–711
Kaneko I, Tanaka A, Tsuge T (2000) REAL, an LTR retrotransposon from the plant pathogenic fungus Alternaria alternata. Mol Gen Genet 263:625–634
Kang S, Lebrun MH, Farrall L, Valent B (2001) Gain of virulence caused by insertion of a Pot3 transposon in a Magnaporthe grisea avirulence gene. Mol Plant Microbe Interact 14:671–674
Katoh I, Yasunaga T, Ikawa Y, Yoshinaka Y (1987) Inhibition of retroviral protease activity by an aspartyl proteinase inhibitor. Nature 329:654–656
Kazazian HH Jr (2004) Mobile elements: drivers of Genome evolution. Science 303:1626–1632
Kim JM, Vanguri S, Boeke JD, Gabriel A, Voytas DF (1998) Transposable elements and genome organization: a comprehensive survey of retrotransposons revealed by the complete Saccharomyces cerevisiae genome sequence. Genome Res 8:464–478
King AM, Adams MJ, Lefkowitz EJ, Carstens EB (2012) Virus taxonomy: classification and nomenclature of viruses. Ninth Report of the International Committee on Taxonomy of Viruses, 9th edn. Elsevier Academic Press, Amsterdam
Kito H, Takahashi Y, Sato J, Fukiya S, Sone T, Tomita F (2003) Occan, a novel transposon in the Fot1 family, is ubiquitously found in several Magnaporthe grisea isolates. Curr Genet 42:322–331
Koonin EV, Zhou S, Lucchesi JC (1995) The chromo superfamily: new members, duplication of the chromo domain and possible role in delivering transcription regulators to chromatin. Nucleic Acids Res 23:4229–4233
Kretschmer M, Leroch M, Mosbach A, Walker AS, Fillinger S, Mernke D, Schoonbeek HJ, Pradier JM, Leroux P, Waard MAD, Hahn M (2009) Fungicide-driven evolution and molecular basis of multidrug resistance in field populations of the grey mould fungus Botrytis cinerea. PLoS Pathog 5:e1000696
Kües U, Navarro-González M (2010) Mating-type orthologous genes in the primarily homothallic Moniliophthora perniciosa, the causal agent of Witches’ Broom Disease in cacao. J Basic Microbiol 50:442–451
Langin T, Capy P, Daboussi MJ (1995) The transposable element impala, a fungal member of the Tc1-mariner superfamily. Mol Gen Genet 246:19–28
Luderer R, Takken FLW, de Wit PJGM, Joosten MHAJ (2002) Cladosporium fulvum overcomes Cf-2-mediated resistance by producing truncated AVR2 elicitor proteins. Mol Microbiol 45:875–884
Ma Z, Proffer TJ, Jacobs JL, Sundin GW (2006) Overexpression of the 14α-demethylase target gene (CYP51) mediates fungicide resistance in Blumeriella jaapii. Appl Environ Microbiol 72:2581–2585
Manning VA, Pandelova I, Dhillon B et al (2013) Comparative genomics of a plant-pathogenic fungus, Pyrenophora tritici-repentis, reveals transduplication and the impact of repeat elements on pathogenicity and population divergence. G3 3:41–63
Marchler-Bauer A, Lu S, Anderson JB et al (2011) CDD: a Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Res 39:D225–D229
Marmorstein R, Harrison SC (1994) Crystal structure of a PPR1-DNA complex: DNA recognition by proteins containing a Zn2Cys6 binuclear cluster. Genes Dev 8:2504–2512
Martinez D, Larrondo LF, Putnam N et al (2004) Genome sequence of the lignocellulose degrading fungus Phanerochaete chrysosporium strain RP78. Nat Biotechnol 22:695–700
Maurer P, Réjasse A, Capy P, Langin T, Riba G (1997) Isolation of the transposable element hupfer from the entomopathogenic fungus Beauveria bassiana by insertion mutagenesis of the nitrate reductase structural gene. Mol Gen Genet 256:195–202
McClintock B (1984) The significance of responses oh the genome to challenge. Science 226:792–801
McHale MT, Roberts IN, Noble SM, Beaumont C, Whitehead MP, Seth D, Oliver RP (1992) CfT-1: an LTR-retrotransposon in Cladosporium fulvum, a fungal pathogen of tomato. Mol Gen Genet 233:337–347
Medina PXL (2006) Isolamento e caracterização dos genes que codificam malato sintase e o repressor Nrg1 em Crinipellis perniciosa, agente causal da vassoura-de-bruxa no cacaueiro (Theobroma cacao). Ph.D. dissertation, Universidade Federal de Viçosa, Brazil
Meinhardt LW, Rincones J, Bailey BA, Aime MC, Griffith GW, Zhang D, Pereira GAG (2008) Moniliophthora perniciosa, the causal agent of witches’ broom disease of cacao: what’s new from this old foe? Mol Plant Pathol 9:577–588
Mes JJ, Haring MA, Cornelissen BJC (2000) Foxy: an active family of short interspersed nuclear elements from Fusarium oxysporum. Mol Gen Genet 263:271–280
Mieczkowski PA, Lemoine FJ, Petes TD (2006) Recombination between retrotransposons as a source of chromosome rearrangements in the yeast Saccharomyces cerevisiae. DNA Repair 5:1010–1020
Mondego JMC, Carazzolle MF, Costa GGL et al (2008) A genome survey of Moniliophthora perniciosa gives new insights into Witches’ Broom Disease of cacao. BMC Genom 9:548
Moore SP, Liti G, Stefanisko KM, Nyswaner KM, Chang C, Louis EJ, Garfinkel DJ (2004) Analysis of a Ty1-less variant of Saccharomyces paradoxus: the gain and loss of Ty1 elements. Yeast 21:649–660
Murata H, Yamada A (2000) marY1, a member of the gypsy group of long terminal repeat retroelements from the ectomycorrhizal basidiomycete Tricholoma matsutake. Appl Environ Microbiol 66:3642–3645
Muszewska A, Hoffman-Sommer M, Grynberg M (2011) LTR retrotransposons in fungi. PloS One 6:e29425
Nakayashiki H, Nishimoto N, Ikeda K, Tosa Y, Mayana S (1999) Degenerate MAGGY elements in a subgroup of Pyricularia grisea: a possible example of successful capture of a genetic invader by a fungal genome. Mol Gen Genet 261:958–966
Nitta N, Farman ML, Leong SA (1997) Genome organization of Magnaporthe grisea: integration of genetic maps, clustering of transposable elements and identification of genome duplications and rearrangments. Theor Appl Genet 95:20–32
Okuda M, Ikeda K, Namiki F, Nishi K, Tsuge T (1998) Tfo1: an Ac-like transposon from the plant pathogenic fungus Fusarium oxysporum. Mol Gen Genet 258:599–607
Osorio-Solano C, Orozco-Castaño CA, López-Gartner GA, Rivera-Páez FA (2012) Variabilidad genética de Moniliophthora perniciosa (Stahel) Aime y Phillips-Mora, comb. nov. (Agaricales—Marasmiaceae) en variedades de cacao (Theobroma cacao L.). Acta Agron 61:93–101
Parlange F, Oberhaensli S, Breen J, Platzer M, Taudien S, Šimková H, Wicker T, Doležel J, Keller B (2011) A major invasion of transposable elements accounts for the large size of the Blumeria graminis f. sp. tritici genome. Funct Integr Genomics 11:671–677
Pereira JF, Araújo EF, Brommonschenkel SH, Queiroz MV (2006) Elementos transponíveis em fungos fitopatogênicos. Rev Anual Patol Plant 14:303–362
Pereira JF, Ignacchiti MDC, Araújo EF, Brommonschenkel SH, Cascardo JCM, Pereira GAG, Queiroz MV (2007) PCR amplification and sequence analyses of reverse transcriptase-like genes in Crinipellis perniciosa isolates. Fitopatol Bras 32:373–380
Pereira JF, Almeida APMM, Cota J, Pamphile JA, Silva GF, Araújo EF, Gramacho KP, Brommonschenkel SH, Pereira GAG, Queiroz MV (2013) Boto, a class II transposon in Moniliophthora perniciosa, is the first representative of the PIF/Harbinger superfamily in a phytopathogenic fungus. Microbiology 159:112–125
Pereira JF, Consoli L, Bombonatto EAS, Bonato ALV, Maciel JLN (2014) Development of genomic SSR markers and molecular characterization of Magnaporthe oryzae isolates from wheat in Brazil. Biochem Genet 52:52–70
Ploetz RC, Schnell RJ, Ying Z, Zheng Q, Olano CT, Montamayor JC, Johnson ES (2005) Analysis of molecular diversity in Crinipellis perniciosa with AFLP markers. Eur J Plant Pathol 111:317–326
Rep M, Van der Does HC, Cornelissen BJC (2005) Drifter, a novel, low copy hAT-like transposon in Fusarium oxysporum is activated during starvation. Fungal Genet Biol 42:546–553
Rincones J, Meinhardt LW, Vidal BC, Pereira GAG (2003) Electrophoretic karyotype analysis of Crinipellis perniciosa, the causal agent of witches’ broom disease of Theobroma cacao. Mycol Res 107:452–458
Rincones J, Mazotti GD, Griffith GW, Pomela A, Figueira A, Leal GA, Queiroz MV, Pereira JF, Pereira GAG, Meinhardt LW (2006) Genetic variability and chromosome-length polymorphisms of the witches’ broom pathogen Crinipellis perniciosa from various plant hosts in South America. Mycol Res 110:821–832
Rincones J, Scarpari LM, Carazzolle MF et al (2008) Differential gene expression between the biotrophic-like and saprotrophic mycelia of the witches’ broom pathogen Moniliophthora perniciosa. Mol Plant Microbe Interact 7:891–908
Rozen S, Skaletsky HJ (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics Methods and Protocols: Methods in Molecular Biology. Humana Press, Totowa, NJ, pp 365–386. http://www-genome.wi.mit.edu/cgibin/primer/primer3.cgi. Accessed 24 April 2006
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Santana MF, Araújo EF, Souza JT, Mizubuti ESG, Queiroz MV (2012) Development of molecular markers based on retrotransposonsfor the analysis of genetic variability in Moniliophthora perniciosa. Eur J Plant Pathol 134:497–507
Schulman AH (2012) Hitching a ride: nonautonomous retrotransposons and parasitism as a lifestyle. In: Grandbastien MA, Casacuberta JM (eds) Plant Transposable Elements, Topics in Current Genetics. Springer, Berlin, Heidelberg, pp 71–88
Shnyreva AV (2003) Transposable elements are the factors involved in various rearragements and modifications of the fungal genomes. Russ J Genet 39:505–518
Shull V, Hamer JE (1996a) Genetic differentiation in the rice blast fungus revealed by the distribution of the Fosbury retrotransposon. Fungal Genet Biol 20:59–69
Shull V, Hamer JE (1996b) Rearrangements at a DNA-fingerprint locus in the rice blast fungus. Curr Genet 30:263–271
Silva JRQ, Figueira A, Pereira GAG, Albuquerque P (2008) Development of novel microsatellites from Moniliophthora perniciosa, causal agent of the witches’ broom disease of Theobroma cacao. Mol Ecol Res 8:783–785
Singh PK, Thakur S, Rathour R, Variar M, Prashanthi SK, Singh AK, Singh UD, Sharma V, Singh NK, Sharma TR (2014) Transposon-based high sequence diversity in Avr-Pita alleles increases the potential for pathogenicity of Magnaporthe oryzae populations. Funct Integr Genomics. doi:10.1007/s10142-014-0369-0
Starnes JH, Thornbury DW, Novikova OS, Rehmeyer CJ, Farman ML (2012) Telomere-targeted retrotransposons in the rice blast fungus Magnaporthe oryzae: agents of telomere instability. Genetics 191:389–406
Tanskanen JA, Sabot F, Vicient C, Schulman AH (2007) Life without GAG: the BARE-2 retrotransposon as a parasite’s parasite. Gene 390:166–174
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensibility of progressive multiple sequence alignment through sequence weighting, position specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Thon MR, Pan HQ, Diener S, Papalas J, Taro A, Mitchell TK, Dean RA (2006) The role of transposable element clusters in genome evolution and loss of synteny in the rice blast fungus Magnaporthe oryzae. Genome Biol 7:R16
Varmus H, Brown P (1989) Retroviruses. In: Berg DE, Howe MM (eds) Mobile DNA. American Society for Microbiology, Washington, pp 53–108
Watkinson JI, Hendricks L, Sioson AA, Heath LS, Bohnert HJ, Grene R (2008) Tuber development phenotypes in adapted and acclimated, drought-stressed Solanum tuberosum ssp. andigena have distinct expression profiles of genes associated with carbon metabolism. Plant Physiol Biochem 46:34–45
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribossomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Wicker T, Sabot F, Hua-Van A et al (2007) A unified classification system for eukaryotic transposable elements. Nat Rev Genet 8:973–982
Xiong Y, Eickbush TH (1990) Origin and evolution of retroelements based upon their reverse transcriptase sequences. EMBO J 9:3353–3362
Xu Z, Wang H (2007) LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res 35:265–268
Zhou E, Jia Y, Singh P, Correll JC, Lee FN (2007) Instability of the Magnaporthe oryzae avirulence gene AVR-Pita alters virulence. Fungal Genet Biol 44:1024–1034
Zhu P, Oudemans PV (2000) A long terminal repeat retrotransposon Cgret from the phytopathogenic fungus Colletotrichum goesporioides on cranberry. Curr Genet 38:241–247
Zolan ME (1995) Chromosome-length polymorphism in fungi. Microbiol Rev 59:686–698
Acknowledgments
We are grateful to CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico), CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior), SEAGRI (Secretaria de Agricultura, Irrigação e Reforma Agrária do Estado da Bahia), CARGILL/FUNCAMP/UNICAMP convenium, and FAPESP (Fundação de Amparo à Pesquisa do Estado de São Paulo) for financial support. We also would like to acknowledge the contributions of the Center for Computational Engineering and Sciences at Universidade Estadual de Campinas, SP, Brazil (FAPESP/CEPID project #2013/08293-7).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by G. Goldman.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pereira, J.F., Araújo, E.F., Brommonschenkel, S.H. et al. MpSaci is a widespread gypsy-Ty3 retrotransposon highly represented by non-autonomous copies in the Moniliophthora perniciosa genome. Curr Genet 61, 185–202 (2015). https://doi.org/10.1007/s00294-014-0469-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-014-0469-3